Abstract
Background
The Biocartis Idylla™ platform is a fully automated, real-time PCR-based diagnostic system. The Idylla™ KRAS and NRAS-BRAF Mutation Tests have been developed for the qualitative detection of mutations in KRAS, NRAS and BRAF genes, facilitating the genomic profiling of patients with colorectal cancer. The aim of the present study was to evaluate clinical performances of these tests in Japan.
Methods
The RAS and BRAF mutation statuses of 253 formalin-fixed paraffin-embedded (FFPE) colorectal cancer tissues were analyzed using the Investigational Use Only Idylla™ KRAS Mutation Test and the Idylla™ NRAS-BRAF Mutation Test and an in vitro diagnostics (IVD) kit (MEBGEN RASKET™-B kit).
Results
The success rate for obtaining a valid mutational data without retest of the Idylla tests was 97.6% (247/253): 111 KRAS mutations (43.8%), 9 NRAS mutations (3.6%), and 36 BRAF V600E mutations (14.2%) were detected using the Idylla tests. Compared with the MEBGEN RASKET-B results, the positive concordance rate was 97.4%, the negative concordance rate was 95.7%, and the overall concordance rate was 95.3% (κ = 0.919, 95% CI 0.871–0.967). The average turnaround time to Idylla™ KRAS and NRAS-BRAF Mutation Test was 5.6 working days (range: 3–11 days).
Conclusion
This result demonstrates a high concordance between the Idylla™ KRAS and NRAS-BRAF Mutation Tests and an existing IVD kit. In this manner, the Idylla™ mutation tests were validated for the detection of clinically significant KRAS, NRAS, and BRAF mutations in FFPE samples from colorectal cancer patients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Colorectal carcinoma (CRC) is one of the most common cancers in the world and is a leading cause of cancer mortality in men and women [1]. The epidermal growth factor receptor (EGFR) and its downstream signaling pathways, including the RAS-RAF-MAPK and phosphatidylinositol 3-kinase (PI3K)-Akt pathways, play important roles in tumor growth in CRC [2]. Anti-EGFR antibodies, such as cetuximab and panitumumab, improve the prognosis of patients with metastatic CRC (mCRC) [3, 4]. However, the therapeutic efficacy of anti-EGFR therapy is limited to patients with wild-type (WT) RAS genotypes: mutations in exons 2, 3, and 4 of the KRAS, NRAS, and exons 15 of the BRAF genes confer resistance to these antibodies [3, 4]. The reduced efficacy of anti-EGFR therapy was initially thought to be the result of mutations within KRAS exon 2 (codons 12 and 13). Subsequently, through two large clinical trials (PRIME and CRYSTAL), additional mutations that conferred resistance to anti-EGFR therapy were discovered [4]. For example, the BRAF V600E mutation was detected in 4–18% of CRC cases and was found to be responsible for a reduced response to EGFR inhibitors and a worse prognosis [5].
Several methods for the detection of KRAS, NRAS, and BRAF mutations in formalin-fixed paraffin-embedded (FFPE) tissues have been reported. These mutation kits have been marketed for colorectal cancer patients. Routine genotyping of DNA extracted from FFPE samples is performed using laboratory-based assays or in vitro diagnostic medical device (IVD) kits. Routine testing methods include Sanger sequencing, pyrosequencing, next-generation sequencing (NGS) [6], immunohistochemistry, and real-time PCR-based testing. Each of these kits has its own assay conditions in terms of sensitivity, specificity, cost, turnaround time, level of automation and multiplexing, special equipment and skilled staff. The turnaround time is a particularly important parameter for patients with rapidly progressing metastases. The turnaround time of the Idylla platform is reported to be shortened to an average of 4.5 working days [7].
