Abstract
Two rice genotypes, Kalanamak 3119 (KN3119) and Pusa Basmati 1(PB1) differing in their optimum nitrogen requirements (30 and 120 kg/ha, respectively) were undertaken to study the expression of both high and low affinity ammonium transporter genes responsible for ammonium uptake. Exposing the roots of the seedlings of both the genotypes to increasing (NH4)2SO4 concentrations revealed that all the three families of rice AMT genes are expressed, some of which get altered in a genotype and concentration specific manner. This indicates that individual ammonium transporter genes have defined contributions for ammonium uptake and plant growth. Interestingly, in response to increasing nitrogen concentrations, a root specific high affinity gene, AMT1;3, was repressed in the roots of KN3119 but not in PB1 indicating the existence of a differential ammonium sensing mechanism. This also indicates that not only AMT1;3 is involved not only in ammonium uptake but may also in ammonium sensing. Further, if it can differentiate and could be used as a biomarker for nitrogen responsiveness. Expression analysis of low affinity AMT genes showed that, both AMT2;1 and AMT2;2 have high levels of expression in both roots and shoots and in KN3119 are induced at low ammonium concentrations. Expressions of AMT3 family genes were higher shoots than in the roots indicating that these genes are probably involved in the translocation and distribution of ammonium ions in leaves. The expression of the only high affinity AMT gene, AMT1;1, along with six low affinity AMT genes in the shoots suggests that low affinity AMTs in the shoots leaves are involved in supporting AMT1;1 to carry out its activities/function efficiently.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Nitrogen is quantitatively the most essential nutrient for plants and thus often a major limiting factor in plant production. The major form of nitrogen that is available for growth of rice plants in paddy fields is NH4 +, requiring NH4 + transport systems at the root plasma membrane. The first step in nitrogen assimilation is the uptake of nitrate and ammonium into root cells from the soil solution. Since ammonium assimilation requires less energy than that of nitrate [1], ammonium is the preferential form of nitrogen uptake when plants are subjected to nitrogen deficiency [2]. However, excessive ammonium uptake into plants can be toxic [3, 4]. Therefore, ammonium uptake and metabolism in plants must be tightly regulated. Ammonium uptake from the soil solution is mediated by ammonium transporters that have been isolated and partially characterized in several plant species.
AMT in rice roots is comprised of four families OsAMT1, OsAMT2, OsAMT3 and OsAMT4. Except for the family OsAMT4 which contain only one member i.e., OsAMT4;1, each of the three families contain three members viz., OsAMT1 (OsAMT1;1, OsAMT1;2 and OsAMT1;3), OsAMT2 (OsAMT2;1, OsAMT2;2 and OsAMT2;3) and OsAMT3 (OsAMT3;1, OsAMT3;2 and OsAMT3;3). The members of OsAMT1s share high sequence similarity to each other and are very dissimilar to the other three OsAMT families and have been characterized as high-affinity transport system (HAT). The other three families have been characterized as low affinity transport system (LAT) [5, 6]. At low NH4 + concentration, uptake is mediated by HATs and exhibits sensitivity to metabolic inhibitors while at high NH4 + concentration (between 1 and 40 mM), uptake is mediated by LATs and is less responsive to metabolic inhibitors [7]. The presence of similar ammonium transporters have also been reported and partially characterized in several plant species, such as Arabidopsis thaliana (AtAMT1;1, AtAMT1;2; AtAMT1;3 and AtAMT2; [2, 8–10] Brassica napus (BnAMT1;2) [11], Lotus japonicas (LjAMT1;1, LJAMT2;1) [12, 13] and Lycopersicon esculentum (LeAMT1;1, LeAMT1;2 and LeAMT1;3) [14–16]. Despite of the knowledge that rice is an ammonium loving plant whose most of the growth period is under flooded conditions in which ammonium ions are the dominant nitrogen species and further large amount of nitrogenous fertilizers are added to the rice fields in order to achieve higher yields, little work is available in the literature till date describing work on the regulation of rice ammonium transporters.
