Abstract
Biochar and compost are seen as two attractive waste management options and are used for soil amendment and pollution remediation. The interaction between biochar and composting may improve the potential benefits of biochar and compost. We investigated soil physicochemical properties, bacterial community, bacterial 16S rRNA, and functional marker genes of nitrogen cycling of the soil remedied with nothing (S), compost (SC), biochar (SB), a mixture of compost and biochar (SBC), composted biochar (SBced), and a composted mixture of biochar and biomass (SBCing). The results were that all amendments (1) increased the bacterial community richness (except SB) and SBCing showed the greatest efficiency; (2) increased the bacterial community diversity (SBCing > SBC > SC > SBced > SB > S); and (3) changed the gene copy numbers of 16S rRNA, nirK, nirS, and nosZ genes of bacteria, ammonia-oxidizing archaea (AOA), and ammonia-oxidizing bacteria (AOB). All amendments (except SB) could increase the gene copy number of 16S rRNA, and SBCing had the greatest efficiency. The changes of soil bacterial community richness and diversity and the gene copy numbers of 16S rRNA, nirK, nirS, nosZ, AOA, and AOB would affect carbon and nitrogen cycling of the ecosystem and also implied that BCing had the greatest efficiency on soil amendment.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Biochar and compost not only are identified as two attractive waste management options and have less negative impacts on the environment (Fan et al. 2008; Tang et al. 2008; Gong et al. 2009; Beesley et al. 2011; Park et al. 2011; Zeng et al. 2013a, 2015) but also generally decrease soil pollution bioavailability and mobility and increase soil organic matter, nutrient availability, microbial activity, and crop yields (Schmidt et al. 2011; Hale et al. 2013; Masiello et al. 2013; Thangarajan et al. 2013; Lu et al. 2014a). Biochar, produced by pyrolysis of biomass under low oxygen conditions, has caught more and more attention as a soil amendment material. Compost is the final product of composting, which is an aerobic bio-decomposition process of organic waste and reduces waste volume by 40 to 50 % (Zeng et al. 2007; Lu et al. 2014b).
In order to maximize the potential benefits of biochar and composting, the interaction between biochar and composting was a research hotspot in recent years (Wei et al. 2014; Wu et al. 2016) and also should be the focus of further study (Karami et al. 2011; Zeng et al. 2015). Biochar could obviously affect the microbial community and physicochemical process during composting as well as the quality and composition of the end product (Dias et al. 2010; Steiner et al. 2011; Jindo et al. 2012). The surface of biochar is changed during the composting process due to abiotic and biotic oxidation and sorption of organic compounds derived from compost (Prost et al. 2013).
Many studies have been conducted on the influences of biochar and compost (or composting) combined amendments (Karami et al. 2011; Xu et al. 2012; Suddick and Six 2013; Beesley et al. 2014; Borchard et al. 2014; Schmidt et al. 2014; Zhu et al. 2014; Mackie et al. 2015; Zeng et al. 2015), but these studies have primarily been focused on the pollution restoration, gas emission, enzymes, and crop growth in soil. However, little is known about the effect of biochar and compost combined amendments (especially composted biochar (Bced, without compost) and a composted mixture of biochar and biomass (SBCing)) on the soil biological properties, particularly the bacterial community structure. Besides, there is also lack of information about the ecological risk of the use of biochar and compost combined amendments in soil amendment and pollution remediation. Soil microorganisms are critical to global ecosystem because of their significant role in regulating global nutrients and carbon cycling via fundamental ecological processes such as mineralization and decomposition (An et al. 2004; Zhang et al. 2007; Hartman et al. 2008; Huang et al. 2008; Hu et al. 2011; Wu et al. 2013). The bacterial community structure is sensitive to local changes in the environmental conditions and anthropogenic activity and serves as a sensitive indicator of environmental changes (Card and Quideau 2010; Wu et al. 2015; Dai et al. 2016). Therefore, the biochar and compost combined amendments will impact soil bacterial community structure as well as nutrient and carbon cycling.
In this study, we used the bacterial community structure and functional marker genes of nitrogen cycling as indicators to investigate the ecological responses of soil microorganisms to the biochar, compost, and combined amendments. The objectives of this study were as follows: (1) to analyze the responses of the soil bacterial community and functional marker genes of nitrogen cycling to biochar, compost, and combined amendments and (2) to determine the ecological risk of using of biochar, compost, and combined amendments in soil amendment and pollution remediation.
