Abstract
Enterococci are commensals of human and other animals’ gastrointestinal tracts. Only making up a small part of the microbiota, they have not played a significant role in research, until the 1980s. Although the exact year is variable according to different geographical areas, this was the decade when vancomycin-resistant enterococci (VRE) were discovered and since then their role as causative agents of human infections has increased. Enterococcus faecium is on the WHO’s list of “bacteria for which new antibiotics are urgently needed,” and with no new antibiotics in development, the situation is desperate. In this review, different aspects of VRE are outlined, including the mortality caused by VRE, antibiotic resistance profiles, animal-modeling efforts, and virulence. In addition, the limitations of current antibiotic treatments for VRE and prospective new treatments, such as bacteriocins, are reviewed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Vancomycin-resistant enterococci (VRE) emerged from the commensal enterococci in the 1980s and have developed from “generally regarded as safe” bacteria to significant nosocomial pathogens [2, 3, 21, 23, 45, 53, 68, 93, 109]. The World Health Organization (WHO) still considers VRE a pathogen with high priority on its global priority list [116]. VRE exist in the gastrointestinal tract of healthy humans and other animals, but may colonize and disseminate if conditions are suitable [1, 15, 57, 71, 97, 111]. Suitable conditions for colonization include treatment with anti-anaerobic antibiotics including vancomycin, which remove colonization resistance and provide a vacant niche for the VRE to invade [15, 84, 116]. In this context, colonization is the establishment of vancomycin-resistant enterococcal populations in the gastrointestinal tract due to displacement of the non-resistant enterococcal counterparts. Dissemination is the spread from the gastrointestinal tract and thereby start of an infection. Enterococci are innately resistant to many classes of antibiotics, but when they acquire additional resistance through, for example, mobile genetic elements, they become increasingly difficult to treat [54, 67, 101]. The factors that contribute to virulence in VRE are not completely characterized, but factors such as enterococcal surface protein, aggregation substance, gelatinase, and collagen adhesin molecule Acm have been implicated in the ability to colonize different tissues [23, 41, 60, 68, 70]. VRE can colonize the gastrointestinal tract of mice after antibiotic treatment and additional treatment with an immunocompromising agent allows dissemination in murine models [40, 79, 120]. Physicians have limited treatment options and resistance to new antibiotics is rising. There is now a pressing need for new classes of antibiotics that can deal with antibiotic resistant pathogens. Small antimicrobial peptides, bacteriocins, may be a new treatment alternative, either as the only treatment or in synergy with existing antimicrobial compounds [27, 114]. In this article, we will review these aspects in light of the significant threat VRE contribute to the era of antibiotic resistance, with special focus on factors facilitating the dissemination of VRE across the gastrointestinal tract to cause systemic infection, our current drug arsenal against VRE infections, and the potential of bacteriocins as alternative drugs and/or supplements in VRE treatments.
VRE Cause Significant Mortality
VRE are increasingly becoming a larger problem for hospitals worldwide, and due to VRE’s capabilities to survive for longer periods on inanimate surfaces (such as benches, beds, implanted surgical devices, and ventilation systems) and role as a commensal, it is increasingly difficult to control their spread [4, 90]. VRE cause a range of infections, from bacteremia, endocarditis, urinary tract infections, intra-abdominal, and pelvic infections, to peritonitis, skin infections, skin-structure infections, and central nervous system infections [29, 84, 90, 116, 128]. The attributable mortality of VRE infections is difficult to determine due to the common contribution of underlying disease, i.e., that the patient often is suffering from another condition prior to infection [104]. Examples of these conditions are cancerous conditions and transplant patients. The pathway of a VRE infection often starts with gastrointestinal colonization due to loss of colonization resistance through destabilization of the gut microbiota [39, 90]. Therefore, risk factors associated with promoting VRE infection include antibiotic exposure [105], comorbid illness [90], and patients that are immunocompromised, like patients undergoing chemotherapy [66]. In addition, factors such as prolonged hospital stay or an indwelling device, such as a catheter, breach a barrier and facilitate a proximity that gives the enterococci opportunity to cause infection [4, 82, 123]. A study among Pakistani hospitalized cancer patients indicated a 12-week mortality rate of 63% for VRE-bacteremia patients [4]. However, earlier studies have found lower mortality rates, for example, a study from several hospitals performed on patients with established VRE infection reported a 19% mortality rate, while a case-control matched study found an VRE attributable mortality of 37% [42, 123]. These studies report VRE mortality between 19% and 63%, and were performed on limited samplings, such as a few hospitals in one geographical area, and have a variability in the patient cohort included. Therefore, a future prospect should be to undertake investigations of a larger area, large cohort and continue to case-control match the patients, in order to gain a full understanding of the mortality burden caused by VRE.
Previous reports have indicated that VRE cause more mortality than infections caused by vancomycin-susceptible enterococci (VSE) [4, 94, 110]. This indicates either that some aspects of the resistant infections are more pathogenic than infections caused by the susceptible pathogen or it may reflect the timespan until effective treatment of the infection is reached. However, two factors may confound the results. Firstly, E. faecium has a higher rate of vancomycin resistance than E. faecalis, and the majority of VRE infections are caused by E. faecium, giving a skewed sampling. Secondly, there is more comorbidity in VRE infections than VSE, which makes determining cause of morbidity difficult [94].
Enterococci and Antibiotic Resistance
The vancomycin-resistant enterococci are facultative anaerobic, oval, Gram-positive cocci with high innate antibiotic resistance as well as significant acquired antibiotic resistance [8]. Intrinsic and acquired resistance differ in that, in general, genes that are present on the chromosome encode intrinsic resistance, while mobile genetic elements, such as plasmids and transposons, encode for acquired resistance [101]. Mobile genetic elements have been shown to play an important role in the acquired resistance of Enterococcus spp., and this has been reviewed elsewhere [54]. Enterococci intrinsically carry resistance to penicillin, aminoglycosides, clindamycin, and cephalosporins. In addition, they may require resistance to other β-lactams and glycopeptide antibiotics such as vancomycin [73, 82]. The different resistance mechanisms of enterococci have been reviewed elsewhere [67]. The most common nosocomial enterococci are E. faecalis and E. faecium. Previously E. faecalis was the most frequently isolated strain of the two, but a shift seems to have caused E. faecium to prevail in recent years [69, 111]. Interestingly, in most cases, E. faecium has a more extensive antibiotic resistance profile than E. faecalis, even though the latter has intrinsic resistance to the streptogramin quinupristin/dalfopristin and is hypothesized as the more virulent of the two [23, 67].
There are surveillance programs in place for monitoring incidences of VRE in both Europe, and other continents. Recent reports indicate an increasing trend of vancomycin resistance among E. faecium in Europe. Northern Europe continues to have a low percentage, but significant travel across European borders likely facilitates the spread of all resistant bacteria. The regional director of the World Health Organization stated that “antimicrobial resistance is increasingly widespread in the WHO European Region as resistant microbes know no borders” [116].
Different van-plasmids carry several genes necessary for vancomycin resistance. These genes encode enzymes that not only produce the alternative cell wall precursor D-Ala-D-Lac or D-Ala-D-Ser, depending on the type of van-plasmid, but also prevent synthesis of the original D-Ala-D-Ala [82]. Carrying the vancomycin resistance trait does not lower fitness, indicating that there is some type of regulatory system to induce the van genes when vancomycin is present [67]. The two most commonly found plasmids are the vanA and vanB plasmids, with vanA conferring the highest resistance [84]. Currently, nine different van-operons conferring different levels of vancomycin resistance are characterized [67, 84]. Vancomycin is a last resort antibiotic, an antibiotic used when other treatments fail due to resistance. It is therefore worrisome that resistance toward this last resort antibiotic spreads through an opportunistic species such as E. faecium and likely results in a relevant clinical impact [124].
Nosocomial Strains of VRE Are Enriched in Putative Virulence Factors
Several studies have investigated the importance of putative virulence genes in VRE, but there are no clear conclusions as to what constituents decisively contribute to the pathogenicity of VRE yet. Although it is difficult to prove the correlation between a virulence factor and the ability of a strain to cause infection, the general trend observed is that nosocomial strains carry the gene in question while indigenous strains do not. The only virulence genes confirmed to be associated with VRE infection are the enterococcal surface protein gene (esp) and the hyaluronidase gene (hyl) [69, 70]. The putative virulence factors include proteins that attack several different constituents of cells, such as cytolysin that targets cell membranes, as well as gelatinase and serine protease that attack various proteins such as collagen, fibrinogen, and insulin. In addition, binding proteins such as collagen-binding protein, enterococcal surface protein, and aggregation substance are putative virulence factors [41, 60, 68, 71]. Furthermore, it has been suggested that the different factors may have different roles in the different resistant Enterococcus species. One example of this is the frequently discussed virulence factor, enterococcal surface protein, Esp. It has been shown that Esp plays a vital role in the establishment of vancomycin resistant E. faecium urinary tract infection in animal models, but the role of Esp in E. faecalis is not clear. Therefore, it is possible to speculate that the Esp protein plays a less important role in E. faecalis, which is supported by the finding of Esp in both indigenous and nosocomial strains of E. faecalis, while the distribution of Esp in E. faecium is mostly limited to nosocomial strains [70]. One study investigating putative virulence factors in enterococci isolated from both the environment, food and nosocomial strains, further supported the differences between E. faecalis and E. faecium [71]. The virulence factors present in E. faecalis varied between strains, and the different strains contained combinations of cytolysin, gelatinase, and surface protein. E. faecium strains only contained surface protein, and no other putative virulence determinants [71]. Another virulence factor involved in adhesion is Acm, which is a member of the microbial surface components recognizing adhesive matrix molecules, MSCRAMMs. This protein is necessary for the bacteria to bind to collagen type 1 in vancomycin-resistant E. faecium isolates. Considering that adhesion to constituents of the extracellular matrix is necessary for the bacteria to colonize and invade tissues, it may be classified as a virulence factor [83].
