Abstract
APOBEC3G is a cellular antiviral protein that restricts retroviral infection. In non-permissive cells infected by Vif-deficient HIV-1, the protein mediates the hypermutation of viral DNA through the enzymatic activity of cytidine deaminase. To counteract the antiviral activity of APOBEC3G, an accessory protein of HIV-1, Vif, forms ubiquitin E3 ligase through assembly with CUL5-RBX2, ELOB-ELOC and CBFβ. Subsequently, Vif recruits APOBEC3G to the complex as a substrate adaptor of ubiquitin E3 ligase and induces poly-ubiquitination of APOBEC3G for its proteasomal degradation (Fig. 1). This review briefly summarizes current understanding of protein–protein interaction between Vif and host factors required for APOBEC3 degradation, based on high resolution structures of APOBEC3 proteins and Vif-CUL5NTD-ELOBC-CBFβ complex.

A schematic model of assembly of Vif ubiquitin E3 ligase and subsequent APOBEC3 degradation. HIV-1 Vif hijacks cellular E3 ligase components containing CUL5, RBX2, ELOB, ELOC and CBFβ, to poly-ubiquitinate antiviral cellular factors, APOBEC3 proteins. Poly-ubiquitinated APOBEC3 proteins are targeted for proteasomal degradation. Ubiquitin is labeled as Ub
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The RNA genome of HIV-1 (Type-1 human immunodeficiency virus) harbors several open reading frames in addition to the retro-viral elements, gag, pol and env. Two additional proteins encoded by rev and tat genes are involved in viral gene regulation. Rev transports unspliced or incompletely spliced HIV mRNA from nucleus to cytoplasm, to regulate the timing of viral gene expression (Cullen 2003). Tat prevents transcription attenuation by recruiting cellular transcription elongation factors such as P-TEFb (Ott et al. 2011). The remaining four accessory proteins are Vif, Vpu, Vpr and Nef. Even though the function of HIV-1 accessory proteins is not fully understood yet, they seem to play a role in providing a virus-favorable environment. Especially, Vif and Vpu help viral spread by neutralizing cellular antiviral factors (Strebel 2013). Virion infectivity factor mediates poly-ubiquitination and subsequent proteasomal degradation of cellular antiviral factors through the assembly of ubiquitin E3 ligase. Vpu releases virus particle by directly blocking tetherin (BST-2, CD164), which holds virions to the plasma membrane (Neil et al. 2008; van Damme et al. 2008; Perez-Caballero et al. 2009).
Virion infectivity factor (Vif) of approximately 23 kDa plays critical roles in HIV-1 spread in non-permissive cell types such as primary HUT78, H9 and peripheral blood lymphocytes, while it is not required in permissive cell types such as SupT1, CEM-SS and Jurkat cells. In other words, non-permissive cells are resistant to viral spread of vif-deficient HIV-1 (HIV-1 Δvif) and produce less infectious virus (Fisher et al. 1987; Gabuzda et al. 1992; Sakai et al. 1993; Sova and Volsky 1993; von Schwedler et al. 1993; Bouyac et al. 1997). Moreover, transient heterokaryons formed by the fusion of non-permissive (HUT78) and permissive (293T) cells retained the restrictive ability against HIV-1 Δvif production. It suggests that Vif may restrict the innate anti-viral activity for HIV-1 replication (Simon et al. 1998). Interestingly, the comparison of mRNA expression between permissive and non-permissive cell types led to identification of APOBEC3G, which is specifically expressed in non-permissive cells. Transient expression of APOBEC3G in permissive cells actually converted them to a non-permissive phenotype that restricts HIV-1 Δvif infectivity (Sheehy et al. 2002).
APOBEC3G (Apolipoprotein B mRNA editing enzyme catalytic peptide 3G; A3G) of 386 amino acids is incorporated into HIV-1 Δvif during viral budding and presents antiviral activity in target cells (Sheehy et al. 2002). It has been suggested that A3G protein restricts HIV-1 spread through DNA editing and non-editing modes. In editing mode, A3G has cytidine deaminase activity that mutates deoxycytidine to deoxyuridine in the minus strand of HIV-1 reverse transcript. The activity results in the accumulation of non-functional proviruses by G to A hyper-mutation in the viral DNA (Harris et al. 2003; Lecossier et al. 2003; Mangeat et al. 2003; Zhang et al. 2003). In non-editing mode, A3G appears to inhibit the HIV-1 cDNA synthesis by reverse transcription and the integration of the proviral DNA into human genome. Indeed, catalytically inactive A3G displays significant anti-retroviral activity, indicating that the deaminase-independent activity of A3G contributes to HIV-1 inhibition (Newman et al. 2005; Guo et al. 2006, 2007; Iwatani et al. 2007; Li et al. 2007; Luo et al. 2007; Mbisa et al. 2007; Bishop et al. 2008; Wang et al. 2012; Belanger et al. 2013). Although the cellular condition that expresses active A3G is unfavorable for viral replication, wild-type HIV-1 can overcome a difficult situation by presenting the anti-antiviral proteins. Vif in HIV-1 counteracts antiviral activities through direct interaction with A3G, resulting in the prevention of its virion incorporation and the recruitment of A3G-targeted ubiquitin–proteasome system (Conticello et al. 2003; Mariani et al. 2003; Marin et al. 2003; Sheehy et al. 2003; Stopak et al. 2003). For the poly-ubiquitination of A3G, Vif reconstitutes ubiquitin E3 ligase by hijacking cellular proteins containing CUL5/RBX2, ELOBC and CBFβ (Yu et al. 2003; Kamura et al. 2004; Jager et al. 2012; Kim et al. 2013).
In human, seven members of the APOBEC3 (A3) protein family, A3A, A3B, A3C, A3DE, A3F, A3G and A3H, are encoded in a tandem array on chromosome 22 and their expression levels vary in different tissues and cell types (Jarmuz et al. 2002; Koning et al. 2009). Although A3G seems to be sufficient to confer a non-permissive phenotype against HIV-1 Δvif to permissive cells, other members of the APOBEC3 family, A3DE, A3F and A3H, have been suggested to contribute to restriction of HIV-1 replication in the absence of Vif (Miyagi et al. 2010; Mulder et al. 2010; Chaipan et al. 2013; Desimmie et al. 2014). And A3A, A3B and A3C have also been reported to inhibit retroviral replication (Langlois et al. 2005; Bishop et al. 2004; Doehle et al. 2005; Aguiar et al. 2008), indicating all of the A3 proteins are involved in restriction of viral replication. Among the APOBEC3 proteins, Vif reduces the expression of A3C, A3DE, A3F and haplotype II A3H as well as A3G through direct interaction, when those proteins are transiently expressed in human cell lines (Wiegand et al. 2004; Langlois et al. 2005; Dang et al. 2006; Zhen et al. 2010).
Since the inhibition of Vif-mediated A3G degradation represents a new strategy for anti-HIV-1 therapy, the binding interface between Vif and APOBEC3 proteins has been studied extensively. Also, the binding interface between Vif and host factors including CUL5, ELOC and CBFβ could be a therapeutic target for inhibition of viral infection. Currently, high-resolution structures of APOBEC3 proteins (A3A, A3C, A3F-CTD and A3G-CTD) and Vif-CUL5NTD-ELOBC-CBFβ complex are available (Chen et al. 2008; Holden et al. 2008; Furukawa et al. 2009; Kitamura et al. 2012; Bohn et al. 2013; Byeon et al. 2013; Siu et al. 2013; Guo et al. 2014). Although the structure of Vif-APOBEC3 complex has not been determined yet, the structural analysis of A3 and Vif combined with biochemical data provides the identification of critical binding motifs that could be targets for viral inhibitors. This brief review aims to summarize current understanding of interactions for assembly of Vif-ubiquitin E3 ligase and recruitment of APOBEC3 proteins, based on the protein structures.
