Abstract
The objective of this study was to characterise the antagonistic activity of cellular components of potential probiotic bacteria isolated from the gut of healthy rohu (Labeo rohita), a tropical freshwater fish, against the fish pathogen, Aeromonas hydrophila. Three potential probiotic strains (referred to as R1, R2, and R5) were screened using a well diffusion, and their antagonistic activity against A. hydrophila was determined. Biochemical tests and 16S rRNA gene analysis confirmed that R1, R2, and R5 were Lactobacillus plantarum VSG3, Pseudomonas aeruginosa VSG2, and Bacillus subtilis VSG1, respectively. Four different fractions of cellular components (i.e. the whole-cell product, heat-killed whole-cell product [HKWCP], intracellular product [ICP], and extracellular product) of these selected strains were effective in an in vitro sensitivity test against 6 A. hydrophila strains. Among the cellular components, the ICP of R1, HKWCP of R2, and ICP of R5 exhibited the strongest antagonistic activities, as evidenced by their inhibition zones. The antimicrobial compounds from these selected cellular components were partially purified by thin-layer and high-performance liquid chromatography, and their properties were analysed. The ranges of pH stability of the purified compounds were wide (3.0–10.0), and compounds were thermally stable up to 90 °C. Considering these results, isolated probiotic strains may find potential applications in the prevention and treatment of aquatic aeromonosis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacterial infections are considered a major cause of mortality in fish hatcheries. Aeromonas hydrophila is a pathogen that affects a wide variety of freshwater fish species [1]. Combined with the problem of antibiotic contamination of aquaculture facilities and livestock, the indiscriminate worldwide use of antibiotics in aquaculture has led to the development of drug-resistant bacteria that are becoming increasingly difficult to control and eradicate [6, 28]. An alternative disease prevention method that involves the use of non-pathogenic bacteria as probiotics is proposed [2]. The role of lactic acid bacteria as probiotics in the digestive tract has been extensively studied in endothermic animals and humans [4, 26]. Lactic acid bacteria were recently described as part of the normal microbiota in freshwater fish [26, 35, 36]. The production of antimicrobial substances by Bacillus species isolated from Japanese costal fish, and their use as a biocontrol agent, is reported [30, 31]. The antagonistic activity of Pseudomonas against a number of pathogens (e.g. Aeromonas and Vibrio sp.) has also been documented [8, 37]. Furthermore, probiotic treatment leads to better protection of fish against multiple diseases [1].
Bacteria in aquatic ecosystems might produce antimicrobial substances inhibiting the growth of other microorganisms [12]. Bacterial strains isolated from culture medium might exhibit antagonistic effects on present pathogenic bacteria [3, 17]. For instance, bacterial strains isolated from the internal organs of Oreochromis niloticus exhibit inhibitory effects against A. hydrophila [1]. Even some bacteria are antagonistic to viruses [9]. Several proposed theories explain the antimicrobial and beneficial effects of probiotics, including their ability to secrete antimicrobial substances such as organic acids, bacteriocins, and peptides [20].
Probiotics is a big business today in Indian aquaculture; it is worth $109 million, and most supplies are imported. Indian fish pathologists are looking at probiotics as a potentially useful disease prevention measure in aquatic farms, and active research is continuing in this regard [24]. However, the efficiency of probiotic isolates from tropical freshwater species is less studied and needs further exploration.
Thanjavur is a tropical district situated at the southern part of Tamil Nadu (N 10°48′0″, E 79°9′0″) and is well known for its marine and freshwater fish production. To our knowledge, there is no report on the probiotic efficiency of intestinal microbiota of tropical freshwater fish. In this study, we explored for the first time in vitro antagonistic activity of the cellular components of potential probiotic bacteria isolated from the gut of tropical freshwater fish species, rohu (Labeo rohita), against A. hydrophila. Furthermore, the purification and partial characterisation of antimicrobial compounds present in the selected cellular components are described.
Materials and Methods
Isolation and Screening of Potential Probiotic Bacteria
Ten L. rohita (mean bodyweight, 154.2 ± 1.01 g; mean length, 16.56 ± 0.1 cm) were collected from the Mannal Aquatic Habitation, Thanjavur, Tamil Nadu. The fishes were killed, and the surfaces were washed with 0.1% benzalkonium chloride for 1 min to remove external bacteria. Under sterile conditions, the gut region was dissected and homogenised with an appropriate volume of sterile 1% peptone water. Serial dilutions were made up to 10−6 dilution, from which 100-μL aliquots were spread on tryptone soya agar plates and incubated for 24 h at 37 °C. A total of 389 colonies were counted and designated as L1–L389. Among them, 140 colonies were selected randomly to test antimicrobial activity against A. hydrophila.
