Abstract
The AMT1 family comprises major ammonium transporters in rice roots. In this study, we utilized AMT1 RNAi mutants (amt1) to explore how AMT1 affects NH4+- and NO3–-mediated morphological development and NH4+-responsive gene expression in roots. In the presence of NH4+, amt1 showed inhibition of NO3–- dependent lateral root development. The inhibitory action of NH4+ on lateral root growth was independent of the NO3– concentrations supplied to amt1 roots. The results of split root assays indicated that NH4+ exerts systemic action in inhibiting NO3–-dependent lateral root development in amt1. Further study with NAA and NOA, a potent auxin flux inhibitor, suggested that perturbation of membrane dynamics might not be the primary cause of the inhibitory action of NH4+ on NO3–-mediated lateral root growth in amt1 mutants. RNA-seq analysis of NH4+-responsive genes showed that approximately half of DEGs observed in wild-type roots were not detected in the DEGs of amt1 roots. Gene ontology enrichment analysis suggested that the expression of specific functional gene groups were affected by amt1 during the early response to NH4+. Auxin-responsive gene expression and root gravity responses were altered in amt1. This study demonstrated that AMT1 affects the interactions not only between ammonium and nitrate in lateral root growth but also between auxin and NH4+ in rice roots.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nitrogen has an essential role in plant growth and development. The initial step in N assimilation is the uptake of nitrate (NO3−) and ammonium (NH4+) from soil solution into root cells, which is primarily facilitated by a specific transporter. NH4+ ions accumulate in cells either by direct uptake from the rhizosphere via ammonium transporters (AMTs) or by reduction of NO3–. Rice plants have developed a high tolerance against NH4+ toxicity compared with other grasses, which depends on an energetically favorable equilibration of NH4+ influx and efflux under conditions of elevated NH4+ levels (Britto et al. 2001). High-affinity NH4+ uptake into root cells is mediated by AMT-type ammonium transporters; in rice, this comprises a family of ten AMT paralogs in four subfamilies (Suenaga et al. 2003; Loque and von Wiren 2004). The paralogs AMT1;1, AMT1;2, and AMT1;3 are most important for physiological and morphological responses. Overexpression of AMT1;1 enhanced NH4+ uptake and improved plant growth and yield production in rice under specialized N-fertilization conditions (Ranathunge et al. 2014). By contrast, overexpression of AMT1;3 resulted in poor growth and reduced NH4+ uptake in rice (Bao et al. 2015).
Extensive studies have reported that early genomic responses of rice and Arabidopsis to exogenous NH4+ triggers multiple specific changes in gene expression, metabolism, hormonal signaling, redox status, and root system architecture (Patterson et al. 2010; Li et al. 2010; Lima et al. 2010; Fernánandez-Crespo et al. 2015; Xuan et al. 2013, Xuan et al. 2019; Moon et al. 2019; Kim et al. 2019). Many of these responses are independent of NH4+ assimilation; therefore, NH4+ has been considered as a signaling molecule and AMT1 acts as a sensor (Gaur et al. 2012; Sonada et al. 2003). In Arabidopsis, AMT1;3 is required for NH4+-dependent lateral root branching (Lima et al. 2010). It has been reported that nitrate transporters act as sensors in NO3−-dependent root growth. NRT1.1 controls the growth of lateral root primordia under conditions of low NO3− (Bouguyon et al. 2016). Under N-limited conditions, AtNRT2.1 may act as a NO3− sensor or signaling component that represses lateral root initiation, which is independent of NO3− uptake activity (Little et al. 2005; Remans et al. 2006). The negative effect of NRT1.1/NRT2.1 on lateral root development may represent a distinct systemic pathway under low nitrate conditions.
Many studies have reported that hormonal signaling pathways are tightly connected with NH4+-related plant growth and stress responses. The auxin-resistant aux1, axr1, and axr2 mutants are insensitive to NH4+-mediated inhibition of root growth in Arabidopsis (Cao et al. 1993). Application of NH4+ to shoots causes the auxin influx carrier AUX1 to inhibit lateral root emergence (Li et al. 2011). ARG1 (ALTERED RESPONSE TO GRAVITY1) is required for normal AUX1 expression and basipetal auxin transport in the root apex, and arg1 mutants are sensitive to NH4+ (Zou et al. 2013). Ethylene production in shoots is associated with NH4+-mediated lateral root inhibition (Li et al. 2013). The activation of ABA signaling reduces NH4+-induced stress in a mutant of AMOS1 (AMMONIUM OVERLY SENSITIVE1)/EGY1 (ETHYLENE-DEPENDENT, GRAVITROPISM-DEFICIENT, AND YELLOW-GREEN-LIKE PROTEIN1) (Li et al. 2012). RAVL1 (RELATED TO ABI3/VP1-LIKE1), a key brassinosteroid (BR) signaling transcription factor in rice, regulates BR-mediated induction of AMT1;2 and NH4+ uptake (Xuan et al. 2017). High concentrations of NH4+ in rice induce primary root coiling in the light, which is rescued by inhibition of NH4+ assimilation (Hirano et al. 2008; Shimizu et al. 2009). However, few studies have explored the relationship between hormones and NH4+.
In this study, we examined NH4+- or NO3−-mediated root growth and gene expression using AMT1 RNA interference (RNAi) mutants (amt1). The results demonstrated that NH4+ systemically suppressed NO3−-dependent lateral root development in amt1 mutants. The inhibitory action of NH4+ on lateral root development (especially the number of lateral roots) was independent of the NO3– concentrations supplied to amt1 roots. We performed RNA-seq and gene ontology (GO) enrichment analysis to evaluate how AMT1 activity affects the expression of NH4+-responsive genes, and examined the expression patterns of auxin-related genes and root gravity responses in amt1. This study demonstrated that AMT1 substantially affected not only the interaction between ammonium and nitrate in lateral root growth but also auxin-responsive gene expression and gravity responses in roots.
Materials and Methods
Construction of AMT1 RNAi Vector
To generate AMT1;1 RNAi transgenic plants, 5′ and 3′ fragments of the AMT1;1 ORF were amplified using the following primer sets: Ri5-F (gagctcggtaccctcgccgcgcacgtcatccag) and Ri5-R (gaattcctgcaggcatgtgcttgaggccgaaga); Ri3-F (gagctcggtaccctcgcggcgcacatcgtgcag) and Ri3-R (gaattcctgcagttacacttggttgttgctgtt), respectively. The PCR products were digested and cloned into EcoRI and SacI sites for the sense orientation insertion and into KpnI and XhoI sites for the antisense orientation insertion in a pBluscript-catalase intron vector. After sequencing, the whole inserts (Fig. S1a) were cloned into SacI and KpnI sites of the PGA1611 binary vector.
Generation of AMT1 RNAi Transgenic Lines
The AMT1 RNAi transgenic lines were generated from japonica rice cultivar ‘Dongjin’ via Agrobacterium (LBA4404 strain)-mediated transformation using calli derived from dry seeds (Chin et al. 1999). The following transgenic lines were selected and propagated: 5′ AMT1 RNAi lines 5–1, 5–2, and 5–3; 3′ AMT1 RNAi lines 3–11, 3–12, 3–13, 3–14, and 3–17.