Structure of HIV-1 Vif
Even though the critical role of Vif in HIV-1 infectivity has been identified, the purification of homogeneous recombinant Vif protein was extremely difficult, resulting in research limitations in the field of protein biochemistry and structure determination (Marcsisin et al. 2011). Breakthrough was achieved by proteomics approaches that allowed the new finding of Vif-binding factor, CBFβ (Jager et al. 2012; Zhang et al. 2012). Additional studies revealed that CBFβ binds to Vif with high binding affinity and cooperatively stabilizes Vif together with ELOBC. Interestingly, the co-expression of Vif, ELOBC and CBFβ facilitated the purification of a hetero-tetrameric complex of Vif-ELOBC-CBFβ. And the hetero-hexameric form of CUL5-RBX2-Vif-ELOBC-CBFβ that has E3 ligase activity was also purified by mixing Vif-ELOBC-CBFβ and CUL5-RBX2 (Jager et al. 2012; Kim et al. 2013).
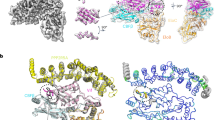
HIV-1 Vif is a highly basic (pI value ≈ 10.0) small protein composed of 192 amino acids. Recently, the crystal structure of CUL5NTD-Vif-ELOBC-CBFβ complex was determined at 3.3 Å resolution by Guo et al. (Guo et al. 2014; Fig. 2a). Consistent with biochemical analysis explained above, the complex structure reveals that Vif mediates the assembly of CUL5, ELOBC and CBFβ through direct binding and CBFβ does not interact with CUL5 or ELOBC, indicating that CBFβ is an additional factor that can regulate Vif function. Vif monomer in the complex structure maintains an elongated shape with positive charge distributions on its surface. In total, 169 residues of Vif (N3 to D171) are traced in the electron density map and its fold represents two domains, α/β domain and α domain. The α/β domain consists of an antiparallel beta-sheet of six beta strands (β1–β6) and three alpha-helices (α1, α2 and α6) that are aligned along the convex side of the beta sheet. The α domain (residues 112–161) is composed of three helices (α3, α4 and α5) and connective flexible loops, and it is inserted between α2 and α6 of the α/β domain. Remarkable features in Vif structure are shown in β1 for CBFβ binding and zinc coordination motif. β1 in Vif partially interacts with β5 through the hydrogen bond but most of it mediates CBFβ binding by forming the beta-sheet with β3 in CBFβ. A zinc atom coordinated by H108 in the α/β domain and H139, C114 and C133 in the α domain appears to contribute to the stabilization of the tertiary structure of Vif by reducing domain flexibility (Fig. 2b).
a Crystal structure of CUL5NTD-Vif-ELOBC-CBFβ complex. There are twelve complexes in asymmetric unit, one of which is drawn as ribbon diagram (PDB id 4N9F; chain C, D, E, F and G). CUL5NTD, Vif, ELOB, ELOC and CBFβ are labeled and colored green, brown, blue, magenta and yellow, respectively. Zinc atom in Vif structure is shown as a red sphere. Vif directly interacts with CBFβ, CUL5 and ELOC and its interaction with ELOB is not observed. b Vif structure in the CUL5NTD-Vif-ELOBC-CBFβ complex. A zinc atom is drawn as a magenta sphere and zinc-coordination residues are depicted as stick model and the residue numbers are labeled. α-helix (α1–α6), β-strand (β1–β6) and loop were colored red, yellow and green, respectively. The secondary structures, N- and C-terminus are labeled. Vif contains a globular domain that consists of β1–β6, α1, α2 and α6 (α/β domain) and a helical domain inserted between α2 and α6 (α domain)
The APOBEC3 structures
The proteins of APOBEC3 family contain either one (A3A, A3C and A3H) or two cytidine deaminase domains (A3B, A3DE, A3F and A3G). Each domain harbors a highly conserved zinc-coordinating motif, HXE(X)23–28CX2–4C. Two cysteine residues and a histidine coordinate a zinc atom, and a glutamate promotes the formation of a nucleophile (Zinc-hydroxide) for deamination reaction. The structure of full-length APOBEC3 proteins that have two catalytic domains has not been determined yet, probably due to poor protein solubility in vitro. To date, high-resolution structures of A3A, A3C, A3F-CTD (A3F-C terminal domain) and A3G-CTD have been determined. All the APOBEC3 structures have a canonical fold of cytidine deaminase that consists of five beta-strands surrounded by six helices (Fig. 3). The major difference of the folds lies in the second beta strand (β2). The β2 region adopts a continuous single strand or β2-bulge-β2′, even in the structure comparison of the same proteins (Harjes et al. 2009). The conformational difference in β2 might be due to differences of experimental methods or its intrinsic flexibility that allows it to adopt multiple conformations.
The structures of A3 proteins. Atomic coordinate files were obtained from PDB (Protein Data Bank). PDB ID of a–f is 2KBO, 2JYW, 3E1U, 2M65, 3VOW and 4J4J, respectively. All the structures are superimposed and are drawn in the same orientation. α-helix, β-strand and loop are colored differently and the secondary structures are labeled in (a). A zinc atom and conserved zinc coordination residues are drawn as a sphere and magenta stick model, respectively. Dotted lines indicate DNA binding modes proposed by the structures. The residues that are suggested to associate with DNA binding are shown as a stick model around dotted lines. e–f Residues on α2, α3 and α4 are drawn as stick model and labeled in a blue circle, blue rectangle and red circle, respectively. The residues are suggested as patches for Vif binding
Among the APOBEC3 protein family members for which structures are available, A3G-CTD and A3A do not mediate Vif binding but contain a catalytically active domain that mutates consecutive deoxy-cytidine residues in a 3′ to 5′ direction (Beale et al. 2004; Yu et al. 2004a; Chelico et al. 2006; Furukawa et al. 2009). Structures of wild-type A3G-CTD were determined by NMR (residues 193–384; Furukawa et al. 2009) and crystallography (residues 197–380; Holden et al. 2008). Though the structures were determined from the same protein with slightly different sequence boundaries, significant conformational differences are observed, probably due to method differences for structure determination. Based on the difference of surface shape and charge distribution and the assay of NMR chemical shift perturbation, three ssDNA (single strand DNA) binding models of A3G were proposed (Fig. 3a–c). First, Chen et al. determined the NMR structure and observed chemical shift perturbation of A3G-CTD-2K3A, which contains five mutations to improve protein solubility. In the brim-domain model they suggest, ssDNA binds to R213, R215, R313 and R320 aligned along the active site (Fig. 3b; Chen et al. 2008). Second, a similar binding mode is suggested from wild-type A3G-CTD. In addition to chemical shift perturbations observed in A3G-CTD-2K3A, perturbations specific to wild-type A3G were observed on R238, G240, L242 and C243, suggesting slightly different DNA binding surfaces (Fig. 3a; Furukawa et al. 2009). Third, Holden et al. determined the crystal structure of A3G-CTD and observed a surface groove perpendicular to NMR structure-based binding modes, suggesting that the surface groove might be an ssDNA binding site (Fig. 3c; Holden et al. 2008). In addition to DNA binding modes of A3G-CTD, another binding mode was proposed, based on the structure of wild-type A3A protein which does not bind to Vif. Byeon et al. determined the NMR structure of wild-type A3A (Byeon et al. 2013) and observed chemical shift perturbations on the surface of α4 and flexible loops around the active site that may be the ssDNA binding region of A3A (Fig. 3d). Thus, the correct DNA binding site on A3 is somewhat elusive and further experiments are required to identify it.
