Abstract
The genus Bacillus is a Gram-positive, aerobic, endospore-forming, rod-shaped bacterium commonly found in the environment that have important industrial, medical, agriculture and environmental values. Here, we report the whole genome sequence analysis of UMX-103 which was isolated from a hydrocarbon contaminated site in Terengganu, Malaysia. An integration of both genomics and chemical approaches were conducted to analyse the biosurfactant production by the strain UMX-103. The genome was assembled using a combination of both de novo and reference-guided assembly methods. The genome size of UMX-103 is 4,234,627 bp with 4399 genes comprising of 4301 protein-coding genes and 98 RNA genes. The mapping results showed 93.44% of genome similarity with B. subtilis strain 168. We have identified 25 genes involved in biosurfactants production. Among the 25 identified genes, 14 genes were involved in surfactin biosynthesis and 11 genes were implicated in surfactin regulation. Fifteen genomic islands were identified which are different from other closely related Bacillus species. In addition, our study also revealed the genetic contents of this bacterium and genes which are involved in biosurfactant production.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Biosurfactants are amphiphilic molecule produced by microorganism that has the ability to lower the surface and interfacial tension between two liquids. They are environmental friendly due to their low toxicity level, extreme biodegradability, stability to high pH and temperature condition which makes them as promising candidate for bioremediation and enhanced oil recovery applications (Mulligan 2009). They are also used in food industry (Nitschke and Costa 2007), pharmaceutical and cosmetics (Banat et al. 2000). Biosurfactants are alternative to the synthetic surfactants; however, the high cost of production limits their application.
There are many bacteria have been reported to produce biosurfactants such as P. aeruginosa N002 (Das et al. 2015), Achromobacter spanius (Alvarez et al. 2015), and Rhodococcus erythroplis (Cai et al. 2015a). Besides that, many strains from genus Bacillus are biosurfactant producers such as B. licheniformis, B. subtilis, and B. pumilus (Płaza et al. 2015a). Bacillus species are genetically diverse and also grow in various environments (Choudhary and Johri 2009). Bacillus species are one of the main sources for producing industrial enzymes such as amylases and protease (Karataş et al. 2013; Kunst et al. 1997).
In the past decade, microbial research was limited to the need to grow bacteria in culture. The revolution of DNA sequencing technology has changed the way scientists think about genetics and genomics information (Lasken and McLean 2014; Zhang et al. 2011). The first complete genome of Bacillus subtilis was published on 20th November 1997 (Kunst et al. 1997). DNA sequencing applications allowed many researchers and scientists to sequence the whole genome or a region of a microbial genome (Kamada et al. 2014; Nishito et al. 2010). To date, the whole genome sequencing application is widely used to analyse microbial biosurfactant producers (Das et al. 2015; Shaligram et al. 2016). Here we report the whole genome sequencing and analysis of the new strain UMX-103, isolated from Terengganu, Malaysia, with the potential for biosurfactant production.
Materials and methods
Sample isolation and preparation
The strain UMX-103 was isolated from a hydrocarbon contaminated site in Terengganu, Malaysia. The bacterium was cultured in Tryptone Soya Agar (TSA) (Merck KGaA, Germany) and incubated overnight at 30 °C. Optimal colony was selected from the cultured bacteria and inoculated in 50 ml of Tryptone Soya Broth (TSB) (Merck KGaA, Germany) using 200 ml conical flask. The broth was incubated overnight at 30 °C in an orbital shaker at 121 rpm.
Determination of bacteria morphology
Gram staining was used to stain the bacteria using GCC Diagnostics (Gainland Chemical Co, UK). Staining was performed according to the manufacture’s protocol. Field Emission Electron Microscope (FESEM)(Quanta 450 FEG, USA) was used to visualize the morphology of the bacteria.
Screening of UMX-103 for capability of producing biosurfactants
The ability of biosurfactant production by UMX-103 was tested using five different methods; (i) hemolytic assay, (ii) oil spreading test, (iii) drop-collapse assay, (iv) emulsification assay, and (v) surface tension measurements. Hemolysis assay on blood agar plates has been widely used as a method to screen surfactant producing bacteria (Banat 1993; Morán et al. 2002; Mulligan et al. 1984; Yonebayashi et al. 2000). In this study, the isolated strain UMX-103 was streaked onto blood agar plates and incubated at 30 °C for 24 h. The plate was visually inspected for clear zone formation around the colonies, which is an indicative of biosurfactant production.
