Abstract
The effect of glucose, chicken manure, and filter mud on the ammonium and nitrate concentrations, ammonia-oxidizing bacterial community and bacterial community in latosolic red soils during the incubation of microcosms was investigated. The soil nitrate concentration was significantly lower in the glucose-treated soil than in the filter mud or chicken manure-treated soil from days 2 and 5 to 21 of incubation. The ammonia-oxidizing bacteria community composition, measured by terminal restriction fragment length polymorphism analysis, was different among the treatments 9 days after incubation, suggesting that the control soil without external fertilization had a low 283-bp (Nitrosospira) fragment relative abundance (27 %) compared with the glucose-treated (62 %), filter mud (73 %) and chicken manure (78 %) samples. Additionally, 491-bp fragments (Nitrosomonas) were detected in all the soil treatments except for the control soil, and 48-bp fragments (from different Nitrosomonas) were detected in the chicken manure-treated soil. The bacterial community structure was markedly changed in the glucose-treated soil on day 9 and in the filter mud-treated soil on day 31, indicating that the effect of filter mud on the bacterial community is delayed compared to the effect of glucose. The chicken manure-treated soil showed less change, similar to that of the control soil. Glucose fertilization greatly increased the soil bacterial abundance and functional diversity; however, the chicken manure and filter mud did not stimulate soil bacterial activity on day 9. These results indicated that nitrification may have been somewhat suppressed in the glucose-treated soils, which was possibly related to the improving ammonia-oxidizing bacterial community, bacterial community and activity via the available carbon application. The filter manure and chicken manure treatments demonstrated fewer effects. These results suggest that organic carbon quality, e.g., increasing the available carbon, regulates the nitrification process and is beneficial to reducing soil nitrogen losses.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Nitrification plays an important role in the retention of nitrogen in ecosystems. Soil organisms are one of the key factors for regulating nitrification. Much of this research has focused on the relative contributions of chemolithotrophic (autotrophic nitrification) and chemo-organotrophic (heterotrophic nitrification) microorganisms (De Boer and Kowalchuk 2001; Kowalchuk and Stephen 2001; He et al. 2012). Ammonia oxidation by autotrophic ammonia-oxidizing bacteria (AOB) plays an essential role in the nitrogen (N) cycle and is stimulated by soil NH3 (Belser 1979; Laanbroek and Woldendorp 1995). Recent studies showed that ammonia-oxidizing archaea (AOA) possess homologs of the bacterial ammonia monooxygenase gene and play an important role in ammonia oxidization (Venter et al. 2004; Treusch et al. 2005; Lehtovirta-Morley et al. 2011). Many factors are involved in regulating soil nitrification, including the soil pH (Stephen et al. 1998; Lehtovirta-Morley et al. 2011; Zhang et al. 2012), fertilization methods (Li et al. 2011; Cai et al. 2012), temperature (Avrahami et al. 2003; Tourna et al. 2008), vegetation composition (Schaffers and Sykora 2000), organic carbon (C) quantity and quality (Priha and Smolander 1999; Gundersen et al. 1998; Strauss and Lamberti 2002).
Organic amendments used in agriculture, including poultry manures and crop residues, are the source of nutrients for plants, and they provide elements to improve the physical, chemical and biological characteristics of the soil, to stimulate soil microbial activity and to alter the community composition for the enhancement of various microbiological soil processes (Garcia et al. 1994; Pascual et al. 1997; Rivero et al. 2004). There have been several research studies related to soil nitrification. Humic acid inhibits changes in the AOB community composition in soil saturated with urea during microcosm incubation (Dong et al. 2009). There was an increase in the AOB population size but a greater decrease in the metabolic activity in mineral N-fertilized soil compared to organic manure-fertilized soil in long-term fields (16 years; Chu et al. 2008). He et al. (2007) found that organic manure significantly alters the composition of the soil AOB in long-term field experiments.
There are clear differences in the nitrification process by various organic amendments. Racz et al. (2010) reported that peptone and glucose had different effects on the composition of the heterotrophic bacteria community and on the AOB community in mixed cultures. Nitrification of stream sediments was completely inhibited by glucose added at a concentration of 30 mg C L−1 in microcosm incubation (Strauss and Lamberti 2002). The stimulation of nitrification by the addition of organic N compounds such as peptone has been proposed as evidence for the occurrence of heterotrophic nitrification (Kreitinger et al. 1985; Adams 1986). The production of superoxide radicals during lignin degradation may accelerate the degradation and lead to the oxidation of simple organic substances, which leads to nitrate formation. Bastida et al. (2009) indicated that a lower availability of suitable substrates results in more efficient nitrification rates. It has been suggested that the stimulatory effect of amendments on soil nitrogen properties is predominantly related to the greater availability of labile C sources (Deng and Tabatabai 1997). Chantigny et al. (2002) indicated that pig slurry and alfalfa had better effects on the stimulation of soil microbial activities than cattle manure and maize residue, and these differences were partly related to the lignin-to-N ratio of the various amendments. The C and N transformations that occurred were in part determined by the C degradability and N availability of the organic amendments. Studies suggest that the effects of soil fertilization on the bacterial and AOB communities may be different depending on the nature of the complex organic amendment fertilization.
Among the available organic fertilizers, filter mud (FM), a rich resource in South China, is an agro-industrial waste from sugarcane mills obtained from the clarification of sugarcane juice. FM is a major source of plant nutrients, such as nitrogen, phosphorus and potassium, and of notable amounts of organic components, such as lignin (Elsayed et al. 2008). Chicken manure (CM) is an organic waste with lower lignin content that contains a considerable amount of nutrients, such as nitrogen, phosphorus, potassium, and other micronutrients. In contrast to FM, rapid mineralization of organic N is expected from CM (Holbeck et al. 2013). There is limited knowledge about the effect of the different characteristics of FM and CM on the dynamic soil nitrification process.
The objectives of this study were to evaluate the effect of the incorporation of glucose, CM and FM into soils on the dynamic nitrification processes over short periods of microcosm incubation and to determine the effects of their incorporation on the soil AOB, the bacterial community structures, the microbial population and functional activity that are possibly involved in microbial nitrification mechanisms.
