Abstract
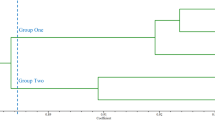
Although lily is the second largest flower crop in cutting flower commodity, only six simple sequence repeats SSRs have been reported. Thus, we developed expressed sequence tag derived-SSRs (EST-SSRs) for the Lilium genus. Among 2,235 unique ESTs, 754 ESTs contained SSR motifs, among which 165 ESTs were amenable to primer design. Among these 165 EST-SSRs, 131 EST-SSRs showed amplification in at least one Lilium species, and 76 EST-SSRs showed amplification in at least nine species. Of the 76 EST-SSRs, 47 showed amplification in all Lilium species analyzed. Using 10 breeding lines, we selected 21 EST-SSRs that had the highest number of alleles and polymorphism information content. The polymorphism information content values of these selected EST-SSRs ranged from 0.49 to 0.94 with an average of 0.76, which are higher than other plant species. The phylogenetic dendrogram derived from the amplification profiles of the 21 high polymorphic EST-SSRs was congruent with the genetic background of the 84 selected lily accessions and hybrids, which are available in commerce. Thus, the developed EST-SSRs will be very useful in germplasm management, genetic diversity analysis, cultivar finger printing, and molecular breeding in the lily.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Abe, M. Nakano, A. Nakatsuka, M. Nakayama, M. Koshioka and M. Yamagishi, Genetic analysis of floral anthocyanin pigmentation traits in Asiatic hybrid lily using molecular linkage maps. Theor. Appl. Genet., 105 (2002), pp. 1175–1182.
Asano Y (1989) Lilium L. In: Tsukumoto Y (ed), The Grand Dictionary of Horticulture, Vol. 5. Sykakukum, Tokyo, pp 198–209 (in Japanese).
Babara-Gonzalez R, Lim K-B, Zhou S, Ramanna MS, and Van Tuyl J (2008) Interspecific hybridization in lily: The use of 2n gametes in interspecific lily hybrids. In Floriculture, ornamental and plant biotechnology, vol V. JAT da Silva (editor), Global Science Books, Japan, pp. 138–145.
Bretting PK and Widrlechner MP (1995) Genetic markers and plant genetic resources management. Plant Breed. Rev. 13: 11–86.
Comber HF (1949) A new classification of the genus Lilium. Lily Yearbook, Royal Hort Soc 13: 86–105
Cuc LM, Mace ES, Crouch JH, Quang VD, Long TD, and Varshney RK (2008) Isolation and characterization of novel microsatellite markers and their application for diversity assessment in cultivated groundnut (Arachis hypogea). BMC Plant Biol. 8:55, doi:10.1186/1471-2229-8-55.
Enoki H, Sato H, and Koinuma K (2002) SSR analysis of genetic diversity among maize inbred lines adapted to cold regions of Japan. Theor. Appl. Genet. 104: 1270–1277.
Gupta PK, Rustgi S, Sharma S, Singh R, Kumar N, Balyan HS (2003) Transferable EST-SSR markers for the study of polymorphism and genetic diversity in bread wheat. Mol. Genet. Genomics 270: 315–323.
Horning ME, Maloney SC, and Webster MS (2003) Isolation and characterization of variable microsatellite loci in Lilum philadelphicum (Liliaeae). Mol. Ecol. Notes 3: 412–413.
Huang Y, Niu B, Gao Y, Fu L, Li W (2010) CD-HIT Suite: a web server for clustering and comparing biological sequences. Bioinformatics. 2010 26:680–682.
Hwang T-Y, Sayama T, Takahashi M, Takada Y, Nakamoto Y, Funatsuki H, Hisano S, Sato S, Tabata S, Kono I, et al. (2009) High-density integrated linkage map based on SSR markers in soybean. DNA Res. 16: 213–225.
Ikinci N and Oberprieler C (2010) Genetic relatiobships among NE Turkish Lilium L. (Liliaceae) species based on a random amplified polymorphic DNA analysis. Plant Syst Evol 284:41–48.
Kim KY (2004) Developing one step program (SSR Manager) for rapid identification of clones with SSRs and primer designing. MS Thesis, Seoul National University, Seoul, Korea.
Liang X, Chen X, Hong Y, Liu H, Zhou G, Li S, and Guo B (2009) Utility of EST-derived SSR in cultivated peanut (Arachis hypogea L.) and Arachis wild species. BMC Plant Biol. 9:35.
Lim KB, Barba-Gonzalez R, Zhou S, Ramanna MS, and van Tuyl JM (2008) Interspecific hybridization in Lily (Lilium): Taxonomy and commercial aspects of using species hybrids in breeding. In Floriculture, ornamental and plant biotechnology, vol V. JAT da Silva (editor), Global Science Books, Japan, pp. 146–151.
Morgante M, Hanafey M, and Powell W (2002) Microsatellites are preferentially associated with nonrepetitive DNA in plant genomes. Nature Genet. 3: 194–200.