Plant Materials and Growth Conditions
The japonica cultivar Dongjin (WT), three 5′ AMT1 RNAi lines (5–1, 5–2, and 5–3), and five 3′ AMT1 RNAi lines (3–11, 3–12, 3–13, 3–14, and 3–17) were utilized in the experiments. Rice seeds were surface-sterilized with 0.05% SPORTEX and then germinated for 3 days in the dark. Uniformly germinated seedlings were selected and cultured hydroponically in different nutrient solutions [¼ MS (Murashige and Skoog), ¼ KB (Kimura B), and ¼ NS (Nutrient Solution)] containing NH4+ or NO3– as the sole nitrogen source. Detailed information on the solution components is given in Table S1. Hydroponic nutrient solutions were replaced with fresh media every 2 days for 2 weeks. Roots were examined after culture for 14 days in a growth chamber under the following conditions: 16/8 h light/dark, light intensity 280 μmol m−2 s−1, temperature 26 °C/18 °C, and 70% humidity. To measure the expression levels of three OsAMT1 (1;1, 1;2, and 1;3) genes, AMT2;1, GS1;2, NADH-GOGAT1, GDH1, and GDH2, seedlings were grown hydroponically in ¼ nutrient medium supplemented with 0.5 mM NO3– or 0.5 mM NH4+ for 7 days. Total cellular RNAs were extracted from roots. For media shift assays, germinated seeds were cultured in modified ¼ NS containing 0.1 mM NH4NO3 for 7 days. The samples were transferred and cultured in the nutrient solution containing 0.1 mM of either NH4NO3, NH4+, or NO3– for an additional 7 days. The same solutions were replaced with fresh medium every 2 days. To investigate the combinatory effects of auxins and inhibitors, germinated seeds were cultured in modified ¼ NS containing 0.01 μM 1-naphthaleneacetic acid (NAA), 0.01 μM 1-naphthoxyacetic acid (NOA), 0.01 μM N-1-naphthylphthalamic acid (NPA), NAA + NOA, or NAA + NPA for 14 days. The same solutions were replaced with fresh medium every 2 days.
Methylammonium (MeA) Treatment
Uniformly germinated seeds were grown hydroponically in modified full nutrient (FN) medium (2 mM NH4NO3, 1 mM KH2PO4, 1 mM MgSO4, 250 mM K2SO4, 250 mM CaCl2, 100 mM NaFe-EDTA, 50 mM KCl, 50 mM H3BO3, 5 mM MnSO4, 1 mM ZnSO4, 1 mM CuSO4, 1 mM NaMoO4, and 1 mM MES, pH 5.8 [KOH]) (Chaudhuri et al. 2008) supplemented with different concentrations of methylammonium (0, 1, 2.5, and 5.0 mM) for 10 days. Root length, shoot height, and dry weight were analyzed.
Measurements of the Seminal, Crown, and Lateral Roots
Seminal root length was manually measured with a scale. Crown and lateral roots were imaged by microscopy (DP70; Olympus, Japan), and their lengths were measured using ImageJ software. Crown roots of less than 0.5 cm in length were counted separately from those with root lengths longer than 0.5 cm. The number of lateral roots was counted within 1.0 cm from the differential zone of seminal roots, where lateral roots can be visibly recognized. The density of lateral roots was calculated using ImageJ software. The ten longest lateral roots were measured to calculate the average length of lateral roots.
RNA Extraction and qRT-PCR
Total cellular RNA was purified using the Qiagen RNeasy Mini Kit (Qiagen, Valencia, CA, USA) according to the manufacturer’s instructions. RNA concentrations were measured using a NanoDrop ND-1000 spectrophotometer, and samples were treated with RQ-RNase-free DNase (Promega, Madison, WI, USA). A reverse transcriptase RNaseH (Toyobo, https://www.toyobo-global.com/) transcription kit was used to synthesize cDNA according to the manufacturer’s instructions (Promega). Then, qRT-PCR was performed using iQ SYBR Green Supermix (Bio-Rad) and gene-specific primers using the CFX Manager software (Bio-Rad) instrument, and values were normalized against UBIQ1 levels in the same samples. A minimum of three biological and two technical replicates were used for each analysis. All primers used for qRT-PCR are presented in Table S2.
Determination of Ammonium Contents
Enzymatic determination of NH4+ content in the roots was performed using an F-kit (Roche) according to the manufacturer’s instructions (Oliveira et al. 2002).
Determination of Glutamine Contents
Enzymatic determination of glutamine contents in roots was performed using an l-glutamine, ammonia Rapid Assay Kit from Megazyme (Megazyme International Ireland Ltd, Co. Wicklow, Ireland) according to the manufacturer’s instructions (Barth et al. 2010).
Split Root Assay
Surface-sterilized seeds were cultured in dH2O for 4 days. Then, crown roots of each plant were divided into two groups, and each group of roots was submerged in one of two split containers filled with solutions of different N nutrients. After 14 days, plants displaying balanced root growth in the split containers were selected to measure lateral root densities and seminal root lengths.
RNA-Sequencing Analysis
Wild-type (WT) japonica rice cv. Dongjin and AMT1 RNAi line 5–2 were utilized for RNA-seq analysis. Sample seedlings were cultured in the following way. After germination, seedlings were grown in distilled water for 14 days in a glasshouse to ensure depletion of endosperm nutrients. These plants were grown in ¼ nutrient medium lacking N for an additional 3 days and then were transferred to the same nutrient solution containing 0.1 mM NH4+ for 0 h or 3 h. Two biological replicates were used for each of the four samples as follows: WT (0 h), WT (3 h), Ri 5–2 (0 h), and Ri 5–2 (3 h). Total RNAs were extracted from roots of the samples using the RNeasy Plant Mini Kit (Qiagen, https://www.qiagen.com/). Construction of cDNA libraries for RNA-seq and data analysis is described in Supplementary Appendix S1.
Root Gravity Analysis
To analyze the gravity response in root tips, 3-day-old WT seedlings grown in water were transferred to water or 0.05 mM (NH4)2SO4 solution, and reoriented so that the root tips were set at an angle 90° away from the direction of gravity. The angle between a horizontal line and the direction of root tip growth was measured every 30 min.
Gene Ontology Enrichment Analysis
GO and rice gene assignments were downloaded from the RiceNetDB Database (https://bis.zju.edu.cn/ricenetdb/) (Liu et al. 2013). A total of 991 upregulated and 395 downregulated gene loci were uploaded in a GO enrichment analysis toolbox, and the biological processes, molecular functions, and cellular components were specified for the analysis.
Image Analysis
To measure the lengths of crown roots and lateral roots, root tissue samples were collected from 2-week-old seedlings, fixed in 70% ethanol, and imaged by microscopy (DP70; Olympus, Japan). The numbers of crown roots and lateral roots were measured by ImageJ.
Statistical Analyses
Statistical calculations were performed using Prism 5 (GraphPad, San Diego, CA, USA). Comparisons between groups were made using one-way analysis of variance (ANOVA; Brady et al. 2015), followed by Bonferroni’s correction for multiple comparisons. Differences in P values < 0.05 were considered as statistically significant. All data are expressed as the mean ± SE or SD.
Results
Generation and Expression of OsAMT1 RNAi Transgenic Rice Lines
OsAMT1 RNAi lines were generated by using two genomic DNA regions as target sequences, the 269 and 277 bp regions from the 5′ and 3′ ends of OsAMT1;1, respectively (Fig. S1a and b). The 5′ target region of OsAMT1;1 showed 88% and 95% sequence identity with the 5′ target regions of AMT1;2 and AMT1;3, respectively (Fig. S1b). The 3′ target region of OsAMT1;1 showed 7.9% and 20% sequence identity with the 3′ target regions of AMT1;2 and AMT1;3, respectively (Fig. S1b). A total of three and seven RNAi lines were generated using the 5′ and 3′ target sequences, respectively. To measure the steady-state expression levels of three AMT1 genes (AMT1;1, AMT1;2, and AMT1;3) in the mutants, germinated seeds were grown hydroponically for 10 d in ¼ NS (Sonoda et al. 2003) containing 0.5 mM NO3– or 0.1 mM NH4+ as a nitrogen source. After 7 days of culture, the expression levels of three AMT1 genes in roots were measured by performing quantitative RT-PCR. The expression of all three AMT1 genes was suppressed in all ten RNAi lines grown in NO3– (Fig. 1a) and NH4+ (Fig. S2) compared with the expression in WT plants. The mRNA levels of AMT1;1, AMT1;2, and AMT1;3 were similar in all RNAi lines. Thus, the RNAi lines were named as AMT1 RNAi lines (or amt1 mutants) instead of AMT1;1 RNAi.