The oil spreading test is to observe a clear zone formation which is a result of dropping a biosurfactant or surfactant solution on an oil–water interface. Approximately, 15 µl of engine oil (10W-40 Shell®) was added to 40 ml of distilled water in a petri dish (150 mm in diameter) to form a thin layer of oil on the surface. Then, 15 µl of culture supernatant was added to the central of the oil layer (Morikawa et al. 1993, 2000; Youssef et al. 2004). Diameter of the clear zone formation on the oil layer was observed and measured after 30 s (Morikawa et al. 2000). We have used (Triton® X-100, USA) as the positive control, while distilled water as negative control (Shoeb et al. 2015).
Drop-collapse qualitative test was conducted on a polystyrene 96-microwell (12.7 × 8.5) plate. Approximately, 2 µl of engine oil (10-40 Shell®) was dropped into the well and equilibrated for 1 h at the room temperature. Then, 5 µl of the culture supernatant was added on the top of oil surface. Water was used as a negative control (Bodour et al. 2003; Shoeb et al. 2015). The shape of the drop on the oil surface was observed after 1 min. In the presence of biosurfactant, the drop will be collapsed in the oil surface.
Emulsification test was performed to check the ability of biosurfactant to emulsify the oil surface. Initially, 5 ml of 50mM Tris buffer (8.0 pH) was added in 30-ml screw-caped test tube. Then, 5 ml of engine oil (10-40 Shell®) was added to the Tris buffer and vortex at room temprature for 2 min. The screw-caped test tube with the mix solution was stabilized for 24 h. The absorbance of aqueous phase was measured by spectrophotometer (Spectroquant® Pharo100, USA) at the wavelength of 400 nm. Distilled water was used as a negative control, while Triton-X as the positive control (Shoeb et al. 2015). The emulsification activity was calculated (Cai et al. 2015b) as stated below:
For the surface tension measurement the bacteria culture was centrifuged at 3000 rpm for 25 min to obtain a cell-free supernatant. The surface tension of the culture supernatant was determined by the Du Nouy ring method (Gudina et al. 2010; Pereira et al. 2013) using interfacial tensiometer (Force Tensiometer, Sigma700, Biolin Scientific) at room temperature. The measurements of the surface tension were repeated three times and an average value was obtained (Cai et al. 2015b; Pereira et al. 2013; Vaz et al. 2012).
DNA preparation and whole genome sequencing
The whole genome sequence of Bacillus subtilis UMX-103 was obtained from Illumina HiSeq 2000 sequencing technology (Illumina, USA). The DNA was extracted using phenol–chloroform method and the quality and quantity of the DNA was measured using QIAxpert (QIAGEN, Germany). The sample was run on 1.2% agarose gel to determine the integrity of genomic DNA. Fragmentation of the DNA was performed using Covaris S220 (Covaris Inc, USA). Ligation to NEBNext adapters was conducted using NEBNext Ultra, while the PCR-enrichment using the DNA Library Prep Kit (NEB, USA). The final library was quantified using KAPA kit (KAPA Biosystem, USA). Library size was confirmed using Agilent Bioanalyzer High Sensitive DNA Chip (Agilent, USA). The prepared library was sequenced using an Illumina flow cell, consisting of 2 × 100 cycles.
De novo assembly by Velvet and mapping the reads to reference genome by BWA
Quality control assessment was performed using Trimmomatic 0.35 (Bolger et al. 2014). The generated dataset was assembled using Velvet 1.2.10 (Zerbino and Birney 2008) which is a de novo assembly software that use de Bruijn graph algorithm. SSPACE-Standard v3.0 (Boetzer et al. 2011) was used for scaffolding the generated contigs from Velvet assembler. GapFiller v1.10 (Boetzer and Pirovano 2012) was used to close the gaps and replaced the unknown nucleotide with known nucleotides. The scaffolds were sorted along with the reference genome (Bacillus subtilis strain 168; accession number NC_000964.3) using Mauve 2.3.1 (Darling et al. 2004). BWA (Li 2013) was used for mapping the reads to the reference genomes.
