Abstract
Breast cancer is among the major frequent types of cancer worldwide, causing a significant death rate every year. It is the second most prevalent malignancy in Egypt. With the increasing number of new cases, it is vital to diagnose breast cancer in its early phases to avoid serious complications and deaths. Therefore, routine screening is important. With the current evolution of deep learning, medical imaging became one of the interesting fields. The purpose of the current work is to suggest a hybrid framework for both the classification and segmentation of breast scans. The framework consists of two phases, namely the classification phase and the segmentation phase. In the classification phase, five different CNN architectures via transfer learning, namely MobileNet, MobileNetV2, NasNetMobile, VGG16, and VGG19, are applied. Aquila optimizer is used for the calculation of the optimal hyperparameters of the different TL architectures. Four different datasets representing four different modalities (i.e., MRI, Mammographic, Ultrasound images, and Histopathology slides) are used for training purposes. The framework can perform both binary- and multi-class classification. In the segmentation phase, five different structures, namely U-Net, Swin U-Net, Attention U-Net, U-Net++, and V-Net, are applied to identify the region of interest in the ultrasound breast images. The reported results prove the efficiency of the suggested framework against current state-of-the-art studies.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Breast cancer affects a huge bulk of women yearly around the world, and it causes fatalities among women. As per the World Health Organization (WHO), breast cancer is an extremely popular sort of cancer worldwide in 2020 (Organization WH 2022) as indicated in Fig. 1. The survival rates vary among different countries, from 80% in North America, 60% in Japan and Sweden, to 40% in low-income nations (Masud et al. 2020). The rates of occurrence shown in Fig. 2 and mortality shown in Fig. 3 differ per country, depending on many circumstances including the environment, the availableness of modern medical care, socioeconomic levels, and so on Francies et al. (2020). As shown in Fig. 4, the mortality rates in nations with a bigger “low to middle” income population are rising every year due to the inability to obtain profitable resources. Several affluent countries, such as Australia, are also seeing an increase in the number of cases. Therefore, raising awareness about breast cancer and encouraging women to be screened is critical because early detection and diagnosis can save lives (Zuluaga-Gomez et al. 2021). As shown in Fig. 5, Egypt is one of the top countries having new cases of breast cancer in Africa in 2020. Breast cancer is the second most prevalent malignancy in Egypt in 2020 as shown in Fig. 6.
Approximate number of novel patients of different types of cancer in 2020 (Organization WH 2022)
Approximate number of novel breast cancer patients distributed by continents in 2020 (Organization WH 2022)
Approximate number of deaths due to breast cancer distributed by continents in 2020 (Organization WH 2022)
Approximate number of deaths due to breast cancer distributed by income in 2020 (Organization WH 2022)
Approximate number of novel breast cancer patients distributed by countries in Africa in 2020 (Organization WH 2022)
Approximate number of novel cancer patients distributed by type in Egypt in 2020 (Organization WH 2022)
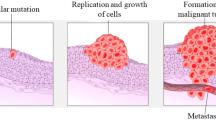
Breast cancer is a disease in which the cells of the breast uncontrollably multiply (For Disease Control 2022). The main elements of the breast are the (1) ducts, (2) lobules, and (3) connective tissues. A duct is a tube that transports breast milk to the nipple. Lobules are the glands responsible for making milk. Connective tissues consist of fiber and fatty tissues and connect the entire components of the breast (Lawrence 2022). Breast cancer is more commonly found in the lobules or the ducts of the breast (Charishma et al. 2020). It begins in the breast tissue. Like other malignant tumors, they can enter and spread to the tissues around the breast. It may also propagate to other organs of the body, leading to the formation of additional tumors, a process called metastasis (Clinic 2022). It is vital to keep in mind that the predominant of breast lumps are not cancerous (i.e., malignant). Non-cancerous breast tumors are irregular masses that remain locally in the breast. Benign breast lipomas are rarely dangerous. However, they do increase the risk of breast cancer in women (Society 2022).
Symptoms of breast cancer can vary according to the affected patient. The majority of people are completely oblivious to any indicators (Melekoodappattu et al. 2022). The frequently obvious sign of breast cancer is a new tumor or mass in the breast tissue (Bakker et al. 2019). A lump in the breast or armpit is the most prevalent symptom. Skin changes, soreness, a nipple that pushes inward, and unusual discharge from the nipple are among the other symptoms (Benson et al. 2020). The risks of acquiring breast cancer rise with age. Every year, more than 80% of women with breast cancer are above 45 years old, with around 43% of women being 65 years old or older (Duffy et al. 2020).
Mammography is the gold standard for routine screening. It is critical to examine the screening data and deliver a diagnosis as correctly and fast as possible after collecting it (Zuluaga-Gomez et al. 2021; Melekoodappattu and Subbian 2020). Experts use mammography and ultrasound pictures to discover malignancies, which necessitate the use of specialist radiologists (Indra and Manikandan 2021). The most commonly used characteristics, such as shape, texture, density, and other characteristics, are characteristics manually configured according to the experience of the physician, that is, subjective characteristics. Although the traditional diagnosis approach is widely utilized, its accuracy can still be improved (Wang et al. 2019). Consequently, computer-aided diagnosis systems (CADs) are now widely employed to assist radiologists in making decisions when diagnosing malignancies (Ahmed et al. 2020). CAD systems can reduce radiologists’ workload and decrease the amount of false-positive and false-negative diagnoses (Elter and Horsch 2009).
With deep learning’s exceptional performance in detecting and recognizing visual items, as well as other applications, deep learning techniques to aid radiologists providing increased accuracy of interpreting mammographic scans have piqued people’s curiosity (Kim et al. 2018; Hamidinekoo et al. 2018). According to recent research, deep learning-based CAD systems perform the same goes for radiation in the standalone mode and even improve radiologists’ performance in the assisted mode (Shen et al. 2019). The Convolutional Neural Network (CNN) is a deep learning algorithm frequently applied in solving challenging problems. It is a representative learning algorithm that can automatically extract meaningful information from the original image without manually designing function descriptors (Khan et al. 2020). It solves the drawbacks of classic machine learning techniques. Traditional machine learning algorithms necessitate feature extraction, which necessitates the assistance of a domain expert (Zheng et al. 2014). In addition, choosing the right function for a specific situation is a daunting task. However, deep learning technology solves the feature selection problem by automatically extracting relevant features from the original input without the need for pre-selected features (Indolia et al. 2018). Due to recent performance improvements in image segmentation, detection, and classification, CNN has been successfully applied in medical imaging challenges (Sohail et al. 2021).
