Abstract
Isolation of high-quality RNA from Dendrobium flowers is challenging because of the high levels of pigment, polysaccharides, and polyphenols. In the present study, an efficient CTAB method for RNA extraction from the pigment-rich flowers of Dendrobium was optimised. The optimised method yielded high quantities of RNA (10.1–12.9 µg/g). Spectrophotometric values of A260/280 in the range of 2.2 to 2.4 and A260/230 values of 2.0 suggested that the isolated RNA was free of polyphenols, polysaccharides, and protein contaminants. RNA integrity numbers determined by microfluidics were in the range of 7.9–8.9 indicative of intact RNA. In the improved method, the addition of 3 M NaCl and 3% PVP-10 in the extraction buffer, followed by an incubation period of 45 min at 65 °C, eliminated most of the polysaccharides, polyphenolic compounds, and denatured protein. Extraction with phenol:chloroform:isoamyl alcohol (125:24:1) effectively removed pigments from the aqueous phase, while the precipitation of RNA with lithium chloride minimised the co-precipitation of protein, DNA, and polysaccharide and resulted in the extraction of high quality of RNA. The suitability of the RNA for downstream processing was confirmed via RT-PCR amplification of Chalcone synthase gene from cDNA prepared from RNA isolated from different developmental stages of the flower of a Dendrobium hybrid. The present method will be highly useful for the isolation of RNA from pigment, polyphenol, and polysaccharide-rich plant tissues.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The floriculture industry is an important contributor to agribusiness with the multiple benefits of meeting the aesthetic needs of consumers, generating employment and income and facilitating foreign exchange (Harisha 2017). Flowers are generally traded both as cut flowers and potted plants and orchids account for around 10% of the global share in floriculture trade (De et al. 2014). The average trade value of orchids in 2018 was reported as US$400 million (https://www.nationthailand.com/breakingnews/30360623). The high trade value of orchids is because of the diverse and exotic colours, shapes, year-round availability, long flowering life as potted plants and long vase life as cut flowers (De and Medhi 2015). Among the orchids, Dendrobium species and hybrids are highly popular both as potted plants (Sarmah et al. 2017) and as cut flowers (Hinsley et al. 2017). Around 70–80% of the over 1200 species of Dendrobium are tropical, growing in diverse habitats throughout Asia, including Malaysia, China, Japan, India, Philippines, Indonesia, Australia, New Guinea, Vietnam, several islands of the Pacific and also in subtropical regions of Africa, and Latin America (Govaerts 2003).
Despite the wide natural diversity of orchid flowers, new varieties and especially hybrids with novel colours, shapes, fragrance, and with long vase life are in high demand in the floral industry (Lau et al. 2015). To support this, there is a need to improve knowledge underlying the molecular genetics of orchid floral development. However, obtaining high-quality RNA from floral tissues is a crucial step that can be challenging, especially where the tissue has a very high flavonoid, pigment, polyphenolic, and polysaccharide content (Gao et al. 2016). As plant tissues vary in chemical composition, the efficiency of any RNA isolation method for a particular plant or tissue will differ, and this requires method testing and optimisation (Liu et al. 2018). A number of RNA isolation methods have been standardized including methods for secondary metabolite-rich tissues (Kiefer et al. 2000), flower of black gram (Raizada and Jegadeesan 2019), leaf and bark of coffee (Huded et al. 2018), leaf and stem of Stevia (Leh et al. 2019), and flower of oil palm (Qadri et al. 2019). These methods use either CTAB/SDS and sodium acetate or isopropanol for precipitation of RNA and have also been found to be effective for the extraction of RNA from orchids with white flowers (Phalaenopsis aphrodite; Su et al. 2011), pink (Paeonia suffruticosa; Gao et al. 2016), or light yellow (Dendrobium huoshanense; Liu et al. 2018) flowers. However, we found the previously reported protocols to result in the extraction of low yields of RNA, contaminated with pigment when applied to flowers of a popular commercial Dendrobium hybrid with vibrant purple flower petals and sepals. For our own research use, and with an aim to develop an effective RNA isolation method for pigment-rich floral tissues, we have developed a simple, economical, and robust RNA isolation method for pigment-rich floral tissues of Dendrobium hybrids, which can successfully yield high-quality RNA from flowers rich in pigment, polysaccharides, and polyphenols.