A3C, A3DE-CTD and A3F-CTD are classified into a different subgroup from A3A and A3G-CTD, based on phylogenetic analysis (LaRue et al. 2009). In the subgroup, crystal structures of A3C and A3F-CTD have been determined (Fig. 3e and f). As A3C and A3F-CTD can directly bind to Vif in contrast to A3G-CTD and A3A, the structures can be useful to identify the Vif binding surface on A3 (Kitamura et al. 2012; Siu et al. 2013; Bohn et al. 2013) as explained below.
Vif-APOBEC3 interaction
Since the structure of Vif-APOBEC3 complex is not available yet, the critical motifs and residues for the Vif-APOBEC3 interaction have been identified through mutagenesis experiments. First, 128-DPD-130 in A3G was identified as Vif binding motif (Mangeat et al. 2004; Schrofelbauer et al. 2004; Bogerd et al. 2004; Santa-Marta et al. 2005; Huthoff and Malim 2007; Zhang et al. 2008; Russell et al. 2009). D128K, P129A and D130K mutants of A3G were not co-immunoprecipitated with Vif and were stably expressed in the presence of Vif, indicating that the mutants are insensitive to Vif (Huthoff and Malim 2007). Second, 289-EFLARH-294 in A3F has been identified as the Vif binding motif (Russell et al. 2009; Smith and Pathak 2010). Especially, E289K mutation in A3F decreased Vif binding in co-immunoprecipitation assays as well as HIV-1 infectivity in the presence of Vif (Smith and Pathak 2010). In the structure model, 128-DPD-130 in A3G lies on the loop between β4 and α4, and 289-EFLARH-294 in A3F forms the C-terminal half of α3 (Fig. 3).
Kitamura et al. determined the crystal structure of A3C and suggested the Vif binding surface on A3C through a mutagenesis assay (Kitamura et al. 2012). Mutations of hydrophobic surface residues on α2 and α3 and negatively charged residues on α3 and α4 reduced Vif binding without affecting virion incorporation of A3C, indicating that the surface areas of α2, α3 and α4 participate in Vif interaction (Fig. 3e). Mutations of conserved residues on A3F-CTD and A3DE-CTD also reduced Vif binding, while those of A3G-NTD did not change Vif sensitivity (Kitamura et al. 2012; Siu et al. 2013). The Vif binding patch on A3F is formed by residues 255–264, 269 and 324, as well as the previously identified residues 289–294. It suggests that the Vif-binding interface conserved among A3C, A3F and A3DE is distinct from that of A3G (Fig. 3e and f).
In addition to Vif binding motifs in APOBEC3 proteins, distinct APOBEC3 binding motifs in Vif have also been suggested through direct binding assays between mutant Vif and APOBEC3 proteins. 40-YRHHY-44 (Russell and Pathak 2007; Yamashita et al. 2008) for A3G binding, 14-DRMR-17 (Russell and Pathak 2007), 74-TGERxW-79 (He et al. 2008) and 171-EDRW-174 (Dang et al. 2010) for A3F binding and 12-QVDRMR-17 for A3C binding (Pery et al. 2009) were identified as specific APOBEC3 binding motifs in Vif. Additionally, mutations of 21-WxSLVK-26 (Dang et al. 2009), 55-VxIPLx4L-64 (Chen et al. 2009) and 81-LGxGxxIxW-89 (Dang et al. 2010) in Vif reduced both A3F and A3G binding. And 69-YxxL-72 in Vif (Pery et al. 2009) was identified as the motif for A3G, A3F and A3C binding.
Because CBFβ binding to Vif facilitates homogeneity of Vif and ubiquitination activity of Vif E3 ligase, the identified A3 binding motifs might be involved in CBFβ binding (Jager et al. 2012; Zhang et al. 2012; Kim et al. 2013). In this respect, it is analyzed whether the binding motifs are surface-exposed for A3 binding or are involved in correct assembly between Vif and CRL5 (Cullin5-RING ligase) containing CUL5, RBX2 and ELOBC, based on the crystal structure of Vif-CUL5NTD-ELOBC-CBFβ complex. First, A3G binding motif, 40-YRHHY-44, is located on the opposite side of CBFβ binding region and the residues are distributed on the Vif surface, indicating that the motif does not overlap with the CBFβ binding site in Vif. Second, A3F binding motifs, 14-DRMR-17, 74-TGERxW-79 and 171-EDRW-174 are also exposed to solvent under CBFβ binding, forming a surface patch near the CBFβ binding region (Fig. 4). In the case of 74-TGERxW-79 located in the loop between β4 and β5, one side is partially involved in CBFβ binding, while the opposite side is exposed to solvent. Third, A3F/A3G binding motifs, 21-WxSLVK-26 and 55-VxIPLx4L-64, are surface motifs that are not buried by CRL5-Vif assembly, while 81-LGxGxxIxW-89 is buried as an internal region of Vif structure. So mutations in 81-LGxGxxIxW-89 motif can disrupt correct protein folding but it does not seem to be a motif for direct binding of A3F/A3G. Fourth, 69-YxxL-72 in Vif for A3C/A3F/A3G binding is located in the loop between β4 and β5. Residues W70 and G71 in the motif are distributed on the surface between A3G and A3F binding motifs, whereas residues Y69 and L72 are involved in CBFβ binding. Taken together, crystal structure of CUL5NTD-Vif-ELOBC-CBFβ complex reveals that the identified A3 binding motifs are partially or fully surface-exposed, except 81-LGxGxxIxW-89 which is an internal region of Vif. Additionally, two A3 binding motifs, 74-TGERxW-79 and 69-YxxL-72, are involved in direct interaction with CBFβ as well as APOBEC3 proteins. In particular, conserved residues Y69 and L72 in 69-YxxL-72 motif are buried by CBFβ binding, indicating that mutations of Y69 and L72 in Vif may reduce APOBEC3 binding by abolishing CBFβ binding but not by disrupting direct interaction with APOBEC3.
A3 binding motifs on Vif. Vif and other proteins in the structure of CUL5NTD-Vif-ELOBC-CBFβ complex are drawn as surface model and ribbon diagram in three different orientations, respectively. The surface structure is colored pink. 14-DRMR-17, 74-TGERxW-79 and 171-EDRW-174 proposed as A3F binding motif are shown in red on the Vif surface model. The other binding motifs, 40-YRHHY-44, 21-WxSLVK-26, 55-VxIPLx4L-64, 69-YxxL-72 and 81-LGxGxxIxW-89 are colored green, cyan, blue, yellow and purple in Vif, respectively. 81-LGxGxxIxW-89 is buried in the surface model as an internal structure
Vif-CBFβ interaction
In the cellular system, CBFβ forms a heterodimer with RUNX protein family members and regulates the transcriptional activity of RUNX proteins as a co-transcription factor (Ogawa et al. 1993). There are three RUNX proteins in mammal, all of which have a conserved Runt domain that mediates both DNA and CBFβ binding (Kagoshima et al. 1993; Tahirov et al. 2001). Three RUNX proteins are associated with mostly developmental processes, including hematopoietic cell, T cell and neuronal cell development and osteoblast differentiation (Okuda et al. 1996; Komori et al. 1997; Otto et al. 1997; Inoue et al. 2002; Levanon et al. 2002; Taniuchi et al. 2002). Recently, direct binding of CBFβ to Vif was found to promote reconstruction of CRL5-Vif and the subsequent poly-ubiquitination of APOBEC3G (Jager et al. 2012; Zhang et al. 2012). Interestingly, co-expression of Vif, CBFβ, ELOB and ELOC in E. coli enables the purification of homogenous Vif complex. In biochemical protein analysis, CBFβ inhibited irregular Vif-oligomerization by binding a hydrophobic surface of Vif (Kim et al. 2013; Guo et al. 2014). The data suggest that CBFβ stabilizes the tertiary structure of Vif without disturbing the assembly of Vif-E3 ligase. The crystal structure of Vif-CUL5NTD-ELOBC-CBFβ complex revealed that a large surface area of Vif and CBFβ mediates their interaction (Figs. 2 and 5a). Binding surfaces between Vif and CBFβ are widely dispersed through residues 1–120 in Vif and whole sequence in CBFβ. More specifically, residues 3–11, 47–50, 69, 72–79, 94–120 and 139 in Vif are involved in direct contact with CBFβ. The first beta strand in Vif (residues 3–11; β1) forms a cross beta sheet with a third beta strand in CBFβ (residues 63–69; β3). And C-terminal region containing α5 and a flexible loop in CBFβ (residues 135–157) binds to a surface pocket near a zinc atom that Vif coordinates (Fig. 5a and c).