In the present work, a novel hybrid framework for the segmentation and classification of breast cancer images is proposed. The framework is composed of two phases, namely the classification phase and the segmentation phase. In the classification phase, the model is used to classify breast images into two categories (i.e., benign or malignant). To train the framework, four different datasets representing different modalities (i.e., MRI, Mammographic, Ultrasound images, and Histopathology slides) are used. The variety of data types ensures that the model can be used with all image types. Each of these datasets is classified into a different number of classes, hence the framework can perform both binary- and multi-class classification. Five pre-trained CNN architectures, namely MobileNet, MobileNetV2, NasNetMobile, VGG16, and VGG19, are used in the classification phase. To refine the performance of the different models, Aquila Optimizer (AO) is used to tune the hyperparameters of the different CNN architectures. During the segmentation phase, five different segmentation models are used, namely U-Net, Swin U-Net, Attention U-Net, U-Net++, and V-Net, to identify the region of interest in the ultrasound breast images.
1.1 Paper contributions
The key contributions of the current study are:
-
Proposing a novel hybrid framework for classification and segmentation of breast cancer images.
-
Using four different datasets for training purposes.
-
The proposed model can be used for MRI, Mammographic, Ultrasound images, and Histopathology slides.
-
The use of five pretrained CNN architectures for breast image classification.
-
AO is used to tune the hyperparameters of the different CNN architectures.
-
Using five models for the segmentation of ultrasound breast cancer images.
1.2 Paper organization
The remaining of the article is divided into six sections. Section 2 gives a survey of the current studies about the use of CNN for detecting breast cancer and the different segmentation techniques. Section 3 presents background about the necessary techniques used in the proposed framework. Section 4 explains in detail the proposed framework for the classification and segmentation of breast cancer images while section 5 gives the experimental results and their discussions. Section 6 is the conclusion, limitations of the current study, and trends for future work.
2 Related studies
This section presents a state-of-the-art survey about the use of CNN in the diagnosis of breast cancer. Then, a survey about the different segmentation techniques applied to breast cancer images is presented.
2.1 Related studies using CNN
Melekoodappattu et al. (2022) developed a system for diagnosing breast cancer using CNN and image texture attribute extraction. They could achieve accuracies of 98% and 97.9% on the MIAS and DDSM repositories, respectively. Wang et al. (2021) proposed a boosted EfficientNet CNN architecture for automatically detecting cancer cells in breast cancer pathology tissue as a solution to low image resolution. Sharma and Kumar (2021) created a deep learning system to identify breast cancer using histopathology photographs. They used the DenseNet201 CNN model for extracting features. Malignant and benign classification tasks are the two categories of classification tasks. Salama and Aly (2021) used images from three different datasets, namely Digital Database for Screening Mammography (DDSM), Mammographic Image Analysis Society (MIAS), and the Curated Breast Imaging Subset of DDSM (CBIS-DDSM). These images are classified as benign and malignant using various models such as DenseNet121, InceptionV3, VGG16, ResNet50, and MobileNetV2. The best-achieved accuracy is 88.87% using InceptionV3 with data augmentation. Chorianopoulos et al. (2020) applied three CNN models, namely MobileNet, VGG16, and AlexNet on two different datasets i.e., ultrasounds and histopathological images. The best accuracy was 96.82% achieved by VGG16 on the ultrasounds dataset. MobileNet achieved the best accuracy of 91.04% on the Invasive Ductal Carcinoma dataset. Hameed et al. (2020) used four distinct CNN models based on the pre-trained VGG16 and VGG19 structures, namely VGG16 Fully Trained, VGG16 Refined, VGG19 Fully Trained, and VGG19 Refined trained to classify histopathological images of noncancerous and cancerous breast cancers using their collected dataset. They found that the best accuracy was 92% achieved by VGG19 Fine Tuned model.
Dabeer et al. (2019) used CNN to identify breast cancer cells into benign or malignant classes. They obtained an accuracy of 99.86%. Alghodhaifi et al. (2019) experimented with two CNN models using depthwise separable convolution (IDCDNet) and standard convolution (IDCNet). Several types of activation functions were investigated, including Sigmoid, TanH, and ReLU. The best achieved accuracy was 87.13% achieved by standard convolution (IDCNet) with ReLU activation function. Saikia et al. (2019) compared multiple fine-tuned transfer learning classification approaches based on CNN to diagnose cell samples. Their suggested method was examined on a dataset containing 212 images of which 113 images are malignant. This dataset was extended to 2120 images of which 1130 images are malignant. Four CNN architectures, namely ResNet50, VGG16, VGG19, and GoogLeNetV3, were used in training. The Fine-tuned GoogLeNetV3 achieved the best accuracy of 96.25%. Ismail et al. (2019) compared the identification of breast cancer using two deep learning model networks, namely VGG16 and ResNet50, and applied the models on the IRMA dataset for classifying benign and malignant tumors. In terms of accuracy, VGG16 outperforms ResNet50 with a score of 94% compared to 91.7% for ResNet50. Mehra (2018) used three popular CNN models, namely ResNet50, VGG16, and VGG19, for both full training and forward learning for classifying histological images of breast cancer. Their best accuracy was 92.60% achieved by VGG16 with Logistic Regression (LR). Gao et al. (2018) proposed a Shallow-Deep CNN to classify patients as benign or cancer from mammography images. The shallow CNN is used to find the recombined images from low-energy images, while a deep CNN is applied to extract the unique features from these images. Their best-achieved accuracy was 90% using their proposed technique.