Materials and methods
Plant material
Flowers from four different Dendrobium hybrids were used in this study. Mature plants of Dendrobium Burmese Ruby × Dendrobium Mae-Klong River (magenta flower) were obtained from Cheah Wah Sang Orchid Farm in Shah Alam, Selangor, Malaysia. Hybrids Dendrobium Burana Jade × (Dendrobium Bertha Chong × Dendrobium Imelda Romualdez) (burgundy flower), Dendrobium Trudy Brandt × Dendrobium Udom Blue Angel (blue–violet flower) and Dendrobium Aridang × Dendrobium Burana Sundae (red–violet flower) were obtained from Lum Chin Orchid Garden Sdn. Bhd, Kuala Lumpur, Malaysia. The plants were maintained at the GP-BSL2 greenhouse (Plant Biotech Facility), University of Malaya, Malaysia under natural light (12 h photoperiod) at a controlled temperature (25 ± 2 °C) and relative humidity (65 ± 5%). Floral buds of Dendrobium Burmese Ruby × Dendrobium Mae-Klong River were collected at five different developmental stages according to Lau et al. (2015) (Stage 1: 0–0.5 cm; Stage 2: 0.51–1.0 cm; Stage 3: 1.1–2.0 cm; Stage 4: 2.1–3.0 cm; and Stage 5: 3.1–4.0 cm). Mature flowers (stage 6) were collected from each of the four Dendrobium hybrids (Fig. 1a–d). Floral tissues of Dendrobium hybrids were rapidly washed with tap water, sterilized with 70% ethanol for 30 s, rinsed thrice with sterile water, blotted dry, and snap-frozen in liquid nitrogen. The frozen samples were stored at – 80 °C until further use.
Flowers of Dendrobium hybrids. a Dendrobium Burmese ruby × Dendrobium Mae-klong River (bar = 1 cm); b Dendrobium Burana Jade × [Dendrobium Bertha Chong × Dendrobium Imelda Romualdez] (scale bar = 1 cm); c Dendrobium Trudy Brandt × Dendrobium Udom Blue Angel) (scale bar = 1 cm); and d Dendrobium Aridang × Dendrobium Burana Sundae (scale bar = 1 cm)
RNA isolation
Five different methods were used in the study including four previously published methods; Kiefer et al. (2000) (Method 1); Su et al. (2011) (Method 2); Gao et al. (2016) (Method 3) and Liu et al. (2018) (Method 4) and an improved CTAB method (Supplementary Table S1, Fig. 2). Plasticware and pipette tips were soaked overnight in 0.01% DEPC-treated distilled water and autoclaved at 121 °C at 15 psi of pressure for a period of 45 min. All chemicals used were of molecular biology grade and the reagents were prepared with 0.01% DEPC-treated distilled water. The workbench was sprayed with RNase Destroyer (Favorgen, Taiwan) spray. The gel electrophoresis accessories were also sprayed with RNase Destroyer spray followed by rinsing with DEPC-treated water before use to prevent contamination from RNase. RNA was isolated from 500 mg FW mature flower starting material for the four different Dendrobium hybrids and from five flower developmental stages of Dendrobium Burmese Ruby × Dendrobium Mae-Klong River. RNA isolated from mature flowers of Dendrobium Burmese Ruby × Dendrobium Mae-Klong River was used to systematically compare the efficiency of the improved method with the four previously published methods (Supplementary Table S1).
