Abstract
Dextranase from Chaetomium gracile is generally considered safe for use in the sugarcane industry. Herein, a truncated and codon-optimised α-dextranase gene from C. gracile was successfully cloned and expressed in Saccharomyces cerevisiae for the first time. The optimum conditions of fermentation was achieved when the maximum dextranase activity reached to 58.45 U/mL after 48 h in shake flasks. The optimal pH and temperature were 5.5 and 60 °C, respectively. The recombinant dextranase remained stable between pH 4 and 6 and temperature between 55 and 60 °C. The findings in the present study could facilitate large-scale production of food-grade recombinant dextranase for use in the sugar industry.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Dextrans are the collective name for high molecular weight polymers composed of d-glucose units connected via α-1,2, α-1,3 and α-1,4 main-chain linkages (Zhang et al. 2017). Dextrans are synthesised from sucrose by the enzyme dextransucrase secreted by bacteria such as Leuconostoc mesenteroides (Fraga Vidal et al. 2011). The presence of dextrans in syrup enhances the viscosity and thereby affects filtration, clarification and crystal formation, leading to economic losses (Eggleston and Monge 2005; Purushe et al. 2012). Physical methods such as ultrafiltration, dialysis and reverse osmosis have proven useful for removing dextrans from sugarcane juice yet are not implemented in the sugarcane industry due to high costs (Mao et al. 2018).
Commercial dextranase has been used in sugarcane factories to degrade dextrans into smaller molecules by endogenously hydrolysing the α-(1,6) linkages, as exemplified in the extraction of sugar from sugar beet (Li et al. 2017). Dextranases are present in a wide variety of fungi and bacteria, and fungi are the main commercial sources (Khalikova et al. 2005). Most commercial dextranases in the USA are produced from either Penicillium spp. or Chaetomium spp. (Eggleston 2009). However, there are safety concerns, and relatively low productivity restricts their application in the sugar-making process (Zhang et al. 2018). To solve these problems, researchers have focused on recombinant enzymes to meet the immense demand for dextranases in the in food, medicine and chemical industries. Although various dextranases of bacterial and fungal origin have been previously reported, few are currently employed in the sugar industry (Khalikova et al. 2005; Bertrand et al. 2014) due to high enzyme cost, cumbersome in dextranase purification and toxic by-products from fungi (Rerngsamran et al. 2014).
A solution is the deployment of engineered strains to produce dextranases, among which Escherichia coli, Pichia pastoris and Saccharomyces cerevisiae are the three most commonly used heterologous expression systems. Dextranase genes from bacteria and fungi have been heterologously expressed in E. coli, often as insoluble inclusion bodies (Purushe et al. 2012; Khalikova et al. 2005). Although the methylotrophic yeast P. pastoris is widely used for the heterologous production of recombinant proteins (Macauley-Patrick et al. 2005), including dextranases (Kang et al. 2009; Chen et al. 2008), volatile toxic compounds such as methanol can impact food safety of products from this organism. S. cerevisiae is easily cultivated, has been used successfully in industrial applications for many years and has Generally Recognised as Safe (GRAS) status (Liu et al. 2013). Kang et al. (2005) reported the cloning and characterisation of a dextranase gene from Lipomyces starkeyi and its heterologous expression in S. cerevisiae, but the activity of the recombinant enzyme was not given, and the optimal temperature of the recombinant enzyme was 37 °C, which is too low for use in the sugarcane industry.
China is the fourth largest producer of sugarcane in the world, producing ~ 10 million tons of sugarcane per year, but residual dextrans in syrup result in large economic losses. Also, the sugar-making process usually requires high temperatures, and hence, thermostable enzymes are needed. In the present study, an optimised dextranase-encoding gene from Chaetomium gracile was expressed in S. cerevisiae, and various enzymatic properties of the recombinant enzyme, including optimal temperature, pH, thermal and pH stabilities, were studied. We also optimised the fermentation conditions to maximise dextranase production in engineered S. cerevisiae strains.