2.2 Related studies using segmentation
Salama and Aly (2021) used a modified U-Net model for segmentation of breast cancerous area from mammographic images. There proposed model could do both segmentation and classification with an accuracy of 98.87% using InceptionV3 plus a modified U-Net model with data augmentation. Byra et al. (2020) proposed a deep learning method using U-Net for breast mass segmentation in ultrasonography. They achieved an overall accuracy of 97.6%. El Adoui et al. (2019) created two CNN using SegNet and U-Net to suggest two deep learning algorithms for automatic segmentation of breast tumors in dynamic contrast-enhanced magnetic resonance imaging. The SegNet architecture achieved a mean intersection over union (i.e., accuracy) of 68.88%, whereas the U-Net architecture achieved 76.14%. Li et al. (2019) used the segmentation of masses for enhancing the accuracy of diagnosing breast cancer and lowering the mortality rate because breast mass is one of the most characteristic markers for the diagnosis of breast cancer. In their experiments, they used U-Net, attention U-Net, and DenseNet for segmentation and could achieve accuracy of 74.37%, 74.83%, and 77.93%, respectively. Alom et al. (2018) applied the Recurrent Residual U-Net for Nuclei segmentation from high-resolution histopathology images to extract the fine features from nuclear morphometrics. They could achieve a dice coefficient of 92.15% segmentation accuracy. Dalmia et al. (2018) compared the accuracy of different algorithms, namely VGGNet, U-Net, and V-Net. They could prove that a larger dataset combined with parameter adjustment would allow the model to generalize to previously unseen examples more efficiently, resulting in better training and validation outcomes. They could achieve an accuracy of 81.6%, 99.5%, and 99.6% for the different models respectively.
3 Background
This section presents background about the techniques used in the proposed framework.
3.1 Convolutional Neural Networks (CNN)
CNN is a class of deep learning used in handling image data (Bingli et al. 2021). It is inspired by the visual cortex in animals (Jogin et al. 2018). It is designed for automatic and adaptive learning of structures, hierarchical and spatial characteristics, and low-level to high-level patterns (Balaha et al. 2021).
3.1.1 CNN layers
CNN is usually made up of three types of layers: convolution layer, pooling layer, and fully connected layer. The earlier two layers (i.e. the convolutional and pooling layers) extract features. On the other hand, the last layer (i.e., fully connected layer) maps the extracted objects to the final output space (Balaha et al. 2021).
3.1.2 Parameters optimization
The choice of the right optimization method and the efficient tuning of the hyperparameters strongly influences the training speed and the final performance of the learned model (Zhang et al. 2021). The current study uses the Adam, AdaGrad, NAdam, AdaDElta, AdaMax, RMSProp, and SGD optimizers. Adaptive Moment Estimation (Adam) Optimizer effective when dealing with a huge problem containing multiple parameters (Kingma and Ba 2014). AdaGrad Optimizer adjusts the learning ratio based on the settings, making smaller updates for settings related to common features and huge updates for settings related to non-features regularly (Luo et al. 2019). Nesterov Adaptive Momentum (NAdam) calculates the velocity before the gradient (Dozat 2016) AdaDelta Optimizer extends AdaGrad as a trial to decrease the rate of excessive and monotonous learning rather than assembling the entire past squared gradients (Dogo et al. 2018). AdaMax Optimizer represents the updated version of Adam (Vani and Rao 2019). RMSProp Optimizer was proposed simultaneously to address Adagrad’s plummeting learning rate. RMSprop is the same as Adadelta’s first update vector (Wu et al. 2016). Stochastic Gradient Descent (SGD) Optimizer is a repetitive technique used for the optimization of the objective function with appropriate regularity properties. Otherwise, it updates the parameter for each input (Bottou 2012).
3.2 Aquila Optimizer (AO)
Aquila’s behavior in the wild while capturing victims is the main inspiration for the AO algorithm. Therefore, the optimization methods of the AO algorithm are presented in 4 methods. The first method is to select a search area by navigating up with vertical tilt. The second method is to explore inside a disparate search area by contour flight with a small glide attack. The third method is to explore inside a convergent search area by low-level flight with a sinking attack for slow prays. The fourth method is walking and grabbing the victim (AlRassas et al. 2021). The selection between the four methods is done based on specific parameters.
3.3 Image segmentation
Image segmentation is one class of digital image processing including splitting an image into various parts on the basis of the image’s properties and qualities (Singh and Singh 2010). The fundamental reason behind image segmentation is to simplify the image for ease in analysis (Norouzi et al. 2014). In diagnosing patients with cancer, the form of cancer cells is important in determining the severity of the cancer disease. The use of image segmentation technologies has had a significant impact in this area so that cancer cells can be correctly and accurately identified (Senthil Kumar et al. 2019).
Segmentation using U-Net Model: U-Net model was created for biological-image segmentation (Ronneberger et al. 2015). The U-Net architecture is essentially a network of encoders followed by a network of decoders (Habijan et al. 2019). Segmentation using Swin U-Net Model: The use of transformers (Vaswani et al. 2017) has extended from natural language processing (NLP) tasks to vision-related and segmentation tasks. Swin U-Net (Cao et al. 2021) implants pure transformer structure into the U-Net architecture for segmentation tasks. Segmentation using Attention U-Net Model: is almost built upon the well-known U-Net (Vaswani et al. 2017). The network consists of a reduced path for extracting features of locality and an extension path for resampling the image map using contextual information (Abraham and Khan 2019). Segmentation using U-Net++ Model: U-Net++ is a general-purpose image segmentation architecture that tries to address the shortcomings of U-Net (Zhou et al. 2019). The U-Net++ is made up of multiple U-Nets of different depths, with the decoders firmly coupled at the same resolution via a revised skip connection (Lu et al. 2021). Segmentation using V-Net Model: The V-Net method consists of two main parts, i.e. left and right sections. The left section contains the compressed path and is divided into various stages that operate at other resolutions with each stage having 1 to 3 convolution layers. On the other hand, the right section compresses the input until an initial size is reached (Abdollahi et al. 2020).