For the method reported as new in the current paper (improved CTAB method, Fig. 2; Supplementary Table S1) tissues were ground into a fine powder using liquid nitrogen in a pre-chilled mortar and pestle. To the finely powdered sample, 12 ml of freshly prepared pre-warmed modified CTAB extraction buffer was added prior to transfer into a 15 ml centrifuge tube. The buffer contained 2% CTAB, 100 mM Tris-HCl (pH 8.0), 3 M NaCl, 25 mM EDTA (pH 8.0), and 3% PVP-10. The mixture was vortexed and incubated at 65 °C for periods of 5, 10, 15, or 45 min to determine the most suitable incubation time. Each sample mixture was then allowed to cool at room temperature for 2 min. The mixture was centrifuged for 5 min at 7000×g, and then, an equal volume of phenol:chloroform:isoamyl alcohol (P:C:I) (125:24:1, pH 4.5) was added to the supernatant and the mixture was vortexed for 3 min. Following that, the tubes were centrifuged at 17,500×g at 4 °C for 15 min and the aqueous (upper) fraction was collected carefully without disturbing the debris (flocculent material containing protein and lysed cell materials) at the interface layer. The aqueous fraction was transferred into a new 2.0 ml microcentrifuge tube and an equal volume of chloroform and isoamyl alcohol (C:I) at a ratio of 24:1 was added, mixed gently by inverting the tube several times and then centrifuged at 17,500×g at 4 °C for 15 min. The resultant aqueous fraction was collected and transferred to fresh tubes. A volume of 8 M LiCl (giving a final concentration of 0.37 M in solution with the sample) representing a third of the recovered supernatant volume was added and mixed well by inverting the tubes. The tubes were then incubated at − 20 °C for 3, 6, 8, 10, 12, or 24 h to determine the optimal precipitation time. Following the incubation, the tubes were thawed on ice and then centrifuged at 17,500×g at 4 °C for 15 min. The supernatant was discarded and the pellet was washed twice with 70% chilled ethanol. The washed RNA pellet was air-dried and resuspended in 25 µl of RNase-free water (Qiagen, Germany). To eliminate any residual genomic DNA from the samples, the extracted samples were treated with DNase I (Qiagen, Germany) according to the manufacturer’s instruction. Following the DNase treatment, the volume of the sample was increased to 500 µl using RNase-free water and incubated for 5 min at room temperature. Then, 1/3 volume of 8 M LiCl was added to the samples and incubated overnight at − 20 °C. Following that, the samples were centrifuged at 4 °C for 30 min at 17,500×g and the supernatant was discarded. The pellet was washed twice with 70% chilled ethanol by centrifugation at 17,500×g at 4 °C for 15 min. Following that, the supernatant was discarded and the RNA pellets were air-dried for 5 min and resuspended in 25 µl of RNase-free water.
Quantification and quality assessment of RNA
The purity and concentration of RNA samples were assessed by Nanophotometer Perl® (Implen, Germany) at 230, 260, and 280 nm absorbance. The quality and integrity of RNA were examined after separation by denaturing agarose gel electrophoresis. The 1% denaturing gel was prepared by melting agarose to MOPS gel buffer [200 mM 3-(N-morpholino) propanesulfonic acid, 50 mM sodium acetate and 10 Mm EDTA, pH 7]. The mixture was cooled to 60−65 °C, and then, 1 ml of 37% formaldehyde and 1 µl ethidium bromide (0.5 μg/ml) were added before pouring onto gel support. The gel was run for around 45 min at 90 V. The bands were visualized and photographed using a Gel documentation system (BioRad, US). The quality of the extracted RNA was analysed with a microfluidics system using the Agilent RNA 6000 Nano LabChip Kit and the Agilent 2100 Bioanalyzer (Agilent Technologies).
cDNA synthesis and PCR analysis
cDNAs were synthesised using 1 µg total RNA using SuperScript® III First-Strand Synthesis System (Invitrogen, USA) (Supplementary Fig. S1). PCR was carried out with Chalcone synthase (CHS) gene-specific primers (Supplementary Fig. S2) using cDNA as a template. The forward (5′GCCCAAATCTCGCATAACTC3′) and reverse (5′TACGCGAGATGGGACTAACC3′) primer of CHS were designed to amplify a 436 bp cDNA fragment based on the GenBank sequence KC345011.1 (Dendrobium hybrid cultivar Sonia Earsakul). PCR reactions were carried out in 25 µl reaction volumes containing 12.5 µl of 2 × GoTaq® Green Master Mix, 1.0 µl cDNA and 0.4 µM of each primer using a Mastercycler Gradient DNA Thermocycler (Eppendorf, Germany). The thermoprofile was as follows: initial denaturation at 95 °C for 3 min, 35 cycles of 94 °C for 30 s, 53 °C for 30 s, and 72 °C for 1 min followed by the final extension for 10 min at 72 °C. The amplified PCR products were separated on a 1.0% agarose gel, stained with ethidium bromide (0.5 μg/ml) and visualized with a gel documentation system (BioRad, USA).