The Idylla™ system (Biocartis, Mechelen, Belgium) is a fully automated, real-time PCR-based molecular diagnostic system [8,9,10,11,12,13,14] and is an example of an in vitro diagnostic medical device (IVD) that can be used for the qualitative detection of mutations. The Idylla™ KRAS and NRAS-BRAF Mutation Tests qualitatively detect mutations in KRAS codons 12, 13, 59, 61, 117, and 146; NRAS codons 12, 13, 59, 61, 117, and 146; and BRAF codon 600 in FFPE human malignant CRC tissue. When performed on the Idylla™ platform, Idylla™ mutation tests are automated sample-to-results solutions that integrate the analytical process into a single cartridge, eliminating the need for time-consuming pretreatment processes, including the steps for FFPE processing. To date, the CE-IVD Idylla™ KRAS Mutation Test and the Idylla™ NRAS-BRAF Mutation Test have been developed to detect KRAS, NRAS, and BRAF mutations in colorectal cancer patients. However, the Idylla™ system has not yet been approved for use in Japan, and a formal clinical performance study conducted in Japan is needed. In the present study, we compared the clinical performances of two Idylla™ tests (the Idylla™ KRAS Mutation Test and the Idylla™ NRAS-BRAF Mutation Test) used in combination with an IVD kit (MEBGEN RASKET™-B kit; Medical & Biological Laboratories Co., Tokyo, Japan) [15, 16] using FFPE tissue samples from colorectal cancer patients.
Materials and methods
Samples and study design
A series of 275 archived FFPE tissues from 275 Japanese patients with colorectal cancer diagnosed at the Kindai University Hospital and Shiga General Hospital was obtained. All cases were staged based on the criteria from the Japanese Classification of Colorectal, Appendiceal, and Anal Carcinoma [17]. And were selected by searching through archival material for the years 2017–2021. The 275 cases included 28 BRAF V600E positive cases obtained by prescreening 250 cases prior to the study. All the patients enrolled in the study had provided informed consent for the use of their resected tissue samples. This study was approved by the ethical committees of the Kindai University Faculty of Medicine and Shiga General Hospital (Authorization Number: R02-312).
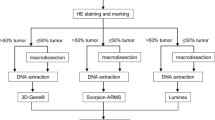
The study design is summarized in Fig. 1. FFPE specimens from 275 colorectal cancer patients were sliced into 5 μm thin slices. Twenty-two cases were excluded from the study, including 21 cases with a tumor percentage of less than 30% based on hematoxylin and eosin (H&E) staining and one demineralized specimen. A total of 253 cases were included in the study. The tumors areas were marked by pathologists, 134 of 253 specimens were manually dissected, and only specimens with tumor percentages ≥ 30% were subjected to the Idylla™ KRAS and NRAS-BRAF Mutation Tests and the MEBGEN RASKET™-B kit.
Procedure for enrolling patient samples in the study. The formalin-fixed paraffin-embedded (FFPE) sample preparation algorithm used prior to Idylla™ KRAS and NRAS-BRAF mutation testing is shown as a pre-analytical process. Histologically confirmed, retrospectively collected FFPE colorectal cancer tissue samples were identified, and tissue Sects. 5 µm thick were sampled as close as possible to the sections used for reference testing. Prior to the analysis, the tumor contents and area were determined using an H&E-stained slide by a consultant histopathologist at both sites. If required, macro-dissections were performed to ensure a tumor nuclei content of ≥ 30% in each sample used for analysis. Mutations in KRAS, NRAS, and BRAF mutations were analyzed using the Idylla™ KRAS Mutation Test and the Idylla™ NRAS-BRAF Mutation Test (maximum three slide each). DNA was extracted from individual slides, and the mutations were analyzed using the MEBGEN RASKET™-B kit. Concordance between the Idylla results and the MEBGEN RASKET-B results was then determined. Discordant cases were analyzed using amplicon deep sequencing
Idylla™ KRAS and NRAS-BRAF mutation tests (Idylla™)
The Idylla™ KRAS Mutation Test is a single-use cartridge-based test designed for the qualitative detection of 21 different mutations. In this study, the Idylla IUO assay was used. The mutations included those in KRAS codons 12, 13, 59, 61, 117 and 146. The Idylla™ NRAS-BRAF Mutation Test utilizes PCR reactions to amplify 15 mutations in codons 12, 13, 59, 61, 117, 146 of the NRAS oncogene and codon 600 of the BRAF oncogene. Table 1 lists the mutations that are detected by the test. Detailed procedures were summarized in Supplementary Method as described previously [18]. The performance evaluation of the Idylla™ KRAS and NRAS-BRAF Mutation Test was performed in the Department of Genome Biology, Kindai University Faculty of Medicine.
