Abstract
In some situations, there is a need for rapid mutation tests for guiding clinical decisions and starting targeted therapies with minimal delays. In this study we evaluated the turnaround time before and after the implementation of a fully automated multiplex assay for KRAS and NRAS/BRAF mutation tests (Idylla™ platform, Biocartis) in metastatic colorectal cancer. The objective of this project was to compare the turnaround times in 2017–2018 with the fully automated multiplex assay to the 2016 results with previous methods. Centers with a number of tests for metastatic colorectal cancer > 100 yearly and a usual turnaround time ≥ 3 weeks for mutation detection were selected. Results of 505 KRAS tests and 369 NRAS/BRAF tests were transmitted by 10 centers. The mean turnaround time from test prescription to reception of results was reduced from 25.8 days in 2016 to 4.5 days in 2017–2018. In conclusion, this pilot project shows that the Idylla™ platform for testing KRAS and NRAS/BRAF mutations allows an optimized turnaround time from test prescription to reception of results.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Colorectal cancer is the 2nd most common cancer in Europe and the 4th most common worldwide [1, 2]. For many years, palliative treatment of colorectal cancer rested on chemotherapy. Recently, the combination of chemotherapy and targeted therapies with human vascular endothelial growth factor (VEGF) or epidermal growth factor receptor (EGFR) monoclonal antibodies improved treatment response and patient survival. Thus, median overall survival in patients with metastatic disease increased from 8 to 12 months to 21–24 months with panitumumab, a monoclonal antibody that targets EGFR [3].
EGFR and its downstream signaling pathway are involved in the development of several tumors including colorectal cancer [4]. The growth of cancer cells is driven by the activation of signaling pathways including the RAS-RAF-BRAF-MAPK and the PI3P-Akt pathways [5]. The monoclonal antibodies panitumumab or cetuximab are efficient on colorectal cancer by disrupting the EGFR signal. A chief condition for the efficacy of these targeted therapies is the absence of mutation in the EGFR signaling pathways [6]. Thus, the presence of mutations in KRAS exon 2 (codon 12 and 13) was shown to reduce strongly the efficacy of panitumumab or cetuximab [7, 8]. Therefore, the exon 2 mutation was used a biomarker to select patients who could benefit from the targeted therapy.
Other mutations causing resistance to anti-EGFR therapy have been evidenced: mutation in KRAS exon 3 (codons 59 and 61) and exon 4 (codons 117 and 146) and in NRAS exon 2 (codons 12 and 13), exon 3 (codons 59 and 61) and exon 4 (codons 117 and 146) [9,10,11]. Therefore, an extensive analysis of RAS mutations is now necessary to select patients eligible to therapies targeting EGFR. Mutations in KRAS (exons 2, 3 and 4) have been reported in 40–45% of patients and NRAS (exons 2, 3 and 4) in 5–10% [12,13,14]. The BRAF V600E mutation is a prognostic factor of outcome and is observed in 4–18% of patients with colon cancer (this mutation is not used to guide anti-EGFR therapy) [13,14,15,16].
Guidelines from the European Society for medical Oncology (ESMO) recommend the genotyping of tumor tissue from primary or metastatic tissue both for KRAS exon 2 and non-exon 2 mutations [17]. A recent study reported that KRAS genotyping results were available within 15 days for 82% of tests in Europe, 51% in Latin America and 98% in Asia [18]. For patients with rapidly progressing disease, shorter turnaround times are needed and systems that perform automatically the time-consuming procedures could be of great value. In France, RAS testing circuit involves several actors: health centers for the management of patients, anatomic pathology laboratories for the preparation of tissue samples and national molecular genetics platforms for genotyping. A recent national French survey reported a median turnaround time for molecular tests (EGFR, RAS, BRAF) of 18 days [19].
In this pilot project we evaluated the turnaround time before and after the implementation of the fully automated multiplex Idylla™ KRAS Mutation Test and Idylla™ NRAS-BRAF Mutation Test in metastatic colorectal cancer.
Materials and Methods
This pilot project was performed in metropolitan France and overseas departments. The objective was to compare the turnaround time in 2017–2018 (with fully automated multiplex assay) compared to the 2016 results (previous methods).
The Idylla™ KRAS Mutation Test (Biocartis, Mechelen, Belgium) detects 21 KRAS mutations: 7 mutations in codons 12 and 13 (exon 2), 9 mutations in codons 59 and 61 (exon 3) and 5 mutations in codons 117 and 146 (exon 4). The Idylla™ NRAS-BRAF Mutation Test detects 23 mutations: 8 NRAS mutations in codons 12 and 13 (exon 2), 6 NRAS mutations in codons 59 and 61 (exon 3), 4 NRAS mutations in codons 117 and 146 (exon 4) and 5 BRAF mutations in codon 600 (exon 15).