Materials and Methods
Microorganisms, Plasmids and Culture Medium
Escherichia coli JM109 was used as the recipient strain for cloning manipulation and plasmid amplification and was cultivated in Luria–Bertani (LB) medium containing tryptone (10 g/L), yeast extract (5 g/L) and NaCl (10 g/L) with or without 100 µg/L of ampicillin. S. cerevisiae CEN. PK2-1B (MATα; his3Δ1, trp1-289, leu2-3, 112, ura3-52, MAL2-8c, SUC2) was obtained from EUROSCARF (Frankfurt, Germany) and used for genetic manipulation. Engineered S. cerevisiae strains were grown in yeast extract–peptone–dextrose (YPD) medium (2% glucose, 2% tryptone, 1% yeast extract) or yeast nitrogen base (YNB) medium (0.17% yeast nitrogen base without amino acids, 0.5% ammonium sulphate, 2% glucose, supplemented with 50 µg/mL of each of the required amino acids). All chemicals were of reagent grade and were obtained from commercial sources.
Construction and Transformation of Recombinant Plasmids
The plasmid T-vector pMD19 (Simple) was purchased from TaKaRa (Dalian, China), and the yeast expression plasmid pY26-TEF1-GPD1 was kindly provided by Li et al. (2007). Plasmid pRS306 was kindly provided by Sikorski and Hieter (1989).
SignalP 4.1 (http://www.cbs.dtu.dk/services/SignalP/) was used to predict the signal peptide sequence of the α-dextranase gene from C. gracile (GenBank: KC707808.1), suggesting the first 54 bp encodes a signal peptide, and the 4–54 bp region was truncated in the codon-optimised dex gene synthesised by Sangon (Shanghai, China). The sequence of the dex gene is provided in Supplementary data.
The synthesised dex gene was cloned into the pY26 and pRS306 vectors to obtain the expression constructs pY26-dex and pRS306-dex, which were verified and transformed into S. cerevisiae CEN.PK2-1B using Fast-Yeast Transformation regent (G-Biosciences, St. Louis, MO) according to the manufacturer’s instructions. Strains harbouring integrated plasmids containing auxotrophic selection markers were screened using corresponding auxotrophic YNB plates and verified by PCR analysis.
Enzyme Assays and Sodium Dodecyl Sulphate–Polyacrylamide Gel Electrophoresis (SDS–PAGE)
In this study, dextranase was produced by S. cerevisiae as an intracellular hydrolase, and snailase was used to destroy the yeast cell wall. Yeast cells were harvested by centrifugation (1500g at 25 °C for 5 min), and cell pellets were resuspended in 5 mL of solution containing 1.0 M sorbitol, 250 µL of 20 × phosphate-buffered saline (PBS) and 1% snailase (Sigma, Shanghai) and incubated at 37 °C for 1 h. Then, the cells were further treated with glass beads (0.4–0.6 mm, OMEGA) by vortex. The supernatant of cell-broken cells was used as crude enzyme.
Dextranase activity was measured using the dinitrosalicylic acid (DNS) method by incubating 100 µL of enzyme with 900 µL of 2% dextran T2000 (Sigma) as previously reported (Li et al. 2006). One unit of dextranase (U/mL) is defined as the amount of enzyme that degrades dextran T2000 to produce reducing sugar per min under assay conditions at 37 °C. Crude dextranase was analysed by SDS–PAGE, and gels were stained with Coomassie Brilliant Blue R-250 (Kang et al. 2009).
Optimisation of Fermentation Conditions
To determine the initial culture pH, we varied the substrate pH range from pH 4.0 to 7.0, and the enzyme activity of the recombinant dextranase was calculated as described above. To investigate the optimum temperature for enzyme production, culturing was carried out at different temperatures between 26 and 34 °C.
In this experiment, we tested the effects of different substrates and substrate concentration on dextranase production. Glucose (1% w/v) present in the basal medium was replaced by sucrose, glycerol, fructose, lactose or starch. Meanwhile, tryptone (2% w/v) present in the basal medium was substituted by organic and inorganic nitrogen sources including casein, ammonium chloride and urea.
Properties of Recombinant Dextranase
To determine the optimum pH of the recombinant dextranase, dextran T-2000 (2%) was dissolved in different substrate solutions at pH 3.0–8.0 using citrate–phosphate buffers. The pH stability was assessed by treating recombinant dextranase at pH 3.0–8.0 for 1 h, and the residual activity at each pH was expressed relative to the highest activity (100%). All experiments were performed in triplicate.
To determine the optimum temperature for dextranase catalysis, enzyme was incubated with dextran T-2000 (2%) at different temperatures between 30 and 80 °C, and thermostability was assessed by incubating dextranase at 30–80 °C for an h. The activity at each temperature was expressed relative to the highest activity (100%). All experiments were performed in triplicate.