Materials and methods
Soil and amendment materials
Soil (clay 24.19 %, silt 45.54 %, and sand 30.27 %) was sourced from beach of the Dongting Lake (28° 30′–29° 38′ N, 112° 18′–113° 15′ E) wetland (Zeng et al. 2015). Dongting Lake, located in the middle reach of Changjiang River (Yangtze River) region, is the second largest fresh lake in China and an important wintering habitat and pathway for East Asian migratory birds (Wu et al. 2013, 2015). The soil was air-dried, sieved to a particle size of <2 mm, and biological debris was removed (Zeng et al. 2015).
Biochar was produced from corn cob according to a former study (Zeng et al. 2015). Compost was produced from rice straw. BC was made of the mixture of compost and biochar with the weight ratio of 1:1 (w/w). Bced was produced as follows: biochar was placed into mesh (0.125 mm) bags and then composted with rice straw, and the bags filled with biochar were taken out after composting was completed (Zeng et al. 2015). BCing was produced as follows: biochar and rice straw were mixed (w/w: 1:1) and placed into the mesh bags and then composted with rice straw according to the above method (Fig. 1) (Zeng et al. 2015). Chemical properties of these amendment materials are mentioned in our previous study (Zeng et al. 2015).
Experimental design and procedure
The soil and amendment materials were thoroughly mixed in the following proportions, which constituted the treatments.
-
S: 500 g soil in each pot
-
SB: 25 g biochar and 500 g soil in each pot
-
SC: 25 g compost and 500 g soil in each pot
-
SBC: 25 g BC and 500 g soil, in each pot
-
SBced: 25 g Bced and 500 g soil in each pot
-
SBCing: 25 g BCing and 500 g soil in each pot
Finally, mixtures of soil and amendment material were placed into 1000-mL pots, and the top of soil in each pot was covered with 50 g fresh soil from the above sampling site to introduce indigenous microbes (they could enter the amended soil through their own mobility and reproduce or by the mobility of soil particles and pore water) and lower the impacts of air-drying, sieving, and mixing, which could have generated additional modifications/disturbances (Wu et al. 2015). Each treatment was conducted in triplicate. Deionized water was added to the soil, so that the fixed moisture was at 60 % water-filled pore space. These pots were then placed in a controlled environment chamber with 28 % relative humidity and at 25 °C for 60 days (Zeng et al. 2015). The moisture of the soil in each pot was checked weekly to maintain the moisture. At day 60, the soil was collected for analyses of soil physicochemical properties and soil microbial properties.
Determination of soil physicochemical properties
The soil total organic carbon (TOC) was determined by the loss-on-ignition method with ashing at 500 °C for 4 h (Wright et al. 2008). Water-extractable organic carbon (WEOC) was obtained according to the method described in our previous report (Zeng et al. 2015). The soil pH was examined using a digital pH meter (for water/soil ratio of 2.5:1). The soil moisture was measured from the mass loss upon drying at 105 °C overnight (Wu et al. 2013). Ammonium nitrogen content (NH+ 4-N) was examined by indophenol blue colorimetry method, and nitrate nitrogen content (NO− 3-N) was determined by phenol disulfonic acid spectrophotometric method (Lu 1999).
High-throughput sequencing and qPCR
DNA was extracted from 0.8 g of soil using a method which has been previously described (Yang et al. 2007). The DNA was divided into two equal parts. One part was used in high-throughput sequencing (the triplicate of each treatment was mixed) and the other part was used in qPCR. The DNA for high-throughput sequencing was sequenced on an Illumina Miseq PE300 platform (Illumina, San Diego, USA). The 338F and 806R, which were the V3 regions of the bacterial-specific fragment 16S rRNA, were chosen as primers (Dennis et al. 2013). Data of high-throughput sequencing were analyzed as follows: (a) any ambiguous reads or low-quality reads were removed; (b) taxonomy of the qualified sequence reads (using RDP classifier, at a 97 % identity threshold) were assigned; (c) singletons and putative chimeras were removed; and (d) diversity and richness indices (Shannon-Weaver index (H), chao1 estimator, and ACE estimator) were calculated to assess the internal (within-sample) complexity of individual microbial populations (Xu et al. 2014).
Gene nirK, nirS, and nosZ served as denitrifying bacteria gene markers. Gene ammonia-oxidizing archaea (AOA) and ammonia-oxidizing bacteria (AOB) served as nitrification functional markers. The qPCR of these genes was completed using an iCycler IQ5 Thermocycler (Bio-Rad, USA). The reactants of qPCR contained 10 μL of 2× SYBR Real-time PCR Premixture (Bioteke, Beijing), 0.2 μM of primer, 1 μL of template DNA, and 8.6 μL of sterile water. PCR conditions and primers used for qPCR are listed in the Supporting Information Table S1. Tenfold serial dilutions of linearized plasmids containing each gene were used to create the standard curves for qPCR (Wu et al. 2015). The initial copy number of each gene was calculated by comparing the threshold cycle values with the standard curve (Wu et al. 2015).
