Abstract
In multicellular organisms, germ cells are an extremely specialized cell type with the vital function of transmitting genetic information across generations. In this respect, they are responsible for the perpetuity of species, and are separated from somatic lineages at each generation. Interestingly, in the past two decades research has shown that germ cells have the potential to proceed along two distinct pathways: gametogenesis or pluripotency. Unequivocally, the primary role of germ cells is to produce gametes, the sperm or oocyte, to produce offspring. However, under specific conditions germ cells can become pluripotent, as shown by teratoma formation in vivo or cell culture-induced reprogramming in vitro. This phenomenon seems to be a general propensity of germ cells, irrespective of developmental phase. Recent attempts at cellular reprogramming have resulted in the generation of induced pluripotent stem cells (iPSCs). In iPSCs, the intracellular molecular networks instructing pluripotency have been activated and override the exclusively somatic cell programs that existed. Because the generation of iPSCs is highly artificial and depends on gene transduction, whether the resulting machinery reflects any physiological cell-intrinsic programs is open to question. In contrast, germ cells can spontaneously shift their fate to pluripotency during in-vitro culture. Here, we review the two fates of germ cells, i.e., differentiation and reprogramming. Understanding the molecular mechanisms regulating differentiation versus reprogramming would provide invaluable insight into understanding the mechanisms of cellular reprogramming that generate iPSCs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mammalian development proceeds unidirectionally from totipotent early embryos to pluripotent cells, to multipotent/unipotent organ-specific stem cells that are the source of terminally differentiated somatic cells. However, advances in science and technology have enabled us to reverse this unidirectional process by inducing cellular reprogramming. Cellular reprogramming can be accomplished artificially by three different methods: nuclear transfer, cell fusion, and direct reprogramming [1]. In particular, the recent success in generating induced pluripotent stem cells (iPSCs) from cells of somatic tissue origin has given rise to a new era of cellular reprogramming research [2–4]. Although iPSC generation and the other two methods of cellular reprogramming require highly artificial manipulations, germ cells intrinsically have the potential to give rise to pluripotent stem cells when placed under appropriate conditions. Germ cells are naturally committed to undergo spermatogenesis in the male or oogenesis in the female while maintaining the expression of several crucial iPSC-reprogramming factors. Studies in the past two decades have revealed the plasticity of germ cells as a source of pluripotency, resulting in a challenge to uncover the underlying mechanisms. In this review, we describe advances resulting from studies of germ cell-derived pluripotent stem cells in mouse and human.
Germ cell development: segregation from pluripotent cells to gametogenesis
In mammals, germ cell fate is not defined by maternally inherited determinants but is rather induced in pluripotent cells by signals from neighboring tissues. Therefore, this process distinguishes germ cells as a lineage with different fates and identities that depart from pluripotency. During mouse embryogenesis, germ cells are first specified from a part of the epiblast in the proximal posterior portion destined to become extraembryonic mesoderm. The specification is a response to WNT3 secreted by the visceral endoderm and BMP4/8b secreted by the extraembryonic ectoderm [5]. The signals give rise to the precursors of germ cells, the primordial germ cells (PGCs), at approximately embryonic day (E) 6.25–6.75 in mouse. The PGC precursors begin to express two transcriptional regulators, Blimp1 (also known as Prdm1) [6] and, shortly thereafter, Prdm14 [7], which act in coordination to suppress somatic programming, re-activate pluripotency-associated genes (e.g., Oct4, Sox2, and Nanog), and induce epigenetic reprogramming [7, 8]. In parallel, a pluripotency-associated gene LIN28 positively regulates Blimp1 expression by inhibiting repressive microRNAs [9]. At E7.25, PGCs can be identified as a cluster of approximately 40 cells with alkaline phosphatase activity, and expressing Stella (also known as PGC7 or Dppa3), Fragilis (also known as Mil1 or Ifitm3) [10], and Nanos3 [11].
After specification, the next step in germ cell development is migration of the PGCs to the genital ridges that will form the future gonads. PGCs migrate along the hindgut and dorsal mesentery while proliferating, and reach the genital ridges by E10.5–12.5. In the course of migration, PGCs receive a variety of exogenous cues, for example stem cell factor (SCF), from somatic cells. PGCs express the receptor C-KIT, which is involved in the motility, directionality, and survival of PGCs [12–15]. The survival of the PGCs is also supported by intracellular proteins, for example OCT4 [16], NANOG [17], and NANOS3 [18]. Another aspect of the development of migrating PGCs is extensive, genome-wide epigenetic reprogramming that leads to alterations of DNA and histone modifications [19, 20], and X chromosome reactivation (in females) [21]. For instance, BLIMP1 forms a protein complex with the arginine methyltransferase PRMT5 and catalyzes H2A/H4R3 methylation by E10.5 [22]. Thereafter, the BLIMP1/PRMT5 complex translocates from the nucleus to the cytoplasm, resulting in the loss of H2A/H4R3 methylation.
Upon colonization of the genital ridges, PGCs undergo dynamic changes in their global gene expression profile [23], genomic imprinting [24], and sex determination [25]. Expression of marker genes characteristic of the migration stage is reduced, as shown by changes in the levels of Blimp1 [26], Prdm14 [7], and Nanos3 [11]. The postmigratory PGCs start to express another set of genes including Vasa [27] and Dazl [28]. Concurrently, genomic imprinting inherited from the parent is erased. The timing of erasure depends on each imprinted gene but is completed by E12.5 [29]. Although the processes described thus far occur in both sexes, germ cells undergo distinct sex-specific development from this point. In the genital ridges, PGCs determine their sex in accordance with that of the surrounding somatic cells; hence, female germ cells enter meiosis, whereas male germ cells undergo mitotic arrest at E13.5. Germ cells in the embryonic ovary become oocytes that initiate meiosis in response to retinoic acid (RA) [30]. Establishment of female imprinting occurs after oocyte growth after birth [31, 32]. In contrast, germ cells in the embryonic testis, now called gonocytes, progressively acquire androgenetic imprinting from E14.5 to spermatogonia at birth [33], while configuring male-type gene expression including Nanos2 [18] and Dnmt3L [34].
Returning to the pluripotent state from lineage-committed PGCs
Testicular teratomas implicate PGC dedifferentiation in vivo
As described above, germ cells are ordinarily committed to a gametogenic fate, but they maintain the potential to return to the pluripotent state, as indicated by the formation of spontaneous or experimental testicular teratomas. Testicular teratoma formation is a rare event in most mouse strains. In the 1950s, Stevens established the 129/SvJ inbred mouse strain in which incidence of spontaneous testicular teratoma formation is approximately 1–10 % [35–37]. Furthermore, incidence of testicular teratoma formation was dramatically higher in spontaneous Ter mutant mice of this line (129/Sv-ter mice)—94 % in the homozygous mice [38, 39]. Importantly, in transplantation experiments, genital ridge tissues depleted of PGCs did not develop into tumors in transplant recipients, suggesting that the PGCs are responsible for the testicular teratomas [40].
Genetic analysis has been conducted to identify the defects that predispose mice to teratoma formation in the 129 strains. Several susceptibility genes have been isolated [35, 41]. Dnd1 proved to be the causative gene for the Ter mutation [42]. A point mutation producing an alternative stop codon was identified within the Dnd1 gene in 129/Sv-ter mice, and the Ter phenotype could be rescued by an intact Dnd1 transgene. How the Dnd1 deficiency leads to teratoma formation remains to be determined. DND1 has an RNA recognition motif that binds the 3′UTR of target mRNAs and protects them from miRNA-associated inhibitory machinery [43]. Therefore, the DND1 target genes may be important suppressors of dedifferentiation. Another study identified Dmrt1 as a suppressor of pluripotency in PGCs. DMRT1 is a transcription factor expressed in the gonads and controls male sex determination [44, 45]. Similar to the 129/Sv-ter mice, the loss of Dmrt1 resulted in a high incidence of testicular teratomas in 129/Sv mice [46]. Dmrt1-deficient PGCs escaped from mitotic arrest in the genital ridges and ectopically overexpressed the pluripotency-associated genes Oct4, Sox2, and Nanog, even at birth. Because the DMRT1 protein binds to the Sox2 promoter region, it could be a direct negative regulator of pluripotency-associated genes.
Both the Dnd1 and Dmrt1 deficiencies are implicated in PGC dedifferentiation, but the phenomenon is restricted to 129 strain mice. What leads to the different susceptibility of PGCs to transformation among different mouse strains is not understood, but one factor may be the sensitivity of PGCs to BAX-mediated apoptosis [47]. In addition, loss of the RNA-binding protein DAZL might release PGCs from repression of the pluripotent state. In Dazl-deficient mice in the C57BL/6 background, germ cells at E15.5 abnormally retained robust expression of Oct4, Sox2, and Nanog in both sexes [48]. However, they did not develop teratomas but instead underwent apoptosis [49]. This observation suggests that prolonged expression of pluripotency-associated genes alone might not be sufficient to cause PGC transformation.