Briefly, A. hydrophila cultures were prepared by pouring 2 mL inoculum (103 CFU/mL) onto Tryptone Soya Agar plates to completely cover the surface of the agar. Excess solution was removed and drained before air-drying for 15 min. Wells 6 mm in diameter were punctured into the agar, and 20 μL of each bacterial inoculum (103 CFU/mL) was carefully transferred into separate wells. Two bacterial isolates were tested per plate in triplicate and incubated for 24–48 h at 37 °C. Only 7 isolates inhibited the growth of A. hydrophila; these 7 isolates were designated R1, R2, R3, R4, R5, R6, and R7. Strains with greater inhibition zone diameters were also selected for further study.
Pathogen Collection and Culture Conditions
Six A. hydrophila strains (ATCC-7965, ATCC-23213, ATCC-23214, and ATCC-21763; MTCC-646 and MTCC-1739) were obtained from the American Type Culture Collection (USA) and the Microbial Type Culture Collection (Chandigarh, India). After 24 h incubation in nutrient broth, A. hydrophila strains were streaked on Rimler–Shott’s medium (Hi-media, Mumbai, India) to determine their purity. The pathogenicity of the strains was checked experimentally by injecting them into L. rohita and by observing the onset of disease in the fish.
The A. hydrophila strains and isolated probiotic bacteria were maintained in the laboratory under standard conditions. Cultures were maintained on tryptone soya agar (Hi-media, Mumbai) slopes at 4 °C. Stock cultures in tryptone soya broth were stored at −70 °C in 0.85% NaCl with 20% glycerol to provide a stable inoculum throughout the study [32].
Preparation of Cellular Components
Four different cellular components were prepared from strains R1, R2, and R5: the whole-cell product (WCP), heat-killed whole-cell product (HKWCP), intracellular product (ICP), and extracellular product (ECP). Pure cultures of the 3 strains were maintained separately under sterile conditions in 400-mL brain heart infusion broth (pH 7.0) (Hi media, India) at 37 °C for 24 h. A pure culture of each strain was divided into 4 equal volumes of 100 mL, and each volume was taken for the preparation to determine the WCP, HKWCP, ICP, and ECP. The protein content of the fractions was estimated, as described by Bardford [5].
WCP Preparation
The WCP was prepared as described by Das et al. [8]. After being incubated for 24 h, 100 mL of brain heart infusion broth of each strain (R1, R2, or R5) was centrifuged at 10,000 rpm for 10 min at 4 °C. The pellet obtained after centrifugation was washed twice with phosphate buffer saline (pH 7.2) and resuspended in 5 mL of the same buffer and then stored at −20 °C prior to use.
HKWCP Preparation
The HKWCP was prepared with minor modification of the procedure outlined by Das et al. [8]. Briefly, the pellet obtained after centrifugation was washed twice with PBS (pH 7.2) and resuspended in 5 mL of the same buffer solution. The buffer solution was put in an 80 °C water bath for 30 min, subsequently cooled, and stored at −20 °C until use.
ICP Preparation
The pellet obtained after centrifuging a 24-h-old broth culture of bacteria was washed twice in PBS (pH 7.2). The pellet was resuspended in PBS and then sonicated at 50 Hz for 5 min (Ultrasonic Processor, AICIL, Chandigarh, India), filtered through a 0.45-μm Millipore filter (Millipore, Bedford, USA), and finally stored at −20 °C for further use.
Collection of the ECP
The ECP was prepared as described by Das et al. [8] with minor modification. Supernatants obtained after the centrifugation of 24-h-old cultures of bacteria were filtered through a 0.22-μm polycarbonate filter (Millipore, USA), further concentrated with a freeze dryer (INDLAB, Chennai, India), and were used as ECP.
Study of Antagonistic Activity Using Cellular Components
Tryptone soya broth cultures were prepared freshly from the A. hydrophila agar slopes (stored at 4 °C) and were incubated on tryptone soya agar plates (diameter, 98 mm) separately using a lawn culture technique [8]. Then, a 6-mm-diameter well was made in each plate with the help of a cork borer. A 100-μL aliquot of different cellular components was charged in the respective wells and incubated at 37 °C for 24 h. The zones of inhibition around the wells were recorded (in millimetres) after the incubation. For each cellular component, averages were calculated for 3 wells. Control plates were simultaneously maintained with sterile phosphate buffer saline (pH 7.2) poured into respective wells prepared as mentioned above. For all cellular components, triplicate plates were maintained along with the PBS control for the biocontrol study.