A connection between a strain carrying virulence traits and a strain being vancomycin resistant has been evaluated with different conclusions. One study found that when investigating clinical strains, there was no correlation between a strain having virulence traits, and a strain being vancomycin resistant [23]. However, other studies have found that such correlation does exist [101], and even indicated that some plasmids carrying resistance genes also carry virulence factors [89].
Key Factors for VRE Infection and Dissemination
The mouse model is a well-established system to study the effect of infection, and this includes infections caused by VRE. Studies have investigated the processes that occur before and during the colonization and possible dissemination of VRE in mice. These studies have considerably increased our understanding of the interaction of VRE with the gut microbiota and the effect of antibiotic administration. Several studies indicate that the effect of antibiotic administration is profound on the composition of the microbiota, the bacteria associated with the human gastrointestinal tract, as well as on the ability of VRE to colonize [12, 40, 79, 115, 120].
The general tendency observed in controlled in vivo experiments, is that antimicrobials targeting anaerobic bacteria, for example metronidazole and β-lactams, promote VRE colonization, while antibiotics with less anti-anaerobic effect have less effect on the ability of VRE to colonize the gastrointestinal tract [40, 115]. In general, the density and diversity of the gastrointestinal microbiota decrease during antibiotic treatments, and although the density increases when the antibiotic treatment is discontinued, the composition is altered. Specifically, the populations of the phyla Lactobacillaceae and Bacteroides are distinctly decreased and the post-antibiotic microbiota consists of a larger number of Clostridium and Enterococcus species [120]. Antibiotics that reduce the population of Enterobacteriaceae and Lactobacillus spp. include kanamycin, ampicillin, metronidazole, neomycin, and vancomycin [79, 120]. Several antimicrobial treatments have an increasing effect on Enterobacteriaceae and Enterococcus spp., most likely due to eradication of competition. This includes metronidazole, vancomycin, and neomycin. The consequence is that administration of these drugs to patients increases the amount of enterococci in the gut, thereby increasing the possibility of dissemination and infection. The administration of an immune suppressant in the absence of antibiotic treatment does not influence the normal intestinal flora, indicating that change in microbiota is not the reason for the increased susceptibility of enterococcal infection in immunocompromised patients [79].
Different antimicrobials have distinct effects on the ability of VRE to colonize the gastrointestinal tract, through their interaction with the microbiota and the inability of the antibiotics to target VRE. Treatment of mice with different combinations of metronidazole, neomycin, kanamycin, and vancomycin increased colonization with VRE in several studies [12, 79, 120]. Anti-anaerobic antibiotics such as piperacillin, tazobactam, cefoxitin, and clindamycin promoted VRE colonization in a mouse model. In the same model, cefepime and aztreonam, which are antibiotics with little anti-anaerobic activity, did not facilitate VRE colonization. The combined antibiotic piperacillin/tazobactam is a drug with anti-enterococcal effect, but it also has potent anti-anaerobic effect. Studies showed that during treatment, VRE colonization was prevented. However, since the anaerobic microbiota was also targeted, VRE was able to colonize after discontinuation of treatment. This indicates that piperacillin/tazobactam is able to prevent VRE colonization to some degree, but that VRE persists in the gastrointestinal tract and is able to re-colonize upon discontinuation of the antibiotic treatment [40, 115].
Colonization is critical in the process of VRE infection. It has been shown that the commensal microbiota influences the ability of opportunistic pathogens to colonize in both a direct and an indirect route. The direct route is that the commensal bacteria occupy the niches that opportunistic species need to colonize, and they are better adapted, so that they displace the opportunists, a process referred to as competitive exclusion [15]. In addition, the commensal bacteria have been shown to interact with the innate immune system of the host, stimulating production of antimicrobial compounds, such as the C-type lectin RegIIIγ [12]. These antimicrobial compounds have minimal effect toward the commensals themselves, but have potent antimicrobial activity toward several Gram-positive opportunists, including the VRE [13].
A few studies have measured VRE dissemination after VRE has colonized the gut, a process believed to be necessary for the ability of VRE to cause systemic infection [12, 55, 79]. Studies have shown that administering a broad-spectrum antibiotic mixture consisting of neomycin, metronidazole, and vancomycin to mice followed by oral inoculation with VRE is sufficient to induce VRE colonization of the gastrointestinal tract and dissemination to other tissues. One study found that broad-spectrum antibiotic treatment and subsequent VRE inoculation were sufficient for VRE to disseminate to the blood of the treated animals [12]. Another study also investigated whether VRE had spread to tissues such as the liver and spleen [79]. In this study, the mice were administered cyclophosphamide in addition to the antibiotic mixture, and therefore were immunocompromised in addition to having their microbiota altered. They discovered VRE dissemination to mesenteric lymph nodes, liver, spleen as well as blood. Whether immunocompromising treatment is necessary for dissemination to tissues is unclear. Retrospective studies in humans provide support for the notion that the individual needs to be immunocompromised in order for the dissemination to occur. For example, one study found that of 216 human patients colonized with VRE, only three developed bloodstream infections, and all patients that developed bloodstream infections were immunocompromised. It might be that immunosuppression is necessary for the dissemination of VRE in humans as well as in experimental animals [66].
Is Esp Involved in Adherence?
Bacteremia with VRE may be dependent on the ability of the bacteria to colonize and translocate the intestinal tract [55]. This would depend on the ability of the bacteria to adhere to and invade the mucosal layer of the intestinal tract. One study in mice showed that the treatment with antibiotics, such as ampicillin, decreased the thickness of the mucus in the mucosal membrane. Colonization with Klebsiella pneumoniae restored the thickness of the colon mucus, but colonization with VRE did not. However, no invasion of the mucosal membrane was found in the antibiotic-treated, VRE-colonized mice [16], in line with previous notion that immunosuppression is necessary in combination with antibiotics for VRE dissemination. Some murine models have investigated the properties of the enterococci that facilitate their ability for high-level gastrointestinal colonization. The enterococcal surface protein (Esp) has been suggested as a factor that might be involved, especially since the encoding gene was enriched in clinical strains [101]. However, studies in mice did not indicate any difference in the colonization properties of neither E. faecalis nor E. faecium whether they had the esp gene or not [56, 95]. Furthermore, no difference was found in adherence to a human cell line in vitro, between an esp-positive E. faecium blood culture isolate, and the same strain with esp deleted [56]. This indicates that esp is not the factor responsible for adherence to human cell lines. In the same study, a significant difference in adherence was shown between a community-acquired esp-negative strain, and the blood culture esp-positive isolate. If one postulates that esp is not responsible for adherence, as indicated by the similar adherence in the same strain with and without esp, it suggests that the community isolates lack some other unknown adherence factors that the nosocomial blood isolates possess [56]. A study has shown that clinical strains of vancomycin-resistant E. faecium have the ability to bind to human intestinal mucus in vitro, but with lower affinity than other commensals of the gastrointestinal tract, indicating that the binding affinity of the different genera may play a part in the colonization resistance of the microbiota [96].
When VRE are able to cross over the gastrointestinal barrier and disseminate, other bacterial species may also spread. Miyazaki et al. found E. coli in the same blood samples as VRE, although outnumbered 100:1, indicating that both have followed the same infection route [79]. This correlates well with the suggestion that VRE might need the presence of other bacteria to cross over the mucus layer of the gastrointestinal tract, considering its limited ability to invade the mucus layer when mono-colonized [16].
Though Effective Treatment Exists, VRE Continues to Cause Significant Mortality and Resistance Arises
VRE are commensals of the gastrointestinal tract, but do not normally colonize this organ [57, 68]. However, colonization of the gastrointestinal tract is believed to be the first step of infection, and therefore it is of great importance [12, 55, 71, 79, 97, 111]. Severe infection with VRE does require treatment, but due to the high antibiotic resistance, and the innate ability of enterococci to develop resistance toward new compounds quickly, there are few effective therapies available [78]. Currently considered effective are quinupristin-dalfopristin, tigecycline, teicoplanin, telavancin, linezolid, and daptomycin (Table 1) [94]. Some of these compounds are only approved as treatment for skin-related infections, and some are in the experimental phases of development (see Table 1). However, resistance to these new antimicrobials has been documented as early as 2001 for quinupristin-dalfopristin, and in fact, none of the new antibiotics are free from enterococcal resistance [67, 123].
Quinupristin-Dalfopristin—Streptogramin Antibiotics
Quinupristin-dalfopristin is a combined antibiotic composed of two synergistically acting constituents that both bind to the 50S ribosomal subunit and interfere with protein translation [7, 67, 72]. It is commonly used against infections with VRE, but several studies have found that resistance is emerging [72, 73, 98]. For example, Maraki et al. found that 17.1% of isolated nosocomial strains were resistant to quinupristin-dalfopristin [73]. The susceptibility to quinupristin-dalfopristin has been determined to depend on the species of VRE. While most E. faecalis strains are resistant to quinupristin-dalfopristin, the antibiotic does have substantial activity toward E. faecium [36, 72]. This difference in susceptibility is likely due to the lsa gene, encoding a putative ABC-transporter that transports the antibiotic away from its 50S rRNA target [67, 127]. E. faecalis therefore has intrinsic resistance to quinupristin-dalfopristin while E. faecium carries acquired resistance due to acetyltransferases that modify the rRNA target, and by genes that encode ABC transporters to efflux the antibiotic [67, 127]. In addition to the occurrence of resistance, treatment with quinupristin-dalfopristin involves side effects such as joint and muscle pain [127].