Conversion of PGCs to pluripotent stem cells
In culture, short-term PGC proliferation is supported by SCF and leukemia inhibitory factor (LIF) [50, 51]. When cultured in combination with fibroblast growth factor 2 (FGF2), PGCs continue to proliferate and dedifferentiate into pluripotent stem cells called “embryonic germ cells” (EGCs) [52, 53]. Unlike the high incidence of teratoma formation in the 129/Sv strains, EGCs seem to arise irrespective of mouse strain. Dedifferentiation from PGCs to EGCs occurs rapidly upon FGF2 stimulation, and PGC gene expression shifts toward pluripotency within the first 24 h of FGF2 treatment [54]. Subsequently, EGCs form ESCs-like colonies by approximately 10 days in culture. Although cytokine requirements to produce EGCs differ from those for ESCs, EGCs no longer require FGF2 and SCF for their maintenance after colony formation. EGCs resemble ESCs in terms of culture conditions, colony morphology, marker gene expression, differentiation capacity in vitro and in vivo, and chimeric mouse formation by blastocyst injection [55, 56]. EGCs can be derived from E8.5–12.5 PGCs, but the efficiency declines later in embryogenesis and no EGCs are obtained from E15.5 PGCs [56].
Although the characteristics of EGCs are basically quite similar to those of ESCs, the epigenomes are different. During embryonic development from which EGCs can originate, PGCs go through extensive epigenetic reprogramming, including genome-wide DNA demethylation, erasure of genomic imprinting, and reactivation of the X chromosome. EGCs inherit these epigenotypes reflecting the parental PGCs in both male and female [56–58]. For example, in EGCs derived from E8.0–8.5 PGCs, approximately half of the cell lines have erased the imprinting of the Igf2r locus, whereas all EGCs lines have lost the Igf2r locus imprinting when established from E12.5 PGCs. It is notable that EGCs not only reflect the epigenotype of the parental PGCs but also still retain the ability to induce demethylation of the somatic genome in EGC-thymic lymphocyte hybrid cells [58]. Consistent with the loss of allele-specific DNA methylation, EGCs have an expression pattern of imprinted genes different from that of ESCs [59], and EGC-derived embryos have developmental abnormalities associated with imprinting aberrations [57]. Furthermore, global gene expression profiling identified approximately 100 genes, in addition to imprinted genes, with different expression in ESC and EGC lines. Whether the different expression of these genes is a secondary effect attributable to the unique epigenomes of the ESCs and EGCs, or reflects some other features of PGC origin, remains to be determined.
Signaling pathways underlying PGC–EGC dedifferentiation
Because the generation of EGCs is well-established as an assay system, different aspects of the molecular basis of PGC–EGC conversion have been addressed. The definitive requirement for FGF2/SCF/LIF treatment has provided important clues about the identity of the intracellular mediators. In LIF signaling, LIF binds to LIF receptor/gp130 heterodimers and the intracellular signal is transmitted via the JAK/STAT3 pathway. Cultured PGCs also utilize these same mediators, in response to the LIF signal, to dedifferentiate into EGCs [60, 61]. Interestingly, this signal is not essential for the first 24 h of PGC culture, and no STAT3 protein is detected in PGCs over this period [61].
Other studies have focused on the role of phosphoinositide-3 kinase (PI3K)/AKT signaling. This signaling pathway is commonly activated by a variety of growth factors and has crucial functions in cell proliferation, survival, and self-renewal [62, 63]. PI3K function is antagonized by the tumor-suppressor protein phosphatase and tensin homolog (PTEN) [64]. Although Pten-deficient mice are embryonic lethal, the heterozygous mice survive and form testicular teratomas, suggesting that PTEN inhibits germ cell dedifferentiation [65]. Indeed, transgenic mice with a PGC-specific Pten ablation form teratomas [66]. Moreover, the efficiency of EGC derivation is dramatically enhanced by PGC-specific Pten depletion. The involvement of PTEN in blocking the generation of EGCs was confirmed by the transduction of Pten antisense oligonucleotides [67]. In addition, AKT activation promoted EGC derivation at levels comparable with those of Pten deficiency [68]. Notably, PGCs with AKT activation became EGCs even in the absence of FGF2. Taken together, these results suggest the PI3K/AKT pathway is likely to be responsible for mediating the FGF2 signal in the PGC–EGC conversion.
What, then, are the downstream targets of the PI3K/AKT pathway? AKT signaling has been shown to inhibit p53 and glycogen synthase kinase-3 (GSK3), directly or indirectly. In fact, AKT activation enhanced the stability of MDM2, a negative regulator of p53, and the inhibitory phosphorylation of GSK3 in PGCs [68]. Furthermore, p53 deficiency not only facilitated efficient EGC derivation but also enabled generation of EGCs in the absence of FGF2. In contrast, the contribution of the WNT/β-catenin pathway, which is blocked by GSK3, remains controversial. WNT/β-catenin signaling has been shown to promote the maintenance and acquisition of pluripotency in ESCs or iPSCs [69]. However, PGC-specific β-catenin stabilization did not cause teratoma formation, and no enhanced EGC derivation was observed in response to GSK3 inhibitor treatment [68, 70]. However, one group has reported that a GSK3 inhibitor in combination with a MEK inhibitor resulted in highly efficient production of EGCs [71]. Because this last finding implicates crosstalk between β-catenin and the MAPK cascade, further investigation will be required to determine how WNT/β-catenin signaling is involved in the PGC–EGC transition. In addition to these signaling pathways, it has been suggested that other pathways contribute to mediation of dedifferentiation signals, because the effects of FGF2 could be mimicked by forskolin or RA [60], and those of SCF could be mimicked by estrogens [67].
Epigenetic regulators and EGC derivation
In addition to signaling molecules, epigenetic factors are involved in EGC derivation. BLIMP1 and PRDM14 are indispensable for reactivation of pluripotency-associated genes and epigenetic reprogramming in PGC specification [7, 8]. Both proteins contain a PR domain, which is similar to the active domain of histone methyltransferases. According to in-vivo studies, PGCs from Prdm14-deficient embryos fail to form EGCs in culture [7]. Sequential analysis of dedifferentiation kinetics also revealed that BLIMP1 and PRMT5 contribute to PGC–EGC conversion. The BLIMP1 protein disappears within 2 days under EGC derivation culture conditions, which leads to the upregulation of target genes such as c-Myc and Klf4 [61]. In contrast, the PRMT5 protein is maintained throughout the culture period, but its intracellular location changes from the nucleus in PGCs to the cytoplasm in EGCs, suggesting that PRMT5 has different functions in each cell type. Although the BLIMP1/PRMT5 complex leads to histone methylation [22], PRMT5 also catalyzes arginine methylation of p53, which, in turn, modulates the target specificity of p53 [72]. Other research has indicated that PRMT5 regulates p53 translation [73]. Given these insights, PRMT5 function in PGC dedifferentiation may be mediated, in part, via p53 regulation. Furthermore, it is likely histone acetylation is associated with EGC derivation, because trichostatin A, an inhibitor of histone deacetylases, can substitute for FGF2 signaling [61].
A PGC subpopulation highly competent to transform into EGCs
Recent studies of cell-surface proteins have revealed that PGCs are a heterogeneous cell population. PGCs can be fractionated by flow cytometry into subpopulations with distinct expression of α6 integrin and C-KIT [74]. PGCs with little or no α6 integrin expression have high rates of apoptosis compared with α6 integrin positive cells in E14.5 female mouse embryos. Similarly, PGCs with little or no C-KIT expression have higher rates of apoptosis than C-KIT positive cells in E12.5–14.5 embryos of either sex. Importantly, there is a correlation between α6 integrin expression and competency to dedifferentiate into EGCs. When PGCs with little or no α6 integrin are cultured under EGC derivation conditions, instead of undergoing apoptosis, these cells become EGCs at rates higher than α6 integrin-positive cells [75]. In addition, higher EGC derivation is correlated with a side-population phenotype of PGCs also. Heterogeneity also arises during EGC formation in the derivation cultures. Although freshly isolated PGCs express FGF receptor 3 (FGFR3) at low levels only, a few cells become strongly positive after 24 h in culture [54]. Eventually, high expression of FGFR3 is observed for all colony-forming cells, consistent with the importance of FGF2 signaling in the generation of EGCs.
