Abstract
Forest precision classification products were the basic data for surveying of forest resource, updating forest subplot information, logging and management of forest. However, due to the diversity of stand structure, complexity of the forest growth environment, it is difficult to discriminate forest tree species using multi-spectral image. The airborne hyper-spectral images can obtain high spatial and spectral resolution imagery of forest canopy, so it may be useful for tree species level classification. The aim of this paper was to test the effective of combining spatial and spectral features in airborne hyper-spectral image classification. The CASI hyper spectral image data were acquired from Liangshui natural reserves area. First the MNF (minimum noise fraction) transform method for to reduce the hyperspectral image dimensionality and highlighting variation. Second, the grey level co-occurrence matrix (GLCM) is used to extract the texture features of forest tree canopy. Thirdly the texture and the spectral features of forest canopy were fused to classify the trees species using support vector machine (SVM) with different kernel functions. The results showed that when using the SVM classifier, MNF and texture-based features combined with linear kernel function can achieve the best overall accuracy which was 85.92 %. It also confirmed the belief that combined the spatial and spectral information can improve the accuracy of tree species classification.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Forest has significant implication on the environment such as protection of biological diversity and climate change. The forest species map is useful information to drive ecosystem’s model, preserve vegetation and management forest (Liang and Zeng 2009; Wang et al. 2010) It is well reported that the biomass and net productivity are quite different for different tree species (Ustin et al. 2010). However, it is difficult and time-consuming to accurately map the distribution of vegetation species based on ground investigation. Remote sensing techniques provide powerful and efficient tools to solve such problems. Several studies have been carried out in this field, analysing the potential of different remote sensing sensors in vegetation classification. Multispectral sensors (like TM of Landsat Satellites) have been widely used for forest classification and analysis (Lu 2005; Lu et al. 2008; Zaw Htun et al. 2011). Regarding classification, due to the different spectral and spatial resolution of multispectral sensors, it is possible to distinguish vegetation with different levels of geometrical detail. Regarding low-resolution multispectral data such as MODIS, the analysis is generally limited to discrimination between forested and non-forested area (Sedano et al. 2005). With medium resolution sensors such as TM, the level of geometrical detail increases, and the analysis on vegetation type classification can be achieved. With high geometrical resolution sensors such as IKONOS, SPOT 5, a more detailed analysis is possible. Especially the very high resolution imagery such as WorldView-2, GeoEye could get more detail about context information (such as textural information or object-based information) about canopy which was used to distinguish the tree species (Immitzer et al. 2012; Leempoel et al. 2013). However, due to the poor spectral information acquired by these multispectral sensors, they do not permit a detailed analysis to distinguish trees at species level (Jose and David 1996; Gong et al. 1998; Chen et al. 2007).
Recently, hyperspectral remote sensing sensors which provide a significant enhancement of spectral measurement capabilities over conventional multi-spectral data have been widely used for detecting vegetation characteristics. Compared to the multispectral data, the hyperspectral data has much higher spectral resolution and shows great potential in vegetation stress (Smith et al. 2004), measuring chlorophyll content and leaf area index (LAI) of vegetation (Zhao et al. 2007), classifying and mapping vegetation species (Clark et al. 2005; Zhao et al. 2007; Hestir et al. 2008; Huang and Asner 2009; Kozoderov and Dmitriev 2011). Concerning classification problems, hyperspectral images have been used in a variety of forest applications, ranging from discrimination between forest and other land covers, to a more detailed analysis dealing with the distinction of different tree species. All the results confirmed that, with hyperspectral data, it is possible to obtain much higher classification accuracies than with multispectral images.
However, it must be noted that analysis of hyperspectral images are much more complex than multispectral data to classify vegetation species. The first task in for processing hyperspectral images for vegetation species classification is to select the suitable features to distinguish the different species. Feature extraction and feature selection methods were used to solve these problems by selecting optimal bands or optimal subset from the hyperspectral data, such as genetic search algorithms (Vaiphasa et al. 2007), principal component analysis (Bajorski 2011), and minimum noise fraction (Jouan 2007). The classification methods such as the maximum likelihood, decision trees, and random forests classifiers are becoming commonly used in tree species classification. The Support Vector Machines (SVM) which were suggested by Vapnik (1998), are one of the latest effectiveness classifiers which can manage classification problems in hyperdimensional features spaces and have been widely applied in tree species classification. But, these methods consider per-pixel spectral information and do not considering the neighbourhoods of a pixel. Because of the large variation in growing conditions caused by difference in geology, lithology, soil, elevation, historic background, local climatic factors and the land abandonment process itself, a large variety in heterogeneous vegetation communities is found in the area. The heterogeneous vegetation communities are challenging to classify using spectral classifiers because the different vegetation may have a very similar spectral response. And the neighbourhood information may useful for classification species especially in high spatial resolution images.