Data analysis
The multivariate relationships, between the bacterial community structure and the soil properties, were determined using Canoco (version 4.5, Centre for Biometry, Wageningen, Netherlands). Detrended correspondence analysis (DCA) was finished. The length of the first DCA ordination axis was 0.970, which clearly indicated a linear species response. Accordingly, redundancy analysis (RDA) with default settings was performed to ordinate the soil bacterial community structure with the soil properties (Lepš and Šmilauer 2003). The distances between centroid points (representative of individual samples) of ordination biplots indicated their level of bacterial community structure similarity (Wu et al. 2013). In the forward selections, a Monte Carlo permutation test with 499 unrestricted permutations was carried out to determine the parameters that significantly affected the bacterial community structure (Lepš and Šmilauer 2003).
A Pearson correlation analysis was completed to test the univariate correlations between each parameter. The mean values among each treatment were compared by one-way analysis of variance (ANOVA), where the post hoc Tukey’s test was used to examine the significant differences (p < 0.05) in the data. All of the analyses were completed using SPSS (version 19).
Accession numbers
Sequence data of high-throughput sequencing have been deposited (accession number SRX1434824) in the NCBI.
Results
Responses of soil properties to amendments
Soil properties of each treatment are shown in Table 1. There was no significant difference among the pH of SB, SBC, and S. The pH of SC, SBced, and SBCing were lower than that of S, and that of SBCing was the lowest one. Comparing with the WEOC of S, SB had no significant change, and others caused an obvious increase (SBCing had the biggest increase). The NH+ 4-N contents of SC and SBCing were significantly higher than that of S. However, there was no significant difference between the NH+ 4-N contents of SB, SBC, and SBced and that of S. The NO− 3-N content of SB was similar with that of S, and the NO− 3-N content of others (SC > SBCing > SBC > SBced) was obviously higher than that of S.
Responses of soil bacterial community to amendments
After demultiplexing and quality filtering, 72,246 high-quality sequences were obtained in total. The ACE estimator and the Chao1 estimator (Table 2) showed that all amendments increased the bacterial community richness (the Chao1 estimator showed that SB decreased the bacterial community richness). SBCing had the greatest efficiency of increasing the bacterial community richness, while SB had the lowest efficiency of that. The Shannon index (Table 2) showed that all amendments increased the bacterial community diversity, and the rule of the samples’ bacterial community diversity was SBCing > SBC > SC > SBced > SB > S. The samples’ bacterial community richness and diversity implied that BCing had the greatest efficiency on soil amendment. The coverage of each sample was above 0.98 (Table 2), which indicated that this study could uncover the true changes of the soil bacterial community. The hierarchical clustering based on OUT information (Fig. 2) was also finished. As presented by the cluster analysis, the samples of SBCing, SBC, and SC were closer to each other and the samples of S, SBced, and SB were closer to each other.
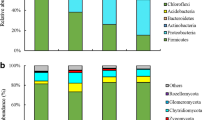
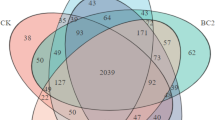
The most abundant phyla were Proteobacteria, Actinobacteria, Acidobacteria, Bacteroidetes, Gemmatimonadetes, Chloroflexi, Verrucomicrobia, and Candidate division TM7, and these taxa accounted for more than 95 % of the bacterial sequences (Fig. 3). All amendments decreased the relative abundance of Proteobacteria from 47.7 to 38.6–46.3 %. The relative abundance of Actinobacteria was highest in SB (31.8 %) and was lowest in SBC (11.5 %). On the contrary, the relative abundance of Bacteroidetes was lowest in SB (5.8 %) and was highest in SBC (21.4 %). The relative abundance of Acidobacteria of SBced (16.7 %) was higher than that of S (16.2 %) and that of others (10.4–13.0 %) was lower than that of S. Similar to the Acidobacteria, the relative abundance of Gemmatimonadetes of SBced (8.1 %) was higher than that of S (7.4 %) and that of others (3.3–5.8 %) was lower than that of S. The relative abundances of Chloroflexi, Verrucomicrobia, and Candidate division TM7 showed no obvious changes under amendment.