3.4 Performance metrics
All learning algorithms require a metric to evaluate performance (Balaha et al. 2021b). The most commonly used performance metrics are TN, TP, FN, FP, Accuracy, Recall (Sensitivity), Precision, F1-score, Specificity, AUC (Area under the curve), IoU Coefficient, Dice Coefficient, Cosine similarity, Hinge, and SquaredHinge (Balaha and Saafan 2021; Abdulazeem et al. 2021). These metrics are: TN is the rightly estimated negative values so that the actual class value is false and the estimated class value is also false. TP is the rightly estimated positive values so that the actual class value is true and the estimated class value is also true. FN is the actual class value is true but the estimated class value is false. FP is the actual class value is false and the estimated class value is true.
Accuracy is the ratio of rightly estimated observations to overall observations. Precision is the ratio of the rightly estimated positive observations to the overall estimated positive observations. Sensitivity (or Recall) is the ratio of rightly estimated positive observations to overall observations. F1-score is the weighted ratio of Recall and Precision. Specificity is the number of cases identified as negative from all the real negative cases. Area Under Curve (AUC) is the area under the Receiver Operating Characteristics Curve (ROC Curve). Intersection over Union (IoU) is also known as the Jaccard index. Dice Coefficient is a measure of similarity of the objects. Cosine Similarity is a similarity measure using Euclidean distance. Hinge Loss is a similarity measure using the loss function. Squared Hinge Loss is a square of the output of the hinge.
4 Methodology
The suggested approach is trained on four different types from four different modalities for versatility. This is important to guarantee the robustness of the model for all types of images. Each type of dataset has a variable number of classes. For this reason, the proposed framework can perform both binary and multi-class classification. The purpose of the current work is to suggest a novel hybrid framework for the classification and segmentation of breast cancer images. The phases of the proposed framework are presented in Fig. 7. The phases of the proposed framework are described in the next subsections.
4.1 Datasets acquisition phase
In the current work, four different datasets with different modalities (i.e., MRI, mammographic, Ultrasound images, and histopathology slides) are used to train the models. Magnetic Resonance Imaging (MRI) is recommended when soft tissue imaging is required. So, it is used for expose lesioned regions (Yurttakal et al. 2020). On the other hand, Mammography is the most commonly used technique for breast cancer diagnosis. It is an accurate technique that uses low-dose X-Ray to display the inner texture of the breast (Maitra et al. 2012). The ultrasound image is preferred due to many advantages including low cost and acceptable accuracy (Feng et al. 2017). On the other hand, the biopsy is defined as the process of extracting a sample or portion of a mass in the human body, usually called the biopsy sample, for further examination (Preetha and Jinny 2021). Histopathology means to analyze the biopsy sample by the specialist, usually called the pathologist. Therefore, histopathology images are microscopic images of the tissues of masses taken from the human body (Aswathy and Jagannath 2017).
The first used dataset is “Breast Cancer Patients MRI’s” from Kaggle which can be retrieved from https://www.kaggle.com/uzairkhan45/breast-cancer-patients-mris. This dataset contains 1,480 MRI images classified into Healthy (Benign) and Sick (Malignant). The second dataset is “Breast Cancer Dataset” from Kaggle which can be retrieved from https://www.kaggle.com/anaselmasry/breast-cancer-dataset. This dataset contains histopathology slides. The third dataset is Dataset_BCD_mammography_images_out downloaded from Kaggle from https://www.kaggle.com/anwarsalem/dataset-bcd-mammography-images-out. This dataset consists of 8 classes representing the severity of breast cancer. The fourth dataset is Breast Ultrasound Images Dataset (Al-Dhabyani et al. 2020) containing ultrasound images of 600 female patients with 780 images classified into normal, benign, and malignant. It can be downloaded from https://www.kaggle.com/aryashah2k/breast-ultrasound-images-dataset. Table 1 presents a summary of the datasets used in the current study. Samples from the acquired datasets are displayed in Fig. 8.
4.2 Pre-processing phase
The used datasets are not found with the same size and hence the datasets are resized equally to the size of (100, 100, 3) for classification and (256, 256, 3) for segmentation in RGB mode. The present work uses 4 distinct scaling techniques. They are (1) normalization, (2) standardization, (3) min–max scaler, and (4) max–abs scaler. The equations for them are shown in Eqs. 1 to 4 respectively where \(X_{input}\) is the input image, \(X_{scaled}\) is the scaled output image, \(\mu\) is the mean of the input image, \(\sigma\) is the standard deviation of the input image. The used datasets are not balanced as shown in Table 1. To overcome this problem, data balancing using data augmentation approach is employed. The present work uses rotation, shifting, shearing, zooming, flipping, and brightness changing augmentation techniques. Table 2 shows the different augmentation techniques and the corresponding configurations.
4.3 Segmentation phase
Segmentation is important to label the area of the tumor (i.e., the region of interest) to facilitate the diagnosis for the physician. Hence, the first processing phase of the framework is to apply segmentation. In the segmentation phase, five different segmentation models (i.e., U-Net Ronneberger et al. 2015, Swin U-Net Cao et al. 2021, Attention U-Net Abraham and Khan 2019, U-Net++ Zhou et al. 2019, and V-Net Abdollahi et al. 2020) are used to identify the region of interest in the ultrasound breast images.
4.4 Classification and hyperparameters optimization phase
Classification of medical images into their correct class helps physicians in their diagnosis. Hence, the second processing phase of the framework is to classify breast images into either benign or malignant. Medical data always suffers from scarcity. Therefore, five different pre-trained architectures (i.e., MobileNet Howard et al. 2017, MobileNetV2 Sandler et al. 2018, NasNetMobile Addagarla et al. 2020, VGG16 Swasono et al. 2019, and VGG19 Carvalho et al. 2017) are used. As mentioned, different optimizers are applied to tune the parameters of the different CNN architectures (i.e., Adam, AdaGrad, NAdam, AdaMax, AdaDelta, RMSProp, and SGD optimizers). The corresponding used equations are Eq. 5 for Adam, Eq. 5 for NAdam, Eq. 7 for AdaGrad, Eq. 8 for AdaDelta, Eq. 9 for AdaMax, Eq. 10 for RMSProp, and Eq. 11 for SGD.
where \(m_t\) is the mean, \(\nu\) is the uncentered variance of the gradients, \(\eta\) is the step size, \(\epsilon\) is a small quantity used to prevent the division by zero, \(G_t \in \mathbb {R}^{(d \times d)}\) is a diagonal matrix, RMS is the root mean squared, \(g_t\) is the gradient of loss function, \(E[g^2]_t\) is the average of squared gradients, \(\beta _1\) and \(\beta _2\) are hyperparameters, \(\gamma _{x-1}\) is the average of squared past gradient, and \(x^{(i)}\) and \(y^{(i)}\) are the input and output pair respectively.