The interaction between Vif and CRL5. a The binding interface between Vif and CBFβ. Ribbon diagram of Vif and CBFβ are colored orange and yellow, respectively. N- and C-termini are labeled. β3, which forms a cross beta-sheet with β1 of Vif, and C-terminal loop, which interacts with surface area near the zinc atom in Vif, are colored dark blue in the CBFβ structure. b The binding interface among CUL5, Vif and ELOC. The structures of Vif, CUL5 and ELOC are colored orange, green, and magenta, respectively. Residues that mediate the interaction between Vif/CUL5 (Vif/ELOC and CUL5/ELOC) are colored red (yellow and blue). c Schematic diagram that represents interaction motifs for assembly of CRL5-Vif. The binding motifs and residues are shown as rectangular boxes on a long bar, which indicates primary structures of each protein, and the sequences of binding motifs are labeled. The same colored motifs indicate a binding interface between the two proteins. For example, the binding motifs for binding between Vif and CBFβ are colored dark blue and those for binding of ELOC and CUL5 are colored green
In the structure of Vif-CUL5NTD-ELOBC-CBFβ complex, the Vif binding surface on CBFβ completely overlaps the RUNX binding surface (Tahirov et al. 2001; Guo et al. 2014). It coincides with previously reported data that Vif and RUNX are mutually exclusive for CBFβ binding (Kim et al. 2013). Kim et al. observed that purified complex of Vif-ELOBC-CBFβ does not interact with Runt1, Runt2 and Runt3 in vitro. Indeed, the transcriptional activity of RUNX1 was reduced when Vif was expressed in a human cell line, and the expression of RUNX1 target genes related to immune response were regulated by Vif expression or HIV infection in the activated T-cell lines (Kim et al. 2013). It suggests that CBFβ hijacking by Vif can result in the regulation of RUNX transcriptional activity as well as the stabilization of Vif structure.
Vif-CUL5-ELOBC interaction
The proteins of Cullin family have elongated, curved and rigid structures formed by three N-terminal helical bundles and a C-terminal globular domain. The first helical bundle at the N-terminus binds substrate adaptor proteins for recruitment of specific substrates, and the globular C-terminal domain mediates the interaction with RING subunit for ubiquitin E2 recruitment (Zheng et al. 2002). CUL2 and CUL5 of Cullin family interact with ELOC adaptor and an additional substrate receptor such as BC box protein to recruit specific substrates. Suppressor of cytokine signaling (SOCS) proteins containing BC box are well-known substrate receptors for the assembly of CRL2 (Cullin2-RING ligase) and CRL5. CRL activation additionally requires the modification of Cullin proteins called neddylation. Covalent attachment of ubiquitin-like protein NEDD8 to Cullin proteins enhances the enzyme activity. A3G ubiquitination by CRL5-Vif is also controlled by NEDD8 attachment (Petroski and Deshaies 2005; Stanley et al. 2012).
HIV-1 Vif recruits the components of CRL5 containing CUL5, RBX2, ELOB and ELOC, to form ubiquitin E3 ligase (Yu et al. 2003; Jager et al. 2012). Like SOCS proteins, it contains a conserved BC box motif (residues 144–158) for ELOBC binding. Deletion of the BC box in Vif abolishes co-immunoprecipitation with ELOBC (Mehle et al. 2004). Crystal structure of Vif BC box complexed with ELOBC (Stanley et al. 2008) shows that hydrophobic residues in Vif BC box participate in ELOC binding, similarly to BC box in cellular SOCS proteins (Bullock et al. 2006, 2007).
Several research groups identified that the HCCH zinc-coordination motif in Vif (residues 100–142 including Hx5Cx17–18Cx3–5H) is critical for selective CUL5 binding (Yu et al. 2004b; Luo et al. 2005; Mehle et al. 2006; Paul et al. 2006; Xiao et al. 2006, 2007a, b). Mutations of conserved hydrophobic residues (I120, A123 and L124) located between two cysteines in the HCCH motif disrupt CUL5 binding ability (Mehle et al. 2006). Consistent with the biochemical data, the structure of Vif-CUL5NTD-ELOBC-CBFβ complex clearly shows a binding interface between Vif and CUL5 (Fig. 5b and c). Third alpha helix (α3) of Vif (residues 121-127) is involved in direct interaction with 52-LWDD-55 of CUL5. The Vif binding motif in CUL5 is not conserved in CUL2, suggesting selective recruitment of CUL5 by Vif for E3 ligase assembly or higher binding affinity with CUL5. Another potential CUL5 binding motif, CUL box (residues 161–169 in Vif), which has been shown to mediate interaction with CUL5 in cellular SOCS-box proteins appears not to participate in any interaction for CRL5-Vif assembly (Kamura et al. 2004).
In addition to Vif-mediated interaction, CUL5 and ELOC bind each other directly. The second helix in CUL5 and additional residues K109 and Q113 mediate ELOC binding. Vif, CUL5 and ELOC seem to interact cooperatively for complex stability and selective assembly (Fig. 5b and c).
Perspectives
HIV-1 Vif recruits A3 proteins to the ubiquitin–proteasome system to counteract the antiviral activity of APOBEC3 proteins. Because the disruption of CRL5-Vif assembly and Vif/A3 interaction can reduce HIV-1 infectivity, their binding surface has been extensively studied as an antiviral target. Currently, multiple binding motifs for CRL5-Vif assembly and Vif/A3 interaction have been suggested and the structures of A3 proteins (A3A, A3C, A3F-CTD and A3G-CTD) and Vif-CUL5NTD-ELOBC-CBFβ complex are available. In this review, the binding motifs were summarized based on available structures. Even though the understanding of CRL5-Vif/A3G assembly has been advanced greatly in the last decade, high-resolution structures of Vif-A3 complex are still required for the identification of exact binding motifs. Additionally, a DNA binding mode of A3 proteins is not clear, even though several research groups have suggested it. Thus, structural studies of the complexes (Vif-A3 and A3-DNA) might be the next challenge to understand the functions of antiviral proteins (A3) and anti-antiviral protein (Vif).
References
Aguiar, R.S., N. Lovsin, A. Tanuri, and B.M. Peterlin. 2008. Vpr. A3A chimera inhibits HIV replication. The Journal of Biological Chemistry 283: 2518–2525.