Results and discussion
Dendrobium flowers contain high levels of protein, carbohydrate and secondary metabolites including flavonoids, pigments, and polyphenols (Moretti et al. 2013). Obtaining pure RNA with high yield from such tissues is a cumbersome process as polysaccharides and polyphenols have similar chemical properties to RNA and tend to co-precipitate with RNA (Asif et al. 2006; Shu et al. 2014). The co-precipitation of these compounds with RNA reduces yield and increases the possibility of rapid degradation which makes the sample unsuitable for further downstream processing. In the current study, we have optimised an efficient RNA isolation method for mature flowers of Dendrobium hybrids. Four previously published methods for secondary metabolite-rich tissues; three CTAB-based RNA isolation methods (Methods 1, 2, and 4) and one SDS-based method (method 3) were initially tested (Supplementary Table S1). Method 1 was reported for RNA isolation from tissues including flowers, rich in polyphenols and polysaccharides (Kiefer et al. 2000), and Method 2 was reported for Phalaenopsis orchids (Su et al. 2011). Method 3, was a combination of SDS and TRIzol reagent (guanidinium thiocyanate–phenol–chloroform), reported for the buds of the tree peony (Paeonia suffruticosa Andr) (Gao et al. 2016) and Method 4 was reported for a pale yellow orchid Dendrobium huoshanense (Liu et al. 2018). However, we found none of the four methods to be adequate for the extraction of high-quality RNA from the pigment, polyphenol, and polysaccharide-rich tissue of Dendrobium hybrid flowers. We chose mature flowers of Dendrobium Burmese Ruby × Dendrobium Mae-Klong River (Fig. 1a) as the sample tissue for optimisation studies, as the mature flower tissue contains the highest level of pigments, and so presents the most challenging tissue for molecular genetic studies using floral RNA (Rodríguez-Ávila et al. 2011).
While performing the isolation of RNA from mature flowers, we noticed that one of the main hurdles was the viscous tissue homogenate that arises during the mixing of ground tissue with the extraction buffer. The high viscosity made both the mixing of the ground sample with buffer and the recovery of the RNA-containing supernatant after centrifugation inefficient. The optimal sample to buffer ratio for nucleic acid extraction has been reported to vary with the species and tissue types with the ideal ratio resulting in a somewhat viscous, but well-dispersed homogenate (Murray and Thompson 1980). By increasing the sample–buffer ratio to almost double that of the other tested methods (Supplementary Table S1), we observed more efficient mixing of the sample with buffer and better recovery of the supernatant containing the RNA (Supplementary Table S2).
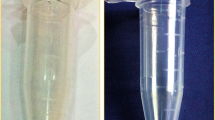
Another hurdle was the removal of pigments from the strongly coloured flower tissues. Method 1, Method 2, and Method 4 failed to remove pigment in the extraction step, leaving the aqueous phase dark purple (Supplementary Fig. S3). Method 3, which is a combination of SDS and TRIzol reagent, and also includes a phenol:chloroform:isoamyl alcohol extraction step, showed better removal of pigments compared to other methods. The improved removal of pigment was likely because of the presence of phenol in the TRIzol reagent (Chomczynski and Sacchi 1987) in addition to the high phenol proportion in the P:C:I extraction included in this method. Methods 1, 2, and 4 lack the use of phenol in the solvent extraction steps (Supplementary Figs. S3, S4). However, the concentration of phenol used in Method 3 was found to be insufficient to obtain a colourless pellet for Dendrobium hybrid flower tissue (Supplementary Fig. S4). To further improve the separation of RNA from pigments, as well as other superfluous cell materials, we added a P:C:I (125:24:1) purification step prior to the two rounds of C:I extraction, in which the proportion of phenol was relatively high. Phenol together with chloroform is widely reported for extraction of high-quality RNA (Li 2015; Lee et al. 2015; Toni et al. 2018). The addition of the P:C:I (125:24:1) extraction step in our improved CTAB method resulted in the comparatively lighter pigmented aqueous phase after P:C:I extraction (Supplementary Fig. S3). Samples also showed A260/230 ratios within the desired range (Supplementary Table S2) and colourless pellets (Supplementary Fig. S4).