Results
Expression of Codon-Optimised Dextranase in S. cerevisiae
To evaluate the expression of dextranase from C. gracile in S. cerevisiae, the codon-optimised dex gene encoding the synthase was expressed in S. cerevisiae CEN. PK2-1B using the episome plasmid pY26-TEF-GPD, resulting in the engineered S. cerevisiae strain XD02. The SDS–PAGE results verified the secretion of the 63 kDa dextranase by the XD02 cells, and no comparable band was visible with the XD01 control strain carrying empty pY26-TEF-GPD (Fig. 1). The DNS results indicated that dextranase production by XD02 in YNB medium reached a maximum of 17.86 U/mL after 72 h of cultivation.
SDS–PAGE analysis of recombinant dextranase expressed in S. cerevisiae. Lane 1, S. cerevisiae CEN. PK-1B (negative control); lane 2, supernatant of S. cerevisiae XD02 wall-broken cells; lane 3, cell suspension of S. cerevisiae XD02 wall-broken cells; lane 4, supernatant of S. cerevisiae XD03 wall-broken cells
We also evaluated the expression of dex following incorporation into the chromosome of S. cerevisiae CEN. PK2-1B using the integrant plasmid pRS306, resulting in engineered S. cerevisiae strain XD03. When cultured in YPD medium, the cell density of S. cerevisiae strain XD03 increased rapidly during the first 36 h and then remained consistent after 48 h. Despite the slower cell growth after 24 h of incubation indicating stationary phase, more dextranase was produced between 24 and 48 h, suggesting cells at stationary phase produced more target enzyme. The maximum dextranase activity of strain XD03 was obtained after 60 h (40.42 U/mL), slightly higher than that after 48 h (40.26 U/mL; Fig. 2). Afterwards, dextranase production declined gradually to the end of the incubation, as cell growth further declined (data not shown). Previous research reported that production of C. gracile dextranase was maximal after incubation for 96 h (Li et al. 2017). Thus, our results indicate that heterologous gene expression in S. cerevisiae is more rapid than the time-consuming fermentation of C. gracile. Since the engineered S. cerevisiae strain XD03 exhibited superior dextranase production compared with strain XD02, XD03 was employed for dextranase production in subsequent experiments.
Cell growth and dextranase production in the engineered S. cerevisiae strain. The solid black and open blue squares represent the absorbance at 600 nm and dextranase activity of strain XD03, respectively. The solid black and blue open triangles represent the absorbance at 600 nm and dextranase activity of strain XD02, respectively. The results are presented as means and standard deviations of at least three independent experiments
Effects of Medium pH and Incubation Temperature on Dextranase Production
Relatively few studies have explored the influence of initial pH and temperature of the medium on yeast intracellular recombinant protein production, although both parameters are known to affect fugal growth. In the present study, the influence of initial pH and culture temperature on dextranase production in engineered S. cerevisiae was examined. Strain XD03 cells were incubated in YPD medium in 500 mL flasks with a working volume of 50 mL and incubated at 30 °C for 60 h. Maximum growth was observed at pH 6.0, at which the absorbance at 600 nm (OD600) reached 10.3. Maximum dextranase production was achieved at pH 5.5, at which dextranase activity reached 42.1 U/mL. These results indicate dextranase production over a broad pH range (pH 4.0–7.0; Fig. 3a). Thus, pH 5.5 was considered the most suitable pH for achieving the highest yield of dextranase in engineered yeast strain XD03.
To address the effect of temperature on dextranase production, XD03 cells were cultured in YPD medium in a 500-mL flask with a working volume of 50 mL at various temperatures from 26 to 34 °C for up to 60 h. Cell density and dextranase production were analysed, and the results are summarised in Fig. 3b. Maximum cell growth rate was achieved at 28 °C, and cell growth decreased gradually thereafter with increasing temperature. The optimal temperature for dextranase production was 30 °C, at which activity reached 42.13 U/mL. Thus, to maximise dextranase production, we selected 30 °C as the optimum temperature in subsequent experiments.