Materials and methods
Soil and organic amendment
Latosolic red soil was collected at a depth of 0–20 cm from an experimental maize farm at the South China Agricultural University, Guangzhou, PR China. Chicken manure (CM) was collected from the South China Agricultural University chicken farm (cage layer system). Filter mud (FM) was obtained from the Guangzhou Sugar Factory. The soil CM and FM samples were air-dried, ground, passed through a 2-mm sieve and thoroughly mixed. The basic characteristics of the soil, CM and FM samples are shown in Table 1. The measurements of the physical and chemical properties of the soils, CM and FM samples were performed as described by Bao (2005). The total organic C was determined by the potassium dichromate external heating method. The pH value was determined using a combined glass electrode in a 1:1 (w:v) ratio of the sample with distilled 1 M KCl. The total N was determined by the Kjeldahl method. The total phosphorus (P) was determined by the sodium hydroxide fusion molybdate colorimetric method. The total potassium (K) was determined by a flame photometer. The electricity conductivity (EC) was determined by an EC meter. The lignin C content was determined by Van Soest and Wine’s method (1967).
Experimental design
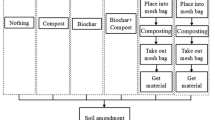
For each treatment, 0.6 kg (dry weight) of the air-dried soil was incubated in a sterile 1 L polyethylene plastic vessel in the dark. Before microcosm incubation, the soil sample was air-dried for 7 days at room temperature. The air-dried soil was adjusted to 70 % water retention capacity and pre-incubated for 7 days at 28 °C to recover the soil microorganisms. After the pre-incubation, the treatments (performed in triplicate) were as follows: (1) CK = no fertilization, (2) GL = 50 mg glucose C kg−1 soil, (3) CM = 50 mg chicken manure C kg−1 soil and (4) FM = 50 mg filter mud C kg−1 soil. Glucose was added to the soil in solution. The chicken manure and filter mud were dried for 72 h at 55 °C and ground to pass through a 2-mm screen before being mixed into the soil. Because the CM and FM originally consisted of organic N, P and K, different concentrations of chemical fertilizers (urea, potassium dihydrogen phosphate and potassium chloride) were added to all the treatments except to the CK soil. The total N, P and K concentrations applied to all the treated soil were 50 mg N, 25 mg P2O5 and 50 mg K2O kg−1 soil. Mineral nutrients and organic carbon were added to the soil as uniformly as possible. The fertilizer concentration was applied following the fertilization level recommended for local maize planting.
The soil moisture was maintained at 70 % water retention capacity, and deionized water was added to maintain the soil water content throughout the entire incubation period. The vessels were incubated for 31 days at 28 °C. On days 0, 1, 2, 3, 5, 7, 9, 13, 21 and 31, a 30 g soil sample was collected from each vessel and analyzed to determine the pH and the NH4 +-N and NO3 −-N concentrations. An additional 2 g of soil from each sample were collected on incubation days 1, 9 and 31 and stored at −20 °C for soil microbial DNA extraction. Soil samples (5 g) were collected on days 1, 9 and 31 for soil microbial population analysis. Soil samples (5 g) were collected from each replicate on incubation day 9 and stored at 4 °C for soil microbial community-level physiological profile analysis.
Soil chemical and general microbial population measurements
The soil pH was determined using a combined glass electrode that was immersed in a 1:1 (w:v) ratio of soil to distilled 1 M KCl. The soil NO3 −-N concentration was determined using the phenol disulfonic acid colorimetric method, and the soil NH4 +-N concentration was determined using the KCl extraction-indophenol blue colorimetric method (Bao 2005). The cultivable bacteria, actinomycetes and fungal populations were determined by soil dilution plating on various agar media (Alef and Nannipieri 1995). Five grams of soil was weighed and added to 95 mL of sterile water, vigorously stirred for 15 min, diluted serially and plated on nutrient agar for total bacterial counts, on Martin agar for total fungal counts, and on Gause No. 1 agar for actinomycete counts.
Soil microbial DNA extraction, PCR and TRFLP analysis
Microbial genomic DNA was extracted from the soil using a Power Soil™ DNA Isolation Kit (MoBio Labs, Solana Beach, CA). Terminal restriction fragment length polymorphism (TRFLP) analysis was performed according to the modified protocol developed by Park and Noguera (2004). Briefly, the 8–27 F and 1,378–1,401 R primers (Heuer et al. 1997) were used to amplify the bacterial 16S rDNA gene, and the amoA-1F and amoA-2R primers (Rotthauwe et al. 1997) were used to amplify the AOB amoA gene. The forward primers were labeled with carboxyfluorescein (6-FAM). The amplification was performed with a master gradient thermocycler (Eppendorf, Hamburg, Germany) under the following conditions: denaturation at 95 °C for 5 min followed by 32 cycles of denaturation at 95 °C for 30 s, annealing at 56 °C for 1 min, and elongation at 72 °C for 1.5 min, with a final elongation at 72 °C for 10 min. The bacterial PCR products were digested with the HaeIII restriction endonuclease, and the amoA PCR products were digested with the TaqI restriction endonuclease (MBI Fermentas, Hanover, MD). Aliquots (1 μL) of the digested PCR products were mixed with 1 μL of GeneScan™ 600 LIZ Size Standard and 20 μL of formamide. The fluorescently labeled terminal fragments were separated by capillary electrophoresis using an ABI PRISM 3130XL Genetic Analyzer (Applied Biosystems, NJ, USA). The TRFLP profiles, which were plotted as the peak area (relative abundance) versus the fragment size, were analyzed using the Peak Scanner software (version 1.0, Applied Biosystems, NJ, USA). Peaks with an area <1 % of the total were reassigned as zero, and the proportion of each remaining peak was recalculated. An identical procedure was used for the amoA-TRFs (terminal restriction fragment size), but only amplicons between 48 and 500 bp were included because a 48-bp TRF could potentially belong to the AOB (Park and Noguera 2004).
Soil microbial community-level physiological profiles
The method described by Garland and Mills (1991) was used to determine the soil microbial community-level physiological profiles (CLPP). Briefly, 5 g of soil was added to 45 mL of sterile H2O and shaken on an orbital shaker for 30 min at 190 rpm. A 150-μL supernatant aliquot from 10−3 dilutions of each soil sample was added to each well of a Biolog EcoPlate™ (Biolog, Hayward, CA, USA). The plates were incubated at 25 °C, and the color development was measured at an absorbance of 595 nm (A595) using a microplate reader (Emax, Molecular Devices, Oxford). The A595 was measured immediately and every 24 h for 5 days. The average well color development (AWCD), the Shannon index for diversity and the richness (the number of positive wells on the EcoPlate™) were calculated based on the bacterial community after EcoPlate™ incubation for 96 h (Zak et al. 1994; Gomez et al. 2006).
