Abstract
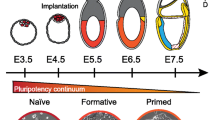
Pluripotency is defined by the ability of a cell to differentiate to the derivatives of all the three embryonic germ layers: ectoderm, mesoderm and endoderm. Pluripotent cells can be captured via the archetypal derivation of embryonic stem cells or via somatic cell reprogramming. Somatic cells are induced to acquire a pluripotent stem cell (iPSC) state through the forced expression of key transcription factors, and in the mouse these cells can fulfil the strictest of all developmental assays for pluripotent cells by generating completely iPSC-derived embryos and mice. However, it is not known whether there are additional classes of pluripotent cells, or what the spectrum of reprogrammed phenotypes encompasses. Here we explore alternative outcomes of somatic reprogramming by fully characterizing reprogrammed cells independent of preconceived definitions of iPSC states. We demonstrate that by maintaining elevated reprogramming factor expression levels, mouse embryonic fibroblasts go through unique epigenetic modifications to arrive at a stable, Nanog-positive, alternative pluripotent state. In doing so, we prove that the pluripotent spectrum can encompass multiple, unique cell states.
Similar content being viewed by others
Main
Somatic cells that have lost their pluripotent properties through the acquisition of differentiation-associated epigenetic marks can be driven to acquire an induced pluripotent cell (iPSC) state by the forced expression of key transcription factors1. iPSCs can fulfil the strictest of murine developmental assays, tetraploid embryo complementation2, to form to all the cells of the embryo proper and the resulting adult animal3. During the reprogramming of somatic cells, it is visibly apparent that there exists a spectrum of distinct cell types. The embryonic stem cell (ESC)-like iPSCs capable of generating healthy mice represent just one end of this spectrum. Many studies describe the successful derivation of iPSCs, however, relatively few studies address the fate of cells that do not reprogram to an ESC-like state. It has been reported that somatic cells expressing the four reprogramming factors1 can stabilize at a Nanog-negative cell state that morphologically resembles ESCs, yet failed to fully acquire an ESC-like expression profile4,5,6. ‘Partially reprogrammed cell’ has become a term to describe any cell that fails to reprogram to an ESC-like state. However, it is likely that a range of cell types exist, whose stable phenotypes and associated epigenetic profiles are different from ESCs.
For somatic cells to acquire an ESC-like state they require extensive genome-wide remodelling, with epigenetic mechanisms regulating cell state transitions throughout the entire reprogramming process. Incomplete remodelling of the somatic epigenome is associated with transgene-dependent cells5 and a functional memory of somatic cell origin7,8. The modulation of epigenetic regulators such as DNA dioxygenases9, histone deacetylases10, H3K36 demethylase (Jhdm1b)11, H3K27 demethylase (Utx)12 and H3K9 demethylases6 greatly influences the efficiency and kinetics of reprogramming towards a ESC-like iPSC state. In particular, vitamin C has been reported to facilitate the transition of cells from a ‘partially reprogrammed state’ to an ESC-like state6,13. In addition to chromatin remodelling, the expression level of reprogramming transcription factors directs cell state. A narrow window of Oct4 expression is required to maintain the ESC state, whereby a twofold perturbation of expression induces cells to transition to a non-ESC state14. During reprograming there are two potential sources of Oct4: the transgene, whose expression has to be high at the beginning, and the endogenous gene, which is reactivated during the process of reprogramming. Towards the end of reprogramming the total expression of these two Oct4 sources has to stabilize within the narrow window required by the ESC-like state. Elevated expression of the four reprogramming factors has the potential to direct cell identity to a non-ESC-like state. In agreement, significant changes in global gene expression are observed when the reprogramming factors are shut down15,16,17.
Somatic-cell-derived epigenetic marks and the conceivable permutations of reprogramming factor expression levels present a unique opportunity to generate novel cell types. Thus, in an experimental approach unbiased by pre-conceptions of what constitutes a reprogrammed cell we characterize the diversity of cell states that arise during somatic cell reprogramming. We define a Nanog-positive cell state (F-class cells) that is stable, occurs frequently, is dependent on high reprogramming factor expression, in which cells do not form typical ESC-like colonies, exhibits advantageous cell culture properties, and yet demonstrates pluripotency.
Reprogramming diversity
To extensively characterize the diversity of cell states arising from embryonic fibroblasts, we initiated reprogramming with the doxycycline-inducible piggyBac transposon system18. Colonies of proliferative cells were picked in a randomized manner, impartial of gene expression and morphological appearance, establishing clonally-derived cell lines (Fig. 1a). Notably, the transgene-expressing cell lines segregated into two distinct cohorts (Fig. 1b), which we had initially classified by morphological appearance as compact colony forming cells (C-class) and fuzzy colony forming cells (F-class). For all 28 cell lines established, the reprogramming genes Oct4 (also known as Pou5f1), Sox2, Klf4 and c-Myc were expressed many fold above ESC levels (Extended Data Fig. 1a), with each clonal cell line exhibiting substantial global gene expression differences when compared to ESCs (Fig. 1b). The majority of genes (67%) that were expressed above ESC levels were also expressed above (>twofold) parental fibroblast levels (Extended Data Fig. 1b, c and Supplementary Information 1), suggesting that these genes were induced upon reprogramming rather than representing a fibroblast memory. 2,959 differentially expressed genes (P < 0.01; false discovery rate (FDR) < 0.05) separated F-class and C-class cells, (Extended Data Fig. 1d, Supplementary Information 2) with the F-class cell lines being particularly intriguing as they expressed Nanog and endogenous Oct4 at ESC levels (Extended Data Fig. 1e, 2a, b), yet did not possess an ESC-like morphology (Fig. 1b). The fuzzy appearance of F-class colonies and low intercellular adhesion was reminiscent of E-cadherin-null ESCs19,20,21 and could be attributed to diminished E-cadherin expression (Extended Data Fig. 1e). When mapped to the previously established PluriNet22 (Extended Data Fig. 1f), F-class cells exhibited significantly reduced expression of many PluriNet genes (Dnmt3b, Zfp42 and Tdgf1), yet they expressed many genes at ESC levels such as Sall4, endogenous Oct4 and Nanog (Supplementary Information 3). In addition, the F-class cells expressed transcription factors associated with lineage commitment including the homeobox protein En2, the helix–loop–helix factor Ngn3 and homeobox protein Nkx2.3.
a, Fibroblasts were transfected with Yamanaka factors in four separate piggyBac transposons (pB) and clonal lines were derived. b, Unsupervised hierarchical clustering and sample distance matrix (Pearson correlation) of gene expression at day 16. Phase contrast images representative of F-class (clone 1) and C-class (clone 23) iPS cell lines. Scale bars, 200 μm.