MEBGEN RASKET™-B kit
Genomic DNA was extracted from FFPE tumor samples using a standard procedure, and 10–20 ng/μl concentrations of DNA were subjected to PCR reactions. In total, 49 KRAS, NRAS, and BRAF mutations were analyzed using the MEBGEN RASKET™-B kit [15, 16]; the PCR reaction reverse sequence-specific oligonucleotide method was used for all the samples. Mutations were determined using multiplex PCR and the xMAP® (Luminex®) technology. The mutations detected by the MEBGEN RASKET™-B kit are listed in Supplementary Method. This procedure was performed in the laboratories of LSI-Medience Co. (Tokyo, Japan) and SRL Inc. (Tokyo, Japan). Finally, all the data were collected and compared.
Sequencing analysis
DNA were purified with the use of an Allprep DNA/RNA FFPE Kit (Qiagen, Valencia, CA, USA) according to the manufacturer’s instructions. Amplicon sequencing were performed using the Ion AmpliSeq Colon and Lung Cancer Panel (Thermo Fisher Scientific, Wilmington, DE, USA). Detailed procedure was described in Supplementary Method.
Turnaround time
The turnaround time was defined as the period from the time of sample registration until reporting by the Idylla™ KRAS Mutation Test or the Idylla™ NRAS-BRAF Mutation Test. Time is shown in working days assuming a Monday to Friday working week.
Statistical analysis
Kappa statistics were used to compare the results of the Idylla™ KRAS and NRAS-BRAF Mutation Tests with the results of the MEBGEN RASKET™-B kit. Categorical variables were compared using the Fisher exact test. Continuous variables were compared between groups with the Mann–Whitney U test. All the statistical analyses were performed using JMP software, version 14.2 (SAS Institute Japan, Tokyo, Japan), and Prism software, version 8.4 (GraphPad Software, San Diego, CA, USA). A p value of < 0.05 was considered statistically significant.
Results
Study population
First, FFPE tissue samples from all 275 patients with colorectal cancer were retrieved for use in this study. All the samples were histologically confirmed as malignant CRC by pathologists. The clinical and pathological characteristics are summarized in Table 2. There were 145 (52. 7%) males and 130 females (47.3%) with a median age of 72 years (range: 32–92). The relative frequencies of histological type are 86.9% for intestinal type and 13.1% for the diffuse type. Tumor stage was 12.7% for stage 1, 34.2% for stage II, 45.5% for stage III, and 7.6% for stage IV. Among the 275 clinical specimens, 253 samples were subjected to both assays (Fig. 1). These include 28 FFPE samples with known BRAF mutations obtained by prescreening. The reasons for the exclusion of the 22 samples were the absence of tumor content or inappropriate tumor proportions (21 cases) as determined using hematoxylin and eosin (H&E) staining; the remaining specimen was judged to be unsuitable for inclusion by the pathologists because of demineralization.
Mutations detected using the Idylla™ KRAS and NRAS-BRAF mutation tests
The success rate for obtaining a valid mutational data without retest of the Idylla™ KRAS and NRAS-BRAF Mutation Tests was 97.6% (247/253). Among the 253 clinical specimens that were examined, 111 KRAS mutations (43.9%) and 9 NRAS (3.6%), and 36 BRAF mutations (14.2%) mutations were detected using the Idylla™ KRAS and NRAS-BRAF Mutation Tests (Fig. 2; details in Table S1). The G12D, G12V, G13D, and G12C KRAS mutations were the most frequently detected (40/111, 36.0%; 25/111, 22.5%, 13/111, 11.7%; and 9/111, 8.1%, respectively). BRAF mutations were detected in 36 specimens (including 28 BRAF mutation positive cases out of 208 prescreened cases) using the Idylla™ NRAS-BRAF Mutation Test. Therefore, BRAF frequency under spontaneous conditions is 17.8% (8/45).
KRAS mutation was associated with female sex (p = 0.0002). BRAF mutation positive cases were older age (p = 0.0198) and showed a higher frequency of poorly differentiated cancers (p < 0.0001) [19]. No differences in age, sex, histological type, or disease stage were observed between samples with and those without a NRAS mutation (Table S2).