Statistical analysis
A two-way analysis of variance (ANOVA) was used to determine the significant changes relative to the sampling day and the significant effects of the organic amendments on the six characteristics analyzed (soil pH, NH4 +–N concentration, NO3 −-N concentration, and cultivable bacteria, actinomycete and fungi populations). A one-way ANOVA was performed to determine the significant variations induced by the amendments during the identical incubation periods. The results were analyzed as a completely randomized design and compared using an ANOVA by Duncan’s multiple range test with SAS software (SAS Institute, Inc., Cary, NC). The soil bacterial 16S rDNA communities profiles and the ammonia-oxidizing bacteria (AOB) communities profiles under the different treatments during microcosm incubation on days 1, 9 and 31 and the soil bacterial community-level physiological profiles (CLPP) on day 9 were analyzed by a principal component analysis (PCA) using ADE-4 software (Thioulouse et al. 1997). These results were graphed with loading plots of the variables and score plots of the gravity center of each treatment at the points. Lines were drawn connecting the samples. A possible correlation between the soil chemical characteristics data and the AOB profiles was measured using Pearson’s correlation coefficient analysis.
Results
Soil pH, ammonium and nitrate concentration
According to the two-way ANOVA, the pH values in the control soil were not significantly different among all the sampling times, and the soil pH values in all the treated soil samples significantly decreased by day 5 and reached the lowest value by day 9, compared with the control soil pH, and subsequently remained stable from day 9 to day 31 (p < 0.01) (Fig. 1a). There was no significant difference among the GL, CM and FM fertilization treatments.
Changes in the soil pH value, ammonium and nitrate concentrations under different fertilization treatments during microcosm incubation. Sampling occurred on days 0, 1, 2, 3, 5, 7, 9, 13, 21 and 31. However, no data on day 9 for ammonium because of experiment error. The bars indicate the standard errors of the means (n = 3). CK, no fertilization; GL, glucose fertilization; CM, chicken manure fertilization; FM, filter mud fertilization. These treatments are the same in the following figure. a Two-way ANOVA significance levels: sampling day: p < 0.01, treatment: p < 0.01. b Two-way ANOVA significance levels: sampling day: p < 0.01, treatment: p < 0.01. c Two-way ANOVA significance levels: sampling day: p < 0.01, treatment: p < 0.01
The soil NH4 + concentrations were significantly different at the various sampling times (p < 0.01) (Fig. 1b). The NH4 + concentration of the control soil was 14.0 mg N kg−1 soil prior to incubation and increased to 18.3 mg N kg−1 soil on day 2, subsequently significantly decreased to 7.0 mg N kg−1 soil on day 7 (p < 0.05), and decreased to 6.7 mg N kg−1 soil by day 31. All the treated samples (GL, CM and FM) had their highest NH4 + concentration on day 3, and the concentration in the CM sample subsequently decreased significantly on day 5 relative to that on day 3. The concentrations in the GL and FM samples remained stable from day 3 to day 5. The concentrations in the GL sample at sampling days 7 and 13 were significantly higher than the values in the CM and FM samples. However, there were no data on day 9 for ammonium because of experiment error. After day 13 of sampling, the soil ammonium concentrations were not significantly different in the control soil and in all the treated samples.
The soil NO3 − concentration with all the treatments increased during the entire incubation period. The NO3 − concentration (Fig. 1c) of the control soil was 9.1 mg N kg−1 soil at the beginning and gradually increased to 38.9 mg N kg−1 soil by day 31. During the entire incubation period, the value of the control soil was the lowest compared with the values of the treated soils. For sampling days 2–21, except for day 3 (possibly because of an experimental error), the soil nitrate concentration in the FM-treated soil was significantly higher than the value in the GL-treated sample (Fig. 1c). The soil NO3 − concentrations on day 31 were not significantly different, and 62.8, 64.1 and 66.1 mg N kg−1 soil were recorded for the GL, CM and FM treatments, respectively.
Soil bacteria, actinomycete and fungi populations
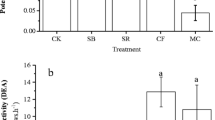
A two-way ANOVA indicated that the soil bacteria populations, as determined by the soil dilution plate method, were significantly different among the treatments (p < 0.01) and were significantly different according to the sampling day (p < 0.01) (Fig. 2a). The bacterial counts showed no differences between the GL, CM, FM soil samples on day 1; the bacterial counts of the GL sample on days 9 and 31 were significantly higher than those of the CM, FM and CK samples. The actinomycete count was significantly different among the treatments (p < 0.05) and significantly different when comparing the count among the sampling days (p < 0.01) (Fig. 2b). The soil actinomycete count of GL sample on day 1 was significantly higher than those of the CM, FM and CK sample; however, it showed no difference between the GL, CM and FM samples on day 9. The soil fungi count (Fig. 2c) was significantly different among the treatments (p < 0.01) and significantly different when comparing the count among the sampling days (p < 0.01). The average fungi counts for all the treatments were ordered as follows: day 31 > day 9 > day 1. The average fungi counts among the treatments were ordered as follows: GL > CM and FM > CK.
The bacteria (a), actinomycetes (b) and fungi (c) population in the soil under different soil treatments following microcosm incubations for 1, 9 and 31 days. The bars indicate the standard errors of the means (n = 3). On the particular sampling day, different letters on the bars indicate the values among different treatments within the same sampling day are significantly different (p < 0.05) according to Duncan’s Multiple Range Test. a Two-way ANOVA significance levels: sampling day: p < 0.01, treatment: p < 0.01. b Two-way ANOVA significance levels: sampling day: p < 0.01, treatment: p < 0.05. c Two-way ANOVA significance levels: sampling day: p < 0.01, treatment: p < 0.01
Response of the soil bacterial community to different fertilization treatments
The bacterial 16S rDNA TRFs histograms were compared among all the treatments on days 1, 9 and 31 (Fig. 3). A total of 21 TRFs were considered: 61, 64, 67, 74, 116, 120, 132, 135, 162, 177, 186, 200, 207, 212, 213, 226, 239, 253, 259, 309 and 409-bp. The 64, 207 and 259 TRFs fragments are predominant and had high relative abundance (64-bp 12 %, 207-bp 17 % and 259-bp 22 % average values in all the treatments and the sampling times). There was no obvious difference in the bacterial TRFs profiles in the CK soil at all the sampling times. There were obvious differences in the bacterial TRFs profiles among the GL, CM and FM treatments on days 1, 9 and 31. One 309-bp TRF (11 % relative abundance) was detected after the FM fertilization treatment on day 1 but not on days 9 and 31. One new 116-bp TRF was detected after the GL treatment on days 9 and 31 and after the CM treatment on day 31, but not following any FM treatment. The PCA of the bacterial TRFs profiles indicates that there is a relationship between the community composition, fertilization and incubation time (Fig. 4). The bacterial community composition for the FM treatment on day 31, with an increase in the relative abundance of 64, 74, 259 and 409-bp TRFs, had the highest first principal component (PC1) values and the lowest second principle component (PC2) values. The value for the FM treatment was clearly different from the GL and CM treatment values. The GL and CM sample values on day 31 were similar to the CM value on day 1 and to the CK sample value on day 1, 9 and 31.