In this study, NH4 + uptake of two rice genotype differing in their optimum nitrogen requirement were investigated by studying the expressions of both high and low affinity ammonium transporters OsAMT1s (HATs-OsAMT1;1-1;3, LATs-AMT2;1-2;3 and AMT3;1-3;3) under different nitrogen conditions to understand the mechanisms associated with differential nitrogen sensing.
Materials and methods
Selection and growth of rice genotypes
Two rice genotypes Pusa Basmati 1 and Kalanamak 3119 were chosen based on their differential response to nitrogen. Kalanamak 3119 is a relatively long (121 cm height), long duration (154 days for maturity) and low average yield (0.8 t/ha) genotype having optimum nitrogen dose of 30 kg/ha. Whereas the Pusa Basmati1 has a relatively shorter height (90–110 cm) and duration (130–135 days) but higher average yield (4.5 t/ha) having optimum nitrogen dose of 120 kg/ha [17]. Seeds of both the genotypes were sterilized in 1% [v/v] NaClO for 20 min and thoroughly rinsed in sterile distilled water, grown hydroponically in tap water for a week, and then seedlings were transferred to nutrient solution (1 mg l−1Na2HPO4·12H2O, 1.19 mg l−1KCl, 1.05 mg l−1 CaCl2·2H2O, 3.05 mg l−1 MgCl2·6H2O, 9.90 mg l−1 FeSO4·7H2O, 2.02 mg l−1 Mn FeSO4·4H2O, 2.86 mg l−1 H3BO4, 0.87 mg l−1 ZnSO4·7H2O, 0.039 mg l−1 CuSO4·7H2O, 0.012 mg l−1 Na2MoO4·2H2O) [5] which do not contain any source of nitrogen for 2 weeks. Plants were then transferred to a nitrogen-free nutrient solution, or to solutions containing increasing concentrations of (NH4)2SO4 as a sole nitrogen source. Five concentrations were used viz., 0.15, 1.5, 15, 150 μM and 1 mM. Plants were grown under light for 2 h, at 60% relative humidity, and a temperature of 30°C.
Selection of stages for RNA isolation and cDNA synthesis
Total RNA was isolated after 2 h from the roots and shoots of 25 days old nitrogen deprived seedlings of both the plants growing under increasing (NH4)2SO4 concentrations. Total RNA was isolated using Concert Plant RNA purification reagent (Invitrogen) followed by on column DNA digestion using RNeasy plant minikit and RNase free DNA digestion kit (Qiagen, Germany). The quality of RNA was checked by running on 1% agarose gel and was quantified using a spectrophotometer. Minimum of three plant replicates were selected for RNA isolation. The quantified individual total RNA replicates were further used to prepare cDNA. 2 μg of total RNA was used to prepare cDNA. The reaction condition was as follows, for 20 μl reaction, 2 μg total RNA, 1 μl (200 U) MMLV reverse transcriptase (Invitrogen), 5× (4 μl) first strand buffer, 2 μl DTT, 0.4 μl dNTP mix (100 mM), 1 μl Superase RNase inhibitor (Ambion) (20 U/μl) and 0.5 μg oligo dT primer (Qiagen) for 60 min at 37°C and a final denaturation step at 70°C for 15 min.
Primer designing
mRNA sequences of all the members of rice AMT gene family were downloaded from NCBI and their gene specific primers were designed using Lasergene DNASTAR software package. The list of primers used to study the expression profiles of rice AMT genes is given in the Table 1 .
Semi-quantitative RT-PCR and cloning of RT-PCR products
RT-PCR using individual cDNAs sets obtained from each of the three replicates was performed for generating the expression profiles of members of AMT genes. Before generating the expression profiles for densitometry analysis, the PCR was optimized by analyzing the PCR products of 10, 20, 30, 35 and 40 cycles. Thirty PCR cycles was found to be optimum in order to avoid plateau of PCR reaction cycles. RT-PCR reaction was performed using aliquots of 1 μl of the cDNA and 12.5 pmol of gene specific primer in a 50 μl reaction volume containing 0.2 mM of each dNTPs, 2 mM MgCl2 and 1 U GoTaq Flexi DNA polymerase (Promega) which is provided with a green buffer containing gel loading dye so that the PCR products can be loaded directly onto the gels. The temperature profiles used for the PCR were 95°C for 2 min initial denaturation followed by 30 cycles of 95°C for 20 s, 58°C for 30 s (primer anneling), 72°C for 30 s and final extension for 10 min. Rice actin primer was taken as internal control. The PCR products were separated on 1% agarose gels and the single specific band of PCR product obtained was cloned into the pGEM-T easy vector (Promega) for sequencing. The nucleotide sequencing analysis revealed almost 98–99% homology of OsAMTs genes with the preexisting sequences available in the NCBI database.