In this paper, we were used the context and spectral features to classification tree species with airborne hyperspectral image using the SVM classifier. The objects of this paper are to: (1) test whether the spectral and context information can promote the accuracy in tree species classification; (2) compare the effectiveness of different kernel function in SVM classifier in tree species classification.
Data Set Description
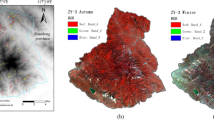
The study site is located in natural reserve area in liangshui, HeiLongJiang province, Northeast China. The forest species are dominated by larch, red pine, birch, conifer and poplar. The hyperspectral image of liangshui was acquired by the Compact Airborne Spectrographic Image (CASI) 1500 hyperspectral sensor August 23, 2009. The CASI imagery provides 144 bands at a 2.3 nm spectral resolution and 1.5 m spatial resolution covering the visible and near-infrared range from 350 to 1,050 nm. A natural colour composite image of the study area is given in Fig. 1. The image was then georeferenced by the position and orientation system (POS) data which including inertial measurement unit (IMU) and global position system (GPS).
Five tree species types, fir, red pine, larch, birch, willow and three other non-forest types, water, built-up areas, cloud were located and marked during ground truth. The tree species classification scheme is shown in Table 1. The sampling unit used in this paper was a pixel and the samples were selected from the CASI Hyperspectral image based on the field survey. There are 18,540 ground truth sample pixels were selected from the CASI hyperspectral image. And 10-fold cross-validation method was used in accuracy estimating. Average overall accuracy was then computed from the confusion matrix with 10th classification.
Methodology
The overall method used in this study is shown as flowchart in Fig. 2. First the minimum noise fraction (MNF) transformation to reduce the dimension of the CASI image and then the grey level co-occurrence matrix (GLCM) is used to extract the textural information. The MNF features are combined with textural features and are used for classification of tree species by SVM with different kernel.
Minimum Noise Fraction Transform
MNF analysis first suggested with Green et al. (1988). MNF transforms were used to determine the inherent dimensionality of image data, to segregate noise in the data, and to reduce the computational requirements for subsequent processing. The MNF transform is essentially two cascaded principal component’s transformations. The first transformation, based on an estimated noise covariance matrix, decorrelates and rescales the noise in the data. This first step results in transformed data in which the noise has unit variance and no band-to-band correlations. The second step is a standard principal components transformation of the noise-whitened data. For the purposes of further spectral processing, the inherent dimensionality of the data is determined by examination of the eigenvalues and the associated images. The data space can be divided into two parts: one part associated with large eigenvalues and corresponding eigenimages, and a complementary part with near zero eigenvalues and noise-dominated images (Jouan 2007; Nielsen 2011). By using only the coherent portions, the noise is separated from the data, thus improving spectral processing results. Based on MNF results, the first 20 eigenvectors which had the cumulative contribution rate up to 95 % were selected and then used as input for the classifiers.
Texture-Based Features
The texture-based features extracted from the grey-level co-occurrence matrix (GLCM). The GLCM represents the distance and angular spatial relationship over an image sub region of the specified size. The GLCM quantifies texture by measuring the spatial frequency of co-occurrence of pixel grey levels in a user-defined moving kernel and forms a co-occurrence of pixel of kernel. During the computation of the GLCM texture measure, consideration should be given to the window size that would best capture the target classes. The optimal window size could be determined through the image spatial resolution and the tree canopy size. In this paper the semi-variograms method was used to determine the optimal windows size (Onojeghuo and Blackburn 2011). The optimal window size for calculating the GLCM measures is 7. A series of GLCM texture measures were calculated according to the following (Onojeghuo and Blackburn 2011):
Where CON is the contrast, i,j are row and col of value in the grey level co-occurrence matrix, DIS is the dissimilarity value, HOM is the homogeneity, ENT is the entropy, ASM is the angular second moment of grey level co-occurrence matrix, COR is the correlation value of grey level co-occurrence matrix.
Classification Method and Accuracy Assessment
The support vector machine (SVM) was used for tree species classification in this paper. SVM classifiers have undergone great development in the last 10 years and have been successfully applied to several remote sensing problems. Let us consider a binary classification problem. And assume that the training set consists of Q vectors x p∈Rq with the corresponding target yp ∈ {−1; +1}, where “+1” and “-1” denote the labels of the considered classes.