The relative abundance of the soil bacterial 16S rDNA terminal restriction fragment size (TRFs) profiles under different soil treatments following microcosm incubations for 1, 9 and 31 days (n = 2). The 21 bacterial TRF fragment sizes were as follows: 61, 64, 67, 74, 116, 120, 132, 135, 162, 177, 186, 200, 207, 212, 213, 226, 239, 253, 259, 309 and 409 bp
The principal component analysis (PCA) of the soil bacterial 16S rDNA TRFs profiles from different soil treatments and incubation periods. a Loading plots of the variables of the 21 bacterial TRFs fragments were made for fragments that were 61, 64, 67, 74, 116, 120, 132, 135, 162, 177, 186, 200, 207, 212, 213, 226, 239, 253, 259, 309 and 409 bp in size under different fertilization and incubation periods, and their involvement in PC1 (X axis) and PC2 (Y axis) is plotted. b Score plots of soil samples on the basis of soil treatment and incubation periods. The small squares and circles linked by lines represent the location of each sample and their means (n = 2). CK1, CK2 and CK3 represent soil sample from no fertilization after 1, 9 and 31 days of microcosm incubation, respectively. GL1, GL2 and GL3 represent soil samples from glucose fertilizations after 1, 9 and 31 days of microcosm incubation, respectively. CM1, CM2 and CM3 represent soil samples from chicken manure fertilizations after 1, 9 and 31 days of microcosm incubation, respectively. FM1, FM2 and FM3 represent soil samples from filter mud fertilizations after 1, 9 and 31 days of microcosm incubation, respectively. The same samples are presented in Fig. 6
Response of the soil AOB community to different fertilization treatments
The amoA gene TRFs histograms were compared among all the treatments on days 1, 9 and 31 (Fig. 5). The 10 AOB TRFs fragments were as follows: 48, 54, 66, 74, 85, 96, 116, 218, 283 and 491-bp. The 54 and 283 TRFs fragments are predominant and had high relative abundance (54-bp 47 % and 283-bp 38 % average value in all the treatments and the sampling times). The amoA gene profiles from all of the treatments were similar on days 1 and 31 (Figs. 5, 6) but different on day 9. New 491-bp fragments were noted by day 9 in all the treatments. The CM sample had a high relative abundance (3 %) of 48-bp fragments on day 9, and additional 85-bp fragments were observed in the GL and CM treatment samples on day 9. The relative abundance of the 54-bp fragments decreased, but the relative abundance of the 283-bp fragments increased by day 9 in the GL, CM and FM treatment samples. The CK soil without external fertilization had a low 283-bp (Nitrosospira) fragment relative abundance (27 %) compared with the GL-treated (62 %), FM (73 %) and CM (78 %) samples The PCAs of the AOB community composition on days 1, 9 and 31, as based on the TRFLPs from the different soil treatments, are shown in Fig. 7. The AOB communities in all the treatments were clearly different, suggesting that, on day 9, the GL-treated soil had high PC1 and PC2 values and that the CM and FM-treated soils had a high PC1 value and a low PC2 value, which was different from the low PC1 value of the CK soil on day 9.
The principal component analysis (PCA) of the soil ammonia-oxidizing bacteria (AOB) TRFs profiles sample from different soil treatments and incubation periods. a Loading plots of variables of the 10 AOB TRFs fragments for the fragments that were 48, 54, 66, 74, 85, 96, 116, 218, 283 and 491 bp in size under different soil treatments and incubation periods; their involvement in PC1 (X axis) and PC2 (Y axis) is plotted. b Score plots of soil samples on the basis of soil treatments and incubation periods
The principal component analysis (PCA) of the soil bacterial community-level physiological profiles (CLPP) from different soil treatments after a 9 days microcosm incubation (n = 3). Loading plots of variables of the utilized 31 carbon resources under different soil treatments; their involvement in PC1 (X axis) and PC2 (Y axis) is plotted. b Score plots of soil samples on the basis of soil treatment after a 9-day microcosm incubation
Response of the soil bacteria community-level physiological profile to different fertilization treatments
The community-level physiological profile of the soil bacteria exposed to different fertilization treatments was measured using a Biolog EcoPlate™ (Table 2) on day 9. There was a difference in the soil bacterial functional diversity among the treatments. The AWCD among the various treatments was ranked as follows: GL > CM and FM > CK. The richness was ranked GL > CM, FM and CK. The Shannon index values were ranked as follows: GL > FM and CK > CM. The PCA values of the soil community-level physiological profiles after 9 days of treatment are shown in Fig. 7 and Table 3. The soil bacterial community substrate metabolism profiles accompanying the GL, CM and FM treatments were extremely different. The GL sample had high PC1 values (Fig. 3), with higher first eigenvector of utilization of amino acids (l-arginine, l-phenylalanine and glycyl-l-glutamic acid), carbohydrates (i-erythritol, d-mannitol, N-acetyl-d-glucosamine, glucose-1-phosphate and d-galactonic acid γ-lactone) and carboxylic acids (d-glucosaminic acid) (Table 3). The FM samples had low PC1 values with higher first eigenvector of utilization of carboxylic acids (pyruvic acid methyl ester) (Table 3). The CM samples had medium PC1 values, which were close to the values observed in the CK soil samples without fertilization.
