Abstract
Tailoring a deep convolutional neural network (CNN) for an implementation is a tedious and time-consuming task especially in image identification. In this study, an optimization scheme based on artificial bee colony (ABC) algorithm so-called optimal deep CNN (ODC) classifier for hyperparameter optimization of deep CNN is proposed for plant species identification. It is implemented to a ready-made leaf dataset namely Folio containing #637 images with 32 different plant species. The images are undergone various image preprocessing such as scaling, segmentation and augmentation so as to improve the efficacy of the ODC classifier. Therefore, the dataset is augmented from #637 to #15,288 leaf images whose #12,103 images is allocated for training phase and the remainder for testing the ODC. Moreover, a validation process on 20% of the training dataset is performed along with the training phase in both optimization and classification stages. The accuracy and loss performance of the ODC are examined over the training and validation results. The achieved ODC is verified through the test phase as well as by a comparison with the results in the literature in terms of performance evaluation metrics such as accuracy, sensitivity, specificity and F1-score. In order to further corroborate the proposed scheme, it is even subjected to a benchmark with optimization-based studies such as genetic, particle swarm and firefly algorithms through MNIST digit-image dataset. The ODC identifies the leaf images and digit-images with the best accuracy of 98.99% and 99.21% surpassing the state of the arts. Therefore, the proposed ODC is effective and useful in achieving an optimal CNN thanks to ABC algorithm.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Automatic plant species identification, which requires an expert analysis, is an important issue for the agronomy, forestry, taxonomy, and particularly weed detection and removal. Progressing with the computer technologies in machine vision, it is possible to identify via artificial intelligence (Hassaballah and Awad 2020). Therefore, visual aspects, morphologies and attributes of the plant such as the leaf, flower, seed and stem gain importance for identification through the artificial intelligence-integrated machine vision (AIMV). Among them, leaf possesses the most descriptive information about the species of the plant. The leaves are hence the most employed organs for the identification of plant species via AIMV (Wäldchen and Mäder 2018). It is worthwhile that the success of the AIMV depends on not only appropriate artificial intelligence algorithm, but also applicable image preprocessing.
Artificial intelligence algorithms can be considered in two categories: traditional algorithms and deep learning algorithms. Artificial neural network (ANN) (McCulloch and Pitts 1943), support vector machine (SVM) (Cortes and Vapnik 1995), K-nearest neighbors (KNN) (Dudani 1976), Random Forest (RF) (Breiman 2001), Naïve Bayes (NB) (Maron 1960) and fuzzy system (Mohammadzadeh and Zhang 2019; Balootaki et al. 2020; Mohammadzadeh et al. 2020; Mittal et al. 2020) are the well-known traditional algorithms. They have been even the most applied algorithms to the plant species classifiers (Lorena et al. 2011; Akbarzadeh et al. 2018; Pacheco and Krohling 2018; Zhang et al. 2018a). Their performance highly depends on how compatible they are for the classification problem. In order to utilize them for the identification, the visual features of the leaves regarding the color, shape, texture and pattern should be properly extracted from the images of the leaves to be classified. Since using redundant features in model of the classifier might give rise to an overfitted and outsized model, determining optimally the features with the minimum number and the most impactive is a crucial investigation in the plant species identification by the traditional algorithms. On the other hand, the classifier models have been trained and tested on various datasets, being able to contain self-taken, self-collected or ready-made leaf images. Ready-made images can be downloaded from open repositories, the most employed open datasets are namely Swedish leaf (Söderkvist 2001), Folio (Munisami et al. 2015b), Flavia (Wu et al. 2007), Leaf12 (Anubha Pearline et al. 2019), Leafsnaps (Kumar et al. 2012), Foliage (Kadir et al. 2011), MalayaKew (Lee et al. 2017), Intelligent Computing Laboratory (ICL) (Hu et al. 2012) and LifeCLEF (Joly et al. 2015).
Several studies based on the traditional algorithms are available in the literature seeking to identify the plant species from leaf images. In (Arribas et al. 2011), leaf classification of sunflower crops was carried out by a modified ANN using extracted features related to color, shape and texture on a self-taken dataset with the accuracy of 85%. SVM and KNN together with bag-of-visual-word scheme were modelled in (Kazmi et al. 2015) for weed detection exploiting the features such as edge shape, affine invariant regions and local vegetation color on plant images of sugar beet and creeping thistle. The SVM detected the weeds with the accuracy of 99.07% by outperforming the detection by KNN. In (Munisami et al. 2015b), plant leaf recognition was performed using the features of the shape and color histogram through KNN on Folio with the accuracy of 87.20%. The plant species were recognized via a classifier depending on local similarity based classification learning (LSCL) utilizing the texture feature on ICL dataset with the accuracy of 81.12% in (Zhang et al. 2017). Weeds in sugar beet were detected in (Bakhshipour and Jafari 2018) by ANN and SVM using the shape, color, moment invariant features of a self-taken dataset with 92.50% and 93.33%, respectively. In (Yigit et al. 2019), an investigation for determining the most effective leaf visual features such as the dimension, color, texture and pattern was comparatively conducted via ANN, NB, RF, KNN and SVM on Folio dataset. SVM was the best accurate classifier with 94.49% in the plant species identification.