A positive control study was carried out using chloramphenicol (30 mcg) and gentamicin (15 mcg). Plates were incubated at 37 °C for 24 h and observed. The zones of inhibition for each cellular component were compared with those of antibiotics against A. hydrophila.
Identification of Bacteria
Three potential probiotic strains (R1, R2, and R5) were identified on the basis of morphological, physiological, and biochemical characterisations as well as 16S rRNA gene sequencing.
For 16S rRNA gene sequencing, chromosomal DNA was extracted and purified using the phenol–chloroform extraction method [29]. PCR amplification was carried out with universal bacterial primers as described previously: forward, 5′-GAGTTTGATCCTGGCTCA-3′ and reverse, 5′-CGGCTACCTTGTTACGACTT-3′ [33]. The PCR products were purified using the QIAquik PCR Purification Kit (Qiagen) and sequenced using automatic ABI 310 DNA Sequencer (Big Dye Terminator Cycle Sequencing Ready Reaction Kit, Perkin Elmer). The Basic Local Alignment Search Tool (BLAST) from the NCBI (http://www.ncbi.nlm.nih.gov/) was used for nucleotide comparison for percentage similarity.
Purification of Antimicrobial Compounds from the Most Effective Cellular Fractions
From each probiotic bacteria, 4 cellular components were prepared and tested for their antagonistic activity. Antagonistic activity was observed to be the highest with the ICPs of R1 and R5; in R2, the HKWCP exhibited the highest activity. Hence, the ICPs of R1 and R5, and HKWCP of R2 were selected for further analysis and were designated ICP-R1, ICP-R5, and HKWCP-R2, respectively. The selected fractions were partially purified by thin-layer chromatography (TLC) using a 1-mm pre-coated TLC silica gel 60 F254 plate (Merck, Darmstadt, Germany). The plates were developed with either n-butanol/acetic acid/water (5:3:1, v/v/v) or chloroform, methanol, and 0.1% trifluoroacetic acid in water (170:120:15, v/v/v). All plates were run in triplicate. The spots were developed by using iodine vapour/ninhydrin (0.1% w/v in water-saturated n-butanol)/silver nitrate and observed under ultraviolet light (254 nm). Each marked spot was scraped off and eluted with methanol/acetone/chloroform. The supernatants obtained after centrifugation were checked for antibacterial activity, concentrated, and stored at 4 °C for further analysis.
The antimicrobial compounds were further purified by reverse-phase high-performance liquid chromatography (HPLC) on a C18 column (11 × 300 mm) (PerkinElmer, Shelton. CT). The concentrated sample obtained from TLC was filtered through a 0.45-μm polytetrafluoroethylene (PTFE) membrane (Schleicher and Schuell, Keene, USA). Fractions of 20 μL were eluted at a flow rate of 0.5 mL/min for 30 min. An isocratic system of water/acetonitrile (60:40) was used as a mobile phase for the ICP-R1 and ICP-R5 fraction, whereas water/methanol (60:40) was used for HKWCP-R2. The eluent patterns of the fractions were monitored at 220–284 nm and collected manually.
Partial Characterisations of Antimicrobial Compounds
Effects of pH, Temperature, and Enzymes on the Activities of Antimicrobial Compounds
To determine the pH stability of these components, purified antimicrobial samples (ICP-R1, ICP-R5, and HKWCP-R2) were incubated in buffer solutions with various pH values (3.0–10) for 1 h at 37 °C. After incubation, the solutions were neutralised to pH 7.0 and residual activities were measured under standard assay conditions.
To determine the effects of temperature, purified antimicrobial samples were incubated independently at 30, 45, 60, 75, and 90 °C for 1 h. Thereafter, their residual activities were measured under standard assay conditions; a non-heated sample was used as a control (100%).
The purified antimicrobial samples were treated with α-amylase (1 mg mL−1), β-amylase (1 mg mL−1), lipase (2 mg mL−1), trypsin (3 mg mL−1), protease K (3 mg mL−1), pepsin (3 mg mL−1), esterase (3 mg mL−1), or lysozyme (3 mg mL−1). All reaction mixtures were incubated at 37 °C for 2 h. The residual activity was determined under standard assay conditions. An untreated sample was used as a control (100%).