We compared the F-class cells to another well-characterized pluripotent stem cell population, epiblast stem cells (EpiSCs), and found that the F-class cells are transcriptionally distinct (Extended Data Fig. 2c, d). Furthermore, F-class cells could not be generated or maintained in EpiSC media (Extended Data Fig. 2e).
An alternative stem-cell state
Differentially expressed genes (P < 0.01; FDR < 0.05) between ESCs and F-class cells are enriched with genes involved in cell adhesion and the extracellular matrix (Fig. 2a, b), which probably contributes to the morphological appearance of F-class cells. Forced expression of Cdh1 induced some cells to acquire an ESC-like morphology; however, it was insufficient for most cells in culture (Extended Data Fig. 3a, b), suggesting that Cdh1 was not the only factor required. Furthermore, elevated Cdh1 expression did not induce the expression of Esrrb and Dppa5, genes that are downregulated in Cdh1-null ESCs20 (Extended Data Fig. 3a). The F-class gene expression profile remained unchanged upon prolonged culture, with cells maintaining a stable transcriptome and no convergence towards an ESC-like state (Fig. 2c). Independent sub-lines exhibited low variance in gene expression, further demonstrating the stable self-renewal of the F-class cell state (Extended Data Fig. 3c). The absence of interspersed Dppa4-expressing cells suggested that cells do not spontaneously progress to an ESC-like state at a detectable rate (Extended Data Fig. 3d). F-class cells possessed a normal karyotype (Extended Data Fig. 3e) and could be expanded exponentially beyond 40 passages. The cells remained in a transgene-dependent state (Extended Data Fig. 3f), whereby turning off transgene expression induced population-wide differentiation within 48 h, demonstrating that cells had not transformed. The self-renewal of F-class cells was independent of LIF or JAK signalling (Extended Data Fig. 4a, b); furthermore, F-class cells can be generated in media supplemented with JAK inhibitor (Extended Data Fig. 4c–f). F-class cells rapidly proliferated to the extent that, when mixed with ESCs, an initial 1% F-class cells became the dominant cell type (>50%) within three passages (Extended Data Fig. 4g). Stable gene expression, rapid proliferation (Extended Data Fig. 4h) and low intercellular adhesion (Extended Data Fig. 4i) confer F-class cells with highly desirable properties for stirred suspension culture.
a, Differentially expressed genes between ESC-like state n = 4 and F-class state n = 6 (Two-tailed Welchs t-test P < 0.01; FDR < 0.05). b, Gene ontology term analysis of differentially expressed genes. c, Two-way scatter plot comparisons of global gene expression (Illumina BeadArray), blue lines represent fourfold differential threshold. d, Histological analysis of teratomas containing differentiated tissues of all three germ layers. Arrowheads denote ciliated epithelia. Scale bars, 100 μm.
Teratomas initiated by pluripotent cells (ESC, ESC-like iPSC and F-class cells) contained well-differentiated (non-dividing) and less differentiated dividing compartments. The teratomas from the F-class cells were indistinguishable from those derived from ESCs, each consisting of complex differentiated tissues representing all three germ layers (Fig. 2d). In vitro, removal of doxycycline in serum-free media initiated efficient neural differentiation of F-class cells, generating multiple neuronal subtypes (Extended Data Fig. 5a–c). Differentiation in serum-based media generated cells representative of the mesoderm (α-SMA+) and endoderm (FoxA2+) lineages (Extended Data Fig. 5d). We then assessed the embryonic developmental potential of F-class cells and found that they do not contribute to the development of chimaeras, nor do they incorporate into blastocysts after injection into the perivitelline space of eight-cell stage embryos (data not shown). In summary, we describe a novel cell state that is distinct from ESCs yet passes criteria used to functionally identify the pluripotent potential of human ESC and iPSC lines, as by the teratoma-forming assay.
Requirement of transgene expression
To determine the influence of transgene expression levels on the establishment of F-class and ESC-like states, we examined three different reprogramming systems: three-factor (3F), which excludes c-Myc; low-expressing four-factor (4F; Col1a1 transgenic secondary system23); and high-expressing four factor, 1B secondary system18 (Extended Data Fig. 6a, b). High-expressing 4F fibroblasts underwent population-wide proliferation and generated distinct colonies within 5 days, which stabilized at a state morphologically and transcriptionally resembling F-class cells (Extended Data Fig. 6c–e). In contrast, 3F and low-expressing 4F fibroblasts sporadically (<0.1%) gave rise to colonies from day 10 onwards, stabilizing at a state that morphologically and transcriptionally resembled ESCs (Extended Data Fig. 6c–e). During low-expressing 4F reprogramming, no morphologically overt F-class cells were observed at any time point, nor were F-class identifier genes expressed at elevated levels (Extended Data Fig. 6f). These observations suggest a model whereby low-transgene-expressing cells do not generate an F-class cell state (Extended Data Fig. 6g). We found that high four-factor expression can also reprogram adult tail skin fibroblasts to the F-class state (Extended Data Fig. 7a–c).
During somatic cell reprogramming retroviral transgenes become silenced and it is thought that this helps stabilize a fully reprogrammed ESC-like state24. Since F-class cells require maintained transgene expression, we questioned whether a retroviral transgene system could give rise to F-class cells. We initially observed rapidly dividing cells possessing an F-class morphology (days 8–16 post-transduction); however, we did not observe these cells beyond day 30. Retrovirus-delivered transgene expression (green fluorescent protein, GFP) was attenuated during transposon-mediated reprogramming to an F-class state and within established F-class cells (Extended Data Fig. 7d, e). We propose that silencing of the retroviral transgenes is not compatible with the F-class cells’ requirement for high transgene expression.
To examine the continued requirement of all four reprogramming factors, F-class cells were generated where three factors are constitutively expressed and the fourth factor is doxycycline-inducible. Doxycycline was removed at day 30 and in all four cases turning off the fourth factor induced a rapid loss of proliferation and a flattening of cell morphology (data not shown). Thus, all four reprogramming factors are needed to maintain the F-class state. The consistent inability to obtain F-class cells with 3F reprogramming indicates that elevated c-Myc expression is necessary. We used the TetO-Myc F-class cells, and found that upon doxycycline removal there was a downregulation of genes involved in growth factor activity and positive regulation of transcription (Extended Data Fig. 8a–d), in accordance with a reduced proliferation. Although cells did not transition to an ESC-like state, a number of ESC-associated genes were upregulated (Extended Data Fig. 8c, Supplementary Information 4), supporting the theory that reprogramming factor expression actively suppresses the final acquisition of an ESC-like state15.