Method correlation agreement analysis
The Idylla™ results for the 253 cases that were examined were compared with the results of the MEBGEN RASKET™-B kit. The number of invalid tests for Idylla™ KRAS Mutation Test, Idylla™ NRAS-BRAF Mutation Test, and MEBGEN RASKET™-B kit were 7, 6, and 4, respectively. Discordances between the results of the Idylla™ KRAS and NRAS-BRAF Mutation Tests and those of the MEBGEN RASKET™-B kit were observed for four specimens. The positive agreement rate was 97.4% (151/155), the negative agreement rate was 95.7% (90/94), and the overall agreement rate was 95.3% (κ = 0.919, 95% CI 0.871–0.967) (Table 3).
Retests
A retest of 17 specimens (13 invalid and 4 discordant specimens) was performed using the Idylla™ KRAS and NRAS-BRAF Mutation Tests. Five of the eight specimens that were retested using the Idylla™ KRAS Mutation Test remained invalid. The result for one case changed from invalid to positive, one case changed from invalid to negative, and one case changed from positive to negative for KRAS mutation. Meanwhile, 2 of the 9 specimens that were retested using the Idylla™ NRAS-BRAF Mutation Test remained invalid. The results for two cases changed from positive to negative for NRAS mutation. Four invalid cases were validated (positive or negative) by retests. The four samples retested using the MEBGEN RASKET™-B kit showed that one case remained invalid, and three cases changed from invalid to positive.
Sequencing analysis for cases with discrepancies
Amplicon sequencing was performed for the four cases with discrepant results between the Idylla™ KRAS and NRAS-BRAF Mutation Tests and the MEBGEN RASKET™-B kit. The gene mutation results detected by sequencing were consistent with the results detected using the MEBGEN RASKET™-B kit. No mutation was detected by the deep sequencing in three cases (K-N-001-NRAS A146T, K-N-151-NRAS A59T, and K-N-221-KRAS Q61H), in which mutations had been detected using the Idylla™ KRAS and NRAS-BRAF Mutation Tests (Table S3). Sequencing analysis identified NRAS Q61H mutations with a variant allele frequency of 22.6% in one case (K-N-214), in which no mutation had been detected using the Idylla™ KRAS and NRAS-BRAF Mutation Tests. This discrepancy is due to the difference in detectable sites between Idylla™ and RASKET. Idylla™ was only designed by NRAS Q61H (c.183A>C), whereas RASKET is designed to detect NRAS Q61H (c.183A>C and c.183A>T). K-N-214 was not detected by Idylla™ because it harbored the NRAS Q61H mutation of c.183A>T nucleotide base substitution.
Turnaround time
The TAT is the time interval between the specimens received in the laboratory to the time of reports of the assay. The average TAT of the Idylla™ KRAS and NRAS-BRAF Mutation Test was 5.61 ± 1.99 working days (range 3–11 days). The TAT of the MEBGEN RASKET™-B kit was not determined because the testing was outsourced.
Discussion
In this study, the success rate of the Idylla tests was 97.6%. This rate was considered a clinically acceptable level. However, the number of tissue specimens submitted was 275, and 22 specimens were excluded at the pre-analytical stage. Appropriate specimen preparation at the pre-analytical stage was considered important for assay success. Highly concordant results were obtained between the Idylla™ KRAS and NRAS-BRAF Mutation Tests and the MEBGEN RASKET™-B kit. The Idylla tests were shown to be reliable with a high positive concordance rate and negative concordance rate relative to the results of the MEBGEN RASKET™-B kit. Deep sequencing of four discordant cases detected no RAS-BRAF mutations in three cases. These results suggest that the mutations detected using the Idylla™ KRAS and NRAS-BRAF Mutation Tests were likely to be false-positive. This number of false-positive cases was considered clinically acceptable.