Statistical Analysis
Data were analysed using one-way analysis of variance (ANOVA) to determine whether significant differences (P < 0.05) existed among treatments. Tukey’s test was used to compare means between individual treatments. The resultant means after one-way ANOVA were used for a subsequent t test. All statistical analyses were carried out using OriginPro software (version 8.5; OriginLab Corporation, Northampton, USA). The results are expressed as mean ± SE of mean (SEM).
Results
Selection of Potential Probiotic Bacteria
Potential probiotic bacteria were screened using a well diffusion method. The average clearance zones of strains R1, R2, and R5 had an average diameter of greater than 10 mm (data not shown). Thus, these 3 strains were used in subsequent experiments.
Protein Estimation of Cellular Fractions
The protein contents of various cellular components of R1, R2, and R5 are summarised in Table 1. The highest protein content was found in the ICP (5.61 mg/mL) of R5, and the lowest protein content was found in the ECP (4.32 mg/mL) of R1. The minimum quantity of protein fraction (100 μL) of the ECP of R1 charged against A. hydrophila was 432 μg. Other fractions were adjusted to 432 μg in 100 μL.
Antagonistic Study Using R1
The results of the antagonistic activity of the WCP, HKWCP, ICP, and ECP of R1 are shown in Table 2. All cellular components exhibited satisfactory bactericidal activity against all A. hydrophila strains. Out of the 4 cellular components, the ICP produced the greatest activity (inhibition zone range, 53–62 mm) in almost all cases, except for ATCC-7965. Similarly, the WCP, ECP, and HKWCP were equally effective against all A. hydrophila strains. The largest inhibition zone was recorded with the ICP against ATCC-21763 (62 ± 0.57 mm), and the smallest inhibition zone was recorded with the ECP against ATCC-7965 (44 ± 0.57 mm). Significant differences (P ≤ 0.05) in the inhibitory activity were observed in all cellular components compared to the positive control.
Antagonistic Activity of R2
The cellular components of R2 were found to be most antagonistic against A. hydrophila (Table 3). The overall bactericidal activity of HKWCP-R2 was the highest among the 4 cellular components tested and produced zones ranging from 55 to 63 mm; meanwhile, the WCP produced an inhibition zone range from 44.6 to 55.5 mm; the ICP from 41.5 to 52.3 mm, and the ECP from 39.5 to 47 mm. The largest inhibition zone was obtained with the HKWCP against MTCC-646 (63 ± 1.52 mm), and the smallest inhibition zone was obtained with the ECP against ATCC-23213 (39.5 ± 1.44 mm). The results were significant (P ≤ 0.05) compared to the positive control.
Antagonistic Study Using R5
The cellular components of R5 were found to be highly antagonistic against A. hydrophila (Table 4). All 4 cellular components produced inhibition zones >38 mm; the greatest bactericidal activity was recorded with the ICP against MTCC-1739 (57 ± 1.52 mm). All 4 components showed significant (P ≤ 0.05) inhibitory activity against 6 strains of A. hydrophila compared to the positive control.
Comparisons Among the Cellular Components of R1, R2, and R5
The results of the t test are summarised in Table 5. The HKWCP from R2 produced a significantly (P ≤ 0.05) larger inhibition zone (59.59 ± 1.11 mm) against A. hydrophila strains compared to the control as well as other components, including those of R1 and R5. ICP-R1 produced the second greatest bactericidal activity (57.33 ± 1.36 mm) against the tested pathogens compared to other components including the ICPs of R2 and R5. On the other hand, the t test revealed that among the cellular components, ECP-R5 was the least effective against the pathogens. Overall, the cellular components of R1 produced more significant results compared with those of the other 2 probiotic strains (Table 5).
Identification of R1, R2, and R5
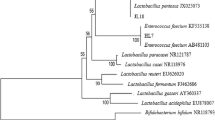
The morphological and biochemical characters of R1, R2, and R5 resembled those of Lactobacillus, Pseudomonas, and Bacillus, respectively. The comparative 16S rRNA gene sequence analysis revealed that isolates R1 (627 bp), R2 (597 bp), and R5 (829 bp) had 98, 99%, and 99% nucleotide base homology with Lactobacillus plantarum, Pseudomonas aeruginosa, and Bacillus subtilis, respectively; the NCBI GenBank accession numbers of the sequences are HQ 141917, HQ 141916, and HQ 127453, respectively.