The cartridges contain reagents necessary to perform sample preparation, real-time PCR amplification and detection starting from formalin-fixed paraffin-embedded tissue sections directly inserted in the cartridges. All operations for deparaffinization, DNA extraction and PCR are performed automatically in the single-use cartridge. Manufacturer specifications and previous studies reported that 2 h approximately were necessary to perform a mutation test using the Idylla™ cartridges [20].
Centers with a number of tests for patients with metastatic colorectal cancer > 100 yearly and a usual turnaround time ≥ 3 weeks for mutation detection were selected. Since the Idylla™ tests are not yet reimbursed by social security, the Idylla™ system was installed in centers that were included in the pilot project and Idylla™ cartridges were provided by Amgen free of charge.
A qualitative survey was conducted in participating pathologists and oncologists in order to evaluate the impact of this molecular testing platform in their daily clinical practice.
Results
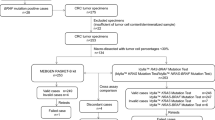
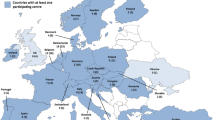
Twelve centers (one public and 11 privates) were selected and 10 of them transmitted results for 505 KRAS tests and 369 NRAS/BRAF tests. The average time from test prescription to reception of results was reduced from 25.8 days in 2016 to 4.5 days in 2017–2018 after implementation of the Idylla™ platform (Table 1).
Ten out the 12 participating pathologists and oncologists completed the survey questionnaire. The satisfaction rates of pathologists on the molecular test circuit increased from 60% (satisfied) to 100% (satisfied plus very satisfied) after the implementation of the project; 80% of pathologists considered that this project had a very positive impact on the organization of the RAS/BRAF testing in their laboratory; 100% of them judged satisfactory (satisfied plus very satisfied) the Idylla™ technology, as well as the RAS testing turnaround time after the implementation of the project.
Regarding oncologists, the satisfaction rates (i.e., very satisfied plus satisfied) on RAS testing turnaround time increased from only 30–100%. Oncologists highlighted that the long turnaround time impacted the therapeutic strategy for the patients (e.g., synchronous metastatic patients, symptomatic patients, patients who need cytoreduction) and led to a delay in the introduction of the targeted therapy (often introduced in the second chemotherapy cycle) or the administration of anti-VEGF regardless of the molecular status. The implementation of the project within the centers had, according to 100% of physicians, a positive impact on the patient’s care (80% very positive and 20% positive). They underlined that patients received more quickly the targeted therapy suited to their Idylla™ RAS status.
Discussion
Our results in real life show that the installation of an automated molecular diagnostic tool resulted in drastic reduction of the mean turnaround time from test prescription to reception of results by the oncologist who prescribed the test (Fig. 1). The specificity of this method and the comparisons with other routine methods have been reported in previous studies for metastatic colorectal cancer [21, 22]. The shortening of turnaround times with automated assay for RAS mutations installed in decentralized anatomic pathology laboratories meets the need for rapid and reliable tests for guiding clinical decisions and starting the most beneficial cancer therapies. A recent survey from the French National Cancer Institute (INCa) showed that the median turnaround time from test prescription to reception of results by the clinician for EGFR, RAS and BRAF molecular tests was 18 days: it was < 1 month for 86% of patients, 1–2 months for 12% and > 2 months for 2% [19]. This turnaround time included 11 days (median) for the management of the sample and the realization of the test itself by the molecular genetic platform. In a study performed in Australia including nine participating laboratories, the major factor that contributed to a long turnaround time was the time needed to retrieve archived tumor block, particularly when they were obtained from sources external to the testing site [23]. Thus, in almost 30% of cases more than 2 weeks elapsed before sample was received by the center that performed the test. Then a result was obtained within 2 weeks in 85% of cases. Overall, the median turnaround time (from ordering of the test to report of the results) was 17 days and in 20% of cases this time was more than 4 weeks for a KRAS test; in only 10% of cases, the results were available within one week.
According to the recommendations of the INCa, the turnaround time should be 2–3 weeks from the prescription date to the transmission of the results to the clinician; therefore, the results of the test should be transmitted with a maximum of 7–10 days after the reception of the sample [24].
For this project, we selected centers where the routine turnaround times exceeded three weeks: the mean turnaround time of the different centers varied from 21.0 to 36.8 days (Table 1). After the implementation of Idylla™ RAS testing in anatomic pathology laboratories, this turnaround time was strongly reduced with times varying from 1.9 to 15.7 days. Generally, mean values of turnaround time did not exceed one week; only one center had turnaround time near two weeks. Overall, these results indicate that a turnaround time below 2 weeks is an objective that can be easily achieved and in most cases this time could be one week.