Effects of Nitrogen and Carbon Source on Dextranase Production
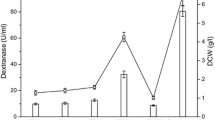
We also studied the effects of different carbon and nitrogen sources on dextranase production at different concentrations. While keeping all other culture conditions unchanged, the sole carbon source was sequentially replaced by sucrose, glucose, glycerol, fructose, lactose and starch. As shown in Fig. 4a, dextranase production and cell growth of strain XD03 on glycerol, fructose and glucose were clearly higher than on sucrose, lactose or starch. Maximal growth of XD03 cells occurred using glycerol, and the highest dextranase yield was achieved using glucose. Thus, glucose was regarded as the best alternative carbon source for dextranase production. With increasing glucose concentration, the growth of yeast cells increased almost proportionately, suggesting the carbon source was growth limiting. We established that the optimal glucose concentration was 25 g/L (Fig. 4b). Previous reports also emphasised that obtaining the maximum biomass does not guarantee successful fermentation and production of target molecules (Martinez-Moreno et al. 2012).
Effects of carbon and nitrogen source on dextranase production and cell growth. a Different carbon sources; b different glucose concentrations; c different nitrogen sources; d different casein concentrations. Columns represent the absorbance variation at 600 nm, and filled blue squares represent dextranase activity changes
We also investigated effect of nitrogen source at different concentrations and found that tryptone and casein stimulated greater enzyme production than other nitrogen sources such as ammonium chloride and urea. Furthermore, the optimal casein concentration was 25 g/L, the same as the optimal glucose concentration (Fig. 4c, d). Consequently, 25 g/L casein was regarded as the best nitrogen source for dextranase production. Using the optimal fermentation conditions, we achieved dextranase production of 58.45 U/mL in 48 h, ~ 1.45-fold higher than the initial fermentation conditions.
Characterisation of Recombinant Dextranase
Temperature and pH are two fundamental factors involved in enzyme catalytic activity. By subjecting recombinant dextranase to different pH conditions ranging from pH 3.0 to 8.0, maximum dextranase activity was observed at pH 5.5 (Fig. 5a). To assess the pH stability of recombinant dextranase, dextranase activity was measured after a 1-h incubation at pH 3.0–8.0. The results showed that the enzyme maintained more than 90% activity at pH 4–6 (Fig. 5b).
Temperature and thermal stability are generally regarded as the primary factors determining whether an enzyme is suitable for industrial applications. To determine the optimal temperature, dextranase activity was measured at temperatures ranging from 30 to 80 °C. As shown in Fig. 5c, enzymatic activity increased with increasing temperature and exhibited maximal activity at 60 °C, but activity decreased sharply thereafter at higher temperatures. Recombinant enzyme retained > 80% of initial activity after incubation at 50–60 °C for 1 h (Fig. 5d). These results were similar to those of previous characterisations of thermostable dextranases from Talaromyces pinophilus (Zhang et al. 2017) and Chaetomium erraticum (Virgen-Ortíz et al. 2015).
Discussion
Given the importance of dextranases in the sugar industry for alleviating the viscosity of syrups and cleaning blocked machines (Chen et al. 2009), much effort has gone into screening high-level dextranase-producing strains, including expressing dextranase genes from Streptococcus rattus and P. minioluteum in E. coli or P. pastoris (Roca et al. 1996; Igarashi et al. 2004). At present, commercial sources of dextranase from Chaetomium spp. require long fermentation times, and enzymes from Penicillium spp. can suffer from food safety issues (Eggleston 2009). Additionally, the main two dextranase producers mentioned above usually utilise expensive dextran as a carbon source. Thus, efficient production of food-grade dextranase from cheap substrates could benefit the sugar-making process. To our knowledge, this is the first report of the production of recombinant dextranase from C. gracile in S. cerevisiae.
S. cerevisiae is a well-characterised eukaryotic model organism for the production of heterologous proteins (Liu et al. 2012), and this organism has been used safely for food production for thousands of years. S. cerevisiae strains have many advantages including a well-understood genetic background, easy cultivation, a cheap substrate spectrum, and many years of use in industrial-scale fermentation processes (Lian et al. 2018). The food safety of dextranase from C. gracile was confirmed in the USA in 1986, it has Generally Recognised as Safe (GRAS) status, and it has been applied industrially (Eggleston and Monge 2005; Virgen-Ortíz et al. 2015). To achieve food-grade dextranase production, we chose S. cerevisiae as host and the C. gracile dextranase gene, and no resistance markers were introduced in the present work. Heterologous gene expression can suffer from low productivity due to the presence of a signal peptide, which has an adverse impact on protein folding (Ljungdahl and Daignan-Fornier 2012). In the present study, we removed the region of the dextranase gene predicted to encode a signal peptide, and the rest of the gene encoding the remaining residues was synthesised using codon optimisation. Following cloning of the resulting gene into appropriate expression vectors, culturing resulted in detectable dextranase production in the engineered S. cerevisiae strains (Figs. 1, 2).