The insoluble RNA pellet obtained from flower tissues using methods 1, 3, and 4 was another hurdle to recover high-quality RNA. The gel-like pellets, with poor solubility, indicated the inefficient separation of contaminants from RNA (Muoki et al. 2012). RNA samples obtained from Method 1 showed A260/230 ratios in the range of 1.6–1.7, and samples from Method 2 gave a value of 1.8, which indicated the presence of polyphenol and/or polysaccharide contamination. Similarly, RNA prepared from Dendrobium hybrid tissue with Method 3 also showed low absorbance ratios, suggesting contamination with protein (ratio of A260/280; 1.6–1.7) and with polyphenol and/or polysaccharides (ratio A 260/230; 1.6–1.7) (Fig. 3; Supplementary Table S2). To address this, we modified the buffer to have higher concentrations of NaCl (3 M), PVP-10 (3% w/v), and ß-mercaptoethanol (3% v/v) (Supplementary Table S1). The higher concentration of NaCl in the extraction buffer was to promote salting-out of the protein while leaving the RNA in solution (El-Ashram et al. 2016). In addition, a higher NaCl content aids in avoiding co-precipitation of polysaccharides with RNA (Fang et al. 1992). The PVP-10 concentration was increased to 3% to facilitate crosslinking with polyphenols and to help in precipitation from the aqueous solution (Carpenter et al. 1976). The relatively high concentration of β-mercaptoethanol in the modified extraction buffer was also aimed at removing polyphenol compounds (Wong et al. 2014) and to inactivate ribonucleases released during cell lysis (Lehninger et al. 2005). We also tested different incubation periods in the extraction buffer to maximise RNA yield. The relatively short incubation times of 5–15 min in extraction buffer used in methods 1–4, resulted in lower RNA recovery along with protein contamination from mature flowers of Dendrobium hybrids (Supplementary Table S3). Hence, we tested longer incubation times of 10, 15, and 45 min to facilitate the salting-out of proteins and separation of RNA from cell debris. An incubation time of 45 min was found to result in a higher yield and good quality of RNA (Supplementary Table S3). Another contributing factor for low pellet solubility might be the use of isopropanol in Methods 1 and 3, because isopropanol promotes the co-precipitation of salts (Choi et al. 2018). It is also difficult to completely remove isopropanol from samples because of low volatility, compared to ethanol, which further deteriorates the quality of RNA (Surzycki 2012). Method 2, which used 8 M LiCl (giving a final concentration of 0.37 M in solution with the sample) for nucleic acid precipitation, was the best among the four methods in terms of pellet solubility. Method 4, also used LiCl but at 10 M (giving a much lower final concentration of 0.04M in solution with the sample). In our modified method, precipitation with 8 M LiCl followed by 75% ethanol wash resulted in soluble pellet allowing good recovery of RNA in the final aqueous solution (Supplementary Table S2). An advantage of using LiCl for RNA recovery is that it does not efficiently co-precipitate protein, DNA, polysaccharide, or salts (Barman et al. 2017). We also tested different LiCl precipitation times to maximise RNA yield. We observed that overnight precipitation (24 h) improved RNA yield compared to the yield after shorter precipitation times of 3–12 h (Supplementary Table S4).
All five RNA isolation methods showed clear and intact bands of 28S rRNA and 18S rRNA from mature flowers of Dendrobium Burmese Ruby × Dendrobium Mae-Klong River, without visible chromosomal DNA contamination (Fig. 4). However, the yields of RNA obtained using each method varied widely: the improved CTAB method resulted in high yields of RNA, in the range of 11–12 µg/g FW (Supplementary Table S2) demonstrating the efficiency of the improved method. Extraction of RNA from mature flowers of four different Dendrobium hybrids with the optimised method resulted in high yields (10.1–12.9 µg/g FW; Table 1, Supplementary Fig. S5) and significantly higher quality of RNA (Fig. 3), demonstrating the efficiency and reproducibility of the improved method.
Agarose gel electrophoresis of total RNA isolated from the mature flower of Dendrobium hybrid. Lanes 1 and 17: 1 kb DNA Ladder (Promega, USA); Lanes 2–4: RNA isolated using Method 1(Kiefer et al. 2000), Lanes 5–7: RNA isolated using Method 2 (Su et al. 2011), Lanes 8–10: RNA isolated using Method 3 (Gao et al. 2016), Lanes 11–13: RNA isolated using Method 4 (Liu et al. 2018), Lanes 14–16: RNA isolated using improved CTAB method
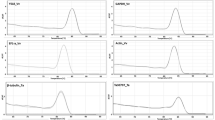
The RNA samples using the improved CTAB method had clear ribosomal peaks and higher RNA Integrity Number (RIN) values (7.9–8.9) than those using the methods 1–4 (4.8–6.8) confirming the suitability of the improved method to isolate RNA from pigment, polyphenol, and polysaccharide-rich tissue (Fig. 5; Table 1). An RIN of ≥ 7 indicates that the RNA is suitable for high stringency applications (Schroeder et al. 2006) such as cDNA library construction or next-generation sequencing (Deepa et al. 2014). To determine the reproducibility of our improved CTAB method for isolating RNA from floral tissues of different developmental stages, we extracted RNA from stage 1–5 flower buds of the Dendrobium Burmese Ruby × Dendrobium Mae-Klong River in addition to the mature flowers. We then tested the integrity of the isolated RNA using RT-PCR of an important orchid floral pigment biosynthetic cDNA, chalcone synthase (CHS). Amplification of a partial coding sequence of CHS, produced a band of the expected size of 436 bp, using cDNA synthesised from RNA extracted from flower buds and mature flowers, demonstrating the suitability of this method to isolate RNA suitable for downstream processes across floral developmental stages (Fig. 6).