Cell-state transitions
We questioned whether re-expressing the reprogramming factors at high levels in the ESC-like state would induce a transition to the F-class state. Reprogramming factor expression was re-activated in the iPSC line 1B18 and cells were transferred to media conditions that are conducive to F-class cells but not ESC-like cells: JAK inhibition in the absence of LIF and feeders (Extended Data Fig. 8e, f). Within 48 h, colonies of cells arose that morphologically resembled F-class cells. These cells maintained expression of some ESC-associated genes (Lin28 and Dnmt3B) yet diminished others such as Dppa5, Dnmt3l and Cdh1 (Extended Data Fig. 8g). Notably, cells upregulated genes expressed by F-class cells, suggesting that elevated reprogramming transgene expression can induce an F-class-like state, with the starting cell type (ESCs or MEFs) leaving a signature on the F-class cell state.
Next, we investigated whether established F-class cells can be induced to transition to an ESC-like state. Exposure to the DNA methyltransferase inhibitor 5-aza-deoxycytidine (Aza) was toxic at active concentrations (>0.05 μM), while vitamin C (ascorbic acid) supplementation and 2i media failed to induce an ESC-like morphology (Fig. 3a and Extended Data Fig. 9a). In contrast, inhibition of histone deacetylases (HDAC) induced F-class cells to acquire an ESC-like morphology (Fig. 3a) and transcriptional profile (Fig. 3b, Extended Data Fig. 9a). To determine whether HDAC inhibition (HDACi) selects for a sub-population of cells, we exposed twelve newly established subclones to HDACi and found that they acquired an ESC-like morphology and consistently upregulate ESC-like markers (Extended Data Fig. 9b). Furthermore, when single cells were treated with HDACi, every subsequent colony possessed elevated expression of ESC-associated genes (Fig. 3c). Direct observation of cells by time-lapse microscopy revealed that HDACi treatment decreased cell proliferation (Extended Data Fig. 9c) with no evidence of cell death (Extended Data Fig. 9d). HDACi-mediated acquisition of an ESC-like state was rapid with transcriptionally silent genes upregulated to ESC expression levels within 72 h (Extended Data Fig. 10a–c, Supplementary Information 5). During the first 24 h of HDACi treatment genes with chromatin and cell-division related ontology were upregulated (Extended Data Fig. 10d). The upregulation of chromatin-related factors possibly facilitated the transcriptional activation of further ESC-associated genes. Following HDACi treatment, cells could be maintained as transgene-independent ESC-like cells capable of contributing to chimaeras and the germ line (Fig. 3d, e). This was not possible before HDACi treatment.
a, Day 30 F-class cells (clone 1) treated for ten days. Scale bar, 100 μm. b, Principal component analysis of 32 cell-state identifier genes (determined by quantitative PCR with reverse transcription, qRT–PCR). C3 and C5 represent C-class clones, F1 and F2 represents F-class clones, maintained in different media. c, F-class cells (clone 1) clonally HDACi-treated (10 nM TSA). Scale bar, 250 μm. Each point represents an individual cell colony profiled by qRT–PCR (n = 10 biological replicates, 3 technical replicates per colony). Ngn3 is also known as Neurog3. d, Chimaeric contribution of HDACi treated F-class cells aggregated with eight-cell stage embryo, visualized by LacZ activity, representative of four embryos. e, Genital ridge dissected from chimaeric embryo (n = 1). GFP represents HDACi-treated clone2 F-class cells and Oct4 represents the germ cells. Scale bar, 100 μm.
Epigenetic forces contribute to F-class state
To identify the epigenetic landmarks associated with the establishment of the F-class cell state, we exploited a high-resolution genome-wide resource that profiles fibroblast reprogramming at the molecular level to both F-class and ESC-like states25. Doxycycline-induced high-level reprogramming factor expression directs 1B secondary fibroblast reprogramming to an F-class transcriptional state (Extended Data Fig. 10e)25,26,27,28. Comparison of primary F-class cell lines and ESC-like cell lines identified 86 genes that exhibited substantial (>fivefold) differential expression (Fig. 4a). For these genes we assessed the status of three major chromatin marks; the activating histone H3K4 trimethylation (H3K4me3), the suppressing histone H3K27 trimethylation (H3K27me3)25 and CpG methylation27 (Supplementary Information 6). Transcriptional activity of 72 of the 86 genes (79%) correlated (Pearson correlation coefficient |r| > 0.5) with at least one epigenetic mark (Fig. 4b). The upregulation of F-class state identifiers, such as Nkx2-3 and Insm1 (Fig. 4c, d), was associated with an active loss of H3K27me3 during the reprogramming process, fitting the model that the F-class state is not an intermediate reprogrammed state but a distinct cell state achieved through active epigenetic changes. Further substantiating this is the observation that genes associated with the ESC-like state (Gbx2, Lefty1, Cldn6) acquired hypermethylation at their genomic loci (Fig. 4e), which is uncharacteristic of the ESC-like state. We further validated a subset of differentially methylated regions (DMRs) within primary F-class cells (Fig. 4f). In summary, fibroblast reprogramming to the F-class state is governed by multiple epigenetic marks, whereby active epigenetic modifications direct cell identity away from both fibroblast and ESC-like state, and repressive epigenetic marks are inherited from the parental cell type (fibroblasts).
a, Differentially expressed genes in primary lines (Welch t-test P < 0.01, FDR < 0.01). Black data points depict >fivefold difference in 1B secondary reprogramming system of F-class and ESC-like cells25. Lines depict fivefold threshold. b, Euler diagram depicting genes (black points in a) whose transcriptional activity corresponds with differential epigenetic marks in 1B secondary reprogramming system derived F-class cells25 (n = 1). c, Unsupervised clustering heat map of H3K27me3 marks identified in b. d, Transcription and histone modifications at the genomic locus of F-class identifier Insm1. e, Unsupervised clustering heat map of differentially methylated regions identified in b. f, Differentially methylated regions observed in the secondary reprogramming system confirmed in primary F-class cells.
