Abstract
Background
Post-translational modification (PTM) of proteins controls various cellular functions of transcriptional regulators and participates in diverse signal transduction pathways in cancer. The thyroid hormone (triiodothyronine, T3) plays a critical role in metabolic homeostasis via its direct interaction with the thyroid hormone receptor beta (TRβ). TRβ is involved in physiological processes, such as cell growth, differentiation, apoptosis, and maintenance of metabolic homeostasis through transcriptional regulation of target genes.
Objective
This study was performed to characterize the specific PTM of TRβ is an active control mechanism for the proteasomal degradation of TRβ in transcriptional signaling pathways in hepatocellular carcinoma cells.
Methods
Based on a previous study, we predicted that the lysine methyltransferase and methylation sites of TRβ by comparing the amino acid sequences of histone H3 and TRβ. Methyl-acceptor site of TRβ was confirmed by point mutation. TRβ protein stability was evaluated by ubiquitination assay with MG132. For glucose starvation, HepG2 cells were incubated in media without D-glucose. Proliferation-related proteins were detected by western blotting. MicroRNA level and autophagy marker were measured by real-time qPCR.
Results
The presence of enhancer of zeste homolog 2 (Ezh2), a methyltransferase of H3 lysine 27, as a methyltransferase of TRβ also revealed that direct lysine methylation and consequent stimulated protein degradation of TRβ underlies the negative correlation between Ezh2 and TRβ. Notably, glucose starvation significantly increased lysine methylation, and methylated TRβ showed further protein instability leading to an increase in the proliferation and growth of hepatocellular carcinoma cells.
Conclusions
TRβ functions as a tumor suppressor in various cancers; therefore, we evaluated the effect of TRβ degradation on oncogenesis during glucose starvation. These data clearly define a functional model and provide a link between metabolism and cancer by regulating methyl-dependent protein levels of tumor suppressors. Taken together, maintaining TRβ against methyl-dependent degradation is considered a possible therapeutic target for cancer progression.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Thyroid hormone receptor beta (TRβ) is a nuclear receptor subfamily of thyroid hormone T3 (Kowalik et al. 2010; Mangelsdorf et al. 1995). TRβ acts as a transcription factor in the genomic or non-genomic signaling pathways (Giammanco et al. 2020; Porlan et al. 2008). TRβ mediates biological mechanisms, such as cell differentiation, growth, apoptosis, and maintenance of homeostasis (Gao et al. 2017; Gu et al. 2015). TRβ is also known as a tumor suppressor that inhibits the migration and expression of oncogenes (Davidson et al. 2021b; Kim et al. 2014). Furthermore, TRβ is dysregulated or mutated in several types of cancer (Singh and Yen 2017). Post-translational modification (PTM) is an enzymatic modification of proteins that occurs after protein biosynthesis and includes processes such as acetylation, phosphorylation, and methylation (Prabakaran et al. 2012). Although the transcriptional regulation function of TRβ is well known, the PTM and protein degradation pathways of TRβ have not been characterized.
Enhancer of zeste homolog 2 (Ezh2) is a subunit of polycomb repressive complex 2 and functions as an epigenetic modifier that catalyzes histone H3 trimethylation at lysine 27 (H3K27me3) (Gan et al. 2018). Ezh2 has six domains: WD8, D1, SANT, D2, CXC, and SET; the SET domain is a catalytic domain of Ezh2 (Tan et al. 2014; Wu et al. 2013). Ezh2 has been proposed to have an oncogenic potential that increases cell migration and proliferation, and is frequently mutated or overexpressed in cancer (Donaldson-Collier et al. 2019). Recently, Ezh2 has been shown to be involved in the direct methylation of non-histone substrates, including the tumor suppressor retinoic acid-related orphan nuclear receptor α (RORα) in breast cancer (Lee et al. 2012). Therefore, inhibiting the oncogenic role exerted by Ezh2-mediated direct methylation is an important therapeutic strategy in cancer therapy.
Cancer cells require a lot of energy and nutrients for their abnormal rapid growth and migration, which could lead to deprivation of nutrients such as amino acids, glutamine, and glucose (Ahmadiankia et al. 2019; Nikolaou and Machesky 2020). The cellular self-digestion autophagy pathway induced by nutrient deprivation has diverse and occasionally opposing roles in cancer progression (He et al. 2018). Autophagy pathways are either antagonistic or agonistic to oncogenesis, depending on the cancer context (Yun and Lee 2018). Several studies have demonstrated that autophagy supplies sufficient nutrients that enable cancer cell growth, and the identification of proteins responsible for autophagy-dependent oncogenesis is important for exploring complex malignant processes (Galluzzi et al. 2014).