For better selection of the different hyperparameters, AO is introduced at the learning phase. As mentioned in Sect. 3.2, it depends on 4 updating methods. They are expanded exploration \((X_1)\), narrowed exploration \((X_2)\), expanded exploitation \((X_3)\), and narrowed exploitation \((X_4)\). For the expanded exploration \((X_1)\) method, Eq. 12 is used. \(X_M(t)\) is calculated using Eq. 13. where \(X_{best}(t)\) is the best location, \(X_M(t)\) is the average location of the entire Aquila’s in the ongoing iteration, t is the number of the ongoing iteration, T is the total count of iterations, N is the population size, and \(r_1\) is an arbitrary number in the range of 0 to 1. For the narrowed exploration \((X_2)\) method, Eq. 14 is used where \(X_R(t)\) is any arbitrary chosen location of Aquila, D is the size of the dimension, and \(r_2\) is an arbitrary value in the range of 0 to 1. LF(D) is the Levy’s flight and is calculated as shown in Eqs. 15 and 16 where s and \(\beta\) are fixed as 0.01 and 1.5, respectively, v and u are arbitrary number in the range of 0 to 1, and x and y represents the helix movement during the search and can be calculated function as shown in Eqs. 17 to 20 where \(r_3\) is the count of search cycles in the range from 1 to 20, \(D_1\) consists of integers in the range from 1 to D, and w is 0.005. For the expanded exploitation \((X_3)\) method, Eq. 21 is used where \(\alpha\) and \(\delta\) are adapting parameters set at 0.1, LB and UB are the lower and upper limits of the problem, and \(r_4\) and \(r_5\) are arbitrary values in the range of 0 to 1. For the narrowed exploitation \((X_4)\) method, Eq. 22 is used. QF(t) is calculated using Eq. 23, \(G_1\) is calculated using Eq. 24, and \(G_2\) is calculated using Eq. 25 where \(r_6\), \(r_7\), and \(r_8\) are arbitrary values in the range of 0 to 1, X(t) is the current location, QF(t) is the value of the quality function used to stabilize the search technique, \(G_1\) is an arbitrary value in the range of − 1 to 1 expressing Aquila’s movement during victim pursuit, and \(G_2\) is linearly reduced from 2 to 0 and represents the slope of flight when hunting victim.
4.5 System evaluation phase
Different performance metrics are used in the current study as mentioned in Sect. 3.4. The corresponding equations for them are accuracy (Eq. 26), (2) precision (Eq. 27), (3) recall (i.e., sensitivity) (Eq. 29), (4) specificity (Eq. 28), (5) F1-score (Eq. 30), (6) AUC, (7) IoU (Eq. 31), (8) dice coef. (Eq. 32), and (9) cosine similarity.
4.6 Pseudocode of the proposed framework
The learning and optimization steps are calculated repeatedly for a predefined number of iterations \(T_{max}\). After the learning iterations are executed, the finest combined configuration can be reported and used in any further systems. Algorithm 1 presents a summary of the introduced overall parameters learning and AO hyperparameters optimization approach.

5 Experiments and discussions
This section presents the results of experiments applied to the proposed framework. Python is the used scripting language. The major used packages are Tensorflow, Keras, keras-unet-collection, NumPy, OpenCV, and Matplotlib. The working environment is Google Colab with GPU (i.e., Intel(R) Xeon(R) CPU @ 2.00GHz, Tesla T4 16 GB GPU, CUDA v.11.2, and 12 GB RAM).
5.1 Segmentation phase experiments and discussion
Table 3 summarizes the common configurations of the segmentation experiments. Table 4 summarizes the segmentation phase experiments results. It is clear that the “Attention U-Net” is better than other models concerning the loss, accuracy, F1-score, precision, specificity, AUC, IoU coef., and dice coef. However, the “V-Net” is better than others concerning the recall (i.e., sensitivity). Figure 9 summarizes the segmentation phase experiments results graphically. Figure 10 shows the result of applying the Attention U-Net on a sample image. It shows that the region of interest from the predicted mask is relatively comparable with the original mask region of interest.
5.2 Classification phase experiments and discussion
Table 5 summarizes the classification phase experiments configurations. The finest associations for every model applied on “Breast Cancer Dataset” dataset are documented in Table 6. It is clear that the Categorical Crossentropy loss is the best choice using two models. The SGD and SGD Nesterov parameters optimizers are the best choice by two models each. Standardization is the best choice using two models. The finest associations for every model applied on “Dataset_BCD_ mammography_images_out” dataset are documented in Table 7. It is clear that the KLDivergence loss is the best choice using three models. The SGD and AdaMax parameters optimizers are the best choice by two models each. The min–max scaler is the best choice using three models. The finest associations for every model applied on “Breast Cancer Patients MRI’s” dataset are documented in Table 8. It is clear that the Categorical Crossentropy and KLDivergence losses are the best choice by two models each. The AdaGrad and SGD Nesterov parameters optimizers are the best choice by two models each. The standardization and min–max scaling are the best choice by two models each. This presents multiple performance indices concerning the “Breast Cancer Dataset” dataset in Table 9. From them, the VGG19 pre-trained model gives the topmost results over other models. This presents multiple performance indices concerning the “Dataset_BCD_ mammography_images_out” dataset in Table 10. From them, the MobileNet pre-trained model gives the topmost results over other models. This presents multiple performance indices concerning the “Breast Cancer Patients MRI’s” dataset in Table 11. From them, the MobileNet, VGG16, and VGG19 pre-trained models are the best model compared to others. Figures 11, 12, and 13 present graphical summarization of the performance metrics concerning the “Breast Cancer Dataset”, “Dataset_BCD_ mammography_images_out”, and “Breast Cancer Patients MRI’s” datasets respectively.