Deep learning, which is the generic name of modern artificial intelligence algorithms, comprises multiple processing layers that allow representation learning of multiple level data abstraction. The deep learning creates and extrapolate new features from raw representations of input data without manually extracting the features as carried out in applying the traditional algorithms. Deep learning has more complex and deeper networks than those of the traditional algorithms, i.e. it is called shallow network. Convolutional Neural Network (CNN) is the premier deep learning framework for the machine vision. There are several pre-built CNN architectures in the open libraries which comprise stacked convolutional layers based on repeatable units such as VGGNet, GoogleNet, AlexNet, CaffeNet, Cifar, ResNet, Overfeat, VGG, and MobileNet (Sewak et al. 2018). However, they allow limited modifications only on their architectures. Therefore, CNN has been increasingly applied to the classifiers to carry its advantages in the machine vision to the plant species identification. In (Grinblat et al. 2016) a CNN model identifies bean leaves on a ready-made image dataset by preprocessing vein patterns with the accuracy of 96.9%. A model of CNN based on CaffeNet was employed in (Sladojevic et al. 2016) to identify plant species on a self-collected image dataset from the internet by preprocessing like resizing, squaring, cropping with the accuracy of 98%. In (Sun et al. 2017), a ResNet based CNN model was used for plant identification in a self-taken natural environment dataset namely BJFU100 and Flavia. Flavia dataset was classified by the model with the accuracy of 99.65%. A CNN built on AlexNet was exploited to extracted features for the identification via ANN (multi-layer perceptron–MLP) and SVM, and the result by ANN was best accurate with 99.4% in (Lee et al. 2017). A CNN with AlexNet, GoogleNet, and VGGnet was applied to plant dentification of LifeCLEF dataset in (Mehdipour Ghazi et al. 2017) by preprocessing such as translation, reflection, rotation and scaling with the accuracy of 80.18%. In (Ferentinos 2018), a CNN based on AlexNet, GoogleNet, Overfeat and VGGnet was implemented to plant disease detection of a ready-made database by preprocessing such as size reduction and cropping with the accuracy of 99.48% by VGGnet. A CNN constructed on GoogleNet and Cifar was used in (Zhang et al. 2018b) to identify maize leaf diseases through self-collected database by resizing with the accuracy of 98.4% by Cifar. In (Anubha Pearline et al. 2019), a CNN with VGG16 and VGG19 was applied to feature extraction of Folio, Swedish leaf, Flavia and Leaf12 databases for various traditional algorithms such as NB, RF and KNN with the best accuracy of 99.41% on Swedish leaf by VGG19. A CNN constructed on AlexNet, GoogleNet, VGG16, ResNet50 and MobileNetV2 was employed in (Esgario et al. 2020) to identify and estimate the stress severity through self-taken image of coffee leaves with the best accuracy of 95.24% by ResNet50. As the literature is reviewed, the success of the suggested approaches is remained limited owing to dependence on the pre-built architectures and the lack of optimization. Though few of them attempted to harmonize the pre-built architectures with the implementation, their performance is slightly improved. Therefore, hyperparameter optimization which aims to tailor the control parameters of CNN to the application should be performed rather than using those architectures which are time-consuming and requires a sophisticated knowledge.
Metaheuristic algorithms of the most eminent optimizations which are genetic algorithm (GA), particle swarm optimization (PSO), artificial bee colony (ABC), differential evolution (DE), firefly algorithm (FA) and tree growth algorithm (TGA) are widely used in engineering optimization applications (Carbas et al. 2021). ABC is one of the recently emerged successful metaheuristic algorithms inspired by foraging behavior of the honey bee (Karaboga and Basturk 2007; Karaboga et al. 2014). Although ABC is relatively new algorithm, it comes to fore among the metaheuristic optimization algorithms thanks to its performance (Toktas and Ustun 2020; Toktas et al. 2020; Toktas 2021). Several of them are also implemented to various hyperparameter optimization of CNN using TGA, FA, PSO and GA (Ijjina and Chalavadi 2016; Sui et al. 2017; Bacanin et al. 2020; Singh et al. 2021). The common features of those studies are that they were tested on MNIST handwritten image database (LeCun et al. 2010) which includes bilevel simple and small size images of 28 × 28 pixel. Moreover, their classification accuracies remain limited. It is reasonable to employ this dataset for benchmarking and comparison.