Here, we identified that TRβ is a non-histone lysine methylation substrate that is methylated by Ezh2, and that lysine methylation of TRβ is regulated by autophagy through glucose starvation. In a previous study, we found that Ezh2 induces lysine methylation in the histone-like RKS sequence in the orphan nuclear receptor RORα, and generates methylation-dependent degradation through the ubiquitin–proteasome pathway (Lee et al. 2012). Compared to histone H3 and RORα, we found that TRβ has a similar RKS sequence in the N-terminus and that Ezh2 increases the lysine methylation of TRβ. In addition, we confirmed that the RKS sequence of TRβ is a lysine methylation site of TRβ through a point mutation construct. Furthermore, polyubiquitination-mediated protein destabilization of TRβ was induced by Ezh2 in a lysine methylation-dependent manner. Notably, glucose starvation increases the lysine methylation of TRβ and facilitates methyl-dependent protein degradation in hepatocellular carcinoma. These findings suggest a previously unrecognized non-histone protein candidate of methyl-degron, the glucose starvation-mediated lysine methyl-degron in TRβ that could stimulate cancer progression.
Materials and methods
Cell lines and cell culture
293 T cells were cultured in Dulbecco’s modified Eagle’s Medium (DMEM, Welgene, LM 001–05) supplemented with 10% Fetal Bovine Serum (FBS, Gibco, 26140–079) and 1% antibiotics (Welgene, LS 203–01) at 37 °C in a humidified 5% CO2 incubator. HepG2 cells were cultured in Roswell Park Memorial Institute Medium 1640 (RPMI1640, Welgene, LM 011–01) supplemented with 10% FBS and 1% antibiotics at 37 °C in a humidified 5% CO2 incubator. Trypsin–EDTA (Welgene, LS 015–01) is used for cell dissociation. For glucose starvation, HepG2 cells were incubated in Dulbecco’s modified Eagle’s Medium without D-glucose (DMEM, Welgene, LM001-56) supplemented with 10% dialyzed FBS (Gibco, 26400–044) and 1% antibiotics at 37 °C in a humidified 5% CO2 incubator.
Cell transfection
293 T cells were transfected with 1 μg/μl polyethyleneimine (PEI, Sigma, 408727) each plasmid.
Western blotting
For western blotting, cells were dissociated from the culture dish with cold phosphate-buffered saline (PBS) and centrifuged at 4 °C 6000 rpm for 3 min. Remove the supernatants and cells were suspended in lysis buffer EBC200 (50 mM Tris–HCl pH 8.0, 200 mM NaCl, 0.5% NP-40, 100 ng/ml aprotinin, 50 ng/ml leupeptin) for 30 min on ice and centrifuged at 4 °C 13,000 rpm for 10 min. Supernatants were transferred to new tubes for subsequent protein assays. Protein samples were separated using SDS polyacrylamide gel electrophoresis (SDS-PAGE) and transferred to a PVDF membrane (Millipore). The transferred membrane was blocked in skim milk for 40 min and was incubated overnight with primary antibodies at 4 °C. Primary antibodies GFP (Santacruz, sc-9996), HA (BioLegend, 901514), Flag (Sigma, F3165), TRβ (Sigma), Ezh2 (BD Bioscience, 612667), pAKT (CST, 4060S), pan-AKT (CST, 9272S), P-p44/42 MAPK (pERK, CST, 4377S), p44/42 MAPK (pan-ERK, CST, 4696S), β-actin (Sigma, A5441) were purchased and used.
Immunoprecipitation (IP) assay
Cells were suspended in 300-700 μl lysis buffer EBC200 for 30 min and centrifuged at 4 °C 13,000 rpm for 10 min, supernatants were divided into Input (10% of supernatants), IgG control and IP (each 50% of extra supernatants). For immunoprecipitation, a primary antibody was added in the IP, and incubated overnight at 4 °C on the rotator. After overnight incubation, immunoprecipitation followed by incubation with 30 μl of Protein A (Sigma, P9424), Protein G (Sigma, P3296), CL4B (Sigma, CL4B200) Sepharose mixture (A:G: CL4B, 1:1:10) for 1 h at 4 °C on the rotator. Before the Sepharose incubation, the Sepharose mixture was washed four times with the lysis buffer. After incubation with lysates (IgG, IP), the Sepharose was washed six times with the lysis buffer. Sepharose was centrifuged at 4 °C 6000 rpm and boiled in an SDS sample buffer. These samples were separated using SDS-PAGE and western blotting. An anti-methylated lysine antibody (abcam, ab23366) was purchased and used for immunoprecipitation.