Gene prediction and functional annotation
Gene prediction for protein-coding genes was conducted using Prodigal (Hyatt et al. 2010). The tRNA and rRNA screenings were performed using tRNAscan-SE v1.3.1 (Lowe and Eddy 1997) and RNAmmer v1.2 (Lagesen et al. 2007), respectively. Gene annotation was conducted using Prokka v1.11 (Seemann 2014). The functional annotation was performed using EggNOG-mapper 4.5.1 database (Huerta-Cepas et al. 2016). The annotated genes were submitted to IslandViewer3 (Dhillon et al. 2015) to identify the genomic islands in the genome. Pan genome analysis and comparison between all the prospective reference genomes were conducted using Roary version 3.6.1 (Larsen et al. 2012).
Phylogenetic and genomic similarity analysis
16S ribosomal DNA was used to identify the bacteria. The 16S rRNA was obtained from the whole genome sequence and aligned with other 16S rRNA genes from different Bacillus strains. The 16S genes were extracted from each reference strains using RNAmmer (Lagesen et al. 2007). Molecular Evolutionary Genetics Analysis (MEGA) version 7 (Kumar et al. 2016) was used to align and construct the distanced phylogenetic tree. The phylogenetic tree and distance were constructed using Neigbour-joing method. The Average Nucleotide Identity of UMX-103 was determined using GGDC 2.1 server (Meier-Kolthoff et al. 2013). Multilocus Sequence Typing (MLST) was predicted using MLST 1.8 server (Larsen et al. 2012).
Results and discussion
Morphology of UMX-103
The Gram staining showed that UMX-103 is a Gram positive strain. FESEM result shows the morphology of the colony which is a rod shape, with size of 1.954 µm length and 540.9 nm width (Fig. 1). Based on the Gram staining and FESEM results we confirmed that UMX-103 belongs to the genus Bacillus.
Biosurfactant activity
The biosurfactant producing capability of UMX-103 was tested in five different assays. The results are shown in Table 1. Bernheimer and Avigad (1970) reported that surfactin produced by Bacillus subtilis lyse the red blood cells. There is an association between hemolysis activity and surfactant production, since then hemolytic assay is recommended as a primary technique to screen biosurfactant producers (Youssef et al. 2004). Therefore, this assay was employed in this research. UMX-103 demonstrated beta lysis as it produced a clear zone around the colony which determines the biosurfactant production by the strain. Biosurfactants are well known to have hemolytic, antibacterial, and antiviral activity, owning a precise mechanism that has impact on the membrane permeability and eventually leading to cell disruption (Heerklotz and Seelig 2007).
Oil spreading assay is based on the formation of a clear zone and a displacement area, in the presence of biosurfactant in the culture supernatant. The diameter of this clear zone on the oil surface correlates to the amount of biosurfactant produced. In this study supernatant of UMX-103 culture formed a clear zone and oil displacement region about 2 cm for as indication of biosurfactant production.
The drop-collapse test depends on the destabilization of liquid droplets by the biosurfactants produced by the bacterial isolate. The stability of drops is dependent on biosurfactant concentration and correlates with the surface and interfacial tension (Sari et al. 2014). In this study, the distilled water was used as a negative control and there is no droplet collapsing was observed. Whereas, the biosurfactant produced by UMX-103 was tested positive, where the droplet was collapsed.
The emulsification test was used to evaluate the emulsification ability of the isolate UMX-103. We have observed a positive activity of the strain where it emulsifies the oil surface. In this study, Triton-X was used as positive control due to it emulsification ability and it have been widely used as positive control (Shoeb et al. 2015).
The measurement of surface tension using Du nouy ring method is based on measurement of the force required to detach the ring from the culture supernatant surface. The detachment force is directly proportional to the interfacial tension. Our test results showed that UMX-103 has a higher reduction ability which is 26.4 ± 0.02 mN/m. While, Triton X showed reduction value of 34.3 ± 0.003 mN/m and the hexane with the lowest reduction value of 18.1 ± 0.06 mN/m.. Summary of the biosurfactant assays is presented in Table 1.