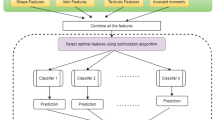
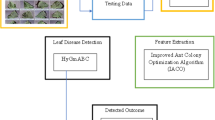
In this study, an optimal deep CNN (ODC) classifier is developed for plant identification from leaf images. Some important hyperparameters of the ODC regarding kernel/filter size and stride are optimized using ABC algorithm on Folio dataset. The working mechanism of the proposed classifier is presented as a flowchart in Fig. 1. In order to enhance the robustness of the classifier, the images are incurred a few image preprocessing such as scaling, segmentation and augmentation. The classifier is trained and tested through randomly selected 80% and 20% of the augmented dataset. The training phase is performed together with a validation process in both optimization and classification stages. The test results of the achieved ODC classifier are compared with the state of the art in terms of some reliable performance metrics. The ODC identifies the plant species with the best metrics accuracy, sensitivity, specificity and weighted average F1-score of 98.99%, 0.9996, 0.9904 and 0.9993, respectively. Furthermore, the proposed scheme is benchmarked with the metaheuristic optimization algorithms used studies through MNIST handwritten digit-image dataset. The ODC also gives the best identification results with 99.21% over the literature.
The study is organized as follows: Sect. 2 introduces the image dataset, describes the steps of image preprocessing applied to the dataset. Section 3 presents design optimization of the ODC classifier. Section 4 conveys the comparative numerical results and discussion on findings. Section 5 expresses a benchmark with literature through MNIST dataset. Section 6 gives the conclusion.
2 Leaf image database and preprocessing
The image preprocessing and computing process regarding the optimization and classification are conducted through MATLAB® platform running on a workstation with Intel Xeon W-2145 128 GB RAM NVIDIA GeForce RTX 2080Ti 11 GB.
2.1 Folio leaf image dataset
The ODC classifier is implemented and optimized for the plant identification for a ready-made image dataset named Folio (Munisami et al. 2015b). The dataset consists of a total number of 637 images with 32 different plant species. The images were taken in daylight via a full HD resolution camera from a farm nearby University of Mauritius. Leaf image examples for each species from Folio dataset are given in Fig. 2.
Image samples for each leaf species in Folio dataset (Munisami et al. 2015a) (For access Folio dataset: https://archive.ics.uci.edu/ml/datasets/Folio)
2.2 Image segmentation
Appropriate image preprocessing techniques to be applied to the images and their sequence are crucial for increasing the performance of the identification. In this study, the images are firstly resized all are to be the same size. The images are then exposed to a series of image preprocessing which is depicted in Fig. 3. The RGB levels of the images are defined in Fig. 3a using ‘imread(I)’ command in MATLAB. The images are then converted to gray scale in Fig. 3b by using ‘imread’, and to binary image in Fig. 3c by the agency of Otsu method (Otsu 1979) using, respectively ‘level = graythresh(I)’ and ‘BW = imbinarize(I,level)’, and then commands ‘strel(‘disk’,100)’ and ‘imclose’ are exploited to recover the erosions and distortions in the images, where disk and 100 are the shape and diameter of the used disk for recovering the images. Since the white background leads to poor performance and increases the processing time in the classification, the backgrounds of the images are therefore converted to the black. The RGB image and the binary image are combined in Fig. 3d by utilizing ‘double(I).*(BW)’, performed for each color layers. Finally, the background is removed in Fig. 3e for decreasing the size of the images.
2.3 Image augmentation
A big dataset is of great importance for performing effective training and test phases for the ODC classifier. Remember that the original number of leaf images are 637. This number of images may not sufficient for a robust CNN (Han et al. 2018). In this respect, increasing the number of the leaf images in Folio dataset would have assisted to construct a more robust classifier. In order to augment the number of images, 360/15 = 24 different images are reproduced from a leaf image by rotating each image with 15 degree as shown in Fig. 4. The number of images in the dataset is herewith augmented from 637 to 15,288 for a tight CNN structure.
3 Construction and optimization of the ODC classifier
Hyperparameters are the control parameters that establish and tune the construction of the ODC classifier. In general, the hyperparameters are determined by an empirical manner which is a tedious and time-consuming task. This study attempts to automatically determine some hyperparameters of ODC using ABC algorithm.
3.1 Data allocation
The augmented dataset with #15,288 leaf images is randomly split into #12,103 leaves (training-dataset) and #3185 leaves (test dataset) which approximately correspond to 80% and 20% of the dataset, respectively. Thereafter, the training-dataset are selected as training-set and validation-set using MATLAB command ‘[training-set, validation-set] = splitEachLabel (training-dataset, 0.8, ‘randomized’)’, where 0.8 implies that 80% of the training-dataset is randomly reserved for the training and the remainders for the validation process. On the other hand, the optimization of the ODC is carried out through #3057 leaf images that correspond to 25% of #12,103 leaves randomly selected from the total augmented dataset in order to speed up the process. This optimization-dataset is likewise split by 80% and 20% for training-set and validation-set, respectively.