Ubiquitination assay
293 T cells were transfected with plasmids including HisMaxC-ubiquitin. After transfection, cells were treated with MG132 (AG scientific, M1157) for 4 h. For ubiquitination assay, cells were harvested in 1 ml cold PBS containing 5 mM N-ethylmaleimide (NEM, Sigma). 100 μl of suspended cells were transferred to a new tube, pelleted and boiled with 30 μl PBS and 30 μl SDS sample buffer (200 mM Tris–HCl pH6.8, 4% SDS, 20% Glycerol, 20 mM EDTA). 800 μl of suspended cells were transferred to 6 ml lysis buffer (6 M Guanidium Chloride, 0.1 M Na2HPO4/NaH2PO4, 0.01 M Tris–HCl pH8.0, 5 mM imidazole, 10 mM β-mercaptoethanol) in a 15 ml conical tube, and incubated with Ni2+-NTA beads (QIAGEN) for overnight at 4 °C. The beads were centrifuged at 2000 rpm for 5 min and sequentially washed with buffer A (6 M Guanidium Chloride, 0.1 M Na2HPO4/NaH2PO4, 0.01 M Tris–HCl pH8.0, 10 mM β-mercaptoethanol), buffer B (8 M Urea, 0.1 M Na2HPO4/NaH2PO4, 0.01 M Tris–HCl pH8.0, 10 mM β-mercaptoethanol), buffer C (8 M Urea, 0.1 M Na2HPO4/NaH2PO4, 0.01 M Tris–HCl pH6.3, 10 mM β-mercaptoethanol, 0.2% Triton X-100) and buffer D (8 M Urea, 0.1 M Na2HPO4/NaH2PO4, 0.01 M Tris–HCl pH6.3, 10 mM β-mercaptoethanol, 0.1% Triton X-100). Ni2+-NTA beads bound proteins were boiled with elution buffer (SDS sample buffer, 0.72 M β-mercaptoethanol, 200 mM imidazole). These samples were separated using SDS-PAGE and western blotting.
RNA isolation and real-time qPCR
RNAs were extracted using TRIzol Reagent (Thermofisher), and reverse transcription was performed using T100 thermal cycler (Biorad) with MystiCq microRNA cDNA synthesis mix (Sigma) and TOPscript cDNA Synthesis kit (Enzynomics). Real-time qPCR was performed using QuantStudio 1 (Applied Biosystem) with SYBR TOPreal qPCR 2X PreMix (Enzynomics). PCR conditions were followed the recommended protocols. The quantity of mRNA was calculated using ddCt method, and U6, ActB was used as control. All sample reactions were performed by triplicate. The following primers were used in this study: Ezh2 Forward 5′-CCC TGA CCT CTG TCT TAC TTG TGG A-3′ Reverse 5′-ACG TCA GAT GGT GCC AGC AAT A-3′, LC3II Forward 5′-GAG AAG CAG CTT CCT GTT CTG G-3′ Reverse 5′-GTG TCC GTT CAC CAA CAG GAA G-3′, ActB Forward 5′-TAG CCA TCC AGG CTG TGC TG-3′ Reverse 5′-CAG GAT CTT CAT GAG GTA GTC-3′, miR21 Forward 5′-ACA CTC CAG CTG GGT AGC TTA TCA GAC TGA-3′ Reverse 5′-TGG TGT CGT GGA GTC G-3′, U6 Forward 5′-CGC TTC GGC AGC ACA TAT ACT AAA ATT GGA AC-3′ Reverse 5′-GCT TCA CGA ATT TGC GTG TCA TCC TTG C-3′.
Results
Ezh2 mediates lysine methylation of TRβ
Ezh2 is a histone lysine methyltransferase enzyme that methylates H3K27 (Cai et al. 2021; Simon and Lange 2008). In our previous study, we confirmed that the lysine of the histone-like RKS sequence could be methylated by the histone modifier Ezh2. We found this sequence by comparing the amino acid sequences of methylated histone and the non-histone substrate RORα. We compared the amino acid sequences of H3K27, RORα, and TRβ; the RKS sequence was also found in the TRβ sequence (Fig. 1a). To investigate whether TRβ could be methylated or not, we performed an immunoprecipitation assay using an anti-methylated lysine antibody and confirmed that TRβ is a lysine methylated substrate (Fig. 1b). Given that Ezh2 was predicted as a putative lysine methyltransferase of TRβ, we overexpressed Ezh2 to confirm the alteration of lysine methylation in TRβ; our results indicated that Ezh2 increased lysine methylation of TRβ (Fig. 1c). Here, we demonstrated that TRβ is a methylated non-histone substrate and that Ezh2 triggers methylation of TRβ as a lysine methyltransferase.