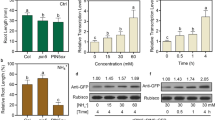
Expression of three AMT1 genes (a) and methylammonium sensitivities of OsAMT1 RNAi lines (b). a Total cellular RNAs from roots were subjected to qRT-PCR analysis to measure the expression levels of AMT1;1, AMT1;2, and AMT1;3. WT plants and ten OsAMT1 RNAi lines were grown hydroponically in ¼ NS containing 0.5 mM NO3– for 7 days. Composition of the nutrient solution is listed in Table S1. The mRNA levels in the samples were normalized against those of ubiquitin mRNA. Error bars are ± SD of the means of three qPCR replicates. b Seedlings were hydroponically cultured on modified full nutrient (FN) medium containing 0, 1.0, 2.5, or 5.0 mM MeA for 10 days after germination. Primary root length (c) and shoot length (d) was measured from 10-day-old seedlings grown in the presence of 0, 1.0, 2.5, or 5.0 mM MeA. Data of (c) and (d) are means ± SE (n > 10 plants per line); different letters indicate significant differences between samples (P < 0.05). Significant differences of seminal root length and shoot height in responses to MeA solution are shown (*P < 0.05)
The expression levels of AMT2;1 and four genes related to NH4+ assimilation were examined, and they displayed similar levels in RNAi and WT lines (Fig. S3a). Those four NH4+ assimilation-related genes included cytosolic glutamine synthetase (GS1;2), NADH glutamate synthase 1 (NADH-GOGAT1), and two glutamate dehydrogenases (GDH1 and GDH2). To investigate the long-term effects of these RNAi mutations on NH4+ uptake, plants were grown hydroponically in modified full nutrient (FN) medium (Chaudhuri et al. 2008) containing 0, 1.0, 2.5, or 5.0 mM methylammonium (MeA) for 10 days (Fig. 1b–d). WT plants treated with MeA exhibited severe growth inhibition in a dose-dependent manner. By contrast, none of the MeA-treated amt1 mutants exhibited any significant retardation of the shoot and root growth. WT plants had much lower dry weight than amt1 mutants (Fig. S3b). These data indicate that mutant roots are substantially inefficient in NH4+ uptake. From the total of ten RNAi lines, we selected two lines from each of the 5′ and the 3′ RNAi lines for subsequent studies. These were Ri 5–1 and Ri 5–2 for the 5′ RNAi lines, and Ri 3–1 and Ri 3–2 for the 3′ RNAi lines. Internal NH4+ levels were measured in the roots of these four RNAi lines grown in ¼ NS containing 0.1 mM NH4+ as the sole nitrogen source for 14 days (Fig. S4). All amt1 mutants contained approximately 30–60% lower levels of NH4+ than WT plants.
NH4+-Mediated Inhibition of Lateral Root Development in OsAMT1 RNAi Roots
The effect of N on plant growth could be modulated by other nutrients in the culture media. Therefore, we compared three well-established media to examine the effect of NH4+ and NO3– on amt1 root growth and development. The media included ¼ MS (¼-strength Murashige and Skoog) (Murashige et al. 1962), ¼ KB (¼-strength Kimura B) (Chen et al. 2006), and ¼ NS (¼-strength Nutrient Solution) (Abiko et al. 2005) (Table S1). When all four AMT1 RNAi lines and WT plants were cultured in dH2O, there was no difference in root growth and development (Fig. S5). To examine the effects of NH4+ and NO3– on root growth, germinating seeds were grown for 14 days in three solutions containing either 0.1 mM NH4+ or NO3– as the sole N source. The length of seminal roots, the number of crown roots, and the density and the average length of lateral roots were measured and compared between mutants and WT plants grown under the same conditions.
Seminal roots in 14-day-old rice plants do not grow any further, and the numbers of lateral roots on seminal roots no longer increase, although lateral and crown roots keep growing when cultured further. The numbers of crown roots increase during longer culture. Crown roots less than 0.5 cm in length were counted separately from those longer than 0.5 cm. At this stage, the average length of WT crown roots is approximately 7–8 cm. The numbers of lateral roots were counted within 1 cm from the differential zone of seminal roots, at which point lateral roots can be visibly recognized. As there were wide variations in lateral root lengths of both WT and mutants, the ten longest lateral roots were counted for the average length of lateral roots. In all three media containing 0.1 mM NO3– as the sole N source, amt1 mutants and WT did not display any differences in these three growth parameters (Fig. S6). However, in all media containing NH4+, amt1 mutants displayed severe growth retardation in roots (Fig. 2). For crown roots cultured in all NH4+-containing media, the total numbers of crown roots were the same in WT and amt1 mutants (Fig. 2b, e). Approximately 20–50% of all crown roots were short (< 0.5 cm) in amt1 mutants, whereas WT did not have short crown roots. Among three culture media, ¼ NS showed the most distinct effect of NH4+ on root growth of amt1 mutants.

source for 14 d. Fresh nutrient solutions were provided every 2 days. a, b, c, d show quantification of seminal root length, crown root numbers, lateral root density, and lateral root length, respectively. Crown roots longer than 0.5 cm were counted separately from those shorter than 0.5 cm. e Crown roots grown in ¼ NS containing NH4+. b Each bar consists of a number of crown roots longer than 0.5 cm (lower part) and those shorter than 0.5 cm (upper part). Red arrows in (e) indicate crown roots shorter than 0.5 cm. Scale bar = 1 cm. Data of (a, b, c, d) are means ± SE (n > 10 plants per line). Different letters indicate significant differences between samples (P < 0.05)
a–d Effect of NH4+ on root development in amt1 mutants cultured in three different media (MS, KB, and NS). e Images of crown roots of plants grown in ¼ NS containing 0.1 mM NH4+. Uniformly germinated seeds of WT and four AMT1 RNAi lines (Ri 5–1, Ri 5–2, Ri 3–1, and Ri 3–2) were cultured in ¼ MS, ¼ KB, and ¼ NS containing 0.1 mM NH4+ as the sole nitrogen
To further examine the N effect on amt1 mutants, seedlings were grown in ¼ NS containing a concentration series (0.01–1 mM) of NH4+, NO3–, and NH4NO3 for 14 days (Fig. 3). The 0.1 mM NH4+ or NH4NO3 concentration was the most stimulating for WT root growth. The 0.1 to 0.3 mM NO3– concentration was optimal for root growth of WT and amt1 mutants. The root morphologies of 14-day-old plants grown in 0.1 and 0.5 mM NH4+, NO3–, or NH4NO3 are shown in Fig. S7. In the presence of NH4+, amt1 mutants showed much lower values of all three growth parameters than WT [Fig. 3a(i), (iv), (vii), and (x)]. There were no growth differences between WT and mutant roots at all NO3– concentrations [Fig. 3a(ii), (v), (viii), and (xi)]. In the presence of NH4NO3, WT and mutant seedlings displayed comparable growth of seminal and crown roots [Fig. 3a(iii) and (vi)], but amt1 mutants displayed severely defective lateral root growth [Fig. 3a(ix) and (xii)]. These data indicate that the total number and average length of amt1 mutant lateral roots in the presence of NH4+ and NH4NO3 were much less than those of WT. For example, at 0.1 mM NH4+ and NH4NO3, lateral root densities in mutants were 27% and 35%, respectively, of those of WT. Detailed morphologies of lateral roots of 14-day-old-plants grown in 0.1 and 0.5 mM NH4+, NO3–, or NH4NO3 are presented in Fig. 3b. These combined results indicate that lateral root development in amt1 is inhibited by NH4+ even in the presence of NO3–. By contrast, seminal and crown roots are not affected by NH4+ in the presence of NO3–.
