Abstract
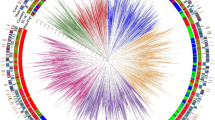
A total of 16,619 ESTs sequences (SSRs) of sesame (Sesamum indicum L.) were mined from Genbank. From sequences, 156 primer pairs were designed and characterized to determine the diversity among 49 sesame accessions. Twenty SSRs were found to be polymorphic and the number of alleles ranged from two to five per locus. The allele size varied from 101 to 399 bp. The average PIC value of the 20 SSR loci was 0.72 ranging from 0.49 (SEM-12-68) to 0.90 (SEM-12-27). Dendrogram analysis grouped the 49 genotypes into five separate clusters exhibiting a genetic similarity coefficient from 0.59 to 1.0. Hence, these EST-derived SSRs markers could be useful in assessing the diversity of sesame accessions and could also help in identifying diverse parents for sesame improvement programs.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Abdellatef E, Sirelkhatem RM, Mohamed Ahmed M, Radwan KH, Khalafalla MM. 2008. Study of genetic diversity in Sudanese sesame (Sesamum indicum L.) germplasm using random amplified polymorphic DNA (RAPD) markers. Afr. J. Biotech. 7: 4423–4427
Ali GM, Yasumoto S, Katsuta MS. 2007. Assessment of genetic diversity in sesame (Sesamum indicum L.) detected by amplified fragment length polymorphism markers. Electron. J. Biotechnol. 10: 12–23
Ashri A. 1998. Sesame breeding. Plant Breed Rev. 16: 179–228
Bedigian D. 1981. Origin, diversity, exploration and collection of sesame. In Sesame: Status and improvement. FAO Plant Production and Protection Paper 29. FAO, Rome, pp 164–169
Bedigian D. 2010. Characterization of sesame (Sesamum indicum L.) germplasm: a critique. Genet. Resour. Crop Evol. 57: 641–647
Bhat KV, Babrekar PP, Lakhanpaul S. 1999. Study of genetic diversity in Indian and exotic sesame (Sesamum indicum L.) germplasm using random amplified polymorphic DNA (RAPD) markers. Euphytica 110: 21–33
Bin WL, Yang ZL, Zhan ZY, Zhen GW, Zhen ZT. 2008. Developing EST-Derived Microsatellites in sesame (Sesamum indicum L.). Acta Agron. Sin. 34: 2077–2084
Bisht IS, Mahajan RK, Loknathan TR, Agrawal RC. 1998. Diversity in Indian sesame collection and stratification of germplasm accessions in different diversity groups. Genet. Resour. Crop Evol. 45: 325–335
Dixit A, Jin MH, Chung JW, Yu JW, Chung HK, Ma KH, Park YJ, Cho EG. 2005. Development of polymorphic microsatellite markers in sesame (Sesamum indicum L.). Mol. Ecol. Notes. 5: 736–738
Ercan AG, Taskin M, Turgut K. 2004. Analysis of genetic diversity in Turkish sesame (Sesamum indicum L.) populations using RAPD markers. Genet. Resour. Crop Evol. 51: 599–607
FAO. Available at: http://faostat.fao.org/ 2009
Fazal Akbar M, Ashiq Rabbani M, Shahid M, Zabta KS. 2011. Genetic diversity of sesame (Sesamum Indicum L.) germplasm from Pakistan using RAPD markers. Pak. J. Bot. 43: 2153–2160
Gupta PK, Rustgi S, Sharma S, Singh R, Kumar N, Balyan HS. 2003. Taransferable EST-SSR markers for the study of polymorphism and genetic diversity in bread wheat. Mol. Genet. Genomics 270: 315–323
Gebremichael DE, Parzies HK. 2010. Genetic variability among landraces of sesame in Ethiopia. Afr. Crop Sci. J. 19: 1–13
Hiremath SC, Patil CG. 1999. Genome homology and the putative progenitor of sesame. J. Cytology Genetics 34: 69–74
Jyothi B. 2009. Molecular mapping and characterization of yield QTL and tagging of wilt resistance gene(s) in sesame (Sesamum indicum L.). Thesis, Acharya NG, Ranga Agricultural University
Kim DH, Zur G, Danin-Poleg Y. 2002. Genetic relationships of sesame germplasm collection as revealed by inter-simple sequence repeats. Plant Breed. 121: 259–262
Kobayashi T. 1981. The wild and cultivated species in the genus Sesamum. Sesame: Status and improvement. Proceedings of Expert Consultation, Rome, Italy, 8–12 December, 1980. FAO Plant Production and Protection Paper 29, pp 157–163
Kumar V, Sharma SN. 2009. Assessment of genetic diversity of sesame (Sesamum indicum L.) genotypes using morphological and RAPD markers. Ind. J. Genet. 69: 209–218
Kumar V, Sharma SN. 2011. Comparative potential of phenotypic, ISSR and SSR markers for characterization of sesame (Sesamum indicum L.) Varieties from India. J. Crop Sci. Biotech. 14: 163–171