Discussion
In the control sample with an initial pH of 4.8 and an ammonium concentration of 14.0 mg N kg−1 soil, the nitrate concentration increased and the AOB community structure measured by the TRFLP analysis remained relatively stable for the 31 incubation days. These results indicated that stimulated nitrification in the control soil did not result from AOB but possibly from AOA or heterotrophic nitrification. Similar observations that ammonia resources generated from the mineralization of organic matter stimulated nitrification have been recorded in soil incubations without external nitrogen addition (Offre et al. 2009; Gubry-Rangin et al. 2010). The low-level ammonia concentrations ranging from 0.00437 to 1.22 mM selectively favored the growth of AOA (He et al. 2012). AOA has been shown to be predominant in soil with pH 4.5 and an initial NH4 + concentration of 0.86 mg kg−1 (Gubry-Rangin et al. 2010) and in soil with pH 4.2 and an initial NH4 + concentration of 39.2 mg kg−1 (Zhang et al. 2012). The control soil is within the concentration range that stimulates AOA nitrification. The organic amendments (GL, CM and FM) soil samples added 50 mg N kg−1 soil; therefore, these samples had higher nitrate concentrations after day 7 than those of the CK samples. Their AOB community structure at sampling day 9 significantly changed and was different from that of the control soil, which indicated that autotrophic nitrification was stimulated. This finding is consistent with the hypothesis that AOB is responsible for the ammonia oxidation supplied by nitrogen fertilizer (Xia et al. 2011), despite the important role of AOA in acidic soil ecosystems (He et al. 2012). Verhamme et al. (2011) reported that soil AOA communities changed significantly at all concentrations of added ammonium (0, 20, 200 mg kg−1 soil), whereas AOB communities changed significantly only at the highest ammonium concentration.
Effect of organic amendment characteristics on soil nitrification
The FM-treated soil had the highest nitrate concentration, the CM-treated soil had a moderate nitrate concentration, and the GL-treated soil had the lowest nitrate concentration and delayed nitrate release during microcosm incubation following N fertilization. On the sampling days 2, 5, 7, 9 and 21, the soil nitrate concentrations of the GL-treated soil were significantly lower than those of the FM-treated soil. These results demonstrate that the organic amendment characteristics have a significant effect on the soil nitrifying process. The glucose delayed nitrate accumulation compared to the filter mud. Several possible theories may explain these results. The first theory hypothesized that soil bacteria typically responded rapidly and increased remarkably after simple C (glucose, etc.) amendment compared to complex C (lignin, etc.) amendment, which resulted in mineral N being assimilated as microbial cell N; thus, the autotrophic nitrification was inhibited by the soil heterotrophic bacteria because of the high competition for ammonium (Strauss and Lamberti 2002; Choi et al. 2006). The evidence in this experiment supporting this theory was that the bacteria count in the glucose soil on day 9 was the highest compared to the chicken manure and filter mud samples. Another possible theory suggested that soil heterotrophic nitrification is most likely relative to the amount of organic nitrogen and organic carbon, especially from lignin, present in acid soil (De Boer et al. 1988; De Boer and Kowalchuk 2001), Nitrification stimulation by the addition of peptone and not by the addition of ammonium salts has been proposed as evidence supporting the theory of fungal soil nitrification (Kreitinger et al. 1985; Adams 1986). Fungal heterotrophic nitrification is positively linked to lignin degradation, and the lignin-degrading basidiomycete Clitocybe metachroa has been shown to possess a fungal nitrification function (Kuyper and Bokeloh 1994; Kusel and Drake 1995; Trap et al. 2009). The third theory is that the AOA preferred to use ammonia produced by organic nitrogen mineralization and stimulated the nitrification (He et al. 2012). Further studies are needed to investigate and verify the phenomenon with additional experiments.
Effect of organic amendment characteristics on soil bacteria count dynamics
The soil bacterial populations in the GL, CM, FM and CK samples were not significantly different on day 1; and the bacterial population of the GL samples, but not those of the FM and CM samples, was significantly increased by day 9. Subsequently, the GL sample bacterial population decreased but was still higher than those of the CM and FM samples on day 31. These results are consistent with a previous study that indicated that the bacterial abundance in soil samples amended with simple C (glucose) and complex C (red clover) for a 24-h microcosm incubation did not cause measurable changes (Miller et al. 2009). Simple C (glucose) and complex C (plant residues, liquid manures or artificial root exudates) used as a soil addendum for 1 month have been shown not to change the soil bacterial abundance (Miller et al. 2008, 2009; Henry et al. 2008). Calbrix et al. (2007) reported that the bacteria population was significantly increased on days 7 and 30 following the application of sewage sludge and turkey manure with 7.6 and 2.4 % lignin contents, respectively, whereas the application of compost (31.6 % lignin content) produced from a mixture of turkey manure and lignin-like waste did not seem to alter the bacterial population. These studies provide evidence that the dynamics of bacteria population, which may have involved soil nitrification functions, were dependent on the sampling day and on the C availability of the organic amendments.
Effect of organic amendment on the soil bacterial community structure
In the TRFLP analysis, peaks with an area <1 % of the total were reassigned as zero. There were 21 TRFs fragments obtained during the treatments. The study revealed that the FM soil treatment exhibited 309-bp fragments on day 1 but not on days 9 and 31 (Figs. 3, 5). These results indicated that the 309-bp fragments that originated from the filter mud could not colonize in the microcosm soil. This finding is consistent with previous study results showing that a number of strains from an external organic amendment could not survive in soil (Innerebner et al. 2006). The high abundance of 64, 74, 259 and 409-bp fragments in the FM sample on day 31 indicated that the filter mud treatment possibly resulted in bacterial community structure changes, and these FM changes were to some extent delayed until day 31 (at the end of the incubation). The bacterial TRF profile of the GL samples with 116, 120 and 64-bp fragment abundance was high on day 9 but not on day 31, which reflects the temporary modification and quick recovery of the bacterial community profile in glucose-treated soil. The CM bacterial TRF profile did not change considerably during the incubation period.
Cytryn et al. (2011) reported that the bacterial community profile of compost-amended soils was a time-related succession from a compost-like (day 1) to an original soil-like profile (days 3–5). Subsequently, the transient presence of the Pseudomonas and Bacteroides strains becomes affiliated with the degradation of the compost-derived persistent organic compounds (days 5–13), and restoration of the baseline community structure takes longer than 50 days. This previous research and our results suggest that modifications of bacterial community structures are dependent on the quality of the organic amendment material; a higher available organic C content results in a stronger but briefer effect on the bacterial communities, while a higher complex C content results in a weaker but longer lasting effect on the bacterial communities.