Real-time PCR
Real-time PCR was done using the 5 Prime Real Master Mix SYBR ROX (Eppendorf India Limited, Chennai, India) according to manufacturer’s instructions. The thermocycler used was eppendorf thermocycler eprealplex. Two-step real-time PCR was carried out using cDNA prepared as mentioned earlier from different developmental stages. The primers for AMT genes and actin genes used were same as earlier described. The reverse transcription efficiencies of AMT and actin genes were almost equal as analyzed by comparing the CT values at different dilutions of cDNA [18]. The amplification program used was: 95°C for 2 min, 40 cycles at 95°C for 30 s, 60°C for 30 s, 72°C for 30 s; 60°C for 15 s and 95°C for 15 s. All samples were amplified in triplicate, and the mean value was considered.
Statistical analysis
Three independent determinations for each parameter were recorded and mean ± SE values were calculated for statistical analysis. CRD was used for analyzing the gel real time PCR data. For comparing two data sets paired ‘t’ test was used.
Results
Expression profiling of members of ammonium transporter genes in roots of PB1 and KN3119
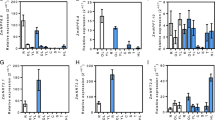
In both the genotypes, the expression of OsAMT1;1 and OsAMT1;2 in the roots was high. However, higher expression was observed in case of KN3119. The real time data showed that there was a slight increase in the expression of AMT1;1 gene in both the genotypes. In case of AMT1;2 in PB1, maximum expression (2.94 fold increase) was observed at 150 μM whereas in KN3119 maximum expression (1.8 fold increase) was observed at 0.15 μM when compared to the control (Figs. 1, 2). Similarly, the expression of OsAMT1;3 was also high in PB1 (Figs. 1, 3) but there was no significant change (P > 0.05) in expression across different nitrogen concentrations. However, in case of KN3119 its expression (Figs. 1, 4) was high at lower nitrogen concentrations that declined sharply beyond 15 μM and at 1 mM only 0.37 of the control was detected (Fig. 4). The expression of the gene OsAMT2;1 was also differential. In PB1 highest expression was detected at the control and 0.15 μM (NH4)2SO4 which declined continuously as the (NH4)2SO4 concentration was increased. In KN3119 the OsAMT2;1 transcripts significantly increased (P < 0.05) (6.68 fold over the control) at 0.15 μM (NH4)2SO4 and reached a constant level (around 3.84 fold over the control) and then declined to a lowest level at 1 mM (NH4)2SO4 (Figs. 1, 4). OsAMT2;2 was highly expressed in the roots of both the genotypes. In PB1 no significant change (P > 0.05) in its expression was detected across different (NH4)2SO4 concentrations whereas in KN3119, its expression sharply increased (10.28 fold as compared to the control) at 0.15 μM but significantly declined (P < 0.05) when the concentration was increased beyond 15 μM. The expression profiles of the gene OsAMT2;3 showed that it is weakly expressed in the roots. Transcripts from this gene could not be detected in the control, however, its expression was significantly induced when nitrogen source was added into the medium. In both the genotypes, OsAMT2;3 transcripts were induced at 0.15 μM (NH4)2SO4 (8.57 fold in PB1 and 6.63 fold in KN3119 as compared to the control) concentration but declined as the concentration was raised beyond 0.15 μM (Figs. 1, 4). A continuous decrease in the expression was observed from 0.15 to 1 mM. In KN3119 the repression of the gene was such that no transcripts could be detected at 150 μM and 1 mM (NH4)2SO4 concentration. In both the genotypes OsAMT3;1 expression was found to decrease with the increase in (NH4)2SO4 concentration. In PB1, the gene was found to express at higher rates than KN3119. In both the genotypes highest level of expression was detected in the controls which declined with increase in concentration. In PB1 the OsAMT3;2 gene was found to be induced at 0.15 μM and increased steadily as the (NH4)2SO4 concentration was increased. Maximum expression (1.13 fold as compared to the controls) of this gene in PB1 was therefore detected at 1 mM (NH4)2SO4 concentration. Unlike PB1 the gene OsAMT3;2 was not induced in KN3119 roots even at higher (NH4)2SO4 concentrations but maintained a low and constant level of expression. OsAMT3;3 was induced in both the genotypes with increase in nitrogen concentration. Induction of 1.4 fold in PB1 and 3.7 fold in KN3119 at 15 and 150 μM, respectively was observed and further increase in the (NH4)2SO4 concentration only slightly decreased the expression.