Linezolid Has Bacteriostatic Effect
Linezolid is used for infections caused by antibiotic resistant Gram-positive organisms and has bacteriostatic effect by targeting the 23S rRNA subunit of the translational machinery [103]. It is reported by surveillance programs that the occurrence of resistance is rare, 1.83% resistance was reported in 2012 [37, 77]. Several studies have investigated the effect of linezolid on enterococcal infections. One large study found that among 138 patients that received linezolid treatment, there was approximately 18% overall mortality [119]. Two other studies reported higher mortality rates, 20.6% and 29.4%, with investigations in smaller patient cohorts, 68 and 34 patients, respectively [28, 74]. Resistance develops with some difficulty due to the fitness loss associated with an altered ribosomal subunit, resulting in less efficient protein translation [76]. Resistance development is normally associated with prolonged use of linezolid or invasive procedures [76]. Several resistance mechanisms have emerged in different types of antibiotic resistant bacteria, including VRE [34, 75, 76]. These resistance mechanisms include chromosomal modifications of the 23S rRNA subunit as well as the L3 and L4 accessory proteins [76]. Resistance may also be caused by the cfr gene which codes for an enzyme belonging to the radical S-adenosyl-L-methionine superfamily. Cfr methylates a carbon atom on the alanine in position 2503 in the 23S rRNA, protecting it from linezolid [34, 127]. It is possible that some strains that are non-susceptible to linezolid, but do not carry modifications in the 23S rRNA, L3 or L4, and do not carry the cfr gene, may contain efflux pumps that recognize linezolid, but this has not been proven yet [76]. The cfr gene is plasmid-located and associated with transposons and other mobile genetic elements; therefore, it is associated with a risk of dissemination [76].
Tigecycline Is a Tetracycline Derivative
Tigecycline is a glycylcycline antibiotic, which is a class of antibiotics derived from tetracycline. It targets the 30S ribosomal subunit and blocks the transfer RNA, hindering protein translation. Tigecycline has an increased affinity for the 30S subunit, hence being more potent compared with tetracycline [50]. Tigecycline monotherapy is not recommended as it has been indicated that the antibiotic cannot achieve high enough serum concentration to achieve sufficient antibacterial effect [7]. The side effects caused by tigecycline can be significant, and in combination with the low serum concentration, it may have limited value as a VRE therapy [127].
Teicoplanin and Telavancin Are Lipoglycopeptides
Teicoplanin and telavancin are semisynthetic derivatives of vancomycin belonging to the class lipoglycopeptides, and teicoplanin has shown more rapid bactericidal effect than vancomycin [38]. However, if the vancomycin resistance is caused by the vanA operon, neither will have any antimicrobial effect through binding to the D-Ala-D-Ala motif, which is the target site for vancomycin. Teicoplanin retains activity toward VRE when the resistance is caused by the vanB operon [67]. Hypothetically, telavancin has a second mode of action through interactions with the bacterial membrane causing depolarization and leakage of solutes. This is independent of the D-Ala-D-Ala binding motif, which means that telavancin may be useful in treatment of infection caused by vancomycin-resistant strains [38, 58].
Daptomycin Belongs to the Most Recently Discovered Class of Antibiotics
Daptomycin differs from other treatments in its mode of action, in that it is bactericidal, and that resistance is very rare [67, 106]. It has been reported that as many as 99.98% of E. faecalis and 99.82% E. faecium isolates are susceptible to daptomycin [106] [88], although reports on resistance range from less than 0.3% to 20% [18, 64, 107]. Daptomycin is believed to associate with the bacterial membrane, causing leakage of cellular solutes, thereby depolarization and cell death [67]. However, daptomycin susceptibility has decreased by mutations in genes such as liaF, gdpD, and cls, both through the occurrence of mutations in resistant strains and site-directed mutagenesis [9, 89, 118]. The mechanism of daptomycin resistance is not clear, but it is believed that the bacterial cell needs to change its membrane or trap the drug in order to divert its effects [118]. When daptomycin is administered in higher doses, there is a concern for the toxicity of the drug, as it has been found to cause increased creatine kinase levels, resulting in muscle toxicity [80].
In summary, the conventional antibiotic treatments for infections caused by VRE are often limited due to the development of resistance, as well as high dosage requirement and severe side effects including muscle and joint pain and nausea. All of these aspects make the advent of new antimicrobial therapies imperative.
Tedizolid—a Novel Oxazolidinone
Tedizolid is a relatively new antibiotic, belonging to the oxazolidinones, like linezolid. However, tedizolid has a 4-fold lower MIC value than linezolid, and it has been shown that some strains resistant to linezolid are susceptible to tedizolid [108, 127]. Tedizolid has bacteriostatic activity against VRE and functions by targeting the 23S rRNA of the 50S subunit and impairs protein translation, much like linezolid [127]. Although clinical data is limited, it is reported that of 163 VRE strains tested, 98.8% were inhibited [108]. However, due to the similar mechanism of action to linezolid, it is likely that cross-resistance may occur, which imposes limitations in the widespread use of this antibiotic [127].
Oritavancin—a Novel Lipoglycopeptide
Oritavancin is possibly the most promising among new antibiotic treatments of VRE infection. It currently holds approval only for treatment of acute bacterial skin and skin structure infections, ABSSI. However, it has shown promising results in animal trials of an endocarditis model [10]. Oritavancin is a semisynthetic lipoglycopeptide. Although its structure is similar to vancomycin, and it possesses the same ability to inhibit transglycosylation, oritavancin possesses an additional mechanism of action; it inhibits transpeptidation and is effective against vancomycin-resistant strains [81, 91]. It is suggested that oritavancin interacts with the bacterial membrane in a similar process as daptomycin, and causes membrane depolarization [38]. One clinical report suggested that oritavancin was successful in curing endocarditis in an elderly patient caused by E. faecium, although the patient experienced side effects from the treatment [62].
Although, these new treatment options are promising, it is likely that resistance mechanisms to these antibiotics will develop, especially if they are frequently used and thereby selective pressure for resistance is maintained.
Bacteriocins Provide New Treatment Options
Development of new antibiotics is important, but there is a growing view that a new type of antimicrobials is required to stagger the ever-developing resistance. Bacteriocins are antimicrobial peptides that are produced by bacteria, often to achieve an advantage over competing bacteria in certain niches.
Most bacteria produce bacteriocins, both Gram-positive and Gram-negative species [102, 126]. There are some different classification schemes, like the 2 categories suggested by Cotter et al. [26], and they broadly agreed with Klaenhammer classification. It has also been suggested that the circular bacteriocins should be a separate class [102]. The first one consists of the lantibiotics, which are small and contain a lanthionine residue, while the second class are also small, but they do not contain a lanthionine residue. This class can be subdivided into four categories: IIa–IId. The third class consists of large heat sensitive peptides [92]. New bacteriocins are frequently discovered [126], but discovering their mechanisms of action have traditionally been more challenging [25]. However, recent advances in receptor identification via, for example, genome sequencing of resistant mutants have significantly increased the ability to elucidate bacteriocin mechanisms [25]. The knowledge of how bacteriocins exert their antibacterial effect is critical in order to further bacteriocins to in vivo treatment of infection.
Bacteriocins in general have many advantages over the traditional antibiotics. Some examples are that they may have broad or narrow spectra, target different parts of the bacterial cell than antibiotics, often have high potency and may be bioengineered because of their gene-encoded nature [27, 86, 114]. Most bacteriocins are membrane-active peptides, targeting specific components, often proteins, in target cells [27, 35, 43, 47, 85, 86]. As shown below, some of these targeted proteins play vital roles in virulence development.
Bacteriocins rarely target the same cell components as antibiotics and therefore often have potent activity against antibiotic resistant strains [27]. This has been shown in murine models where different bacteriocins have been used to treat infections caused by other resistant bacteria, for example methicillin resistant Staphylococcus aureus, MRSA [11, 32, 33, 121, 122]. Therefore, bacteriocins may be a potential treatment for many types of bacterial infection. However, there is a lack of new research into how bacteriocins can be used in vivo. Recently the bacteriocin AS-48 was thoroughly investigated, with positive results in a preclinical study [19]. Hence, more bacteriocins need to be put through such in depth in vivo investigations in order to promote bacteriocins to relevant treatment options.
Although the use of bacteriocins as in vivo treatment is still limited, for now, the use of bacteriocins as additives in food has been recognized for some time, especially with nisin [100, 126]. As the food industry currently requires more food preservatives with a natural origin, bacteriocins represent promising possibilities [112]. However, bacteriocins have been under-utilized and further relevant research is required.
Combination Therapy: Bacteriocins and Antibiotics
Bacteriocins are promising treatment options alone but may be extra potent in combination treatment with synergistic antibiotics. Recently Hayes et al. published results that indicate that erythromycin and nisin have synergistic effect against strains of group B streptococcus [52]. Nisin also exhibits synergy with polymyxin B against Acinetobacter baumannii infections, which are nosocomial infections that are increasingly problematic [117]. Further, several combinations of nisin and antibiotics have been shown to be effective against Salmonella, both in vitro and in vivo in a murine model [113]. Chi and Holo described synergy between the bacteriocin garvicin KS and farnesol or polymyxin B against a range of bacteria, indicating that nisin is not the only bacteriocin that has synergy with the traditional antibiotics [20]. Hanchi et al. investigated synergy between durancin 61A and several traditional antibiotics, such as vancomycin and tetracycline. Durancin/vancomycin was favorably synergistic against Staphylococcus aureus, another critical antibiotic-resistant pathogen [51]. The synergy of antibiotics, bacteriocins, and other novel antimicrobials was described in a mini-review by Wolska et al., describing how combinatorial therapy has implications for many fields, such as the food industry, agriculture, and medicine [125]. Despite these examples, there are relatively few studies in this important area, as in other aspects of clinical bacteriocin research, that it is necessary to deal with in order to fully utilize bacteriocins and their potential.