a Seminal, crown, and lateral root growth of amt1 mutants cultured in ¼ nutrient solution containing a concentration series of NH4+, NO3–, and NH4NO3. b Lateral roots grown in 0.1 and 0.5 mM NH4+, NO3–, and NH4NO3. c Glutamine contents of WT and amt1 (Ri 5–2 and Ri 3–1) roots grown in 0.1 mM NH4+, 0.1 mM NO3−, 0.1 mM NH4NO3, or 0.1 mM NH4+ + 1.0 mM NO3−. a Uniformly germinated seeds of WT and two AMT1 RNAi lines (Ri 5–2 and Ri 3–1) were cultured in ¼ NS containing 0, 0.01, 0.05, 0.1, 0.2, 0.3, 0.5, and 1.0 mM of NH4+ [(i), (iv), (vii), and (x)], NO3– [(ii), (v), (viii), and (xi)], and NH4NO3 [(iii), (vi), (ix), and (xii)] for 14 days. Fresh nutrient solutions were provided every 2 days. After 14 days, root growth and development were analyzed. Seminal root length shown in (i), (ii), and (iii); crown root numbers shown in (iv), (v), and (vi). Only crown roots longer than 0.5 cm were counted for the measurements. Lateral root density is shown in (iii), (vi), and (ix); average length of ten lateral roots shown in (x), (xi), and (xii). The average length of lateral roots is measured using the ten longest ones. b Lateral roots on seminal roots of plants grown in ¼ NS containing 0.1 and 0.5 mM NH4+, NO3–, and NH4NO3 for 14 days. Scale bar = 1 mm. c Uniformly germinated seeds of WT and two AMT1 RNAi lines (Ri 5–2 and Ri 3–1) were cultured in ¼ NS containing 0.1 mM NH4+, 0.1 mM NO3−, 0.1 mM NH4NO3, or 0.1 mM NH4+ + 1.0 mM NO3− for 14 d. Fresh nutrient solutions were provided every 2 days. After 14 days, whole roots were ground in liquid nitrogen and the glutamine content per gram of fresh weight was measured using the L-Glutamine, Ammonia Rapid Assay Kit (Megazyme Ltd.) at 340 nm. Values are means ± SE of three independent replicates. Different letters indicate significant differences between samples (P < 0.05)
To examine whether the growth defect of mutant lateral roots grown in NH4NO3 but not in NO3– might result from low efficiency of N assimilation, cellular levels of glutamine, the first amino acid assimilated from exogenous NO3– and NH4+, were measured in roots. Wild type and amt1 were grown in 0.1 mM NH4+, 0.1 mM NO3–, 0.1 mM NH4NO3, or 0.1 mM NH4+ + 1.0 mM NO3– (Fig. 3c). When grown in NH4+, amt1 roots accumulated much lower levels of glutamine than WT roots, whereas when grown in NO3–, amt1 roots acuminated similar levels of glutamine to the WT roots. When cultured in NH4NO3, amt1 roots contained less glutamine than the wild type. However, the glutamine content of the amt1 roots grown in NH4NO3 was similar to that of amt1 roots grown in NO3–. These results strongly suggest that the lateral root developmental defect of amt1 mutants grown in NH4NO3 is unlikely to be due to inefficient N assimilation or low N nutrients.
Dominant and Systemic Effect of NH4+ on NO3–-Dependent Lateral Root Development in amt1 Mutants
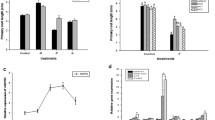
To further explore the relationship between NH4+ and NO3− in amt1 mutant lateral root development, three experiments were performed. First, seedlings were grown in ¼ NS containing 0.1 mM NH4NO3 for 7 days, and then shifted to the same solution containing 0.1 mM of either NH4NO3, NH4+, or NO3– for another 7 days. After 14 days, lateral root densities and lengths were measured and compared (Fig. 4a, b). The lateral root densities of amt1 mutants continuously cultured in NH4NO3 were 53% of those of WT grown under the same conditions. The amt1 mutants shifted to NH4+ for the last 7 days of culture developed 38% of lateral root densities of those of WT cultured under the same conditions. By contrast, amt1 mutants shifted to NO3– had 90% of the lateral root density of WT plants. The average lateral root lengths of amt1 mutants grown only in NH4NO3 were longer than those of mutants shifted to NH4+, but were shorter than those of mutants shifted to NO3– (Fig. 4b). The morphologies of lateral roots grown in the three N culture conditions are shown in Fig. S8a. These combined results indicate that suppression of lateral root growth and development in the presence of NH4NO3 could result from a dominant effect of NH4+ over NO3– in regulating lateral root development in amt1 mutants.
a, b Lateral root densities (a) and lengths (b) in plants shifted to NH4NO3, NH4+, or NO3– after growing in NH4NO3 for 7 days. c, d Effect of NH4+ on lateral root densities (c) and lengths (d) in plants grown in a series of NO3– concentrations. a, b Seedlings grown in 0.1 mM NH4NO3 for 7 days were transferred and cultured in nutrient solution containing 0.1 mM of NH4NO3, NH4+, or NO3– for another 7 days. After 14 days culture, lateral root densities (a) and lengths (b) were measured for WT, Ri 5–2, and Ri 3–1 plants. c, d Seedlings were cultured in nutrient solution containing 0.1, 1.0, 2.5, 5.0, or 10 mM NO3– along with 0.1 mM NH4+. After 14 d of culture, lateral root densities (c) and lengths (d) were measured for WT, Ri 5–2, and Ri 3–1 plants. Values are means ± SE (n > 10 plants per line). Different letters indicate significant differences between samples (P < 0.05)
Second, to evaluate whether the effect of NH4+ on NO3–-dependent lateral root growth might be influenced by NO3– concentration, mutants were incubated in media containing a series (0.1, 1.0, 2.5, 5.0, and 10 mM) of NO3– concentrations along with 0.1 mM NH4+. In the presence of 0.1 mM NH4+, all amt1 mutants grown in various concentrations of NO3– displayed similarly defective lateral root growth (Fig. 4c, d). Lateral root densities were essentially constant among mutants grown in different NO3– concentrations, although those of WT plants were slightly reduced as NO3– concentrations increased. Lateral root lengths slightly increased in amt1 mutants as NO3– concentrations increased (Fig. 4d). The morphologies of lateral roots grown in 0.1, 1.0, or 5.0 mM NO3– along with 0.1 mM NH4+ are shown in Fig. S8b. WT plants and amt1 mutants cultured only in NO3– displayed normal lateral root growth under various NO3– concentrations (Fig. S9a). Seminal and crown root growth in amt1 mutants were essentially identical to those of WT grown in media containing different NO3– concentrations with or without NH4+ (Fig. S9b). Therefore, the specific suppression effect of NH4+ on NO3–-dependent lateral root growth in amt1 mutants is independent of NO3– concentration.
Third, we performed split root assays to evaluate whether the suppression effect of NH4+ could be systemic and specific to NO3– during lateral root development in amt1 mutants. Germinated seeds cultured in dH2O for 4 d displayed well-developed primary root and crown roots. For split root assays, uniformly developed roots were divided into two equivalent parts and placed in two separate nutrient media solutions (Fig. 5). To examine the systemic effect of NH4+ on NO3–-dependent lateral root development, one side of the split root system was cultured in NO3– and the other side was cultured in either NH4+, NO3–, or no N for 14 days (Fig. 5a). Seminal roots were placed in the NO3– containers and were examined for lateral root densities and lengths (Fig. 5a). In WT, lateral root density in the split roots cultured in media containing NO3– was slightly adversely affected when the other half of roots was cultured in NH4+, compared with the density when the other half of roots was cultured in either NO3– or no N. By contrast, in amt1 mutants, the lateral root density when cultured in NO3– displayed 49% reduction when the other half of roots was cultured in NH4+, compared with the density when the other half of roots was cultured in either NO3– or no N. In amt1 mutants, lateral root length in NH4+-containing media was reduced by 35–40% compared with lateral root lengths when the other half of roots was cultured in media containing either NO3– or no N, respectively. To examine the specificity of NH4+-mediated suppression of NO3–-dependent lateral root development, one side of the split root system was cultured in media lacking a N source and the other side was cultured in media containing NH4+, NO3–, or no N (Fig. 5b). Seminal roots were placed in media lacking a N source, and then were inspected for lateral root density and length. WT plants and amt1 mutants displayed essentially identical lateral root densities and lengths in media lacking N for all three split root assays when the other half of roots was cultured in NH4+, NO3–, or no N (Fig. 5b). These combined results indicate that the suppression effect of NH4+ is specific to NO3–-dependent lateral root development. These data strongly suggest that NH4+ has a dominant systemic signaling activity that specifically suppresses NO3–-dependent lateral root development in amt1 mutants.