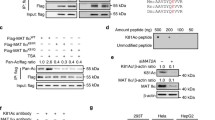
TRβ is a lysine methylation substrate and Ezh2 is a lysine methyltransferase of TRβ. a Comparison of histone-like RKS sequence containing proteins. TRβ has an RKS sequence in the amino acid sequence. b Immunoprecipitation assay was performed with an anti-methylated lysine antibody in 293 T cells. SDS-PAGE was performed for protein separation, followed by immunoblotting with a GFP antibody. TRβ is a lysine methylated substrate. c Immunoprecipitation assay was performed with an anti-methylated lysine antibody in the presence of HA-Ezh2 in 293 T cells. Ezh2 overexpression increased lysine methylation of TRβ
K40 of TRβ is a methyl acceptor site
The lysine in the RKS sequence of TRβ is the 40th amino acid (K40). We introduced a TRβ K40R point mutation to determine whether TRβ K40 is a methyl-acceptor site. An immunoprecipitation assay revealed that K40R decreased TRβ lysine methylation, indicating that K40 of TRβ is a methyl-acceptor site (Fig. 2a). Next, we investigated the binding affinity between Ezh2 with TRβ wild type (WT) and K40R. Ezh2 binds to both TRβ WT and K40R (Fig. 2b). Taken together, Ezh2 directly binds to TRβ WT and induces methylation of K40, but there was no detectable alteration in the methylation signal of TRβ K40R by Ezh2.
K40 is a methyl acceptor site of TRβ. a GFP-TRβ WT and K40R mutant were immunoprecipitated with anti-methylated lysine antibody in 293 T cells. Lysine methylation of TRβ decreased in K40R. b Flag-TRβ WT and K40R mutant were immunoprecipitated with anti-Flag antibodies to detect the interaction between the HA-Ezh2 in 293 T cells. Ezh2 binds to both TRβ WT and K40R
TRβ lysine methylation activates proteasomal degradation
The PTM of proteins plays a crucial role in biological mechanisms as it regulates protein activity, localization, protein–protein interaction, and protein stability. PTMs, such as phosphorylation and methylation, are well-known prerequisite modifications of ubiquitin-mediated protein degradation pathways in vivo (Swaney et al. 2013; Yang et al. 2009). The ubiquitination-proteasome pathway is a representative protein degradation pathway in which polyubiquitinated proteins are degraded by proteasomes (Lecker et al. 2006). In the methylation-dependent proteasomal degradation pathway, the recognized motif of the target protein is called methyl-degron (Yang and Bedford 2012).
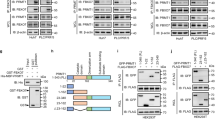
To examine the possibility that TRβ serves methyl-degron for further degradation, we performed a TRβ ubiquitination assay to confirm the alteration in TRβ stability according to the lysine methylation status of TRβ. In the TRβ ubiquitination assay with Ezh2 WT and Ezh2 ΔSET, only Ezh2 WT increased TRβ polyubiquitination, indicating that upregulation of TRβ polyubiquitination by Ezh2 requires methyltransferase activity of Ezh2 (Fig. 3a). In addition, the TRβ K40R was more stable than the TRβ WT (Fig. 3b). These data suggest that Ezh2-mediated TRβ lysine methylation destabilizes and degrades TRβ through activation of the ubiquitination-proteasome pathway.
Lysine methylation-dependent degradation of TRβ. a Ni2+-NTA pull-down assay was performed for endogenous TRβ in the presence of Ezh2 WT and Ezh2 ΔSET in 293 T cells incubated with MG132. Polyubiquitination of TRβ was increased by Ezh2 WT overexpression, but Ezh2 ΔSET overexpression did not increase polyubiquitination of TRβ. b Ni2+-NTA pull-down assay was performed for Flag-TRβ WT and K40R mutant to compare the stability of TRβ WT and K40R in 293 T cells incubated with MG132. Polyubiquitination of TRβ WT is higher than TRβ K40R
Glucose starvation increases TRβ lysine methylation and degradation
TRβ is involved in various nutrient pathways, such as lipid and glucose metabolism (Brenta 2011; Sinha et al. 2018). In the cancer microenvironment, the demand for nutrients such as glucose and amino acids is increased because of abnormal rapid cell proliferation; therefore, nutrient deprivation could occur in the cancer microenvironment (Altea-Manzano et al. 2020; Jo et al. 2020; Lieu et al. 2020). Nutrient deprivation triggers a self-digestion mechanism called autophagy, and proteins undergo PTM regulation under autophagy conditions (Russell et al. 2014; Wang and Wang 2019).
To further identify the lysine methylation mechanism and biological effect of TRβ lysine methylation under metabolic changes, we tested glucose starvation-mediated TRβ lysine methylation in hepatocellular carcinoma. HepG2 cells were incubated for 6 h in conditioned media with or without glucose. Lysine methylation of TRβ was observed to be increased by glucose starvation in these cells (Fig. 4a). In addition, we confirmed the time-dependent alteration in TRβ protein levels by glucose starvation; TRβ decreased upon glucose starvation (Fig. 4b). These results indicate that TRβ undergoes a lysine methylation-dependent proteasomal degradation pathway in autophagy signals.