Purification of Antimicrobial Compounds from the Cellular Fractions
From Table 5, ICP-R1, HKWCP-R2, and ICP-R5 exhibited the greatest antagonistic activities in their respective strains. The TLC spots of the ICP-R1 and ICP-R5 fractions were developed by ninhydrin and iodine reactions, indicating the presence of peptides or lipopeptidal compounds. The HKWCP-R2 spot was developed by silver nitrate, indicating the presence of phenolic compounds. The antimicrobial substances of ICP-R1 and ICP-R5 were eluted with methanol, while that of HKWCP-R2 was eluted with chloroform. The R f values of the antagonistically active spots of ICP-R1, HKWCP-R2, and ICP-R5 were 0.6, 0.8, and 0.51, respectively.
The antagonistic substances that were partially purified by TLC were subjected to HPLC on an 18C reverse-phase column. In the case of ICP-R1, 3 major fractions were found; the third fraction (retention time, 22 min) exhibited the greatest antimicrobial activity against A. hydrophila (data not shown) and was pooled and collected. Similarly, for HKWCP-R2 and ICP-R5, only the fraction showing the greatest antimicrobial activity was pooled and collected.
Partial Characterisations of Antimicrobial Compounds
Effects of pH, Temperature, and Enzymes on the Activity of Antimicrobial Components
The antimicrobial fractions purified from ICP-R1, HKWCP-R2, and ICP-R5 were tested for their sensitivity to pH, temperature, and enzymes; the results are summarised in Table 6.
All fractions were highly active over a wide pH range of 3.0–10.0. The HKWCP-R2 and ICP-R5 fractions were more stable in alkaline pH than in acidic pH (Table 6).
The ICP-R1 and HKWCP-R2 fractions exhibited greater thermal stability than the ICP-R5 fraction. The HKWCP-R2 fraction retained 81% of its initial activity after 1 h incubation at 90 °C, whereas ICP-R5 retained only 63% of its initial activity after the same incubation period (Table 6).
The antibacterial activity of ICP-R1 was completely abolished after treatment with proteolytic enzymes, whereas amylase and lipase had no effect. The antimicrobial activity of the purified HKWCP-R2 fraction was partially abolished when treated with protease K and pepsin. The antibacterial activity of the ICP-R5 fraction was completely abolished with lipase, protease K, and trypsin but was retained (79%) with esterase.
Discussion
From this study, we identified R1, R2, and R5 as Lactobacillus plantarum VSG3, Pseudomonas aeruginosa VSG2, and Bacillus subtilis VSG1, respectively.
The common modes of action of probiotics include (1) stimulation of immune responses, (2) modification of the metabolism of bacterial pathogens by changing their enzyme levels, and (3) competitive exclusion through the production of inhibitory compounds that are antagonistic towards pathogens [13]. Antagonistic compounds produced by bacteria include antibiotics, organic acids, hydrogen peroxide [35], bacteriocins [16, 18] carbon dioxide, and siderophores [38]. The inhibition due to such compounds is highly dependent on the experimental conditions, which are different in vitro and in vivo [14]. Probiotic B. subtilis BT23 controls the growth of pathogenic Vibrio harveyi both in vitro and in vivo, and reduces shrimp mortality by 90% [34]. The use of Bacillus strains as probiotics against bacterial pathogen among fishes is reported [1, 30, 31, 34]. Ghosh et al. [15] report the presence of Bacillus spp. in the gut of L. rohita. In the case of B. subtilis VSG1, the ICP most strongly inhibited A. hydrophila.
In the present investigation, L. plantarum VSG3, P. aeruginosa VSG2, and B. subtilis VSG1 were isolated from the gut of rohu, L. rohita. The presence of different bacterial strains in fish intestines and their possible probiotic activity has been studied [1, 11, 15, 17, 24, 30, 31]. The WCP, ECP, ICP, and HKWCP of all bacterial species were significantly effective against all tested A. hydrophila strains as revealed by their inhibition zones. Antibiotics such as chloramphenicol (30 mcg) and gentamicin (15 mcg) produced smaller inhibition zones than the above-mentioned cellular components. We found that the HKWCP of P. aeruginosa produced the largest inhibition zone (63 ± 1.52 mm) against A. hydrophila. The in vitro antagonistic effects of the 4 cellular components of Pseudomonas spp. against A. hydrophila were reported previously [8]. Pseudomonas strains have the properties of a biocontrol agent for use in shrimp hatcheries and farms [7].
