Abstract
Novel cry genes are potential candidates for resistance management strategies, due to their different structures and modes of action. Therefore, it is desirable to clone and express novel cry genes from several new isolates of Bacillus thuringiensis (Bt). In the present study, 28 Bt strains were characterized at morphological and molecular level. All these strains are Gram positive, endospore forming and had shown different crystal morphologies when viewed under the microscope. The ARDRA (16S rDNA PCR-RFLP technique) with AluI, HaeIII, HinfI and TaqI produced unique and distinguishable restriction patterns used for the molecular characterization of these isolates. Based on UPGMA clustering analysis, Bt strains showed significant molecular diversity and the dendrogram obtained differentiated 28 Bt strains into 1 major cluster at a similarity coefficient 0.56. PCR analysis demonstrated that the Bt strains showed diverse cry gene profiles with several genes per strain. The Bt strain G3C1 showed the presence of maximum cry-type genes by PCR. The toxicological characterization of these cry genes will have huge importance in transgenic technology and will be useful in transgenesis of crop plants for better resistance management.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacillus thuringiensis (Bt), is a Gram-positive, spore-forming soil bacterium that forms parasporal insecticidal crystal proteins during the stationary and sporulation phase of its growth cycle. These proteins are termed delta-endotoxins because of their intracellular location and have been used for many years as successful biological insecticides (Schnepf et al. 1998). Commercial Bt-based bio-insecticides used worldwide are applied at 10–50 g/acre or about 1020 molecules/acre, while chemical pesticides such as organophosphates and synthetic pyrethroids are applied about 8 × 1024 molecules/acre and 3 × 1022 molecules/acre, respectively. Thus, molecular potency of these toxins is 80,000 times better than organophosphates and 300 times greater than synthetic pyrethroids (Feitelson et al. 1992). Search for novel Bt strains may lead to the discovery of novel insecticidal proteins with higher toxicity which will be important for providing alternatives to cope up with the emergence of resistant insect populations. Therefore, there is a need of isolation of large number of Bt strains from diverse geographical conditions and cloning of many new types of insecticidal crystal proteins genes (Ramalakshmi and Udayasuriyan 2010).
The polymerase chain reaction (PCR) has been widely used to characterize the Bt strain collections (Ceron et al. 1995; Bravo et al. 1998; Ferrandis et al. 1999; Beron et al. 2005; Vidal-Quist et al. 2009). This technique is a highly sensitive method for rapid detection and identification of target cry gene sequences requires very small amounts of DNA and allows simultaneous screening of many Bt strains to classify them and predict their insecticidal activities. The efficacy of PCR in identifying the large family of cry genes is based on the presence of conserved regions. Another strategy for the screening is based on the multiplex PCR which uses more than two primers in a mixture of the same reaction (Juarez-Perez et al. 1997).
Continuous exposure to a single kind of Bt toxin can lead to resistance development in insects. Routine replacement of cry genes or pyramiding of cry genes could be useful for effective control of insect pests by Bt transgenic plants. Variation of a single amino acid can significantly influence the level of toxicity in Cry proteins (Udayasuriyan et al. 1994). The applications of Bt products as biopesticides are limited by their narrow host range, low toxicity to the targeted insects and the resistance from insects. Therefore, it is necessary to continuously screen novel cry genes and perform rational design based on a known Cry toxin (Lin et al. 2008). In general, the type of cry and cyt genes present in a strain correlates to some extent with its insecticidal activity (Porcar and Juárez-Pérez 2003). Since India is very rich in biodiversity and genetic resources, different Bt strains available in the country are valuable source for identification of indigenous, novel Bt genes, which could encode more potent toxins due to sequence variations (Jain et al. 2006). In the present study, 28 strains of Bt were characterized and screened for novel cry genes for better resistance management.
Materials and methods
Bacterial strains, chemicals and oligonucleotide primers
The Bt were isolated from the northeast region of India (Table 1). PCR chemicals, oligonucleotide primers and restriction enzymes were procured from Bangalore Genei, Pvt. Ltd., India, and AB genes Pvt. Ltd., UK. Microscopic examinations of the isolated strains were used for the characterization of Bt by physiological methods of staining such as Gram, endospore and crystal staining.