Discussion
In this study, we observed that reprogramming somatic cells, in the presence of elevated reprogramming factor expression, could stabilize at a Nanog-positive fuzzy colony forming (F-class) state. Previous studies may have overlooked this state as the F-class cells highly express Nanog without completing one of the early reprogramming events, the mesenchymal to epithelial transition16,29. Chan and colleagues previously described a human reprogrammed cell type (type II cells) that is Nanog-positive and persists in a state that represents an intermediate stage of somatic cell reprogramming30. In contrast to the human type II cells, the murine F-class cells do not morphologically resemble ESCs, nor do they transcriptionally or epigenetically represent an intermediate cell state that reprogramming cells transit through as they acquire ESC-like state. Two central observations support the notion that the F-class cell state is not representative of an intermediate state. First, F-class cells upregulate a cohort of genes that were not observed during reprogramming without c-Myc (3F) or with low-level four-factor (Oct4, Klf4, Sox2 and c-Myc) expression. Second, the expression of these genes in F-class cells is associated with the loss of repressive epigenetic marks (H3K27me3 and/or DNA methylation) that are typically present in the parental fibroblasts and the ESC-like state. The loss of these repressive marks suggests that, during sustained reprogramming factor expression, cell identity is diverted away from the molecular pathway that leads to an ESC-like state (Fig. 5). This is further supported by the observation that ESC-associated genes (Lefty1, Cldn6, Gbx2) actually acquire inhibitory DNA methylation in the F-class state. To our knowledge, this is the first report to identify dynamic epigenetic changes that actively propel reprogramming cells towards an alternative pluripotent cell state. In conclusion, the F-class cells represent an acquired state and not an intermediate state that all reprogramming cells transition through on the way to an ESC-like state.
HDACi denotes histone deacetylation inhibition, 4F denotes the four Yamanaka factors, 3F denotes the four Yamanaka factors minus c-Myc.
We propose that the F-class cell state is stably maintained as a consequence of high reprogramming factor expression and multiple epigenetic determinants. Through elevated expression of the four reprogramming factors we showed that F-class cells could be generated from both fibroblasts and ESC-like iPSCs. Notably, the cell type of origin leaves distinct signatures on the resultant F-class cells, as an imprint of their respective origin.
The ability to reprogram cells to novel cell states, such as the F-class state, can be harnessed to create a variety of artificial cells that possess desirable properties for regenerative medicine and drug discovery, such as the ability for scalable expansion in bioreactors and reproducible differentiation. ESCs are themselves an artificial in vitro cell state, captured during a brief developmental window and require specific culture conditions for their maintenance. The F-class cell state can be considered to be a distant pluripotent relative of the ESC state. The frequency at which F-class cells arise in transposon-based reprogramming, in combination with their advantageous properties, presents the opportunity to study and utilize a novel pluripotent cell type in biology, medical research and future medicine.
Methods
Cell culture
All cell lines were established in-house with full pathogen testing performed and maintained in a mycoplasma-free facility. Mouse embryonic fibroblasts (MEF) were isolated as previously described18. 15.5 days post coitum ROSA26-rtTA-IRES-GFP embryos (JAX 005572)31 were decapitated, eviscerated, dissociated with 0.25% trypsin, 0.1% EDTA and plated in DMEM, 10% FBS, penicillin–streptomycin and GlutaMAX. MEFs were reprogrammed within 4 passages of derivation. Tail-tip fibroblasts (TTFs) were obtained from 8-week-old mice. Tail-tips were mechanically dissociated with 0.25% trypsin and 1,000 U ml−1 collagenase (Type XI-S).
A standardised transfection protocol was established to electroporate fibroblasts (Neon, Invitrogen) with piggyBac transposons encoding the four reprogramming factors. In brief, 2 × 106 MEFs were electroporated with 4 μg of plasmid (0.5 μg PBase transposon and 3.5 μg factors), using optimized parameters (2 pulses, 1,200 V). Electroporated fibroblasts were plated in serum-based mouse ESC media32 supplemented with 1.5 μg ml−1 doxycycline on gelatinized (0.1%) plates, at a density of 1.5 × 104 cells per cm2. Cells were fed every three days with doxycycline-containing media (1.5 μg ml−1). Colonies were clonally picked and expanded in a 96-well format. Unless stated otherwise, clonal cell lines were maintained in mouse ESC media supplemented with 1.5 μg ml−1 doxycycline. ROSA26-rtTA-IRES-GFP ESCs were used as control cells. 2i media conditions represent serum-free media consisting of DMEM:F12 supplemented with 15% Knockout serum replacement (Gibco), 3 μM CHIR99021 (GSK3β inhibitor) and 1 μM PD0325901 (MEK inhibitor) as previously described33.
Transgene independent ESC-like iPSCs were obtained from F-class cells by exposure to sodium butyrate (0.25 mM) for seven days (plus doxycycline). Cells were then maintained in 2i media in the absence of sodium butyrate (plus doxycycline) for five days and then doxycycline was removed. Cells were furthermore maintained in either serum-based ESC media or 2i media.
EpiSCs were maintained in X-vivo base media (Lonza) supplemented with 10 mM β-mercaptoethanol (Sigma), 1 mM MEM-NEAA (Invitrogen), 2 mM GlutaMAX (Invitrogen), 20 ng ml−1 Activin A (R&D Systems), and 20 ng ml−1 basic fibroblast growth factor (R&D Systems). EpiSCs were passaged every 3–4 days as single cells in TrypLE (Invitrogen) and plated on wells pre-coated with Matrigel.
For retrovirus mediated reprogramming, retroviral packaging of pMX constructs and subsequent transduction of cells was performed as previously described34.
Stirred suspension culture
Adherent cells were trypsinized and seeded into spinner flasks at 2 × 104 cells per ml. 30-ml culture volumes were maintained at constant stirring speed of 85 r.p.m. at 37 °C and 10% CO2. Every three days cell numbers were quantified and suspension cultures reset to 2 × 104 cells per ml. One-half of the culture medium was replaced every two days.
In vitro neural differentiation
Cells were plated on geltrex (1:100 PBS dilution) coated plates at 5,000 cells per cm2. 24 h after plating cells, ESC media was changed to serum-free media that consisted of DMEM:F12 supplemented with N2 (Gibco), B27 (Gibco), and 4 μg ml−1 insulin. Doxycycline was removed by washing cells three times with PBS to remove all traces of doxycycline. Differentiation media was changed every three days.
