Abstract
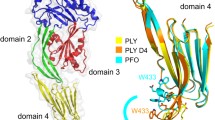
Cholesterol-dependent cytolysins (CDC) are pore forming toxins. A prototype of the CDC family members is perfringolysin O (PFO), which directly binds to the cell membrane enriched in cholesterol, causing cell lysis. However, an exception of this general observation is intermedilysin (ILY) of Streptococcus intermedius, which requires human CD59 as a receptor in addition to cholesterol for its hemolytic activity. A possible explanation of this functional difference is the conformational variation between the C-terminal domains of the two toxins, particularly in the highly conserved undecapeptide termed tryptophan rich motif. Here, we present the crystal structure of suilysin, a CDC toxin from the infectious swine pathogen Streptococcus suis. Like PFO, suilysin does not require a host receptor for hemolytic activity; yet the crystal structure of suilysin exhibits a similar conformation in the tryptophan rich motif to ILY. This observation suggests that the current view of the structure-function relationship between CDC proteins and membrane association is far from complete.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Adams, P.D., Grosse-Kunstleve, R.W., Hung, L.W., Ioerger, T.R., McCoy, A.J., Moriarty, N.W., Read, R.J., Sacchettini, J.C., Sauter, N.K., and Terwilliger, T.C. (2002). PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr D Biol Crystallogr 58, 1948–1954.
Bahadur, R.P., Chakrabarti, P., Rodier, F., and Janin, J. (2004). A dissection of specific and non-specific protein-protein interfaces. J Mol Biol 336, 943–955.
Bailey, S. (1994). The CCP4 suite: programs for protein crystallography. Acta Crystallographica D50, 760–763.
Bork, P., Holm, L., and Sander, C. (1994). The immunoglobulin fold. Structural classification, sequence patterns and common core. J Mol Biol 242, 309–320.
Bourdeau, R.W., Malito, E., Chenal, A., Bishop, B.L., Musch, M.W., Villereal, M.L., Chang, E.B., Mosser, E.M., Rest, R.F., and Tang, W. J. (2009). Cellular functions and X-ray structure of anthrolysin O, a cholesterol-dependent cytolysin secreted by Bacillus anthracis. J Biol Chem.
Davis, I.W., Leaver-Fay, A., Chen, V.B., Block, J.N., Kapral, G.J., Wang, X., Murray, L.W., Arendall, W.B., 3rd, Snoeyink, J., Richardson, J.S., et al. (2007). MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res 35, W375–383.
DeLano, W.L. (2002). The PyMOL Molecular Graphics System. DeLano Scientific, Palo Alto, CA, USA.
Emsley, P., and Cowtan, K. (2004). Coot: model-building tools for molecular graphics. Acta Crystallogr D Biol Crystallogr 60, 2126–2132.
French G.S. and Wilson K.S. (1978). On the treatment of negative intensity observations. Acta Cryst A34, 517–525.
Giddings, K.S., Johnson, A.E., and Tweten, R.K. (2003). Redefining cholesterol’s role in the mechanism of the cholesterol-dependent cytolysins. Proc Natl Acad Sci U S A 100, 11315–11320.
Giddings, K.S., Zhao, J., Sims, P.J., and Tweten, R.K. (2004). Human CD59 is a receptor for the cholesterol-dependent cytolysin intermedilysin. Nat Struct Mol Biol 11, 1173–1178.
Gottschalk, M., and Segura, M. (2000). The pathogenesis of the meningitis caused by Streptococcus suis: the unresolved questions. Vet Microbiol 76, 259–272.
Jacobs, A.A., Loeffen, P.L., van den Berg, A.J., and Storm, P.K. (1994). Identification, purification, and characterization of a thiolactivated hemolysin (suilysin) of Streptococcus suis. Infect Immun 62, 1742–1748.
Laskowski, R.A., MacArthur, M.W., Moss, D.S., and Thornton, J.M. (1993). PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Crystallogr 26, 283–291.
Matthews, B.W. (1968). Solvent contents of protein crystals. J Mol Biol 33, 491–497.
Nagamune, H., Ohkura, K., Sukeno, A., Cowan, G., Mitchell, T.J., Ito, W., Ohnishi, O., Hattori, K., Yamato, M., Hirota, K., et al. (2004). The human-specific action of intermedilysin, a homolog of streptolysin O, is dictated by domain 4 of the protein. Microbiol Immunol 48, 677–692.
Nagamune, H., Ohnishi, C., Katsuura, A., Fushitani, K., Whiley, R.A., Tsuji, A., and Matsuda, Y. (1996). Intermedilysin, a novel cytotoxin specific for human cells secreted by Streptococcus intermedius UNS46 isolated from a human liver abscess. Infect Immun 64, 3093–3100.