5.3 Comparative study
The results of the proposed framework against the related studies are shown in Table 12. As seen from these results, BCSF could achieve a 100% classification accuracy on MRI data, which is higher than the recorded accuracy. For segmentation, the achieved accuracy is better than most of the recent studies.
6 Conclusion and future work
This study introduced a hybrid framework for both classification and segmentation of breast images to diagnose breast cancer using CNN. The framework included two phases. The first phase is the segmentation phase. During this phase, the area of the tumor is detected to facilitate the diagnosis for the physician. Five different segmentation models are used, namely U-Net, Swin U-Net, Attention U-Net, U-Net++, and V-Net, to identify the region of interest in the ultrasound breast images. The used performance metrics are Accuracy, Recall, Precision, Specificity, F1-score, AUC, Sensitivity, IoU coef., dice coef., Hinge, and Squared Hinge. The second phase is the classification phase, in which breast images are classified. Five pretrained CNN architectures, namely MobileNet, MobileNetV2, NasNetMobile, VGG16, and VGG19 are applied. The choice of the different parameters for the used CNN architectures is done using seven different optimizers, namely Adam, AdaGrad, NAdam, AdaMax, AdaDelta, RMSProp, and SGD optimizers. On the other hand, Aquila Optimizer is used for the choice of the different hyperparameters of the various CNN architectures. For training purposes, three different datasets with different modalities are used to allow the diagnosis of breast cancer. Due to the differences in the datasets, a different number of classes for each type is available. Therefore, the suggested framework can perform both binary- and multi-class classification. The used performance metrics are Accuracy, Recall, Precision, Specificity, F1-score, AUC, Sensitivity, IoU coef., dice coef., TP, TN, FP, FN, and cosine similarity. the proposed framework can achieve a classification accuracy of 100% on MRI images. From the different segmentation methods, the best-recorded segmentation accuracy is 95.58% using Attention U-Net. However, the main limitations of the current work are: (1) limitation of data for training and (2) segmentation was applied to ultrasonic data only. As future work, we will apply the segmentation techniques to other types of images, namely MRI images. We also hope to apply other different optimization techniques such as the Red Deer algorithm (RDA) and Sine Cosine Algorithm (SCA). We also aim to use the suggested hybrid framework in other different medical imaging problems.
References
Abdollahi A, Pradhan B, Alamri A (2020) Vnet: an end-to-end fully convolutional neural network for road extraction from high-resolution remote sensing data. IEEE Access 8:179424–179436
Abdulazeem Y, Balaha HM, Bahgat WM, Badawy M (2021) Human action recognition based on transfer learning approach. IEEE Access
Abraham N, Khan NM (2019) A novel focal tversky loss function with improved attention u-net for lesion segmentation. In: 2019 IEEE 16th international symposium on biomedical imaging (ISBI 2019). IEEE, pp. 683–687
Addagarla SK, Chakravarthi GK, Anitha P (2020) Real time multi-scale facial mask detection and classification using deep transfer learning techniques. Int J 9(4):4402–4408
Ahmed L, Iqbal MM, Aldabbas H, Khalid S, Saleem Y, Saeed S (2020) Images data practices for semantic segmentation of breast cancer using deep neural network. J Ambient Intell Humaniz Comput 1–17
Al-Dhabyani W, Gomaa M, Khaled H, Fahmy A (2020) Dataset of breast ultrasound images. Data Brief 28:104863
Alghodhaifi H, Alghodhaifi A, Alghodhaifi M (2019) Predicting invasive ductal carcinoma in breast histology images using convolutional neural network. In: 2019 IEEE national aerospace and electronics conference (NAECON). IEEE, pp. 374–378
Alom MZ, Yakopcic C, Taha TM, Asari VK (2018) Nuclei segmentation with recurrent residual convolutional neural networks based u-net (r2u-net). In NAECON 2018-IEEE national aerospace and electronics conference. IEEE, pp. 228–233
AlRassas AM, Al-qaness MA, Ewees AA, Ren S, Abd Elaziz M, Damaševičius R, Krilavičius T (2021) Optimized anfis model using aquila optimizer for oil production forecasting. Processes 9(7):1194
Aswathy M, Jagannath M (2017) Detection of breast cancer on digital histopathology images: present status and future possibilities. Inform Med Unlocked 8:74–79
Bakker MF, de Lange SV, Pijnappel RM, Mann RM, Peeters PH, Monninkhof EM, Emaus MJ, Loo CE, Bisschops RH, Lobbes MB et al (2019) Supplemental MRI screening for women with extremely dense breast tissue. N Engl J Med 381(22):2091–2102
Balaha HM, Ali HA, Youssef EK, Elsayed AE, Samak RA, Abdelhaleem MS, Tolba MM, Shehata MR, Mahmoud MR, Abdelhameed MM, et al. (2021b) Recognizing Arabic handwritten characters using deep learning and genetic algorithms. Multimed Tools Appl 1–37
Balaha HM, Saafan MM (2021) Automatic exam correction framework (aecf) for the mcqs, essays, and equations matching. IEEE Access 9:32368–32389
Balaha HM, Ali HA, Saraya M, Badawy M (2021) A new Arabic handwritten character recognition deep learning system (ahcr-dls). Neural Comput Appl 33(11):6325–6367
Balaha HM, El-Gendy EM, Saafan MM (2021) Covh2sd: a covid-19 detection approach based on Harris hawks optimization and stacked deep learning. Expert Syst Appl 186:115805
Benson R, Mallick S, Rath GK (2020) Case carcinoma breast. Practical radiation oncology. Springer, New York, pp 201–209