Diploid aggregation generation of chimaeras
Cells were maintained for two passages in 2i media with cell clumps of ∼8–15 cells collected from gelatinized dishes by gentle trypsinization. For diploid chimaeras, 2.5 d.p.c. Hsd:ICR(CD-1) or C57BL/6 embryos were aggregated with in-vitro-derived cell clumps and cultured overnight at 37 °C in 5% CO2 in KSOM medium33. All embryos were transferred into pseudopregnant recipient ICR females 24 h later. For LacZ detection, pregnant dams were fed doxycycline food and water (0.2 mg ml−1 doxycycline; 5% sucrose in water) 24 h before dissection to activate β-geo expression in iPSC-derived cells. All mouse procedures were performed in accordance with Toronto Centre for Phenogenomics animal care committee.
LacZ staining
As described in ref. 18 cells and embryos were fixed with 0.25% glutaraldehyde, rinsed in wash buffer (2 mM MgCl2, 0.01% sodium deoxycholate, and 0.02% Nonidet-P40 in PBS) and stained overnight (∼16 h) in LacZ staining solution: 20 mM MgCl2, 5 mM K3Fe(CN)6, 5 mM K4Fe(CN)6 and 1 mg ml−1 X-gal in PBS. Embryos were embedded in paraffin, sectioned and counterstained with neutral red.
Teratoma formation
Cells were trypsinized and suspended in DMEM:Matrigel mix (1:1) with 100 μl of 1 × 106 cells injected subcutaneously into the dorsal flanks of nude mice (CByJ.Cg-Foxn1nu/J females, 6 weeks of age) anaesthetized with isoflurane. 4–6 weeks after injection, teratomas were dissected and fixed overnight in 4% formalin. Tissue was embedded in paraffin, sectioned and stained with haematoxylin and eosin.
Immunostaining and flow cytometry
Cells were washed once with PBS, fixed in 4% PFA for 15 min at room temperature and permeabilized with 0.1% Triton X-100 in PBS for 10 min. Primary antibody was added overnight at 4 °C: anti-α-SMA (C6198, Sigma), anti-Nanog (RCAB0002P, Reprocell), anti-DPPA4 (AF3730, R&D Systems), anti-FoxA2 (ab40874, Abcam) anti-SSEA1 (MAB4301, Millipore), anti-Sox2 (MAB2018, R&D Systems), anti-Oct3/4 (611203, BD), anti-GFP (6673, Abcam), anti-βIII-tubulin (TUJ1, Covance), anti-tyrosine hydroxylase (AB152, Millipore), anti-VGAT (131103, SYSY), anti-VGLUT1 (135302, SYSY). Secondary antibody (Jackson immune research cy3 IgG, 1:200; Alexa488 IgG or IgM, 1:400; Alexa594 IgG, 1:400) was added for 1 h at room temperature. Cell nuclei were stained with Hoechst 33342 (5 μg ml−1) for 15 min.
Flow cytometry
Cells were trypsinized and fixed in 4% PFA for 15 min at room temperature. Cells were washed and then stained with 0.1% Triton X-100 in PBS (2% FBS), incubated with primary antibody (Nanog 1:200) for 1 h on ice, washed twice in PBS (2% FBS), incubated with secondary antibody for 30 min on ice, washed twice and resuspended in PBS with 2% FBS for analysis on a FACS-Calibur. Cells were gated on the basis of forward scatter and side scatter.
Cell viability assay
Cell samples were trypsinized, resuspended in Annexin V buffer (10 mM HEPES, 140 mM NaCl, and 2.5 mM CaCl2, pH 7.4) and then incubated with Sytox AADvanced for 5 min and Annexin V for 10 min. Cellular fragments and debris were excluded from analysis using forward-scatter and side-scatter selection.
G-band karyotyping
G-banding was performed on actively dividing cells at the TCAG facility (Toronto, Canada). Cells were incubated with 0.2 μg ml−1 colcemid for 2 h at 37 °C and dissociated with 0.25% trypsin-EDTA. After pipetting a single-cell suspension was resuspended in pre-warmed (37 °C) 75 mM KCl for 15 min. Cells were then fixed with methanol:glacial acetic acid (1:3) and dropped onto glass slides. The slides containing cells were stained in Giemsa solution for 3 min, with 20 metaphases counted and scored for karyotyping.
Quantitative RT–PCR
Cells for RNA preparation were passaged on gelatin-coated plates. Total RNA was extracted from cells using a RNeasy kit (Qiagen). 1 μg of DNase treated RNA was used as template to generate cDNA by QuantiTect reverse transcription kit (Qiagen). For quantitative RT–PCR we used LuminoCt SYBR Green qPCR ReadyMix (Sigma) with JANUS automated liquid handling robot (PerkinElmer) loading the 384-well plates for RT-qPCR. 384 plates were run on a CFX384 (Bio-Rad) with an annealing temperature of 58 °C for all primers. Primer pairs were all assessed for efficiency and melt curves performed. All PCR reactions were performed in triplicate. Primer sequences are listed in Supplementary Information 7.
Illumina BeadChip
Total RNA was assessed for quality and quantity on a Bioanalyzer and global gene expression profiling performed with the Illumina microarray. Purified and labelled RNA was hybridized to MouseRef-8 v2 expression BeadChips (Illumina) according to the manufacturer’s instructions. Bead intensities were mapped to gene information using BeadStudio 3.2 (Illumina). Background correction was performed using the Affymetrix Robust Multi-array Analysis and data log2-scaled with gene expression quantile normalized in the lumi package of Bioconductor.
Bisulphite sequencing
Bisulphite conversion was performed on genomic DNA sample (1 μg) using the EpiTect Bisulfite Kit (QIAGEN). Bisulphite-treated genomic DNA was amplified by EpiTaq HS (Takara) using previously published bisulphite-specific primers35 and novel primers (Supplementary Information 4), with a PCR protocol consisting of an initial 1 min denaturation step followed by 35 cycles of 95 °C for 15 s, 55 °C for 30 s and 72 °C for 30 s. The resultant PCR amplicons were cloned in to pGemTeasy and sequenced at the Centre for Applied Genomics (Toronto, Canada).
Statistical analysis
Unless otherwise stated, all data presented are representative of at least three independent experiments. Hierarchical clustering, principal component analysis and gene distance matrices were performed with Multiexperiment Viewer. Statistical analysis was performed with either Prism (Graphpad) or Multiexperiment viewer (http://www.tm4.org/index.html). Gene ontology term analysis was performed with DAVID (Database for Annotation, Visualization and Integrated Discovery, http://david.abcc.ncifcrf.gov). Gene network association analysis was performed with GeneMANIA (http://www.genemania.org). Genes in the network analysis were chosen based on their membership of the PluriNet network22 and statistically significant differential expression between F-class samples and ESC samples. Differential expression was assessed using the limma package, P values were adjusted using the Benjamini–Hochberg method and significance cut-off set at 0.05.
