Abstract
Overuse and misuse of antibiotics have led bacteria to acquire several mechanisms of resistance. In response to this, researchers have identified natural antimicrobial peptides as promising candidates to fight against multiresistant bacteria. However, their mode of action is still unclear, and a better understanding of the mode of action of these peptides is of primary importance to develop new peptides displaying high antibacterial activity and low hemolytic activity. One of the main features that defines the mechanism of action is the membrane topology of the peptide. Among the spectroscopic techniques, solid-state NMR is the technique of choice for determining the location of the peptide within the membrane. It can be achieved by performing experiments with oriented samples. In the literature, the two most common types of oriented samples are bicelles and phospholipids mechanically oriented between glass plates. The mode of perturbation of the membrane-active peptide can be studied by phosphorus-31 and deuterium NMR. On the other hand, several experiments such as nitrogen-15 and fluorine solid-state NMR, that require labeled peptides, can give valuable information on the membrane topology of the peptide. The combination of the latter techniques allows the determination of a precise topology, thus a better knowledge of the molecular determinants involved in the membrane interactions of antimicrobial peptides.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The increase of infections involving multi-resistant bacteria is currently a major threat, especially in hospital environments. Indeed, since the early 1980s, the number of infections caused by methicillin-resistant Staphylococcus aureus, vancomycin-resistant Enterococcus and fluoroquinolone-resistant Pseudomonas aeruginosa has steadily increased (Levy and Marshall 2004; Taubes 2008). In order to reverse this trend, researchers are trying to develop new molecules with an antibacterial activity resulting from novel mechanisms of action. Among the promising alternatives are cationic antimicrobial peptides which are a key component in the innate immune system from lower to higher organisms (Hancock and Chapple 1999). In general, these peptides share common characteristics which are a short length (12–45 amino acids), a net positive charge (+2 to +9) and a marked amphiphilic character (Hancock and Lehrer 1998). An interesting feature about these peptides is that they do not alter the functioning of a specific target such as conventional antibiotics. Instead, they target the bacterial cellular membrane where they induce diverse perturbations.
The use of these peptides as antibacterial agents presents several advantages over current antibiotics. Indeed, they have a large spectrum of activity (Gram-positive and Gram-negative bacteria), they kill bacteria rapidly and they are less prone to drug resistance. In addition to their antibacterial activity, some of the peptides are also potent against fungi, parasites, and cancer cells (Jenssen et al. 2006). On the other hand, the lack of selectivity represents a disadvantage as they are often toxic towards eukaryotic cells (hemolytic activity).
In the literature, several mechanisms of action have been proposed and widely cited (Fig. 1) (Brogden 2005; Chan et al. 2006; Nguyen et al. 2011; Shai 1999). For all the mechanisms, the initial step is the electrostatic binding of the peptides on the membrane surface. Depending on the specific molecular determinants of the peptide, it can act by either a detergent-like mechanism, better known as the micellization mechanism, or a pore-forming mechanism. For the latter, the peptides induce the formation of pores or defects that will eventually lead to cell death. By looking at the different mechanisms of action, the membrane topology is one of the main features that characterize each mechanism. For example, the peptide has a transmembrane topology in the barrel-stave and toroidal pore mechanisms, whereas the peptide lies on the membrane surface in the case of the sinking-raft mechanism.
Examples of mechanisms of action proposed for natural antimicrobial peptides. Peptides can either destroy the bacterial cell membrane by micellization or induce pore formation. Adapted from Chan et al. 2006 and reproduced with permission
To develop new synthetic antimicrobial peptides that are efficient in targeting and killing bacteria and viable from a pharmacological point of view, the knowledge of the mechanisms of action is of primary importance. In order to achieve this, several spectroscopic techniques can be used. Overall, solid-state NMR is the most suited technique because of its atomic resolution and the possibility to study membrane-active peptides in interaction with hydrated bilayers (Hong et al. 2012). In addition, this technique provides information on both the lipids and the peptides depending on which nucleus is investigated. In comparison with solution-state NMR, solid-state NMR suffers from a lack of resolution due to anisotropic interactions such as chemical shift, dipolar coupling and quadrupolar coupling. The main strategies employed to increase the resolution are to perform experiments with oriented samples or to cancel the anisotropic interactions by spinning the sample rapidly at the magic angle (Drechsler and Separovic 2003). The first strategy has been extensively used to study the membrane topology of both membrane proteins and cationic antimicrobial peptides. This technique relies on aligning all the phospholipids in the same direction relative to the magnetic field, thus the membrane-interacting peptides will also be aligned relative to the magnetic field.
The most common methods to prepare oriented samples are glass plates and bicelles (Fig. 2). Glass plate samples are prepared by mechanically aligning phospholipids between thin glass plates (Hallock et al. 2002; Yamaguchi et al. 2002). Bicelles are obtained by mixing long- and short-chain phospholipids. The phospholipids will auto-assemble to form disk-shaped particles in which the long-chain phospholipids are localized in the planar region and the short-chain phospholipids are preferentially localized in the rim (Ram and Prestegard 1988). However, their morphology is still being debated in the literature and the reader is referred to these papers for a more exhaustive review (Durr et al. 2013; Harroun et al. 2005; Triba et al. 2005). When the bicelles are placed in an external magnetic field, they spontaneously orient with their normal perpendicular relative to the magnetic field. This is due to the negative diamagnetic susceptibility (∆χ) of the phospholipids that forces them to align their principal axis perpendicular to the magnetic field (Marcotte et al. 2006). It is important to mention that the formation and orientation of bicelles in the magnetic field is only possible if the sample is prepared according to specific conditions of long-chain/short-chain phospholipids molar ratio (q) and hydration level (Raffard et al. 2000). Furthermore, the temperature of analysis is also important for the formation and alignment of bicelles. Orientation of the bicelles relative to the external magnetic field can be flipped from perpendicular to parallel by adding components that will confer a positive anisotropic magnetic susceptibility (∆χ) to the phospholipids such as the amphiphilic aromatic 1-naphtol or salts of lanthanides (Prosser et al. 1998; Sanders et al. 1993). More recently, bicelles have been designed to orient with their normal parallel to the magnetic field without adding external components. To achieve this, the long-chain phospholipids have been replaced by chemically modified phospholipids having a biphenyl group in one of the two acyl chains (Diller et al. 2009; Loudet et al. 2007, 2010; Park et al. 2008; Tan et al. 2002). There are some advantages to using bicelles over glass plates (De Angelis and Opella 2007). Indeed, the sample preparation is easier, the hydration level is higher and they are not prone to dehydration during the acquisition time. However, glass plate samples allow more flexibility in the lipid composition, quantity of peptides and temperature of analysis.