3.2 The considered raw structure for the ODC classifier
For reducing the computational burden, kernel/filter sizes and stride among the hyperparameters, regarding the convolution and pooling operations are merely optimized. The kernel/filter sizes and stride are optimally selected within [1, 5] and [0, 4], respectively. The other hyperparameters are set as given in Table 1. The batch normalization in each layer is carried out, normalizing each input channel across a mini batch so as to reduce the sensitivity of network initialization and to improve the speed and stability of the network. The structure of CNN considered for the ODC classifier, which is empirically determined, consists of five convolution, four max pooling layers and a fully connected layer. The max pooling layer allows to reduce the dimension of the extracted feature vector and the feature map is converted to one-dimensional (1D) feature vector by the fully connected layer.
3.3 Hyperparameter optimization of ODC using ABC
A semantic block diagram for representation of the hyperparameter optimization of the ODC classifier is revealed in Fig. 5 (Karaboga and Basturk 2007). ABC is sequentially processing across three phases: employed bee (EmBee), onlooker bee (OnBee) and scout bee (ScBee) whose names come from the variety of honey bees. A natural bee colony consists of two groups of bees: EmBee and OnBee. Each bee works in related with a nectar source. All EmBees work as the ScBees and initially they randomly fly vicinity of the hive to explore the positions of new nectar sources. The EmBees then start to exploit the nectar sources and search new positions of quality nectar sources around the initial positions. Whenever the EmBees come back to their hive, the OnBees in the hive determine the prolificacy of the nectar sources by observing actions of the EmBees. The OnBees then start to exploit the new nectar sources surrounding the previous ones. Meantime, whether a nectar source is consumed after several essays by an EmBee, and then this bee is assigned as a ScBee to explore another nectar source as performed in the initial phase.
In ABC algorithm, candidate solutions \({x}_{ij}\) are randomly produced within two specified constraints \(\left[{x}_{j}^{max}, {x}_{j}^{min}\right]\) as follows
and then the objective value \(V\_loss\) which is the validation loss of the ODC and respectively the fitness value \({fit}_{i}=1/\left(1+{V\_loss}_{i}\right)\) of each solution are calculated. In the EmBee phase, the following operator reproduces new solutions \({v}_{ij}\) around the previous ones.
where k, j and i (i ≠ k) are random positive integer numbers, and \({\phi }_{ij}\) is a random factor within [− 1, 1]. Similarly, the objective value \(V\_loss\) is obtained from the ODC and fitness value for each solution are calculated again, and probability is, respectively computed using the fitness values as given below:
In the OnBee phase, new solutions are reproduced from the greedy selected ones according to the probability values using (3). Thereafter, the objective value \(V\_loss\) is get from the ODC and the fitness value is achieved as calculated in the initial phase to provide the data for selection the best solution. Whether a solution could not be improved after a defined number of essays (limit), a totally new solution is produced instead of this solution by a ScBee using (1). Finally, the best solution achieved so far is recorded. These phases except for the initial phase are sequentially processing to a defined maximum number of iterations (MNI). The control parameters of ABC related to the number of populations, MNI and limit are set as 20, 50 and 180, respectively.
3.4 The achieved structure of the ODC classifier
The accomplished structure of ODC classifier is depicted in Fig. 6, its kernel/filter sizes and stride (shown with red) are optimally found by ABC through the optimization scheme demonstrated in Fig. 5. Recall that the considered CNN structure is not based on any pre-built architectures. Therefore, it is particularly tailored for the plant identification. From the Fig. 5, the preprocessed images are input to the ODC classifier and hence undergone various operations layer by layer such as convolution and max pooling. In convolution, a multiplication is performed between the elements of the leaf images and the kernel by sliding the kernel window as the stride over the images, and then their sum is added as a single pixel to form another reduced image. By max pooling, the filter is slid as the stride size across the image, finding out the maximum pixel level remains within the area of the filter. A new image is then constituted by respectively inserting the maximum pixel levels into the image. The output of the convolution and pooling varies with the sizes of the kernel/filter and the stride. Their size highly depends on the image characteristics and the implementation. e.g. larger size should be used for global, high-level and representative information, whereas smaller size should be chosen as much as possible for local and sensitive information. It is apparent that there is a tradeoff between large and small size. Therefore, they should be optimized for the implementation.