Mapping reads to the reference genome
A total of 565,068,437 paired-end reads with a length of 101 bp were generated, with average insertion size of 534 bp. Low quality bases and reads were filtered to get an optimal quality score of 30 or higher at each base. The reads were mapped to Bacillus subtilis strain 168, where results showed that 93.44% of the generated reads are mapped to the reference genome.
De novo assembly and scaffold sorting
Velvet assembler generates a total of 69 contigs with an average length of 61,362 bp and with maximum and minimum length of 869,096 and 137 bp, respectively (Table 2). All contigs generated by Velvet were used to generate the scaffolds using SSPACE-Standard software (Boetzer et al. 2011). Scaffolding process generated a total of 39 scaffolds, with average scaffold size of 108,565 bp and with maximum and minimum scaffold sizes of 1,059,836 and 144 bp, respectively (Table 2). Then, GapFiller software (Boetzer and Pirovano 2012) was used to close the gaps in the generated scaffolds. Total of 34 gaps from 41 gaps were closed. In addition, the result after gap closing shows a total of 39 scaffold with an average size of 108,580 bp (Table 2). The scaffolds were sorted according to the reference genome using MAUVE (Rissman et al. 2009).
Gene prediction and functional annotation
The strain UMX-103 contains a single circular chromosome of 4,234,627 bp with an average G+C content of 43.41% (Table 3). The assembled genome consists of 39 scaffolds with an average scaffold size of 108,580 bp (Table 2). By using a combination of several gene-prediction software and manual inspection, a total of 4301 protein-coding genes and 98 RNA genes were identified in this strain (Fig. 2). The annotated genes which identified using Prokka software were used for functional annotation analysis. The functional annotation was conducted using EggNOG-mapper, the summary of functional categories of annotated genes is shown in Table 4. The result revealed existing of biosynthetic gene cluster of genes which are known for coding surfactin. This gene cluster belongs to Nonribosomal Peptide Synthetase (NRPS) family, particularly to the microbial surfactants group. These genes usually present in secondary metabolites biosynthesis, transport and catabolism (Doroghazi et al. 2014).
Genome features of UMX-103: the two outmost concentric circles denote the predicted protein-coding genes represent as forward strand (external blue circle) and the reverse strand (internal grey). The third concentric circle (purple) represents tRNAs while the fourth concentric circle (light brown) shows rRNAs genes. The fifth concentric (green and purple) represents the GC content. The green colour shows GC content more than the average while the purple colour shows the GC content below average. Purple and silver in the last inner concentric represent GC skew
Phylogenetic and genomic similarity
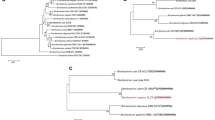
The 16S rRNA from UMX-103 was used to carry out the phylogenetic analysis where it was aligned with other 16S rRNA genes of Bacillus strains which including; B. subtilis LM 4-2, B. subtilis BEST7003, B. subtilis KCTC 1028, B. subtilis 168, B. subtilis RO-NN-1, B. amyloliquefaciens DSM7, B. licheniformis, B. pumilus GR-8, and Paenibacillus macerans (Fig. 3). Average Nucleotide Identity (ANI) of UMX-103 was determined by comparing the whole genome with the selected references (Table 5). The highest ANI was detected with KCTC 1028 and 168 strains which is 89%. The seven housekeeping genes used to determine the species of the UMX-103 were detected using MLSTserver 1.8 (Larsen et al. 2012) (Supplementary Table 1). All of the housekeeping genes in UMX-103 are highly identical with the housekeeping genes of Bacillus subtilis.
Phylogenetic analysis based on 16S rRNA. Phylogenetic reconstruction was performed based on the sequence of 16S rRNA gene using MEGA7 (Kumar et al. 2016). The 16S rRNA genes sequence of Bacillus pumilus GR-8 and Paenibacillus macerans was used as outgroup
Genomic Islands and horizontal genes transfer
Genomic islands is widely used to compare bacteria strains and identify essential genes in bacterial genome (Dobrindt et al. 2004; Langille et al. 2010). Basically genomic islands associate with horizontal gene transfer (HGT) which is also known as mobile genetic elements. The strain UMX-103 has witnessed a number of Horizontal Gene Transfer events. There are 15 genomic islands in UMX-103 that was predicted by IslandViewer 3 (Dhillon et al. 2015) and the localization of the predicted genomic islands is shown in (Fig. 4). The 15 predicted genomic islands consist of 331 genes (Supplementary Table 2). These genomic islands are possibly having a significant role in adapting and surviving the bacteria to different abiotic stress and antimicrobial resistance, which may occur after the bacteria exposed to different environment including the hydrocarbon contaminated soil. Features of the genomic islands in the strain UMX-103 are given in (Supplementary Table 3).