Expression profiles of high and low affinity ammonium transporter genes in the roots of Pusa Basmati1 (PB1) and Kalanamak 3119 (KN3119). Total RNA was isolated after 2 h from roots of 25 days old nitrogen deprived seedlings of PB1 and KN3119 exposed to increasing (NH4)2SO4 concentration. 2 μg of total RNA was use to prepare cDNA and expression of individual genes were detected using gene specific primers. Lanes: A control, B 0.15 μM, C 1.5 μM, D 15 μM, E 150 μM and F 1 mM
Expression profiling of ammonium transporter genes in the shoots of 25 days old rice seedling of PB1 and KN3119
As reported earlier and confirmed in the present study, unlike roots the genes OsAMT1;2 and OsAMT1;3 do not express in the leaves of the 25 days old seedling. While the gene OsAMT1;1 was strongly expressed in the shoot leaves of both the genotypes and was induced by the addition of (NH4)2SO4 in the medium (Fig. 5). In PB1 OsAMT1;1 transcript was significantly (P < 0.05) induced (2.49 fold increase) at 1.5 μM (NH4)2SO4 concentration which further increased with higher (NH4)2SO4 concentration (11.79 fold at 150 μM (NH4)2SO4). In KN3119 significant induction (P < 0.05) of OsAMT1;1 was also observed but at a higher (NH4)2SO4 concentration than PB1 i.e., at 150 μM (NH4)2SO4 (5.77 fold). Unlike the OsAMT1 family, transcripts all the members of the OsAMT2 family i.e., OsAMT2;1, OsAMT2;2 and OsAMT2;3 were found to be expressed in the shoot leaves of both the genotypes. In PB1, OsAMT2;1 was induced (2.78 fold increase) at 0.15 μM (NH4)2SO4 and maximum expression was detected at 1 mM (NH4)2SO4 concentration (6.21 folds as compared to the control) (Figs. 5, 3). In contrast to PB1 maximum transcript levels of OsAMT2;1 in KN3119 was detected in the control which declined slowly as the (NH4)2SO4 concentration was increased to 1 mM. OsAMT2;2 was found to be differentially expressed. In PB1 the expression of OsAMT2;2 was significantly induced (P < 0.05) (4.03 fold increase) at 1.5 μM (NH4)2SO4 concentration which did not change with increase in (NH4)2SO4 concentration. However, in KN3119 the transcript levels was high in the control which steadily decreased as the (NH4)2SO4 concentration was increased to 1 mM (Figs. 5, 6). The gene OsAMT2;3 was found to be weakly expressed in the shoots of both the genotypes studied. In PB1 its expression in gel could only be detected at 1.5 μM (NH4)2SO4 concentration (which correspond to 4.62 fold increase) which declined when the (NH4)2SO4 concentration was raised above 150 μM. In KN3119 the expression of OsAMT2;3 was slightly induced at 0.15–150 (NH4)2SO4 concentration and similar to PB1 also declined at 1 mM (NH4)2SO4 concentration (Figs. 5, 6). Similar to OsAMT2 gene family, all the members of OsAMT3 family i.e., OsAMT3;1, OsAMT3;2 and OsAMT3;3 were found to express in the shoots leaves of both the genotypes and were affected by varying nitrogen concentrations. In PB1 OsAMT3;1 was strongly induced at 1.5–150 μM (NH4)2SO4 concentration (14.68 fold increase in expression at 150 (NH4)2SO4) but was weakly induced at 1 mM (NH4)2SO4 concentration. In KN3119 no significant change in expression was detected up to 150 μM (NH4)2SO4 concentration but similar to PB1 declined at 1 mM. Expression of OsAMT3;2 was induced in both the genotypes but at varying concentrations of (NH4)2SO4. In PB1, OsAMT3;2 was induced (4.01 fold as compared to the control) at 1.5 μM (NH4)2SO4 while in KN3119 it was induced at 0.15 μM. In both the genotypes further increase in (NH4)2SO4 concentration slightly increased the expression. Similar to OsAMT3;2 the expression of OsAMT3;3 was also induced in the shoot leaves of both the genotypes. Like OsAMT3;2, OsAMT3;3 in PB1 was also induced (5.79 fold as compared to the control) at 1.5 μM (NH4)2SO4 but in KN3119 it was induced (6.06 fold as compared to the control) at 15 μM and further increase in (NH4)2SO4 concentration slightly increased the expression in PB1 but not in KN3119.