Accession codes
Primary accessions
Gene Expression Omnibus
Data deposits
Microarray data have been deposited on Stemformatics (http://www.stemformatics.org) and in the Gene Expression Omnibus database under accession number GSE49940.
Change history
10 December 2014
A minor addition was made to the Acknowledgements in the HTML and PDF versions.
References
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006)
Nagy, A., Rossant, J., Nagy, R., Abramow-Newerly, W. & Roder, J. C. Derivation of completely cell culture-derived mice from early-passage embryonic stem cells. Proc. Natl Acad. Sci. USA 90, 8424–8428 (1993)
Zhao, X.-Y. et al. iPS cells produce viable mice through tetraploid complementation. Nature 461, 86–90 (2009)
Fussner, E. et al. Constitutive heterochromatin reorganization during somatic cell reprogramming. EMBO J. 30, 1778–1789 (2011)
Sridharan, R. et al. Role of the murine reprogramming factors in the induction of pluripotency. Cell 136, 364–377 (2009)
Chen, J. et al. H3K9 methylation is a barrier during somatic cell reprogramming into iPSCs. Nature Genet. 45, 34–42 (2013)
Kim, K. et al. Epigenetic memory in induced pluripotent stem cells. Nature 467, 285–290 (2010)
Polo, J. M. et al. Cell type of origin influences the molecular and functional properties of mouse induced pluripotent stem cells. Nature Biotechnol. 28, 848–855 (2010)
Hu, X. et al. Tet and TDG mediate DNA demethylation essential for mesenchymal-to-epithelial transition in somatic cell reprogramming. Cell Stem Cell 14, 512–522 (2014)
Mali, P. et al. Butyrate greatly enhances derivation of human induced pluripotent stem cells by promoting epigenetic remodeling and the expression of pluripotency-associated genes. Stem Cells 28, 713–720 (2010)
Wang, T. et al. The histone demethylases Jhdm1a/1b enhance somatic cell reprogramming in a vitamin-C-dependent manner. Cell Stem Cell 9, 575–587 (2011)
Mansour, A. A. et al. The H3K27 demethylase Utx regulates somatic and germ cell epigenetic reprogramming. Nature 488, 409–413 (2012)
Esteban, M. A. et al. Vitamin C enhances the generation of mouse and human induced pluripotent stem cells. Cell Stem Cell 6, 71–79 (2009)
Niwa, H., Miyazaki, J. & Smith, A. G. Quantitative expression of Oct-3/4 defines differentiation, dedifferentiation or self-renewal of ES cells. Nature Genet. 24, 372–376 (2000)
Golipour, A. et al. A late transition in somatic cell reprogramming requires regulators distinct from the pluripotency network. Cell Stem Cell 11, 769–782 (2012)
Samavarchi-Tehrani, P. et al. Functional genomics reveals a BMP-driven mesenchymal-to-epithelial transition in the initiation of somatic cell reprogramming. Cell Stem Cell 7, 64–77 (2010)
Polo, J. M. et al. A molecular roadmap of reprogramming somatic cells into iPS cells. Cell 151, 1617–1632 (2012)
Woltjen, K. et al. piggyBac transposition reprograms fibroblasts to induced pluripotent stem cells. Nature 458, 766–770 (2009)
Soncin, F. et al. Abrogation of E-cadherin-mediated cell–cell contact in mouse embryonic stem cells results in reversible LIF-independent self-renewal. Stem Cells 27, 2069–2080 (2009)
Soncin, F. et al. E-cadherin acts as a regulator of transcripts associated with a wide range of cellular processes in mouse embryonic stem cells. PLoS ONE 6, e21463 (2011)
Larue, L. et al. A role for cadherins in tissue formation. Development 122, 3185–3194 (1996)
Müller, F.-J. et al. Regulatory networks define phenotypic classes of human stem cell lines. Nature 455, 401–405 (2008)
Wernig, M. et al. A drug-inducible transgenic system for direct reprogramming of multiple somatic cell types. Nature Biotechnol. 26, 916–924 (2008)
Okita, K., Ichisaka, T. & Yamanaka, S. Generation of germline-competent induced pluripotent stem cells. Nature 448, 313–317 (2007)
Hussein, S. M. I. et al. Genome-wide characterization of the routes to induced pluripotency. Nature http://dx.doi.org/10.1038/nature14046 (2014)
Benevento, M. et al. Proteome adaptation in cell reprogramming proceeds via distinct transcriptional networks. Nature Commun. http://dx.doi.org/10.1038/ncomms6613 (2014)
Lee, D. S. et al. An epigenomic roadmap to induced pluripotency reveals DNA methylation as a reprogramming modulator. Nature Commun. http://dx.doi.org/10.1038/ncomms6619 (2014)
Clancy, J. L. et al. Small RNA changes en route to distinct cellular states of induced pluripotency. Nature Commun. http://dx.doi.org/10.1038/ncomms6522 (2014)
Li, R. et al. A mesenchymal-to-epithelial transition initiates and is required for the nuclear reprogramming of mouse fibroblasts. Cell Stem Cell 7, 51–63 (2010)
Chan, E. M. et al. Live cell imaging distinguishes bona fide human iPS cells from partially reprogrammed cells. Nature Biotechnol. 27, 1033–1037 (2009)
Belteki, G. et al. Conditional and inducible transgene expression in mice through the combinatorial use of Cre-mediated recombination and tetracycline induction. Nucleic Acids Res. 33, e51 (2005)
Nagy, A., Gertsenstein, M., Vintersten, K. & Behringer, R. R. Manipulating the Mouse Embryo: A Laboratory Manual 3rd edn (Cold Spring Harbor Laboratory Press, 2003)
Gertsenstein, M. et al. Efficient generation of germ line transmitting chimeras from C57BL/6N ES cells by aggregation with outbred host embryos. PLoS ONE 5, e11260 (2010)
Hussein, S. M. et al. Copy number variation and selection during reprogramming to pluripotency. Nature 471, 58–62 (2011)
Imamura, M. et al. Transcriptional repression and DNA hypermethylation of a small set of ES cell marker genes in male germline stem cells. BMC Dev. Biol. 6, 34 (2006)
Han, D. W. et al. Epiblast stem cell subpopulations represent mouse embryos of distinct pregastrulation stages. Cell 143, 617–627 (2010)
Acknowledgements
We are grateful for A. Bang’s expertise and assistance regarding flow cytometry. We thank M. Gertsenstein and M. Pereira for chimaera production, and K. Harpal for teratoma sectioning. We would also like to acknowledge the assistance and support of lab colleagues, collaborators and all the members of the Project Grandiose Consortium who are too numerous to name individually but who made a positive impact on this research. A.N. is Tier 1 Canada Research Chair in Stem Cells and Regeneration. This work was supported by grants awarded to A.N. and I.M.R. from the Ontario Research Fund Global Leadership Round in Genomics and Life Sciences grants (GL2), to A.N. from the Canadian stem cell network (9/5254 (TR3)) and Canadian Institutes of Health Research (CIHR MOP102575), to J.-S.S. by the South Korean Ministry of Knowledge Economy (no. 10037410), SNUCM research fund (grant no. 0411-20100074), and Macrogen Inc. (no. MGR03-11 and 12), to S.G. from the Australian Research Council (no. SR110001002), and to C.A.W. by a Queensland government Smart Futures Fellowship and an ARC by Stem Cells Australia and to T.P. grants from NHMRC and ARC. S.M.I.H. received a fellowship from the McEwen Centre of Regenerative Medicine.