Genomic comparisons with closely related bacteria
Comparative genomics of UMX-103 with six related genomes (Table 6) showed that UMX-103 genome is closely related to Bacillus subtilis KCTC 1028 and Bacillus subtilis 168 with the genome sequence similarity of 93.99 and 93.44%, respectively. Analysis showed that UMX-103 has the largest genome size compared to the other bacteria strains studied. The genome contains the highest number of genes and the lowest GC contents which is 43.41%. Pangenome analysis resulted in the identification of 3434 core genes which present in all the Bacillus strains studied (Fig. 5). Pangenome composed of the essential genes in species and it also used as a method in identification of unknown bacteria (Lasken and McLean 2014). The genomic islands comparison (Table 7) showed that UMX-103 has the same number of genomic islands with Bacillus subtilis LM 4-2; however, the total number of genes in the genomic islands of UMX-103 is 322 genes, where only 108 genes were found in Bacillus subtilis LM 4-2.
Biosurfactant production genes
Based on the gene prediction and annotation analyses, we have identified 25 genes in UMX-103 which are involved in biosurfactant production. The list of the identified genes is presented in Supplementary Table 4. These genes involved in the biosynthesis and regulation of surfactin, which is a type of biosurfactant produced by Bacillus subtilis with high industrial value (Płaza et al. 2015a). The genes that involved in the biosynthesis of biosurfactant are including; 4-phosphopantetheinyl transferase (sfp), glucose-1-phosphate thymidyly transferase (rmlA), dTDP-glucose 4,6-dehydratase (rmlB), dTDP-4-dehydrorhamnose 3,5-epimerase (rmlC), dTDP-4-dehydrorhamnose reductase (rmlD) (Das et al. 2015), and non-ribosomal peptide synthetase (dhbF) (May et al. 2001).
In this study, we have also identified two operons srfA (Nakano et al. 1991) and pps (Coutte et al. 2010) which are involved in coding of the non-ribosomal peptide synthetase (NRPS) subunits, that catalyse the incorporation of the seven amino acid form surfactin (Coutte et al. 2010; Peypoux et al. 1999). The srfA operon contains four genes; srfAA, srfAB, srfAC, and srfAD, while the operon pps contains five genes; ppsA, ppsB, ppsC, ppsD, and ppsE. The srfA operon encodes surfactin synthetase subunits (Płaza et al. 2015a). Surfactin is made of seven amino acids which are (Glu–Leu–(D)Leu–Val–Asp–(D)Leu–Leu) (Cosmina et al. 1993). The gene srfAA encodes the peptide synthetize subunit which involved in the makeup of amino acids Glu, Leu and D-leu. Whereas, srfAB encodes the subunit that implicate in catalysis of Val, Asp, and D-leu. The third gene in the operon srfAC functions in the foundation of Leu amino acid. The surfactin synthase thioesterase subunit is produced by srfAD (Marahier et al. 1993).The activating enzyme sfp plays essential role in surfactin biosynthesis, as it tranforms the inactive protein that changes surfactin synthetase into an active form (Nakano et al. 1992; Płaza et al. 2015b).
In addition, we have also identified genes implicated in regulation of surfactin; comA and comP which comprise a signal transduction system that involved in the competence development pathway and is required for the transcription of srfA (Marahier et al. 1993; Nakano et al. 1992). The rest of the genes are involved in DNA-binding response, sporulation, phosphate regulon transcription, carbon storage, and sensory transduction protein (Supplementary Table 4).
Our results suggested that presence of biosurfactant genes in UMX-103 has the ability to produce surfactin which is lipopeptide biosurfactant. These results are in agreement with our earlier screening assays.