The transaction of images through the ODC classifier is hereby addressed. First, the leaf image with size of 1024 × 1024 × 3 (red, green and blue) is taken as an input to the ODC classifier using activation function ReLU all over the convolution layers. It is then convoluted using kernel 3 × 3, stride 2 and 32 filters, and the resultant images with 512 × 512 × 32 are processed max pooling layer with filter 2 × 2 and stride 4. Afterwards, the images with 128 × 128 × 32 are undergone a convolution layer across kernel 3 × 3, stride 2 and 64 filters, and the outcome images with 64 × 64 × 64 are operated max pooling layer utilizing filter 2 × 2 and stride 4. Thereafter, the images with 32 × 32 × 64 are imposed a convolution layer over kernel 3 × 3, stride 1 and 64 filter, and the output images are transacted max pooling layer using filter 2 × 2 and stride 2. Then, the images with 16 × 16 × 64 are convoluted employing kernel 3 × 3, stride 1 and filter 64, and the resultant images 16 × 16 × 64 are performed max pooling layer through filter 2 × 2 and stride 2. Afterwards, the images with 8 × 8 × 64 are subjected to a convolution layer utilizing kernel 3 × 3, stride 1 and filter 32, and the outcome images enter fully connected layer with 2048 size. Eventually, it is undergone classification softmax layer with 32 sizes.
4 The numerical results
The ODC classifier is optimally constructed in Fig. 6 with the aid of ABC in the previous section. In this section, the ODC is incurred a training process (with validation) and then a test process and the test results are compared with the state of the arts using Folio dataset in terms of a statistical analysis using well-known metrics.
4.1 Training and validation results
The ODC classifier is trained through the training dataset with #12,103 leaves for a number of 20 epoch. The accuracy and loss results regarding the training and validation versus epoch are given in Fig. 7. Although the training is beginning (at the end of 1st epoch) with training accuracy/validation accuracy and training loss/validation loss of 75.00%/70.68% and 1.3655/1.3044, respectively. They are 98.44%/95.21% and 0.0199/0.0272 only at the end of 5th epoch. Eventually, they reach to 98.56%/98.28% and 0.00252/0.00358 at the end of 20th epoch. The results therefore demonstrate that the ODC is not only an accurate but also a stable classifier.
4.2 Comparative test results
In order to verify the ODC classifier, it is elaborately compared with the best results reported elsewhere for Folio dataset (Yigit et al. 2019; Anubha Pearline et al. 2019) in terms of the evaluation performance metrics sensitivity, specificity and F1-score values (Erkan 2021) in addition to the accuracy, which are given below:
where TP, TN, FP and FN are true positive, true negative, false positive and false negative, respectively. The comparative test results are tabulated in Table 2. In our study, F1-score is obtained as weighted average of F1-score of each class that computed in Eq. (7). Note that the study in (Yigit et al. 2019) was carried out for the feature selection of the leaf images, the best classification results were obtained with an accuracy of 94.49% by using SVM. The other study in (Anubha Pearline et al. 2019) was performed on various datasets, the best classification results on Folio dataset were achieved with an accuracy 96.53% by a CNN built on VGG19 with logistic regression (LR). On the other hand, the ODC classifies the testing-dataset with a surpassing accuracy of 98.99%, and with sensitivity, specificity and weighted average F1-score of 0.9996, 0.9904 and 0.9993, respectively. It is evident that these results are better than those of 0.9700 and 0.9700 for sensitivity and F1-score in (Anubha Pearline et al. 2019) considering that the ideal results of the metrics such as sensitivity, specificity and F1-score are the unit (one) for a fully accurate classification. Since the other results of the metrics are not available (N/A) in the studies, they could not be used for the comparison.
One may wonder that how many and which leaf species images are misclassified by the ODC. The number of 33 images with a variety of 13 species are misclassified from the total test dataset #3185 images. The most three species of them are Betel, Sweet Olive and Barbados Cherry with the number of 9, 5 and 3 misclassifications, respectively. The main reason of that, several species is very similar to each other in the view of visual features such as the color, shape, texture and pattern as seen from Fig. 8.
5 A benchmark with literature through MNIST dataset
In order to further corroborate the proposed scheme, it is implemented to a well-known dataset so-called MNIST handwritten image dataset (LeCun et al. 2010) for measuring and benchmarking with the state of the arts. One sample from every digit is given in Fig. 9. That is why, this dataset is the most utilized database in optimization of CNN structure. The MNIST image dataset consists of handwritten decimal digits including #60,000 and #10,000 samples for training-dataset and testing-dataset samples, respectively. These of training-dataset are randomly split into 80% and 20% as training-set and validation-set. In order for speeding up the optimization process, the ODC is optimized with 15,000-digit images which corresponds to 25% of #60,000 digits. The images which are bilevel (black and white) images are normalized by anti-aliasing technique and shaping to be fixed 28 × 28 pixel as conserving aspect ratio. They also are centered by using the computation of the mass center of the pixels and translating the image so as to position this point at the center of the field. Since the dataset includes simple and small size images, any preprocessing is not applied. Through MNIST dataset, the accomplished structure of ODC classifier is revealed in Fig. 10. As performed for Folio dataset, its kernel/filter sizes and stride (shown with red) are optimized by ABC. The considered CNN structure is also optimized for MNIST dataset.