a Disk-shaped bicelles aligned with their normal perpendicular relative to the external magnetic field. The long-chain phospholipids (DMPC) are localized in the flat region whereas the short-chain phospholipids (DHPC) are localized in the rim region. b Phospholipids mechanically oriented between glass plates. There are thousands of bilayers of lipids between two glass plates and there is a water layer between the bilayers
Phosphorus-31
Phosphorus is a convenient nucleus to study due its 100 % natural abundance and its spin of ½ (Seelig 1978; Seelig and Seelig 1980). Because each phospholipid contains a phosphorus nucleus in their polar headgroup, 31P solid-state NMR is a useful technique for monitoring the effects induced by the interaction with cationic antimicrobial peptides. However, experiments performed on unoriented samples such as multilamellar vesicles give information on the dynamics of the polar headgroups and the shape of the vesicles. In order to characterize the perturbations or defects induced by the peptides, experiments must be performed on glass plate samples. In comparison with multilamellar vesicle samples, these samples are less hydrated and there is the presence of electrostatic interactions between the glass plates and the first layers of phospholipids. Consequently, both the bilayer fluidity and lipid dynamics are decreased and this helps to stabilize transient membrane deformations or defects induced by membrane active peptides (Bertelsen et al. 2012; Kim et al. 2009; Wi and Kim 2008). The spectral lineshapes can be indicative of the mode of perturbation and can help to determine the mechanism of action of the peptide.
For example, our group has studied some cationic analogues designed from a base 14-mer peptides and the results have shown that both the analogues having either a α-helical conformation or forming intermolecular β-sheet structures significantly perturb the alignment of the phospholipids mechanically oriented between glass plates (Fillion et al. 2014). For the β-aggregated peptides, the results were unexpected because these analogues are unable to induce the release of calcein confined within dimyristoylphosphatidylcholine (DMPC) liposomes (Lorin et al. 2011). In order to better characterize the defects induced by the peptides, Kim et al. have developed a spectral simulation approach that can be applied to two types of membrane deformations, namely toroidal pores and membrane thinning (Wi and Kim 2008). We have performed spectral simulations by using their approach, and these helped to identify the type of membrane deformations induced by the peptides. In the case of the α-helical peptides, the experimental spectra were adequately simulated with defects that consist of a pore radius of 10 Å whereas, for the aggregated peptides, the experimental spectra were simulated with an ellipsoid deformation (Picard et al. 1999). These results are in agreement with previous studies indicating that only the α-helical peptides have a pore-forming ability in interaction with DMPC vesicles (Lorin et al. 2011). This type of experiment has been conducted by the group of Vosegaard to characterize the type of deformations induced by novicidin and alamethicin (Bertelsen et al. 2012), by the group of Hong to study tachyplesin and its linear derivatives (Doherty et al. 2006), by the group of Ramamoorthy to study MSI-78 and MSI-694 which derive from magainin 2 and melittin, respectively (Ramamoorthy et al. 2006), and by the group of Separovic to study antimicrobial peptides, isolated from different frog species, in interaction with bicelles (Marcotte et al. 2003).
Nitrogen-15
15N solid-state NMR has been extensively used to determine the membrane topology of both membrane proteins and membrane-active peptides. However, labeling of the peptides is required because of the low natural abundance of this nucleus. In the case of peptides adopting a α-helical conformation, 15N chemical shift values obtained with oriented samples can be correlated with peptide topology because of the orientational dependence of the chemical shift interaction, as shown in Fig. 3. Indeed, for a peptide that sits on the membrane surface, the 15N chemical shift value is approximately 70 ppm whereas, for a peptide having a transmembrane topology, the 15N chemical shift value is approximately 200 ppm (Bechinger and Sizun 2003). The rationale is that the 15N chemical shift tensor element σ33 makes an angle of 17° relative to the N–H bond and that the N–H bonds are approximately parallel relative to the principal axis of the helix (Bechinger et al. 2004; Strandberg and Ulrich 2004). In order to determine a more precise topology, it is possible to perform experiments with a separated-local-field (SLF) method such as 2D 15N/1H PISEMA (Ramamoorthy et al. 2004), as shown in Fig. 4. Depending on the membrane topology of the α-helical peptide, the spectral pattern, which is called the PISA wheel, is unique and well known for each orientation (Franzin and Marassi 2005). In contrast with the conventional 1D experiment, this experiment can give helpful insights on the membrane topology of peptides or proteins having a β-sheet conformation. Analogously to nitrogen-15, solid-state NMR experiments can be performed with peptides having a 13C-labeled carbonyl group in order to determine the membrane topology of α-helical peptides because of the orientational dependence of the 13C chemical shift (Smith et al. 1994).