Conclusion
We conclude that, the new strain UMX-103 belongs to Bacillus subtilis species. It is a Gram positive bacteria, rod shape and with a length of 1.954 µm and a diameter of 540.9 nm. All the five different biosurfactant producing tests showed that UMX-103 has the capability to produce biosurfactant. In addition, we have successfully assembled the genome of the strain UMX-103 using a combination of both de novo and reference-guided assembly methods. The genome was assembled into 39 scaffolds with a size of 4,234,627 bp. Interestingly, we identified 25 genes which are involved in biosurfactant production, where 14 genes involved in biosynthesis and 11 genes associated with the gene regulation. Genomic analysis revealed that UMX-103 has the genes which promote biosurfactant production. Future work will be conducted to characterize the unknown function genes as well as biosurfactant genes using various Omics approaches.
References
Alvarez VM, Jurelevicius D, Marques JM, de Souza PM, de Araújo LV, Barros TG, de Souza RO, Freire DM, Seldin L (2015) Bacillus amyloliquefaciens TSBSO 3.8, a biosurfactant-producing strain with biotechnological potential for microbial enhanced oil recovery. Colloids Surf B 136:14–21
Banat IM (1993) The isolation of a thermophilic biosurfactant producing Bacillus sp. Biotechnol Lett 15:591–594
Banat IM, Makkar RS, Cameotra S (2000) Potential commercial applications of microbial surfactants. Appl Microbiol Biotechnol 53:495–508
Bernheimer A, Avigad LS (1970) Nature and properties of a cytolytic agent produced by Bacillus subtilis. Microbiology 61:361–369
Bodour AA, Drees KP, Maier RM (2003) Distribution of biosurfactant-producing bacteria in undisturbed and contaminated arid southwestern soils. Appl Environ Microbiol 69:3280–3287
Boetzer M, Pirovano W (2012) Toward almost closed genomes with GapFiller. Genome Biol 13:R56
Boetzer M, Henkel CV, Jansen HJ, Butler D, Pirovano W (2011) Scaffolding pre-assembled contigs using SSPACE. Bioinformatics 27:578–579
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. doi:10.1093/bioinformatics/btu170
Cai Q, Zhang B, Chen B, Song X, Zhu Z, Cao T (2015a) Screening of biosurfactant-producing bacteria from offshore oil and gas platforms in North Atlantic Canada. Environ Monit Assess 187:1–12
Cai Q, Zhang B, Chen B, Song X, Zhu Z, Cao T (2015b) Screening of biosurfactant-producing bacteria from offshore oil and gas platforms in North Atlantic Canada. Environ Monit Assess 187:1–12
Choudhary DK, Johri BN (2009) Interactions of Bacillus spp. and plants–with special reference to induced systemic resistance (ISR). Microbiol Res 164:493–513
Cosmina P, Rodriguez F, Ferra F, Grandi G, Perego M, Venema G, Sinderen D (1993) Sequence and analysis of the genetic locus responsible for surfactin synthesis in Bacillus subtilis. Mol Microbiol 8:821–831
Coutte F et al (2010) Effect of pps disruption and constitutive expression of srfA on surfactin productivity, spreading and antagonistic properties of Bacillus subtilis 168 derivatives. J Appl Microbiol 109:480–491
Darling AC, Mau B, Blattner FR, Perna NT (2004) Mauve: multiple alignment of conserved genomic sequence with rearrangements. Genome Res 14:1394–1403
Das D, Baruah R, Roy AS, Singh AK, Boruah HPD, Kalita J, Bora TC (2015) Complete genome sequence analysis of Pseudomonas aeruginosa N002 reveals its genetic adaptation for crude oil degradation. Genomics 105:182–190
Dhillon BK, Laird MR, Shay JA, Winsor GL, Lo R, Nizam F, Pereira SK, Waglechner N, McArthur AG, Langille MG et al (2015) IslandViewer 3: more flexible, interactive genomic island discovery, visualization and analysis. Nucleic Acids Res. doi:10.1093/nar/gkv401
Dobrindt U, Hochhut B, Hentschel U, Hacker J (2004) Genomic islands in pathogenic and environmental microorganisms. Nat Rev Microbiol 2:414–424