Discussion
In the present investigation, attempts were undertaken to study the expressions of genes involved nitrogen uptake in two rice genotypes differing in their level of optimum nitrogen requirement. The optimum nitrogen requirement of these two rice genotypes has been experimentally determined that KN3119 is a low nitrogen requiring non basmati scented rice genotype having optimum nitrogen requirement of 30 kg/ha while the optimum nitrogen requirement of PB1 is 120 kg/ha. On increasing the nitrogen dose of KN3119 there was a substantial reduction in the yield thereby making the genotype fit for organic farming [17]. The difference of optimum nitrogen requirement in these two rice genotypes provides excellent plant materials to study nitrogen responsiveness in rice genotypes. To further understand the molecular basis of responsiveness of these two contrasting rice genotypes to different nitrogen inputs at the nitrogen uptake level, a laboratory experiment was conducted to study how members of ammonium transporter gene family responds to different nitrogen conditions. Semiquantitative RT-PCR was initially used to study the expression of patterns of rice OsAMT genes and was further confirmed by quantitative RT-PCR.
Low and high affinity OsAMT genes are differentially expressed in the roots and shoots
In a similar experiment using the rice genotype ‘Nipponbare’, workers reported constitutive expression in shoots and roots for OsAMT1;1, root-specific and ammonium-inducible expression for OsAMT1;2 and root specific and NH4 +-derepressed expression for OsAMT1;3 [5]. In the present investigation similar results were found with OsAMT1;1 in roots where it was found to be constitutive and was induced in both roots and shoots leaves of both the genotypes which was however seen at different nitrogen conditions. This indicates that the nitrogen in the soil solution is being sensed and its transportation starts within 2 h of application of nitrogen fertilizers reaching to soil solution. This induction of OsAMT1;1 in the shoots is may be due to the transport of NH4 + ions from the roots to the shoots. Although, it has been known that NO3 − enters in the xylem, there had been some debate in the past as to whether NH4 + is also translocated in the xylem. Loque and von Wiren [6] using positron emitting tracer imaging found 13N from root-applied 13N labelled ammonium in the shoot tissue within less than 2 min suggesting that root-to-shoot translocation of ammonium or of ammonium-derived nitrogen is rapid. Also recently, another group reported that the xylem sap NH4 + concentrations increased markedly 1 day after nitrogen application further suggesting direct transportation of NH4 + from the rice roots to the above-ground parts [19]. These results indicate the possible involvement of the ammonium transporters in uptake/downloading of NH4 + from the xylem sap into the leaf tissues similar to roots for further assimilation through the cytosolic GS1/GOGAT pathway [6, 20, 21]. Interestingly, in PB1 this induction was observed even at the lowest NH4 + concentration supplied (0.15 μM) compared to KN3119 where the induction was observed at a higher NH4 + concentration. However, the high affinity OsAMT1;2 did not showed NH4 + inducibility in the roots of both the genotypes but was found to be expressing higher in the roots of KN3119 compared to PB1 indicating that ammonium uptake at the root level through this transporter is higher in KN3119.