a 15N chemical shift tensor elements σ11, σ22 and σ33 for an amino acid. b 15N chemical shift value for a α-helical peptide that adopts a transmembrane topology and 15N chemical shift value for a α-helical peptide that sits on the membrane surface. Adapted from Bechinger and Sizun 2003 and reproduced with permission
a Resonance patterns in 2D PISEMA experiments for a α-helical peptide tilted at different angles relative to the bilayer normal. b Resonance patterns in 2D PISEMA experiments for a peptide adopting a β-sheet conformation tilted at different angles relative to the bilayer normal. Adapted from Franzin and Marassi 2005 and reproduced with permission
This technique has been applied for the base 14-mer peptide in interaction with DMPC oriented between glass plates and the chemical shift measured suggests that the peptide sits on the membrane surface (Ouellet et al. 2007). 15N 1D experiment has been used to determine the qualitative orientation of cationic antimicrobial peptides in membrane such as pleurocidin (Mason et al. 2006), dermadistinctin (Verly et al. 2009), maximin-4 (Heinzmann et al. 2011). When a peptide has several amino acids that are 15N labeled, it is possible to perform SLF experiments. This has been done by De Angelis et al. (2011) for the cationic antimicrobial peptides piscidin 1 (De Angelis et al. 2011). This peptide displays antimicrobial and hemolytic activity and it is a candidate of great interest because of its tolerance to high salt concentrations. Membrane topology of this peptide has been determined in samples of phospholipids mechanically oriented between glass plates and bicelles. They have shown that the membrane interaction and the topology of this peptide are different whether the model membranes mimic the membrane of eukaryotic (zwitterionic) or bacterial cells (anionic). Indeed, analysis of the results obtained by 15N NMR with piscidin 1 in interaction with zwitterionic bicelles demonstrates that the peptide is tilted at 57° relative to the bilayer normal and/or is in equilibrium between bound and unbound states. On the other hand, analysis of the results obtained with piscidin 1 in interaction with anionic bicelles reveals that the peptide is located on the membrane surface. In addition, the membrane topology of piscidin 1 determined in bicelles is similar to the one previously determined with glass plates, thus demonstrating the viability of bicelles to study membrane active peptides. Other examples include a peptide segment of cathelicidin LL-37 (Thennarasu et al. 2010), arenicin (Salnikov et al. 2011) and ampulosporin A (Salnikov et al. 2009).
Nitrogen-14 is a high abundant nucleus with a spin of 1. Therefore, 14N solid-state NMR experiments do not require specific labeling and the spectra are dominated by the anisotropic quadrupolar interaction. For phospholipids having a choline polar head group, the 14N quadrupolar splitting is correlated with the electrostatic potential on the membrane surface. Consequently, performing 14N solid-state NMR experiments with aligned samples allows to investigate both electrostatic interactions between phosphatidylcholine (PC) phospholipids and cationic peptides and the oligomerization process of peptides, thus shedding light on the mechanism of action (Ramamoorthy et al. 2008).
Fluorine-19
As for the phosphorus-31 nucleus, fluorine-19 has a 100 % natural abundance and a spin of ½ but its gyromagnetic ratio is close to that of the proton (Chen et al. 2013). In addition, fluorine-19 is well suited for in-cell NMR due to the absence of background signal in biological samples (Koch et al. 2012). The high sensitivity of this nucleus allows studying antimicrobial peptides over a large range of concentrations, and this is an important point since the membrane topology can be dependent on peptide concentration. Because fluorine atoms are absent in natural amino acids, the latter must be chemically modified to contain fluorine atoms. In general, the strategy relies on replacing a CH3 group by a CF3 group or simply adding a CF3 group. In the literature, there are several examples of fluorinated amino acids that can be incorporated into the primary sequence during the peptide synthesis (Durr et al. 2008; Grage et al. 2008; Mikhailiuk et al. 2006; Tkachenko et al. 2013). In the precise case of macroscopically oriented samples, the spectral lineshape will depend on the substitution pattern of the labeled amino acid (Koch et al. 2012). For a peptide having an amino acid with only one fluorine atom, the interaction observed is the chemical shift anisotropy (CSA). Because the CSA is non-axially symmetric with a mono-fluorine substituent, it is impossible to describe the CSA by a unique angle. Instead, it is more convenient to incorporate 19F-labeled amino acids containing a CF3 group. On the spectrum, the CF3 group gives rise to a triplet, thus allowing the determination of both the axially symmetric dipolar coupling and the axially symmetric CSA. Measuring these interactions give angular constraints that are useful for determining the membrane topology of membrane-active peptides. This technique has been employed to study the membrane topology of several antimicrobial peptides such as PGLa (Afonin et al. 2008; Ieronimo et al. 2010), gramicidin S (Grage et al. 2006), MSI-103 (Strandberg et al. 2008; Toke et al. 2004) and BP100 (Wadhwani et al. 2014). More recently, Ulrich’s group have used fluorine-19 solid-state NMR spectroscopy to study the re-alignment behavior of PGLa, an antimicrobial peptide from the magainin family and gramicidin S, an antimicrobial peptide isolated from Bacillus brevis (Afonin et al. 2014). More specifically, they have shown that the re-alignement was dependent on the phospholipid/peptide molar ratio and the temperature for both peptides. Moreover, the re-alignement of these peptides was influenced by other factors such as the length of the acyl chains, the presence of charged lipids and the presence of cholesterol, thus demonstrating the importance of the nature of the phospholipids in the membrane interaction.
Deuterium
Deuterium NMR is a well-established method to probe the changes occurring in the hydrophobic core of the bilayer. Due to the low natural abundance of deuterium, hydrogen atoms of the acyl chains must be replaced by deuteron atoms. For this nucleus, the dominant interaction is the quadrupolar interaction which results from the coupling between the nucleus quadropolar moment and electric field gradients. In the specific case of a spin-1 nucleus, there are two spin transitions possible that give rise to a doublet of resonances on the spectrum. The separation between these peaks is termed the quadropolar splitting (∆νQ), and its value gives information on the order of the acyl chains. In general, an increase in the value of the quadrupolar splitting is associated with order in the acyl chains. Phospholipids in hydrated bilayers are animated with axially symmetric motions and, therefore, the quadrupolar splitting is given by:
where e2qQ/h is the quadrupolar coupling constant (~170 kHz for aliphatic C-D) (Davis 1983), θ is the angle between the bilayer normal and the magnetic field, and S CD is the order parameter of a deuterium bond vector. The latter is the product of several contributions such as trans-gauche isomerizations and anisotropic reorientation of the whole phospholipid molecules. In addition, there is evidence that the bilayer thickness can be correlated with the order parameter S CD (Salnikov et al. 2009). Performing deuterium experiments with oriented samples leads to a better spectral resolution, allowing the possibility to obtain information on each segment of the acyl chain. However, it is possible to obtain a similar resolution with multilamellar vesicle samples by applying the dePaking technique (Bloom et al. 1981; Lafleur et al. 1989; Sternin et al. 1983).