Beale, R.C., S.K. Petersen-Mahrt, I.N. Watt, R.S. Harris, C. Rada, and M.S. Neuberger. 2004. Comparison of the differential context-dependence of DNA deamination by APOBEC enzymes: correlation with mutation spectra in vivo. Journal of Molecular Biology 337: 585–596.
Belanger, K., M. Savoie, M.C. Rosales Gerpe, J.F. Couture, and M.A. Langlois. 2013. Binding of RNA by APOBEC3G controls deamination-independent restriction of retroviruses. Nucleic Acids Research 41: 7438–7452.
Bishop, K.N., R.K. Holmes, A.M. Sheehy, N.O. Davidson, S.J. Cho, and M.H. Malim. 2004. Cytidine deamination of retroviral DNA by diverse APOBEC proteins. Current Biology 14: 1392–1396.
Bishop, K.N., M. Verma, E.Y. Kim, S.M. Wolinsky, and M.H. Malim. 2008. APOBEC3G inhibits elongation of HIV-1 reverse transcripts. PLoS Pathogens 4: e1000231.
Bogerd, H.P., B.P. Doehle, H.L. Wiegand, and B.R. Cullen. 2004. A single amino acid difference in the host APOBEC3G protein controls the primate species specificity of HIV type 1 virion infectivity factor. Proceedings of the National Academy of Sciences of the United States of America 101: 3770–3774.
Bohn, M.F., S.M. Shandilya, J.S. Albin, T. Kouno, B.D. Anderson, R.M. Mcdougle, M.A. Carpenter, A. Rathore, L. Evans, A.N. Davis, J. Zhang, Y. Lu, M. Somasundaran, H. Matsuo, R.S. Harris, and C.A. Schiffer. 2013. Crystal structure of the DNA cytosine deaminase APOBEC3F: the catalytically active and HIV-1 Vif-binding domain. Structure 21: 1042–1050.
Bouyac, M., F. Rey, M. Nascimbeni, M. Courcoul, J. Sire, D. Blanc, F. Clavel, R. Vigne, and B. Spire. 1997. Phenotypically Vif- human immunodeficiency virus type 1 is produced by chronically infected restrictive cells. Journal of Virology 71: 2473–2477.
Bullock, A.N., J.E. Debreczeni, A.M. Edwards, M. Sundstrom, and S. Knapp. 2006. Crystal structure of the SOCS2-elongin C-elongin B complex defines a prototypical SOCS box ubiquitin ligase. Proceedings of the National Academy of Sciences of the United States of America 103: 7637–7642.
Bullock, A.N., M.C. Rodriguez, J.E. Debreczeni, Z. Songyang, and S. Knapp. 2007. Structure of the SOCS4-ElonginB/C complex reveals a distinct SOCS box interface and the molecular basis for SOCS-dependent EGFR degradation. Structure 15: 1493–1504.
Byeon, I.J., J. Ahn, M. Mitra, C.H. Byeon, K. Hercik, J. Hritz, L.M. Charlton, J.G. Levin, and A.M. Gronenborn. 2013. NMR structure of human restriction factor APOBEC3A reveals substrate binding and enzyme specificity. Nature Communications 4: 1890.
Chaipan, C., J.L. Smith, W.S. Hu, and V.K. Pathak. 2013. APOBEC3G restricts HIV-1 to a greater extent than APOBEC3F and APOBEC3DE in human primary CD4+ T cells and macrophages. Journal of Virology 87: 444–453.
Chelico, L., P. Pham, P. Calabrese, and M.F. Goodman. 2006. APOBEC3G DNA deaminase acts processively 3′ → 5′ on single-stranded DNA. Nature Structural & Molecular Biology 13: 392–399.
Chen, G., Z. He, T. Wang, R. Xu, and X.F. Yu. 2009. A patch of positively charged amino acids surrounding the human immunodeficiency virus type 1 Vif SLVx4Yx9Y motif influences its interaction with APOBEC3G. Journal of Virology 83: 8674–8682.
Chen, K.M., E. Harjes, P.J. Gross, A. Fahmy, Y. Lu, K. Shindo, R.S. Harris, and H. Matsuo. 2008. Structure of the DNA deaminase domain of the HIV-1 restriction factor APOBEC3G. Nature 452: 116–119.
Conticello, S.G., R.S. Harris, and M.S. Neuberger. 2003. The Vif protein of HIV triggers degradation of the human antiretroviral DNA deaminase APOBEC3G. Current Biology 13: 2009–2013.
Cullen, B.R. 2003. Nuclear mRNA export: Insights from virology. Trends in Biochemical Sciences 28: 419–424.
Dang, Y., R.W. Davis, I.A. York, and Y.H. Zheng. 2010. Identification of 81LGxGxxIxW89 and 171EDRW174 domains from human immunodeficiency virus type 1 Vif that regulate APOBEC3G and APOBEC3F neutralizing activity. Journal of Virology 84: 5741–5750.
Dang, Y., X. Wang, W.J. Esselman, and Y.H. Zheng. 2006. Identification of APOBEC3DE as another antiretroviral factor from the human APOBEC family. Journal of Virology 80: 10522–10533.
Dang, Y., X. Wang, T. Zhou, I.A. York, and Y.H. Zheng. 2009. Identification of a novel WxSLVK motif in the N terminus of human immunodeficiency virus and simian immunodeficiency virus Vif that is critical for APOBEC3G and APOBEC3F neutralization. Journal of Virology 83: 8544–8552.
Desimmie, B.A., K.A. Delviks-Frankenberrry, R.C. Burdick, D. Qi, T. Izumi, and V.K. Pathak. 2014. Multiple APOBEC3 restriction factors for HIV-1 and one Vif to rule them all. Journal of Molecular Biology 426: 1220–1245.
Doehle, B.P., A. Schafer, and B.R. Cullen. 2005. Human APOBEC3B is a potent inhibitor of HIV-1 infectivity and is resistant to HIV-1 Vif. Virology 339: 281–288.
Fisher, A.G., B. Ensoli, L. Ivanoff, M. Chamberlain, S. Petteway, L. Ratner, R.C. Gallo, and F. Wong-Staal. 1987. The sor gene of HIV-1 is required for efficient virus transmission in vitro. Science 237: 888–893.
Furukawa, A., T. Nagata, A. Matsugami, Y. Habu, R. Sugiyama, F. Hayashi, N. Kobayashi, S. Yokoyama, H. Takaku, and M. Katahira. 2009. Structure, interaction and real-time monitoring of the enzymatic reaction of wild-type APOBEC3G. The EMBO Journal 28: 440–451.
Gabuzda, D.H., K. Lawrence, E. Langhoff, E. Terwilliger, T. Dorfman, W.A. Haseltine, and J. Sodroski. 1992. Role of vif in replication of human immunodeficiency virus type 1 in CD4+ T lymphocytes. Journal of Virology 66: 6489–6495.
Guo, F., S. Cen, M. Niu, J. Saadatmand, and L. Kleiman. 2006. Inhibition of formula-primed reverse transcription by human APOBEC3G during human immunodeficiency virus type 1 replication. Journal of Virology 80: 11710–11722.
Guo, F., S. Cen, M. Niu, Y. Yang, R.J. Gorelick, and L. Kleiman. 2007. The interaction of APOBEC3G with human immunodeficiency virus type 1 nucleocapsid inhibits tRNA3Lys annealing to viral RNA. Journal of Virology 81: 11322–11331.