In contrast OsAMT1;1 and OsAMT1;2 the expression of OsAMT1;3 was differential among the two genotypes studied. Expression of OsAMT1;3 in rice roots has been reported to be high under nitrogen limiting conditions that repress when the nitrogen concentration of the medium increase [5]. In the present study similar repression of OsAMT1;3 was observed when KN3119 roots were exposed to increasing nitrogen concentration but in contrast was unchanged in PB1. This clearly indicates that the repression of OsAMT1;3 with increase in nitrogen input may not be a universal mechanism but is dependent on genotype and probably the level of nitrogen requirement or probably much higher concentration of NH4 + is required i.e., higher than 1 mM to repress OsAMT1;3 gene in PB1 roots. Interestingly, the rice genotype ‘Nipponbare’ is a low nitrogen requiring cultivar like KN3119 used in the present study [5, 19]. If in general the expression of OsAMT1;3 distinguishes rice genotypes requiring low and high doses of nitrogen it could be used as an biomarker in determining the optimum nitrogen doses of rice varieties. The subsequent prove to validate OsAMT1;3 as biomarker for optimization of N-dose can be obtained by screening large number of genotypes requiring different N-doses and also analyzing the mapping populations derived from the hybridizations resulting from low and high nitrogen responsive genotypes. These differences in induction of high affinity AMT genes are may be due to the differences in nitrogen sensing and signaling. The induction or repression of transporter-gene expression has been reported to require some N-sensing system(s) within the cell, perhaps in the nucleus or at the cell surface. Membrane-associated proteins have been identified as possible sensors of soil N availability [22, 23]. These sensors located at the root/soil interface may be involved in sensing flux through the transporter protein and/or availability of particular forms of N at the cell surface. These sensors may have a role in regulating cellular N pools and/or detecting available pools of N both inside cells and in the soil around the root [24].
In addition to the high affinity OsAMT genes, all the low affinity OsAMTs except OsAMT4, were also found to express in both roots and shoots of both the genotypes studied. Even after testing several primers sets the expression of OsAMT4 could not be studied. This may be due to very low levels of its expression or maybe it is expressed under certain conditions. Expression of all the low affinity OsAMTs in both roots and shoots suggests that low affinity OsAMTs are also involved in ammonium uptake and its translocation to ariel parts of the plant. The expression profiling of members of OsAMT2 and OsAMT3 family showed interesting and distinct tissue specific level of expression. The expression level of OsAMT2;1 and OsAMT2;2 was almost similar in roots and shoots while the expression of all the members of OsAMT3 gene family was higher in the shoots than in the roots. On the basis of expression, only OsAMT2;1 of the OsAMT2 family has yet been characterized as a constitutively expressed gene in the roots and shoots irrespective of the nitrogen supply [25]. It was interesting to observe that in PB1 which is a high nitrogen responsive genotype expressions of both the OsAMT2 family genes, OsAMT2;1 and OsAMT2;2 were high even in the controls compared to KN3119 which was low but was induced at lower nitrogen conditions and repressed at higher conditions. Low expression levels of OsAMT2;3 in both roots and shoots in all the nitrogen conditions applied indicate that its role in ammonium uptake and translocation is limited. Taken together it can be assumed that all the OsAMT2 family genes in both roots and shoots of low nitrogen responsive genotype are repressed under high nitrogen. In contrast to the OsAMT2 family genes higher expression of members of AMT3 family genes in the shoots than in the roots indicate that their major roles are in the leaves for translocation and distribution of ammonium ions brought from the roots through the xylem. Similar to OsAMT2;1, only OsAMT3;1 of the OsAMT3 family has been characterized. In ‘Nipponbare’ OsAMT3;1 is weakly expressed in the roots and shoots [25]. The present study not only confirm that OsAMT3;1 is weakly expressed in the roots but also report that the other two members of this family i.e., OsAMT3;2 and OsAMT3;3 are also weakly expressed in rice roots. In contrast to root, all the members of AMT3 family were induced in the shoots of both the genotypes except OsAMT3;1 in KN3119. Differential expression of the members of OsAMT2 and OsAMT3 family genes further suggests differential nitrogen sensing mechanism in the genotypes differing in their nitrogen responsiveness. It can also be seen that in low nitrogen responsive genotype many of the low affinity OsAMT genes are repressed under high nitrogen conditions. The expression pattern of almost all the low affinity OsAMT genes were also affected by the changes in nitrogen conditions provided to the plants indicate the optimization of nitrogen uptake by varying the expression levels of AMT genes. These results along with the expression profiles of OsAMT1;3 suggests a probable relation between the nitrogen responsiveness of rice genotypes and the expression patterns of OsAMT genes. Further, the expression of the only high affinity ammonium transporter gene OsAMT1;1 along with six low affinity OsAMT genes in leaves probably provides a support to the high affinity OsAMT to carry out its full function.