As an example, the Separovic group has studied the membrane interactions of aurein 1.2, citropin 1.1 and maculatin 1.1 (Balla et al. 2004). These antimicrobial peptides, isolated from Australian tree frogs, adopt a α-helical amphipathic conformation. They have performed deuterium NMR experiments on oriented samples and noticed that the shorter peptides, aurein 1.2 and citropin 1.1, trigger a decrease of the quadrupolar splitting whereas the longer peptide, maculatin 1.1, does not significantly perturb the quadrupolar splitting. These results give insights into the membrane topology. More specifically, the shorter peptides may be located on the membrane surface whereas the longer peptide may adopt a transmembrane topology. Deuterium NMR experiments performed on oriented samples have also been useful in determining the membrane interaction of synthetic 14- and 21-mer peptides (Ouellet et al. 2006), protegrin-1 (Buffy et al. 2004), magainin 2 (Kim et al. 2009) and aurein 3.3 (Kim et al. 2009). More recently, pulse sequences based on SLF experiments such as 2D HIMSELF/HERSELF have been developed. More specifically, these pulse sequences may be used to correlate the 13C chemical shift and the 13C-1H dipolar coupling for each 13C site of the phospholipids. They require no isotopic labeling (13C natural abundance), and they provide information on both the phospholipid structure and the perturbations induced by antimicrobial peptides (Dvinskikh et al. 2007).
Deuterium NMR can also be used to obtain information on the peptide. In order to do this, a deuterated alanine residue is incorporated in the sequence during the peptide synthesis. Both the conformation and the membrane topology of the peptide can be studied because the methyl group is linked with the peptide backbone and the Cα–Cβ bond is oriented at a precise angle relative to the principal axis of the helix (Bechinger and Salnikov 2012). The methyl group gives rise to a doublet of resonance because the three deuterium atoms are chemically equivalent due to free rotation around the Cα–Cβ bond (Batchelder et al. 1983). This type of experiment has notably been conducted on MSI-103, a synthetic peptide designed from PGLa and displaying a high antibacterial activity (Strandberg et al. 2012). Orientation of this peptide has been determined in different membrane compositions by varying the length of acyl chain, degree of saturation, nature of lipid headgroup and phospholid/peptide molar ratio. The results indicate that the membrane topology of MSI-103 is dependent on the spontaneous curvature of the lipids. Indeed, in membrane mimicking systems made of unsaturated lipids, the peptide lies on the membrane surface at all phospholipid/peptide molar ratios. In contrast, the peptide in interaction with mimicking systems made of saturated lipids adopts a tilted state at higher phospholipid/peptide molar ratios. This technique has also been used to study the synergistic transmembrane insertion of two antimicrobial peptides from the magainin family, PGLa and maganin 2 (Strandberg et al. 2009), and to study the membrane topology of mastoparan X in interaction with bicelles (Whiles et al. 2001).
In addition, the determination of a precise membrane topology can be achieved by combining this technique with 15N NMR. Analysis of the results can reveal the combination of the pairs of tilt and rotational pitch angles that are possible. This has been done on the cationic antimicrobial peptide phylloseptin-2 (PS-2) (Bechinger et al. 2011). More specifically, 15N and 2H NMR experiments were performed on oriented samples and the combination of the results revealed that there were 5 possible combinations which are individually associated with a specific topology. In order to determine the right topology, other measurements have been performed to restrict even more the rotational pitch angle values and the results indicate that the peptide is preferentially located on the membrane surface. Other valuable examples include the study of the peptaibol alamethicin (Bertelsen et al. 2009), the antibiotic heterodimeric peptide distinctin (Resende et al. 2009) and some derivatives of the transmembrane model peptide WALP23 (Vostrikov et al. 2011). Along with oriented solid-state NMR, other techniques that require oriented or supported lipid bilayers may be used to study the membrane interaction of antimicrobial peptides such as surface plasmon resonance (SPR), dual polarization interferometry (DPI) and oriented circular dichroism (OCD) (Besenicar et al. 2006; Hall et al. 2014; Lee et al. 2010). In particular, OCD has been shown to be useful for determining the membrane topology of α-helical peptides (Burck et al. 2008; Sani et al. 2012).
Concluding remarks
The membrane topology is of primary importance to better understand the mechanism of action of peptides displaying antimicrobial activity. Both the membrane topology and the perturbations induced by membrane-active peptides can be studied using solid-state NMR. The versatility of the technique has been demonstrated for many systems such as bicelles and glass plates, and comes with the possibility to exploit diverse experiments involving different nuclei (Hong and Su 2011). The 31P and 2H nuclei are useful for determining the local perturbations induced by the peptides in interaction with phospholipids. The location of the peptide in the membrane can be assessed by performing several solid-state NMR experiments. The 15N chemical shift measured with labeled peptides is a good indicator of the orientation of the peptide relative to the bilayer normal. In addition, it is possible to estimate the peptide tilt angle with the PISEMA pulse sequence. More recently, determination of peptide membrane topology has been done by measuring both the axially symmetric dipolar coupling and the axially symmetric CSA of the highly sensitive 19F nucleus. Furthermore, combination of several experiments can give valuable information on the orientation and depth of insertion of the peptides, thus allowing the determination of a more precise topology.