MNIST dataset including 70,000 handwritten decimal digits (LeCun et al. 2010)
Kernel/filter sizes and stride from the hyperparameters related to the convolution and pooling layers are also optimized here. The kernel/filter sizes and stride are optimally selected within [1, 5] and [0, 4], respectively. The other hyperparameters are even set as given in Table 1. From the table, the ODC is optimized with epoch is set 3 during optimization, then it is taken as 20 for training. The training begins with training accuracy/validation accuracy and training loss/validation loss of 98.44%/97.18% and 0.066/0.1088 at the end of first epoch, respectively. At the end of 5th epoch, they are respectively 100.00%/98.77% and 0.0143/0.0378 only. At the end of 20th epoch, eventually they reach to 100.00%/99.28% and 0.0004/0.0313, respectively.
The comparative test accuracy results for benchmarking the optimization algorithms used studies are tabulated in Table 3. From the table, ODC classifies the digit images with the best accuracy of 99.21%. The other results of ODC regarding sensitivity, specificity and weighted average F1-score are 0.9917, 0.9990 and 0.9916, respectively. Therefore, proposed optimization scheme is even efficient and successful in addition to the leaf image classification.
6 Conclusion
In this study, a hyperparameter optimization scheme using ABC algorithm was proposed to fit ODC so as to identify the plant species from the leaf images. The ODC classifier was applied to a ready-made leaf dataset referred to as Folio having #637 images with 32 different plant species. The images were preprocessed by scaling, segmentation and augmentation for achieving a robust ODC classifier. After the preprocessing, there would be #15,288 augmented leaf images ready for the implementation, and thus the ODC was trained on #12,103 images from the augmented dataset. The training phase was integrated with a validation process on 20% of the training dataset in both optimization and classification stages. The ODC was diagnosed through the training and validation results with regard to the accuracy and loss rates. The ODC classifier was then verified through the test phase on #3185 leaf images by a comparison with the state-of-the-art results in terms of well-known performance metrics such as accuracy, sensitivity, specificity and weighted average F1-score. Moreover, in order to corroborate the proposed scheme with ABC, it was exposed to a benchmark with similar optimization employed studies through MNIST handwritten digit-image dataset. The results show that the ODC, respectively, classifies the plant species and digit images with best accuracy of 98.99% and 99.21% digit-images which outperforms the previously reported results. Besides, the ODC plant identification metrics such as sensitivity, specificity and weighted average are 0.9996, 0.9904 and 0.9993, respectively. Therefore, the proposed ODC with ABC is a handy and successful for the hyperparameter optimization of a CNN.
References
Akbarzadeh S, Paap A, Ahderom S et al (2018) Plant discrimination by Support Vector Machine classifier based on spectral reflectance. Comput Electron Agric 148:250–258. https://doi.org/10.1016/j.compag.2018.03.026
Anubha Pearline S, Sathiesh Kumar V, Harini S (2019) A study on plant recognition using conventional image processing and deep learning approaches. Journal of Intelligent and Fuzzy Systems. https://doi.org/10.3233/JIFS-169911
Arribas JI, Sánchez-Ferrero GV, Ruiz-Ruiz G, Gómez-Gil J (2011) Leaf classification in sunflower crops by computer vision and neural networks. Comput Electron Agric 78:9–18. https://doi.org/10.1016/j.compag.2011.05.007
Bacanin N, Bezdan T, Tuba E et al (2020) Optimizing convolutional neural network hyperparameters by enhanced swarm intelligence metaheuristics. Algorithms 13:67. https://doi.org/10.3390/a13030067
Bakhshipour A, Jafari A (2018) Evaluation of support vector machine and artificial neural networks in weed detection using shape features. Comput Electron Agric 145:153–160. https://doi.org/10.1016/j.compag.2017.12.032
Balootaki MA, Rahmani H, Moeinkhah H, Mohammadzadeh A (2020) On the Synchronization and Stabilization of fractional-order chaotic systems: recent advances and future perspectives. Phys A Stat Mech Appl 551:124203
Breiman L (2001) Random forests. Mach Learn 45:5–32. https://doi.org/10.1023/A:1010933404324