Molecular characterization using ARDRA of the 16S rDNA region
The total genomic DNA of Bt strains were isolated according to standard protocols (Kalman et al. 1993). The total DNA pattern of all Bt strains were analyzed on agarose gel. Genomic DNA isolated from Bt strains were used as template for the PCR amplification. Amplification of 16S rDNA region was performed with universal primers, 27F (5′AGAGTTTGATCCTGGCTCAG3′) and 1492R (5′ACGGCTACCTTGTTACGACTT3′). Each 40 µl reaction mixture contained 50 ng of genomic DNA of Bt strain, 50 ng of forward and reverse primers each, each dNTP at a final concentration of 200 µM and 1 U of Taq polymerase in 1× Taq buffer (with 15 µM MgCl2). Amplification was accomplished with the thermal cycler (Eppendorf, Germany). The PCR was performed with Taq polymerase for 30 cycles as follows: 94 °C for 1 min, 60 °C for 45 s and 72 °C for 1 min., the final extension was performed for 10 min at 72 °C. The 16S rDNA amplicons were digested using four different restriction endonucleases viz HinfI, HaeIII, AluI and TaqI (Cihan et al. 2012). The restriction-digested products were analyzed on 2% agarose gel prepared in 1× TAE buffer containing 0.5 µg/ml of ethidium bromide. Electrophoresis was carried out at 100 V for 3 h in 1× TAE electrophoresis buffer. Data analysis was done using NTSYS-PC (Numerical Taxonomy and Multivariate Analysis System) software and SIMUQUAL Jaccard’s similarity coefficient (Rohlf 1997).
PCR-based cry gene screening in indigenous Bt strains
Genomic DNA of the Bt was used in PCR with cry gene-specific screening primers (Table 2). The PCR was accomplished using an Eppendorf thermal cycler as prescribed by Jain et al. (2012).
Results and discussion
Characterization of indigenous Bt strains
All the Bt isolates were found to be Gram positive and endospore forming. Similarly different crystal morphologies were seen blue under the white background and showed bi-pyramidal, spherical, cuboidal, rectangular, irregular crystals, etc. The shape of the crystalline inclusions varied among the 28 Bt strains. Shishir et al. (2012) identified Bt isolates based on their hemolytic activity, presence of parasporal crystal proteins and crystal protein profile and observed five different types of parasporal crystal proteins such as spherical, bi-pyramidal, irregular pointed, cuboidal and irregular shaped which formed irregular white colonies with a pink background among the isolates which indicates the diversity of the local Bt isolates. Similarly, Unalmis et al. (2015) isolated Bt-like colonies and characterized them on the basis of Gram staining, spore straining and crystal staining. From 60 bacterial colonies observed through bright field microscopy, 28 isolates were identified as Bt based on the presence of crystalline inclusions.
Molecular characterization of indigenous Bt strains using ARDRA
Amplified rDNA (Ribosomal DNA) restriction analysis (ARDRA) is the extension of the technique of RFLP (restriction fragment length polymorphism) to the gene encoding the small ribosomal subunit (16S) of bacteria. The technique involves an enzymatic amplification using primers directed at the conserved regions at the ends of the 16S rDNA, followed by digestion using restriction enzymes. The pattern obtained is said to be representative of the species analyzed and important for their molecular characterization. In the present study ARDRA produced a fingerprint based on length polymorphism for molecular characterization of native Bt strains. Four restriction endonucleases viz., HinfI, HaeIII, AluI and TaqI were used for restriction fragment analysis of amplified 16S rDNA. The banding patterns of the representative Bt are shown in Fig. 1, with standard molecular weight marker. Totally 20 bands of varying sizes were observed in all the 28 strains when digested with four restriction enzymes. When digested with HaeIII, 7 different DNA fragments were obtained, whereas 3 different DNA fragments were obtained with HinfI endonuclease, TaqI digestion resulted 6 different DNA fragments while AluI resulted in 4 different DNA fragments.