Abbreviations
- ∆χ:
-
Magnetic susceptibility
- AMP:
-
Antimicrobial peptides
- CSA:
-
Chemical shift anisotropy
- DHPC:
-
Dihexanoylphosphatidylcholine
- DMPC:
-
Dimyristoylphosphatidylcholine
- DPI:
-
Dual polarization interferometry
- NMR:
-
Nuclear magnetic resonance
- OCD:
-
Oriented circular dichroism
- PISEMA:
-
Polarization inversion spin exchange at the magic angle
- SLF:
-
Separated-local-field
- SPR:
-
Surface plasmon resonance
References
Afonin S, Grage SL, Ieronimo M, Wadhwani P, Ulrich AS (2008) Temperature-dependent transmembrane insertion of the amphiphilic peptide PGLa in lipid bilayers observed by solid state 19F NMR spectroscopy. J Am Chem Soc 130:16512–16514. doi:10.1021/ja803156d
Afonin S, Glaser RW, Sachse C, Salgado J, Wadhwani P, Ulrich AS (2014) 19F NMR screening of unrelated antimicrobial peptides shows that membrane interactions are largely governed by lipids. Biochim Biophys Acta 1838:2260–2268. doi:10.1016/j.bbamem.2014.03.017
Balla MS, Bowie JH, Separovic F (2004) Solid-state NMR study of antimicrobial peptides from Australian frogs in phospholipid membranes. Eur Biophys J 33:109–116. doi:10.1007/s00249-003-0342-7
Batchelder LS, Niu CH, Torchia DA (1983) Methyl reorientation in polycrystalline amino acids and peptides: a deuteron NMR spin–lattice relaxation study. J Am Chem Soc 105:2228–2231. doi:10.1021/ja00346a021
Bechinger B, Salnikov ES (2012) The membrane interactions of antimicrobial peptides revealed by solid-state NMR spectroscopy. Chem Phys Lipids 165:282–301. doi:10.1016/j.chemphyslip.2012.01.009
Bechinger B, Sizun C (2003) Alignment and structural analysis of membrane polypeptides by 15N and 31P solid-state NMR spectroscopy. Concepts Magn Reson A 18A:130–145. doi:10.1002/cmr.a.10070
Bechinger B, Aisenbrey C, Bertani P (2004) The alignment, structure and dynamics of membrane-associated polypeptides by solid-state NMR spectroscopy. Biochim Biophys Acta 1666:190–204. doi:10.1016/j.bbamem.2004.08.008
Bechinger B, Resende JM, Aisenbrey C (2011) The structural and topological analysis of membrane-associated polypeptides by oriented solid-state NMR spectroscopy: established concepts and novel developments. Biophys Chem 153:115–125. doi:10.1016/j.bpc.2010.11.002
Bertelsen K et al (2009) Residue-specific information about the dynamics of antimicrobial peptides from 1H-15N and 2H solid-state NMR spectroscopy. J Am Chem Soc 131:18335–18342. doi:10.1021/ja908604u
Bertelsen K, Dorosz J, Hansen SK, Nielsen NC, Vosegaard T (2012) Mechanisms of peptide-induced pore formation in lipid bilayers investigated by oriented 31P solid-state NMR spectroscopy. PLoS ONE 7:e47745. doi:10.1371/journal.pone.0047745
Besenicar M, Macek P, Lakey JH, Anderluh G (2006) Surface plasmon resonance in protein-membrane interactions. Chem Phys Lipids 141:169–178. doi:10.1016/j.chemphyslip.2006.02.010
Bloom M, Davis JH, Mackay AL (1981) Direct determination of the oriented sample nmr spectrum from the powder spectrum for systems with local axial symmetry. Chem Phys Lett 80:198–202. doi:10.1016/0009-2614(81)80089-9
Brogden KA (2005) Antimicrobial peptides: pore formers or metabolic inhibitors in bacteria? Nat Rev Microbiol 3:238–250. doi:10.1038/nrmicro1098. This is a review on cationic antimicrobial peptides that focuses on the mechanisms of action of these peptides. This article also provides information on the role of these peptides in the innate immune response by altering physiological pathways. Therefore, other mechanisms of action (intracellular targets) are described along with examples from the literature
Buffy JJ, McCormick MJ, Wi S, Waring A, Lehrer RI, Hong M (2004) Solid-state NMR investigation of the selective perturbation of lipid bilayers by the cyclic antimicrobial peptide RTD-1. Biochemistry 43:9800–9812. doi:10.1021/bi036243w
Burck J, Roth S, Wadhwani P, Afonin S, Kanithasen N, Strandberg E, Ulrich AS (2008) Conformation and membrane orientation of amphiphilic helical peptides by oriented circular dichroism. Biophys J 95:3872–3881. doi:10.1529/biophysj.108.136085
Chan DI, Prenner EJ, Vogel HJ (2006) Tryptophan- and arginine-rich antimicrobial peptides: structures and mechanisms of action. Biochim Biophys Acta 1758:1184–1202. doi:10.1016/j.bbamem.2006.04.006
Chen H, Viel S, Ziarelli F, Peng L (2013) 19F NMR: a valuable tool for studying biological events. Chem Soc Rev 42:7971–7982. doi:10.1039/c3cs60129c
Davis JH (1983) The description of membrane lipid conformation, order and dynamics by 2H-NMR. Biochim Biophys Acta 737:117–171
de Angelis AA, Opella SJ (2007) Bicelle samples for solid-state NMR of membrane proteins. Nat Protoc 2:2332–2338. doi:10.1038/nprot.2007.329
de Angelis AA, Grant CV, Baxter MK, McGavin JA, Opella SJ, Cotten ML (2011) Amphipathic antimicrobial piscidin in magnetically aligned lipid bilayers. Biophys J 101:1086–1094. doi:10.1016/j.bpj.2011.07.015