Human EGCs
Although establishment of mouse ESCs greatly preceded generation of mouse EGCs, generation of human EGCs was reported at approximately the same time as that of human ESCs [76, 77]. In mice, EGCs have cytological features quite similar to those of ESCs. In contrast, studies have revealed that human EGCs are quite different from human ESCs in several respects [78]. First, culture conditions for derivation of human EGCs are nearly identical with those for mouse EGCs [77, 79–81], but generation of human and mouse ESCs requires different cytokines for each species. The efficiency of human EGC derivation is reduced by withdrawal of LIF or FGF2 [77]. Conversely, in the presence of feeder cells overexpressing LIF, human EGCs tend to express pluripotency marker genes at higher levels than controls [81]. Second, human ESCs and EGCs can easily be discriminated by their colony morphology: the former form flat colonies whereas the latter form tightly compact, dome-shaped colonies. Third, the self-renewal capacity of human EGCs in culture is not infinite. The proliferation activity of human EGCs declines and expression of pluripotency markers is attenuated as the number of cell passages increases. Why human EGCs cannot maintain their proliferation and undifferentiated state is currently unknown, but long-term culture of human EGCs may require conditions different from those currently in use. Fourth, human EGCs express SSEA1 on their cell surface, in contrast with human ESCs, which do not express SSEA1. Thus, human EGCs have unique characteristics distinct from those of both human ESCs and mouse EGCs. Further characterization of these cells is needed, however, because of the limited number of studies on human EGCs.
Why are PGCs per se not pluripotent?
An open question is how PGCs are insulated from pluripotency while expressing pluripotency-associated genes, including the iPSCs-reprogramming factors Oct4, Sox2, and Nanog. Indeed, PGCs cannot be incorporated into chimeras even by transplantation into blastocysts [54, 82]. Although Oct4 and Nanog are critical for self-renewal in ESCs and reprogramming in iPSCs, conditional depletion of these factors in PGCs resulted in apoptosis, suggesting they are involved in PGC survival [16, 17] and specification [83], rather than in self-renewal and pluripotency. Furthermore, global gene expression analysis revealed different expression profiles for OCT4 target genes in PGCs and ESCs [84]. In particular, expression of Klf4 and Tbx3, which are both core factors in the pluripotency network [85], is very much lower in PGCs than in ESCs. Thus, OCT4 may have different target genes in PGCs than in ESCs. In addition, PGC dedifferentiation may require other signals, for example those related to cellular transformation, to superimpose pluripotency on the germ cell program. This speculation is supported by the higher rate of EGC formation in cells depleted of tumor suppressors (Pten and p53) and the absence in PGCs of the expression of ERas [86], which is an oncogenic Ras expressed in ESCs [87].
Pluripotency in spermatogenic cells
Male germline stem cells as a spermatogonial stem cell line
Mammalian spermatogenesis starts at puberty and continues throughout life because of the self-renewal and differentiation of spermatogonial stem cells (SSCs). In 2003, long-term cultivation of SSCs was achieved by use of mouse neonatal testis [88]. The cells, called “germline stem cells” (GSCs), expand stably and clonally in culture and contribute to normal spermatogenesis when transplanted into the seminiferous tubules of mouse testis. DNA methylation [89], transcription [90], and protein [91] profiles of GSCs are clearly distinguishable from those of ESCs. The cultured GSCs depend predominantly on stimulation with glial cell-line-derived neurotrophic factor (GDNF) [92], which is secreted by Sertoli cells in the testis and regulates SSC self-renewal [93, 94]. Since these pioneering findings, equivalent cell lines have been established by use of adult mouse [95], rat [96, 97], hamster [98], and rabbit [99] testes. Because SSCs are only a small fraction of testicular cells, examining the molecular composition of these cells had long been difficult. Successful derivation of GSCs will enable researchers to investigate the properties of SSCs by use of conventional molecular and cellular biological techniques. Moreover, by taking advantage of the SSC activity of GSCs, the feasibility of using such cells as a tool for genetic modification has been demonstrated using a variety of vectors, for example plasmids [97, 100–102], viruses [100, 101, 103], and transposons [104, 105].
Because GSCs are spermatogenic cells, androgenetic genomic imprinting is observed—DNA hypermethylation of paternally imprinted genes and hypomethylation of maternally imprinted genes [89, 106]. Imprinting status and karyotype were stable in GSCs cultured for more than 2 years [107]. In contrast, GSCs derived from E12.5–18.5 fetal gonocytes have a significant imprinting defect [108]. The embryonic GSCs (eGSCs) are indistinguishable from postnatal GSCs in cell morphology, proliferation, marker gene expression, and spermatogenic potential. Furthermore, DNA methylation status of imprinted genes and repetitive sequences are normal for eGSCs. However, the histone modification status of imprinted genes is altered, possibly because of different expression of histone modifiers, and the eGSC-derived offspring have aberrant DNA methylation of imprinted genes and growth abnormalities. Furthermore, the imprinting defect is heritable. Nevertheless, neither GSCs nor eGSCs develop teratomas, indicating that they are definitely unipotential stem cells for spermatogenesis.
Derivation of pluripotent stem cells of spermatogenic origin
Considering that EGCs have not been successfully derived from cells older than E12.5, postnatal SSCs were not thought to be capable of pluripotency. However, evidence was first presented in 2004 showing that SSCs can give rise to pluripotent ESC-like cells. In the course of culturing neonatal testis to generate GSCs, epiblast-like colonies occasionally emerged [106]. Under ESC culture conditions, these cells developed ESC-like morphology, continued to grow and express pluripotency marker genes, and differentiated into all three germ layers. Thus, these cells were designated “multipotent GSCs” (mGSCs). Despite their origin from GSC derivation cultures, mGSCs could not be stably maintained in an undifferentiated state under GSC culture conditions; they required LIF, but not GDNF, to self-renew [109]. Clonal tracking has shown that unipotent GSCs and pluripotent mGSCs may actually share identical SSC origin [109]. Nonetheless, mGSCs are different from GSCs in genomic imprinting. Whereas GSCs clearly have androgenetic imprinting, the paternally imprinted genes H19 and Meg3IG are partially demethylated in mGSCs [106].
After the generation of mGSCs from neonatal SSCs it was discovered that pluripotent stem cells could be obtained from adult mouse SSCs also [110–114]. The first adult testis-derived ESCs-like colonies were generated by culturing purified SSCs in the presence of GDNF and serum [110]. The resulting cells satisfied the pluripotency criteria: expansion in a manner similar to ESCs, expression of pluripotency marker genes, and differentiation into teratomas and chimeras. These cells were designated “multipotent adult GSCs” (maGSCs) to distinguish them from mGSCs. Global and comparative analysis has revealed that maGSCs are very similar to ESCs in their pattern of DNA methylation [115] and histone modification [116], and microRNA [117, 118], mRNA [119], and protein [120] expression. However, maGSCs do not simply correspond to the adult counterpart of mGSCs, because significant differences, for example much higher derivation efficiency and cytoplasmic location of OCT4, have been detected [121]. Remarkably, cultivated SSCs could contribute to both spermatogenesis in the testes and embryogenesis in blastocysts, which had been regarded as mutually exclusive.
Subsequently, a number of independently established pluripotent stem cell lines from neonatal and adult testes have been described. Interestingly, not only were different culture procedures used, but also the properties of the cells obtained were slightly different from each other [122, 123]. Seandel et al. [111] basically followed the mGSCs derivation but used a G-protein-coupled receptor, GPR125, to purify spermatogonial progenitor cells from adult testis. In this study, ESC-like cells capable of forming teratomas and chimeras could be derived from GPR125-positive GSC cultures. However, comparison of global gene expression in the ESC-like cells with ESCs and GSCs revealed different expression profiles, although GPR125 and Oct4 were expressed in all three cell types. In contrast, Izadyar et al. [112] identified OCT4+/C-KIT+GSCs subpopulations with pluripotent characteristics by use of an Oct4-GFP reporter. These cells formed GSCs-like colonies, expressed germ cell markers and pluripotency markers, and had androgenetic imprinting. Unexpectedly, they contributed neither to spermatogenesis nor teratoma formation, but could be incorporated into chimeras. Similarly, Huang et al. [113] found a cell population with high alkaline phosphatase activity and a pluripotent phenotype in GSC cultures. Ko et al. [114, 124] have reported a highly reproducible method for establishing ESC-like cells from clonal adult GSCs. These cells were quite similar to mGSCs, but the genomic imprinting pattern was completely androgenetic.
Molecular insights into the pluripotency in spermatogenic cells
What are the crucial mechanisms that confer pluripotency on SSCs? One candidate is the enhanced self-renewal of SSCs. Transgenic mice overexpressing GDNF frequently developed malignant testicular germ cell tumors (TGCTs) by 1 year of age [125, 126]. However, the TGCTs were classified as seminomas consisting of germ cells only; they were not teratomas. Similarly, downstream signaling of GDNF is not likely to be related to pluripotency reacquisition in SSCs. The PI3K/AKT pathway, which is activated by GDNF stimulation, is of crucial importance in self-renewal of GSCs: a PI3K inhibitor prevented GSC self-renewal whereas forced-activation of AKT supported it [127]. In addition, PTEN is involved in Nanog repression in GSCs [128]. However, although the PI3K/AKT pathway promotes the dedifferentiation of PGCs into EGCs, this is not observed for GSCs. Another study showed that constitutive activation of the Ras/cyclin D2 pathway enables GSCs to grow in a GDNF-independent manner [129]. When transplanted into testes, GSCs with activated Ras/cyclin D2 formed TGCTs, but similar to GDNF-overexpressing mice, the tumors were seminomas not teratomas.