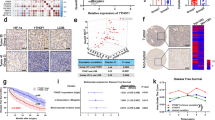
Glucose starvation increases tumorigenesis by TRβ degradation. a Immunoprecipitation assay was performed with an anti-methylated lysine antibody in glucose-starved (GS) HepG2 cells. Lysine methylation of TRβ was increased by glucose starvation. b Immunoblotting of GS HepG2 cells was performed with TRβ antibody. TRβ protein was decreased by glucose starvation. c Immunoblotting with proliferation marker in GS HepG2 cells. pAKT/pERK was increased by glucose starvation. d Real-time qPCR of Ezh2, miR21, LC3II, and ATG12 was performed in HepG2 cells. Ezh2 and miR21 were increased by glucose starvation
Glucose starvation stimulates oncogenesis in hepatocellular carcinoma
TRβ is well known as a tumor suppressor and is frequently dysregulated in cancer (Bolf et al. 2020). Autophagy can promote or suppress oncogenesis depending on the tumor type or progression. Glucose starvation-mediated TRβ degradation may affect oncogenic characteristics. AKT/ERK signaling pathways promote proliferation and are aberrantly activated in cancer, and TRβ inhibits these pathways (Davidson et al. 2021a; Suarez et al. 2010). Thus, we investigated whether glucose starvation modulates these pathways. Phosphorylated AKT/ERK (pAKT/pERK) was upregulated, which indicated oncogenesis stimulated by glucose starvation (Fig. 4c). MicroRNAs (miRNAs) play a significant role in cancer, and thyroid hormone-associated miRNAs affect hepatocellular carcinoma (Huang et al. 2019). Several studies have reported miR21 having an oncogenic potential that enhances the migration, invasion, and drug resistance of hepatocellular carcinoma (Huang et al. 2013). We performed real-time qPCR to confirm the expression of Ezh2 and miR21; the expression levels of miR21 were observed to be increased by glucose starvation. We also examined LC3II and ATG12 together as markers of autophagy induction (Fig. 4d). Together, we suggest that the glucose starvation-mediated TRβ lysine methyl-degron could promote cancer progression.
Discussion
In this study, we identified a signal integration pathway, lysine methylation of nuclear receptor TRβ, in the modulation of oncogenic Ezh2-dependent signaling pathways in hepatocellular carcinoma. Given that in a previous study, the RORα methyl-degron was found to be generated by Ezh2 (Lee et al. 2012), we explored the possible roles of lysine methylation through Ezh2 in the regulation of another non-histone substrate TRβ on a histone-like RKS sequence. We demonstrated here that Ezh2 triggers methylation of TRβ at K40 and that methylation of TRβ underlies post-translational protein instability in the regulation of metabolism in cancer cells. In hepatocellular carcinoma, in the event of glucose starvation, increased lysine methylation and further protein degradation of TRβ are required steps for the coordinated increase in oncogenic transcriptional target genes and cancer-promoting miRNA. Given that oncogenic Ezh2 inhibits the tumor suppressor TRβ under metabolic changes, it is tempting to speculate that autophagy signals, including nutrient starvation, might promote tumorigenesis in hepatocellular carcinoma, a process requiring the degradation of tumor suppressor TRβ.
Our data show that lysine methylation, a PTM regulation process of proteins, is responsible for the strong control of the protein stability of TRβ on a subset of target genes in the crosstalk between metabolism and cancer (Fig. 5). It is noteworthy that in the in vitro methylation assay only a tiny fraction of the TRβ was observed to be methylated, although lysine methylation has a large impact on the modulation of proteasomal degradation, transcriptional activity, and regulation of cancer metabolism. Recently, epigenetic modulation control had been reported as an important strategy for cancer therapy (Majchrzak-Celinska et al. 2021). Epigenetic drugs that target enzymes modulating lysine methylation have been approved by the Food and Drug Administration (Ghasemi 2020). Among the drugs regulating lysine methyltransferase, tazemetostat is the first approved Ezh2 inhibitor (Simeone et al. 2021). Currently, Ezh2 inhibitors are approved for limited therapeutic applications such as epithelioid sarcomas; however, considering the methyl-degron mechanism via Ezh2, there may be a possible broadening of the applicable therapeutic area of Ezh2 inhibitor.