Diller A et al (2009) Bicelles: a natural ‘molecular goniometer’ for structural, dynamical and topological studies of molecules in membranes. Biochimie 91:744–751. doi:10.1016/j.biochi.2009.02.003
Doherty T, Waring AJ, Hong M (2006) Peptide-lipid interactions of the beta-hairpin antimicrobial peptide tachyplesin and its linear derivatives from solid-state NMR. Biochim Biophys Acta 1758:1285–1291. doi:10.1016/j.bbamem.2006.03.016
Drechsler A, Separovic F (2003) Solid-state NMR structure determination. IUBMB Life 55:515–523. doi:10.1080/15216540310001622740
Durr UH, Grage SL, Witter R, Ulrich AS (2008) Solid state 19F NMR parameters of fluorine-labeled amino acids. Part I: Aromat Substituents. J Magn Reson 191:7–15. doi:10.1016/j.jmr.2007.11.017
Durr UH, Soong R, Ramamoorthy A (2013) When detergent meets bilayer: birth and coming of age of lipid bicelles. Prog Nucl Magn Reson Spectrosc 69:1–22. doi:10.1016/j.pnmrs.2013.01.001
Dvinskikh SV, Durr UH, Yamamoto K, Ramamoorthy A (2007) High-resolution 2D NMR spectroscopy of bicelles to measure the membrane interaction of ligands. J Am Chem Soc 129:794–802. doi:10.1021/ja065536k
Fillion M, Noël M, Lorin A, Voyer N, Auger M (2014) Investigation of the mechanism of action of novel amphipathic peptides: insights from solid-state NMR studies of oriented lipid bilayers. Biochim Biophys Acta 1838:2173–2179. doi:10.1016/j.bbamem.2014.01.029
Franzin CM, Marassi FM (2005) NMR structure determination of proteins in bilayer lipid membranes: The FXYD family proteins. In: Ottova-Leitmannova A (ed) Advances in Planar Lipid Bilayers and Liposomes, vol 2. Academic, Burlington, pp 77–93. doi:http://dx.doi.org/10.1016/S1554-4516(05)02003-X
Grage SL, Suleymanova AV, Afonin S, Wadhwani P, Ulrich AS (2006) Solid state NMR analysis of the dipolar couplings within and between distant CF3-groups in a membrane-bound peptide. J Magn Reson 183:77–86. doi:10.1016/j.jmr.2006.07.012
Grage SL, Durr UH, Afonin S, Mikhailiuk PK, Komarov IV, Ulrich AS (2008) Solid state 19F NMR parameters of fluorine-labeled amino acids. Part II: aliphatic substituents. J Magn Reson 191:16–23. doi:10.1016/j.jmr.2007.11.016
Hall K, Lee TH, Mechler AI, Swann MJ, Aguilar MI (2014) Real-time measurement of membrane conformational states induced by antimicrobial peptides: balance between recovery and lysis. Sci Rep 4:5479. doi:10.1038/srep05479
Hallock KJ, Henzler Wildman K, Lee DK, Ramamoorthy A (2002) An innovative procedure using a sublimable solid to align lipid bilayers for solid-state NMR studies. Biophys J 82:2499–2503. doi:10.1016/S0006-3495(02)75592-6
Hancock RE, Chapple DS (1999) Peptide antibiotics. Antimicrob Agents Chemother 43:1317–1323
Hancock RE, Lehrer R (1998) Cationic peptides: a new source of antibiotics. Trends Biotechnol 16:82–88
Harroun TA, Koslowsky M, Nieh MP, de Lannoy CF, Raghunathan VA, Katsaras J (2005) Comprehensive examination of mesophases formed by DMPC and DHPC mixtures. Langmuir 21:5356–5361. doi:10.1021/la050018t
Heinzmann R, Grage SL, Schalck C, Burck J, Banoczi Z, Toke O, Ulrich AS (2011) A kinked antimicrobial peptide from Bombina maxima. II. Behavior in phospholipid bilayers. Eur Biophys J 40:463–470. doi:10.1007/s00249-010-0668-x
Hong M, Su Y (2011) Structure and dynamics of cationic membrane peptides and proteins: insights from solid-state NMR. Protein Sci 20:641–655. doi:10.1002/pro.600
Hong M, Zhang Y, Hu F (2012) Membrane protein structure and dynamics from NMR spectroscopy. Annu Rev Phys Chem 63:1–24. doi:10.1146/annurev-physchem-032511-143731
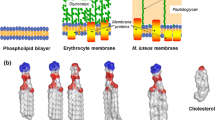
Ieronimo M, Afonin S, Koch K, Berditsch M, Wadhwani P, Ulrich AS (2010) 19F NMR analysis of the antimicrobial peptide PGLa bound to native cell membranes from bacterial protoplasts and human erythrocytes. J Am Chem Soc 132:8822–8824. doi:10.1021/ja101608z
Jenssen H, Hamill P, Hancock RE (2006) Peptide antimicrobial agents. Clin Microbiol Rev 19:491–511. doi:10.1128/CMR. 00056-05
Kim C, Spano J, Park EK, Wi S (2009) Evidence of pores and thinned lipid bilayers induced in oriented lipid membranes interacting with the antimicrobial peptides, magainin-2 and aurein-3.3. Biochim Biophys Acta 1788:1482–1496. doi:10.1016/j.bbamem.2009.04.017
Koch K, Afonin S, Ieronimo M, Berditsch M, Ulrich A (2012) Solid-state 19F-NMR of peptides in native membranes. In: Chan JCC (ed) Solid State NMR, vol 306. Topics in Current Chemistry. Springer, Berlin, pp 89–118. doi:10.1007/128_2011_162. This book chapter provides valuable information to better understand the basis of 19F solid-state NMR with labeled peptides. In addition, an overview of model membranes is introduced with an emphasis on using biomembranes for in cell-NMR.