KRAS and NRAS mutations are known to be the most frequent actionable mutations observed in colorectal cancer. In the present study, the Idylla tests detected 111 KRAS mutations (43.9%) and 9 NRAS mutations (3.6%) in 253 specimens. Among the KRAS mutations, mutations in codons 12 and 13 of KRAS were detected at a high frequency. The frequencies of KRAS and NRAS mutations were similar to those in previous reports [20, 21]. We also detected BRAF V600E mutations. Mutations in the BRAF gene occur in approximately 12% of mCRC patients, with reported frequencies ranging from as low as 5% to as high as 21% [21,22,23,24]. The majority of BRAF mutations are V600E substitutions [25]. In our sample cohort, BRAF V600E mutations were detected in 36 samples, including 28 samples with known mutations as a result of prescreening. Therefore, eight additional BRAF V600E mutations were detected in 45 naïve specimens (17.8%). The design of the present study did not allow for an accurate estimation of the BRAF mutation frequency. Although the Idylla™ tests were not designed to detect other genotypes of BRAF mutations, the mutation frequencies for such genotypes are very low, and the present performance was considered to be clinically acceptable. When differences among histological types were examined, the BRAF V600E mutation was detected at a significantly higher frequency in poorly differentiated adenocarcinomas, compared with other histological types (p < 0.0001). These results were consistent with those of a previous paper reported by Yokota et al. They also reported that BRAF V600E mutations were frequently observed in mucinous adenocarcinoma of the colon in Japanese patients [19], but no BRAF mutations were observed in the mucinous types in our sample cohort. Further study might be necessary.
The TAT of the Idylla tests was 5.6 days, and no difference was seen between the Idylla™ KRAS Mutation Test and the Idylla™ NRAS-BRAF Mutation Test. Therefore, no time loss occurred when both kits were used together. The TAT of 5.6 days was caused by the working period for the preparation of the tumor tissue at the pre-analytical stage, including the sectioning of the samples and the confirmation of the tumor contents using hematoxylin and eosin staining. After sample preparation, there were hands-on in required for the Idylla tests were less than 2 min, excluding the nucleic acid extraction procedure; the time required from sample application until the end result was approximately 2 h. Therefore, the Idylla™ platform is expected to shorten the TAT in real-world clinical practice. In addition, since only one slide is necessary per Idylla test, it is clinically advantageous that only two slides for two tests are required.
In this study, we examined the clinical performance of the Idylla™ platform and found that its performance was comparable to that described in previous overseas reports [7, 26, 27]. This formal clinical performance study supports the IVD approval of this test in Japan. Currently, an Idylla™ system for circulating tumor DNA in blood samples have already been developed for BRAF, KRAS, NRAS, and EGFR [27, 28]. Liquid biopsy tests are also available using the same assay system as tumor tissue samples. To promote liquid biopsy testing in Japan, formal clinical performance tests are warranted. The usefulness of the Idylla™ system is good news for Japanese colorectal cancer patients, but the system might also be applicable to other types of cancer, such as non-small cell lung cancer.
In conclusion, the Idylla™ KRAS and NRAS-BRAF Mutation Tests for colorectal patients are reliable, simple, and rapid sample-to-result solutions for the detection of clinically important KRAS, NRAS, and BRAF mutations without the need for molecular expertise or infrastructure.
Abbreviations
- 95% CI:
-
95% Confidence interval
- CRC:
-
Colorectal carcinoma
- EGFR:
-
Epidermal growth factor receptor
- FFPE:
-
Formalin-fixed paraffin-embedded
- H&E:
-
Hematoxylin and eosin
- IUO:
-
Investigational use only
- IVD:
-
In vitro diagnostics
- TAT:
-
Turnaround time
- WT:
-
Wild-type
References