Effect of organic application on soil AOB community structure
A total of 10 TRFs fragments were obtained among the treatments by the TRFLP analysis, which indicated that 283-bp fragments (identified as Nitrosospira) predominated in the studied soil. These results are in accordance with previously published evidence that Nitrosospira are the most commonly detected AOB genus in acidic soils (Kowalchuk et al. 1997; Stephen et al. 1998; Nugroho et al. 2005). These results are also consistent with the hypothesis that low pH conditions select for Nitrosospira. This finding is consistent with previous observations that Nitrosospira recovery is more efficient following MPN enrichment using a slightly acidic medium (Kowalchuk et al. 2000). Nitrosospira or Nitrosospira-like species have been shown to be predominant in 12 treatments in forest soil with a pH range of 4.12–5.20 (Long et al. 2012).
In addition, 491-bp fragments (Nitrosomonas) were noted in all the treatment samples on day 9, and the relative abundance of 491-bp fragments had a strong relationship with the soil NH4 + concentration (r = 0.05, p < 0.05) and pH (r = −0.05, p < 0.05). An analysis revealed that 48-bp fragments from an additional Nitrosomonas species were detected in the CM treatment soil on days 9 and 31. This change might occur in response to increased ammonium concentration in the latosolic red soil because Nitrosomonas are frequently detected in high N environments (Wang et al. 2009).
Effect of organic carbon application on soil bacterial activity
In this study, the soil AWCD, richness and Shannon index values were significantly higher (p < 0.05) in the GL samples than in the FM and CM samples (Table 2). The GL samples had a higher utilization of amino acids and carbohydrates (Table 3; Fig. 7), which was related to the soil C and N cycling. Soil microbes normally respond rapidly in terms of activity to the addition of readily available C (Hatch et al. 2000; Jackson et al. 2004; Carranca et al. 2009). Bengtsson et al. (2003) reported that gross nitrification rates were negatively related to ATP content. The results of these studies suggest that the nitrification inhibition in the GL-treated soil may be also related to the increasing activity of the microbial community.
The CM sample had significantly lower Shannon index values compared with the FM sample (Table 2). The soil bacterial community-level physiological profile from the CM sample was more similar to that of the control soil compared with the FM sample (Fig. 7). It is possible that this finding results from the high toxicity effect of EC or other toxin in chicken manure. The EC in the chicken manure is higher than that in the filter mud in this study. Other studies have reported that poultry manure has also a high toxicity from CO3 2− content and lower fatty acids (Materechera and Mkhabela 2008; Azeez et al. 2010). Further research is recommended to elucidate this phenomenon.
Conclusion
In this study, the soil nitrate concentration was significantly lower in the GL-treated soil than in the FM and CM-treated soil from incubation days 2 and 5–21. This finding is possibly linked to the C availability and organic N. GL greatly increased the soil bacterial population and soil bacterial functional diversity, especially from the higher utilization of carbohydrates and amino acids, which are involved in soil C and N cycling; however, the CM and FM treatments did not stimulate an increase in the soil bacterial population and function. The AOB community composition was significantly different among the treatment samples on day 9; and the CK sample had the lowest relative abundance of 283-bp fragments (Nitrosospira, 27 %), compared with the GL (62 %), FM (73 %) and CM (78 %) treatments. In addition, 48-bp fragments (another Nitrosomonas species) were detected in the CM-treated soil on days 9 and 31. These results indicate that organic carbon has a considerable and instantaneous effect on the soil AOB community. In conclusion, nitrification may have been suppressed to some extent in the GL samples, which was possibly related to the improving AOB community, bacterial community composition and activity and to increasing bacterial abundance via the available C fertilization. It is possible to regulate the dynamic soil nitrification process by optimizing the characteristics of the organic amendment used (e.g., increasing the available carbon), and this optimization may be helpful in reducing nitrogen losses in soil.
References
Adams JA (1986) Nitrification and ammonification in acid forest litter and humus as affected by peptone and ammonium-N amendment. Soil Biol Biochem 18:45–51
Alef K, Nannipieri P (1995) Methods in applied soil microbiology and biochemistry. Academic Press, London
Avrahami S, Liesack W, Conrad R (2003) Effects of temperature and fertilizer on activity and community structure of soil ammonia oxidizers. Environ Microbiol 5:691–705
Azeez JO, Van Averbeke W, Okorogbona AOM (2010) Differential responses in yield of pumpkin (Cucurbita maxima L.) and nightshade (Solanum retroflexum Dun.) to the application of three animal manures. Bioresour Technol 101:2499–2505
Bao SD (2005) Soil and agricultural chemistry analysis. Chinese Agricultural Publishing House, Beijing
Bastida F, Pérez-de-Mora A, Babic K, Hai B, Hernández T, García C, Schloter M (2009) Role of amendments on N cycling in Mediterranean abandoned semiarid soils. Appl Soil Ecol 41:195–205
Belser LW (1979) Population ecology of nitrifying bacteria. Annu Rev Microbiol 33:309–333
Bengtsson G, Bengtson P, Månsson KF (2003) Gross nitrogen mineralization-, immobilization-, and nitrification rates as a function of soil C/N ratio and microbial activity. Soil Biol Biochem 35:143–154
Cai YF, Peng C, Zhao SQ, Cai AH, Li YT (2012) Balanced fertilization improved soil ammonia-oxidizing bacterial community in latosolic red soil. Acta Agric Scand Sect B Soil Plant Sci 62:531–540