Author information
Authors and Affiliations
Contributions
P.D.T. and A.N. conceived and designed the experiments, and wrote the manuscript. P.D.T. and A.J.C. derived all iPSC lines, performed real-time PCR analysis and bisulphite sequencing analysis of DNA methylation. P.D.T., C.M. and I.P.M. performed the in vivo characterization of the iPS lines (teratomas and chimaera formation). O.K., J.L.C. and T.P. assisted in data analysis. P.D.T., J.C.M. and C.A.W. performed microarray analysis. M.L., M.C.P., S.M.I.H. and I.M.R. performed pull-downs for genome-wide ChiP-seq. D.-S.L., J.-Y.S., and J.-S.S. performed genome-wide MethylC-seq and ChiP-seq. N.C., D.L.W., M.E.G. and S.M.G. performed RNA-seq.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Expression profile of F-class cells.
a, Quantitative RT–PCR analysis of total reprogramming factor expression in day 16 F-class (n = 6) and C-class (n = 22), non-parametric t-test. b, Differentially expressed genes (two-tailed Welch t-test P < 0.01, FDR < 0.01) between transgene-expressing reprogrammed lines (n = 28) and ESC-like lines (n = 3). c, Genes highly expressed in b compared against parental fibroblasts. Genes >twofold higher than fibroblasts classified as reprogramming induced. d, Scatter plot of differentially expressed genes (Welch’s t-test P < 0.01; FDR < 0.05). e, Quantitative RT–PCR profiling of cells in a. Non-parametric t-test between the F- and C-class lines (n = 28) ; *P < 0.05, **P < 0.01, ***P < 0.001. f, Expression of PluriNet genes were compared between ESC-like state and F-class state (P values < 0.05, adjusted using the Benjamini–Hochberg method). GeneMANIA interaction network of known gene co-expression and physical interactions. Black nodes represent input genelist, grey nodes represent connecting genes, red nodes represent non-PluriNet genes identified by GeneMANIA that are downregulated in F-class cells.
Extended Data Figure 2 Comparison to epiblast stem cells.
a, Flow cytometric analysis of Nanog expression in F- and C-class primary cell lines after 21 days of transgene expression. Graphs show one of n = 2 experiments. b, Immunofluorescent staining of F-class cells (clone 2) after 30 days of transgene expression. Blue represents Hoechst DNA stain. Scale bars, 100 μm. c, Unsupervised hierarchical clustering of gene expression. EpiSC and ESC populations from ref. 36, with all other cell lines described in Fig. 1b. d, Quantitative RT–PCR analysis of F-class cells (day 30) grown in EpiSC media for 7 days. Graphs show one of n = 2 biological replicates, with 3 technical replicates each. e, Proliferation of established F-class cells (day 30) plated in different media compositions, 1,000 cells plated per cm2, n = 3 technical replicates from one experiment.
Extended Data Figure 3 A stable stem-cell state.
a, Schematic of Cdh1 overexpressing sleeping beauty transposon. IR depicts sleeping beauty inverted repeats. Quantitative RT–PCR of gene expression after 7 days Cdh1 overexpression. n = 3 technical replicates from one experiment. b, Images of Cdh1 overexpressing F-class cells. Scale bars, 100 μm. c, Quantitative RT–PCR analysis of 12 sub-lines derived from clone 1 F-class cells. Average ± s.d. d, DPPA4 immunofluorescence of clone 1 (sub-line-1) after 30 days of transgene expression. Scale bars, 200 μm. e, G-banded karyotype on diploid metaphases of F-class clones. f, Ability to maintain a reprogrammed state in the absence of transgene expression, doxycycline removed after day 21, n = 3 technical replicates from one experiment. Clonal lines ordered as in Fig. 1b.
Extended Data Figure 4 F-class expansion in absence of LIF signalling.
a, Nanog immunofluorescence of single-cell-derived colonies (Day 5). b, Clonal efficiency of F-class cells and ESCs treated with JAK inhibitor (data shown is the mean from n = 3 biological replicates, with 3 technical replicates each, average ± s.d.). c, 1B secondary fibroblast reprogramming25 initiated by doxycycline treatment of fibroblasts in either JAKi-supplemented media (no LIF) or LIF-supplemented media (standard serum-based ESC media). Scale bars, 200 μm. d, Cell expansion of c during 10 days of reprogramming (data shown is the mean from 3 technical replicates from one experiment). e, f, Gene expression analysis (qRT–PCR) of Day 16 reprogramming in JAKi and LIF media (c). Assessment of F-class markers (e) and ESC markers (f) (data shown is the mean from n = 2 biological replicates with 3 technical replicates each). g, DsRed ESCs were mixed with GFP F-class cells. Flow cytometric analysis of population composition before and after passaging. h, Proliferation of F-class and ESC cells grown as suspension culture. i, Phase contrast image of cells grown in suspension for 9 days. Scale bars, 200 μm.