In the present study, we provide evidence that autophagy induced by glucose starvation in hepatocellular carcinoma is crucial for oncogenic transition. Given that the mechanism of autophagy in cancer is complex and requires further study, we speculate that the methylation status of certain protein substrates is a critical modulator of cancer progression via autophagy. Furthermore, determining the upstream signal for lysine methylation of these proteins may shed light on the role of lysine methylation in the crosstalk between autophagy and cancer. Autophagy is known to promote or inhibit cancer progression. Here, we identified that autophagy could induce a specific protein turnover through PTM and ubiquitination. Recent studies have reported that Ezh2 and autophagy were related to drug resistance in cancer, but the underlying mechanism of resistance acquisition varied according to the type of cancer (Bai et al. 2019; Chang and Zou 2020; Sun et al. 2016). Like epigenetic drugs, inducers and inhibitors regulating autophagy have been clinically approved by the Food and Drug Administration (Liu et al. 2020). To overcome the drug resistance of cancer, combination therapy that combines epigenetic drugs and autophagy modulating agents is required in cancer therapy (Bayat Mokhtari et al. 2017). Based on our results, we suggest that a combination of an approved Ezh2 inhibitor and an autophagy inhibitor may be an efficient strategy for inhibiting hepatocellular carcinoma progression by regulating the protein levels of TRβ via the methyl-degron.
References
Ahmadiankia N, Bagheri M, Fazli M (2019) Nutrient deprivation modulates the metastatic potential of breast cancer cells. Rep Biochem Mol Biol 8(2):139–146
Altea-Manzano P, Cuadros AM, Broadfield LA, Fendt SM (2020) Nutrient metabolism and cancer in the in vivo context: a metabolic game of give and take. EMBO Rep. https://doi.org/10.15252/embr.202050635
Bai Y, Zhang Z, Cheng L, Wang R, Chen X, Kong Y, Feng F, Ahmad N, Li L, Liu X (2019) Inhibition of enhancer of zeste homolog 2 (EZH2) overcomes enzalutamide resistance in castration-resistant prostate cancer. J Biol Chem 294(25):9911–9923. https://doi.org/10.1074/jbc.RA119.008152
Bayat Mokhtari R, Homayouni TS, Baluch N, Morgatskaya E, Kumar S, Das B, Yeger H (2017) Combination therapy in combating cancer. Oncotarget 8(23):38022–38043. https://doi.org/10.1863/oncotarget.16723
Bolf EL, Gillis NE, Davidson CD, Rodriguez PD, Cozzens L, Tomczak JA, Frietze S, Carr FE (2020) Thyroid hormone receptor beta induces a tumor-suppressive program in anaplastic thyroid cancer. Mol Cancer Res 18(10):1443–1452. https://doi.org/10.1158/1541-7786.MCR-20-0282
Brenta G (2011) Why can insulin resistance be a natural consequence of thyroid dysfunction? J Thyroid Res. https://doi.org/10.4061/2011/152850
Cai Y, Zhang Y, Loh YP, Tng JQ, Lim MC, Cao Z, Raju A, Lieberman Aiden E, Li S, Manikandan L, Tergaonkar V, Tucker-Kellogg G, Fullwood MJ (2021) H3K27me3-rich genomic regions can function as silencers to repress gene expression via chromatin interactions. Nat Commun 12(1):719. https://doi.org/10.1038/s41467-021-20940-y
Chang H, Zou Z (2020) Targeting autophagy to overcome drug resistance: further developments. J Hematol Oncol 13(1):159. https://doi.org/10.1186/s13045-020-01000-2
Davidson CD, Bolf EL, Gillis NE, Cozzens LM, Tomczak JA, Carr FE (2021a) Thyroid hormone receptor beta inhibits PI3K-Akt-mTOR signaling axis in anaplastic thyroid cancer via genomic mechanisms. J Endocr Soc. https://doi.org/10.1210/jendso/bvab102
Davidson CD, Gillis NE, Carr FE (2021b) Thyroid hormone receptor beta as tumor suppressor: untapped potential in treatment and diagnostics in solid tumors. Cancers (basel). https://doi.org/10.3390/cancers13174254
Donaldson-Collier MC, Sungalee S, Zufferey M, Tavernari D, Katanayeva N, Battistello E, Mina M, Douglass KM, Rey T, Raynaud F, Manley S, Ciriello G, Oricchio E (2019) EZH2 oncogenic mutations drive epigenetic, transcriptional, and structural changes within chromatin domains. Nat Genet 51(3):517–528. https://doi.org/10.1038/s41588-018-0338-y