Cheap substrates can reduce the cost of fermentation processes and thereby decrease product price. Much research has been conducted on optimising the production of valuable products using low-cost substrates in fermentation processes in S. cerevisiae (Kwak and Jin 2017). Carbon sources are indispensable nutrients for microbial growth and metabolite accumulation and can impact on microbial growth and metabolism, and thus enzyme production (Luo et al. 2018). Nitrogen sources are also vital for the synthesis of microbial cell proteins and nucleic acids and can have an important influence on cell growth and the accumulation of metabolites (Armando et al. 2013). In the present study, we evaluated the influence of different carbon and nitrogen sources on dextranase production in S. cerevisiae, and optimum concentrations were also determined (Fig. 4). Optimising the culture medium in this way resulted in a 44.6% increase compared with the initial medium. Moreover, the fermentation time was shortened from more than 96 h in Chaetomium spp. to 48 h in engineered S. cerevisiae cells.
In general, weak acidic conditions can decrease non-specific reactions and the accumulation of undesirable by-products that may be accrued under alkaline and high-temperature conditions (Chen et al. 2018). In the present work, the recombinant dextranase exhibited high activity and stability under weak acid conditions (Fig. 5). Also, industrial production is often carried out at high temperatures; in the sugar-making process, the temperature is generally > 60 °C (Eggleston and Monge 2005). The recombinant dextranase produced in the present work exhibited high enzyme activity and thermal stability at high temperatures (Fig. 5).
In summary, a truncated dextranase from C. gracile was recombinantly expressed in S. cerevisiae, and the enzyme exhibited high pH and thermal stability, making it suitable for use in industrial applications. Dextranase production by the engineered S. cerevisiae strain XD03 was increased significantly following optimisation of the fermentation conditions. This work provides a strategy for expression dextranase genes from C. gracile in recombinant S. cerevisiae for dextranase production. Although the engineered dextranase displayed good food safety and thermostability credentials, further studies are needed to assess the large-scale application of this enzyme.
References
Armando, M.R., M.A. Galvagno, C.A. Dogi, P. Cerrutti, A.M. Dalcero, and L.R. Cavaglieri. 2013. Statistical optimization of culture conditions for biomass production of probiotic gut-borne Saccharomyces cerevisiae strain able to reduce fumonisin B1. Journal of Applied Microbiology 114(5): 1338–1346. https://doi.org/10.1111/jam.12144.
Bertrand, E., G. Pierre, C. Delattre, C. Gardarin, N. Bridiau, T. Maugard, A. Strancar, and P. Michaud. 2014. Dextranase immobilization on epoxy CIM (R) disk for the production of isomaltooligosaccharides from dextran. Carbohydrate Polymers 111: 707–713. https://doi.org/10.1016/j.carbpol.2014.04.100.
Chen, Z.W., W. Xu, W.L. Zhang, T. Zhang, B. Jiang, and W.M. Mu. 2018. Characterization of a thermostable recombinant l-rhamnose isomerase from Caldicellulosiruptor obsidiansis OB47 and its application for the production of l-fructose and l-rhamnulose. Journal of the Science of Food and Agriculture 98(6): 2184–2193. https://doi.org/10.1002/jsfa.8703.
Chen, L., C. Yu, X.S. Zhou, and Y.X. Zhang. 2009. Rational introduction of disulfide bond to enhance optimal temperature of Lipomyces starkeyi alpha-dextranase expressed in Pichia pastoris. Journal of Microbiology and Biotechnology 19(12): 1506–1513. https://doi.org/10.4014/jmb.0902.0096.
Chen, L., X.S. Zhou, W.M. Fan, and Y.X. Zhang. 2008. Expression, purification and characterization of a recombinant Lipomyces starkey dextranase in Pichia pastoris. Protein Expression and Purification 58(1): 87–93. https://doi.org/10.1016/j.pep.2007.10.021.
Eggleston, Gillian. 2009. Application of dextranases in sugarcane factory: Overcoming practical problems. Sugar Tech 11(2): 135–141. https://doi.org/10.1007/s12355-009-0020-x.