Torre LA, Bray F, Siegel RL et al (2015) Global cancer statistics. CA Cancer J Clin 65(2):87–108. https://doi.org/10.3322/caac.21262
Zenonos K, Kyprianou K (2013) RAS signaling pathways, mutations and their role in colorectal cancer. World J Gastrointest Oncol 5(5):97–101. https://doi.org/10.4251/wjgo.v5.i5.97
Bokemeyer C, Kohne CH, Ciardiello F et al (2015) FOLFOX4 plus cetuximab treatment and RAS mutations in colorectal cancer. Eur J Cancer 51(10):1243–1252. https://doi.org/10.1016/j.ejca.2015.04.007
Douillard JY, Siena S, Cassidy J et al (2014) Final results from PRIME: randomized phase III study of panitumumab with FOLFOX4 for first-line treatment of metastatic colorectal cancer. Ann Oncol 25(7):1346–1355. https://doi.org/10.1093/annonc/mdu141
Pietrantonio F, Petrelli F, Coinu A et al (2015) Predictive role of BRAF mutations in patients with advanced colorectal cancer receiving cetuximab and panitumumab: a meta-analysis. Eur J Cancer 51(5):587–594. https://doi.org/10.1016/j.ejca.2015.01.054
Sakai K, Tsurutani J, Yamanaka T et al (2015) Extended RAS and BRAF mutation analysis using next-generation sequencing. PLoS ONE 10(5):e0121891. https://doi.org/10.1371/journal.pone.0121891
Rossat S, Perrier H, Lefevre M et al (2020) Drastic reduction of turnaround time after implementation of a fully automated assay for RAS-BRAF mutations in colorectal cancer: a pilot prospective study in real-life conditions. Pathol Oncol Res 26(4):2469–2473. https://doi.org/10.1007/s12253-020-00818-y
Colling R, Wang LM, Soilleux E (2016) Automated PCR detection of BRAF mutations in colorectal adenocarcinoma: a diagnostic test accuracy study. J Clin Pathol 69(5):398–402. https://doi.org/10.1136/jclinpath-2015-203345
de Biase D, de Luca C, Gragnano G et al (2016) Fully automated PCR detection of KRAS mutations on pancreatic endoscopic ultrasound fine-needle aspirates. J Clin Pathol. https://doi.org/10.1136/jclinpath-2016-203696
De Luca C, Gragnano G, Pisapia P et al (2017) EGFR mutation detection on lung cancer cytological specimens by the novel fully automated PCR-based Idylla EGFR Mutation Assay. J Clin Pathol 70(4):295–300. https://doi.org/10.1136/jclinpath-2016-203989
Janku F, Claes B, Huang HJ et al (2015) BRAF mutation testing with a rapid, fully integrated molecular diagnostics system. Oncotarget 6(29):26886–26894. https://doi.org/10.18632/oncotarget.4723
Melchior L, Grauslund M, Bellosillo B et al (2015) Multi-center evaluation of the novel fully-automated PCR-based Idylla BRAF mutation test on formalin-fixed paraffin-embedded tissue of malignant melanoma. Exp Mol Pathol 99(3):485–491. https://doi.org/10.1016/j.yexmp.2015.09.004
Schiefer AI, Parlow L, Gabler L et al (2016) Multicenter evaluation of a novel automated rapid detection system of BRAF status in formalin-fixed, paraffin-embedded tissues. J Mol Diagn 18(3):370–377. https://doi.org/10.1016/j.jmoldx.2015.12.005
Yeo MK, Jung MK, Lee SY et al (2017) The usefulness of a novel fully automated PCR-based Idylla test for detection of the BRAF V600E mutation in thyroid tissue: comparison with PNA-clamping PCR, real-time PCR and pyrosequencing. J Clin Pathol 70(3):260–265. https://doi.org/10.1136/jclinpath-2016-204025
Taniguchi H, Okamoto W, Muro K et al (2018) Clinical validation of newly developed multiplex kit using luminex xMAP technology for detecting simultaneous RAS and BRAF mutations in colorectal cancer: results of the RASKET-B study. Neoplasia 20(12):1219–1226. https://doi.org/10.1016/j.neo.2018.10.004
Yoshino T, Muro K, Yamaguchi K et al (2015) Clinical validation of a multiplex kit for RAS mutations in colorectal cancer: results of the RASKET (RAS key testing) prospective multicenter study. EBioMedicine 2(4):317–323. https://doi.org/10.1016/j.ebiom.2015.02.007
Japenese society for cancer of the colon and rectum (2019) Japanese classification of colorectal, appendiceal, and anal carcinoma: the 3d English edition [secondary publication]. J Anus Rectum Colon 3(4):175–195. https://doi.org/10.23922/jarc.2019-018