Calbrix R, Barray S, Chabrerie O, Fourrie L, Laval K (2007) Impact of organic amendments on the dynamics of soil microbial biomass and bacterial communities in cultivated land. Appl Soil Ecol 35:511–522
Carranca C, Oliveira A, Pampulha ME, Torres MO (2009) Temporal dynamics of soil nitrogen, carbon and microbial activity in conservative and disturbed fields amended with mature white lupine and oat residues. Geoderma 151:50–59
Chantigny MH, Angers DA, Rochette P (2002) Fate of carbon and nitrogen from animal manure and crop residues in wet and cold soils. Soil Biol Biochem 34:509–517
Choi WJ, Lee SM, Han GH, Yoon KS, Jung JW, Lim SS, Kwak JH (2006) Available organic carbon controls nitrification and immobilization of ammonium in an acid loam-textured soil. Agric Chem Biotechnol 49:28–32
Chu H, Fujii T, Morimoto S, Lin X, Yagi K (2008) Population size and specific nitrification potential of soil ammonia-oxidizing bacteria under long-term fertilizer management. Soil Biol Biochem 40:1960–1963
Cytryn E, Kautsky L, Ofek M, Mandelbaum RT, Minz D (2011) Short-term structure and functional changes in bacterial community composition following amendment with biosolids compost. Appl Soil Ecol 48:160–167
De Boer W, Kowalchuk GA (2001) Nitrification in acid soils: micro-organisms and mechanisms. Soil Biol Biochem 33:853–866
De Boer W, Duyts H, Laanbroek HJ (1988) Autotrophic nitrification in a fertilized heath soil. Soil Biol Biochem 20:845–850
Deng SP, Tabatabai MA (1997) Effect of tillage and residue management on enzyme activities in soils III: phosphatases and arylsulfatase. Biol Fertil Soils 24:141–146
Dong LH, Córdova-Kreylos AL, Yang JS, Yuan HL, Scow KM (2009) Humic acids buffer the effects of urea on soil ammonia oxidizers and potential nitrification. Soil Biol Biochem 41:1612–1621
Elsayed MT, Babiker MH, Abdelmalik ME, Mukhtar ON, Montange D (2008) Impact of filter mud applications on the germination of sugarcane and small-seeded plants and on soil and sugarcane nitrogen contents. Bioresour Technol 99:4164–4168
Garcia C, Hernandez T, Costa F, Ceccanti B (1994) Biochemical parameters in soil regenerated by the addition of organic wastes. Waste Manag Res 12:457–466
Garland JL, Mills AL (1991) Classification and characterization of heterotrophic microbial communities on the basis of patterns of community-level sole-carbon-source utilization. Appl Environ Microbiol 57:2351–2359
Gomez E, Ferreras L, Toresani S (2006) Soil bacterial functional diversity as influenced by organic amendment application. Bioresour Technol 97:1484–1489
Gubry-Rangin C, Nicol GW, Prosser JI (2010) Archaea rather than bacteria control nitrification in two agricultural acidic soils. FEMS Microbiol Ecol 74:566–574
Gundersen P, Callesen I, de Vries W (1998) Nitrate leaching in forest ecosystems is controlled to forest floor C/N ratios. Environ Pollut 102:403–407
Hatch DJ, Bhogal A, Lovell RD, Shepherd MA, Jarvis SC (2000) Comparison of different methodologies for field measurement of net nitrogen mineralization in pasture soils under different soil conditions. Biol Fertil Soils 32:287–293
He JZ, Shen JP, Zhang LM, Zhu YG, Zheng YM, Xu MG, Di HJ (2007) Quantitative analyses of the abundance and composition of ammonia-oxidizing bacteria and ammonia-oxidizing archaea of a Chinese upland red soil under long-term fertilization practices. Environ Microbiol 9:2364–2374
He JZ, Hu HW, Zhang LM (2012) Current insights into the autotrophic thaumarchaeal ammonia oxidation in acidic soils. Soil Biol Biochem 55:146–154
Henry S, Texier S, Hallet S, Bru D, Dambreville C, Chèneby D, Bizouard F, Germon JC, Philippot L (2008) Disentangling the rhizosphere effect on nitrate reducers and denitrifiers: insight into the role of root exudates. Environ Microbiol 10:3082–3092
Heuer H, Krsek M, Baker P, Smalla K, Wellington EMH (1997) Analysis of actinomycete communities by specific amplification of genes encoding 16S rRNA and gel-electrophoretic separation in denaturing gradients. Appl Environ Microbiol 63:3233–3241
Holbeck B, Amelung W, Wolf A, Sudekum KH, Schloter M, Welp G (2013) Recoveries of 15N-labelled fertilizers (chicken manure, mushroom compost and potassium nitrate) in arable topsoil after autumn application to winter cover crops. Soil Tillage Res 130:120–127
Innerebner G, Knapp B, Vasara T, Romantschuk M, Insam H (2006) Traceability of ammonia-oxidizing bacteria in compost-treated soils. Soil Biol Biochem 38:1092–1100
Jackson LE, Ramirez I, Yokota R, Fennimore SA, Koike ST, Henderson DM, Chaney WE, Caldéron FJ, Klonsky K (2004) On-farm assessment of organic matter and tillage management on vegetable yield, soil, weeds, pests, and economics in California. Agric Ecosyst Environ 103:443–463
Kowalchuk GA, Stephen JR (2001) Ammonia-oxidizing bacteria: a model for molecular microbial ecology. Annu Rev Microbiol 55:485–529
Kowalchuk GA, Stephen JR, De Boer W, Prosser JI, Embley TM, Woldendorp JW (1997) Analysis of β-proteobacteria ammonia oxidizing bacteria in coastal sand dunes using denaturing gradient gel electrophoresis and sequencing of PCR amplified 16S rDNA fragments. Appl Environ Microbiol 63:1489–1497
Kowalchuk GA, Stienstra AW, Heilig GHJ, Stephen JR, Woldendorp JW (2000) Molecular analysis of ammonia-oxidising bacteria in soil of successional grasslands of the Drentsche A (The Netherlands). FEMS Microbiol Ecol 31:207–215
Kreitinger JP, Klein TM, Novick NJ, Alexander M (1985) Nitrification and characteristics of nitrifying microorganisms in an acid forest soil. Soil Sci Soc Am J 49:1407–1410
Kusel K, Drake HL (1995) Effects of environmental parameters on the formation and turnover of acetate by forest soils. Appl Environ Microbiol 61:3667–3675
Kuyper TW, Bokeloh DJ (1994) Ligninolysis and nitrification in vitro by a nitrotolerant and a nitrophobic decomposer basidiomycete. Oikos 70:417–420