Extended Data Figure 5 In vitro differentiation to three germ layers.
a, TUJ1-positive neurons generated by F-class cells upon doxycycline withdrawal in serum-free media (day 30). b, Multiple neuronal subtypes generated by F-class cells, (clone 1, sub-line 1). c, Quantitative RT–PCR analysis of gene expression during neural differentiation. Clone 1 F-class cells (black line) in comparison to ESC differentiation (grey line). 3 biological replicates (average ± s.d.). d, Doxycycline withdrawal induced differentiation of day 35 F-class cells in 15% serum-based media for 8 days. Immunofluorescent staining of cells representing endoderm (FoxA2) and mesoderm (α-SMA). Scale bars, 200 μm.
Extended Data Figure 6 Transgene expression levels direct reprogramming.
a, Schematic representation of three assessed reprogramming systems. b, Quantitative RT–PCR of factor expression of clonal lines, day 25–day 35. Each point represents a clonal reprogramming colony with n = 6 biological replicates and 3 technical replicates each. c, Representative images of transgene-expressing cells. The day 14 images are representative of 3F and low-expressing 4F reprogramming to highlight the appearance of ESC-like colonies. Scale bars, 200 μm. d, Nanog expression in day 30 colonies. High-expressing 4F cells (1B) exhibit the F-class cell morphology. Scale bars, 100 μm. e, Principal component analysis of quantitative RT–PCR values (32 genes). 3F, 4F low and 4F high cell lines are described in a and b. Cell-state landmarks are F-class clones 1 and 5 (red squares), and C-class clones 5, 10 and 23 (blue squares). f, Quantitative RT–PCR analysis of low-expressing 4F (Col1a1, grey line) and high-expressing 4F (2° 1B, black line) reprogramming, n = 1. g, Schematic model of proposed cell reprogramming routes.
Extended Data Figure 7 Adult tail tip derived F-class cells.
a, Tail-tip fibroblast-derived F-class cells. Scale bars, 200 μm. b, Quantitative RT–PCR analysis of gene expression (day 25 of transgene expression) in clonal tail tip fibroblast reprogrammed cell lines n = 7 biological replicates. c, Principal component analysis of gene expression profile (quantitative RT–PCR, 32 genes). d, Retroviral silencing during transposon mediated reprogramming to F-class state. Quantitative PCR analysis of retroviral copy number (genomic DNA levels) and RNA transcription (n = 3 technical replicates from one experiment). e, Retroviral silencing in established F-class cells (n = 3 technical replicates from one experiment).
Extended Data Figure 8 Requirement of four reprogramming factors.
a, Phase contrast images of F-class cells (CAG-3F + tetO Myc cells). Scale bars, 200 μm. b, Quantitative RT–PCR analysis of reprogramming factor expression, two independent cell lines (data are from n = 2 biological replicates with 3 technical replicates each). c, Genes exhibiting >twofold change upon doxycycline removal (Illumina BeadArray, two independent clones). d, Gene ontology term enrichment of differential gene expression. e, Reprogramming factor expression was activated in ESC-like cells (1B primary iPS cell line). f, Quantitative RT–PCR of reprogramming factor expression in cell lines (n = 10) established from F-class colonies picked in e. g, Quantitative RT–PCR expression of ESC and F-class gene identifiers in cell lines (n = 10) established from F-class colonies picked in e.
Extended Data Figure 9 HDACi-induced transition to ESC-like state.
a, Quantitative RT–PCR of gene expression in F-class cells (day 30, doxycycline-supplemented) that were either maintained in 2i media or exposed to HDAC inhibitors for 10 days. Two cell lines representative of six F-class lines. Data are from n = 3 technical replicates from one experiment. b, Quantitative RT–PCR of F-class (clone 1) sub-lines (n = 12) treated with 10 nM trichostatin A for 6 days. Line denotes average. c, Cell division rate of HDACi-treated cells, as determined directly by time-lapse analysis. d, Flow cytometric analysis of cell viability upon HDACi treatment (10 nM trichostatin A).
Extended Data Figure 10 Temporal effect of HDACi.
a, Schematic representation of HDACi treatment. b, Quantitative RT–PCR analysis of gene expression during HDACi treatment. Red bars depict time points of HDACi exposure (10 nM TSA). Data are from n = 3 biological replicates with 3 technical replicates each (average ± s.d.). c, Principal component analysis of gene expression (Illumina BeadArray). d, Gene ontology term enrichment analysis of genes during HDACi treatment. e, Unsupervised hierarchical clustering of gene expression (Illumina BeadArray) corresponding to primary reprogrammed clones after 16 days of transgene expression and day 16 cells from the 1B secondary reprogramming system (1BD16).
Supplementary information
Supplementary Information
This file contains Supplementary Information sets 1-6. Supplementary Information 1 contains differentially expressed genes (twotailed Welch t-test p<0.01, FDR<0.01) between 28 transgene-expressing reprogrammed lines and 3 ESC-like lines. Supplementary Information 2 contains differentially expressed genes (Welchs t-test p<0.01 FDR<0.05) between F-class and C-class cells. Supplementary Information 3 contains plurinet genes of statistically significant differential expression between F-class samples and ESC samples. Supplementary Information 4 contains differential gene expression upon inactivation of c-Myc expression in F-class cells. Supplementary Information 5 contains differential gene expression upon treatment of F-class cells with HDAC inhibitors. Supplementary Information 6 contains epigenetic status of loci that exhibited differential gene expression (>5 fold) between F-class state and ESC-like cell state. The status of three epigenetic marks are examined; H3K4 trimethylation (H3K4me3), H3K27 trimethylation (H3K27me3) and CpG methylation. Supplementary Information 7 contains primer sequences for quantitative RT-PCR analysis of gene expression. (XLSX 8992 kb)
Rights and permissions
About this article
Cite this article
Tonge, P., Corso, A., Monetti, C. et al. Divergent reprogramming routes lead to alternative stem-cell states. Nature 516, 192–197 (2014). https://doi.org/10.1038/nature14047
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14047
- Springer Nature Limited
This article is cited by
-
Cell-type dependent regulation of pluripotency and chromatin remodeling genes by hydralazine
Stem Cell Research & Therapy (2023)
-
Determining epigenetic memory in kidney proximal tubule cell derived induced pluripotent stem cells using a quadruple transgenic reprogrammable mouse
Scientific Reports (2022)
-
Imprinting fidelity in mouse iPSCs depends on sex of donor cell and medium formulation
Nature Communications (2022)
-
Here and there a trophoblast, a transcriptional evaluation of trophoblast cell models
Cellular and Molecular Life Sciences (2022)
-
Porcine Primordial Germ Cell-Like Cells Generated from Induced Pluripotent Stem Cells Under Different Culture Conditions
Stem Cell Reviews and Reports (2022)