Johnston L, Power M, Sloan P et al (2018) Clinical performance evaluation of the Idylla NRAS-BRAF mutation test on retrospectively collected formalin-fixed paraffin-embedded colorectal cancer tissue. J Clin Pathol 71(4):336–343. https://doi.org/10.1136/jclinpath-2017-204629
Yokota T, Ura T, Shibata N et al (2011) BRAF mutation is a powerful prognostic factor in advanced and recurrent colorectal cancer. Br J Cancer 104(5):856–862. https://doi.org/10.1038/bjc.2011.19
Kawazoe A, Shitara K, Fukuoka S et al (2015) A retrospective observational study of clinicopathological features of KRAS, NRAS, BRAF and PIK3CA mutations in Japanese patients with metastatic colorectal cancer. BMC Cancer 15:258. https://doi.org/10.1186/s12885-015-1276-z
Peeters M, Kafatos G, Taylor A et al (2015) Prevalence of RAS mutations and individual variation patterns among patients with metastatic colorectal cancer: a pooled analysis of randomised controlled trials. Eur J Cancer 51(13):1704–1713. https://doi.org/10.1016/j.ejca.2015.05.017
Barras D, Missiaglia E, Wirapati P et al (2017) BRAF V600E mutant colorectal cancer subtypes based on gene expression. Clin Cancer Res 23(1):104–115. https://doi.org/10.1158/1078-0432.CCR-16-0140
Davies H, Bignell GR, Cox C et al (2002) Mutations of the BRAF gene in human cancer. Nature 417(6892):949–954. https://doi.org/10.1038/nature00766
Yaeger R, Chatila WK, Lipsyc MD et al (2018) Clinical sequencing defines the genomic landscape of metastatic colorectal cancer. Cancer Cell 33(1):125–136. https://doi.org/10.1016/j.ccell.2017.12.004
Kayhanian H, Goode E, Sclafani F et al (2018) Treatment and survival outcome of BRAF-mutated metastatic colorectal cancer: a retrospective matched case-control study. Clin Colorectal Cancer 17(1):e69–e76. https://doi.org/10.1016/j.clcc.2017.10.006
Haiduk T, Brockmann M, Tillmann RL et al (2021) Comparison of biocartis IDYLLA cartridge assay with Qiagen genereader NGS for detection of targetable mutations in EGFR, KRAS/NRAS, and BRAF genes. Exp Mol Pathol 120:104634. https://doi.org/10.1016/j.yexmp.2021.104634
Holm M, Andersson E, Osterlund E et al (2020) Detection of KRAS mutations in liquid biopsies from metastatic colorectal cancer patients using droplet digital PCR, Idylla, and next generation sequencing. PLoS ONE 15(11):e0239819. https://doi.org/10.1371/journal.pone.0239819
Vessies DCL, Greuter MJE, van Rooijen KL et al (2020) Performance of four platforms for KRAS mutation detection in plasma cell-free DNA: ddPCR, Idylla, COBAS z480 and beaming. Sci Rep 10(1):8122. https://doi.org/10.1038/s41598-020-64822-7
Acknowledgements
This study was sponsored by Nichirei Biosciences Inc. This means that the publication and performance of this study was conducted by the investigators and is not intended to promote the Idylla™ mutation testing platform. We thank Ms. Tomoko Hashimoto (Department of Surgery, Kindai University Faculty of Medicine) for technical assistance provided during the study.
Funding
This study is funded from Nichirei Biosciences Inc. (Tokyo, Japan) for TS, HY, KN, and JK.
Author information
Authors and Affiliations
Contributions
JK, KN, YM, and KS contributed conceptualization. YM, KS, MY, TW TC, TS, YI, and HY contributed investigations. HK and LZ wrote the paper and performed the literature review. YT, MI, TH, and ST contributed to the conception and design of the manuscript and critically revised the manuscript. All authors read and approved the final version of the manuscript. YM and KS contributed data curation. MD contributed formal analysis, methodology, and writing. YM, KS, and KN wrote the original draft. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
Takao Satou, Hidekazu Yamamoto, Kazuto Nishio, and Junichiro Kawamura received funding from Nichirei Biosciences Inc. (Tokyo, Japan) for this study. No other financial support was provided.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Makutani, Y., Sakai, K., Yamada, M. et al. Performance of Idylla™ RAS-BRAF mutation test for formalin-fixed paraffin-embedded tissues of colorectal cancer. Int J Clin Oncol 27, 1180–1187 (2022). https://doi.org/10.1007/s10147-022-02167-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10147-022-02167-z