Another key finding was demethylation of paternal imprinted genes in mGSCs [106], which suggested the potential involvement of DNA methylation in SSC reprogramming. To address this question, genetically manipulated GSCs having Dnmt1 knockdown, Dnmt3a/3b knockout, or Dnmt3L overexpression were constructed [130]. The Dnmt1 knockdown resulted in apoptosis, whereas Dnmt3a/3b-knockout or Dnmt3L-overexpressing GSCs grew normally. GSCs with Dnmt3a/3b-knockout or Dnmt3L-overexpression developed DNA hypomethylation or hypermethylation of repetitive sequences, respectively. However, neither expressed pluripotency marker genes nor formed teratomas after transplantation. Thus, DNA demethylation in mGSCs might not be a resultant event but rather than a causal event of SSC dedifferentiation. Anyway, because the study did not assess mGSC derivation from GSCs with modified Dnmt expression, further rigorous studies should be conducted to verify the implication of DNA methylation for SSC reprogramming.
The microenvironments in SSC cultures have also been examined. ESC-like cells appeared only when GSCs were plated at 5 to 20-fold lower densities than in regular expansion cultures [114]. The ordinary culture conditions may not support SSC dedifferentiation because of cell density or passage frequency. In other work, an immortalized cell line of CD34-positive mouse testicular stromal cells was established to support the derivation and proliferation of GSCs, and the maintenance of maGSCs and derivation of EGCs [131]. However, the effect on maGSC derivation was not validated. Subsequently, by use of a cytokine antibody array, potential niche factors were screened by using the supernatant of testicular stromal cell cultures [113]. Eventually, insulin-like growth factor 1 (IGF1) was identified as a factor secreted by Leydig cells that enhanced the formation of pluripotent cell colonies. The IGF1 effect increased in a dose-dependent manner, whereas neutralizing antibody against the IGF1 receptor blocked colony formation.
Once established, GSCs never convert to mGSCs. However, mGSCs did appear in cultures of p53-deficient GSCs [106]. In addition to activity as a tumor suppressor, p53 binds the Nanog promoter and represses Nanog expression in ESCs [132] and GSCs [128]. The Nanog promoter also contains an OCT4/SOX2-binding site close to the p53-binding sites, and the Nanog promoter DNA is hypermethylated in a spermatogenic cell-specific manner [89, 133, 134] but becomes demethylated by reprogramming [114, 135]. These observations suggest that Nanog reactivation may be a key event in SSC reprogramming. However, forced expression of Nanog and/or Sox2 was not sufficient to confer pluripotency on GSCs [109]. Thus, it is likely that SSCs also have potent cell-autonomous pluripotency-inhibitor programs. At present, no significant changes have been observed in pluripotency-associated gene expression or overall histone modifications between wild-type and p53-deficient GSCs [109]. Comprehensive and thorough studies of p53-deficient GSCs will be needed to reveal the molecular mechanisms underlying GSC-mGSC reprogramming.
Pluripotent stem cells derived from human adult testis
In primates, including humans, no counterparts of mouse GSCs have yet been established, except for the short-term culture of gonocytes and spermatogonial cells [136, 137]. Two groups have reported isolation of GSC-like cells from human testes, but characterization was not sufficient to definitively identify them as GSCs [138, 139]. However, as in mice, pluripotent stem cells have been derived from the adult human testis. Human testicular cells were cultivated with GDNF or LIF for 4 days, and then spermatogonia were purified on the basis of α6 integrin expression [140]. The α6 integrin-positive cells were further cultured with LIF under conditions optimized for mouse ESCs. The cells formed multilayered colonies, expressed pluripotency genes, and led to the development of teratomas when injected subcutaneously into immunodeficient mice. Their derivation and proliferation predominantly depends on LIF, not GDNF or FGF2. This is in contrast with human ESC cultures, which require FGF2 and TGFβ signals. Although their global gene expression profile has been reported to be similar to that of human ESCs, other researchers have claimed that the cells were more similar to fibroblasts [141].
However, others have demonstrated that pluripotent stem cells can be derived and maintained under human ESC culture conditions with FGF2 alone or in combination with TGFβ [142–144]. Interestingly, testis-derived pluripotent stem cells are negative for SSEA1, similarly to human ESCs, indicating that they are clearly distinguishable from PGC-derived EGCs, which express the SSEA1 antigen [144]. Furthermore, DNA methylation analysis has shown that the H19 locus becomes demethylated, in contrast to its androgenetic hypermethylation in spermatogenic cells [143, 144]. Likewise, in the OCT4 and NANOG regulatory regions DNA is demethylated but methylation levels are still much higher than in ESCs. Considering the lower expression of pluripotency-associated genes [144], the reprogramming of these adult testis pluripotent cells may not be sufficient to establish cells comparable with human ESCs.
Female germ cell-derived stem cells
The potential existence of oogonial stem cells
Unlike male GSCs, whether any stem cells contribute to oogenesis in the mammalian ovary remains controversial. It has long been believed that mammalian females lose the ability to make new germ cells during fetal life. However, advances in the past decade have challenged this notion of the function of stem cells in the postnatal ovary. In pioneering work in 2004 mitotically active germ cells were found in the ovarian surface epithelium of juvenile and adult mice, [145, 146]. This finding raised the possibility that the mammalian postnatal ovary might harbor oocyte stem cells. This possibility was further supported by evidence that premeiotic germ cells could be identified in ovaries of aged mice [147]. The premeiotic germ cells remained quiescent in aged ovaries but could be reactivated to produce oocytes when transplanted into young mouse ovaries. In addition, subsequent studies showed that bone marrow and peripheral blood could be sources of putative oogonial stem cells [148–150]. On the basis of these studies it was proposed that the putative stem cells were to replenish the oocyte pool in the ovary. However, the possibility of oocyte neogenesis is still under debate, because others have disputed oocyte renewal in the postnatal ovary [151–153].
In addition to the indirect in-vivo evidence above, attempts have been made to isolate oogonial stem cells in in-vitro studies. In an attempt to generate female GSCs (FGSCs), ovarian cells from neonatal mice were cultured by the same procedures and conditions used to generate male GSCs. Eventually, proliferative colony-forming cells were isolated but these were thecal stem cells [154]. However, an attempt to purify VASA-positive cells from neonatal and adult mouse ovaries by cell sorting [155] produced VASA-positive cells that, when cultured with LIF, GDNF, EGF, and FGF2, with serum, displayed BrdU uptake. These cells were capable of growing for months and enduring cryopreservation, and were therefore termed FGSCs. The FGSCs had a unique gene-expression profile: positive for Oct4, Blimp1, Stella, Vasa, and Dazl but negative for Nanog, Sox2, c-Kit, Figlα, Scp1/2/3, and Zp3. Direct evidence that these cells are oogonial stem cells was provided by transplantation into a mouse ovary, where the FGSCs contributed to oogenesis and to offspring. Subsequently, FGSCs were found to be highly enriched by sorting FRAGILIS-positive cells instead of VASA-positive cells [156] and transgenic mice were produced by manipulating these FGSCs [157]. Forced-expression or knockdown of particular genes in FGSCs was performed and the cells were then transplanted into mouse ovaries. Pups carrying the transgenes were obtained. Independently, in a report using Oct4-GFP transgenic mice, GFP-expressing cells were collected from the ovarian surface epithelium and cultivated as ovarian GSCs [158]. Unlike those in former studies, they had an epithelial morphology and expressed OCT4, NANOG, and C-KIT, but did not form teratomas. However, they spontaneously gave rise to oocyte-like cells during culture. These studies have indicated that the mammalian ovary contains a stem cell population that can produce oocytes. Nevertheless, whether the FGSCs are truly derived from oogonial stem cells of the postnatal ovary is currently unknown.
Studies of human oogenesis have demonstrated that putative stem cells exist in the surface epithelium of the human ovary also. When the epithelium was isolated from the ovaries of postmenopausal women, the putative stem cells, which are small and spherical, could be expanded, and produced early oocyte-like cells spontaneously in vitro [159]. The putative stem cells expressed OCT4, SOX2, NANOG, and C-KIT and did not form teratomas on injection into immunodeficient mice. A further study also showed the presence of oocyte-like cells and the development of blastocyst-like structures, possibly by parthenogenetic activation [160]. Very recently, by using cell-sorting techniques, proliferative human FGSC lines were established from ovaries of reproductive-age women [161]. Similar to mouse FGSCs obtained by the same procedure, the human FGSCs were small and round in shape, and expressed germ cells marker genes but not oocyte markers. The human FGSCs spontaneously produced oocytes in culture. Moreover, human FGSCs could form follicles after injection into human ovarian biopsies, which were xenografted into immunodeficient mice. Thus, these findings suggested that oogonial stem cells are present in the human ovary, and in mouse.