Those genera, which have a relative abundance above 1.5 % in at least two samples, were selected for analysis. As shown in Table 3, at the genus level the relative abundances of Lysobacter, Subgroup 6 norank, and Thermomonas of S were higher than 10 %. All amendment decreased their abundances. The relative abundances of Tamlana and Parapedobacter of S, SB, and SBced were 0, while that of amendments with material containing composted products (SC, SBC, and SBCing) were 1.4–5.0 and 0.9–5.0 %, respectively. The relative abundances of Luteimonas of S, SB, and SBced were 0, while that of amendments with material contain composted products (SC, SBC, and SBCing) was 1.5–2.6 %. The relative abundance of Aquamicrobium of S, SB, and SBced was 0, while that of amendments with material containing composted products (SC, SBC, and SBCing) was 1.1–2.2 %.
The RDA biplot of the soil bacterial community structure and the investigated soil properties are shown in Fig. 4. All soil properties could explain 71.7 % of the variation in the species data (Supporting Information Table S2). The community variation explained by the soil properties decreased as follows: NH+ 4-N > NO− 3-N > WEOC > pH. Moreover, NH+ 4-N, NO− 3-N, and WEOC could be used to explain 43.6 % of the variation in the species data. As shown in Supporting Information Table S3, the ACE estimator, Chao1 estimator, and Shannon index were strongly correlated with WEOC (p < 0.05). The Shannon index was correlated with (not strongly) NH+ 4-N (p = 0.06) and NO− 3-N (p = 0.06). All of the aforementioned results suggested that NH+ 4-N, WEOC, and NO− 3-N were the dominant influence factors of the soil bacterial community structure.
Redundancy analysis for bacterial community structure and soil properties. Samples of soil with addition of nothing (S), compost (SC), biochar (SB), mixture of biochar and compost (SBC), composted biochar (SBced), and a composted mixture of biochar and biomass (SBCing) are indicated by open circles. Soil properties are represented by solid lines with filled arrows
Responses of 16S rRNA and functional marker genes to amendments
Gene copy numbers of bacteria 16S rRNA, nirK, nirS, nosZ, AOA, and AOB are shown in Fig. 5. There were significant differences among each treatment. The gene copy numbers of 16S rRNA of S and SB were similar. However, other amendments obviously increased the gene copy number and SBCing had the greatest efficiency, which implied that BCing had the greatest efficiency on soil amendment. Besides, Pearson’s correlation coefficient showed that the gene copy number of 16S rRNA was strongly correlated with WEOC but not correlated with TOC. Similar to 16S rRNA, the gene copy numbers of AOA of S, SC, and SB were similar and other amendments obviously increased the gene copy number. The gene copy numbers of AOB of SB and SBCing were lower than those of S, and those of SC, SBC, and SBced were obviously higher than those of S. The gene copy numbers of nirS of SC, SB, and SBC were lower than those of S, and those of SBced and SBCing were higher than those of S obviously. All amendments increased the gene copy numbers of nirK and SBced had the greatest efficiency. On the contrary, compared to nirK, all amendments decreased the gene copy numbers of nosZ and SB had the greatest efficiency.
Gene copy numbers of bacterial 16S rRNA, AOA, AOB, nirK, nirS, and nosZ in the soil with addition of nothing (S), compost (SC), biochar (SB), mixture of biochar and compost (SBC), composted biochar (SBced), and a composted mixture of biochar and biomass (SBCing). Different letters of the same indicator indicate significant difference (p < 0.05) between each treatment
Discussion
Responses of soil properties to amendments
Compost addition, whether alone (SC) or combined with biochar (SBC, SBced, and SBCing), led to the reduction of soil pH. This is because of humic acids isolated from organic materials of compost (Zeng et al. 2015). Other studies also found that Bced reduced the pH of neutral soil while biochar did not change the pH (Zeng et al. 2013b; Borchard et al. 2014). The soil WEOC with biochar increased less than that of other amendments, because carbon pool of biochar is relatively stable and insoluble (Beesley et al. 2014; Zeng et al. 2015). Other studies also found that biochar caused an obvious change in the concentration of WEOC (Clemente et al. 2012; Jones et al. 2012). The changes of TOC and WEOC caused by combined amendments (SBC, SBced, and SBCing) were the results of combined impact of compost and biochar (Zeng et al. 2015). Compost could provide nutrients (such as NH+ 4-N and NO− 3-N) to soil, and biochar could adsorb nutrients from soil. Moreover, the changes of NH+ 4-N and NO− 3-N were the results of combined impacts of them.