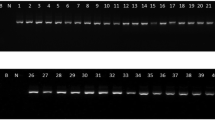
Agarose gel electrophoresis of a PCR amplification of 16S rDNA region from native Bt strains. b ARDRA patterns of 16S rDNA sequence of Bt strains by restriction enzyme AluI. M1 500 bp DNA ladder, Bt strains 1 S4C1, 2 S16C2, 3 S4C2, 4 D4C1, 5 T5C1, 6 S20C4, 7 G1C1, 8 S19C1, 9 S22C1, 10 S20C3, 11 D9C1, 12 T8C2, 13 T2C2, 14 S21C1, 15 D1C1, 16 G3C1, 17 J11C1, 18 D12C1, 19 T3C2, 20 G2C1, 21 D5C1, 22 S29C2, 23 J11C3, 24 T7C1, 25 S23C1, 26 S6C3, 27 S11C1, 28 J10C1
Genetic similarity estimates the variations based on ARDRA banding patterns, which were calculated using method of Jaccard’s coefficient analysis. The similarity coefficient matrix generated was subjected to algorithm “Unweighted Pair Group Method for Arithmetic Average (UPGMA)” to generate clusters using NTSYS 2.02 pc program. The pairwise comparison of ARDRA patterns based on both shared and unique amplification products was made to generate a similarity matrix. Similarity indices established on the basis 20 bands of 4 restriction enzymes ranged from 0.11 to 0.93.
The dendrogram (Fig. 2) is a close representation of the values obtained in the similarity matrix. The dendrogram depicted the relationship among the Bt strains and clearly divided into one major cluster at a similarity coefficient 0.56. The dendrogram clearly indicated that S6C3 and S23C1 were different from remaining strains, hence these are more diverse. The first (A) cluster was divided into two sub-clusters A1 and A2. Sub-cluster A1 included 24 strains and further divided into two sub-clusters A1a and A1b. Sub-cluster A1a included strains J11C3, G2C1, S20C4, T7C1, S21C1, T8C2, D12C1, S20C3, D9C1, D5C1, S22C1, T5C1, J11C1, D4C1, and S4C1. S4C1, D4C1 and J11C1 and sub-cluster A1b comprises only one strain S11C1. Sub-cluster A2 comprises 2 strains S16C2 and S4C2.
Molecular techniques have helped to develop easy and rapid methods to perform microbial characterization at genus, species and even at strain level. Molecular markers have been found to be strain specific and have proved to be valuable tools in the characterization and evaluation of genetic diversity within and between species and populations. Identification of Bacillus species using conventional sequencing methods can divulge their taxonomic affiliation, but there are certain groups of Bacillus where alternate methods like ARDRA and PCR fingerprinting can expose the exact lineage of the species rapidly.
Saadaoui et al. (2009) reported a new Bacillus thuringiensis kurstaki strain BLB1, isolated from a Tunisian soil sample showed the same 16S rDNA ARDRA profile than HD1 using TaqI, AluI, MboII, and MspI restriction enzymes. Gowdaman et al. (2014) reported that a collection of 171 soil bacterial isolates was analyzed for the occurrence of genus Bacillus using group-specific primers and ARDRA was performed for the Bacillus-positive isolates with standard Bacillus strains. Sangeetha et al. (2016) analyzed 15 bacterial isolates by ARDRA and used eight restriction endonucleases (CfoI, HinfI, RsaI, DdeI, Sau3AI, AluI, HaeIII, and MspI) forming two heterogenous main clusters after analysis by unweighted pair-group method using arithmetic averages.
PCR-based screening of cry genes in native Bt strains
Amplification of expected size of PCR products in different primer pairs (Fig. 3) indicated the presence of the above-mentioned cry-type genes in Bt strains (Table 3). The primers cry2, cry3, cry5 and cry7–8 did not show any amplification in PCR-based cry gene screening. The strains containing cry1-type genes were the most abundant (100%) in the indigenous Bt strains since all the strains were harboring these genes followed by cry4 (84.14%), cry9 (64.28%), cry11 (39.28%), cyt2 (39.28%), and vip3A (25%) (Table 4). Agarose gel electrophoresis showed non-specific amplification along with specific partial cry gene amplicon which was also observed in many published research findings using the same screening primers.