The inhibition profiles revealed that the cellular components of L. plantarum VSG3 were the most effective among the 3 isolates against the tested A. hydrophila stains; the ICP of this strain produced the largest inhibition zone against the tested pathogens. The cell-free supernatants of Lactobacillus spp. JK-8 and JK-11 had remarkable antimicrobial activities against target pathogens (e.g. Vibrio parahaemolyticus, Vibrio harveyi, and Edwardsiella tarda) [21] and removed the pathogens within 4 min of exposure [22]. Balcázar et al. [4] report that the neutralised supernatants of Lactococcus lactis CLFP-101 can inhibit the growth of fish pathogens, A. hydrophila, A. salmonicida, Vibrio anguillarum, and Yersinia ruckeri. Bacillus pumilus might be usable as a probiotic for controlling A. hydrophila infection in Nile tilapia [1]. The cellular components of lactic acid bacteria are reported to have the ability to inhibit the growth of various fish pathogens in vitro conditions [1, 3, 4, 8, 11]. Hence, lactic acid bacterial strains can be considered a very promising alternative to the use of chemotherapeutic agents [3].
Bacteria can produce a variety of antimicrobial substances [22]. The antimicrobial compounds produced by L. plantarum (R1), P. aeruginosa (R2), and B. subtilis (R5) were purified by TLC and HPLC. These compounds had different compositions, because they exhibited different migration rates on TLC and gave different colours after reaction with ninhydrin/iodine vapour/silver nitrate. In the present study, a single ninhydrin spot (R f = 0.6) was observed for ICP-R1, indicating the presence of a protein moiety. A bacteriocin with R f = 0.6 was purified from Lactobacillus paracasei HL32 [25]. HKWCP-R2 from P. aeruginosa gave a brown colour with silver nitrate (R f = 0.8), indicating that the substance is a phenolic compound. According to the ninhydrin and iodine reactions [19], ICP-R5 (R f = 0.51) could be peptidal with lipid moieties. Lipopeptide antibiotics and surfactins with R f = 0.37 and 0.51 were purified from B. subtilis [23].
In an effort to characterise the partial biochemical properties of antimicrobial substances produced by the 3 strains, we tested their sensitivity towards some commercial enzymes. The antimicrobial activity of purified ICP-R1 was completely inhibited by proteolytic enzymes, while amylase and lipase had no effect. These results indicate that ICP-R1 is a kind of peptide that does not contain lipid or carbohydrate groups. Again, ICP-R1 exhibited high thermal and pH stability. In this case, the secreted organic acids can be ruled out since the pH of the growth medium was always in the neutral range (7.0–7.5). These properties are similar to that of a bacteriocin produced by Lactobacillus sake C2 [16]. The purified antimicrobial substance of P. aeruginosa (HKWCP-R2) was partially inactivated by protease K and pepsin, suggesting that a protein moiety is involved in its activity. In addition, HKWCP-R2 exhibited heat and pH stability. The heat- and pH-stable antimicrobial components of Pseudomonas spp. were reported earlier [18, 27]. There is more than one explanation for the observed resistance to proteolytic enzymes such as the presence of unusual amino acids in their structures or cyclic N- and/or C-terminally blocked peptides. Cyclic peptides can be resistant to hydrolysis by proteases, because their cyclic structure renders them relatively inflexible, which may make cleavage sites inaccessible due to steric hindrance [10]. The purified antimicrobial substance of B. subtilis (ICP-R5) completely lost its activity with proteinase K, trypsin, and lipase but not with esterase (Table 6), indicating that it could be a biodegradable antibiotic such as a peptidal compound containing fatty acids [19]. The isolation and characterisation of antimicrobial components from Bacillus strains have been reported previously [19, 22, 30, 31]. The antimicrobial components secreted by probiotic strains may help to avoid pathogen colonisation of fish intestines. Again, these antimicrobial components may find applications as food preservatives and in clinical studies.
This is the first report of characterising the cellular components of L. plantarum VSG3, P. aeruginosa VSG2, and B. subtilis VSG1 newly isolated from the gut of L. rohita, a tropical freshwater fish. The ICPs of R1 and R5 and the HKWCP of R2 most effectively inhibited the growth of A. hydrophila strains. The partially purified antibacterial compounds were not only biodegradable, but also stable in a wide range of pH values (3.0–10.0) and temperatures (30–90 °C). Isolated probiotic strains may find potential application in the prevention and treatment of aeromonosis, which is a major problem in freshwater fishes. However, this needs to be confirmed in future in in vivo experiments. In this study, we report the partial characterisations of the above-mentioned purified antimicrobial compounds; however, the identification and chemical characterisations of these compounds must be carried out to elucidate their complete structure.