Responses of soil bacterial community to amendments
The results of this study showed that all the amendments could increase the bacterial community richness (the Chao1 estimator showed that SB decreased the bacterial community richness) and SBCing had the greatest efficiency while SB had the lowest efficiency. This was because all amendment materials provided nutrients for soil bacterial growth, but most of biochar was unlikely used by soil microbes (Karami et al. 2011; Beesley et al. 2014; Xu et al. 2014; Zeng et al. 2015). Due to the same reason, all amendments increased the bacterial community diversity, and the rule of the samples’ bacterial community diversity was SBCing > SBC > SC > SBced > SB > S. Xu et al. also found that biochar could not cause the obvious increase in soil bacterial community diversity (Xu et al. 2014). The soil bacterial communities of some treatments (such as S, SBced, and SB) were similar. It was maybe because of their similar nutrient content and ratio (such as C/N, which was found to be significant in shaping the soil microbial community composition) (Xu et al. 2014).
The genera information from the present study also supported this view, such as the following: (1) The relative abundances of Tamlana and Parapedobacter of S, SB, and SBced were 0, while those of SC, SBC, and SBCing were 1.4–5.0 and 0.9–5.0 %, respectively; (2) the relative abundances of Luteimonas of S, SB, and SBced were 0, while those of SC, SBC, and SBCing were 1.5–2.6 %; and (3) the relative abundances of Aquamicrobium of S, SB, and SBced were 0, while those of SC, SBC, and SBCing were 1.1–2.2 %. All above illustrated that amendments could increase the bacterial community diversity; soil bacterial communities of SBCing, SBC, and SC were closer to each other; and the soil bacterial communities of S, SBced, and SB were closer to each other.
Our results also showed that NH+ 4-N, WEOC, and NO− 3-N were the dominant factors that influenced the soil bacterial community structure. This may be because these soil properties can regulate the nutrient content in the soil, which further affect the soil bacterial community structure (Wu et al. 2013, 2015). We also found that the ACE estimator, Chao1 estimator, and Shannon index were strongly correlated with WEOC, which suggested that soil bacterial abundance and diversity were strongly correlated with WEOC. However, other studies argued that soil microbial biomass and bacterial community diversity were strongly correlated with TOC (Wu et al. 2013, 2015). This difference was because in most of biochar, carbon pool of biochar was relatively insoluble and stable, unlike what was used by soil microbes (Karami et al. 2011; Beesley et al. 2014; Xu et al. 2014; Zeng et al. 2015).
Responses of 16S rRNA and functional marker genes to amendments
This study showed that the gene copy numbers of 16S rRNA of S and SB were similar. It means that the bacterial abundances at the DNA level of S and SB were similar. This result was in accordance with other studies (Harter et al. 2014; Xu et al. 2014). Other amendments obviously increased the gene copy number, and SBCing had the greatest efficiency. It means that other amendments could obviously increase the bacterial abundance and SBCing had the greatest efficiency, which was the result of these amendment materials providing nutrients for soil bacterial growth. The change in the bacterial abundance determined the bacterial metabolic potential, which was consistent with the change in the soil CO2 efflux (Poret-Peterson et al. 2007; Li et al. 2010; Wu et al. 2013). The change in the gene copy numbers of 16S rRNA would affect carbon exchange throughout the ecosystem. Further study is needed to determine how the carbon exchange of the entire ecosystem would be affected. However, in general, such a large increase in the gene copy numbers of 16S rRNA would reduce the carbon storage of the wetland.
The gene copy numbers of AOA, AOB, nirK, nirS, and nosZ could indicate the activity and abundance of nitrifiers and denitrifiers. The results of this study demonstrated the significant differences of the gene copy numbers of these functional marker genes between each treatment. Part of the result was in agreement with the previous studies. For example, Xu et al. (2014) found that biochar could not cause obvious changes in the gene copy numbers of AOA, nirK, and nirS. The significant changes in the gene copy numbers of these functional marker genes in other treatments could affect the gas (e.g., N2O, NO, and N2) emissions and nitrogen cycling of the ecosystem. Further study is needed to determine how the N2O emission and nitrogen cycling of the entire ecosystem would be affected.