The expression of many ammonium transporter genes in the shoots raises one obvious question that, ‘where do the AMT proteins localize in the leaf tissue?’ It is known that the ammonium released from the photorespiratory cycle inside the mitochondria is assimilated by chloroplastic/plastidic form of glutamine synthetase GS2 [26, 27] and for this process to occur, NH4 + released from the mitochondria has to enter chloroplast for its assimilation which might be facilitated by the ammonium transporters present in the membranes of these organelles. Although computer simulation have predicted with considerable likelihood the existence of N-terminal signal peptides for chloroplastic localization (TargetP, www.cbs.dtu.dk) in Arabidopsis and Tomato ammonium transporters (AtAMT1;1, AtAMT1;2, LeAMT1;1, and LeAMT1;3), chloroplast import assays showed that in vitro translated radio-labelled proteins were not incorporated into isolated chloroplasts [28]. A similar study with all rice AMTs (using TargetP, www.cbs.dtu.dk) have also predicted with considerable likelihood the existence of N-terminal signal peptides for chloroplastic localization but its functional validation remains awaited (unpublished data).
Abbreviations
- NUE:
-
Nitrogen use efficiency
- AMT:
-
Ammonium transporter
- RT-PCR:
-
Reverse transcriptase polymerase chain reaction
References
Bloom AJ, Sukrapanna SS, Warner RL (1992) Root respiration associated with ammonium and nitrate absorption by barley. Plant Physiol 99:1294–1301
Gazzarrini S, Lejay L, Gojon A, Ninnemann O, Frommer WB, von WireÂn N (1999) Three functional transporters for constitutive, diurnally regulated, and starvation-induced uptake of ammonium into Arabidopsis roots. Plant Cell 11:937–947
Britto DT, Kronzucker HJ (2001) Constancy of nitrogen turnover kinetics in the plant cell: insights into the integration of subcellular N fluxes. Planta 213:175–181
Kronzucker HJ, Britto DT, Davenport RJ, Tester M (2001) Ammonium toxicity and the real cost of transport. Trends Plant Sci 6:335–337
Sonoda Y, Ikeda A, Saiki S, von Wirén N, Yamaya T, Yamaguchi J (2003) Distinct expression and function of three ammonium transporter genes (OsAMT1;1-1;3) in rice. Plant Cell Physiol 44(7):726–734
Loqué D, von Wiré N (2004) Regulatory levels for the transport of ammonium in plant roots. J Exp Bot 55:1293–1305
Wang MY, Glass ADM, Shaff JE, Kochian LV (1994) Ammonium uptake by rice roots. III. Electrophysiology. Plant Physiol 104:899–906
Ninnemann O, Jauniaux JC, Frommer WB (1994) Identification of a high-affinity NH4 + transporter from plants. EMBO J 13:3464–3471
Sohlenkamp C, Shelden M, Howitt S, Udvardi M (2000) Characterization of Arabidopsis AtAMT2, a novel ammonium transporter in plants. FEBS Lett 467:273–278
Kaiser BN, Rawat SR, Siddiqi MY, Masle J, Glass ADM (2002) Functional analysis of an Arabidopsis T-DNA ‘Knockout’ of the high-affinity NH4 + transporter AtAMT1;1. Plant Physiol 130:1263–1275
Pearson CJ, Finnegan PM, Schjoerring JK (2002) Regulation of the high-affinity ammonium transporter (BnAMT1;2) in the leaves of Brassica napus by nitrogen status. Plant Mol Biol 49:483–490
Salvemini F, Marini AM, Riccio A, Patriarca EJ, Chiurazzi M (2001) Functional characterization of an ammonium transporter gene from Lotus japonicus. Gene 270:237–243
Simon-Rosin U, Wood C, Udvardi MK (2003) Molecular and cellular characterization of LjAMT2;1, ammonium transporter from the model legume Lotus japonicus. Plant Mol Biol 51:99–108