Enterococci as Bacteriocin Producers and Bacteriocin Targets
Enterococci are common bacteriocin producers as well as being bacteriocin targets. Some are well characterized both in their antibacterial activity and their therapeutic potential. One example of this is the bacteriocin enterocin A + B. This bacteriocin shows antibacterial activity against Staphylococcus aureus, Listeria monocytogenes, and Escherichia coli [6]. Another two-peptide bacteriocin that is promising against Clostridium perfringens is the DD14 bacteriocin, which was shown identical to bacteriocin MR10. This bacteriocin in addition did not indicate any cytotoxicity against the cell line IPEC-1, indicating that this bacteriocin may be used in vivo [17]. Very few bacteriocins have undergone significant in vivo testing, but recently Cebrián et al. described the bacteriocin AS-48, produced by different strains of enterococci. They report low haemolytic effect, lack of toxicity and pro-inflammatory effect in a murine model, taking the promise of infection treatment with bacteriocin to the next level [19]. Another example of an enterococcal bacteriocin is EntV with activity against Candida albicans, indicating that bacteriocin also can be used against fungal infections [49]. These bacteriocins indicate the extensive range of bacteriocins produced and their respective targets, in enterococci. This diversity clearly illustrates the untapped potential of the bacteriocins.
As described, the enterococci produce a diverse group of bacteriocins, but they are also a suitable target for other bacteriocins. Keeping in mind the issues with VRE and antibiotic resistance raised in this article, one could say that they are not only suitable targets, but that bacteriocins are necessary to combat this pathogen. There are several bacteriocins that have in vitro activity against E. faecalis and/or E. faecium, such as nisin, bacteriocin EF478, enterocin P, enterocin K1, and more [22, 78, 87, 93]; some of these will be further treated below.
Nisin, Garvicin KS, and Bacteriocin EF478 Are Examples of Broad-Spectrum Bacteriocins
Studies have shown that nisin has been able to reduce the viability of both vancomycin-resistant E. faecium and E. faecalis in vitro, and that the supplement of nisin producing bacteria to VRE-colonized mice reduced the colonization [78]. Nisin targets the lipid II molecule of the bacterial membrane, using it as a docking molecule, and creates pores in the membrane and disrupts the proton motive force [121]. However, treating infection with nisin has not been attempted [78].
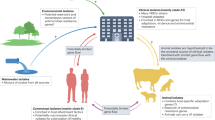
Garvicin KS is a multi-peptide bacteriocin produced by a Lactococcus garvieae strain isolated from contaminated raw milk [86]. It has a broad inhibitory spectrum including many Gram-positive pathogens such as MRSA and VRE, and the food-borne pathogens Listeria and Bacillus (Fig. 1). Resistance rate of Lactococcus lactis toward garvicin KS is quite low and the bacteriocin seems to target the PspC-mediated stress response network (unpublished data). In E. coli, pspC (also known as ythA) is important for the integrity and function of bacterial cell envelope [30]. In Yersinia enterocolitica, a pspC null mutant is virulent in a mouse model of infection [31]. Whether garvicin KS targets such a PspC-mediated stress response network in enterococci remains to be investigated.
Enterococcus faecium is intrinsically resistant to β-lactams and aminoglycosides, represented here by penicillin G and kanamycin. In addition, they may acquire resistance to antibiotics such as glycopeptides, represented here by vancomycin. VRE is a global issue, and has the ability to cause life-threatening infection. Bacteriocins represent potential new powerful treatment modalities against antibiotic resistant bacteria. Enterocin K1, enterocin EJ97, and garvicin KS are presented in this figure. These bacteriocins show potent activity against E. faecium (left: LMG3593, right: LMG3104) [72]. K1 = 10 μg enterocin K1, EJ97 = 10 μg enterocin EJ97, KS = 10 μg garvicin KS, PenG = 10 μg penicillin G, Kan = 5 μg kanamycin, Van = 5 μg vancomycin
Bacteriocin EF478 is a newly discovered bacteriocin showing potent activity against both E. faecalis and E. faecium [93]. Analysis indicates that this bacteriocin is a serine protease. This type of protein is known to be excreted as a toxin by other bacterial species. In addition, this bacteriocin demonstrated favorable chemical and thermodynamic qualities, indicating that it could be stable in an in vivo setting, and provide a promising new treatment option if developed [93].
Enterocins K1 and EJ97 Have Narrow Spectrum Activity Against Enterococcal Species
We have previously shown that enterocins K1 and EJ97 are bacteriocins that have potent and relatively narrow spectrum activity against E. faecium and E. faecalis, respectively (Fig. 1). They have been shown to target both vancomycin-susceptible and vancomycin-resistant nosocomial E. faecium and E. faecalis strains [87]. The bacteriocins are parts of the LsbB group of leaderless bacteriocins and have a conserved PWE motif in the C-terminus, which is important for the activity [88]. The current view of the mechanism of action is that enterocin K1 binds to the membrane-bound protein RseP, creating pores that cause leakage of solutes and other cellular constituents, thereby disrupting the proton motive force and killing the bacterial cell [87]. A structure–function relationship has also been studied for these peptides [87, 88]. They have similar structures, all with an alpha-helical motif at the N-terminal half and a non-structured part at the C-terminal half. The alpha-helical part has amphiphilic property and hence is believed to be involved in pore-forming, while the C-terminal part was demonstrated to be involved in receptor binding.
RseP Is an Achilles’ Heel
RseP (regulator of sigma-E protease) is a membrane-bound Zn-dependent protease involved in stress response through a process called regulated intramembrane proteolysis (RIP) [14, 24]. In E. coli, B. subtilis, E. faecalis, and other bacteria, RseP performs the second cleavage of an anti-σ factor after cleavage by a site-1 protease [5, 44]. The release of the alternative sigma factor is crucial for bacterial response to environmental stress [99] (Fig. 2). The RseP-mediated stress response process is remarkably similar in both Gram-positive and Gram-negative bacteria [59, 63, 65].
Gene activation through regulated intramembrane proteolysis [RIP] of anti-sigma factor in B. subtilis. Stress factors activate site-1 protease (PrsW) which cleaves anti-sigma factor RsiW at a periplasmic site followed by the second cleavage of RsiW which is carried by RasP (RseP) in the membrane. The cleaved sigma-factor undergoes further trimming by ClpXP in the cytoplasm before acting on to activate stress response genes [49]
Since RseP is crucial for the bacterial stress response in enterococci, bacteriocins targeting RseP (K1 and EJ97) not only kill sensitive bacteria but also leave resistant bacteria (with mutated RseP) to be killed by numerous environmental stressors. One such environmental stress factor is temperature. In fact, addition of K1 and EJ97 to enterococcal cells at different temperatures—comfortable (30 °C) and stressful (45 °C)—has shown that mutants appear at comfortable, but not at stressful temperatures [87]. The rseP gene has also been studied in vivo.
Frank et al. (2011) found that expression of rseP was increased during early infection, indicating that rseP is upregulated for infection establishment. However, deletion of rseP in E. faecalis OG1RF severely attenuated infection in an endocarditis model [46]. This indicates that the loss of rseP function affects the virulence of E. faecalis. Therefore, RseP mutants will likely not be able to establish infection, which is significant since bacteria with a functional RseP are eliminated by K1/EJ97 [87].
Concluding Remarks and Future Perspectives
The combined effect of enterococcal intrinsic and acquired antibiotic resistance results in dangerous opportunist pathogens. The general tendency of reduced potency of existing antibiotics and very limited development of new therapeutic agents, which are often synthetic derivatives, is emphasized by increased resistance development and cross-resistance. Considering the significant economic and social burden imposed by VRE infection, it is significant to develop new treatment, as well as limit the spread of the opportunists. Bacteriocins offer new possibilities in therapy, with significant advantages that ought not to be overlooked.
Hundreds of bacteriocins have been reported in literature; however, most of them are studied as natural food preservatives or probiotics, applications which require relatively few or less strict legal regulations. This is especially the case for those produced by lactic acid bacteria which are found in diverse fermented food products and also because they are common inhabitants in our gut flora. In fact, they are often referred to as “generally regarded as safe” or GRAS. However, for medical use, bacteriocins or any new drugs are exposed to more strict regulations as they need to be carefully assessed not only for potency but also for toxicity, delivery efficacy, and other physiological and immunological parameters, in both preclinical and clinical settings. One important field in bacteriocin research which is still lagging behind is the mode of action. It is about how a bacteriocin finds the target cells at the molecular level, the following interactions between a bacteriocin and a receptor or a docking molecule, and how these interactions eventually lead to the killing of the target cells. Detailed knowledge from this field is crucial to help develop bacteriocins into drug formulations that can kill target cells efficiently, without collateral effects and development of resistant cells. We and others have identified different membrane-located proteins that are required for the sensitivity to the bacteriocins; most likely these proteins serve as receptors. The majority of them are involved in transport of sugars or amino acids across the membrane. Others are involved in stress response against antimicrobials affecting membrane integrity. We currently study the interactions between enterocins K1 and EJ97 and their receptor RseP by crystallography which hopefully will share light into their mode of action in the near future.
Bacteriocins are gene-encoded, hence their sequences can be genetically modified that may lead to new properties [61, 129]. Other modification approaches are emerging. Peptidomimetics is modification on an existing peptide that can lead to advantageous properties, such as increased stability or broadened biological activity. Interestingly, some natural bacteriocins are post-translationally glycosylated in which the glycosyl group and the target cell’s sugar transporter PST are important for the antimicrobial activity [48]. Although it has not been experimentally demonstrated yet, it is tempting to speculate the attached glycosyl group might be used as a decoy so that the entire bacteriocin can enter the target cell via the sugar-PTS, a strategy resembling the Trojan horse strategy. Future research should include such modifications on natural bacteriocins, with sugars or other chemical groups, to seek for new and favorable properties, e.g., redirecting or broadening of the target spectrum and increased diffusion.