For split root assays, one side of the split roots was cultured in media containing NO3– (a) or no N source (b), and the other side of the split roots was cultured in NH4+, NO3–, or no N. Roots of WT and Ri 5–2 seedlings that were grown in dH2O for 4 days were split into two separate nutrient media. One side was cultured in media containing NO3– (a) and no N (b), whereas the other side was cultured in NH4+, NO3–, or no N. After 14 d culture, lateral root densities and lengths on seminal roots cultured in NO3– (a) or no N (b) were measured for plants with the other side cultured in NO3–, no N, or NH4+. White, red, and blue columns indicate lateral root densities of roots when the other half of roots were cultured in NO3–, no N, or NH4+, respectively. Values are means of ± SE (n > 10 plants per line). Different letters indicate significant differences between samples (P < 0.05). White arrow heads in the left panels mark the seminal roots
NAA/NOA Exerts Similar, But Not the Same, Action as NH4+ on Lateral Root Growth in amt1 Mutants
Exposure to external NH4+ causes more dramatic changes in apoplastic pH or membrane polarization in amt1 roots than in WT roots (Liu and von Wiren 2017; Wang et al. 1994). To evaluate these dramatic effects on membrane dynamics in amt1 root cells, mutant roots were grown in the presence of both NAA and the potent auxin transport inhibitor NOA. NOA modulates overall auxin transport (both influx and efflux) across the plasma membrane (Lankova et al. 2010). Germinating seeds were grown for 14 days in nutrient media containing 0.01 µM NAA and 0.01 µM NOA. As controls, germinating seeds were grown for 14 days in nutrient media containing 0.01 µM of either a combination of NAA and its efflux inhibitor NPA, or only NAA, NOA, or NPA (Fig. 6a). The root parameters of WT and amt1 mutant plants did not significantly differ in the presence of only NAA, NOA, or NPA. By contrast, mutant lateral roots showed hypersensitive response to the combination of NAA/NOA and NAA/NPA (Fig. 6a). In the presence of NAA/NOA, lateral root density in amt1 mutants was reduced to approximately 30% of that of WT. In the presence of NAA/NPA, lateral root density in amt1 mutants was approximately 75% of that of WT. However, there were no differences in lateral root length between amt1 mutants and WT exposed to either NAA/NOA or NAA/NPA [Fig. 6a(ii)]. The morphologies of WT and mutant lateral roots grown in the presence of NAA, NAA/NOA, or NAA/NPA are presented in Fig. 6a(iii). By contrast, primary and crown root growth were the same in mutant and WT plants exposed to both combinations of auxin and inhibitors (Fig. S10). These results suggest that amt1 mutants became hypersensitive to changes of plasma membrane environments, which might lead to suppression of lateral root development.
a Suppression of amt1 mutant lateral root growth by NAA/NOA or NAA/NPA treatments. b No effect of NAA/NOA on NO3–-dependent lateral root growth. c Suppression effect of NH4NO3 on NO3–-dependent lateral root growth in the presence of NAA/NOA. a Germinated seeds were cultured in ¼ NS containing 0.01 μM NAA, 0.01 μM NOA, 0.01 μM NPA, NAA + NOA, and NAA + NPA. After 14 days culture, lateral root densities (i) and lengths (ii) were measured. Values are means ± SE (n > 10 plants per line). Different letters indicate significant differences between samples (P < 0.05). Photographs show lateral roots on seminal roots of WT and mutants (Ri 5–2 and Ri 3–1) grown in the presence of NAA, NAA/NOA, or NAA/NPA (iii). Scale bar = 1 mm. b Germinated seeds were cultured in ¼ NS containing either 0.1 mM NO3– and 0.01 μM of NAA/NOA or only 0.1 mM NO3−. After 14 days culture, lateral root densities and lengths were measured. Values are means ± SE (n > 10 plants per line). Different letters indicate significant differences between samples (P < 0.05). c Germinated seeds were cultured in ¼ NS containing either 0.1 mM NH4NO3 and 0.01 μM of NAA/NOA, or only 0.1 mM NH4NO3. After 14 days culture, lateral root densities and lengths were measured. Values are means ± SE (n > 10 plants per line). Different letters indicate significant differences between samples (P < 0.05)
To evaluate whether NO3–-dependent lateral root growth is affected by cellular environments induced by NAA/NOA, seedlings were grown for 14 days in ¼ NS with the following components: (1) 0.01 µM NAA/NOA and 0.1 mM NO3–, (2) only 0.01 µM NAA/NOA, or (3) only 0.1 mM NO3– (Fig. 6b). After 14 days, the primary, crown, and lateral roots were measured. The results showed that there were no differences in all root development parameters between mutants and WT grown in all three treatments (Fig. 6b for lateral roots and Fig. S11 for primary and crown roots). Next, we examined whether NAA/NOA affected the inhibitory action of NH4+ on NO3–-dependent lateral root growth (Fig. 6c). Mutants were grown in solutions containing both NH4NO3 and NAA/NOA. The results showed that NH4+ inhibited lateral root density and length even in the presence of NAA/NOA (Fig. 6c). These combined results strongly suggest that perturbation of membrane dynamics may not be the primary cause of the inhibitory action of NH4+ on NO3–-mediated lateral root growth in amt1 mutants.
Comparative Analysis of Transcriptomic Profiling Between AMT1 and amt1 Roots
Our previous work showed that many genes show dramatic expression changes in roots within 3 h after NH4+ treatment (Xuan et al. 2013). To estimate how many NH4+-responsive genes are affected by AMT1 function, we performed transcriptomic profiling using roots of amt1 mutants and WT that were exposed to 0.1 mM NH4+ for 0 and 3 h. Sample seedlings were cultured as follows. After germination, seedlings were grown in distilled water for 14 days in a glasshouse to ensure the depletion of endosperm nutrients. These plants were grown in ¼ nutrient medium lacking N for an additional 3 days, and were then transferred to the same nutrient solution containing 0.1 mM NH4+ for 0 or 3 h. Under these culture conditions, mutant roots did not show any morphological defects. The total cellular RNA samples from roots were used for RNA-seq. Based on our analysis of RNA-seq reads and comparative analysis of transcriptomic profiles, differentially expressed genes (DEGs) were identified in roots of WT and mutant plants. NH4+-responsive DEGs showed at least > twofold differences in expression levels when compared between 0 and 3 h treatments of WT or mutant plants. Based on AMT1 function, the NH4+-responsive DEGs were classified into two groups, class I and II. Class I represents ‘AMT1-dependent’ NH4+-responsive genes that showed twofold differences in expression levels between 0 and 3 h after NH4+ treatment in WT but not mutant roots (Fig. 7a). Among a total of 991 upregulated and 395 downregulated genes identified in WT roots, 467 and 223, respectively, failed to respond to NH4+ treatment in mutant roots. These genes are listed in Table S3. Class II is called ‘AMT1-independent’ NH4+-responsive genes. They showed at least twofold differences in expression levels between WT and mutant plants before and after NH4+ treatment (Fig. 7a). A total of 524 upregulated and 172 downregulated genes in WT roots showed similar expression patterns in amt1 mutant roots (Table S3). We performed qRT-PCR to confirm the expression patterns of some transcription factor genes of class I and II (Fig. 7b).