Eggleston, G., and A. Monge. 2005. Optimization of sugarcane factory application of commercial dextranases. Process Biochemistry 40(5): 1881–1894. https://doi.org/10.1016/j.procbio.2004.06.025.
Fraga Vidal, Reinaldo, Aidin Martinez, Claire Moulis, Pierre Escalier, Sandrine Morel, Magali Remaud-Simeon, and Pierre Monsan. 2011. A novel dextransucrase is produced by Leuconostoc citreum strain B/110-1-2: An isolate used for the industrial production of dextran and dextran derivatives. Journal of Industrial Microbiology and Biotechnology 38(9): 1499–1506. https://doi.org/10.1007/s10295-010-0936-x.
Igarashi, T., H. Morisaki, and N. Goto. 2004. Molecular characterization of dextranase from Streptococcus rattus. Microbiology and Immunology 48(3): 155–162. https://doi.org/10.1111/j.1348-0421.2004.tb03501.x.
Kang, H.K., S.H. Kim, J.Y. Park, X.J. Jin, D.K. Oh, S.S. Kang, and K. Doman. 2005. Cloning and characterization of a dextranase gene from Lipomyces starkeyi and its expression in Saccharomyces cerevisiae. Yeast 22(15): 1239–1248. https://doi.org/10.1002/yea.1311.
Kang, H.K., J.Y. Park, J.S. Ahn, S.H. Kim, and D. Kim. 2009. Cloning of a gene encoding dextranase from Lipomyces starkeyi and its expression in Pichia pastoris. Journal of Microbiology and Biotechnology 19(2): 172–177. https://doi.org/10.4014/jmb.0802.100.
Khalikova, E., P. Susi, and T. Korpela. 2005. Microbial dextran-hydrolyzing enzymes: Fundamentals and applications. Microbiology and Molecular Biology Reviews 69(2): 306–325. https://doi.org/10.1128/Jmrb.69.2.306-325.2005.
Kwak, Suryang, and Yong-Su Jin. 2017. Production of fuels and chemicals from xylose by engineered Saccharomyces cerevisiae: A review and perspective. Microbial Cell Factories 16: 82. https://doi.org/10.1186/s12934-017-0694-9.
Li, X., S.H. Millson, R.D. Coker, and I.H. Evans. 2006. Cloning and expression of Penicillium minioluteum dextranase in Saccharomyces cerevisiae and its exploitation as a reporter in the detection of mycotoxins. Biotechnology Letters 28(23): 1955–1964. https://doi.org/10.1007/s10529-006-9183-7.
Li, Aimin, Zengshan Liu, LuYu. Qianxue Li, Dacheng Wang, and Xuming Deng. 2007. Construction and characterization of bidirectional expression vectors in Saccharomyces cerevisiae. FEMS Yeast Research 8(1): 6–9. https://doi.org/10.1111/j.1567-1364.2007.00335.x.
Li, K., H.Q. Lu, F.X. Hang, S.B. Li, and J.D. Liu. 2017. Improved dextranase production by Chaetomium gracile through optimization of carbon source and fermentation parameters. Sugar Tech 19(4): 432–437. https://doi.org/10.1007/s12355-016-0476-4.
Lian, Jiazhang, Shekhar Mishra, and Huimin Zhao. 2018. Recent advances in metabolic engineering of Saccharomyces cerevisiae: New tools and their applications. Metabolic Engineering. https://doi.org/10.1016/j.ymben.2018.04.011.
Liu, Z.H., K.E.J. Tyo, J.L. Martinez, D. Petranovic, and J. Nielsen. 2012. Different expression systems for production of recombinant proteins in Saccharomyces cerevisiae. Biotechnology and Bioengineering 109(5): 1259–1268. https://doi.org/10.1002/bit.24409.
Liu, J.D., W.P. Zhang, G.C. Du, J. Chen, and J.W. Zhou. 2013. Overproduction of geraniol by enhanced precursor supply in Saccharomyces cerevisiae. Journal of Biotechnology 168(4): 446–451. https://doi.org/10.1016/j.jbiotec.2013.10.017.
Ljungdahl, Per O., and Bertrand Daignan-Fornier. 2012. Regulation of amino acid, nucleotide, and phosphate metabolism in Saccharomyces cerevisiae. Genetics 190(3): 885–929. https://doi.org/10.1534/genetics.111.133306.