New sources for antimicrobials to combat antibiotic resistance are now a global demand. It is no doubt that bacteriocins represent a great potential in therapeutic treatments although they are currently underexploited. With better understanding of their mode of action and new technologies to modify and increase their usefulness, bacteriocins can be the next wave of drugs or supplements for therapeutic use. Finally, but not the least important, bacteriocins are superior to antibiotics in terms of environmental-friendliness. The former are of peptides and have therefore relatively short life in nature while the latter, especially those synthetic ones, are more difficult to degrade, often leaving a long-lasting footprint in nature.
Change history
25 November 2021
A Correction to this paper has been published: https://doi.org/10.1007/s12602-021-09873-6
References
Aarestrup FM, Agerso Y, Gerner-Smidt P, Madsen M, Jensen LB (2000) Comparison of antimicrobial resistance phenotypes and resistance genes in Enterococcus faecalis and Enterococcus faecium from humans in the community, broilers, and pigs in Denmark. Diagn Microbiol Infect Dis 37(2):127–137
Abele-Horn M, Vogel U, Klare I, Konstabel C, Trabold R, Kurihara R, Witte W, Kreth W, Schlegel PG, Claus H (2006) Molecular epidemiology of hospital-acquired vancomycin-resistant enterococci. J Clin Microbiol 44(11):4009–4013. https://doi.org/10.1128/JCM.00195-06
Adams DJ, Eberly MD, Goudie A, Nylund CM (2016) Rising vancomycin-resistant Enterococcus infections in hospitalized children in the United States. Hosp Pediatr 6(7):404–411. https://doi.org/10.1542/hpeds.2015-0196
Akhtar N, Sultan F, Nizamuddin S, Zafar W (2016) Risk factors and clinical outcomes for vancomycin-resistant Enterococcus bacteraemia in hospitalised cancer patients in Pakistan: a case-control study. J Pak Med Assoc 66(7):829–836
Akiyama Y, Kanehara K, Ito K (2004) RseP (YaeL), an Escherichia coli RIP protease, cleaves transmembrane sequences. EMBO J 23(22):4434–4442. https://doi.org/10.1038/sj.emboj.7600449
Ankaiah D, Palanichamy E, Antonyraj CB, Ayyanna R, Perumal V, Ahamed SIB, Arul V (2018) Cloning, overexpression, purification of bacteriocin enterocin-B and structural analysis, interaction determination of enterocin-a, B against pathogenic bacteria and human cancer cells. Int J Biol Macromol 116:502–512. https://doi.org/10.1016/j.ijbiomac.2018.05.002
Arias CA, Contreras GA, Murray BE (2010) Management of multidrug-resistant enterococcal infections. Clin Microbiol Infect 16(6):555–562. https://doi.org/10.1111/j.1469-0691.2010.03214.x10.1111/j.1198-743X.2010.03214.x
Arias CA, Murray BE (2012) The rise of the Enterococcus: beyond vancomycin resistance. Nat Rev Microbiol 10(4):266–278. https://doi.org/10.1038/nrmicro2761
Arias CA, Panesso D, McGrath DM, Qin X, Mojica MF, Miller C, Diaz L, Tran TT, Rincon S, Barbu EM, Reyes J, Roh JH, Lobos E, Sodergren E, Pasqualini R, Arap W, Quinn JP, Shamoo Y, Murray BE, Weinstock GM (2011) Genetic basis for in vivo daptomycin resistance in enterococci. N Engl J Med 365(10):892–900. https://doi.org/10.1056/NEJMoa1011138
Brade KD, Rybak JM, Rybak MJ (2016) Oritavancin: a new lipoglycopeptide antibiotic in the treatment of Gram-positive infections. Infect Dis Ther 5(1):1–15. https://doi.org/10.1007/s40121-016-0103-4
Brand AM, de Kwaadsteniet M, Dicks LM (2010) The ability of nisin F to control Staphylococcus aureus infection in the peritoneal cavity, as studied in mice. Lett Appl Microbiol 51(6):645–649. https://doi.org/10.1111/j.1472-765X.2010.02948.x
Brandl K, Plitas G, Mihu CN, Ubeda C, Jia T, Fleisher M, Schnabl B, DeMatteo RP, Pamer EG (2008) Vancomycin-resistant enterococci exploit antibiotic-induced innate immune deficits. Nature 455(7214):804–807. https://doi.org/10.1038/nature07250
Brandl K, Plitas G, Schnabl B, DeMatteo RP, Pamer EG (2007) MyD88-mediated signals induce the bactericidal lectin RegIII gamma and protect mice against intestinal Listeria monocytogenes infection. J Exp Med 204(8):1891–1900. https://doi.org/10.1084/jem.20070563
Brown MS, Ye J, Rawson RB, Goldstein JL (2000) Regulated intramembrane proteolysis: a control mechanism conserved from bacteria to humans. Cell 100(4):391–398
Buffie CG, Pamer EG (2013) Microbiota-mediated colonization resistance against intestinal pathogens. Nat Rev Immunol 13(11):790–801. https://doi.org/10.1038/nri3535
Caballero S, Carter R, Ke X, Susac B, Leiner IM, Kim GJ, Miller L, Ling L, Manova K, Pamer EG (2015) Distinct but spatially overlapping intestinal niches for vancomycin-resistant Enterococcus faecium and carbapenem-resistant Klebsiella pneumoniae. PLoS Pathog 11(9):e1005132. https://doi.org/10.1371/journal.ppat.1005132
Caly DL, Chevalier M, Flahaut C, Cudennec B, Al Atya AK, Chataigne G, D'Inca R, Auclair E, Drider D (2017) The safe enterocin DD14 is a leaderless two-peptide bacteriocin with anti-Clostridium perfringens activity. Int J Antimicrob Agents 49(3):282–289. https://doi.org/10.1016/j.ijantimicag.2016.11.016
Casapao AM, Kullar R, Davis SL, Levine DP, Zhao JJ, Potoski BA, Goff DA, Crank CW, Segreti J, Sakoulas G, Cosgrove SE, Rybak MJ (2013) Multicenter study of high-dose daptomycin for treatment of enterococcal infections. Antimicrob Agents Chemother 57(9):4190–4196. https://doi.org/10.1128/AAC.00526-13
Cebrián R, Rodríguez-Cabezas ME, Martín-Escolano R, Rubiño S, Garrido-Barros M, Montalbán-López M, Rosales MJ, Sánchez-Moreno M, Valdivia E, Martínez-Bueno M, Marín C, Galvez J, Maqueda M (2019) Preclinical studies of toxicity and safety of the AS-48 bacteriocin. J Adv Res. https://doi.org/10.1016/j.jare.2019.06.003
Chi H, Holo H (2018) Synergistic antimicrobial activity between the broad spectrum bacteriocin garvicin KS and nisin, farnesol and polymyxin B against Gram-positive and Gram-negative bacteria. Curr Microbiol 75(3):272–277. https://doi.org/10.1007/s00284-017-1375-y
Chuang YC, Wang JT, Lin HY, Chang SC (2014) Daptomycin versus linezolid for treatment of vancomycin-resistant enterococcal bacteremia: systematic review and meta-analysis. BMC Infect Dis 14:687. https://doi.org/10.1186/s12879-014-0687-9
Cintas LM, Casaus P, Havarstein LS, Hernandez PE, Nes IF (1997) Biochemical and genetic characterization of enterocin P, a novel sec-dependent bacteriocin from Enterococcus faecium P13 with a broad antimicrobial spectrum. Appl Environ Microbiol 63(11):4321–4330
Comerlato CB, Resende MC, Caierao J, d'Azevedo PA (2013) Presence of virulence factors in Enterococcus faecalis and Enterococcus faecium susceptible and resistant to vancomycin. Mem Inst Oswaldo Cruz 108(5):590–595
Cook LC, Federle MJ (2014) Peptide pheromone signaling in Streptococcus and Enterococcus. FEMS Microbiol Rev 38(3):473–492. https://doi.org/10.1111/1574-6976.12046
Cotter PD (2014) An ‘Upp’-turn in bacteriocin receptor identification. Mol Microbiol 92(6):1159–1163. https://doi.org/10.1111/mmi.12645
Cotter PD, Hill C, Ross RP (2005) Bacteriocins: developing innate immunity for food. Nat Rev Microbiol 3(10):777–788. https://doi.org/10.1038/nrmicro1273
Cotter PD, Ross RP, Hill C (2013) Bacteriocins—a viable alternative to antibiotics? Nat Rev Microbiol 11(2):95–105. https://doi.org/10.1038/nrmicro2937
Crank CW, Scheetz MH, Brielmaier B, Rose WE, Patel GP, Ritchie DJ, Segreti J (2010) Comparison of outcomes from daptomycin or linezolid treatment for vancomycin-resistant enterococcal bloodstream infection: a retrospective, multicenter, cohort study. Clin Ther 32(10):1713–1719. https://doi.org/10.1016/j.clinthera.2010.09.008
Crouzet L, Derrien M, Cherbuy C, Plancade S, Foulon M, Chalin B, van Hylckama Vlieg JET, Grompone G, Rigottier-Gois L, Serror P (2018) Lactobacillus paracasei CNCM I-3689 reduces vancomycin-resistant Enterococcus persistence and promotes Bacteroidetes resilience in the gut following antibiotic challenge. Sci Rep 8(1):5098. https://doi.org/10.1038/s41598-018-23437-9
Darwin AJ (2013) Stress relief during host infection: the phage shock protein response supports bacterial virulence in various ways. PLoS Pathog 9(7):e1003388. https://doi.org/10.1371/journal.ppat.1003388
Darwin AJ, Miller VL (1999) Identification of Yersinia enterocolitica genes affecting survival in an animal host using signature-tagged transposon mutagenesis. Mol Microbiol 32(1):51–62