References
Aly SM, Abd-El-Rahman AM, John G, Mohamed MF (2008) Characterization of some bacteria isolated from Oreochromis niloticus and their potential use as probiotics. Aquaculture 277:1–6
Austin B, Stuckey LE, Robertson PAW, Effendi I, Griffith DRW (1995) A probiotic strain of Vibrio alginolyticus effective in reducing disease caused by Aeromonas salmonicida, Vibrio anguillarum and Vibrio ordalli. J Fish Dis 18:93–96
Balcázar JL, Vendrell D, Blas ID, Ruiz-Zarzuela I, Girones O, Muzquiz JL (2007) In vitro competitive adhesion and production of antagonistic compounds by lactic acid bacteria against fish pathogens. Vet Microbiol 122:373–380
Balcázar JL, Vendrell D, Blas ID, Ruiz-Zarzuela I, Muzquiz JL, Girones O (2008) Characterization of probiotic properties of lactic acid bacteria isolated from intestinal microbiota of fish. Aquaculture 278:188–191
Bradford MM (1976) A rapid and sensitive method for the quantification of microgram quantities of protein utilizing the principle dye binding. Anal Biochem 72:248–254
Bruun MS, Schmidt AS, Madsen L, Dalsgaard I (2000) Antimicrobial resistance patterns in Danish isolates of Flavobacterium psychrophilum. Aquaculture 187:201–212
Chythanya R, Karunasagar I, Karunasagar I (2002) Inhibition of shrimp pathogenic vibrios by a marine Pseudomonas I-2 strain. Aquaculture 208:1–10
Das BK, Samal SK, Samantaray BR, Sethi S, Pattnaik P, Mishra BK (2006) Antagonistic activity of cellular components of Pseudomonas species against Aeromonas hydrophila. Aquaculture 253:17–24
Direkbusarakom S, Yoshimizu M, Ezura Y, Ruangpan L, Danayadol Y (1998) Vibrio species, the dominant flora in shrimp hatchery against some fish pathogenic viruses. J Marine Biotechnol 6:266–267
Eckart K (1994) Mass spectrometry of cyclic peptides. Mass spectrometry Rev 13:23–55
EI-Rhman AMA, Khattab YAE, Shalaby AME (2009) Micrococcus luteus and Pseudomonas species as probiotics for promoting the growth performance and health of Nile-tilapia, Oreochromis niloticus. Fish Shellfish Immunol 27:175–180
Fabregas J, Munoz A, Otero A, Barja JL, Roruaris M (1991) A preliminary study on antimicrobial activities of some bacteria isolated from the marine environment. Nippon Suisan Gakkaishi 57:1377–1380
Fuller R (1987) A review, probiotics in man and animals. J Appl Bacteriol 66:365–378
Gatesoupe FJ (1999) The use of probiotics in aquaculture. Aquaculture 180:147–165
Ghosh S, Sinha A, Sahu C (2007) Isolation of putative probionts from the intestines of Indian major carps. The Israeli J Aquaculture—Bamidgeh 59(3):127–132
Goa Y, Jia S, Goa Q, Tan Z (2010) A novel bacteriocin with a broad inhibitory spectrum produced by Lactobacillus sake C2, isolated from traditional Chinese fermented cabbage. Food Cont 21:76–81
Hjelm M, Bergh Ø, Riaza A, Nielsen J, Melchiorsen J, Jensen S, Duncan H, Ahrens P, Birkbeck H, Gram L (2004) Selection and identification of autochthonous potential probiotic bacteria from turbot larvae (Scophtalmus maximus) rearing units. Syst Appl Microbiol 27:360–371
Hubert E, Lobos O, Brevis P, Padilla C (1998) Purification and characterization of the bacteriocin PsVP-10 produced by Pseudomonas sp. J Appl Microbiol 84:910–913
Lee NK, Yeo IC, Park JW, Kang BS, Hahm YT (2010) Isolation and characterization of a novel analyte from Bacillus subtilis SC-8 antagonistic to Bacillus cereus. J Biosc Bioeng 110(3):298–303
Ljungh A, Wadström T (2006) Lactic acid bacteria as probiotics. Curr Issues Intest Microbiol 7:73–89
Ma CW, Cho YS, Oh KH (2009) Removal of pathogenic bacteria and nitrogens by Lactobacillus spp. JK-8 and JK-11. Aquaculture 287:266–270
Motta AS, Cannavan FS, Tsai SM, Brandelli A (2007) Characterization of a broad range antibacterial substance from a new Bacillus species isolated from Amazon basin. Arch Microbiol 188:367–375