The linear SVM approach consists of mapping the data into a higher dimensional feature space to separate the two classes by means of an optimal hyperplane defined by a weight vector w and a bias b. The optimal hyperplane is the one that minimizes a cost function, which expresses a combination of two criteria: margin maximization and error minimization. It is defined as (7) and (8)
Where ξ p are the so-called slack variables and ξ p ≥ 0.
The constant C which called cost parameter represents a regularization parameter that controls the shape of the discriminant function, and consequently, the decision boundary when data are nonseparable. The above optimization problem can be reformulated through a Lagrange functional for which the Lagrange multipliers can be found by means of a dual optimization leading to a quadratic programming solution. According to the nonlinear case, the SVM uses the kernel functions to generalize the non-linear decision boundaries. Commonly use SVM kernels include polynomial, radial basis function (RBF) and sigmoid kernels. The SVM classifier was also easily extended to multiclass problems with One-Against-One and One-Against-All methods (Vapnik 1998).
Several SVM programs have been developed and made publicly available. In this study, we used the LIBSVM program developed by Hsu et al. (Hsu et al. 2001). We choose the linear, quadratic polynomial, cubic polynomial, sigmoid and RBF kernel to test the effect of different kernels in tree species discrimination. The parameters that are needed in the LIBSVM program were predefined as suggested in Hsu et al. (2001). The SVM need two type of parameters: 1) the kernel function type and its parameters; 2) the cost parameter C. For each kernel function, the kernel parameters are not the same. The Table 2 list the parameters for each kernel function. The appropriate values for these parameters were determined with the guidance of Hsu et al. (2001). Specifically, the values for γ, r, d and C was systematically change from low to high. For each combination of γ, r, d and C, the prediction accuracy of the trained SVM model was estimated through cross-validation. The combination giving the highest prediction accuracy was used to tree species classification.
In order to evaluate the effectiveness of the proposed tree species classification strategy and achieve the goal of this paper, there are three level experiments were defined: 1) tree species classification with SVM using MNF features; 2) tree species classification with SVM using MNF and texture-based features; 3) SVM classifier with 5 different kernels such as linear, quadratic polynomial, cubic polynomial, sigmoid and RBF kernel.
Results
The overall accuracies of SVM classification method with different kernels and features are given in Table 3. From the table, it can be see that the best classification result of all combinations is the linear kernel function with MNF and texture-based features. The Fig. 3 shows the classification results using SVM with linear kernel function and MNF and texture-based features.
The classification accuracy with SVM is different when kernel function changes. On average, the linear kernel function gives the best classification results, followed by RBF and sigmoid kernel. Polynomial kernel functions give the worst classification results. This indicates that the polynomial kernels are not good for MNF and texture-based features in tree species classification in this case.
Considering the features in SVM algorithm, we can find that the over accuracy in MNF combined with texture-based features is higher than that only with MNF features, but the overall accuracy increase is low with all kernel functions in SVM.
The SVM method with linear, RBF and sigmoid kernel functions all perform well in tree species classification with MNF and MNF combined texture-based features when using CASI hyperspectral image. However, the kernel function also influences the classification results. How to select the kernel function maybe has the relationship with the feature types. In this paper, we find that when we use the MNF and textures based features, the linear kernel function has the best the result. The features types may also influence the hyperspectral images classification results. The spectral feature combined with the context feature extracted from hyperspectral images can promote the classification. In this paper, it is found that MNF features combined with texture based features increase the accuracy of the classification though the increase is low.
Discussion
The tree species classification at crown level in forests with high tree species diversity is a big problem with only spectral or textural information. Airborne hyperspectral sensor provides data with both high spatial and spectral resolution imagery which has the huge advantages in tree species classification. The research results in this paper showed that the hyperspectral information combined with textural information can promote the accuracy in tree species classification. And the results are consistent with other researcher’s reports (Immitzer et al. 2012). Overall classification accuracies of 75–90 % are achieved by several groups of researchers in tree species classification. The accuracy in this paper (85.92 %) is in line with the accuracies reported in comparable studies (Clark et al. 2005; Hestir et al. 2008; Zhang et al. 2006; White et al. 2010).