De Kwaadsteniet M, Doeschate KT, Dicks LM (2009) Nisin F in the treatment of respiratory tract infections caused by Staphylococcus aureus. Lett Appl Microbiol 48(1):65–70. https://doi.org/10.1111/j.1472-765X.2008.02488.x
de Kwaadsteniet M, van Reenen CA, Dicks LM (2010) Evaluation of nisin F in the treatment of subcutaneous skin infections, as monitored by using a bioluminescent strain of Staphylococcus aureus. Probiotics Antimicrob Proteins 2(2):61–65. https://doi.org/10.1007/s12602-009-9017-8
Deshpande LM, Ashcraft DS, Kahn HP, Pankey G, Jones RN, Farrell DJ, Mendes RE (2015) Detection of a new cfr-like gene, cfr(B), in Enterococcus faecium isolates recovered from human specimens in the United States as part of the SENTRY antimicrobial surveillance program. Antimicrob Agents Chemother 59(10):6256–6261. https://doi.org/10.1128/AAC.01473-15
Diep DB, Skaugen M, Salehian Z, Holo H, Nes IF (2007) Common mechanisms of target cell recognition and immunity for class II bacteriocins. Proc Natl Acad Sci U S A 104(7):2384–2389. https://doi.org/10.1073/pnas.0608775104
Dina J, Malbruny B, Leclercq R (2003) Nonsense mutations in the lsa-like gene in Enterococcus faecalis isolates susceptible to lincosamides and Streptogramins A. Antimicrob Agents Chemother 47(7):2307–2309
Dobbs TE, Patel M, Waites KB, Moser SA, Stamm AM, Hoesley CJ (2006) Nosocomial spread of Enterococcus faecium resistant to vancomycin and linezolid in a tertiary care medical center. J Clin Microbiol 44(9):3368–3370. https://doi.org/10.1128/JCM.00850-06
Domenech O, Francius G, Tulkens PM, Van Bambeke F, Dufrene Y, Mingeot-Leclercq MP (2009) Interactions of oritavancin, a new lipoglycopeptide derived from vancomycin, with phospholipid bilayers: effect on membrane permeability and nanoscale lipid membrane organization. Biochim Biophys Acta 1788(9):1832–1840. https://doi.org/10.1016/j.bbamem.2009.05.003
Donskey CJ (2004) The role of the intestinal tract as a reservoir and source for transmission of nosocomial pathogens. Clin Infect Dis 39(2):219–226. https://doi.org/10.1086/422002
Donskey CJ, Hanrahan JA, Hutton RA, Rice LB (2000) Effect of parenteral antibiotic administration on the establishment of colonization with vancomycin-resistant Enterococcus faecium in the mouse gastrointestinal tract. J Infect Dis 181(5):1830–1833. https://doi.org/10.1086/315428
Dupont H, Vael C, Muller-Serieys C, Chosidow D, Mantz J, Marmuse JP, Andremont A, Goossens H, Desmonts JM (2008) Prospective evaluation of virulence factors of enterococci isolated from patients with peritonitis: impact on outcome. Diagn Microbiol Infect Dis 60(3):247–253. https://doi.org/10.1016/j.diagmicrobio.2007.10.006
Edmond MB, Ober JF, Dawson JD, Weinbaum DL, Wenzel RP (1996) Vancomycin-resistant enterococcal bacteremia: natural history and attributable mortality. Clin Infect Dis 23(6):1234–1239
Ekblad B, Nissen-Meyer J, Kristensen T (2017) Whole-genome sequencing of mutants with increased resistance against the two-peptide bacteriocin plantaricin JK reveals a putative receptor and potential docking site. PLoS One 12(9):e0185279. https://doi.org/10.1371/journal.pone.0185279
Ellermeier CD, Losick R (2006) Evidence for a novel protease governing regulated intramembrane proteolysis and resistance to antimicrobial peptides in Bacillus subtilis. Genes Dev 20(14):1911–1922. https://doi.org/10.1101/gad.1440606
Fantin B, Leclercq R, Garry L, Carbon C (1997) Influence of inducible cross-resistance to macrolides, lincosamides, and streptogramin B-type antibiotics in Enterococcus faecium on activity of quinupristin-dalfopristin in vitro and in rabbits with experimental endocarditis. Antimicrob Agents Chemother 41(5):931–935
Frank KL, Barnes AM, Grindle SM, Manias DA, Schlievert PM, Dunny GM (2012) Use of recombinase-based in vivo expression technology to characterize Enterococcus faecalis gene expression during infection identifies in vivo-expressed antisense RNAs and implicates the protease Eep in pathogenesis. Infect Immun 80(2):539–549. https://doi.org/10.1128/IAI.05964-11
Gabrielsen C, Brede DA, Nes IF, Diep DB (2014) Circular bacteriocins: biosynthesis and mode of action. Appl Environ Microbiol 80(22):6854–6862. https://doi.org/10.1128/AEM.02284-14
Garcia De Gonzalo CV, Denham EL, Mars RA, Stulke J, van der Donk WA, van Dijl JM (2015) The phosphoenolpyruvate:sugar phosphotransferase system is involved in sensitivity to the glucosylated bacteriocin sublancin. Antimicrob Agents Chemother 59(11):6844–6854. https://doi.org/10.1128/AAC.01519-15
Graham CE, Cruz MR, Garsin DA, Lorenz MC (2017) Enterococcus faecalis bacteriocin EntV inhibits hyphal morphogenesis, biofilm formation, and virulence of Candida albicans. Proc Natl Acad Sci U S A 114(17):4507–4512. https://doi.org/10.1073/pnas.1620432114
Greer ND (2006) Tigecycline (Tygacil): the first in the glycylcycline class of antibiotics. Proc (Bayl Univ Med Cent) 19(2):155–161
Hanchi H, Hammami R, Gingras H, Kourda R, Bergeron MG, Ben Hamida J, Ouellette M, Fliss I (2017) Inhibition of MRSA and of Clostridium difficile by durancin 61A: synergy with bacteriocins and antibiotics. Future Microbiol 12:205–212. https://doi.org/10.2217/fmb-2016-0113
Hayes K, Cotter L, O'Halloran F (2019) In-vitro synergistic activity of erythromycin and nisin against clinical Group B Streptococcus isolates. J Appl Microbiol. https://doi.org/10.1111/jam.14400
Hefazi M, Damlaj M, Alkhateeb HB, Partain DK, Patel R, Razonable RR, Gastineau DA, Al-Kali A, Hashmi SK, Hogan WJ, Litzow MR, Patnaik MM (2016) Vancomycin-resistant Enterococcus colonization and bloodstream infection: prevalence, risk factors, and the impact on early outcomes after allogeneic hematopoietic cell transplantation in patients with acute myeloid leukemia. Transpl Infect Dis 18(6):913–920. https://doi.org/10.1111/tid.12612
Hegstad K, Mikalsen T, Coque TM, Werner G, Sundsfjord A (2010) Mobile genetic elements and their contribution to the emergence of antimicrobial resistant Enterococcus faecalis and Enterococcus faecium. Clin Microbiol Infect 16(6):541–554. https://doi.org/10.1111/j.1469-0691.2010.03226.x
Heikens E, Bonten MJ, Willems RJ (2007) Enterococcal surface protein Esp is important for biofilm formation of Enterococcus faecium E1162. J Bacteriol 189(22):8233–8240. https://doi.org/10.1128/JB.01205-07
Heikens E, Leendertse M, Wijnands LM, van Luit-Asbroek M, Bonten MJ, van der Poll T, Willems RJ (2009) Enterococcal surface protein Esp is not essential for cell adhesion and intestinal colonization of Enterococcus faecium in mice. BMC Microbiol 9:19. https://doi.org/10.1186/1471-2180-9-19
Hendrickx AP, van Wamel WJ, Posthuma G, Bonten MJ, Willems RJ (2007) Five genes encoding surface-exposed LPXTG proteins are enriched in hospital-adapted Enterococcus faecium clonal complex 17 isolates. J Bacteriol 189(22):8321–8332. https://doi.org/10.1128/JB.00664-07
Higgins DL, Chang R, Debabov DV, Leung J, Wu T, Krause KM, Sandvik E, Hubbard JM, Kaniga K, Schmidt DE Jr, Gao Q, Cass RT, Karr DE, Benton BM, Humphrey PP (2005) Telavancin, a multifunctional lipoglycopeptide, disrupts both cell wall synthesis and cell membrane integrity in methicillin-resistant Staphylococcus aureus. Antimicrob Agents Chemother 49(3):1127–1134. https://doi.org/10.1128/AAC.49.3.1127-1134.2005
Ho TD, Ellermeier CD (2012) Extra cytoplasmic function sigma factor activation. Curr Opin Microbiol 15(2):182–188. https://doi.org/10.1016/j.mib.2012.01.001
Ispirli H, Demirbas F, Dertli E (2015) Characterization of functional properties of Enterococcus faecium strains isolated from human gut. Can J Microbiol 61(11):861–870. https://doi.org/10.1139/cjm-2015-0446
Johnsen L, Fimland G, Eijsink V, Nissen-Meyer J (2000) Engineering increased stability in the antimicrobial peptide pediocin PA-1. Appl Environ Microbiol 66(11):4798–4802. https://doi.org/10.1128/aem.66.11.4798-4802.2000
Johnson JA, Feeney ER, Kubiak DW, Corey GR (2015) Prolonged use of oritavancin for vancomycin-resistant Enterococcus faecium prosthetic valve endocarditis. Open Forum Infect Dis 2(4):ofv156. https://doi.org/10.1093/ofid/ofv156
Jordan S, Hutchings MI, Mascher T (2008) Cell envelope stress response in Gram-positive bacteria. FEMS Microbiol Rev 32(1):107–146. https://doi.org/10.1111/j.1574-6976.2007.00091.x
Kamboj M, Cohen N, Gilhuley K, Babady NE, Seo SK, Sepkowitz KA (2011) Emergence of daptomycin-resistant VRE: experience of a single institution. Infect Control Hosp Epidemiol 32(4):391–394. https://doi.org/10.1086/659152