Agarose gel electrophoresis of a DNA isolation from native Bt strains. b PCR amplification of partial cry 1 gene from Bt strains. c PCR amplification of partial cry4 gene from Bt strains. M1 500 bp DNA ladder, Bt strains 1 S4C1, 2 S16C2, 3 S4C2, 4 D4C1, 5 T5C1, 6 S20C4, 7 G1C1, 8 S19C1, 9 S22C1, 10 S20C3, 11 D9C1, 12 T8C2, 13 T2C2, 14 S21C1, 15 D1C1, 16 G3C1, 17 J11C1, 18 D12C1, 19 T3C2, 20 G2C1, 21 D5C1, 22 S29C2, 23 J11C3, 24 T7C1, 25 S23C1, 26 S6C3, 27 S11C1, 28 J10C1
Jain et al. (2012) observed the frequency of cry-type genes in eight Bt strains (IS1–IS8) with the result that the cry1 type genes were most abundant in the indigenous isolates since all the strains were harboring these genes, followed by vip3A (87.5%), cry2 (75%), cry9 (62%), cry3 (50%), cry11 (37.5%), cry7–8 (37.5%), cry5, 12, 14, 21 (25%), cyt1 (25%), cry4 (12.5%) and cyt2 (12.5%) as detected by PCR. Patel et al. (2012) reported the diversity of cry genes from different soil types and climatic environments and reported the presence of cry1, cry2, cry3, 7, 8, cry4, cry5, 12, 14, 21, cry11, cry13 and cyt1 genes from Bt, whereas absence of cry3 and cry13 genes were reported in the isolates of non-agricultural samples. Similarly, Salekjalali et al. (2012) identified isolates harboring different cry-type genes through PCR and found 47% of the strains amplified with the cry1 primer, 29% with cry3 and 13% with cry4. Salama et al. (2015) reported that cry1 gene is the most abundant in these isolates (83.33%) among tested cry-type genes, followed by cry1 gene subfamilies (cry1B and cry1C) with percentage of 38.88 and 77.77%, respectively. The tested isolates showed the presence of cry2A gene, but not all of these isolates were positive for cry2 gene (55.55%). Only 27.77 and 16.66% of the tested isolates harbor cry4 and cry3 genes, respectively.
The use of PCR has greatly improved cry gene detection; however, this method is mostly limited to members of previously described gene families and requires a large number of primers. The results of the present study suggest the presence of diversity in the native Bt isolates. Further studies on cloning and characterization of those novel cry genes from these new isolates of Bt will be useful and open new opportunities in the area of integrated pest management for sustainable agriculture.
Abbreviations
- Bt:
-
Bacillus thuringiensis
- Cry ptrotein:
-
Crystal protein
- PCR:
-
Polymerase chain reaction
- ARDRA:
-
Amplified ribosomal DNA restriction analysis
- UPGMA:
-
Unweighted pair-group method for arithmetic average
References
Beron CM, Curatti L, Salerno GL (2005) New strategy for identification of novel Cry-type genes from Bacillus thuringiensis strains. App Environ Microbiol 71(2):761–765
Bravo A, Sarabia S, Lopez L, Ontiveros H, Abarca C, Ortiz A, Ortiz M, Lina L, Villalobos FJ, Peña G, Nuñez-Valdez ME, Soberón M, Quintero R (1998) Characterization of cry genes in a Mexican Bacillus thuringiensis strain collection. App Environ Microbiol 64(12):4965–4972
Ceron J, Ortiz A, Quintero R, Guereca L, Bravo A (1995) Specific PCR primers directed to identify cry I and cry III genes within a Bacillus thuringiensis strain collection. App Environ Microbiol 61(11):3826–3831
Cihan C, Tekin N, Ozcan C, Cokmus C (2012) The genetic diversity of genus Bacillus and the related genera revealed by 16 s rRNA gene sequences and ARDRA analyses isolated from geothermal regions of turkey. Br J Microbiol 43:309–324
Feitelson JS, Payne J, Kim L (1992) Bacillus thuringiensis: insects and beyond. Nat Biotechnol 10(3):271–275
Ferrandis MD, Juárez-Pérez VM, Frutos R, Bel Y, Ferré J (1999) Distribution of cryI, cryII and cryV genes within Bacillus thuringiensis isolates from Spain. Syst App Microbiol 22(2):179–185
Gowdaman V, Kumar RM, Venkatachalam S, Prabagaran SR (2014) Comparison of DNA fingerprinting analysis for identification of Bacillus species. Int J Res Adv Technol 2:278–288