Galluzzi L, Pietrocola F, Levine B, Kroemer G (2014) Metabolic control of autophagy. Cell 159(6):1263–1276. https://doi.org/10.1016/j.cell.2014.11.006
Gan L, Yang Y, Li Q, Feng Y, Liu T, Guo W (2018) Epigenetic regulation of cancer progression by EZH2: from biological insights to therapeutic potential. Biomark Res 6:10. https://doi.org/10.1186/s40364-018-0122-2
Gao X, Lee HY, Li W, Platt RJ, Barrasa MI, Ma Q, Elmes RR, Rosenfeld MG, Lodish HF (2017) Thyroid hormone receptor beta and NCOA4 regulate terminal erythrocyte differentiation. Proc Natl Acad Sci USA 114(38):10107–10112. https://doi.org/10.1073/pnas.1711058114
Ghasemi S (2020) Cancer’s epigenetic drugs: where are they in the cancer medicines? Pharmacogenomics J 20(3):367–379. https://doi.org/10.1038/s41397-019-0138-5
Giammanco M, Di Liegro CM, Schiera G, Di Liegro I (2020) Genomic and non-genomic mechanisms of action of thyroid hormones and their catabolite 3,5-diiodo-l-thyronine in mammals. Int J Mol Sci. https://doi.org/10.3390/ijms21114140
Gu G, Gelsomino L, Covington KR, Beyer AR, Wang J, Rechoum Y, Huffman K, Carstens R, Ando S, Fuqua SA (2015) Targeting thyroid hormone receptor beta in triple-negative breast cancer. Breast Cancer Res Treat 150(3):535–545. https://doi.org/10.1007/s10549-015-3354-y
He L, Zhang J, Zhao J, Ma N, Kim SW, Qiao S, Ma X (2018) Autophagy: the last defense against cellular nutritional stress. Adv Nutr 9(4):493–504. https://doi.org/10.1093/advances/nmy011
Huang YH, Lin YH, Chi HC, Liao CH, Liao CJ, Wu SM, Chen CY, Tseng YH, Tsai CY, Lin SY, Hung YT, Wang CJ, Lin CD, Lin KH (2013) Thyroid hormone regulation of miR-21 enhances migration and invasion of hepatoma. Cancer Res 73(8):2505–2517. https://doi.org/10.1158/0008-5472.CAN-12-2218
Huang PS, Wang CS, Yeh CT, Lin KH (2019) Roles of thyroid hormone-associated micrornas affecting oxidative stress in human hepatocellular carcinoma. Int J Mol Sci. https://doi.org/10.3390/ijms20205220
Jo H, Lee J, Jeon J, Kim SY, Chung JI, Ko HY, Lee M, Yun M (2020) The critical role of glucose deprivation in epithelial-mesenchymal transition in hepatocellular carcinoma under hypoxia. Sci Rep 10(1):1538. https://doi.org/10.1038/s41598-020-58124-1
Kim WG, Zhao L, Kim DW, Willingham MC, Cheng SY (2014) Inhibition of tumorigenesis by the thyroid hormone receptor beta in xenograft models. Thyroid 24(2):260–269. https://doi.org/10.1089/thy.2013.0054
Kowalik MA, Perra A, Pibiri M, Cocco MT, Samarut J, Plateroti M, Ledda-Columbano GM, Columbano A (2010) TRbeta is the critical thyroid hormone receptor isoform in T3-induced proliferation of hepatocytes and pancreatic acinar cells. J Hepatol 53(4):686–692. https://doi.org/10.1016/j.jhep.2010.04.028
Lecker SH, Goldberg AL, Mitch WE (2006) Protein degradation by the ubiquitin-proteasome pathway in normal and disease states. J Am Soc Nephrol 17(7):1807–1819. https://doi.org/10.1681/ASN.2006010083
Lee JM, Lee JS, Kim H, Kim K, Park H, Kim JY, Lee SH, Kim IS, Kim J, Lee M, Chung CH, Seo SB, Yoon JB, Ko E, Noh DY, Kim KI, Kim KK, Baek SH (2012) EZH2 generates a methyl degron that is recognized by the DCAF1/DDB1/CUL4 E3 ubiquitin ligase complex. Mol Cell 48(4):572–586. https://doi.org/10.1016/j.molcel.2012.09.004
Lieu EL, Nguyen T, Rhyne S, Kim J (2020) Amino acids in cancer. Exp Mol Med 52(1):15–30. https://doi.org/10.1038/s12276-020-0375-3
Liu T, Zhang J, Li K, Deng L, Wang H (2020) Combination of an autophagy inducer and an autophagy inhibitor: a smarter strategy emerging in cancer therapy. Front Pharmacol 11:408. https://doi.org/10.3389/fphar.2020.00408
Majchrzak-Celinska A, Warych A, Szoszkiewicz M (2021) Novel approaches to epigenetic therapies: from drug combinations to epigenetic editing. Genes (basel). https://doi.org/10.3390/genes12020208
Mangelsdorf DJ, Thummel C, Beato M, Herrlich P, Schutz G, Umesono K, Blumberg B, Kastner P, Mark M, Chambon P, Evans RM (1995) The nuclear receptor superfamily: the second decade. Cell 83(6):835–839. https://doi.org/10.1016/0092-8674(95)90199-x