Kanehara K, Akiyama Y, Ito K (2001) Characterization of the yaeL gene product and its S2P-protease motifs in Escherichia coli. Gene 281(1–2):71–79
Kara A, Devrim I, Bayram N, Katipoglu N, Kiran E, Oruc Y, Demiray N, Apa H, Gulfidan G (2015) Risk of vancomycin-resistant enterococci bloodstream infection among patients colonized with vancomycin-resistant enterococci. Braz J Infect Dis 19(1):58–61. https://doi.org/10.1016/j.bjid.2014.09.010
Kristich CJ, Rice LB, Arias CA (2014) Enterococcal infection—treatment and antibiotic resistance. In: Gilmore MS, Clewell DB, Ike Y, Shankar N (eds) Enterococci: from commensals to leading causes of drug resistant infection, Boston. https://www.ncbi.nlm.nih.gov/books/NBK190424/
Lam MM, Seemann T, Bulach DM, Gladman SL, Chen H, Haring V, Moore RJ, Ballard S, Grayson ML, Johnson PD, Howden BP, Stinear TP (2012) Comparative analysis of the first complete Enterococcus faecium genome. J Bacteriol 194(9):2334–2341. https://doi.org/10.1128/JB.00259-12
Leavis HL, Willems RJ, van Wamel WJ, Schuren FH, Caspers MP, Bonten MJ (2007) Insertion sequence-driven diversification creates a globally dispersed emerging multiresistant subspecies of E. faecium. PLoS Pathog 3(1):e7. https://doi.org/10.1371/journal.ppat.0030007
Leendertse M, Heikens E, Wijnands LM, van Luit-Asbroek M, Teske GJ, Roelofs JJ, Bonten MJ, van der Poll T, Willems RJ (2009) Enterococcal surface protein transiently aggravates Enterococcus faecium-induced urinary tract infection in mice. J Infect Dis 200(7):1162–1165. https://doi.org/10.1086/605609
Lindenstrauss AG, Pavlovic M, Bringmann A, Behr J, Ehrmann MA, Vogel RF (2011) Comparison of genotypic and phenotypic cluster analyses of virulence determinants and possible role of CRISPR elements towards their incidence in Enterococcus faecalis and Enterococcus faecium. Syst Appl Microbiol 34(8):553–560. https://doi.org/10.1016/j.syapm.2011.05.002
Lopez F, Culebras E, Betriu C, Rodriguez-Avial I, Gomez M, Picazo JJ (2010) Antimicrobial susceptibility and macrolide resistance genes in Enterococcus faecium with reduced susceptibility to quinupristin-dalfopristin: level of quinupristin-dalfopristin resistance is not dependent on erm(B) attenuator region sequence. Diagn Microbiol Infect Dis 66(1):73–77. https://doi.org/10.1016/j.diagmicrobio.2008.06.004
Maraki S, Samonis G, Dimopoulou D, Mantadakis E (2014) Susceptibility of glycopeptide-resistant enterococci to linezolid, quinupristin/dalfopristin, tigecycline and daptomycin in a tertiary Greek hospital. Infect Chemother 46(4):253–256. https://doi.org/10.3947/ic.2014.46.4.253
Mave V, Garcia-Diaz J, Islam T, Hasbun R (2009) Vancomycin-resistant enterococcal bacteraemia: is daptomycin as effective as linezolid? J Antimicrob Chemother 64(1):175–180. https://doi.org/10.1093/jac/dkp154
McLaughlin M, Malczynski M, Qi C, Barajas G, Radetski J, Zembower T, Scheetz MH (2013) Virulence of vancomycin-resistant Enterococcus faecium according to linezolid resistance and clinical outbreak status. Antimicrob Agents Chemother 57(8):3923–3927. https://doi.org/10.1128/AAC.00192-13
Mendes RE, Deshpande LM, Jones RN (2014) Linezolid update: stable in vitro activity following more than a decade of clinical use and summary of associated resistance mechanisms. Drug Resist Updat 17(1–2):1–12. https://doi.org/10.1016/j.drup.2014.04.002
Mendes RE, Flamm RK, Hogan PA, Ross JE, Jones RN (2014) Summary of linezolid activity and resistance mechanisms detected during the 2012 LEADER surveillance program for the United States. Antimicrob Agents Chemother 58(2):1243–1247. https://doi.org/10.1128/AAC.02112-13
Millette M, Cornut G, Dupont C, Shareck F, Archambault D, Lacroix M (2008) Capacity of human nisin- and pediocin-producing lactic acid bacteria to reduce intestinal colonization by vancomycin-resistant enterococci. Appl Environ Microbiol 74(7):1997–2003. https://doi.org/10.1128/AEM.02150-07
Miyazaki S, Fujikawa T, Kobayashi I, Matsumoto T, Tateda K, Yamaguchi K (2001) Development of systemic bacteraemia after oral inoculation of vancomycin-resistant enterococci in mice. J Med Microbiol 50(8):695–701. https://doi.org/10.1099/0022-1317-50-8-695
Moise PA, Hershberger E, Amodio-Groton MI, Lamp KC (2009) Safety and clinical outcomes when utilizing high-dose (> or = 8 mg/kg) daptomycin therapy. Ann Pharmacother 43(7):1211–1219. https://doi.org/10.1345/aph.1M085
Munch D, Engels I, Muller A, Reder-Christ K, Falkenstein-Paul H, Bierbaum G, Grein F, Bendas G, Sahl HG, Schneider T (2015) Structural variations of the cell wall precursor lipid II and their influence on binding and activity of the lipoglycopeptide antibiotic oritavancin. Antimicrob Agents Chemother 59(2):772–781. https://doi.org/10.1128/AAC.02663-14
Murray BE (2000) Vancomycin-resistant enterococcal infections. N Engl J Med 342(10):710–721. https://doi.org/10.1056/NEJM200003093421007
Nallapareddy SR, Weinstock GM, Murray BE (2003) Clinical isolates of Enterococcus faecium exhibit strain-specific collagen binding mediated by Acm, a new member of the MSCRAMM family. Mol Microbiol 47(6):1733–1747
O'Driscoll T, Crank CW (2015) Vancomycin-resistant enterococcal infections: epidemiology, clinical manifestations, and optimal management. Infect Drug Resist 8:217–230. https://doi.org/10.2147/IDR.S54125
Oppegard C, Kjos M, Veening JW, Nissen-Meyer J, Kristensen T (2016) A putative amino acid transporter determines sensitivity to the two-peptide bacteriocin plantaricin JK. Microbiologyopen 5(4):700–708. https://doi.org/10.1002/mbo3.363
Ovchinnikov KV, Chi H, Mehmeti I, Holo H, Nes IF, Diep DB (2016) Novel group of leaderless multipeptide bacteriocins from Gram-positive bacteria. Appl Environ Microbiol 82(17):5216–5224. https://doi.org/10.1128/AEM.01094-16
Ovchinnikov KV, Kristiansen PE, Straume D, Jensen MS, Aleksandrzak-Piekarczyk T, Nes IF, Diep DB (2017) The leaderless bacteriocin enterocin K1 is highly potent against Enterococcus faecium: a study on structure, target spectrum and receptor. Front Microbiol 8:774. https://doi.org/10.3389/fmicb.2017.00774
Ovchinnikov KV, Kristiansen PE, Uzelac G, Topisirovic L, Kojic M, Nissen-Meyer J, Nes IF, Diep DB (2014) Defining the structure and receptor binding domain of the leaderless bacteriocin LsbB. J Biol Chem 289(34):23838–23845. https://doi.org/10.1074/jbc.M114.579698
Palmer KL, Kos VN, Gilmore MS (2010) Horizontal gene transfer and the genomics of enterococcal antibiotic resistance. Curr Opin Microbiol 13(5):632–639. https://doi.org/10.1016/j.mib.2010.08.004
Patel R, Gallagher JC (2015) Vancomycin-resistant enterococcal bacteremia pharmacotherapy. Ann Pharmacother 49(1):69–85. https://doi.org/10.1177/1060028014556879
Patti GJ, Kim SJ, Yu TY, Dietrich E, Tanaka KS, Parr TR Jr, Far AR, Schaefer J (2009) Vancomycin and oritavancin have different modes of action in Enterococcus faecium. J Mol Biol 392(5):1178–1191. https://doi.org/10.1016/j.jmb.2009.06.064
Perez RH, Zendo T, Sonomoto K (2014) Novel bacteriocins from lactic acid bacteria (LAB): various structures and applications. Microb Cell Fact 13(Suppl 1):S3. https://doi.org/10.1186/1475-2859-13-S1-S3
Phumisantiphong U, Siripanichgon K, Reamtong O, Diraphat P (2017) A novel bacteriocin from Enterococcus faecalis 478 exhibits a potent activity against vancomycin-resistant enterococci. PLoS One 12(10):e0186415. https://doi.org/10.1371/journal.pone.0186415
Prematunge C, MacDougall C, Johnstone J, Adomako K, Lam F, Robertson J, Garber G (2016) VRE and VSE bacteremia outcomes in the era of effective VRE therapy: a systematic review and meta-analysis. Infect Control Hosp Epidemiol 37(1):26–35. https://doi.org/10.1017/ice.2015.228
Pultz NJ, Shankar N, Baghdayan AS, Donskey CJ (2005) Enterococcal surface protein Esp does not facilitate intestinal colonization or translocation of Enterococcus faecalis in clindamycin-treated mice. FEMS Microbiol Lett 242(2):217–219. https://doi.org/10.1016/j.femsle.2004.11.006
Pultz NJ, Vesterlund S, Ouwehand AC, Donskey CJ (2006) Adhesion of vancomycin-resistant enterococcus to human intestinal mucus. Curr Microbiol 52(3):221–224. https://doi.org/10.1007/s00284-005-0244-2
Purohit G, Gaind R, Dawar R, Verma PK, Aggarwal KC, Sardana R, Deb M (2017) Characterization of vancomycin resistant enterococci in hospitalized patients and role of gut colonization. J Clin Diagn Res 11(9):DC01–DC05. https://doi.org/10.7860/JCDR/2017/25988.10548