Pluripotent stem cells from ovarian cell culture
Similar to testicular germ cells, it is likely that the ovary contains cells that are capable of pluripotency. ESC-like colonies have been reported in mouse ovarian cell cultures cultured under the same conditions as mouse ESCs [162]. The ESC-like cells grew stably, expressed pluripotency markers, and produced teratomas. However, the genomic imprinting pattern was different not only from that of ordinary ESCs but also from that of parthenogenetic ESCs: H19 and Gtl2 were highly methylated whereas Peg3 and Snrpn were completely demethylated. This unusual genomic imprinting was observed in testis-derived ESC-like cells also [106], suggesting that pluripotent stem cells derived from postnatal gonads may undergo unique epigenetic reprogramming that differs from that of embryonic cells. It is not clear whether the origin of the ESC-like cells is attributable to ovarian germ cells such as oocytes and oogonial stem cells.
Female germ cell-derived stem cells are a new topic of investigation in germ cell research and much controversy surrounds the putative origins of these cells. Despite great interest in the possibility of female GSCs, our current insight is quite limited and more extensive efforts should be made to clarify our understanding of ovarian stem-like cells.
Induced pluripotent stem cells: an analogy with PGC specification?
iPSC technology exploited a new way of making pluripotent stem cells from cells of somatic origin by gene transduction into the target cells. As a result, the kinds of physiological state the iPSCs reflect are unclear. However, PGCs emerge from the differentiating epiblast by inhibiting the somatic differentiation program and reactivating pluripotency-associated genes [86]. The process is similar to iPSC reprogramming, in which cells repress somatic genes and subsequently upregulate pluripotency markers [163]. Thus, PGC specification could be regarded as an “in vivo reprogramming event” to re-acquire pluripotency.
On the basis of this idea, the reprogramming activity of crucial PGC specification factors was assessed by use of iPSC technology. As candidate genes, Blimp1, Prdm14, and Prmt5 were co-transfected into mouse embryonic fibroblasts carrying a Nanog-GFP reporter in combination with known iPSCs-reprogramming factors (Oct4, Sox2, Klf4, and c-Myc) [164]. Among the combinations tested, two sets of genes (Blimp1/Prdm14/c-Myc and Prmt5/Oct4/Klf4) successfully generated Nanog-GFP-positive cells. However, the reprogramming activity of Blimp1/Prdm14/c-Myc was quite limited: the frequency of occurrence of Nanog-GFP positive cells was low and could not be increased owing to growth arrest. In contrast, Nanog-GFP-positive cells from Prmt5/Oct4/Klf4 transfected cells had characteristics similar to those of iPSCs and contributed to chimeric mice. The iPSCs retained the parental genomic imprinting, suggesting that Prmt5/Oct4/Klf4 did induce iPSCs but not EGCs, even though PRMT5 is essential for PGC specification. Furthermore, the contribution of PRMT5 to iPSC generation was confirmed by a knockdown, which reduced the emergence of Nanog-GFP positive colonies.
To date, this is the only report to investigate the potential association between PGC specification and iPSC generation. However, in both PGCs and GSCs, p53 acts to suppress the induction of pluripotency [68, 106], as is the case in iPSC generation: suppression of the p53/p21 pathway increases the efficiency of iPSC generation [165–168]. Thus, repression of p53 may be a universal event required for non-pluripotent cells to acquire pluripotency. Additionally, protection against cellular senescence is a key step in iPSC generation [169]. Because GSCs and FGSCs with high telomerase activity are not susceptible to cellular senescence, this may be one factor contributing to the higher pluripotency potential of germ cells. Generation of iPSCs from germ cells such as GSCs could provide a means of investigating the stability of genomic imprinting and the pluripotency-inhibitory mechanisms innate in germ cells.
Conclusion
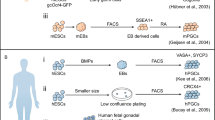
To ultimately produce offspring, germ cells must undergo tightly regulated development. Faithful execution of developmental programs is crucial for determining their fate as gametogenetic cells and ensuring that a pluripotent fate is blocked. Nevertheless, germ cells retain the potential to re-acquire pluripotency, as is evident by the derivation, in vitro, of pluripotent stem cells from germ cells at different stages (Fig. 1). The generation of such cells demonstrates that germ cells have a cell-intrinsic property enabling them to be pluripotent. In this regard, the reprogramming of germ cells contrasts greatly with the generation of iPSCs, which requires multiple gene transductions. Thus, the spontaneous induction of pluripotency in germ cells is a significant counterpart of iPSCs generation that will enable us to understand the cellular reprogramming machinery. Forthcoming techniques will enable us to determine global molecular profiles using a very small number of cells or, ultimately, a single cell. Using comprehensive and high-throughput techniques, comparative studies of germ cell reprogramming and iPSC generation will accelerate not only our understanding of cellular reprogramming but also the development of translational applications for areas such as regenerative medicine.
Overview of mammalian germline development and resulting pluripotent stem cell line cultures. Blue arrows indicate sequential and unidirectional development of the germline in vivo. During early embryogenesis, the fertilized egg is totipotent and develops into the pluripotent inner cell mass (ICM) and then into the pluripotent epiblast of the egg cylinder. Sexually bipotential primordial germ cells (PGCs) develop from differentiating epiblast cells, and migrate into genital ridges. Thereafter, PGCs follow sex-specific programs toward spermatogenesis or oogenesis. Lineage-committed germline stem cells can be maintained in vitro with gametogenetic potential (germ cells in culture). On the other hand, pluripotent stem cells maintained in vitro are also derived from several sources of the germline (pluripotent stem cells in culture). The pluripotent ICM of the blastocyst and epiblast of the egg cylinder give rise to embryonic stem cells (ESCs) and epiblast stem cells (EpiSCs) respectively (yellow column). After incorporation into germ cell development, primordial germ cells (PGCs) and spermatogenic cells can dedifferentiate to produce a variety of pluripotent stem cells including embryonic germ cells (EGCs), multipotent germline stem cells (mGSCs), and multipotent adult germline stem cells (maGSCs) (green and blue columns). Despite their unknown origin, ESCs-like cells also occur in ovarian cell culture (orange column). In contrast, induced pluripotent stem cells (iPSCs) are generated from non-germline somatic cells by artificial gene transduction (grey column)
References
Takahashi K. Direct reprogramming 101. Dev Growth Differ. 2010;52(3):319–33.
Takahashi K, Yamanaka S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell. 2006;126(4):663–76.
Takahashi K, et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell. 2007;131(5):861–72.
Yu J, et al. Induced pluripotent stem cell lines derived from human somatic cells. Science. 2007;318(5858):1917–20.
Ohinata Y, et al. A signaling principle for the specification of the germ cell lineage in mice. Cell. 2009;137(3):571–84.
Ohinata Y, et al. Blimp1 is a critical determinant of the germ cell lineage in mice. Nature. 2005;436(7048):207–13.
Yamaji M, et al. Critical function of Prdm14 for the establishment of the germ cell lineage in mice. Nat Genet. 2008;40(8):1016–22.
Kurimoto K, et al. Complex genome-wide transcription dynamics orchestrated by Blimp1 for the specification of the germ cell lineage in mice. Genes Dev. 2008;22(12):1617–35.
West JA, et al. A role for Lin28 in primordial germ-cell development and germ-cell malignancy. Nature. 2009;460(7257):909–13.
Saitou M, Barton SC, Surani MA. A molecular programme for the specification of germ cell fate in mice. Nature. 2002;418(6895):293–300.
Yamaji M, et al. Functional reconstruction of NANOS3 expression in the germ cell lineage by a novel transgenic reporter reveals distinct subcellular localizations of NANOS3. Reproduction. 2010;139(2):381–93.
Gu Y, et al. Steel factor controls primordial germ cell survival and motility from the time of their specification in the allantois, and provides a continuous niche throughout their migration. Development. 2009;136(8):1295–303.
Gu Y, et al. Membrane-bound steel factor maintains a high local concentration for mouse primordial germ cell motility, and defines the region of their migration. PLoS ONE. 2011;6(10):e25984.
Molyneaux KA, et al. The chemokine SDF1/CXCL12 and its receptor CXCR4 regulate mouse germ cell migration and survival. Development. 2003;130(18):4279–86.
Ara T, et al. Impaired colonization of the gonads by primordial germ cells in mice lacking a chemokine, stromal cell-derived factor-1 (SDF-1). Proc Natl Acad Sci USA. 2003;100(9):5319–23.