However, classification with MNF features combined the texture features didn’t give much improvement in overall accuracy. The result is not same with the result of Onojeghuo and Blackburn (2011) which indicate that the texture information highly increases the overall accuracy. The mainly reason is that tree species type in these two studies are not similarity. In our experiments, fir, red pine, larch are all coniferous trees, and the appearance is almost the same. And also the same condition in the birch and willow which are broadleaved trees. However, in Onojeghuo and Blackburn (2011) paper, the textural information was used to distinguish the broadleaved, coniferous, grassland and reedbeds, and these four types had the significant difference appearance. Whether textural features increase a little or big accuracy in tree classification may depend on the type of vegetation. If the species are in the same type, the textural features may not give much improvement and if the species are in different type, the textural features may improve much.
SVM is an advanced machine learning algorithms for classification, but the classification accuracy with SVM is different when kernel function changes. How to select the suitable kernel function is depend on the number of features, number of samples and the distribution of the feature (Keerthi and Lin 2003). According to the Keerthi and Lin (2003), if the feature distribution didn’t known, the RBF kernel is a reasonable first choice and this kernel nonlinearly maps samples into a higher dimensional space, so it can handle the case when the relation between class labels and attributes is nonlinear. Furthermore, the linear kernel is special case of RBF; in addition, the sigmoid kernel behaves like RBF for certain parameters (Lin and Lin 2003). Compared with the RBF kernel, the polynomial kernel has more hyper-parameters which will influence the complexity of computation in SVM. The difference of kernel parameters with each kernel function can be seen in Table 2. So, in this paper the accuracy of RBF kernel, linear kernel and sigmoid kernel is almost the same and higher than the polynomial kernel. Another possible reason for different accuracy with different kernel function is the input features. In this paper, the feature number is 26 and the sample number is 18,540. The sample number is much larger than the feature number, so in hyper-plane the linear kernel may classify the 8 class types.
Conclusion
The results indicate that hyperspectral images provide the ability for effective forest species recognition. The spectral and context features were used as input for SVM classifier and compared the effective of different kernel function in SVM classifier for forest tree species classification. The classification results indicate that the SVM method with linear, RBF and sigmoid kernel functions all perform well in tree species classification when using CASI hyperspectral image and the linear kernel function has the best result. MNF features combined with texture based features increase the accuracy of the classification.
References
Bajorski, P. (2011). Statistical inference in PCA for hyperspectral images. IEEE Journal of Selected Topics in Signal Processing, 5(3), 438–445.
Chen, E. X., Li, Z. Y., Tan, B. X., Liang, Y. Z., & Zhang, Z. L. (2007). Validation of statistic based forest types classification methods using hyerspectral data. Scientia Silvae Sinicae, 43(1), 84–89.
Clark, M. L., Roberts, D. A., & Clark, D. B. (2005). Hyperspectral discrimination of tropical rain forest tree species at leaf to crown scales. Remote Sensing of Environment, 96(3–4), 375–398.
Gong, P., Pu, R. L., & Yu, B. (1998). Conifer species recognition with seasonal hyperspectral data. Journal of Remote Sensing, 2(3), 211–217.
Green, A. A., Berman, M., Switzer, P., & Craig, M. D. (1988). A transformation for ordering multispectral data in terms of image quality with implications for noise removal,”. IEEE Transaction on Geoscience and Remote Sensing, GRS-26(1), 65–74.
Hestir, E. L., Khanna, S., Andrew, M. E., Santos, M. J., Viers, J. H., Greenberg, J. A., et al. (2008). Identification of invasive vegetation using hyperspectral remote sensing in the California delta ecosystem. Remote Sensing of Environment, 112(11), 4034–4047.
Hsu, C.W., Chang, C. C., & Lin, C. J. (2001). A practical guide to support vector classification
Huang, C. Y., & Asner, G. P. (2009). Applications of remote sensing to alien invasive plant studies. Sensors, 9, 4869–4889.
Immitzer, M., Atzberger, C., & Koukal, T. (2012). Tree species classification with random forest using very high spatial resolution 8-band worldview-2 satellite data. Remote Sensing, 4, 2661–2693.
Jose, P. H., & David, A. L. (1996). Classification of remote sensing image having high spectral resolution. Remote Sensing of Environment, 57, 119–126.
Jouan, A. (2007). FastICA(MNF) for feature generation in hyperspectral imagery, 10th International Conference on Information Fusion, 1–8
Keerthi, S. S., & Lin, C. J. (2003). Asymptotic behaviors of support vector machines with gaussian kernel. Neural Computation, 15(7), 1667–1689.
Kozoderov, V. V., & Dmitriev, E. V. (2011). Remote sensing of soils and vegetation: quantitative parameters retrieval using pattern-recognition techniques and forest stand structure assessment. International Journal of Remote Sensing, 32(20), 5699–5717.