Lauter FR, Ninnemann O, Bucher M, Riesmeier JW, Frommer WB (1996) Preferential expression of an ammonium transporter and of two putative nitrate transporters in root hairs of tomato. Proc Natl Acad Sci USA 93:8139–8144
Von WireÂn N, Gazzarrini S, Gojon A, Frommer WB (2000) The molecular physiology of ammonium uptake and retrieval. Curr Opin Plant Biol 3:254–261
Becker D, Stahnke R, Fendrick I, Frommer WB, Vanderleyden J, Kaiser WM, Hedrich R (2002) Expression of the ammonium transporter gene LeAMT1;2 is induced in tomato roots upon association with N2 fixing bacteria. Planta 215:424–429
Singh RK, Singh US, Khush GS, Rohilla R (2000) Genetics and biotechnology quality traits in aromatic rices. In: Singh RK, Singh US, Khush GS (eds) Aromatic rices. Oxford & IBH Publishing Co. Pvt. Ltd., New Delhi, pp 47–69
Livak KJ, Schmitten TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25:402–408
Kentaro H, Syuntaro H, Satoru I, Isamu N (2008) Ammonia exchange between rice leaf blades and the atmosphere: effect of broadcast urea and changes in xylem sap and leaf apoplastic ammonium concentrations. Soil Sci Plant Nutr 54:807–818
Kiyomiya S, Nakanishi H, Uchida H, Tsuji A, Nishiyama S, Futatsubashi M, Tsukada H, Ishioka N, Watanabe S, Ito S, Mizuniwa T, Osa C, Matsuhashi AS, Hashimoto S, Sekine T, Mori S (2001) Real time visualization of N13-translocation in rice under different environmental conditions using positron emitting tracer imaging system. Plant Physiol 125:1743–1753
Kronzucker HJ, Schjoerring JK, Erner Y, Kirk GJD, Siddiqi MY, Glass ADM (1998) Dynamic interactions between root NH4 + influx and long-distance N translocation in rice: insights into feedback processes. Plant Cell Physiol 39:1287–1293
Redinbaugh M, Campbell W (1991) Higher plant responses to environmental nitrate. Physiol Plant 82:640–650
Forde BG, Clarkson DT (1999) Nitrate and ammonium nutrition of plants: physiological and molecular perspectives. Adv Bot Res 30:1–90
Miller AJ, Cramer MD (2004) Root nitrogen acquisition and assimilation. Plant Soil 274:1–36
Suenaga A, Moriya K, Sonoda Y, Ikeda A, Von Wirén N, Hayakawa T, Yamaguchi J, Yamaya T (2003) Constitutive expression of a novel-type ammonium transporter OsAMT2 in rice plants. Plant Cell Physiol 44(2):206–211
Blackwell RD, Murray AJS, Lea PJ (1987) Inhibition of photosynthesis in barley with decreased levels of chloroplastic glutamine synthetase activity. J Exp Bot 38:1799–1809
Wallsgrove RM, Turner JC, Hall NP, Kendall AC, Bright SWJ (1987) Barley mutants lacking chloroplast glutamate synthetase: biochemical and genetic analysis. Plant Physiol 83:155–158
Shelden MC, De Bruxelles GL, Whelan J, Ryan PR, Howitt SM, Udvardi MK (2001) Arabidopsis ammonium transporter, AtAMT1;1 and AtAMT1;2, have different biochemical properties and functional roles. Plant Soil 231:151–160
Acknowledgments
The present investigation was a part of the DBT (Department of Biotechnology), Govt. of India supported JRF programme. Financial assistance provided by DBT to Vikram Singh Gaur is duly acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Gaur, V.S., Singh, U.S., Gupta, A.K. et al. Understanding the differential nitrogen sensing mechanism in rice genotypes through expression analysis of high and low affinity ammonium transporter genes. Mol Biol Rep 39, 2233–2241 (2012). https://doi.org/10.1007/s11033-011-0972-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-011-0972-2