Kehler J, et al. Oct4 is required for primordial germ cell survival. EMBO Rep. 2004;5(11):1078–83.
Yamaguchi S, et al. Conditional knockdown of Nanog induces apoptotic cell death in mouse migrating primordial germ cells. Development. 2009;136(23):4011–20.
Tsuda M, et al. Conserved role of nanos proteins in germ cell development. Science. 2003;301(5637):1239–41.
Seki Y, et al. Cellular dynamics associated with the genome-wide epigenetic reprogramming in migrating primordial germ cells in mice. Development. 2007;134(14):2627–38.
Seki Y, et al. Extensive and orderly reprogramming of genome-wide chromatin modifications associated with specification and early development of germ cells in mice. Dev Biol. 2005;278(2):440–58.
Sugimoto M, Abe K. X chromosome reactivation initiates in nascent primordial germ cells in mice. PLoS Genet. 2007;3(7):e116.
Ancelin K, et al. Blimp1 associates with Prmt5 and directs histone arginine methylation in mouse germ cells. Nat Cell Biol. 2006;8(6):623–30.
Sabour D, et al. Identification of genes specific to mouse primordial germ cells through dynamic global gene expression. Hum Mol Genet. 2011;20(1):115–25.
Arnaud P. Genomic imprinting in germ cells: imprints are under control. Reproduction. 2010;140(3):411–23.
Bowles J, Koopman P. Sex determination in mammalian germ cells: extrinsic versus intrinsic factors. Reproduction. 2010;139(6):943–58.
Ohinata Y, et al. A comprehensive, non-invasive visualization of primordial germ cell development in mice by the Prdm1-mVenus and Dpp a3-ECFP double transgenic reporter. Reproduction. 2008;136(4):503–14.
Toyooka Y, et al. Expression and intracellular localization of mouse Vasa-homologue protein during germ cell development. Mech Dev. 2000;93(1–2):139–49.
Xu EY, Moore FL, Pera RA. A gene family required for human germ cell development evolved from an ancient meiotic gene conserved in metazoans. Proc Natl Acad Sci USA. 2001;98(13):7414–9.
Lee J, et al. Erasing genomic imprinting memory in mouse clone embryos produced from day 11.5 primordial germ cells. Development. 2002;129(8):1807–17.
Bowles J, et al. Retinoid signaling determines germ cell fate in mice. Science. 2006;312(5773):596–600.
Lucifero D, et al. Gene-specific timing and epigenetic memory in oocyte imprinting. Hum Mol Genet. 2004;13(8):839–49.
Hiura H, et al. Oocyte growth-dependent progression of maternal imprinting in mice. Genes Cells. 2006;11(4):353–61.
Li JY, et al. Timing of establishment of paternal methylation imprints in the mouse. Genomics. 2004;84(6):952–60.
Sakai Y, et al. Co-expression of de novo DNA methyltransferases Dnmt3a2 and Dnmt3L in gonocytes of mouse embryos. Gene Expr Patterns. 2004;5(2):231–7.
Jiang LI, Nadeau JH. 129/Sv mice—a model system for studying germ cell biology and testicular cancer. Mamm Genome. 2001;12(2):89–94.
Stevens LC, Little CC. Spontaneous testicular teratomas in an inbred strain of mice. Proc Natl Acad Sci USA. 1954;40(11):1080–7.
Stevens LC, Hummel KP. A description of spontaneous congenital testicular teratomas in strain 129 mice. J Natl Cancer Inst. 1957;18(5):719–47.
Stevens LC. A new inbred subline of mice (129-terSv) with a high incidence of spontaneous congenital testicular teratomas. J Natl Cancer Inst. 1973;50(1):235–42.
Noguchi T, Noguchi M. A recessive mutation (ter) causing germ cell deficiency and a high incidence of congenital testicular teratomas in 129/Sv-ter mice. J Natl Cancer Inst. 1985;75(2):385–92.
Stevens LC. Origin of testicular teratomas from primordial germ cells in mice. J Natl Cancer Inst. 1967;38(4):549–52.
Lam MY, Nadeau JH. Genetic control of susceptibility to spontaneous testicular germ cell tumors in mice. APMIS. 2003;111(1):184–90. (discussion 191).
Youngren KK, et al. The Ter mutation in the dead end gene causes germ cell loss and testicular germ cell tumours. Nature. 2005;435(7040):360–4.
Kedde M, et al. RNA-binding protein Dnd1 inhibits microRNA access to target mRNA. Cell. 2007;131(7):1273–86.
Raymond CS, et al. Dmrt1, a gene related to worm and fly sexual regulators, is required for mammalian testis differentiation. Genes Dev. 2000;14(20):2587–95.
Matson CK, et al. DMRT1 prevents female reprogramming in the postnatal mammalian testis. Nature. 2011;476(7358):101–4.
Krentz AD, et al. The DM domain protein DMRT1 is a dose-sensitive regulator of fetal germ cell proliferation and pluripotency. Proc Natl Acad Sci USA. 2009;106(52):22323–8.
Cook MS, et al. BAX-mediated cell death affects early germ cell loss and incidence of testicular teratomas in Dnd1(Ter/Ter) mice. Dev Biol. 2009;328(2):377–83.
Gill ME, et al. Licensing of gametogenesis, dependent on RNA binding protein DAZL, as a gateway to sexual differentiation of fetal germ cells. Proc Natl Acad Sci USA. 2011;108(18):7443–8.
Lin Y, Page DC. Dazl deficiency leads to embryonic arrest of germ cell development in XY C57BL/6 mice. Dev Biol. 2005;288(2):309–16.
Matsui Y, et al. Effect of steel factor and leukaemia inhibitory factor on murine primordial germ cells in culture. Nature. 1991;353(6346):750–2.
De Felici M, Dolci S. Leukemia inhibitory factor sustains the survival of mouse primordial germ cells cultured on TM4 feeder layers. Dev Biol. 1991;147(1):281–4.
Resnick JL, et al. Long-term proliferation of mouse primordial germ cells in culture. Nature. 1992;359(6395):550–1.
Matsui Y, Zsebo K, Hogan BL. Derivation of pluripotential embryonic stem cells from murine primordial germ cells in culture. Cell. 1992;70(5):841–7.
Durcova-Hills G, et al. The role of exogenous fibroblast growth factor-2 on the reprogramming of primordial germ cells into pluripotent stem cells. Stem Cells. 2006;24(6):1441–9.
Stewart CL, Gadi I, Bhatt H. Stem cells from primordial germ cells can reenter the germ line. Dev Biol. 1994;161(2):626–8.
Labosky PA, Barlow DP, Hogan BL. Mouse embryonic germ (EG) cell lines: transmission through the germline and differences in the methylation imprint of insulin-like growth factor 2 receptor (Igf2r) gene compared with embryonic stem (ES) cell lines. Development. 1994;120(11):3197–204.
Tada T, et al. Epigenotype switching of imprintable loci in embryonic germ cells. Dev Genes Evol. 1998;207(8):551–61.
Tada M, et al. Embryonic germ cells induce epigenetic reprogramming of somatic nucleus in hybrid cells. EMBO J. 1997;16(21):6510–20.
Sharova LV, et al. Global gene expression profiling reveals similarities and differences among mouse pluripotent stem cells of different origins and strains. Dev Biol. 2007;307(2):446–59.
Koshimizu U, et al. Functional requirement of gp130-mediated signaling for growth and survival of mouse primordial germ cells in vitro and derivation of embryonic germ (EG) cells. Development. 1996;122(4):1235–42.
Durcova-Hills G, et al. Reprogramming primordial germ cells into pluripotent stem cells. PLoS ONE. 2008;3(10):e3531.
Brazil DP, Yang ZZ, Hemmings BA. Advances in protein kinase B signalling: AKTion on multiple fronts. Trends Biochem Sci. 2004;29(5):233–42.
Takahashi K, Murakami M, Yamanaka S. Role of the phosphoinositide 3-kinase pathway in mouse embryonic stem (ES) cells. Biochem Soc Trans. 2005;33(Pt 6):1522–5.
Cantley LC, Neel BG. New insights into tumor suppression: PTEN suppresses tumor formation by restraining the phosphoinositide 3-kinase/AKT pathway. Proc Natl Acad Sci USA. 1999;96(8):4240–5.
Suzuki A, et al. High cancer susceptibility and embryonic lethality associated with mutation of the PTEN tumor suppressor gene in mice. Curr Biol. 1998;8(21):1169–78.
Kimura T, et al. Conditional loss of PTEN leads to testicular teratoma and enhances embryonic germ cell production. Development. 2003;130(8):1691–700.
Moe-Behrens GH, et al. Akt/PTEN signaling mediates estrogen-dependent proliferation of primordial germ cells in vitro. Mol Endocrinol. 2003;17(12):2630–8.