Mutaz A, Sheikh MA, Ahmed Z, Hasnain S (2007) Production of surfactin from Bacillus subtilis MZ-7 grown on pharmamedia commercial medium. Microb Cell Fac 6:1–8
Panigrahi A, Azad IS (2007) Microbial intervention for better fish health in aquaculture: the Indian scenario. Fish Physiol Biochem 33:429–440
Pangsomboon K, Kaewnopparat S, Pitakpornpreecha T, Srichana T (2006) Antibacterial activity of a bacteriocin from Lactobacillus paracasei HL32 against Porphyromonas gingivalis. Arch Oral Biol 51:784–793
Robertson PAW, O’Dowd C, Burrells C, Williams P, Austin B (2000) Use of Carnobacterium sp as a probiotic for Atlantic salmon (Salmo salar L.) and rainbow trout (Oncorhynchus mykiss, Walbaum). Aquaculture 185:235–243
Roberta F, Spada JC, Silveira ST, Tsai SM, Brandelli A (2009) Purification and characterization of an antimicrobial peptide produced by Pseudomonas sp. strain 4B. World J Microbiol Biotechnol 25:205–213
Sahul H, Balasubramanian G (2000) Antibiotic resistance in bacteria isolated from Artemia nauplii and efficacy of formaldehyde to control bacterial load. Aquaculture 183:195–205
Shah K, Mody K, Keshri J, Jha B (2010) Purification and characterization of a solvent, detergent and oxidizing agent tolerant protease from Bacillus cereus isolated from the Gulf of Khambhat. J Mol Catal B: Enzym 67:85–91
Sugita H, Matsuo N, Shibuya K, Deguchi Y (1996) Production of antimicrobial substances by intestinal bacteria isolated from coastal crab and fish species. J Mar Biotechnol 4:220–223
Sugita H, Hirose Y, Matsuo N, Deguchi Y (1998) Production of antibacterial substances by Bacillus sp. strain NM 12, an intestinal bacterium of Japanese coastal fish. Aquaculture 165:269–280
Swain SM, Singh C, Arul V (2009) Inhibitory activity of probiotics Streptococcus phocae PI80 and Enterococcus faecium MC13 against Vibriosis in shrimp Penaeus monodon. World J Microbiol Biotechnol 25:697–703
Weisburg WG, Barn SM, Pelletier DA, Lane DJ (1991) Ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703
Vaseeharan B, Ramasamy P (2003) Control of pathogenic Vibrio spp. by Bacillus subtilis BT23, a possible probiotic treatment for black tiger shrimp Penaeus monodon. Lett in Appl Microbiol 36:83–87
Vázquez JA, González MP, Murado MA (2005) Effects of lactic acid bacteria cultures on pathogenic microbiota from fish. Aquaculture 245:149–161
Vendrell D, Balcázar JL, Blas ID, Ruiz-Zarzuela I, Gironés I, Múzquiz JL (2008) Protection of rainbow trout (Oncorhynchus mykiss) from lactococcosis by probiotic bacteria. Comp Immunol Microbiol Infect Dis 31:337–345
Vijayan KK, Bright Singh IS, Jayaprakash NS, Alavandi SV, Pai SS, Preetha R, Rajan JJS, Santiago TC (2006) A brackishwater isolate of Pseudomonas PS-102, a potential antagonistic bacterium against pathogenic vibrios in penaeid and non-penaeid rearing systems. Aquaculture 251:192–200
Yoshida T, Hayashi KI, Ohmoto H (2002) Dissolution of iron hydroxides by marine bacterial siderophore. Chem Geol 184:1–9
Acknowledgments
S. S. Giri gratefully acknowledge the Department of Science and Technology, Ministry of Science and Technology, Govt. of India for awarding “Innovation in Science Pursuit for Inspired Research [INSPIRE] Fellowship” [IF10575] to pursue Doctoral research.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Giri, S.S., Sukumaran, V., Sen, S.S. et al. Antagonistic Activity of Cellular Components of Potential Probiotic Bacteria, Isolated from the Gut of Labeo rohita, Against Aeromonas hydrophila . Probiotics & Antimicro. Prot. 3, 214–222 (2011). https://doi.org/10.1007/s12602-011-9078-3
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12602-011-9078-3