References
An Y-J, Joo Y-H, Hong L-Y, Ryu H-W, Cho K-S (2004) Microbial characterization of toluene-degrading denitrifying consortia obtained from terrestrial and marine ecosystems. Appl Microbiol Biotechnol 65:611–619. doi:10.1007/s00253-004-1596-4
Beesley L, Moreno-Jiménez E, Gomez-Eyles JL, Harris E, Robinson B, Sizmur T (2011) A review of biochars’ potential role in the remediation, revegetation and restoration of contaminated soils. Environ Pollut 159:3269–3282. doi:10.1016/j.envpol.2011.07.023
Beesley L, Inneh OS, Norton GJ, Moreno-Jimenez E, Pardo T, Clemente R, Dawson JJ (2014) Assessing the influence of compost and biochar amendments on the mobility and toxicity of metals and arsenic in a naturally contaminated mine soil. Environ Pollut 186:195–202. doi:10.1016/j.envpol.2013.11.026
Borchard N, Spokas K, Prost K, Siemens J (2014) Greenhouse gas production in mixtures of soil with composted and noncomposted biochars is governed by char-associated organic compounds. J Environ Qual 43:971. doi:10.2134/jeq2013.07.0290
Card SM, Quideau SA (2010) Microbial community structure in restored riparian soils of the Canadian prairie pothole region. Soil Biol Biochem 42:1463–1471. doi:10.1016/j.soilbio.2010.05.010
Clemente R, Walker DJ, Pardo T, Martinez-Fernandez D, Bernal MP (2012) The use of a halophytic plant species and organic amendments for the remediation of a trace elements-contaminated soil under semi-arid conditions. J Hazard Mater 223–224:63–71. doi:10.1016/j.jhazmat.2012.04.048
Dai J, Wu H, Zhang C, Zeng G, Liang J, Guo S, Li X, Huang L, Lu L, Yuan Y (2016) Responses of soil microbial biomass and bacterial community structure to closed-off management (an ecological natural restoration measures): a case study of Dongting Lake wetland, middle China. J Biosci Bioeng. doi:10.1016/j.jbiosc.2016.03.001
Dennis KL, Wang Y, Blatner NR, Wang S, Saadalla A, Trudeau E, Roers A, Weaver CT, Lee JJ, Gilbert JA, Chang EB, Khazaie K (2013) Adenomatous polyps are driven by microbe-instigated focal inflammation and are controlled by IL-10–producing T cells. Cancer Res 73:5905–5913
Dias BO, Silva CA, Higashikawa FS, Roig A, Sanchez-Monedero MA (2010) Use of biochar as bulking agent for the composting of poultry manure: effect on organic matter degradation and humification. Bioresource Technol 101:1239–1246
Fan T, Liu Y, Feng B, Zeng G, Yang C, Zhou M, Zhou H, Tan Z, Wang X (2008) Biosorption of cadmium(II), zinc(II) and lead(II) by Penicillium simplicissimum: isotherms, kinetics and thermodynamics. J Hazard Mater 160:655–661
Gong JL, Wang B, Zeng GM, Yang CP, Niu CG, Niu QY, Zhou WJ, Liang Y (2009) Removal of cationic dyes from aqueous solution using magnetic multi-wall carbon nanotube nanocomposite as adsorbent. J Hazard Mater 164:1517–1522. doi:10.1016/j.jhazmat.2008.09.072
Hale SE, Jensen J, Jakob L, Oleszczuk P, Hartnik T, Henriksen T, Okkenhaug G, Martinsen V, Cornelissen G (2013) Short-term effect of the soil amendments activated carbon, biochar, and ferric oxyhydroxide on bacteria and invertebrates. Environ Sci Technol 47:8674–8683. doi:10.1021/es400917g
Harter J, Krause HM, Schuettler S, Ruser R, Fromme M, Scholten T, Kappler A, Behrens S (2014) Linking N2O emissions from biochar-amended soil to the structure and function of the N-cycling microbial community. ISME J 8:660–674
Hartman WH, Richardson CJ, Vilgalys R, Bruland GL (2008) Environmental and anthropogenic controls over bacterial communities in wetland soils. Proc Natl Acad Sci U S A 105:17842–17847. doi:10.1073/pnas.0808254105
Hu X, Wang J, Liu Y, Li X, Zeng G, Bao Z, Zeng X, Chen A, Long F (2011) Adsorption of chromium (VI) by ethylenediamine-modified cross-linked magnetic chitosan resin: Isotherms, kinetics and thermodynamics. J Hazard Mater 185:306–314
Huang D, Zeng G, Feng C, Hu S, Jiang X, Tang L, Su F, Zhang Y, Zeng W, Liu H (2008) Degradation of lead-contaminated lignocellulosic waste by Phanerochaete chrysosporium and the reduction of lead toxicity. Environ Sci Technol 42(13):4946–4951
Jindo K, Sánchez-Monedero MA, Hernández T, García C, Furukawa T, Matsumoto K, Sonoki T, Bastida F (2012) Biochar influences the microbial community structure during manure composting with agricultural wastes. Sci Total Environ 416:476–481