Kimura T, et al. AKT signaling promotes derivation of embryonic germ cells from primordial germ cells. Development. 2008;135(5):869–79.
Miki T, Yasuda SY, Kahn M. Wnt/beta-catenin signaling in embryonic stem cell self-renewal and somatic cell reprogramming. Stem Cell Rev. 2011;7(4):836–46.
Kimura T, et al. The stabilization of beta-catenin leads to impaired primordial germ cell development via aberrant cell cycle progression. Dev Biol. 2006;300(2):545–53.
Leitch HG, et al. Embryonic germ cells from mice and rats exhibit properties consistent with a generic pluripotent ground state. Development. 2010;137(14):2279–87.
Jansson M, et al. Arginine methylation regulates the p53 response. Nat Cell Biol. 2008;10(12):1431–9.
Scoumanne A, Zhang J, Chen X. PRMT5 is required for cell-cycle progression and p53 tumor suppressor function. Nucleic Acids Res. 2009;37(15):4965–76.
Morita-Fujimura Y, Tokitake Y, Matsui Y. Heterogeneity of mouse primordial germ cells reflecting the distinct status of their differentiation, proliferation and apoptosis can be classified by the expression of cell surface proteins integrin alpha6 and c-Kit. Dev Growth Differ. 2009;51(6):567–83.
Matsui Y, Tokitake Y. Primordial germ cells contain subpopulations that have greater ability to develop into pluripotential stem cells. Dev Growth Differ. 2009;51(7):657–67.
Thomson JA, et al. Embryonic stem cell lines derived from human blastocysts. Science. 1998;282(5391):1145–7.
Shamblott MJ, et al. Derivation of pluripotent stem cells from cultured human primordial germ cells. Proc Natl Acad Sci USA. 1998;95(23):13726–31.
Turnpenny L, et al. Evaluating human embryonic germ cells: concord and conflict as pluripotent stem cells. Stem Cells. 2006;24(2):212–20.
Turnpenny L, et al. Derivation of human embryonic germ cells: an alternative source of pluripotent stem cells. Stem Cells. 2003;21(5):598–609.
Liu S, et al. Human embryonic germ cells isolation from early stages of post-implantation embryos. Cell Tissue Res. 2004;318(3):525–31.
Li F, et al. Leukemia inhibitory factor-expressing human embryonic lung fibroblasts as feeder cells for human embryonic germ cells. Cells Tissues Organs. 2007;186(4):221–8.
Labosky PA, Barlow DP, Hogan BL. Embryonic germ cell lines and their derivation from mouse primordial germ cells. Ciba Found Symp. 1994;182:157–68. Discussion 168–178.
Okamura D, et al. Requirement of Oct3/4 function for germ cell specification. Dev Biol. 2008;317(2):576–84.
Mise N, et al. Differences and similarities in the developmental status of embryo-derived stem cells and primordial germ cells revealed by global expression profiling. Genes Cells. 2008;13(8):863–77.
Niwa H, et al. A parallel circuit of LIF signalling pathways maintains pluripotency of mouse ES cells. Nature. 2009;460(7251):118–22.
Yabuta Y, et al. Gene expression dynamics during germline specification in mice identified by quantitative single-cell gene expression profiling. Biol Reprod. 2006;75(5):705–16.
Takahashi K, Mitsui K, Yamanaka S. Role of ERas in promoting tumour-like properties in mouse embryonic stem cells. Nature. 2003;423(6939):541–5.
Kanatsu-Shinohara M, et al. Long-term proliferation in culture and germline transmission of mouse male germline stem cells. Biol Reprod. 2003;69(2):612–6.
Imamura M, et al. Transcriptional repression and DNA hypermethylation of a small set of ES cell marker genes in male germline stem cells. BMC Dev Biol. 2006;6:34.
Fujino RS, et al. Capillary morphogenesis gene (CMG)-1 is among the genes differentially expressed in mouse male germ line stem cells and embryonic stem cells. Mol Reprod Dev. 2006;73(8):955–66.
Kurosaki H, et al. A comparison study in the proteomic signatures of multipotent germline stem cells, embryonic stem cells, and germline stem cells. Biochem Biophys Res Commun. 2007;353(2):259–67.
Kubota H, Avarbock MR, Brinster RL. Growth factors essential for self-renewal and expansion of mouse spermatogonial stem cells. Proc Natl Acad Sci USA. 2004;101(47):16489–94.
Meng X, et al. Regulation of cell fate decision of undifferentiated spermatogonia by GDNF. Science. 2000;287(5457):1489–93.
He Z, et al. Gfra1 silencing in mouse spermatogonial stem cells results in their differentiation via the inactivation of RET tyrosine kinase. Biol Reprod. 2007;77(4):723–33.
Ogawa T, et al. Derivation and morphological characterization of mouse spermatogonial stem cell lines. Arch Histol Cytol. 2004;67(4):297–306.
Ryu BY, et al. Conservation of spermatogonial stem cell self-renewal signaling between mouse and rat. Proc Natl Acad Sci USA. 2005;102(40):14302–7.
Hamra FK, et al. Self renewal, expansion, and transfection of rat spermatogonial stem cells in culture. Proc Natl Acad Sci USA. 2005;102(48):17430–5.
Kanatsu-Shinohara M, et al. Long-term culture of male germline stem cells from hamster testes. Biol Reprod. 2008;78(4):611–7.
Kubota H, et al. Glial cell line-derived neurotrophic factor and endothelial cells promote self-renewal of rabbit germ cells with spermatogonial stem cell properties. FASEB J. 2011;25(8):2604–14.
Kanatsu-Shinohara M, et al. Homologous recombination in rat germline stem cells. Biol Reprod. 2011;85(1):208–17.
Kanatsu-Shinohara M, et al. Production of knockout mice by random or targeted mutagenesis in spermatogonial stem cells. Proc Natl Acad Sci USA. 2006;103(21):8018–23.
Kanatsu-Shinohara M, Toyokuni S, Shinohara T. Genetic selection of mouse male germline stem cells in vitro: offspring from single stem cells. Biol Reprod. 2005;72(1):236–40.
Takehashi M, et al. Adenovirus-mediated gene delivery into mouse spermatogonial stem cells. Proc Natl Acad Sci USA. 2007;104(8):2596–601.
Ivics Z, et al. Sleeping Beauty transposon mutagenesis in rat spermatogonial stem cells. Nat Protoc. 2011;6(10):1521–35.
Ivics Z, et al. Sleeping Beauty transposon mutagenesis of the rat genome in spermatogonial stem cells. Methods. 2011;53(4):356–65.
Kanatsu-Shinohara M, et al. Generation of pluripotent stem cells from neonatal mouse testis. Cell. 2004;119(7):1001–12.
Kanatsu-Shinohara M, et al. Genetic and epigenetic properties of mouse male germline stem cells during long-term culture. Development. 2005;132(18):4155–63.
Lee J, et al. Heritable imprinting defect caused by epigenetic abnormalities in mouse spermatogonial stem cells. Biol Reprod. 2009;80(3):518–27.
Kanatsu-Shinohara M, et al. Pluripotency of a single spermatogonial stem cell in mice. Biol Reprod. 2008;78(4):681–7.
Guan K, et al. Pluripotency of spermatogonial stem cells from adult mouse testis. Nature. 2006;440(7088):1199–203.
Seandel M, et al. Generation of functional multipotent adult stem cells from GPR125+ germline progenitors. Nature. 2007;449(7160):346–50.
Izadyar F, et al. Generation of multipotent cell lines from a distinct population of male germ line stem cells. Reproduction. 2008;135(6):771–84.
Huang YH, et al. Pluripotency of mouse spermatogonial stem cells maintained by IGF-1-dependent pathway. FASEB J. 2009;23(7):2076–87.
Ko K, et al. Induction of pluripotency in adult unipotent germline stem cells. Cell Stem Cell. 2009;5(1):87–96.
Zechner U, et al. Comparative methylation profiles and telomerase biology of mouse multipotent adult germline stem cells and embryonic stem cells. Mol Hum Reprod. 2009;15(6):345–53.
Khromov T, et al. Global and gene-specific histone modification profiles of mouse multipotent adult germline stem cells. Mol Hum Reprod. 2011;17(3):166–74.
Zovoilis A, et al. Multipotent adult germline stem cells and embryonic stem cells have similar microRNA profiles. Mol Hum Reprod. 2008;14(9):521–9.
Zovoilis A, et al. Embryonic stem cell-related miRNAs are involved in differentiation of pluripotent cells originating from the germ line. Mol Hum Reprod. 2010;16(11):793–803.
Meyer S, et al. Pluripotent embryonic stem cells and multipotent adult germline stem cells reveal similar transcriptomes including pluripotency-related genes. Mol Hum Reprod. 2010;16(11):846–55.
Dihazi H, et al. Multipotent adult germline stem cells and embryonic stem cells: comparative proteomic approach. J Proteome Res. 2009;8(12):5497–510.