Luo, G., J.H. Tian, H.Q. Huang, and L. An. 2018. Improving heterologous expression of porcine follicle-stimulating hormone in Pichia pastoris by integrating molecular strategies and culture condition optimization. Applied Microbiology and Biotechnology 102(20): 8867–8882. https://doi.org/10.1007/s00253-018-9260-6.
Macauley-Patrick, S., M.L. Fazenda, B. McNeil, and L.M. Harvey. 2005. Heterologous protein production using the Pichia pastoris expression system. Yeast 22(4): 249–270. https://doi.org/10.1002/yea.1208.
Mao, H.Y., M.H. Qiu, X.F. Chen, H. Verweij, and Y.Q. Fan. 2018. Fabrication and in situ fouling mitigation of a supported carbon nanotube/gamma-alumina ultrafiltration membrane. Journal of Membrane Science 550: 26–35. https://doi.org/10.1016/j.memsci.2017.12.050.
Martinez-Moreno, R., P. Morales, R. Gonzalez, A. Mas, and G. Beltran. 2012. Biomass production and alcoholic fermentation performance of Saccharomyces cerevisiae as a function of nitrogen source. FEMS Yeast Research 12(4): 477–485. https://doi.org/10.1111/j.1567-1364.2012.00802.x.
Purushe, Shweta, Divya Prakash, Neelu N. Nawani, Prashant Dhakephalkar, and Balasaheb Kapadnis. 2012. Biocatalytic potential of an alkalophilic and thermophilic dextranase as a remedial measure for dextran removal during sugar manufacture. Bioresource Technology 115: 2–7. https://doi.org/10.1016/j.biortech.2012.01.002.
Rerngsamran, P., P. Temjitpukdee, N. Assavasirijinda, S. Chareonpornwattana, and S. Thaniyavarn. 2014. Cloning, characterization, and heterologous expression of a dextranase gene from Penicillium pinophilum SMCU3-14. Scienceasia 40(6): 405–413. https://doi.org/10.2306/scienceasia1513-1874.2014.40.405.
Roca, H., B. Garcia, E. Rodriguez, D. Mateu, L. Coroas, J. Cremata, R. Garcia, T. Pons, and J. Delgado. 1996. Cloning of the Penicillium minioluteum gene encoding dextranase and its expression in Pichia pastoris. Yeast 12(12): 1187–1200. https://doi.org/10.1002/(Sici)1097-0061(19960930)12:12%3c1187:Aid-Yea986%3e3.0.Co;2-U.
Sikorski, Robert S., and Philip Hieter. 1989. A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics 122: 19–27.
Virgen-Ortíz, J.J., V. Ibarra-Junquera, P. Escalante-Minakata, J. de Ornelas-Paz, J.A. Osuna-Castro, and A. González-Potes. 2015. Kinetics and thermodynamic of the purified dextranase from Chaetomium erraticum. Journal of Molecular Catalysis B: Enzymatic 122: 80–86. https://doi.org/10.1016/j.molcatb.2015.08.020.
Zhang, Yu-Qi, Ruo-Han Li, Hong-Bin Zhang, Wu Min, and Hu Xue-Qin. 2017. Purification, characterization, and application of a thermostable dextranase from Talaromyces pinophilus. Journal of Industrial Microbiology and Biotechnology 44(2): 317–327. https://doi.org/10.1007/s10295-016-1886-8.
Zhang, Z.D., J.D. Liu, S.Y. Ma, H.Q. Lu, F.X. Hang, P. Huang, and K. Li. 2018. Enhancement of catalytic performance of alpha-dextranase from Chaetomium gracile through optimization and suitable shear force. Sugar Tech 20(1): 78–87. https://doi.org/10.1007/s12355-017-0540-8.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31460026), the National Natural Science Foundation of Guangxi Province (2018GXNSFAA050126), the Bosch Young Teachers Innovation Training Project (BRP180215) and the Fundamental Research Fund of Guangxi University (XJZ140293).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical Statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed Consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, J., Sun, Q., Yin, H. et al. Optimal Fermentation of Saccharomyces cerevisiae Expressing a Dextranase from Chaetomium gracile. Sugar Tech 22, 171–178 (2020). https://doi.org/10.1007/s12355-019-00746-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12355-019-00746-5