Lafleur M, Fine B, Sternin E, Cullis PR, Bloom M (1989) Smoothed orientational order profile of lipid bilayers by 2H-nuclear magnetic resonance. Biophys J 56:1037–1041. doi:10.1016/S0006-3495(89)82749-3
Lee TH et al (2010) The membrane insertion of helical antimicrobial peptides from the N-terminus of Helicobacter pylori ribosomal protein L1. Biochim Biophys Acta 1798:544–557. doi:10.1016/j.bbamem.2010.01.014
Levy SB, Marshall B (2004) Antibacterial resistance worldwide: causes, challenges and responses. Nat Med 10:S122–129. doi:10.1038/nm1145
Lorin A et al (2011) Revisiting peptide amphiphilicity for membrane pore formation. Biochemistry 50:9409–9420. doi:10.1021/bi201335t
Loudet C, Manet S, Gineste S, Oda R, Achard MF, Dufourc EJ (2007) Biphenyl bicelle disks align perpendicular to magnetic fields on large temperature scales: a study combining synthesis, solid-state NMR, TEM, and SAXS. Biophys J 92:3949–3959. doi:10.1529/biophysj.106.097758
Loudet C, Diller A, Grelard A, Oda R, Dufourc EJ (2010) Biphenyl phosphatidylcholine: a promoter of liposome deformation and bicelle collective orientation by magnetic fields. Prog Lipid Res 49:289–297. doi:10.1016/j.plipres.2010.02.002
Marcotte I et al (2003) Interaction of antimicrobial peptides from Australian amphibians with lipid membranes. Chem Phys Lipids 122:107–120
Marcotte I, Belanger A, Auger M (2006) The orientation effect of gramicidin A on bicelles and Eu3+ -doped bicelles as studied by solid-state NMR and FT-IR spectroscopy. Chem Phys Lipids 139:137–149. doi:10.1016/j.chemphyslip.2005.12.002
Mason AJ, Chotimah IN, Bertani P, Bechinger B (2006) A spectroscopic study of the membrane interaction of the antimicrobial peptide Pleurocidin. Mol Membr Biol 23:185–194. doi:10.1080/09687860500485303
Mikhailiuk PK et al (2006) Conformationally rigid trifluoromethyl-substituted α-amino acid designed for peptide structure analysis by solid-state 19F NMR spectroscopy. Angew Chem Int Ed 45:5659–5661. doi:10.1002/anie.200600346
Nguyen LT, Haney EF, Vogel HJ (2011) The expanding scope of antimicrobial peptide structures and their modes of action. Trends Biotechnol 29:464–472. doi:10.1016/j.tibtech.2011.05.001
Ouellet M, Bernard G, Voyer N, Auger M (2006) Insights on the interactions of synthetic amphipathic peptides with model membranes as revealed by 31P and 2H solid-state NMR and infrared spectroscopies. Biophys J 90:4071–4084. doi:10.1529/biophysj.105.077339
Ouellet M, Doucet JD, Voyer N, Auger M (2007) Membrane topology of a 14-mer model amphipathic peptide: a solid-state NMR spectroscopy study. Biochemistry 46:6597–6606. doi:10.1021/bi0620151
Park SH, Loudet C, Marassi FM, Dufourc EJ, Opella SJ (2008) Solid-state NMR spectroscopy of a membrane protein in biphenyl phospholipid bicelles with the bilayer normal parallel to the magnetic field. J Magn Reson 193:133–138. doi:10.1016/j.jmr.2008.04.033
Picard F, Paquet MJ, Levesque J, Belanger A, Auger M (1999) 31P NMR first spectral moment study of the partial magnetic orientation of phospholipid membranes. Biophys J 77:888–902. doi:10.1016/S0006-3495(99)76940-7
Prosser RS, Hwang JS, Vold RR (1998) Magnetically aligned phospholipid bilayers with positive ordering: a new model membrane system. Biophys J 74:2405–2418. doi:10.1016/S0006-3495(98)77949-4
Raffard G, Steinbruckner S, Arnold A, Davis JH, Dufourc EJ (2000) Temperature−composition diagram of dimyristoylphosphatidylcholine−dicaproylphosphatidylcholine “bicelles” self-orienting in the magnetic field. A solid state 2H and 31P NMR study. Langmuir 16:7655–7662. doi:10.1021/la000564g. This article presents a good overview on bicelles in terms of experimental preparation and spectral patterns. More specifically, temperature-composition diagrams are depicted in order to determine conditions such as lipid composition, temperature and hydration that are required to form bicelles that align spontaneously in the magnetic field
Ram P, Prestegard JH (1988) Magnetic field induced ordering of bile salt/phospholipid micelles: new media for NMR structural investigations. Biochim Biophys Acta 940:289–294
Ramamoorthy A, Wei Y, Lee D-K (2004) PISEMA Solid-State NMR Spectroscopy. In: Webb GA (ed) Annual Reports on NMR Spectroscopy, vol 52. Academic, Burlington, pp 1–52. doi:http://dx.doi.org/10.1016/S0066-4103(04)52001-X
Ramamoorthy A, Thennarasu S, Lee DK, Tan A, Maloy L (2006) Solid-state NMR investigation of the membrane-disrupting mechanism of antimicrobial peptides MSI-78 and MSI-594 derived from magainin 2 and melittin. Biophys J 91:206–216. doi:10.1529/biophysj.105.073890
Ramamoorthy A, Lee DK, Santos JS, Henzler-Wildman KA (2008) Nitrogen-14 solid-state NMR spectroscopy of aligned phospholipid bilayers to probe peptide-lipid interaction and oligomerization of membrane associated peptides. J Am Chem Soc 130:11023–11029. doi:10.1021/ja802210u
Resende JM et al (2009) Membrane structure and conformational changes of the antibiotic heterodimeric peptide distinctin by solid-state NMR spectroscopy. Proc Natl Acad Sci U S A 106:16639–16644. doi:10.1073/pnas.0905069106
Salnikov ES et al (2009) Structure and alignment of the membrane-associated peptaibols ampullosporin A and alamethicin by oriented 15N and 31P solid-state NMR spectroscopy. Biophys J 96:86–100. doi:10.1529/biophysj.108.136242
Salnikov ES, Aisenbrey C, Balandin SV, Zhmak MN, Ovchinnikova TV, Bechinger B (2011) Structure and alignment of the membrane-associated antimicrobial peptide arenicin by oriented solid-state NMR spectroscopy. Biochemistry 50:3784–3795. doi:10.1021/bi1018732
Sanders CR 2nd, Schaff JE, Prestegard JH (1993) Orientational behavior of phosphatidylcholine bilayers in the presence of aromatic amphiphiles and a magnetic field. Biophys J 64:1069–1080. doi:10.1016/S0006-3495(93)81473-5