Jain D, Udayasuriyan V, Arulselvi P, Dev S, Sangeetha P (2006) Cloning, characterization, and expression of a new cry2Ab gene from Bacillus thuringiensis strain 14-1. App Biochem Biotechnol 128(3):185–194
Jain D, Kachhwaha S, Jain R, Kothari S (2012) PCR based detection of cry genes in indigenous strains of Bacillus thuringiensis isolated from the soils of Rajasthan. Indian J Biotechol 11:491–494
Juarez-Perez VM, Ferrandis MD, Frutos R (1997) PCR-based approach for detection of novel Bacillus thuringiensis cry genes. App Environ Microbiol 63(8):2997–3002
Kalman S, Kiehne KL, Libs JL, Yamamoto T (1993) Cloning of a novel cry I C-type gene from a strain of Bacillus thuringiensis subsp. galleriae. App Environ Microbiol 59(4):1131–1137
Lin Y, Fang G, Cai F (2008) The insecticidal crystal protein Cry2Ab10 from Bacillus thuringiensis: cloning, expression, and structure simulation. Biotechnol Lett 30(3):513–519
Patel KD, Purani S, Ingle SS (2012) Distribution and diversity analysis of Bacillus thuringiensis cry genes in different soil types and geographical regions of India. J Invertebr Pathol 112:116–121
Porcar M, Juárez-Pérez V (2003) PCR-based identification of Bacillus thuringiensis pesticidal crystal genes. FEMS Microbiol Rev 26(5):419–432
Ramalakshmi A, Udayasuriyan V (2010) Diversity of Bacillus thuringiensis isolated from Western Ghats of Tamil Nadu State. India Curr Microbiol 61(1):13–18
Rohlf F (1997) NTSYS-pc: numerical taxonomy and multivariate analysis system, version 2.0. Exeter Software, Setauket, New York
Saadaoui I, Rouis S, Jaoua S (2009) A new Tunisian strain of Bacillus thuringiensis kurstaki having high insecticidal activity and δ endotoxin yield. Arch Microbiol 191:341–348
Salama HS, El-Ghany NM, Saker MM (2015) Diversity of Bacillus thuringiensis isolates from Egyptian soils as shown by molecular characterization. J Genet Eng Biotechnol 13:101–109
Salekjalali M, Barzegari A, Jafari B (2012) Isolation, PCR detection and diversity of native Bacillus thuringiensis strains collection isolated from diverse Arasbaran Natural Ecosystems. World App Sci J 18:1133–1138
Sangeetha BG, Jayaprakas CA, Siji JV, Rajitha M, Shyni B, Mohandas C (2016) Molecular characterization and amplified ribosomal DNA restriction analysis of entomopathogenic bacteria associated with Rhabditis (Oscheius) spp. Biotechnol 6:32. doi:10.1007/s13205-015-0326-1
Schnepf E, Crickmore N, Van Rie J, Lereclus D, Baum J, Feitelson J, Zeigler DR, Dean DH (1998) Bacillus thuringiensis and its pesticidal crystal proteins. Microbiol Mol Biol Rev 62(3):775–806
Shishir A, Akter A, Hassan MH, Kibria G, Ilias M, Khan SN, Hoq MM (2012) Characterization of locally isolated Bacillus thuringiensis for the development of eco-friendly biopesticides in Bangladesh. J Biopest 5:216–222
Udayasuriyan VA, Nakamura A, Mori A, Masaki H, Uozumi T (1994) Cloning of a new crylA(a), gene from B. thuringiensis strain FU-2-7 and analysis of chimeric cry1A(a) proteins of toxicity. Biosci Biotechnol Biochem 58:830–835
Unalmis S, Ayvaz A, Yilmaz S, Azizoglu U (2015) Molecular screening and bioactivity of native Bacillus thuringiensis isolates. J Agric Sci Technol 17:1197–1207
Vidal-Quist JC, Castañera P, González-Cabrera J (2009) Diversity of Bacillus thuringiensis strains isolated from citrus orchards in Spain and evaluation of their insecticidal activity against Ceratitis capitata. J Microbiol Biotechnol 19(8):749–759
Acknowledgements
Financial assistance from AINP on soil biodiversity and Biofertilizers is highly acknowledged.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All the authors declare that there is no conflict of interest.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Jain, D., Sunda, S.D., Sanadhya, S. et al. Molecular characterization and PCR-based screening of cry genes from Bacillus thuringiensis strains. 3 Biotech 7, 4 (2017). https://doi.org/10.1007/s13205-016-0583-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-016-0583-7