Laurentin H, Karlovsky P. 2006. Genetic relationship and diversity in a sesame (Sesamum indicum L.) germplasm collection using amplified fragment length polymorphism. BMC-Genet. 7: 10
Li G, Quiros CF. 2001. Sequence-related amplified polymorphism (SRAP), a new markersystem based on a simple PCR reaction: its application to mapping and genetagging in Brassica. Theor. Appl. Genet. 103: 455–461
Li YY, Shen JX, Wang TH, Fu TD, Ma CZ. 2007. Construction of a linkage map using SRAP, SSR and AFLP markers in Brassica napus L. Sci. Agric. Sinica 40: 1118–1126 (in Chinese)
Liu BH. 1997. Statistical Genomics: Linkage, Mapping and QTL Analysis. CRC Press, Boca Raton, FL, pp 648
Laurentin HE, Karlovsky P. 2007. AFLP fingerprinting of sesame (Sesamum indicum L.) cultivars: identification, genetic relationship and comparison of AFLP informativeness parameters. Genet. Resour. Crop Evol. 54: 1437–1446
Nayar NM, Mehra KL. 1970. Sesame: Its uses: botany, cytogenetics, and origin. Econ. Bot. 24: 20–31
Nweke FN, Ubi BE, Kunert K. 2012. Application of microsatellite polymorphisms to study the diversity in seed oil content and fatty acid composition in Nigerian sesame (Sesamum indicum L.) accessions. Afr. J. Biotechnol. 11: 8820–8830
Parsaeian M, Mirlohi A, Saeidi G. 2011. Study of genetic variation in sesame (Sesamum indicum L.) using agromorphological traits and ISSR markers. Russ. J. Genet. 47: 314–321
Pathirana R. 1994. Natural cross-pollination in sesame (Sesamum indicum L.). Plant Breed. 112: 167–170
Pham TD, Geleta M, Bui TM, Bui TC, Merker A, Carlsson AS. 2011. Comparative analysis of genetic diversity of sesame (Sesamumindicum L.) from Vietnam and Cambodia using agro-morphological and molecular markers. Hereditas 148: 28–35
Rao PVR, Anuradha G, Sridhar S, Gouri Shankar V, Raja Reddy K, Eswara Reddy NP, Siddiq EA. 2010. Standardization of DNA extraction protocol in sesame (Sesamum indicum L.). Int. J. Trop. Agric. 28:3–8, 329–333
Rheenen HAV. 1980. Aspects of natural cross-fertilization in season. (Sesamum indicum L.). Trop. Agric. 57: 53–59
Rohlf FJ. 2000. NTSYS-PC Numerical taxonomy and multivariate analysis system. In FJ Rohlf, NTSYS-PC: Numerical Taxonomy and Multivariate Analysis System. Applied Biostatistics, Exerter Publishing Ltd., New York, Software, ISBN 0-925031-18-6
Sun J, Tu YQ, Zhang XR. 2007. DNA fingerprint analysis in space-induced sesame mutant lines. Sci. Agric. Sin. 40: 2696–2701 (in Chinese)
Spandana B, Prathap Reddy V, John Prasanna G, Anuradha G, Sivaramakrishnan S. 2012. Development and characterization of microsatellite markers (SSR) in sesamum (Sesamum indicum L.) species. Appl. Biochem. biotechnol. 168: 1594–1607
Squirrell J, Hollingsworth PM, Woodhead M, Russell J, Lowe AJ, Gibby M, Powell W. 2003. How much effort is required to isolate nuclear microsatellites from plants?. Mol. Ecol. 12: 1339–1348
Zane L, Bargelloni L, Patarnello T. 2002. Strategies for microsatellite isolation: a review. Mol. Ecol. 11: 1–11
Wei W, Qi XQ, Wang LH, Zhang YX, Hua W, Li DH, Lv HX, Zang XR. 2011. Characterization of sesame (Sesamumindicum L.) global transcriptome using Illumina pair-end sequencing and development EST-SSR markers. BMC Genomics 12: 451
Zhang H, Wei L, Miao H, Zhang T, Wang C. 2012. Development and validation of genic-SSR markers in sesame by RNA-seq. BMC Genomics 13: 316
Zhang XR, Chen KR, Peng J, Xu ZY. 2004. The RAPD analysis and genetic diversity of selected sesame germplasm. Chinese J. Oil Crop Sci. 26: 34–37
Zhang YX, Zhang XR, Hua W, Wang LH, Che Z. 2010. Analysis of genetic diversity among indigenous landraces from sesame (Sesamum indicum L.) core collection in China as revealed by SRAP and SSR markers. Genes Genomics 32: 207–215
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yepuri, V., Surapaneni, M., Kola, V.S.R. et al. Assessment of genetic diversity in sesame (Sesamum indicum L.) genotypes, using EST-derived SSR markers. J. Crop Sci. Biotechnol. 16, 93–103 (2013). https://doi.org/10.1007/s12892-012-0116-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12892-012-0116-9