Agarose gel electrophoresis of PCR product amplified with Chalcone synthase-specific primers. Lanes 1 and 9: 1 kb DNA Ladder (Promega, USA); Lanes 2–8 are PCR products after cDNA synthesis from RNA isolated from flower buds of six different developmental stages of Dendrobium hybrid flowers. Lane 2: RNA isolated from stage 1; Lane 3: RNA isolated from stage 2; Lane 4: RNA isolated from stage 3; Lane 5: RNA isolated from stage 4; Lane 6: RNA isolated from stage 5; Lane 7: RNA isolated from stage 6 sepals; Lane 8: RNA isolated from stage 6 petals; Lane 10: negative control (no template)
Conclusions
An improved CTAB-based method of RNA isolation from the pigment, polyphenol, and polysaccharide-rich tissues was developed and tested using flowers from four different pigment-rich Dendrobium hybrids. We demonstrated that RNA of high quality and yield was isolated from six different developmental stages of Dendrobium hybrid flowers, from immature bud to fully open flower and that the RNA was suitable for reverse transcription and qPCR amplification of a partial coding sequence of chalcone synthase. This protocol is an easy, efficient, and reproducible method for RNA isolation from flowers rich in secondary metabolites, and it may result in improved RNA extraction from other highly pigmented flowers.
Abbreviations
- bp:
-
Base pair
- cDNA:
-
Complimentary DNA
- CHS:
-
Chalcone synthase
- C:I:
-
Chloroform:isoamyl alcohol
- P:C:I:
-
Phenol:chloroform:isoamyl alcohol
- CTAB:
-
Cetyltrimethylammonium bromide
- °C:
-
Degrees Celsius
- cm:
-
Centimetre
- DEPC:
-
Diethyl pyrocarbonate
- DNase:
-
Deoxyribonuclease
- EDTA:
-
Ethylenediaminetetraacetic acid
- EtBr:
-
Ethidium bromide
- FW:
-
Fresh weight
- LiCl:
-
Lithium chloride
- M:
-
Molarity
- ml:
-
Millilitre
- mM:
-
Millimolar
- min:
-
Minutes
- NaCl:
-
Sodium chloride
- psi:
-
Pound-force per square inch
- PVP-10:
-
Polyvinylpyrrolidone-10
- RNase:
-
Ribonuclease
- rRNA:
-
Ribosomal RNA
- RT-PCR:
-
Reverse transcription polymerase chain reaction
- s:
-
Second
- SDS:
-
Sodium dodecyl sulfate
- TBE:
-
Tris-borate-EDTA
- Tris–HCl:
-
Tris hydrochloride
- µl:
-
Microlitre
- ng:
-
Nanogram
- w/v:
-
Weight per volume
- v/v:
-
Volume per volume
- %:
-
Percentage
References
Asif M, Trivedi P, Solomos T, Tucker M (2006) Isolation of high-quality RNA from apple (Malus domestica) fruit. J Agric Food Chem 54:5227–5229. https://doi.org/10.1021/jf053137n
Barman P, Choudhary AK, Geeta R (2017) A modified protocol yields high-quality RNA from highly mucilaginous Dioscorea tubers. 3 Biotech 7:150. https://doi.org/10.1007/s13205-017-0775-9
Carpenter A, Siggia S, Carter S (1976) Separation and/or concentration of phenolic materials from dilute aqueous solutions. Anal Chem 48:225–228
Choi C, Yoon S, Moon H, Bae YU, Kim CB, Diskul-Na-Ayudthaya P, Ngu TV, Munir J, Han J, Park SB, Moon JS (2018) mirRICH, a simple method to enrich the small RNA fraction from over-dried RNA pellets. RNA Biol 28:1
Chomczynski P, Sacchi N (1987) Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal Biochem 162:156–159
De LC, Medhi RP (2015) Orchid–a diversified component of farming systems for profitability and livelihood security of small and marginal farmers. J Glob Biosci 4:1393–1406
De LC, Pathak P, Rao AN, Rajeevan PK (2014) Commercial orchids National Research Centre for Orchids, India
Deepa K, Sheeja TE, Santhi R, Sasikumar B, Cyriac A, Deepesh PV, Prasath D (2014) A simple and efficient protocol for isolation of high quality functional RNA from different tissues of turmeric (Curcuma longa L.). Physiol Mol Biol Plants 20:263–267. https://doi.org/10.1007/s12298-013-0218-y