a Classification of DEGs identified by RNA-seq analysis. b Quantitative RT-PCR analysis to verify genes identified by RNA-seq in WT and amt1 mutants. c GO enrichment analysis of class I and II genes. a Genes that were differentially expressed by NH4+ treatment in WT roots were classified based on their expression patterns in amt1 mutant roots. Left and right graphs indicate the total numbers of DEGs that were upregulated and downregulated, respectively, in WT roots treated with NH4+ for 3 h. Gray color indicates class I DEGs that did not respond to NH4+ in amt1 mutants. Yellow color indicates class II DEGs that showed similar expression patterns in both WT and amt1 mutant roots. Numbers within the box are the numbers of DEGs that belong to the corresponding classes. b Total cellular RNAs from roots of WT and amt1 mutants were used for qRT-PCR analysis. Some of class I ‘AMT1-dependent’ (a, b, e, f) and class II ‘AMT1-independent’ (c, d, g, h) genes were examined by qRT-PCR to measure mRNA levels before and 3 h after NH4+ treatment. UBQ1 was used as a control to normalize the expression data. Error bars represent ± SD of the means of three qPCR replicates. c GO enrichment of a total of 1386 genes of class I and II was analyzed with respect to the following three terms: biological process (i and ii), cellular component (iii and iv), and molecular function (v and vi). Each class was divided into two groups of upregulated and downregulated genes. GO terms that were enriched in class I genes but not in class II genes are indicated with dotted boxes. Further information about the genes is presented in Table S3
We used bioinformatics tools provided by bis.zju.edu.cn and pantherdb.org to perform GO enrichment analysis of 1386 NH4+-responsive genes identified in WT roots. GOs of class I ‘AMT1-dependent’ genes were compared with GOs of class II ‘AMT1-independent’ genes for the biological process [Fig. 7c(i) and (ii)], cellular component [Fig. 7c(iii) and (iv)], and molecular function [Fig. 7c(v) and (vi)]. The most interesting observation is that certain GOs in class I ‘AMT1-dependent’ genes are absent in class II ‘AMT1-independent’ genes. Otherwise, the distributions and frequencies of the remaining genes were similar in class I and class II in all three GO terms. In the biological process term, upregulated NH4+-responsive genes of class II are missing in the GO classes of metabolism, response to abiotic stimulus, response to endogenous stimulus, signal transduction, postembryonic development, and signal transduction [Fig. 7c(i)]. Similarly, among the downregulated genes of class II, GOs of macromolecular metabolism, protein metabolism/modification, and transport were missing [Fig. 7c(ii)]. In the cellular process term, the only mitochondrial group was absent in the GOs of class II upregulated genes [Fig. 7c(iii)], whereas three GOs (membrane, plasma membrane, and thylakoid) were absent in class II downregulated genes [Fig. 7c(iv)]. In the molecular function term, the following GOs were absent in class II upregulated genes: transferase activity, hydrolase activity, DNA binding, transcription factor activity, and transcription regulator activity [Fig. 7c(v)]. For class II downregulated genes, transferase activity, protein binding, nucleotide binding, transport activity, DNA binding, and kinase activity were absent [Fig. 7c(vi)]. The GO analysis clearly demonstrates that AMT1 activity profoundly impacts the expression of specific functional gene groups during the early stages of root response to NH4+.
Alteration of Auxin-Responsive Gene Expression and Root Gravity in amt1 Mutants
Ammonium affects root architecture, auxin transport, and gravity responses in roots (Zou et al. 2012; Liu and von Wiren 2017). The RNA-seq analysis recognized some auxin-related genes as ‘AMT1-dependent’ genes (Table S4). To further examine the relationship between AMT1 and those auxin-related genes, ammonium- and auxin-induction kinetics of these genes were compared by qPCR with RNAs of WT and amt1 roots treated with NH4+ or NAA. Sample seedlings were grown in the same way as those used for the RNA-seq experiments. Whole roots were harvested at 0, 1, 3, 6, 12, and 24 h after administration of NH4+ or NAA. qRT-PCR analyses were performed to determine expression kinetics induced by NH4+ or NAA. Comparisons of NH4+ and auxin-induction kinetics of some of auxin-related genes in WT and amt1 roots are presented in Fig. 8. The auxin-induction kinetics of these genes were dependent on AMT1 function. The expression kinetics of these genes showed similar patterns in both auxin- and NH4+-treated samples. For example, the expression of LOC_Os09g37330 became hypersensitive to both NH4+ and NAA in the mutant compared with that of WT (Fig. 8a, b). Other genes became less sensitive to both NH4+ and NAA in mutant roots than in WT roots (Fig. 8c–h). To further examine the involvement of AMT1 in interactions between auxin and NH4+, the gravity response was examined in WT and mutant roots. Three-day-old seedlings grown in dH2O were subjected to a 90° change in orientation with respect to gravity in two nutrient media containing 0.1 mM NH4+ or without NH4+. Root tip angles were recorded every 30 min. From 120 to 150 min after changing the gravity direction and in the presence of NH4+, root tips showed significantly wider angles of curvature in amt1 mutants than in WT plants (Fig. 9). However, without NH4+, the roots of both WT and mutants showed similar bending kinetics to the gravity change. The same experiment was performed with roots treated with NO3–, but NO3– did not alter the root gravity responses of mutants (Fig. S12). These results clearly demonstrated that amt1 mutation altered the gene expression and gravity response interactions between auxin and NH4+.
Expression kinetics of auxin-related genes that were induced by NH4+ and NAA. Total RNAs from roots of WT and two amt1 mutant lines (Ri 5–2 and Ri 3–1) were used for qRT-PCR analysis. The gene expression levels were examined at 0, 1, 3, 6, 12, and 24 h after addition of NH4+ or NAA. UBQ1 was used as a control to normalize the expression data. Error bars represent ± SD of the means of three qPCR replicates
Gravity responses of WT and amt1 roots in the presence (a) and absence (b) of NH4+. Two-day-old seedlings grown in water were transferred to a ¼ nutrient medium containing 0.1 mM NH4+ or no N source. The gravity direction was changed 90°, and bending angles of root tips were measured at various time points. The right graphs are magnifications of the dotted boxes in the left graphs. The experiments were repeated at least three times, and values represent means ± SE (n > 10). Significant differences in gravity responses in 0.05 mM (NH4)2SO4 solution are shown (*P < 0.05)
Discussion
The exogenous NH4+ supply profoundly impacts root system architecture (for review, Britto and Kronzucker 2002; Li et al. 2010; Liu et al. 2013; Araya et al. 2016; Liu et al. 2017). In this study, amt1 mutants were analyzed to estimate the role of AMT1 in NH4+-mediated root development. The main objective of this study was to explore prominent phenotypes of amt1 roots, which could elucidate possible interactions between AMT1 and N-dependent root development.
This study showed that NH4+ or NAA/NOA treatments specifically inhibited lateral root growth in amt1 mutants. In mutant root cells with low AMT1 activity, NH4+ dramatically changed apoplastic pH and membrane polarization (Husted and Schjoerring 1995). The combined action of NAA and NOA is expected to disrupt intracellular membrane trafficking and apoplasmic accumulation of auxin (Lankova et al. 2010; Imhoff et al. 2000). Under these disturbances of cellular membrane processes, amt1 failed to maintain normal development of lateral roots. Therefore, our work strongly suggests the possibility that AMT1 might have a role in lateral root growth by supporting membrane dynamics and integrity.