Doroghazi JR, Albright JC, Goering AW, Ju KS, Haines RR, Tchalukov KA, Labeda DP, Kelleher NL, Metcalf WW (2014) A roadmap for natural product discovery based on large-scale genomics and metabolomics. Nat Chem Biol 10:963–968
Gudina EJ, Teixeira JA, Rodrigues LR (2010) Isolation and functional characterization of a biosurfactant produced by Lactobacillus paracasei. Colloids Surf B 76:298–304
Heerklotz H, Seelig J (2007) Leakage and lysis of lipid membranes induced by the lipopeptide surfactin. Eur Biophys J 36:305–314
Huerta-Cepas J, Forslund K, Szklarczyk D, Jensen LJ, von Mering C, Bork P (2016) Fast genome-wide functional annotation through orthology assignment by eggNOG-mapper. bioRxivorg. doi:10.1101/076331
Hyatt D, Chen G-L, LoCascio PF, Land ML, Larimer FW, Hauser LJ (2010) Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinform 11:119
Kamada M, Hase S, Sato K, Toyoda A, Fujiyama A, Sakakibara Y (2014) Whole genome complete resequencing of Bacillus subtilis natto by combining long reads with high-quality short reads. PLoS ONE 9:e109999
Karataş H, Uyar F, Tolan V, Baysal Z (2013) Optimization and enhanced production of α-amylase and protease by a newly isolated Bacillus licheniformis ZB-05 under solid-state fermentation. Ann Microbiol 63:45–52
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol Biol Evol. doi:10.1093/molbev/msw054
Kunst F, Ogasawara N, Moszer I, Albertini AM, Alloni G, Azevedo V, Bertero MG, Bessières P, Bolotin A, Borchert S et al (1997) The complete genome sequence of the gram-positive bacterium Bacillus subtilis. Nature 390:249–256
Lagesen K, Hallin P, Rødland EA, Stærfeldt H-H, Rognes T, Ussery DW (2007) RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res 35:3100–3108
Langille MG, Hsiao WW, Brinkman FS (2010) Detecting genomic islands using bioinformatics approaches. Nat Rev Microbiol 8:373–382
Larsen MV, Cosentino S, Rasmussen S, Friis C, Hasman H, Marvig RL, Jelsbak L, Sicheritz-Pontén T, Ussery DW, Aarestrup FM et al (2012) Multilocus sequence typing of total genome sequenced bacteria. J Clin Microbiol. doi:10.1128/JCM.06094-11
Lasken RS, McLean JS (2014) Recent advances in genomic DNA sequencing of microbial species from single cells. Nat Rev Genet 15:577–584
Li H (2013) Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv preprint arXiv:13033997
Lowe TM, Eddy SR (1997) tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res 25:0955–0964
Marahier M, Nakano M, Zuber P (1993) Regulation of peptide antibiotic production in Bacillus. Mol Microbiol 7:631–636
May JJ, Wendrich TM, Marahiel MA (2001) The dhb operon of Bacillus subtilisEncodes the biosynthetic template for the catecholic siderophore 2,3-dihydroxybenzoate-glycine-threonine trimeric ester Bacillibactin. J Biol Chem 276:7209–7217
Meier-Kolthoff JP, Auch AF, Klenk H-P, Göker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform 14:1
Morán AC, Martinez MA, Siñeriz F (2002) Quantification of surfactin in culture supernatants by hemolytic activity. Biotechnol Lett 24:177–180
Morikawa M, Daido H, Takao T, Murata S, Shimonishi Y, Imanaka T (1993) A new lipopeptide biosurfactant produced by Arthrobacter sp. strain MIS38. J Bacteriol 175:6459–6466
Morikawa M, Hirata Y, Imanaka T (2000) A study on the structure–function relationship of lipopeptide biosurfactants. Biochim Biophys Acta 1488:211–218
Mulligan CN (2009) Recent advances in the environmental applications of biosurfactants. Curr Opin Interface Sci 14:372–378
Mulligan CN, Cooper DG, Neufeld RJ (1984) Selection of microbes producing biosurfactants in media without hydrocarbons. J Ferment Technol 62:311–314
Nakano M, Magnuson R, Myers A, Curry J, Grossman A, Zuber P (1991) srfA is an operon required for surfactin production, competence development, and efficient sporulation in Bacillus subtilis. J Bacteriol 173:1770–1778