Guo, Y., L. Dong, X. Qiu, Y. Wang, B. Zhang, H. Liu, Y. Yu, Y. Zang, M. Yang, and Z. Huang. 2014. Structural basis for hijacking CBF-beta and CUL5 E3 ligase complex by HIV-1 Vif. Nature 505: 229–233.
Harjes, E., P.J. Gross, K.M. Chen, Y. Lu, K. Shindo, R. Nowarski, J.D. Gross, M. Kotler, R.S. Harris, and H. Matsuo. 2009. An extended structure of the APOBEC3G catalytic domain suggests a unique holoenzyme model. Journal of Molecular Biology 389: 819–832.
Harris, R.S., K.N. Bishop, A.M. Sheehy, H.M. Craig, S.K. Petersen-Mahrt, I.N. Watt, M.S. Neuberger, and M.H. Malim. 2003. DNA deamination mediates innate immunity to retroviral infection. Cell 113: 803–809.
He, Z., W. Zhang, G. Chen, R. Xu, and X.F. Yu. 2008. Characterization of conserved motifs in HIV-1 Vif required for APOBEC3G and APOBEC3F interaction. Journal of Molecular Biology 381: 1000–1011.
Holden, L.G., C. Prochnow, Y.P. Chang, R. Bransteitter, L. Chelico, U. Sen, R.C. Stevens, M.F. Goodman, and X.S. Chen. 2008. Crystal structure of the anti-viral APOBEC3G catalytic domain and functional implications. Nature 456: 121–124.
Huthoff, H., and M.H. Malim. 2007. Identification of amino acid residues in APOBEC3G required for regulation by human immunodeficiency virus type 1 Vif and Virion encapsidation. Journal of Virology 81: 3807–3815.
Inoue, K., S. Ozaki, T. Shiga, K. Ito, T. Masuda, N. Okado, T. Iseda, S. Kawaguchi, M. Ogawa, S.C. Bae, N. Yamashita, S. Itohara, N. Kudo, and Y. Ito. 2002. Runx3 controls the axonal projection of proprioceptive dorsal root ganglion neurons. Nature Neuroscience 5: 946–954.
Iwatani, Y., D.S. Chan, F. Wang, K.S. Maynard, W. Sugiura, A.M. Gronenborn, I. Rouzina, M.C. Williams, K. Musier-Forsyth, and J.G. Levin. 2007. Deaminase-independent inhibition of HIV-1 reverse transcription by APOBEC3G. Nucleic Acids Research 35: 7096–7108.
Jager, S., D.Y. Kim, J.F. Hultquist, K. Shindo, R.S. Larue, E. Kwon, M. Li, B.D. Anderson, L. Yen, D. Stanley, C. Mahon, J. Kane, K. Franks-Skiba, P. Cimermancic, A. Burlingame, A. Sali, C.S. Craik, R.S. Harris, J.D. Gross, and N.J. Krogan. 2012. Vif hijacks CBF-beta to degrade APOBEC3G and promote HIV-1 infection. Nature 481: 371–375.
Jarmuz, A., A. Chester, J. Bayliss, J. Gisbourne, I. Dunham, J. Scott, and N. Navaratnam. 2002. An anthropoid-specific locus of orphan C to U RNA-editing enzymes on chromosome 22. Genomics 79: 285–296.
Kagoshima, H., K. Shigesada, M. Satake, Y. Ito, H. Miyoshi, M. Ohki, M. Pepling, and P. Gergen. 1993. The Runt domain identifies a new family of heteromeric transcriptional regulators. Trends in Genetics 9: 338–341.
Kamura, T., K. Maenaka, S. Kotoshiba, M. Matsumoto, D. Kohda, R.C. Conaway, J.W. Conaway, and K.I. Nakayama. 2004. VHL-box and SOCS-box domains determine binding specificity for Cul2-Rbx1 and Cul5-Rbx2 modules of ubiquitin ligases. Genes & Development 18: 3055–3065.
Kim, D.Y., E. Kwon, P.D. Hartley, D.C. Crosby, S. Mann, N.J. Krogan, and J.D. Gross. 2013. CBFbeta stabilizes HIV Vif to counteract APOBEC3 at the expense of RUNX1 target gene expression. Molecular Cell 49: 632–644.
Kitamura, S., H. Ode, M. Nakashima, M. Imahashi, Y. Naganawa, T. Kurosawa, Y. Yokomaku, T. Yamane, N. Watanabe, A. Suzuki, W. Sugiura, and Y. Iwatani. 2012. The APOBEC3C crystal structure and the interface for HIV-1 Vif binding. Nature Structural & Molecular Biology 19: 1005–1010.
Komori, T., H. Yagi, S. Nomura, A. Yamaguchi, K. Sasaki, K. Deguchi, Y. Shimizu, R.T. Bronson, Y.H. Gao, M. Inada, M. Sato, R. Okamoto, Y. Kitamura, S. Yoshiki, and T. Kishimoto. 1997. Targeted disruption of Cbfa1 results in a complete lack of bone formation owing to maturational arrest of osteoblasts. Cell 89: 755–764.
Koning, F.A., E.N. Newman, E.Y. Kim, K.J. Kunstman, S.M. Wolinsky, and M.H. Malim. 2009. Defining APOBEC3 expression patterns in human tissues and hematopoietic cell subsets. Journal of Virology 83: 9474–9485.
Langlois, M.A., R.C. Beale, S.G. Conticello, and M.S. Neuberger. 2005. Mutational comparison of the single-domained APOBEC3C and double-domained APOBEC3F/G anti-retroviral cytidine deaminases provides insight into their DNA target site specificities. Nucleic Acids Research 33: 1913–1923.
Larue, R.S., V. Andresdottir, Y. Blanchard, S.G. Conticello, D. Derse, M. Emerman, W.C. Greene, S.R. Jonsson, N.R. Landau, M. Lochelt, H.S. Malik, M.H. Malim, C. Munk, S.J. O’brien, V.K. Pathak, K. Strebel, S. Wain-Hobson, X.F. Yu, N. Yuhki, and R.S. Harris. 2009. Guidelines for naming nonprimate APOBEC3 genes and proteins. Journal of Virology 83: 494–497.
Lecossier, D., F. Bouchonnet, F. Clavel, and A.J. Hance. 2003. Hypermutation of HIV-1 DNA in the absence of the Vif protein. Science 300: 1112.
Levanon, D., D. Bettoun, C. Harris-Cerruti, E. Woolf, V. Negreanu, R. Eilam, Y. Bernstein, D. Goldenberg, C. Xiao, M. Fliegauf, E. Kremer, F. Otto, O. Brenner, A. Lev-Tov, and Y. Groner. 2002. The Runx3 transcription factor regulates development and survival of TrkC dorsal root ganglia neurons. The EMBO Journal 21: 3454–3463.
Li, X.Y., F. Guo, L. Zhang, L. Kleiman, and S. Cen. 2007. APOBEC3G inhibits DNA strand transfer during HIV-1 reverse transcription. The Journal of Biological Chemistry 282: 32065–32074.
Luo, K., T. Wang, B. Liu, C. Tian, Z. Xiao, J. Kappes, and X.F. Yu. 2007. Cytidine deaminases APOBEC3G and APOBEC3F interact with human immunodeficiency virus type 1 integrase and inhibit proviral DNA formation. Journal of Virology 81: 7238–7248.
Luo, K., Z. Xiao, E. Ehrlich, Y. Yu, B. Liu, S. Zheng, and X.F. Yu. 2005. Primate lentiviral virion infectivity factors are substrate receptors that assemble with cullin 5-E3 ligase through a HCCH motif to suppress APOBEC3G. Proceedings of the National Academy of Sciences of the United States of America 102: 11444–11449.