Carbas S, Toktas A, Ustun D (eds) (2021) Nature-ınspired metaheuristic algorithms for engineering optimization applications. Springer Singapore
Cortes C, Vapnik V (1995) Support-vector networks. Mach Learn 20:273–297. https://doi.org/10.1023/A:1022627411411
Dudani SA (1976) The distance-weighted k-nearest-neighbor rule. IEEE Trans Syst Man Cybern SMC-6:325–327. https://doi.org/10.1109/TSMC.1976.5408784
Erkan U (2021) A precise and stable machine learning algorithm: eigenvalue classification (EigenClass). Neural Comput Appl 33(10):5381–5392. https://doi.org/10.1007/s00521-020-05343-2
Esgario JGM, Krohling RA, Ventura JA (2020) Deep learning for classification and severity estimation of coffee leaf biotic stress. Comput Electron Agric 169:105162. https://doi.org/10.1016/j.compag.2019.105162
Ferentinos KP (2018) Deep learning models for plant disease detection and diagnosis. Comput Electron Agric 145:311–318. https://doi.org/10.1016/j.compag.2018.01.009
Grinblat GL, Uzal LC, Larese MG, Granitto PM (2016) Deep learning for plant identification using vein morphological patterns. Comput Electron Agric 127:418–424. https://doi.org/10.1016/j.compag.2016.07.003
Han D, Liu Q, Fan W (2018) A new image classification method using CNN transfer learning and web data augmentation. Expert Syst Appl 95:43–56. https://doi.org/10.1016/j.eswa.2017.11.028
Hassaballah M, Awad AI (2020) Deep learning in computer vision: principles and applications. CRC Press, NW
Hu R, Jia W, Ling H, Huang D (2012) Multiscale distance matrix for fast plant leaf recognition. IEEE Trans Image Process 21:4667–4672. https://doi.org/10.1109/TIP.2012.2207391
Ijjina EP, Chalavadi KM (2016) Human action recognition using genetic algorithms and convolutional neural networks. Pattern Recognit 59:199–212. https://doi.org/10.1016/j.patcog.2016.01.012
Joly A, Göeau H, Glotin H et al (2015) LifeCLEF 2015: multimedia life species identification challenges. Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics). Springer Verlag, Cham, pp 462–483
Kadir A, Nugroho LE, Susanto A, Santosa PI (2011) Signal & image processing. An Int J. https://doi.org/10.5121/sipij.2011.2301
Karaboga D, Basturk B (2007) A powerful and efficient algorithm for numerical function optimization: artificial bee colony (ABC) algorithm. J Glob Optim 39:459–471. https://doi.org/10.1007/s10898-007-9149-x
Karaboga D, Gorkemli B, Ozturk C, Karaboga N (2014) A comprehensive survey: artificial bee colony (ABC) algorithm and applications. Artif Intell Rev 42:21–57. https://doi.org/10.1007/s10462-012-9328-0
Kazmi W, Garcia-Ruiz F, Nielsen J et al (2015) Exploiting affine invariant regions and leaf edge shapes for weed detection. Comput Electron Agric 118:290–299. https://doi.org/10.1016/j.compag.2015.08.023
Kumar N, Belhumeur PN, Biswas A et al (2012) Leafsnap: a computer vision system for automatic plant species identification. European Conference on Computer Vision (ECCV). Springer Berlin Heidelberg, Berlin, Heidelberg, pp 502–516
LeCun Y, Cortes C, Burges CJC (2010) MNIST Handwritten Digit Database. In: AT&T Labs
Lee SH, Chan CS, Mayo SJ, Remagnino P (2017) How deep learning extracts and learns leaf features for plant classification. Pattern Recognit 71:1–13. https://doi.org/10.1016/j.patcog.2017.05.015
Lorena AC, Jacintho LFO, Siqueira MF et al (2011) Comparing machine learning classifiers in potential distribution modelling. Expert Syst Appl 38:5268–5275. https://doi.org/10.1016/j.eswa.2010.10.031
Maron ME (1960) Automatic indexing: an experimental inquiry. J ACM 8:404–414. https://doi.org/10.1145/321075.321084
McCulloch WS, Pitts W (1943) A logical calculus of the ideas immanent in nervous activity. Bull Math Biophys 5:115–1333. https://doi.org/10.1007/BF02478259
Mehdipour Ghazi M, Yanikoglu B, Aptoula E (2017) Plant identification using deep neural networks via optimization of transfer learning parameters. Neurocomputing 235:228–235. https://doi.org/10.1016/j.neucom.2017.01.018
Mittal K, Jain A, Vaisla KS et al (2020) A comprehensive review on type 2 fuzzy logic applications: past, present and future. Eng Appl Artif Intell 95:103916. https://doi.org/10.1016/j.engappai.2020.103916
Mohammadzadeh A, Zhang W (2019) Dynamic programming strategy based on a type-2 fuzzy wavelet neural network. Nonlinear Dyn 95:1661–1672. https://doi.org/10.1007/s11071-018-4651-x