Nikolaou S, Machesky LM (2020) The stressful tumour environment drives plasticity of cell migration programmes, contributing to metastasis. J Pathol 250(5):612–623. https://doi.org/10.1002/path.5395
Porlan E, Vidaurre OG, Rodriguez-Pena A (2008) Thyroid hormone receptor-beta (TR beta 1) impairs cell proliferation by the transcriptional inhibition of cyclins D1, E and A2. Oncogene 27(19):2795–2800. https://doi.org/10.1038/sj.onc.1210936
Prabakaran S, Lippens G, Steen H, Gunawardena J (2012) Post-translational modification: nature’s escape from genetic imprisonment and the basis for dynamic information encoding. Wiley Interdiscip Rev Syst Biol Med 4(6):565–583. https://doi.org/10.1002/wsbm.1185
Russell RC, Yuan HX, Guan KL (2014) Autophagy regulation by nutrient signaling. Cell Res 24(1):42–57. https://doi.org/10.1038/cr.2013.166
Simeone N, Frezza AM, Zaffaroni N, Stacchiotti S (2021) Tazemetostat for advanced epithelioid sarcoma: current status and future perspectives. Future Oncol 17(10):1253–1263. https://doi.org/10.2217/fon-2020-0781
Simon JA, Lange CA (2008) Roles of the EZH2 histone methyltransferase in cancer epigenetics. Mutat Res 647(1–2):21–29. https://doi.org/10.1016/j.mrfmmm.2008.07.010
Singh BK, Yen PM (2017) A clinician’s guide to understanding resistance to thyroid hormone due to receptor mutations in the TRalpha and TRbeta isoforms. Clin Diabetes Endocrinol 3:8. https://doi.org/10.1186/s40842-017-0046-z
Sinha RA, Singh BK, Yen PM (2018) Direct effects of thyroid hormones on hepatic lipid metabolism. Nat Rev Endocrinol 14(5):259–269. https://doi.org/10.1038/nrendo.2018.10
Suarez J, Scott BT, Suarez-Ramirez JA, Chavira CV, Dillmann WH (2010) Thyroid hormone inhibits ERK phosphorylation in pressure overload-induced hypertrophied mouse hearts through a receptor-mediated mechanism. Am J Physiol Cell Physiol 299(6):C1524-1529. https://doi.org/10.1152/ajpcell.00168.2010
Sun Y, Jin L, Liu JH, Sui YX, Han LL, Shen XL (2016) Interfering EZH2 expression reverses the cisplatin resistance in human ovarian cancer by inhibiting autophagy. Cancer Biother Radiopharm 31(7):246–252. https://doi.org/10.1089/cbr.2016.2034
Swaney DL, Beltrao P, Starita L, Guo A, Rush J, Fields S, Krogan NJ, Villen J (2013) Global analysis of phosphorylation and ubiquitylation cross-talk in protein degradation. Nat Methods 10(7):676–682. https://doi.org/10.1038/nmeth.2519
Tan JZ, Yan Y, Wang XX, Jiang Y, Xu HE (2014) EZH2: biology, disease, and structure-based drug discovery. Acta Pharmacol Sin 35(2):161–174. https://doi.org/10.1038/aps.2013.161
Wang R, Wang G (2019) Protein modification and autophagy activation. Adv Exp Med Biol 1206:237–259. https://doi.org/10.1007/978-981-15-0602-4_12
Wu H, Zeng H, Dong A, Li F, He H, Senisterra G, Seitova A, Duan S, Brown PJ, Vedadi M, Arrowsmith CH, Schapira M (2013) Structure of the catalytic domain of EZH2 reveals conformational plasticity in cofactor and substrate binding sites and explains oncogenic mutations. PLoS ONE 8(12):e83737. https://doi.org/10.1371/journal.pone.0083737
Yang Y, Bedford MT (2012) Titivated for destruction: the methyl degron. Mol Cell 48(4):487–488. https://doi.org/10.1016/j.molcel.2012.11.007
Yang XD, Lamb A, Chen LF (2009) Methylation, a new epigenetic mark for protein stability. Epigenetics 4(7):429–433. https://doi.org/10.4161/epi.4.7.9787
Yun CW, Lee SH (2018) The roles of autophagy in cancer. Int J Mol Sci. https://doi.org/10.3390/ijms19113466
Acknowledgements
This work was supported by Basic Science Research Program NRF-2021R1C1C1008780 to J.M.L from the National Research Foundation (NRF) grant funded by the Korean government (MSIT). The figures were generated using BioRender with a paid license to publish.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Park, S.C., Lee, J.M. Ezh2 promotes TRβ lysine methylation-mediated degradation in hepatocellular carcinoma. Genes Genom 44, 369–377 (2022). https://doi.org/10.1007/s13258-021-01196-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-021-01196-8