The most significant finding of this study is the observation that amt1 mutants exhibit NH4+-induced suppression of NO3–-dependent lateral root elongation. NAA/NOA do not affect NO3–-dependent lateral root growth. This indicates that NO3–-dependent cellular processes for lateral root development are not affected by the disruption of membrane dynamics. These data suggest that the mechanism of NH4+ inhibition of lateral root development in amt1 mutants is different from that of NAA/NOA. As amt1 mutants accumulated lower levels of cellular NH4+ than WT plants, it is very unlikely that intracellular toxicity of NH4+ is related to the suppression of lateral root growth. One might suspect that NH4+ could interfere with NO3– uptake, which subsequently results in retardation of lateral root elongation. It has been reported that NH4+ can inhibit NO3– uptake in rice, barley, and Arabidopsis (Kronzucker et al. 1999a, b; Cerezo et al. 2001). However, in barley and Arabidopsis, inhibition of total NO3– uptake by NH4+ was significant under low NO3– conditions, suggesting that the high-affinity transport system (HATS) is involved. The following observations strongly suggest that NH4+ triggers indirect and systemic signaling, rather than directly interfering with NO3– uptake, in inhibiting NO3–-mediated lateral root growth. (1) NH4+-induced inhibition of lateral root development (especially lateral root numbers) is independent of the NO3– concentrations applied to mutant roots. (2) NH4+ can systemically prevent lateral root development. (3) NH4+ does not affect NO3–-dependent crown root development. Depending on the NO3– concentration, different NO3– signaling pathways have different effects on the development of lateral roots (for review, Sun et al. 2017). In general, low NO3– exerts stimulatory or inhibitory effects on lateral root development, whereas high NO3– supply has an inhibitory effect on lateral root development. In our study, there were no preferential ranges of NO3– concentrations at which NH4+ most effectively inhibited lateral root development (especially lateral root numbers). There was no difference in growth defect severity among roots exposed to a wide range (from 0.1 mM to 10 mM) of NO3–. Glutamine contents were similar in mutant roots grown in either NO3–, or NH4NO3, suggesting that the inhibition of lateral root development should not be due to an N nutrient effect that results from inefficient NO3– uptake. NH4+ can exert systemic action without direct contact with lateral roots. Our data strongly suggest that NH4+ can trigger a secondary cellular messenger that prevents NO3–-mediated lateral root elongation. This study supports the notion that N signaling for lateral root elongation is interactive between NH4+ and NO3–.
It has been reported that more than 2,000 genes have been identified in rice root in response to NH4+ treatment within 3 h (Xuan et al. 2013). Our RNA-seq analysis found that AMT1 function is required for the early response to NH4+ by half of the NH4+-responsive genes. GO enrichment analysis showed that specific functional gene groups required AMT1 function for the early response to NH4+ treatment. This work showed that AMT1 activity was necessary for auxin induction of some NH4+-responsive genes. The requirement of AMT1 function for the expression of these genes can be explained in three ways. (1) These genes are sensitive to differences in NH4+-dependent acidification of apoplasts between amt1 roots and WT roots (Patterson et al. 2010). (2) NH4+ assimilation and subsequent metabolite production could be essential processes for the expression of these genes. (3) AMT1 might act as a sensor to regulate the expression of these genes in the presence of NH4+. Further work is required to distinguish these possibilities. This study also showed that amt1 became hypersensitive to gravity in the presence of NH4+. It has been reported that excess NH4+ reduces the gravity response of roots by affecting auxin distribution and the activity of a potassium transporter in Arabidopsis (Zou et al. 2012). The gravity response of rice root tips is delayed in the presence of NH4+ (Xuan et al. 2018). Compared with WT, amt1 showed a hypersensitive gravity response in the presence of NH4+. These data suggest that AMT1 is involved in NH4+-triggered gravity response in roots.
Currently, little is known about cellular chemicals or messengers that can convey signals triggered or induced by NH4+ or AMT1 for root growth or gene expression. Our data on NH4+-triggered phenotypes of amt1 mutants might provide a basic platform to identify these signals and to explore molecular or cellular mechanisms underlying the interactions between NH4+ and AMT1 in root development.
References
Abiko T, Obara M, Ushioda A, Hayakawa T, Hodges M, Yamaya T (2005) Localization of NAD-isocitrate dehydrogenase and glutamate dehydrogenase in rice roots: candidates for providing carbon skeletons to NADH-glutamate synthase. Plant Cell Physiol 46:1724–1734
Araya T, Kubo T, von Wirén N, Takahashi H (2016) Statistical modeling of nitrogen-dependent modulation of root system architecture in Arabidopsis thaliana. J Integr Plant Biol 58:254–265
Bao AZ, Liang Z, Zhao CH (2015) Overexpressing of OsAMT1-3, a high affinity ammonium transporter gene, modifies rice growth and carbon-nitrogen metabolic status. Int J Mol Sci 16(5):9037–9063
Barth C, Gouzd ZA, Steele HP, Imperio RM (2010) A mutation in GDP-mannose pyrophosphorylase causes conditional hypersensitivity to ammonium, resulting in Arabidopsis root growth inhibition, altered ammonium metabolism, and hormone homeostasis. J Exp Bot 61(2):379–394
Bouguyon E, Perrine-Walker F, Pervent M, Rochette J, Cuesta C, Benkova E, Nacry P (2016) Nitrate controls root development through posttranscriptional regulation of the NRT1.1/NPF6.3 transporter/sensor. Plant Physiol 172(2):1237–1248. https://doi.org/10.1104/pp.16.01047
Brady SM, Burow M, Busch W, Carlborg Ö, Denby KJ, Glazebrook J, Hamilton ES, Harmer SL, Haswell ES, Maloof JN (2015) Reassess the t test: interact with all your data via ANOVA. Plant Cell 27:2088–2094
Britto DT, Kronzucker HJ (2002) NH4+ toxicity in higher plants: a critical review. J Plant Physiol 159:567–584
Britto DT, Siddiqi MY, Glass AD, Kronzucker HJ (2001) Futile transmembrane NH4+ cycling: a cellular hypothesis to explain ammonium toxicity in plants. Proc Natl Acad Sci USA 98(7):4255–4258
Cao Y, Glass AD, Crawford NM (1993) Ammonium inhibition of Arabidopsis root growth can be reversed by potassium and by auxin resistance mutations aux1, axr1, and axr2. Plant Physiol 102:983–989
Cerezo M, Tillard P, Gojon A, Primo-Millo E, Garcia-Agustin P (2001) Characterization and regulation of ammonium transport systems in Citrus plants. Planta 214:97–105
Chaudhuri B, Hormann F, Lalonde S, Brady SM, Orlando DA, Benfey P, Frommer WB (2008) Protonophore- and pH-insensitive glucose and sucrose accumulation detected by FRET nanosensors in Arabidopsis root tips. Plant J 56:948–962
Chen RF, Shen RF, Gu P, Dong XY, Du CW, Ma JF (2006) Response of rice (Oryza sativa) with root surface iron plaque under aluminium stress. Ann Bot 98(2):389–395
Chin HC, Choe MS, Lee SH, Park SH, Park SH, Koo JC, Kim NY, Lee JJ, Oh BG, Yi GH, Kim SC, Choi HC, Cho MJ, Han CD (1999) Molecular analysis of rice plants harboring an Ac/Ds transposable element-mediated gene trapping system. Plant J 19(5):615–623
Fernández-Crespo E, Scalschi L, Llorens E, García-Agustín P, Camañes G (2015) NH4+ protects tomato plants against Pseudomonas syringae by activation of systemic acquired acclimation. J Exp Bot 66:6777–6790