Mangeat, B., P. Turelli, G. Caron, M. Friedli, L. Perrin, and D. Trono. 2003. Broad antiretroviral defence by human APOBEC3G through lethal editing of nascent reverse transcripts. Nature 424: 99–103.
Mangeat, B., P. Turelli, S. Liao, and D. Trono. 2004. A single amino acid determinant governs the species-specific sensitivity of APOBEC3G to Vif action. The Journal of Biological Chemistry 279: 14481–14483.
Marcsisin, S.R., P.S. Narute, L.A. Emert-Sedlak, M. Kloczewiak, T.E. Smithgall, and J.R. Engen. 2011. On the solution conformation and dynamics of the HIV-1 viral infectivity factor. Journal of Molecular Biology 410: 1008–1022.
Mariani, R., D. Chen, B. Schrofelbauer, F. Navarro, R. Konig, B. Bollman, C. Munk, H. Nymark-Mcmahon, and N.R. Landau. 2003. Species-specific exclusion of APOBEC3G from HIV-1 virions by Vif. Cell 114: 21–31.
Marin, M., K.M. Rose, S.L. Kozak, and D. Kabat. 2003. HIV-1 Vif protein binds the editing enzyme APOBEC3G and induces its degradation. Nature Medicine 9: 1398–1403.
Mbisa, J.L., R. Barr, J.A. Thomas, N. Vandegraaff, I.J. Dorweiler, E.S. Svarovskaia, W.L. Brown, L.M. Mansky, R.J. Gorelick, R.S. Harris, A. Engelman, and V.K. Pathak. 2007. Human immunodeficiency virus type 1 cDNAs produced in the presence of APOBEC3G exhibit defects in plus-strand DNA transfer and integration. Journal of Virology 81: 7099–7110.
Mehle, A., J. Goncalves, M. Santa-Marta, M. Mcpike, and D. Gabuzda. 2004. Phosphorylation of a novel SOCS-box regulates assembly of the HIV-1 Vif-Cul5 complex that promotes APOBEC3G degradation. Genes & Development 18: 2861–2866.
Mehle, A., E.R. Thomas, K.S. Rajendran, and D. Gabuzda. 2006. A zinc-binding region in Vif binds Cul5 and determines cullin selection. The Journal of Biological Chemistry 281: 17259–17265.
Miyagi, E., C.R. Brown, S. Opi, M. Khan, R. Goila-Gaur, S. Kao, R.C. Walker Jr, V. Hirsch, and K. Strebel. 2010. Stably expressed APOBEC3F has negligible antiviral activity. Journal of Virology 84: 11067–11075.
Mulder, L.C., M. Ooms, S. Majdak, J. Smedresman, C. Linscheid, A. Harari, A. Kunz, and V. Simon. 2010. Moderate influence of human APOBEC3F on HIV-1 replication in primary lymphocytes. Journal of Virology 84: 9613–9617.
Neil, S.J., T. Zang, and P.D. Bieniasz. 2008. Tetherin inhibits retrovirus release and is antagonized by HIV-1 Vpu. Nature 451: 425–430.
Newman, E.N., R.K. Holmes, H.M. Craig, K.C. Klein, J.R. Lingappa, M.H. Malim, and A.M. Sheehy. 2005. Antiviral function of APOBEC3G can be dissociated from cytidine deaminase activity. Current Biology 15: 166–170.
Ogawa, E., M. Inuzuka, M. Maruyama, M. Satake, M. Naito-Fujimoto, Y. Ito, and K. Shigesada. 1993. Molecular cloning and characterization of PEBP2 beta, the heterodimeric partner of a novel Drosophila runt-related DNA binding protein PEBP2 alpha. Virology 194: 314–331.
Okuda, T., J. Van Deursen, S.W. Hiebert, G. Grosveld, and J.R. Downing. 1996. AML1, the target of multiple chromosomal translocations in human leukemia, is essential for normal fetal liver hematopoiesis. Cell 84: 321–330.
Ott, M., M. Geyer, and Q. Zhou. 2011. The control of HIV transcription: keeping RNA polymerase II on track. Cell Host & Microbe 10: 426–435.
Otto, F., A.P. Thornell, T. Crompton, A. Denzel, K.C. Gilmour, I.R. Rosewell, G.W. Stamp, R.S. Beddington, S. Mundlos, B.R. Olsen, P.B. Selby, and M.J. Owen. 1997. Cbfa1, a candidate gene for cleidocranial dysplasia syndrome, is essential for osteoblast differentiation and bone development. Cell 89: 765–771.
Paul, I., J. Cui, and E.L. Maynard. 2006. Zinc binding to the HCCH motif of HIV-1 virion infectivity factor induces a conformational change that mediates protein-protein interactions. Proceedings of the National Academy of Sciences of the United States of America 103: 18475–18480.
Perez-Caballero, D., T. Zang, A. Ebrahimi, M.W. Mcnatt, D.A. Gregory, M.C. Johnson, and P.D. Bieniasz. 2009. Tetherin inhibits HIV-1 release by directly tethering virions to cells. Cell 139: 499–511.
Pery, E., K.S. Rajendran, A.J. Brazier, and D. Gabuzda. 2009. Regulation of APOBEC3 proteins by a novel YXXL motif in human immunodeficiency virus type 1 Vif and simian immunodeficiency virus SIVagm Vif. Journal of Virology 83: 2374–2381.
Petroski, M.D., and R.J. Deshaies. 2005. Function and regulation of cullin-RING ubiquitin ligases. Nature Reviews Molecular Cell Biology 6: 9–20.
Russell, R.A., and V.K. Pathak. 2007. Identification of two distinct human immunodeficiency virus type 1 Vif determinants critical for interactions with human APOBEC3G and APOBEC3F. Journal of Virology 81: 8201–8210.
Russell, R.A., J. Smith, R. Barr, D. Bhattacharyya, and V.K. Pathak. 2009. Distinct domains within APOBEC3G and APOBEC3F interact with separate regions of human immunodeficiency virus type 1 Vif. Journal of Virology 83: 1992–2003.
Sakai, H., R. Shibata, J. Sakuragi, S. Sakuragi, M. Kawamura, and A. Adachi. 1993. Cell-dependent requirement of human immunodeficiency virus type 1 Vif protein for maturation of virus particles. Journal of Virology 67: 1663–1666.
Santa-Marta, M., F.A. Da Silva, A.M. Fonseca, and J. Goncalves. 2005. HIV-1 Vif can directly inhibit apolipoprotein B mRNA-editing enzyme catalytic polypeptide-like 3G-mediated cytidine deamination by using a single amino acid interaction and without protein degradation. The Journal of Biological Chemistry 280: 8765–8775.
Schrofelbauer, B., D. Chen, and N.R. Landau. 2004. A single amino acid of APOBEC3G controls its species-specific interaction with virion infectivity factor (Vif). Proceedings of the National Academy of Sciences of the United States of America 101: 3927–3932.
Sheehy, A.M., N.C. Gaddis, J.D. Choi, and M.H. Malim. 2002. Isolation of a human gene that inhibits HIV-1 infection and is suppressed by the viral Vif protein. Nature 418: 646–650.
Sheehy, A.M., N.C. Gaddis, and M.H. Malim. 2003. The antiretroviral enzyme APOBEC3G is degraded by the proteasome in response to HIV-1 Vif. Nature Medicine 9: 1404–1407.
Simon, J.H., N.C. Gaddis, R.A. Fouchier, and M.H. Malim. 1998. Evidence for a newly discovered cellular anti-HIV-1 phenotype. Nature Medicine 4: 1397–1400.