Sani MA, Whitwell TC, Separovic F (2012) Lipid composition regulates the conformation and insertion of the antimicrobial peptide maculatin 1.1. Biochim Biophys Acta 1818:205–211. doi:10.1016/j.bbamem.2011.07.015
Seelig J (1978) 31P nuclear magnetic resonance and the head group structure of phospholipids in membranes. Biochim Biophys Acta 515:105–140
Seelig J, Seelig A (1980) Lipid conformation in model membranes and biological membranes. Q Rev Biophys 13:19–61
Shai Y (1999) Mechanism of the binding, insertion and destabilization of phospholipid bilayer membranes by alpha-helical antimicrobial and cell non-selective membrane-lytic peptides. Biochim Biophys Acta 1462:55–70
Smith R, Separovic F, Milne TJ, Whittaker A, Bennett FM, Cornell BA, Makriyannis A (1994) Structure and orientation of the pore-forming peptide, melittin, in lipid bilayers. J Mol Biol 241:456–466. doi:10.1006/jmbi.1994.1520
Sternin E, Bloom M, Mackay AL (1983) de-Pake-ing of NMR spectra. J Magn Reson 55:274–282. doi:10.1016/0022-2364(83)90239-1
Strandberg E, Ulrich AS (2004) NMR methods for studying membrane-active antimicrobial peptides. Concepts Magn Reson A 23A:89–120. doi:10.1002/cmr.a.20024. This is a comprehensive article that reviews the most used techniques for studying the interactions between membrane-active peptides and model membranes by both solution and solid-state NMR. For each technique, several examples of the literature are presented and annotated with experimental details
Strandberg E, Kanithasen N, Tiltak D, Burck J, Wadhwani P, Zwernemann O, Ulrich AS (2008) Solid-state NMR analysis comparing the designer-made antibiotic MSI-103 with its parent peptide PGLa in lipid bilayers. Biochemistry 47:2601–2616. doi:10.1021/bi701944r
Strandberg E, Tremouilhac P, Wadhwani P, Ulrich AS (2009) Synergistic transmembrane insertion of the heterodimeric PGLa/magainin 2 complex studied by solid-state NMR. Biochim Biophys Acta 1788:1667–1679. doi:10.1016/j.bbamem.2008.12.018
Strandberg E, Tiltak D, Ehni S, Wadhwani P, Ulrich AS (2012) Lipid shape is a key factor for membrane interactions of amphipathic helical peptides. Biochim Biophys Acta Biomembr 1818:1764–1776. doi:10.1016/j.bbamem.2012.02.027
Tan C, Fung BM, Cho G (2002) Phospholipid bicelles that align with their normals parallel to the magnetic field. J Am Chem Soc 124:11827–11832
Taubes G (2008) The bacteria fight back. Science 321:356–361. doi:10.1126/science.321.5887.356
Thennarasu S, Tan A, Penumatchu R, Shelburne CE, Heyl DL, Ramamoorthy A (2010) Antimicrobial and membrane disrupting activities of a peptide derived from the human cathelicidin antimicrobial peptide LL37. Biophys J 98:248–257. doi:10.1016/j.bpj.2009.09.060
Tkachenko AN, Radchenko DS, Mykhailiuk PK, Afonin S, Ulrich AS, Komarov IV (2013) Design, synthesis, and application of a trifluoromethylated phenylalanine analogue as a label to study peptides by solid-state 19F NMR spectroscopy. Angew Chem Int Ed 52:6504–6507. doi:10.1002/anie.201301344
Toke O, O’Connor RD, Weldeghiorghis TK, Maloy WL, Glaser RW, Ulrich AS, Schaefer J (2004) Structure of (KIAGKIA)3 aggregates in phospholipid bilayers by solid-state NMR. Biophys J 87:675–687. doi:10.1529/biophysj.103.032714
Triba MN, Warschawski DE, Devaux PF (2005) Reinvestigation by phosphorus NMR of lipid distribution in bicelles. Biophys J 88:1887–1901. doi:10.1529/biophysj.104.055061
Verly RM et al (2009) Structure and membrane interactions of the antibiotic peptide dermadistinctin K by multidimensional solution and oriented 15N and 31P solid-state NMR spectroscopy. Biophys J 96:2194–2203. doi:10.1016/j.bpj.2008.11.063
Vostrikov VV, Grant CV, Opella SJ, Koeppe RE 2nd (2011) On the combined analysis of 2H and 15N/1H solid-state NMR data for determination of transmembrane peptide orientation and dynamics. Biophys J 101:2939–2947. doi:10.1016/j.bpj.2011.11.008
Wadhwani P, Strandberg E, van den Berg J, Mink C, Burck J, Ciriello RA, Ulrich AS (2014) Dynamical structure of the short multifunctional peptide BP100 in membranes. Biochim Biophys Acta 1838:940–949. doi:10.1016/j.bbamem.2013.11.001
Whiles JA, Brasseur R, Glover KJ, Melacini G, Komives EA, Vold RR (2001) Orientation and effects of mastoparan X on phospholipid bicelles. Biophys J 80:280–293. doi:10.1016/S0006-3495(01)76013-4
Wi S, Kim C (2008) Pore structure, thinning effect, and lateral diffusive dynamics of oriented lipid membranes interacting with antimicrobial peptide protegrin-1: 31P and 2H solid-state NMR study. J Phys Chem B 112:11402–11414. doi:10.1021/jp801825k
Yamaguchi S, Hong T, Waring A, Lehrer RI, Hong M (2002) Solid-state NMR investigations of peptide-lipid interaction and orientation of a beta-sheet antimicrobial peptide, protegrin. Biochemistry 41:9852–9862
Compliance with Ethical Standards
Funding
This study was funded by the Natural Sciences and Engineering Research Council of Canada (NSERC) (grant number RGPIN/184177-2010), the Fonds de recherche du Québec - Nature et Technologies (FRQ-NT), the Regroupement québécois de recherche sur la fonction, la structure et l’ingénierie des protéines (PROTEO), the Centre de recherche sur les matériaux avancés (CERMA) and the Centre québécois sur les matériaux fonctionnels (CQMF). M.F. would like to acknowledge graduate scholarships from NSERC and FRQ-NT.
Conflict of interest
Matthieu Fillion declares that he has no conflict of interest. Michèle Auger declares that she has no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Fillion, M., Auger, M. Oriented samples: a tool for determining the membrane topology and the mechanism of action of cationic antimicrobial peptides by solid-state NMR. Biophys Rev 7, 311–320 (2015). https://doi.org/10.1007/s12551-015-0167-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12551-015-0167-5