Laanbroek HJ, Woldendorp JW (1995) Activity of chemolithotrophic nitrifying bacteria under stress in natural soils. In: Jones JG (ed) Advances in microbial ecology. Plenum Press, New York, pp 275–304
Lehtovirta-Morley LE, Stoecker K, Vilcinskas A, Prosser JI, Nicol GW (2011) Cultivation of an obligate acidophilic ammonia oxidizer from a nitrifying acid soil. Proc Natl Acad Sci USA 108:15892–15897
Li XX, Ying JY, Chen Y, Zhang LM, Gao YS, Bai YF (2011) Effects of nitrogen addition on the abundance and composition of soil ammonia oxidizers in Inner Mongolia Grassland. Acta Ecol Sin 31:174–178
Long X, Chen C, Xu Z, Oren R, He JZ (2012) Abundance and community structure of ammonia-oxidizing bacteria and archaea in a temperate forest ecosystem under ten-years elevated CO2. Soil Biol Biochem 46:163–171
Materechera SA, Mkhabela TS (2008) The effectiveness of lime, chicken manure and leaf litter ash in ameliorating acidity in a soil previously under black wattle (Acacia mearnsii) plantation. Bioresour Technol 85:9–16
Miller MN, Zebarth BJ, Dandie CE, Burton DL, Goyer C, Trevors JT (2008) Crop residue influence on denitrification, N2O emissions and denitrifier community abundance in soil. Soil Biol Biochem 40:2553–2562
Miller MN, Zebarth BJ, Dandie CE, Burton DL, Goyer C, Trevors JT (2009) Influence of liquid manure on soil denitrifier abundance, denitrification and N2O emissions. Soil Sci Soc Am J 73:760–768
Nugroho RA, Röling WFM, Laverman AM, Zoomer HR, Verhoef HA (2005) Presence of Nitrosospira cluster 2 bacteria corresponds to N transformation rates in nine acid Scots pine forest soils. FEMS Microbiol Ecol 53:473–481
Offre P, Prosser JI, Nicol GW (2009) Growth of ammonia-oxidizing archaea in soil microcosms is inhibited by acetylene. FEMS Microbiol Ecol 70:99–108
Park HD, Noguera DR (2004) Evaluating the effect of dissolved oxygen on ammonia-oxidizing bacterial communities in activated sludge. Water Res 38:3275–3286
Pascual JA, Hernandez T, Ayuso M, Garcia C (1997) Changes in the microbial activity of arid soils amended with urban organic wastes. Biol Fertil Soils 24:429–434
Priha O, Smolander A (1999) Nitrogen transformations in soil under Pinus sylvestris, Picea abies and Betula pendula at two forest sites. Soil Biol Biochem 31:965–977
Racz L, Datta T, Goel R (2010) Effect of organic carbon on ammonia oxidizing bacteria in a mixed culture. Bioresour Technol 101:6454–6460
Rivero C, Chirenje T, Ma LQ, Martinez G (2004) Influence of compost on soil organic matter quality under tropical conditions. Geoderma 123:355–361
Rotthauwe JH, Witzel KP, Liesack W (1997) The ammonia monooxygenase structural gene amoA as a functional marker: molecular fine-scale analysis of natural ammonia-oxidizing populations. Appl Environ Microbiol 63:704–712
Schaffers AP, Sykora KV (2000) Reliability of Ellenberg indicator values for moisture, nitrogen and soil reaction: a comparison with field measurements. J Veg Sci 11:225–244
Stephen JR, Kowalchuk GA, Bruns MAV, McCaig AE, Phillips CJ, Embley TM, Prosser JI (1998) Analysis of β-subgroup proteobacterial ammonia oxidizer populations in soil by denaturing gradient gel electrophoresis analysis and hierarchical phylogenic probing. Appl Environ Microbiol 64:2958–2965
Strauss EA, Lamberti GA (2002) Effect of dissolved organic carbon quality on microbial decomposition and nitrification rates in stream sediments. Freshw Biol 47:65–74
Thioulouse J, Chessel D, Doledec S, Olivier JM (1997) ADE-4: a multivariate analysis and graphical display software. Stat Comput 7:75–83
Tourna M, Freitag TE, Nicol GW, Prosser JI (2008) Growth, activity and temperature responses of ammonia-oxidizing archaea and bacteria in soil microcosms. Environ Microbiol 10:1357–1364
Trap J, Bureau F, Vinceslas-Akpa M, Chevalier R, Aubert M (2009) Changes in soil N mineralization and nitrification pathways along a mixed forest chronosequence. For Ecol Manag 258:1284–1292
Treusch AH, Leininger S, Kietzin A, Schuster SC, Klenk HP, Schleper C (2005) Novel genes for nitrite reductase and amo-related proteins indicate a role of uncultivated mesophilic crenarchaeota in nitrogen cycling. Environ Microbiol 7:1985–1995
Van Soest PJ, Wine RH (1967) Use of detergents in the analysis of fibrous feeds. IV. Determination of cell-wall constituents. J AOAC 50:50–55
Venter JC, Remington K, Heidelberg JF, Halpern AL, Rusch D, Eisen JA (2004) Environmental genome shotgun sequencing of the Sargasso Sea. Science 304:66–74
Verhamme DT, Prosser JI, Nicol GW (2011) Ammonia concentration determines differential growth of ammonia-oxidising archaea and bacteria in soil microcosms. ISME J 5:1067–1071
Wang Y, Ke X, Wu L, Lu Y (2009) Community composition of ammonia-oxidizing bacteria and archaea in rice field soil as affected by nitrogen fertilization. Syst Appl Microbiol 32:27–36
Xia WW, Zhang CX, Zeng XW, Feng YZ, Weng JH, Lin XG, Zhu JG, Xiong ZQ, Xu J, Cai ZC, Jia ZJ (2011) Autotrophic growth of nitrifying community in an agricultural soil. ISME J 5:1226–1236
Zak JC, Willig MR, Moorhead DL, Wildman HG (1994) Functional diversity of microbial communities: a quantitative approach. Soil Biol Biochem 26:1101–1108
Zhang LM, Hu HW, Shen JP, He JZ (2012) Ammonia-oxidizing archaea have more important role than ammonia-oxidizing bacteria in ammonia oxidation of strongly acidic soils. ISME J 6:1032–1045
Acknowledgments
This research was supported by the National Natural Science Foundation of China (Nos. 41171209, 41171210, 41240007, and 41071176), the Science and Technology Research Project of Guangdong Province (2011B020308014), and the Team Project of the Natural Science Foundation of Guangdong Province (S2011030002882 and S2012010009023).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Zhao, X., Cai, Y., Peng, C. et al. Regulation of nitrification in latosolic red soils by organic amendment. Environ Earth Sci 71, 3865–3878 (2014). https://doi.org/10.1007/s12665-013-2773-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12665-013-2773-5