Another survey of the INCa evidenced that a mutation test had not been performed in 4.8% of patients with colorectal cancer although they were at the metastatic stage [19]. These results raise the question of an equitable access to targeted therapies. One of the objectives of the French Cancer Plan for 2014–2019 was that all cancer patients in metropolitan France and overseas departments would benefit from access to genetic molecular tests regardless their localization or medical structure where they are managed [25]. The Idylla™ Mutation Assay meets these requirements since it is a fully-automated and one-step solution. Therefore, specialized infrastructure for molecular testing and highly qualified staff are not necessary. Frequently, laboratories do not perform in-house assays for molecular testing and send samples to external specialized centers where samples are often tested in batch in order to optimize costs. This step is avoided with the automatic Idylla™ system which is implemented in cancer centers where patients are managed.
In conclusion, this pilot project shows that the Idylla™ platform for testing KRAS and NRAS/BRAF mutations allows optimizing turnaround time from test prescription to reception of results.
References
Kuipers EJ, Grady WM, Lieberman D, Seufferlein T, Sung JJ, Boelens PG, van de Velde CJ, Watanabe T (2015) Colorectal cancer. Nat Rev Dis Primers 1:15065
Ferlay J, Steliarova-Foucher E, Lortet-Tieulent J, Rosso S, Coebergh JW, Comber H, Forman D, Bray F (2013) Cancer incidence and mortality patterns in Europe: estimates for 40 countries in 2012. Eur J Cancer 49:1374–1403
Hocking CM, Price TJ (2014) Panitumumab in the management of patients with KRAS wild-type metastatic colorectal cancer. Therap Adv Gastroenterol 7:20–37
Prenen H, Tejpar S, Van Cutsem E (2010) New strategies for treatment of KRAS mutant metastatic colorectal cancer. Clin Cancer Res 16:2921–2926
Siena S, Sartore-Bianchi A, Di Nicolantonio F, Balfour J, Bardelli A (2009) Biomarkers predicting clinical outcome of epidermal growth factor receptor-targeted therapy in metastatic colorectal cancer. J Natl Cancer Inst 101:1308–1324
Er TK, Chen CC, Bujanda L, Herreros-Villanueva M (2014) Current approaches for predicting a lack of response to anti-EGFR therapy in KRAS wild-type patients. Biomed Res Int 2014:591867
Amado RG, Wolf M, Peeters M, Van Cutsem E, Siena S, Freeman DJ, Juan T, Sikorski R, Suggs S, Radinsky R, Patterson SD, Chang DD (2008) Wild-type KRAS is required for panitumumab efficacy in patients with metastatic colorectal cancer. J Clin Oncol 26:1626–1634
Karapetis CS, Khambata-Ford S, Jonker DJ, O’Callaghan CJ, Tu D, Tebbutt NC, Simes RJ, Chalchal H, Shapiro JD, Robitaille S, Price TJ, Shepherd L, Au HJ, Langer C, Moore MJ, Zalcberg JR (2008) K-ras mutations and benefit from cetuximab in advanced colorectal cancer. N Engl J Med 359:1757–1765
Douillard JY, Oliner KS, Siena S, Tabernero J, Burkes R, Barugel M, Humblet Y, Bodoky G, Cunningham D, Jassem J, Rivera F, Kocakova I, Ruff P, Blasinska-Morawiec M, Smakal M, Canon JL, Rother M, Williams R, Rong A, Wiezorek J, Sidhu R, Patterson SD (2013) Panitumumab-FOLFOX4 treatment and RAS mutations in colorectal cancer. N Engl J Med 369:1023–1034
Hecht JR, Douillard JY, Schwartzberg L, Grothey A, Kopetz S, Rong A, Oliner KS, Sidhu R (2015) Extended RAS analysis for anti-epidermal growth factor therapy in patients with metastatic colorectal cancer. Cancer Treat Rev 41:653–659
Sorich MJ, Wiese MD, Rowland A, Kichenadasse G, McKinnon RA, Karapetis CS (2015) Extended RAS mutations and anti-EGFR monoclonal antibody survival benefit in metastatic colorectal cancer: a meta-analysis of randomized, controlled trials. Ann Oncol 26:13–21
Zenonos K, Kyprianou K (2013) RAS signaling pathways, mutations and their role in colorectal cancer. World J Gastrointest Oncol 5:97–101