Raad II, Hanna HA, Hachem RY, Dvorak T, Arbuckle RB, Chaiban G, Rice LB (2004) Clinical-use-associated decrease in susceptibility of vancomycin-resistant Enterococcus faecium to linezolid: a comparison with quinupristin-dalfopristin. Antimicrob Agents Chemother 48(9):3583–3585. https://doi.org/10.1128/AAC.48.9.3583-3585.2004
Raivio TL, Silhavy TJ (2001) Periplasmic stress and ECF sigma factors. Annu Rev Microbiol 55:591–624. https://doi.org/10.1146/annurev.micro.55.1.591
Ramu R, Shirahatti PS, Devi AT, Prasad A, J K, M S L, F Z, B LD, M N N (2015) Bacteriocins and their applications in food preservation. Crit Rev Food Sci Nutr 0. https://doi.org/10.1080/10408398.2015.1020918
Rathnayake IU, Hargreaves M, Huygens F (2012) Antibiotic resistance and virulence traits in clinical and environmental Enterococcus faecalis and Enterococcus faecium isolates. Syst Appl Microbiol 35(5):326–333. https://doi.org/10.1016/j.syapm.2012.05.004
Riley MA, Wertz JE (2002) Bacteriocins: evolution, ecology, and application. Annu Rev Microbiol 56:117–137. https://doi.org/10.1146/annurev.micro.56.012302.161024
Roger C, Roberts JA, Muller L (2018) Clinical pharmacokinetics and pharmacodynamics of oxazolidinones. Clin Pharmacokinet 57(5):559–575. https://doi.org/10.1007/s40262-017-0601-x
Rosa RG, Schwarzbold AV, Dos Santos RP, Turra EE, Machado DP, Goldani LZ (2014) Vancomycin-resistant Enterococcus faecium bacteremia in a tertiary care hospital: epidemiology, antimicrobial susceptibility, and outcome. Biomed Res Int 2014:958469. https://doi.org/10.1155/2014/958469
Rosko AE, Corriveau M, Suwantarat N, Arfons L, Treasure M, Parker P, Jacobs M, Fu P, Salata R, Lazarus HM (2014) Vancomycin-resistant enterococci infection: not just for the transplanted. Leuk Lymphoma 55(6):1320–1325. https://doi.org/10.3109/10428194.2013.842983
Sader HS, Farrell DJ, Flamm RK, Jones RN (2014) Daptomycin activity tested against 164457 bacterial isolates from hospitalised patients: summary of 8 years of a worldwide surveillance programme (2005–2012). Int J Antimicrob Agents 43(5):465–469. https://doi.org/10.1016/j.ijantimicag.2014.01.018
Sader HS, Jones RN (2009) Antimicrobial susceptibility of Gram-positive bacteria isolated from US medical centers: results of the Daptomycin surveillance program (2007–2008). Diagn Microbiol Infect Dis 65(2):158–162. https://doi.org/10.1016/j.diagmicrobio.2009.06.016
Sahm DF, Deane J, Bien PA, Locke JB, Zuill DE, Shaw KJ, Bartizal KF (2015) Results of the surveillance of Tedizolid activity and resistance program: in vitro susceptibility of Gram-positive pathogens collected in 2011 and 2012 from the United States and Europe. Diagn Microbiol Infect Dis 81(2):112–118. https://doi.org/10.1016/j.diagmicrobio.2014.08.011
Saleh-Mghir A, Lefort A, Petegnief Y, Dautrey S, Vallois JM, Le Guludec D, Carbon C, Fantin B (1999) Activity and diffusion of LY333328 in experimental endocarditis due to vancomycin-resistant Enterococcus faecalis. Antimicrob Agents Chemother 43(1):115–120
Salgado CD, Farr BM (2003) Outcomes associated with vancomycin-resistant enterococci: a meta-analysis. Infect Control Hosp Epidemiol 24(9):690–698. https://doi.org/10.1086/502271
Sillanpaa J, Nallapareddy SR, Singh KV, Prakash VP, Fothergill T, Ton-That H, Murray BE (2010) Characterization of the ebp(fm) pilus-encoding operon of Enterococcus faecium and its role in biofilm formation and virulence in a murine model of urinary tract infection. Virulence 1(4):236–246
Silva CCG, Silva SPM, Ribeiro SC (2018) Application of bacteriocins and protective cultures in dairy food preservation. Front Microbiol 9:594. https://doi.org/10.3389/fmicb.2018.00594
Singh AP, Prabha V, Rishi P (2013) Value addition in the efficacy of conventional antibiotics by nisin against Salmonella. PLoS One 8(10):e76844. https://doi.org/10.1371/journal.pone.0076844
Steckbeck JD, Deslouches B, Montelaro RC (2014) Antimicrobial peptides: new drugs for bad bugs? Expert Opin Biol Ther 14(1):11–14. https://doi.org/10.1517/14712598.2013.844227
Stiefel U, Pultz NJ, Helfand MS, Donskey CJ (2004) Increased susceptibility to vancomycin-resistant Enterococcus intestinal colonization persists after completion of anti-anaerobic antibiotic treatment in mice. Infect Control Hosp Epidemiol 25(5):373–379. https://doi.org/10.1086/502408
Surveillance of antimicrobial resistance in Europe 2016. (2017). Surveillance of antimicrobial resistance in Europe 2016
Thomas VM, Brown RM, Ashcraft DS, Pankey GA (2019) Synergistic effect between nisin and polymyxin B against pandrug-resistant and extensively drug-resistant Acinetobacter baumannii. Int J Antimicrob Agents 53(5):663–668. https://doi.org/10.1016/j.ijantimicag.2019.03.009
Tran TT, Panesso D, Mishra NN, Mileykovskaya E, Guan Z, Munita JM, Reyes J, Diaz L, Weinstock GM, Murray BE, Shamoo Y, Dowhan W, Bayer AS, Arias CA (2013) Daptomycin-resistant Enterococcus faecalis diverts the antibiotic molecule from the division septum and remodels cell membrane phospholipids. MBio 4(4). https://doi.org/10.1128/mBio.00281-13
Twilla JD, Finch CK, Usery JB, Gelfand MS, Hudson JQ, Broyles JE (2012) Vancomycin-resistant Enterococcus bacteremia: an evaluation of treatment with linezolid or daptomycin. J Hosp Med 7(3):243–248. https://doi.org/10.1002/jhm.994
Ubeda C, Taur Y, Jenq RR, Equinda MJ, Son T, Samstein M, Viale A, Socci ND, van den Brink MRM, Kamboj M, Pamer EG (2010) Vancomycin-resistant Enterococcus domination of intestinal microbiota is enabled by antibiotic treatment in mice and precedes bloodstream invasion in humans. J Clin Invest 120(12):4332–4341. https://doi.org/10.1172/Jci43918
van Staden AD, Brand AM, Dicks LM (2012) Nisin F-loaded brushite bone cement prevented the growth of Staphylococcus aureus in vivo. J Appl Microbiol 112(4):831–840. https://doi.org/10.1111/j.1365-2672.2012.05241.x
van Staden AP, Heunis T, Smith C, Deane S, Dicks LM (2016) Efficacy of lantibiotic treatment of Staphylococcus aureus-induced skin infections, monitored by in vivo bioluminescent imaging. Antimicrob Agents Chemother 60(7):3948–3955. https://doi.org/10.1128/AAC.02938-15
Vergis EN, Hayden MK, Chow JW, Snydman DR, Zervos MJ, Linden PK, Wagener MM, Schmitt B, Muder RR (2001) Determinants of vancomycin resistance and mortality rates in enterococcal bacteremia: a prospective multicenter study. Ann Intern Med 135(7):484–492
Willems RJ, Top J, van Santen M, Robinson DA, Coque TM, Baquero F, Grundmann H, Bonten MJ (2005) Global spread of vancomycin-resistant Enterococcus faecium from distinct nosocomial genetic complex. Emerg Infect Dis 11(6):821–828. https://doi.org/10.3201/eid1106.041204
Wolska KI, Grzes K, Kurek A (2012) Synergy between novel antimicrobials and conventional antibiotics or bacteriocins. Pol J Microbiol 61(2):95–104
Yang SC, Lin CH, Sung CT, Fang JY (2014) Antibacterial activities of bacteriocins: application in foods and pharmaceuticals. Front Microbiol 5:241. https://doi.org/10.3389/fmicb.2014.00241
Yim J, Smith JR, Rybak MJ (2017) Role of combination antimicrobial therapy for vancomycin-resistant Enterococcus faecium infections: review of the current evidence. Pharmacotherapy 37(5):579–592. https://doi.org/10.1002/phar.1922
Zasowski EJ, Claeys KC, Lagnf AM, Davis SL, Rybak MJ (2016) Time is of the essence: the impact of delayed antibiotic therapy on patient outcomes in hospital-onset enterococcal bloodstream infections. Clin Infect Dis 62(10):1242–1250. https://doi.org/10.1093/cid/ciw110
Zhou L, van Heel AJ, Montalban-Lopez M, Kuipers OP (2016) Potentiating the activity of nisin against Escherichia coli. Front Cell Dev Biol 4:7. https://doi.org/10.3389/fcell.2016.00007
Funding
This work was supported by grants from the Research Council of Norway (project numbers 275190 and 273646) and from the Bio-based Industries Joint Undertaking under the European Union’s Horizon 2020 research and innovation programme under grant agreement no. 790507.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original version of this article was revised due to Retrospective Open Access.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Reinseth, I.S., Ovchinnikov, K.V., Tønnesen, H.H. et al. The Increasing Issue of Vancomycin-Resistant Enterococci and the Bacteriocin Solution. Probiotics & Antimicro. Prot. 12, 1203–1217 (2020). https://doi.org/10.1007/s12602-019-09618-6
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12602-019-09618-6