Mohammadzadeh A, Sabzalian MH, Zhang W (2020) An interval type-3 fuzzy system and a new online fractional-order learning algorithm: theory and practice. IEEE Trans Fuzzy Syst 28:1940–1950. https://doi.org/10.1109/TFUZZ.2019.2928509
Munisami T, Ramsurn M, Kishnah S, Pudaruth S (2015a) Folio Data Set UCI Machine Learning Repository. https://archive.ics.uci.edu/ml/datasets/Folio
Munisami T, Ramsurn M, Kishnah S, Pudaruth S (2015b) plant leaf recognition using shape features and colour histogram with K-nearest neighbour classifiers. Procedia Comput Sci 58:740–747. https://doi.org/10.1016/j.procs.2015.08.095
Otsu N (1979) A threshold selection method from gray-level histograms. IEEE Trans Syst Man Cybern 9:62–66. https://doi.org/10.1109/TSMC.1979.4310076
Pacheco AGC, Krohling RA (2018) Aggregation of neural classifiers using Choquet integral with respect to a fuzzy measure. Neurocomputing 292:151–164. https://doi.org/10.1016/j.neucom.2018.03.002
Sewak M, Karim R, Pujari P (2018) Practical convolutional neural networks. Packt Publishing
Singh P, Chaudhury S, Panigrahi BK (2021) Hybrid MPSO-CNN: Multi-level Particle Swarm optimized hyperparameters of Convolutional Neural Network. Swarm Evol Comput 63:100863. https://doi.org/10.1016/j.swevo.2021.100863
Sladojevic S, Arsenovic M, Anderla A et al (2016) Deep neural networks based recognition of plant diseases by leaf image classification. Comput Intell Neurosci. https://doi.org/10.1155/2016/3289801
Söderkvist O (2001) Computer vision classification of leaves from swedish trees. Master’s Thesis, Linkoping University
Sui C, Bennamoun M, Togneri R (2017) Deep feature learning for dummies: a simple auto-encoder training method using Particle Swarm Optimisation. Pattern Recognit Lett 94:75–80. https://doi.org/10.1016/j.patrec.2017.03.021
Sun Y, Liu Y, Wang G, Zhang H (2017) Deep learning for plant identification in natural environment. Comput Intell Neurosci. https://doi.org/10.1155/2017/7361042
Toktas A (2021) Multi-objective design of multilayer microwave dielectric filters using artificial bee colony algorithm. In: Carbas S, Toktas A, Ustun D (eds) Nature-ınspired metaheuristic algorithms for engineering optimization applications. Springer Singapore
Toktas A, Ustun D (2020) A triple-objective optimization scheme using butterfly-integrated ABC algorithm for design of multilayer RAM. IEEE Trans Antennas Propag 68:5602–5612. https://doi.org/10.1109/TAP.2020.2981728
Toktas A, Ustun D, Tekbas M (2020) Global optimisation scheme based on triple-objective ABC algorithm for designing fully optimised multi-layer radar absorbing material. IET Microwaves Antennas Propag 14:800–811. https://doi.org/10.1049/iet-map.2019.0868
Wäldchen J, Mäder P (2018) Plant species identification using computer vision techniques: a systematic literature review. Arch Comput Methods Eng 25:507–543. https://doi.org/10.1007/s11831-016-9206-z
Wu SG, Bao FS, Xu EY et al (2007) A leaf recognition algorithm for plant classification using probabilistic neural network. In: ISSPIT 2007-2007 IEEE ınternational symposium on signal processing and ınformation technology, pp 11–16
Yigit E, Sabanci K, Toktas A, Kayabasi A (2019) A study on visual features of leaves in plant identification using artificial intelligence techniques. Comput Electron Agric 156:369–377. https://doi.org/10.1016/j.compag.2018.11.036
Zhang S, Wang H, Huang W (2017) Two-stage plant species recognition by local mean clustering and Weighted sparse representation classification. Cluster Comput 20:1517–1525. https://doi.org/10.1007/s10586-017-0859-7
Zhang S, Zhang C, Wang Z, Kong W (2018a) Combining sparse representation and singular value decomposition for plant recognition. Appl Soft Comput J 67:164–171. https://doi.org/10.1016/j.asoc.2018.02.052
Zhang X, Qiao Y, Meng F et al (2018b) Identification of maize leaf diseases using improved deep convolutional neural networks. IEEE Access 6:30370–30377. https://doi.org/10.1109/ACCESS.2018.2844405
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Erkan, U., Toktas, A. & Ustun, D. Hyperparameter optimization of deep CNN classifier for plant species identification using artificial bee colony algorithm. J Ambient Intell Human Comput 14, 8827–8838 (2023). https://doi.org/10.1007/s12652-021-03631-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12652-021-03631-w