Jones DL, Rousk J, Edwards-Jones G, DeLuca TH, Murphy DV (2012) Biochar-mediated changes in soil quality and plant growth in a three year field trial. Soil Biol Biochem 45:113–124. doi:10.1016/j.soilbio.2011.10.012
Karami N, Clemente R, Moreno-Jimenez E, Lepp NW, Beesley L (2011) Efficiency of green waste compost and biochar soil amendments for reducing lead and copper mobility and uptake to ryegrass. J Hazard Mater 191:41–48. doi:10.1016/j.jhazmat.2011.04.025
Lepš J, Šmilauer P (2003) Multivariate analysis of ecological data using CANOCO. Cambridge University Press, Cambridge
Li Y-L, Wang L, Zhang W-Q, Zhang S-P, Wang H-L, Fu X-H, Le Y-Q (2010) Variability of soil carbon sequestration capability and microbial activity of different types of salt marsh soils at Chongming Dongtan. Ecol Eng 36:1754–1760. doi:10.1016/j.ecoleng.2010.07.029
Lu R (1999) Soil agricultural chemical analysis method. China’s agricultural science and technology press, Beijing
Lu L, Huggins T, Jin S, Zuo Y, Ren ZJ (2014a) Microbial metabolism and community structure in response to bioelectrochemically enhanced remediation of petroleum hydrocarbon-contaminated soil. Environ Sci Technol 48:4021–4029. doi:10.1021/es4057906
Lu L, Zeng G, Fan C, Zhang J, Chen A, Chen M, Jiang M, Yuan Y, Wu H, Lai M, He Y (2014b) Diversity of two-domain laccase-like multicopper oxidase genes in Streptomyces spp.: identification of genes potentially involved in extracellular activities and lignocellulose degradation during composting of agricultural waste. Appl Environ Microbiol 80:3305–3314. doi:10.1128/aem.00223-14
Mackie KA, Marhan S, Ditterich F, Schmidt HP, Kandeler E (2015) The effects of biochar and compost amendments on copper immobilization and soil microorganisms in a temperate vineyard. Agric Ecosyst Environ 201:58–69. doi:10.1016/j.agee.2014.12.001
Masiello CA, Chen Y, Gao X, Liu S, Cheng HY, Bennett MR, Rudgers JA, Wagner DS, Zygourakis K, Silberg JJ (2013) Biochar and microbial signaling: production conditions determine effects on microbial communication. Environ Sci Technol 47:11496–11503. doi:10.1021/es401458s
Park JH, Lamb D, Paneerselvam P, Choppala G, Bolan N, Chung J-W (2011) Role of organic amendments on enhanced bioremediation of heavy metal(loid) contaminated soils. J Hazard Mater 185:549–574. doi:10.1016/j.jhazmat.2010.09.082
Poret-Peterson AT, Ji B, Engelhaupt E, Gulledge J (2007) Soil microbial biomass along a hydrologic gradient in a subsiding coastal bottomland forest: implications for future subsidence and sea-level rise. Soil Biol Biochem 39:641–645. doi:10.1016/j.soilbio.2006.09.016
Prost K, Borchard N, Siemens J, Kautz T, Sequaris JM, Moller A, Amelung W (2013) Biochar affected by composting with farmyard manure. J Environ Qual 42:164–172. doi:10.2134/jeq2012.0064
Schmidt MW, Torn MS, Abiven S, Dittmar T, Guggenberger G, Janssens IA, Kleber M, Kögel-Knabner I, Lehmann J, Manning DA, Nannipieri P, Rasse DP, Weiner S, Trumbore SE (2011) Persistence of soil organic matter as an ecosystem property. Nature 478:49–56. doi:10.1038/nature10386
Schmidt H-P, Kammann C, Niggli C, Evangelou MWH, Mackie KA, Abiven S (2014) Biochar and biochar-compost as soil amendments to a vineyard soil: influences on plant growth, nutrient uptake, plant health and grape quality. Agric Ecosyst Environ 191:117–123
Steiner C, Melear N, Harris K, Das KC (2011) Biochar as bulking agent for poultry litter composting. Carbon Manag 2:227–230
Suddick EC, Six J (2013) An estimation of annual nitrous oxide emissions and soil quality following the amendment of high temperature walnut shell biochar and compost to a small scale vegetable crop rotation. Sci Total Environ 465:298–307. doi:10.1016/j.scitotenv.2013.01.094
Tang L, Zeng G, Shen G, Li Y, Zhang Y, Huang D (2008) Rapid detection of picloram in agricultural field samples using a disposable immunomembrane-based electrochemical sensor. Environ Sci Technol 42(4):1207–1212
Thangarajan R, Bolan NS, Tian G, Naidu R, Kunhikrishnan A (2013) Role of organic amendment application on greenhouse gas emission from soil. Sci Total Environ 465:72–96. doi:10.1016/j.scitotenv.2013.01.031