Malumbres M, Barbacid M (2003) RAS oncogenes: the first 30 years. Nat Rev Cancer 3:459–465
Fearon ER (2011) Molecular genetics of colorectal cancer. Annu Rev Pathol 6:479–507
Rajagopalan H, Bardelli A, Lengauer C, Kinzler KW, Vogelstein B, Velculescu VE (2002) Tumorigenesis: RAF/RAS oncogenes and mismatch-repair status. Nature 418:934
Oikonomou E, Koustas E, Goulielmaki M, Pintzas A (2014) BRAF vs RAS oncogenes: are mutations of the same pathway equal? Differential signalling and therapeutic implications. Oncotarget 5:11752–11777
Van Cutsem E, Cervantes A, Nordlinger B, Arnold D, Group EGW (2014) Metastatic colorectal cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann Oncol 25(Suppl 3):iii1–i9
Ciardiello F, Tejpar S, Normanno N, Mercadante D, Teague T, Wohlschlegel B, Van Cutsem E (2011) Uptake of KRAS mutation testing in patients with metastatic colorectal cancer in Europe, Latin America and Asia. Target Oncol 6:133–145
Institut National du Cancer. Accès aux tests moléculaires EGFR, RAS et BRAF. Résultats d’une enquête dans 5 régions françaises. Janvier 2016
de Biase D, de Luca C, Gragnano G, Visani M, Bellevicine C, Malapelle U, Tallini G, Troncone G (2016) Fully automated PCR detection of KRAS mutations on pancreatic endoscopic ultrasound fine-needle aspirates. J Clin Pathol
Solassol J, Vendrell J, Markl B, Haas C, Bellosillo B, Montagut C, Smith M, O’Sullivan B, D’Haene N, Le Mercier M, Grauslund M, Melchior LC, Burt E, Cotter F, Stieber D, Schmitt FL, Motta V, Lauricella C, Colling R, Soilleux E, Fassan M, Mescoli C, Collin C, Pages JC, Sillekens P (2016) Multi-Center Evaluation of the Fully Automated PCR-Based Idylla KRAS Mutation Assay for Rapid KRAS Mutation Status Determination on Formalin-Fixed Paraffin-Embedded Tissue of Human Colorectal Cancer. PLoS One 11:e0163444
Johnston L, Power M, Sloan P, Long A, Silmon A, Chaffey B, Lisgo AJ, Little L, Vercauteren E, Steiniche T, Meyer T, Simpson J (2018) Clinical performance evaluation of the Idylla NRAS-BRAF mutation test on retrospectively collected formalin-fixed paraffin-embedded colorectal cancer tissue. J Clin Pathol 71:336–343
Scott RJ, Fox SB, Desai J, Grieu F, Amanuel B, Garrett K, Harraway J, Cheetham G, Pattle N, Haddad A, Byron K, Rudzki B, Waring P, Iacopetta B (2014) KRAS mutation testing of metastatic colorectal cancer in Australia: where are we at? Asia Pac J Clin Oncol 10:261–265
INCa (Bonnes pratiques pour la recherche à visée théranostique demutations somatiques dans les tumeurs solides. Août 2010) https://www.e-cancer.fr/content/download/63175/568709/file/OUTTHERANOS10.pdf
INCa. Plan Cancer 2014–2019 https://www.e-cancer.fr/Expertises-et-publications/Catalogue-des-publications/Plan-Cancer-2014-2019
Funding
Amgen France.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
Stéphane ROSSAT: No conflict of interest.
Hervé PERRIER : Amgen, Roche, Servier, Sanofi
Marine LEFEVRE: No conflict of interest
Christophe LOUVET: MSD, Roche, Halozyme, Servier, AstraZeneca
Nathalie LE BERRE: No conflict of interest
Jérôme CHAMOIS : Astellas, Ipsen, Janssen, Sanofi Aventis
Maryline DOREL: No conflict of interest
Daniel VAQUE: Amgen, Roche, Janssen
Angélique GUILLAUDEAU: No conflict of interest
Dominique GENET: Novartis, Pfizer, Roche
Evelyne MAILLET: No conflict of interest
Simon TRIBY: Amgen employee
Jean-Christophe SABOURIN: Amgen, Merck Serono, Pfizer, Bayer, BMS, MSD, AstraZeneca, Boehringer Ingelheim
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Rossat, S., Perrier, H., Lefevre, M. et al. Drastic Reduction of Turnaround Time After Implementation of a Fully Automated Assay for RAS-BRAF Mutations in Colorectal Cancer: A Pilot Prospective Study in Real-life Conditions. Pathol. Oncol. Res. 26, 2469–2473 (2020). https://doi.org/10.1007/s12253-020-00818-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12253-020-00818-y