Otwinowski, Z., and Minor, W. (1997). Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol 276, 307–326.
Painter, J., and Merritt, E.A. (2006). Optimal description of a protein structure in terms of multiple groups undergoing TLS motion. Acta Crystallogr D Biol Crystallogr 62, 439–450.
Polekhina, G., Giddings, K.S., Tweten, R.K., and Parker, M.W. (2005). Insights into the action of the superfamily of cholesterol-dependent cytolysins from studies of intermedilysin. Proc Natl Acad Sci U S A 102, 600–605.
Ponting, C.P. (1999). Chlamydial homologues of the MACPF (MAC/perforin) domain. Curr Biol 9, R911–913.
Ramachandran, R., Tweten, R.K., and Johnson, A.E. (2004). Membrane-dependent conformational changes initiate cholesteroldependent cytolysin oligomerization and intersubunit beta-strand alignment. Nat Struct Mol Biol 11, 697–705.
Rosado, C.J., Buckle, A.M., Law, R.H., Butcher, R.E., Kan, W.T., Bird, C.H., Ung, K., Browne, K.A., Baran, K., Bashtannyk-Puhalovich, T. A., et al. (2007). A common fold mediates vertebrate defense and bacterial attack. Science 317, 1548–1551.
Rossjohn, J., Feil, S.C., McKinstry, W.J., Tweten, R.K., and Parker, M. W. (1997). Structure of a cholesterol-binding, thiol-activated cytolysin and a model of its membrane form. Cell 89, 685–692.
Rossjohn, J., Polekhina, G., Feil, S.C., Morton, C.J., Tweten, R.K., and Parker, M.W. (2007). Structures of perfringolysin O suggest a pathway for activation of cholesterol-dependent cytolysins. J Mol Biol 367, 1227–1236.
Schneider, T.R., and Sheldrick, G.M. (2002). Substructure solution with SHELXD. Acta Crystallogr D Biol Crystallogr 58, 1772–1779.
Shatursky, O., Heuck, A.P., Shepard, L.A., Rossjohn, J., Parker, M. W., Johnson, A.E., and Tweten, R.K. (1999). The mechanism of membrane insertion for a cholesterol-dependent cytolysin: a novel paradigm for pore-forming toxins. Cell 99, 293–299.
Shepard, L.A., Heuck, A.P., Hamman, B.D., Rossjohn, J., Parker, M. W., Ryan, K.R., Johnson, A.E., and Tweten, R.K. (1998). Identification of a membrane-spanning domain of the thiol-activated pore-forming toxin Clostridium perfringens perfringolysin O: an alpha-helical to beta-sheet transition identified by fluorescence spectroscopy. Biochemistry 37, 14563–14574.
Soltani, C.E., Hotze, E.M., Johnson, A.E., and Tweten, R.K. (2007a). Specific protein-membrane contacts are required for prepore and pore assembly by a cholesterol-dependent cytolysin. J Biol Chem 282, 15709–15716.
Soltani, C.E., Hotze, E.M., Johnson, A.E., and Tweten, R.K. (2007b). Structural elements of the cholesterol-dependent cytolysins that are responsible for their cholesterol-sensitive membrane interactions. Proc Natl Acad Sci U S A 104, 20226–20231.
Thompson, J.D., Gibson, T.J., Plewniak, F., Jeanmougin, F., and Higgins, D.G. (1997). The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25, 4876–4882.
Tilley, S.J., Orlova, E.V., Gilbert, R.J., Andrew, P.W., and Saibil, H.R. (2005). Structural basis of pore formation by the bacterial toxin pneumolysin. Cell 121, 247–256.
Tweten, R.K. (2005). Cholesterol-dependent cytolysins, a family of versatile pore-forming toxins. Infect Immun 73, 6199–6209.
Tweten, R.K., Parker, M.W., and Johnson, A.E. (2001). The cholesterol-dependent cytolysins. Curr Top Microbiol Immunol 257, 15–33.
Zhang, X., and Matthews, B.W. (1995). EDPDB: A multifunctional tool for protein structure analysis. J Appl Crystallogr 28, 624–630.
Author information
Authors and Affiliations
Corresponding authors
Additional information
These authors contributed equally to this work.
Rights and permissions
About this article
Cite this article
Xu, L., Huang, B., Du, H. et al. Crystal structure of cytotoxin protein suilysin from Streptococcus suis. Protein Cell 1, 96–105 (2010). https://doi.org/10.1007/s13238-010-0012-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13238-010-0012-3