Nakano MM, Corbell N, Besson J, Zuber P (1992) Isolation and characterization of sfp: a gene that functions in the production of the lipopeptide biosurfactant, surfactin, in Bacillus subtilis. Mol Gen Genet 232:313–321
Nishito Y, Osana Y, Hachiya T, Popendorf K, Toyoda A, Fujiyama A, Itaya M, Sakakibara Y (2010) Whole genome assembly of a natto production strain Bacillus subtilis natto from very short read data. BMC Genom 11:1
Nitschke M, Costa S (2007) Biosurfactants in food industry. Trends Food Sci Technol 18:252–259
Pereira JF, Gudiña EJ, Costa R, Vitorino R, Teixeira JA, Coutinho JA, Rodrigues LR (2013) Optimization and characterization of biosurfactant production by Bacillus subtilis isolates towards microbial enhanced oil recovery applications. Fuel 111:259–268
Peypoux F, Bonmatin J, Wallach J (1999) Recent trends in the biochemistry of surfactin. Appl Microbiol Biotechnol 51:553–563
Płaza G, Chojniak J, Rudnicka K, Paraszkiewicz K, Bernat P (2015a) Detection of biosurfactants in Bacillus species: genes and products identification. J Appl Microbiol 119:1023–1034
Płaza G, Chojniak J, Rudnicka K, Paraszkiewicz K, Bernat P (2015b) Detection of biosurfactants in Bacillus species: genes and products identification. J Appl Microbiol 119:1023–1034
Rissman AI, Mau B, Biehl BS, Darling AE, Glasner JD, Perna NT (2009) Reordering contigs of draft genomes using the Mauve aligner. Bioinformatics 25:2071–2073
Sari M, Kusharyoto W, Artika IM (2014) Screening for biosurfactant-producing yeast: confirmation of biosurfactant production. Biotechnology 13:106
Seemann T (2014) Prokka: rapid prokaryotic genome annotation. Bioinformatics. doi:10.1093/bioinformatics/btu153
Shaligram S, Kumbhare SV, Dhotre DP, Muddeshwar MG, Kapley A, Joseph N, Purohit HP, Shouche YS, Pawar SP (2016) Genomic and functional features of the biosurfactant producing Bacillus sp. AM13. Funct Integr Genom 16:557–566
Shoeb E, Ahmed N, Akhter J, Badar U, Siddiqui K, Ansari FA, Waqar M, Imtiaz S, Akhtar N, Shaikh QA et al (2015) Screening and characterization of biosurfactant-producing bacteria isolated from the Arabian Sea coast of Karachi. Turk J Biol 39:210–216
Vaz DA, Gudiña EJ, Alameda EJ, Teixeira JA, Rodrigues LR (2012) Performance of a biosurfactant produced by a Bacillus subtilis strain isolated from crude oil samples as compared to commercial chemical. Colloids Surf B 89:167–174
Yonebayashi H, Yoshida S, Ono K, Enomoto H (2000) Screening of microorganisms for microbial enhanced oil recovery processes. Sekiyu Gakkai Shi 43:59–69
Youssef NH, Duncan KE, Nagle DP, Savage KN, Knapp RM, McInerney MJ (2004) Comparison of methods to detect biosurfactant production by diverse microorganisms. J Microbiol Methods 56:339–347
Zerbino DR, Birney E (2008) Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res 18:821–829
Zhang J, Chiodini R, Badr A, Zhang G (2011) The impact of next-generation sequencing on genomics. J Genet Genom 38:95–109
Acknowledgements
This research was funded under University of Malaya Research Grant (UMRG: RG353-15AFR) and Postgraduate Research Grant (PG195-2016A).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
YAA, TM, SM and AFM declare that there is no conflict of interest.
Ethical approval
The article does not contain any studies with human participants performed by any of the authors.
Additional information
Data accessibility: The complete genome sequences are deposited at DDBJ/EMBL/GenBank under the accession number BDCV01000000.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Abdelhafiz, Y.A., Manaharan, T., Mohamad, S.B. et al. Whole genome sequencing and functional features of UMX-103: a new Bacillus strain with biosurfactant producing capability. Genes Genom 39, 877–886 (2017). https://doi.org/10.1007/s13258-017-0550-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-017-0550-7