El-Ashram S, Al Nasr I, Suo X (2016) Nucleic acid protocols: extraction and optimization. Biotechnol Rep 12:33–39. https://doi.org/10.1016/j.btre.2016.10.001
Fang G, Hammar S, Grumet R (1992) A quick and inexpensive method for removing polysaccharides from plant genomic DNA. Biotechniques 13(52–54):56
Gao Y, Guangqi Z, Jiang C, Yao S, Kang YE, Shucheng F (2016) Comparison of Different Methods for RNA Extraction from Floral Buds of Tree Peony (Paeonia suffruticosa Andr.). Notulae Botanicae Horti Agrobotanici Cluj-Napoca 44:418–422
Govaerts R (2003) World checklist of monocotyledons database in ACCESS: 1-71827 The Board of Trustees of the Royal Botanic Gardens, Kew
Harisha BN (2017) An economic analysis of floriculture in India. Int J Acad Res Dev. https://doi.org/10.22271/academic
Hinsley A, de Boer HJ, Fay MF, Gale SW, Gardiner LM, Gunasekara RS, Kumar P, Masters S, Metusala D, Roberts DL, Veldman S (2017) A review of the trade in orchids and its implications for conservation. Bot J Linn Soc 186:435–455
Huded AK, Jingade P, Mishra MK (2018) A rapid and efficient SDS-based RNA isolation protocol from different tissues of coffee. 3 Biotech 8:183
Kiefer E, Heller W, Ernst D (2000) A simple and efficient protocol for isolation of functional RNA from plant tissues rich in secondary metabolites. Plant Mol Biol Report 18:33–39. https://doi.org/10.1007/BF02825291
Lau S-E, Schwarzacher T, Othman RY, Harikrishna JA (2015) dsRNA silencing of an R2R3-MYB transcription factor affects flower cell shape in a Dendrobium hybrid. BMC Plant Biol 15:194. https://doi.org/10.1186/s12870-015-0577-3
Lee WS, Gudimella R, Wong GR, Tammi MT, Khalid N (2015) Harikrishna JA (2015) Transcripts and microRNAs responding to salt stress in Musa acuminata Colla (AAA Group) cv Berangan roots. PLoS One 10(5):e0127526
Leh TY, Yong CS, Nulit R, Abdullah JO (2019) Efficient and high-quality RNA isolation from metabolite-rich tissues of Stevia rebaudiana, an important commercial crop. Trop Life Sci Res 30:149
Lehninger AL, Nelson DL, Cox MM (2005) Lehninger principles of biochemistry. WH Freeman, New York
Li R (2015) Forensic biology. CRC Press, Boca Raton
Liu L, Han R, Yu N, Zhang W, Xing L, Xie D, Peng D (2018) A method for extracting high-quality total RNA from plant rich in polysaccharides and polyphenols using Dendrobium huoshanense. PLOS One 13:e0196592. https://doi.org/10.1371/journal.pone.0196592
Moretti M, Cossignani L, Messina F, Dominici L, Villarini M, Curini M, Marcotullio MC (2013) Antigenotoxic effect, composition and antioxidant activity of Dendrobium speciosum. Food Chem 140:660–665. https://doi.org/10.1016/j.foodchem.2012.10.022
Muoki RC, Paul A, Kumari A, Singh K, Kumar S (2012) An improved protocol for the isolation of RNA from roots of tea (Camellia sinensis (L.) O. Kuntze). Mol Biotechnol 52:82–88. https://doi.org/10.1007/s12033-011-9476-5
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4326
Qadri R, Iqbal A, Wu Y, Li J, Nisar N, Azam M, Yang Y (2019) A modified protocol for total RNA isolation from different oil palm (Elaeis guineensis) tissues using cetyltrimethylammonium bromide. Curr Sci 116:479–482
Raizada A, Jegadeesan S (2019) An improved method for rapid isolation of DNA and RNA from leaves, flowers and roots of blackgram [Vigna mungo (L.) Hepper] for detection of begomovirus infection and RT-PCR. Electron J Plant Breed 10:167–176