Bingli L, Kanzaki T, Vargas DV (2021) Towards understanding the space of unrobust features of neural networks. In 2021 5th IEEE international conference on cybernetics (CYBCONF). IEEE, pp. 091–094
Bottou L (2012) Stochastic gradient descent tricks. Neural networks: tricks of the trade. Springer, New York, pp 421–436
Byra M, Jarosik P, Szubert A, Galperin M, Ojeda-Fournier H, Olson L, O’Boyle M, Comstock C, Andre M (2020) Breast mass segmentation in ultrasound with selective kernel u-net convolutional neural network. Biomed Signal Process Control 61:102027
Cao H, Wang Y, Chen J, Jiang D, Zhang X, Tian Q, Wang M (2021) Swin-unet: Unet-like pure transformer for medical image segmentation. arXiv preprint arXiv:2105.05537
Carvalho T, De Rezende ER, Alves MT, Balieiro FK, Sovat RB (2017) Exposing computer generated images by eye’s region classification via transfer learning of vgg19 cnn. In 2017 16th IEEE international conference on machine learning and applications (ICMLA). IEEE, pp. 866–870
Charishma GCG, Kusuma PNKPN, Narendra JBNJB (2020) Review on breast cancer and its treatment. Int J Indigenous Herbs Drugs, pp 21–26
Chorianopoulos AM, Daramouskas I, Perikos I, Grivokostopoulou F, Hatzilygeroudis I (2020) Deep learning methods in medical imaging for the recognition of breast cancer. In 2020 11th international conference on information, intelligence, systems and applications (IISA). IEEE, pp. 1–8, pp. 1–8
Clinic C (2022) Our commitment to safe care. https://my.clevelandclinic.org. Accessed 19 Jan 2022
Dabeer S, Khan MM, Islam S (2019) Cancer diagnosis in histopathological image: Cnn based approach. Inform Med Unlocked 16:100231
Dalmia A, Kakileti ST, Manjunath G (2018) Exploring deep learning networks for tumour segmentation in infrared images. In 14th quantitative infrared thermography conference
Dogo E, Afolabi O, Nwulu N, Twala B, Aigbavboa C (2018) A comparative analysis of gradient descent-based optimization algorithms on convolutional neural networks. In: 2018 international conference on computational techniques, electronics and mechanical systems (CTEMS). IEEE, pp. 92–99
Dozat T (2016) Incorporating nesterov momentum into adam
Duffy SW, Vulkan D, Cuckle H, Parmar D, Sheikh S, Smith RA, Evans A, Blyuss O, Johns L, Ellis IO et al (2020) Effect of mammographic screening from age 40 years on breast cancer mortality (uk age trial): final results of a randomised, controlled trial. Lancet Oncol 21(9):1165–1172
El Adoui M, Mahmoudi SA, Larhmam MA, Benjelloun M (2019) Mri breast tumor segmentation using different encoder and decoder cnn architectures. Computers 8(3):52
Elter M, Horsch A (2009) Cadx of mammographic masses and clustered microcalcifications: a review. Med Phys 36(6Part1):2052–2068
Feng X, Guo X, Huang Q (2017) Systematic evaluation on speckle suppression methods in examination of ultrasound breast images. Appl Sci 7(1):37
For Disease Control, C., & Prevention (2022) COVID-19. https://www.cdc.gov. Accessed 19 Jan 2022
Francies FZ, Hull R, Khanyile R, Dlamini Z (2020) Breast cancer in low-middle income countries: Abnormality in splicing and lack of targeted treatment options. Am J Cancer Res 10(5):1568
Gao F, Wu T, Li J, Zheng B, Ruan L, Shang D, Patel B (2018) Sd-cnn: a shallow-deep cnn for improved breast cancer diagnosis. Comput Med Imaging Graph 70:53–62
Habijan M, Leventić H, Galić I, Babin D (2019) Whole heart segmentation from ct images using 3d u-net architecture. In 2019 international conference on systems, signals and image processing (IWSSIP). IEEE, pp. 121–126
Hameed Z, Zahia S, Garcia-Zapirain B, Javier Aguirre J, María Vanegas A (2020) Breast cancer histopathology image classification using an ensemble of deep learning models. Sensors 20(16):4373
Hamidinekoo A, Denton E, Rampun A, Honnor K, Zwiggelaar R (2018) Deep learning in mammography and breast histology, an overview and future trends. Med Image Anal 47:45–67
Howard AG, Zhu M, Chen B, Kalenichenko D, Wang W, Weyand T, Andreetto M, Adam H (2017) Mobilenets: efficient convolutional neural networks for mobile vision applications. arXiv preprint arXiv:1704.04861
Indolia S, Goswami AK, Mishra SP, Asopa P (2018) Conceptual understanding of convolutional neural network-a deep learning approach. Proc Comput Sci 132:679–688
Indra P, Manikandan M (2021) Multilevel tetrolet transform based breast cancer classifier and diagnosis system for healthcare applications. J Ambient Intell Humaniz Comput 12(3):3969–3978
Ismail NS, Sovuthy C (2019) Breast cancer detection based on deep learning technique. In 2019 international UNIMAS STEM 12th engineering conference (EnCon). IEEE, pp. 89–92
Jogin M, Madhulika M, Divya G, Meghana R, Apoorva S, et al (2018) Feature extraction using convolution neural networks (cnn) and deep learning. In 2018 3rd IEEE international conference on recent trends in electronics, information & communication technology (RTEICT). IEEE, pp. 2319–2323
Khan A, Sohail A, Zahoora U, Qureshi AS (2020) A survey of the recent architectures of deep convolutional neural networks. Artif Intell Rev 53(8):5455–5516
Kim E-K, Kim H-E, Han K, Kang BJ, Sohn Y-M, Woo OH, Lee CW (2018) Applying data-driven imaging biomarker in mammography for breast cancer screening: preliminary study. Sci Rep 8(1):1–8
Kingma DP, Ba J (2014) Adam: a method for stochastic optimization. arXiv preprint arXiv:1412.6980
Lawrence RA (2022) Anatomy of the breast. Breastfeeding. Elsevier, Amsterdam, pp 38–57
Li S, Dong M, Du G, Mu X (2019) Attention dense-u-net for automatic breast mass segmentation in digital mammogram. IEEE Access 7:59037–59047