Siu, K.K., A. Sultana, F.C. Azimi, and J.E. Lee. 2013. Structural determinants of HIV-1 Vif susceptibility and DNA binding in APOBEC3F. Nature Communications 4: 2593.
Smith, J.L., and V.K. Pathak. 2010. Identification of specific determinants of human APOBEC3F, APOBEC3C, and APOBEC3DE and African green monkey APOBEC3F that interact with HIV-1 Vif. Journal of Virology 84: 12599–12608.
Sova, P., and D.J. Volsky. 1993. Efficiency of viral DNA synthesis during infection of permissive and nonpermissive cells with vif-negative human immunodeficiency virus type 1. Journal of Virology 67: 6322–6326.
Stanley, B.J., E.S. Ehrlich, L. Short, Y. Yu, Z. Xiao, X.F. Yu, and Y. Xiong. 2008. Structural insight into the human immunodeficiency virus Vif SOCS box and its role in human E3 ubiquitin ligase assembly. Journal of Virology 82: 8656–8663.
Stanley, D.J., K. Bartholomeeusen, D.C. Crosby, D.Y. Kim, E. Kwon, L. Yen, N.C. Cartozo, M. Li, S. Jager, J. Mason-Herr, F. Hayashi, S. Yokoyama, N.J. Krogan, R.S. Harris, B.M. Peterlin, and J.D. Gross. 2012. Inhibition of a NEDD8 cascade restores restriction of HIV by APOBEC3G. PLoS Pathogens 8: e1003085.
Stopak, K., C. De Noronha, W. Yonemoto, and W.C. Greene. 2003. HIV-1 Vif blocks the antiviral activity of APOBEC3G by impairing both its translation and intracellular stability. Molecular Cell 12: 591–601.
Strebel, K. 2013. HIV accessory proteins versus host restriction factors. Current opinion in Virology 3: 692–699.
Tahirov, T.H., T. Inoue-Bungo, H. Morii, A. Fujikawa, M. Sasaki, K. Kimura, M. Shiina, K. Sato, T. Kumasaka, M. Yamamoto, S. Ishii, and K. Ogata. 2001. Structural analyses of DNA recognition by the AML1/Runx-1 Runt domain and its allosteric control by CBFbeta. Cell 104: 755–767.
Taniuchi, I., M. Osato, T. Egawa, M.J. Sunshine, S.C. Bae, T. Komori, Y. Ito, and D.R. Littman. 2002. Differential requirements for Runx proteins in CD4 repression and epigenetic silencing during T lymphocyte development. Cell 111: 621–633.
Van Damme, N., D. Goff, C. Katsura, R.L. Jorgenson, R. Mitchell, M.C. Johnson, E.B. Stephens, and J. Guatelli. 2008. The interferon-induced protein BST-2 restricts HIV-1 release and is downregulated from the cell surface by the viral Vpu protein. Cell Host & Microbe 3: 245–252.
Von Schwedler, U., J. Song, C. Aiken, and D. Trono. 1993. Vif is crucial for human immunodeficiency virus type 1 proviral DNA synthesis in infected cells. Journal of Virology 67: 4945–4955.
Wang, X., Z. Ao, L. Chen, G. Kobinger, J. Peng, and X. Yao. 2012. The cellular antiviral protein APOBEC3G interacts with HIV-1 reverse transcriptase and inhibits its function during viral replication. Journal of Virology 86: 3777–3786.
Wiegand, H.L., B.P. Doehle, H.P. Bogerd, and B.R. Cullen. 2004. A second human antiretroviral factor, APOBEC3F, is suppressed by the HIV-1 and HIV-2 Vif proteins. The EMBO Journal 23: 2451–2458.
Xiao, Z., E. Ehrlich, K. Luo, Y. Xiong, and X.F. Yu. 2007a. Zinc chelation inhibits HIV Vif activity and liberates antiviral function of the cytidine deaminase APOBEC3G. FASEB Journal 21: 217–222.
Xiao, Z., E. Ehrlich, Y. Yu, K. Luo, T. Wang, C. Tian, and X.F. Yu. 2006. Assembly of HIV-1 Vif-Cul5 E3 ubiquitin ligase through a novel zinc-binding domain-stabilized hydrophobic interface in Vif. Virology 349: 290–299.
Xiao, Z., Y. Xiong, W. Zhang, L. Tan, E. Ehrlich, D. Guo, and X.F. Yu. 2007b. Characterization of a novel Cullin5 binding domain in HIV-1 Vif. Journal of Molecular Biology 373: 541–550.
Yamashita, T., K. Kamada, K. Hatcho, A. Adachi, and M. Nomaguchi. 2008. Identification of amino acid residues in HIV-1 Vif critical for binding and exclusion of APOBEC3G/F. Microbes and infection/Institut Pasteur 10: 1142–1149.
Yu, X., Y. Yu, B. Liu, K. Luo, W. Kong, P. Mao, and X.F. Yu. 2003. Induction of APOBEC3G ubiquitination and degradation by an HIV-1 Vif-Cul5-SCF complex. Science 302: 1056–1060.
Yu, Q., R. Konig, S. Pillai, K. Chiles, M. Kearney, S. Palmer, D. Richman, J.M. Coffin, and N.R. Landau. 2004a. Single-strand specificity of APOBEC3G accounts for minus-strand deamination of the HIV genome. Nature Structural & Molecular Biology 11: 435–442.
Yu, Y., Z. Xiao, E.S. Ehrlich, X. Yu, and X.F. Yu. 2004b. Selective assembly of HIV-1 Vif-Cul5-ElonginB-ElonginC E3 ubiquitin ligase complex through a novel SOCS box and upstream cysteines. Genes & Development 18: 2867–2872.
Zhang, H., B. Yang, R.J. Pomerantz, C. Zhang, S.C. Arunachalam, and L. Gao. 2003. The cytidine deaminase CEM15 induces hypermutation in newly synthesized HIV-1 DNA. Nature 424: 94–98.
Zhang, W., G. Chen, A.M. Niewiadomska, R. Xu, and X.F. Yu. 2008. Distinct determinants in HIV-1 Vif and human APOBEC3 proteins are required for the suppression of diverse host anti-viral proteins. PLoS One 3: e3963.
Zhang, W., J. Du, S.L. Evans, Y. Yu, and X.F. Yu. 2012. T-cell differentiation factor CBF-beta regulates HIV-1 Vif-mediated evasion of host restriction. Nature 481: 376–379.
Zhen, A., T. Wang, K. Zhao, Y. Xiong, and X.F. Yu. 2010. A single amino acid difference in human APOBEC3H variants determines HIV-1 Vif sensitivity. Journal of Virology 84: 1902–1911.
Zheng, N., B.A. Schulman, L. Song, J.J. Miller, P.D. Jeffrey, P. Wang, C. Chu, D.M. Koepp, S.J. Elledge, M. Pagano, R.C. Conaway, J.W. Conaway, J.W. Harper, and N.P. Pavletich. 2002. Structure of the Cul1-Rbx1-Skp1-F boxSkp2 SCF ubiquitin ligase complex. Nature 416: 703–709.
Acknowledgments
This work was supported by the 2013 Yeungnam University Research Grant, Korea Basic Science Institute Grant (T34415, H.S.Jung) and Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Science, ICT & Future Planning (NRF-2014R1A1A1002064).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kim, D.Y. The assembly of Vif ubiquitin E3 ligase for APOBEC3 degradation. Arch. Pharm. Res. 38, 435–445 (2015). https://doi.org/10.1007/s12272-014-0519-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12272-014-0519-x