Gaur VS, Singh US, Gupta AK, Kumar A (2012) Understanding the differential nitrogen sensing mechanism in rice genotypes through expression analysis of high and low affinity ammonium transporter genes. Mol Biol Rep 39:2233–2241. https://doi.org/10.1007/s11033-011-0972-2
Hirano T, Satoh Y, Ohki A, Takada R, Arai T, Michiyama H (2008) Inhibition of ammonium assimilation restores elongation of seminal rice roots repressed by high levels of exogenous ammonium. Physiol Plant 134:183–190
Husted S, Schjoerring JK (1995) Apoplastic pH and ammonium concentration in leaves of Brassica napus L. Plant Physiol 109(4):1453–1460
Imhoff V, Muller P, Guern J, Delbarre A (2000) Inhibitors of the carrier-mediated influx of auxin in suspension-cultured tobacco cells. Planta 210:580–588
Kim EJ, Kim YJ, Hong WJ, Jeon JS, Jung KH (2019) Genome-wide analysis of root hair preferred RBOH genes suggests that three RBOH genes are associated with auxin-mediated root hair development in rice. J Plant Biol 62:229–238
Kronzucker HJ, Siddiqi MY, Glass ADM, Kirk GJD (1999a) Nitrate ammonium synergism in rice: a subcellular analysis. Plant Physiol 119:1041–1046
Kronzucker HJ, Glass AD, Yaeesh Siddiqi M (1999b) Inhibition of nitrate uptake by ammonium in barley. Analysis component fluxes. Plant Physiol 120(1):283–292. https://doi.org/10.1104/pp.120.1.283
Lankova M, Smith R, Pesek B, Kubes M, Zazimalova E, Petrasek J (2010) Auxin influx inhibitors 1-NOA, 2-NOA and CHPAA interfere with membrane dynamics in tobacco cells. J Exp Bot 61:3589–3598. https://doi.org/10.1093/jxb/erq172
Li Q, Li BH, Kronzucker HJ, Shi WM (2010) Root growth inhibition by NH4+ in Arabidopsis is mediated by the root tip and is linked to NH4+ efflux and GMPase activity. Plant Cell Environ 33:1529–1542
Li B, Li Q, Su Y, Chen H, Xiong L, Mi G, Kronzucker HJ, Shi W (2011) Shoot-supplied ammonium targets the root auxin influx carrier AUX1 and inhibits lateral root emergence in Arabidopsis. Plant Cell Environ 34:933–946
Li B, Li Q, Xiong L, Kronzucker HJ, Kramer U, Shi W (2012) Arabidopsis plastid AMOS1/EGY1 integrates abscisic acid signaling to regulate global gene expression response to ammonium stress. Plant Physiol 160:2040–2051
Li G, Li B, Dong G, Feng X, Kronzucker HJ, Shi W (2013) Ammonium-induced shoot ethylene production is associated with the inhibition of lateral root formation in Arabidopsis. J Exp Bot 64:1413–1425
Lima JE, Kojima S, Takahashi H, von Wirén N (2010) Ammonium triggers lateral root branching in Arabidopsis in an AMMONIUM TRANSPORTER1;3-dependent manner. Plant Cell 22:3621–3633
Little DY, Rao H, Oliva S, Daniel-Vedele F, Krapp A, Malamy JE (2005) The putative high-affinity nitrate transporter NRT2.1 represses lateral root initiation in response to nutritional cues. Proc Natl Acad Sci USA 102:13693–13698
Liu Y, von Wiren N (2017) Ammonium as a signal for physiological and morphological responses in plants. J Exp Bot 68(10):2581–2592
Liu L, Mei Q, Yu Z, Sun T, Zhang Z, Chen M (2013) An integrative bioinformatics framework for genome-scale multiple level network reconstruction of rice. J Integr Bioinf 10(2):223
Loque D, von Wiren N (2004) Regulatory levels for the transport of ammonium in plant roots. J Exp Bot 55(401):1293–1305
Moon S, Chandran AKN, Kim YJ, Gho Y, Hong WJ, An G, Lee C, Jung KH (2019) Rice RHC encoding a putative cellulase is essential for normal root hair elongation. J Plant Biol 62:82–91
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Plant Physiol 15:473–497
Oliveira IC, Brears T, Knight TJ, Clark A, Coruzzi GM (2002) Overexpression of cytosolic glutamine synthetase. relation to nitrogen, light, and photorespiration. Plant Physiol 129(3):1170–1180.
Patterson K, Cakmak T, Cooper A, Lager I, Rasmusson AG, Escobar MA (2010) Distinct signalling pathways and transcriptome response signatures differentiate ammonium- and nitrate-supplied plants. Plant Cell Environ 33:1486–1501
Ranathunge KA, El-Kereamy S, Gidda Y, Bi M, Rothstein SJ (2014) AMT1;1 transgenic rice plants with enhanced NH4+ permeability show superior growth and higher yield under optimal and suboptimal NH4+ conditions. J Exp Bot 65(4):965–979
Remans T, Nacry P, Pervent M, Filleur S, Diatloff E, Mounier E et al (2006) The Arabidopsis NRT1.1 transporter participates in the signaling pathway triggering root colonization of nitrate-rich patches. Proc Natl Acad Sci USA 103:19206–19211
Shimizu H, Tanabata T, Xie X, Inagaki N, Takano M, Shinomura T, Yamamoto KT (2009) Phytochrome-mediated growth inhibition of seminal roots in rice seedlings. Physiol Plantarum 137:289–297
Sonoda Y, Ikeda A, Saiki S, von Wiren N, Yamaya T, Yamaguchi J (2003) Distinct expression and function of three ammonium transporter genes (OsAMT1;1–1;3) in rice. Plant Cell Physiol 44(7):726–734
Suenaga A, Moriya K, Sonoda IA, Von Wiren N, Hayakawa T, Yamaguchi J, Yamaya T (2003) Constitutive expression of a novel-type ammonium transporter OsAMT2 in rice plants. Plant Cell Physiol 44(2):206–211
Sun CH, Yu JQ, Hu DG (2017) A crucial signal during lateral roots development. Front Plant Sci 8:448
Wang MY, Glass ADM, Shaff JE, Kochian LV (1994) Ammonium uptake by rice roots (III. Electrophysiology). Plant Physiol 104(3):899–906
Xuan YH, Priatama RA, Huang J, Je BI, Liu JM, Park SJ, Piao HL, Son DY, Lee JJ, Park SH, Jung KH, Kim TH, Han CD (2013) Indeterminate domain 10 regulates ammonium-mediated gene expression in rice roots. New Phytol 197:791–804
Xuan YH, Duan FY, Je BI, Kim CM, Li TY, Liu JM, Park SJ, Cho JH, Kim TH, von Wiren N, Han CD (2017) Related to ABI3/VP1-Like 1 (RAVL1) regulates brassinosteroid-mediated activation of AMT1;2 in rice (Oryza sativa). J Exp Bot 68(3):727–737
Xuan YH, Kumar V, Zhu XF et al (2018) IDD10 is involved in the interaction between NH4+ and auxin signaling in rice roots. J Plant Biol 61:72–79
Xuan YH, Kumar V, Han X, Kim SH, JeongJH KCM, Gao Y, Han CD (2019) CBL-INTERACTING PROTEIN KINASE 9 regulates ammonium-dependent root growth downstream of IDD10 in rice (Oryza sativa). Ann Bot 124:947–960
Zou N, Li B, Dong G, Kronzucker HJ, Shi W (2012) Ammonium induced loss of root gravitropism is related to auxin distribution and TRH1 function, and is uncoupled from the inhibition of root elongation in Arabidopsis. J Exp Bot 63:3777–3788
Zou N, Li B, Chen H, Su Y, Kronzucker HJ, Xiong L, Baluska F, Shi W (2013) GSA-1/ARG1 protects root gravitropism in Arabidopsis under ammonium stress. New Phytol 200:97
Acknowledgements
This research was supported by grants from the Next-Generation BioGreen 21 Program (PJ01326601), the Rural Development Administration, Republic of Korea, and from Support Plan for Innovative Talents in Colleges and Universities of Liaoning Province [LR2017037].
Author information
Authors and Affiliations
Contributions
CDH and YHX experimental design, data analysis and interpretation, manuscript editing. VK and SHK data generation and analysis, image analysis, data presentation, manuscript writing. RAP, JHJ, MRA, BAS material generation and analysis, DNA extraction, qPCR analysis. KHJ bioinformatics analysis of RNA-seq data. CMK, BIJ, and SJP data analysis and interpretation. KMK material propagation.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kumar, V., Kim, S.H., Priatama, R.A. et al. NH4+ Suppresses NO3–-Dependent Lateral Root Growth and Alters Gene Expression and Gravity Response in OsAMT1 RNAi Mutants of Rice (Oryza sativa). J. Plant Biol. 63, 391–407 (2020). https://doi.org/10.1007/s12374-020-09263-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12374-020-09263-5