Rodríguez-Ávila NL, Narváez-Zapata JA, Aguilar-Espinosa M, Rivera-Madrid R (2011) Regulation of pigment-related genes during flower and fruit development of Bixa orellana. Plant Mol Biol Report 29:43–50
Sarmah D, Kolukunde S, Sutradhar M, Singh BK, Mandal T, Mandal N (2017) A review on: in vitro cloning of orchids. Int J Curr Microbiol Appl Sci 6:1909–1927
Schroeder A, Mueller O, Stocker S, Salowsky R, Leiber M, Gassmann M, Lightfoot S, Menzel W, Granzow M, Ragg T (2006) The RIN: an RNA integrity number for assigning integrity values to RNA measurements. BMC Mol Biol 1:3. https://doi.org/10.1186/1471-2199-7-3
Shu C, Sun S, Chen J, Chen J, Zhou E (2014) Comparison of different methods for total RNA extraction from sclerotia of Rhizoctonia solani. Electron J Biotechnol 17:50–54
Su CL, Chao YT, Alex Chang YC, Chen WC, Chen CY, Lee AY, Hwa KT, Shih MC (2011) De novo assembly of expressed transcripts and global analysis of the Phalaenopsis aphrodite transcriptome. Plant Cell Physiol 52:1501–1514. https://doi.org/10.1093/pcp/pcr097
Surzycki S (2012) Basic techniques in molecular biology. Springer Science & Business Media, Berlin
The nation Thailand (2018) https://www.nationthailand.com/breakingnews/30360623. Accessed 5 Sept 2019
Toni LS, Garcia AM, Jeffrey DA, Jiang X, Stauffer BL, Miyamoto SD, Sucharov CC (2018) Optimization of phenol-chloroform RNA extraction. MethodsX 5:599–608
Wong L-M, Silvaraj S, Phoon L-Q (2014) An optimised high-salt CTAB protocol for both DNA and RNA isolation from succulent stems of Hylocereus sp. J Med Bioeng 3:236–240
Acknowledgements
This work was supported by the Frontier Research Grant (FRG 2017, FG026-17AFR) and CEBAR Research University Grants (RU006-2017 and RU006-2018).
Author information
Authors and Affiliations
Contributions
MAKA, PM, S-EL, and TTT conducted experimental work; PM and JAH made the experimental design. All authors contributed to manuscript preparation and to editing of both original and revised manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
13205_2019_1898_MOESM2_ESM.pptx
Supplementary Fig. S1 Flow diagram of cDNA synthesis from total RNA (SuperScript® III First-Strand Synthesis system (Invitrogen, USA)
13205_2019_1898_MOESM3_ESM.pptx
Supplementary Fig. S2 Chalcone synthase (CHS) gene sequence of Dendrobium hybrid cultivar Sonia Earsakul (KC345011.1) with primer binding sites (Orange letter denotes introns and blue- bold letter denotes primer binding sites)
13205_2019_1898_MOESM4_ESM.pptx
Supplementary Fig. S3 Aqueous fraction collected after P:C:I steps of RNA isolation method. M1: RNA isolation method of Kiefer et al. 2000, M2: RNA isolation method of Su et al. 2011, M3: RNA isolation method of Gao et al. 2016, M4: RNA isolation method of Liu et al. 2018, and M5: improved CTAB method
13205_2019_1898_MOESM6_ESM.pptx
Supplementary Fig. S5 Agarose gel electrophoresis of total RNA isolated from the mature flower of four Dendrobium hybrid flowers. Lanes 1–3: Dendrobium Burmese ruby × Dendrobium Mae-klong River; Lanes 4–6: Dendrobium Burana Jade × [Dendrobium Bertha Chong × Dendrobium Imelda Romualdez]), Lanes 7–9: Dendrobium Trudy Brandt × Dendrobium Udom Blue Angel, Lanes 10–12: Dendrobium Aridang × Dendrobium Burana Sundae and Lane 13: 1 kb DNA Ladder (Promega, USA)
Rights and permissions
About this article
Cite this article
Khairul-Anuar, MA., Mazumdar, P., Lau, SE. et al. High-quality RNA isolation from pigment-rich Dendrobium flowers. 3 Biotech 9, 371 (2019). https://doi.org/10.1007/s13205-019-1898-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-019-1898-y