Leempoel, K., Bourgeois, C., Zhang, J., Wang, J., Chen, M., Satyaranayana, B., Bogaert, J., & Dahdouh-Guebas, F. (2013). Spatial heterogeneity in mangroves assessed by GeoEye-1 satellite data: a case-study in Zhanjiang Mangrove National Nature Reserve (ZMNNR). China, Biogeosciences Discussion, 10, 2591–2615.
Liang, Y. Q., & Zeng, H. (2009). Application of hyperspectral remote sensing in identification of vegetation characteristics. World Forestry Research, 22(1), 41–47.
Lin, H. T., & Lin, C. J. (2003). A study on sigmoid kernels for SVM and the training of non-PSD kernels by SMO-type methods. National Taiwan University: Technical report, Department of Computer Science.
Lu, D. (2005). Integration of vegetation inventory data and landsat TM image for vegetation classification in the western Brazilian amazon,”. Forest Ecology and Management, 213, 369–383.
Lu, D. S., Batistella, M., Moran, E., & de Miranda, E. E. (2008). A comparative study of landsat TM and SPOT HRG images for vegetation classification in the Brazilian amazon. Photogrammetric Engineering and Remote Sensing, 74(3), 311–321.
Nielsen, A. A. (2011). Kernel maximum autocorrelation factor and minimum noise fraction transformations. IEEE Transaction on Geoscience and Remote Sensing, 20(3), 612–624.
Onojeghuo, A. O., & Blackburn, G. A. (2011). Mapping reedbed habitats using texture-based classification of quickbird imagery. International Journal of Remote Sensing, 32(23), 8121–8138.
Sedano, F., Gong, P., & Ferrao, M. (2005). Land cover assessment with MODIS imagery in southern African miombo ecosystems. Remote Sensing of Environment, 98(4), 429–441.
Smith, K. L., Steven, M. D., & Colls, J. J. (2004). Use of hyperspectral derivative ratios in the red-edge region to identify plant stress responses to gas leaks. Remote Sensing Environment, 92, 207–17.
Ustin, Susan, L., John, A., & Gamon. (2010). Remote sensing of plant functional types. New Phytologist, 186(4), 795–816.
Vaiphasa, C., Skidmore, A. K., Boer, W. F., & Vaiphasa, T. (2007). A hyperspectral band selector for plant species discrimination. ISPRS Journal of Photogrammetry and Remote Sensing, 62, 225–235.
Vapnik, V. N. (1998). Statistical Learning Theory. Hoboken: Wiley.
Wang, K., Steven, E., Franklin, X. G., & Cattet, M. (2010). Remote sensing of ecology, biodiversity and conservation: a review from the perspective of remote sensing specialists. Sensors, 10(11), 9647–9667.
White, J. C., Gómez, C., Wulder, M. A., & Coops, N. C. (2010). Characterizing temperate forest structural and spectral diversity with Hyperion EO-1 data. Remote Sensing of Environment, 114, 1576–1589.
Zaw Htun, N., Mizoue, N., & Yoshida, S. (2011). Classifying tropical deciduous vegetation: a comparison of multiple approaches in popa mountain park, Myanmar. International Journal of Remote Sensing, 32(24), 8935–8948.
Zhang, J., Rivard, B., Sánchez-Azofeifa, A., & Castro-Esau, K. (2006). Intra- and inter-class spectral variability of tropical tree species at La selva, Costa Rica: implications for species identification using HYDICE imagery. Remote Sensing of Environment, 105, 129–141.
Zhao, D., Huang, L., Li, J., & Qi, J. (2007). A comparative analysis of broadband and narrowband derived vegetation indices in predicting LAI and CCD of a cotton canopy. ISPRS Journal of Photogrammetry and Remote Sensing, 62(1), 25–33.
Acknowledgments
This research was supported by National High Technology Research and Development Program of China (863 Program) (grant No. 2012AA120906). Additional funding and supporting were also provided by the Fundamental Research Funds for the Central Universities (grant No. 2014QC018) and Key Laboratory for National Geographic Census and Monitoring, National Administration of Surveying, Mapping and Geoinformation (grant No. 2013NGCM05)
Author information
Authors and Affiliations
Corresponding author
About this article
Cite this article
Dian, Y., Li, Z. & Pang, Y. Spectral and Texture Features Combined for Forest Tree species Classification with Airborne Hyperspectral Imagery. J Indian Soc Remote Sens 43, 101–107 (2015). https://doi.org/10.1007/s12524-014-0392-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12524-014-0392-6