Lu Y, Qin X, Fan H, Lai T, Li Z (2021) Wbc-net: a white blood cell segmentation network based on unet++ and resnet. Appl Soft Comput 101:107006
Luo L, Xiong Y, Liu Y, Sun X (2019) Adaptive gradient methods with dynamic bound of learning rate. arXiv preprint arXiv:1902.09843
Maitra IK, Nag S, Bandyopadhyay SK (2012) Technique for preprocessing of digital mammogram. Comput Methods Programs Biomed 107(2):175–188
Masud M, Rashed AEE, Hossain MS (2020) Convolutional neural network-based models for diagnosis of breast cancer. Neural Comput Appl, pp 1–12
Mehra R et al (2018) Breast cancer histology images classification: training from scratch or transfer learning? ICT Express 4(4):247–254
Melekoodappattu JG, Subbian PS (2020) Automated breast cancer detection using hybrid extreme learning machine classifier. J Ambient Intell Humanized Comput, pp 1–10
Melekoodappattu JG, Dhas AS, Kandathil BK, Adarsh K (2022) Breast cancer detection in mammogram: Combining modified CNN and texture feature based approach. J Ambient Intell Humanized Comput, pp 1–10
Norouzi A, Rahim MSM, Altameem A, Saba T, Rad AE, Rehman A, Uddin M (2014) Medical image segmentation methods, algorithms, and applications. IETE Tech Rev 31(3):199–213
Organization WH (2022) GLOBAL CANCER OBSERVATORY. https://gco.iarc.fr/today/home. Accessed 19 Jan 2022
Preetha R, Jinny SV (2021) Early diagnose breast cancer with pca-lda based fer and neuro-fuzzy classification system. J Ambient Intell Humaniz Comput 12(7):7195–7204
Ronneberger O, Fischer P, Brox T (2015) U-net: Convolutional networks for biomedical image segmentation. In International Conference on Medical image computing and computer-assisted intervention. Springer, New York, pp. 234–241
Saikia AR, Bora K, Mahanta LB, Das AK (2019) Comparative assessment of cnn architectures for classification of breast fnac images. Tissue Cell 57:8–14
Salama WM, Aly MH (2021) Deep learning in mammography images segmentation and classification: automated cnn approach. Alex Eng J 60(5):4701–4709
Sandler M, Howard A, Zhu M, Zhmoginov A, Chen LC (2018) Mobilenetv2: Inverted residuals and linear bottlenecks. In Proceedings of the IEEE conference on computer vision and pattern recognition, pp. 4510–4520
Senthil Kumar K, Venkatalakshmi K, Karthikeyan K (2019) Lung cancer detection using image segmentation by means of various evolutionary algorithms. Comput Math Methods Med 2019
Sharma C, Kumar R (2021) Histopathology images and deep cnn based approach for breast cancer detection. Turkish J Physiother Rehabilit 32:3
Shen L, Margolies LR, Rothstein JH, Fluder E, McBride R, Sieh W (2019) Deep learning to improve breast cancer detection on screening mammography. Sci Rep 9(1):1–12
Singh KK, Singh A (2010) A study of image segmentation algorithms for different types of images. Int J Comput Sci Issues (IJCSI) 7(5):414
Society AC (2022) Get cancer information now. https://www.cancer.org. Accessed 19 Jan 2022
Sohail A, Khan A, Wahab N, Zameer A, Khan S (2021) A multi-phase deep cnn based mitosis detection framework for breast cancer histopathological images. Sci Rep 11(1):1–18
Swasono DI, Tjandrasa H, Fathicah C (2019) Classification of tobacco leaf pests using vgg16 transfer learning. In: 2019 12th International conference on information & communication technology and system (ICTS). IEEE, pp. 176–181
Vani S, Rao TM (2019) An experimental approach towards the performance assessment of various optimizers on convolutional neural network. In: 2019 3rd international conference on trends in electronics and informatics (ICOEI). IEEE, pp. 331–336
Vaswani A, Shazeer N, Parmar N, Uszkoreit J, Jones L, Gomez AN, Kaiser Ł, Polosukhin I (2017) Attention is all you need. In: Advances in neural information processing systems, pp. 5998–6008
Wang Z, Li M, Wang H, Jiang H, Yao Y, Zhang H, Xin J (2019) Breast cancer detection using extreme learning machine based on feature fusion with cnn deep features. IEEE Access 7:105146–105158
Wang J, Liu Q, Xie H, Yang Z, Zhou H (2021) Boosted efficientnet: detection of lymph node metastases in breast cancer using convolutional neural networks. Cancers 13(4):661
Wu L, Shen C, Hengel Avd (2016) Personnet: Person re-identification with deep convolutional neural networks. arXiv preprint arXiv:1601.07255
Yurttakal AH, Erbay H, İkizceli T, Karaçavuş S (2020) Detection of breast cancer via deep convolution neural networks using MRI images. Multimed Tools Appl 79(21):15555–15573
Zhang B, Rajan R, Pineda L, Lambert N, Biedenkapp A, Chua K, Hutter F, Calandra R (2021) On the importance of hyperparameter optimization for model-based reinforcement learning. In: International conference on artificial intelligence and statistics. PMLR, pp. 4015–4023
Zheng B, Yoon SW, Lam SS (2014) Breast cancer diagnosis based on feature extraction using a hybrid of k-means and support vector machine algorithms. Expert Syst Appl 41(4):1476–1482
Zhou Z, Siddiquee MMR, Tajbakhsh N, Liang J (2019) Unet++: redesigning skip connections to exploit multiscale features in image segmentation. IEEE Trans Med Imaging 39(6):1856–1867
Zuluaga-Gomez J, Al Masry Z, Benaggoune K, Meraghni S, Zerhouni N (2021) A cnn-based methodology for breast cancer diagnosis using thermal images. Comput Methods Biomech Biomed Eng Imaging Vis 9(2):131–145
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Balaha, H.M., Antar, E.R., Saafan, M.M. et al. A comprehensive framework towards segmenting and classifying breast cancer patients using deep learning and Aquila optimizer. J Ambient Intell Human Comput 14, 7897–7917 (2023). https://doi.org/10.1007/s12652-023-04600-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12652-023-04600-1