Kanatsu-Shinohara M, Shinohara T. The germ of pluripotency. Nat Biotechnol. 2006;24(6):663–4.
de Rooij DG, Mizrak SC. Deriving multipotent stem cells from mouse spermatogonial stem cells: a new tool for developmental and clinical research. Development. 2008;135(13):2207–13.
Geijsen N, Hochedlinger K. gPS navigates germ cells to pluripotency. Cell Stem Cell. 2009;5(1):3–4.
Ko K et al (2011) Autologous pluripotent stem cells generated from adult mouse testicular biopsy. Stem Cell Rev
Meng X, et al. Promotion of seminomatous tumors by targeted overexpression of glial cell line-derived neurotrophic factor in mouse testis. Cancer Res. 2001;61(8):3267–71.
Sariola H, Meng X. GDNF-induced seminomatous tumours in mouse–an experimental model for human seminomas? APMIS. 2003;111(1):192–6. Discussion 196.
Lee J, et al. Akt mediates self-renewal division of mouse spermatogonial stem cells. Development. 2007;134(10):1853–9.
Kuijk EW, et al. PTEN and TRP53 independently suppress Nanog expression in spermatogonial stem cells. Stem Cells Dev. 2010;19(7):979–88.
Lee J, et al. Genetic reconstruction of mouse spermatogonial stem cell self-renewal in vitro by Ras-cyclin D2 activation. Cell Stem Cell. 2009;5(1):76–86.
Takashima S, et al. Abnormal DNA methyltransferase expression in mouse germline stem cells results in spermatogenic defects. Biol Reprod. 2009;81(1):155–64.
Kim J, et al. CD34+ testicular stromal cells support long-term expansion of embryonic and adult stem and progenitor cells. Stem Cells. 2008;26(10):2516–22.
Lin T, et al. p53 induces differentiation of mouse embryonic stem cells by suppressing Nanog expression. Nat Cell Biol. 2005;7(2):165–71.
Farthing CR, et al. Global mapping of DNA methylation in mouse promoters reveals epigenetic reprogramming of pluripotency genes. PLoS Genet. 2008;4(6):e1000116.
Western PS, et al. Male fetal germ cell differentiation involves complex repression of the regulatory network controlling pluripotency. FASEB J. 2010;24(8):3026–35.
Jung YH, et al. Glial cell line-derived neurotrophic factor alters the growth characteristics and genomic imprinting of mouse multipotent adult germline stem cells. Exp Cell Res. 2010;316(5):747–61.
He Z, et al. Isolation, characterization, and culture of human spermatogonia. Biol Reprod. 2010;82(2):363–72.
Tu J, et al. Stem cell factor affects fate determination of human gonocytes in vitro. Reproduction. 2007;134(6):757–65.
Lee DR, et al. Isolation of male germ stem cell-like cells from testicular tissue of non-obstructive azoospermic patients and differentiation into haploid male germ cells in vitro. Hum Reprod. 2006;21(2):471–6.
Sadri-Ardekani H, et al. Propagation of human spermatogonial stem cells in vitro. JAMA. 2009;302(19):2127–34.
Conrad S, et al. Generation of pluripotent stem cells from adult human testis. Nature. 2008;456(7220):344–9.
Ko K, et al. Human adult germline stem cells in question. Nature. 2010;465(7301):E1. discussion E3.
Golestaneh N, et al. Pluripotent stem cells derived from adult human testes. Stem Cells Dev. 2009;18(8):1115–26.
Kossack N, et al. Isolation and characterization of pluripotent human spermatogonial stem cell-derived cells. Stem Cells. 2009;27(1):138–49.
Mizrak SC, et al. Embryonic stem cell-like cells derived from adult human testis. Hum Reprod. 2010;25(1):158–67.
Johnson J, et al. Germline stem cells and follicular renewal in the postnatal mammalian ovary. Nature. 2004;428(6979):145–50.
Kerr JB, et al. Quantification of healthy follicles in the neonatal and adult mouse ovary: evidence for maintenance of primordial follicle supply. Reproduction. 2006;132(1):95–109.
Niikura Y, Niikura T, Tilly JL. Aged mouse ovaries possess rare premeiotic germ cells that can generate oocytes following transplantation into a young host environment. Aging (Albany NY). 2009;1(12):971–8.
Johnson J, et al. Oocyte generation in adult mammalian ovaries by putative germ cells in bone marrow and peripheral blood. Cell. 2005;122(2):303–15.
Lee HJ, et al. Bone marrow transplantation generates immature oocytes and rescues long-term fertility in a preclinical mouse model of chemotherapy-induced premature ovarian failure. J Clin Oncol. 2007;25(22):3198–204.
Tilly JL, Niikura Y, Rueda BR. The current status of evidence for and against postnatal oogenesis in mammals: a case of ovarian optimism versus pessimism? Biol Reprod. 2009;80(1):2–12.
Eggan K, et al. Ovulated oocytes in adult mice derive from non-circulating germ cells. Nature. 2006;441(7097):1109–14.
Begum S, Papaioannou VE, Gosden RG. The oocyte population is not renewed in transplanted or irradiated adult ovaries. Hum Reprod. 2008;23(10):2326–30.
Kerr JB et al (2012) The primordial follicle reserve is not renewed after chemical or gamma-irradiation mediated depletion. Reproduction
Honda A, et al. Isolation, characterization, and in vitro and in vivo differentiation of putative thecal stem cells. Proc Natl Acad Sci USA. 2007;104(30):12389–94.
Zou K, et al. Production of offspring from a germline stem cell line derived from neonatal ovaries. Nat Cell Biol. 2009;11(5):631–6.
Zou K, et al. Improved efficiency of female germline stem cell purification using fragilis-based magnetic bead sorting. Stem Cells Dev. 2011;20(12):2197–204.
Zhang Y, et al. Production of transgenic mice by random recombination of targeted genes in female germline stem cells. J Mol Cell Biol. 2011;3(2):132–41.
Pacchiarotti J, et al. Differentiation potential of germ line stem cells derived from the postnatal mouse ovary. Differentiation. 2010;79(3):159–70.
Virant-Klun I, et al. Putative stem cells with an embryonic character isolated from the ovarian surface epithelium of women with no naturally present follicles and oocytes. Differentiation. 2008;76(8):843–56.
Virant-Klun I, et al. Parthenogenetic embryo-like structures in the human ovarian surface epithelium cell culture in postmenopausal women with no naturally present follicles and oocytes. Stem Cells Dev. 2009;18(1):137–49.
White YA, et al. Oocyte formation by mitotically active germ cells purified from ovaries of reproductive-age women. Nat Med. 2012;18(3):413–21.
Gong SP, et al. Embryonic stem cell-like cells established by culture of adult ovarian cells in mice. Fertil Sterili. 2010;93(8):2594-601, 2601 e1-9.
Stadtfeld M, et al. Defining molecular cornerstones during fibroblast to iPS cell reprogramming in mouse. Cell Stem Cell. 2008;2(3):230–40.
Nagamatsu G, et al. A germ cell-specific gene, Prmt5, works in somatic cell reprogramming. J Biol Chem. 2011;286(12):10641–8.
Zhao Y, et al. Two supporting factors greatly improve the efficiency of human iPSC generation. Cell Stem Cell. 2008;3(5):475–9.
Hong H, et al. Suppression of induced pluripotent stem cell generation by the p53–p21 pathway. Nature. 2009;460(7259):1132–5.
Marion RM, et al. A p53-mediated DNA damage response limits reprogramming to ensure iPS cell genomic integrity. Nature. 2009;460(7259):1149–53.
Kawamura T, et al. Linking the p53 tumour suppressor pathway to somatic cell reprogramming. Nature. 2009;460(7259):1140–4.
Utikal J, et al. Immortalization eliminates a roadblock during cellular reprogramming into iPS cells. Nature. 2009;460(7259):1145–8.
Acknowledgments
This work was supported by grants from: the Ministry of Education, Culture, Sports, Science, and Technology of Japan (MEXT); the Ministry of Health, Labor, and Welfare; the Japan Society for the Promotion of Science (JSPS); the National Institute of Biomedical Innovation; the Strategic Research Program for Brain Sciences and the Leading Project for Realization of Regenerative Medicine, MEXT; the Project for Realization of Regenerative Medicine, MEXT; the Funding Program for World-leading Innovative R&D in Science and Technology (FIRST); and a Keio University Grant-in-Aid for the Encouragement of Young Medical Scientists.
Author information
Authors and Affiliations
Corresponding author
About this article
Cite this article
Imamura, M., Lin, Z.YC. & Okano, H. Cell-intrinsic reprogramming capability: gain or loss of pluripotency in germ cells. Reprod Med Biol 12, 1–14 (2013). https://doi.org/10.1007/s12522-012-0131-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12522-012-0131-z