Wei L, Shutao W, Jin Z, Tong X (2014) Biochar influences the microbial community structure during tomato stalk composting with chicken manure. Bioresource Technol 154:148–154. doi:10.1016/j.biortech.2013.12.022
Wright AL, Wang Y, Reddy KR (2008) Loss-on-ignition method to assess soil organic carbon in calcareous everglades wetlands. Commun Soil Sci Plan 39:3074–3083. doi:10.1080/00103620802432931
Wu H, Zeng G, Liang J, Zhang J, Cai Q, Huang L, Li X, Zhu H, Hu C, Shen S (2013) Changes of soil microbial biomass and bacterial community structure in Dongting Lake: impacts of 50,000 dams of Yangtze River. Ecol Eng 57:72–78. doi:10.1016/j.ecoleng.2013.04.038
Wu H, Zeng G, Liang J, Guo S, Dai J, Lu L, Wei Z, Xu P, Li F, Yuan Y, He X (2015) Effect of early dry season induced by the three gorges Dam on the soil microbial biomass and bacterial community structure in the Dongting Lake wetland. Ecol Indic 53:129–136. doi:10.1016/j.ecolind.2015.01.041
Wu H, Lai C, Zeng G, Liang J, Chen J, Xu J, Dai J, Li X, Liu J, Chen M, Lu L, Hu L, Wan J (2016) Interaction of composting and biochar and its implication for soil amendment and pollution remediation—a review. Crit Rev Biotechnol (Accepted on Apr 01, 2016)
Xu P, Zeng GM, Huang DL, Feng CL, Hu S, Zhao MH, Lai C, Wei Z, Huang C, Xie GX, Liu ZF (2012) Use of iron oxide nanomaterials in wastewater treatment: a review. Sci Total Environ 424:1–10. doi:10.1016/j.scitotenv.2012.02.023
Xu HJ, Wang XH, Li H, Yao HY, Su JQ, Zhu YG (2014) Biochar impacts soil microbial community composition and nitrogen cycling in an acidic soil planted with rape. Environ Sci Technol 48:9391–9399. doi:10.1021/es5021058
Yang ZH, Xiao Y, Zeng GM, Xu ZY, Liu Y (2007) Comparison of methods for total community DNA extraction and purification from compost. Appl Microbiol Biotechnol 74:918–925. doi:10.1007/s00253-006-0704-z
Zeng G, Huang D, Huang G, Hu T, Jiang X, Feng C, Chen Y, Tang L, Liu H (2007) Composting of lead-contaminated solid waste with inocula of white-rot fungus. Bioresource Technol 98:320–326. doi:10.1016/j.biortech.2006.01.001
Zeng G, Chen M, Zeng Z (2013a) Risks of neonicotinoid pesticide. Science 340(6139):1403
Zeng G, Chen M, Zeng Z (2013b) Shale gas: surface water also at risk. Nature 499:154. doi:10.1038/499154d
Zeng G, Wu H, Liang J, Guo S, Huang L, Xu P, Liu Y, Yuan Y, He X, He Y (2015) Efficiency of biochar and compost (or composting) combined amendments for reducing Cd, Cu, Zn and Pb bioavailability, mobility and ecological risk in wetland soil. RSC Adv 5:34541–34548. doi:10.1039/c5ra04834f
Zhang Y, Zeng G, Tang L, Huang D, Jiang X, Chen Y (2007) A hydroquinone biosensor based on immobilizing laccase to modified core-shell magnetic nanoparticles supported on carbon paste electrode. Biosens Bioelectron 22:2121–2126
Zhu K, Christel W, Bruun S, Jensen LS (2014) The different effects of applying fresh, composted or charred manure on soil N2O emissions. Soil Biol Biochem 74:61–69. doi:10.1016/j.soilbio.2014.02.020
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Funding
This research was financially supported by the CRSRI Open Research Program (CKWV2015203/KY), the National Natural Science Foundation of China (51521006, 51479072, 51378190, 51509014, 51579094, and 51508177), the New Century Excellent Researcher Award Program from the Ministry of Education of China (NCET-08-0181), and the Program for Changjiang Scholars and Innovative Research Team in University (IRT-13R17).
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors. The article is an original paper, is not under consideration by another journal, and has not been published previously. All authors read and approved the final manuscript.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 217 kb)
Rights and permissions
About this article
Cite this article
Wu, H., Zeng, G., Liang, J. et al. Responses of bacterial community and functional marker genes of nitrogen cycling to biochar, compost and combined amendments in soil. Appl Microbiol Biotechnol 100, 8583–8591 (2016). https://doi.org/10.1007/s00253-016-7614-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-016-7614-5