Moyib OK, Odunola OA, and Dixon AGO (2007) SSR markers reveal genetic variation between improved cassava cultivars and landraces within a collection of Nigerian cassava germplasm. Afric. J. Biotechnol. 8: 2666–2674.
Nishikawa T, Okazaki K, Uchino T, Arakawa K, and Nagamine T (1999) A molecular phylogeny of Lilium in the transcribed spacer region of nuclear ribosomal DNA. J. Mol. Evol. 49: 238–249.
Nishikawa T, Okazaki K, Arakawa K and Nagamine T (2001) Phylogenetic analysis of section Sinomartagon in Genus Lilium using sequences of the internal transcribed spacer region in nu clear ribosomal DNA. Breed. Sci. 51:39–46
Park Y-J, Lee JK, and Kim N-S (2009) Simple sequence repeat polymorphisms (SSRPs) for evaluation of molecular diversity and germplasm classification of minor crops. Molecules 14: 4546–4569; doi:10.3390/molecules14114546.
Peruzzi L, Leitch IJ, and Caparelli KF (2009) Chromosome diversity and evolution in Liliaceae. Ann. Bot. 103: 450–475.
Poncet V, Rondeau M, Tranchant C, Cayrel A, Hamon S, de Kochoko A, and Hamon P (2006). SSR mining in coffee tree EST databases: potential use of EST-SSRs as markers for the Coffea genus. Mol. Gen. Genomics 276: 436–449.
Roholf FJ (1989) NTSYS-pc Numerical Taxonomy and Multivariate Analysis System. Exter, New York, USA.
Selkoe KA and Toonen RJ (2006) Microsatellites for ecologists: A practical guide to using and evaluating microsatellite markers. Ecol. Lett. 9: 615–617.
Shahin A, Arens P, van Heusden AW, van der Linden G, van Kaauwen M, Khan N, Schouten HJ, van de Weg W, Visser RGF and van Tuyl JM (2011) Genetic mapping in Lilium: mapping of major genes and quantitative trait loci for several ornamental trait and disease resistances. Plant Breeding 130:372–382.
Shete S, Tiwari H, and Elston RC (2000) On estimating the heterozygosity and polymorphism information content value. Theor. Pop. Biol. 57: 265–271.
Smyth DR, Kongsuwan K, and Wisudharomn S (1989) A survey of C-banded patterns in chromosomes of Lilium (Liliacea). Pl. Syst. Evol. 163: 53–69.
Synge PM (1980) Lilies.-London: Batsford.
Temnykh S, DeClerck G, Lukashova A, Lipovich L, Cartinhour S, and McCouch S (2001) Computational and experimental analysis of microsatellites in rice (Oryza sativa L.): frequency, length variation, transposon associations, and genetic marker potential. Genome Res. 11:1441–1452.
Thomas RG (1960) Peliotropism and the developmental changes correlated with flowering. Genetica 31: 329–339.
Van Heusden AW, Jongerius MC, van Tuyl JM, staarhof TP and Mes JJ (2002) Molecular assisted breeding for disease resistance in lily Act Hortic 572:131–138.
Van Tienderen, PH, de Haan AA, van der Linden, and Vosman B (2002) Biodiversity assessment using markers for ecologically important traits. Trends Ecol. Evol. 17: 577–582.
Victoria FC, da Maia LC, and de Oliveira AC (2011) In silico comparative analysis of SSR markers in plants. BMC Plant Biol. 11:15.
Wang Z, Weber JL, Zhong G, and Tanksley SD (1994) Survey of plant short tandem DNA repeats. Theor. Appl. Genet. 88: 1–6.
White G and Powell W (1997) Cross-species amplification of SSR loci in Meliaceae family. Mol. Ecol. 6: 1195–1197.
Yamagushi M (1995) Detection of section-specific random amplified polymorphic DNA (RAPD) markers in Lilium. Theor. Appl. Genet. 91: 830–835.
Yamagushi M, Abe H, Nakano M, and Nakatsuka A (2002) PCR-based molecular markers in Asiatic hybrid lily. Scient. Hort. 96: 225–234.
Zheng S, Xiao G, Guo J, Fei Z, Xu Y, Roe BA, and Wang Y (2010) Development of a EST dataset and characterization of EST-SSRs in a trandiational Chinese medicinal plant, Epimedium sagittatum (Sieb. Et. Zucc.) Maxim. BMC Genomics 11: 94.
Author information
Authors and Affiliations
Corresponding authors
Additional information
S.-I. Lee, K.-C. Park, and Y.-S. Song contributed equally to this work.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Lee, SI., Park, KC., Song, YS. et al. Development of expressed sequence tag derived-simple sequence repeats in the genus Lilium . Genes Genom 33, 727–733 (2011). https://doi.org/10.1007/s13258-011-0203-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-011-0203-1