Abstract
Efficient bacterial strain was isolated from the dye contaminated area and identified as Bacillus stratosphericus SCA1007 based on 16S rRNA gene sequence (GenBank under accession number KY992944). This isolate was selected based on its potential to efficiently decolorize reactive orange 16 dye which is extensively used in textile industries. Various culture conditions like dye concentration, temperature, pH, salinity, and additional nitrogen source were optimized in the present study. The optimal conditions for decolorization of reactive orange 16 was found to be: dye concentration 150 mg/L, pH 7, temperature 35 °C, and yeast extract as nitrogen source. The isolate was also resistant to 4% saline culture condition. Decolorization and degradation of dye were confirmed through UV–visible spectroscopy, Fourier transform infrared (FTIR) and liquid chromatography-mass spectrometry analysis (LC–MS). Toxicity studies were performed on Escherichia coli and Vigna radiata to confirm the non-toxic nature of the degraded metabolites. This is the first study demonstrating complete decolorization of reactive orange 16 dye by Bacillus stratosphericus SCA1007 at high salinity within 10 h of incubation under optimized conditions.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The growth of polluting industries is causing tremendous stress on the environment and the ecological system. Textile industry is one of the most polluting industries (Pritchard and Costa 1991). Textile processing industry uses a large amount of chemicals dissolved in water which ultimately makes its way into the environment leading to manifold increase in land and water pollution. Synthetic dyes are one such class of chemicals. Apart from textile industry, these are extensively utilized in leather tanning, paper industries, agriculture research, food technologies, photo electrochemical cells, light harvesting arrays, and hair colors (Popli and Patel 2015). Textile industry alone produces 2,80,000 tonnes of dyes which are released every year all around the world (Elango et al. 2017). Once the processing of product is done, discharge of colored effluent is mixed with water streams. This mixing of highly colored effluent leads to water pollution (Ojekunle and Lateef 2017). Appearance of color in the water is the main visible indication of water being polluted (Ali 2010). The wastewater enters agricultural land, which has serious implications for living organisms. Moreover, synthetic dye reduces light penetration through water, thereby reducing photosynthesis carried out by hydrophytes, thus disturbing the food chain of aquatic eco system (Yaseen and Scholz 2018).
Dyes containing azo group are the most versatile class of man-made dyes which are applied widely because of their ease of use, varied color and cost efficiency (Tomczak and Gorecki 2012). They are highly mutagenic and carcinogenic in nature (Pillai 2017). The azo bonds are resistant to breakdown, resulting in accumulation in the environment. Therefore, it is essential to develop efficient and economical solution for textile effluents (Park et al. 2007). Among the azo dyes, monoazo dye reactive orange 16 is an extensively used textile dye (Kapdan and Oztekin 2003). It is brilliantly colored and easily dissolves in water which makes it even more challenging for conventional wastewater treatment methods to remediate dye.
Numerous physicochemical techniques are applied for the remediation of dye effluents, which include chemical oxidation, adsorption, coagulation, precipitation, electrolysis, bleaching, membrane filtration, and ozonation (Geetha and Velmani 2015). The major disadvantages of all these methods are that they are expensive, less efficient and lead to production of sludge, which creates further pollution (Sharma and Roy 2015). Therefore, biological methods are considered an attractive solution for mineralization of dyes (Bhatia et al. 2017), as these are environmental friendly and cost-effective (Megha et al. 2015). Generally, dye decolorization is the result of two processes either by adsorption or degradation. During adsorption, parent structure of dye remains unchanged as dyes are being adsorbed onto microbial cell surfaces, whereas, in case of degradation, naive structure of the dye is changed or broken down to smaller fragments or converted to CO2, H2O and salts of inorganic origin (Zhou and Zimmermann 1993).
Several taxonomic groups of microbes like bacteria, fungi, and yeast algae can decolorize and sometimes mineralize reactive orange 16. These are Pseudomonas sp., Nocardiopsis alba, Micrococcus luteus, Mycobacterium sp., Bacillus flexus, Fundalia trogii, and Irpex lacteus (Kalyani et al. 2009; Ali 2010; Shobana and Hamgam 2012; Saha and Rao 2019). It is found that bacterial decolorization is normally faster as compared to fungal decolorization. The present study aims to isolate and characterize a bacterium which is able to decolorize and degrade reactive orange 16.
Material and methods
Chemicals
The chemicals used in the study were of analytical grade. Reactive orange 16 was obtained from Sigma-Aldrich, Bangalore, India. Mineral salt medium (MSM) [in g/L: potassium dihydrogen phosphate 1.0, sodium chloride 1.0, magnesium sulfate heptahydrate 0.5, calcium chloride 0.1, sodium hydrogenphosphate 1.0, yeast extract 4.0, agar (solid media)] and nutrient media were retrieved from HiMedia, Mumbai, India.
Isolation, screening, and identification of dye decolorizing bacteria
Soil samples were sourced from areas of Jharkhand (23.3441° N, 85.3096° E) and Bihar (25.5941° N, 85.1376° E) where dyeing is prevalent and were serially diluted in 0.9% (w/v) saline solution, following which, sample was spread over nutrient agar plates (Habib et al. 2018). Morphologically distinct colonies were selected and purified on nutrient agar slant. Pure colonies were kept at 4 °C for further use.
Screening was carried out in mineral salt medium agar plates containing reactive orange 16 dye to isolate bacteria with decolorizing ability for reactive orange 16 dye (25 mg/L). Bacterial colonies, showing clear zones, were identified for further studies. Initial experiments were performed to study the dye decolorization efficiency of each isolated bacteria, using flask (250 mL) containing 100 mL mineral salt medium along with dye (150 mg/L). Freshly grown bacterium was used to inoculate the media. The bacterial isolates which showed the best potential for decolorization of reactive orange 16 were screened to be used for further investigations (Zin et al. 2020).
Biochemical, morphological, and molecular characterizations were performed to identify the selected bacterium. Gram reaction, motility, shape, and endospore staining were the basis of morphological characterization (Breed et al. 1957). Following biochemical tests were done, namely starch hydrolysis, gelatinase, lipase, carbohydrate fermentation, catalase, oxidase, indole test, methyl red, voges-proskauer, nitrate reduction, hydrogen sulfide production, and citrate utilization. Genus was identified in accordance to Bergey’s Manual of Determinative Bacteriology (Breed et al. 1957). Bacterial genomic DNA was isolated from the suspended pellet using Zymo Research Fungal/Bacterial DNA MicroPrep™ following the standard protocol prescribed by the manufacturer. DNA samples were subjected to PCR amplification of 16S rRNA gene using the universal primers pA (5′-AGAGTTTGATCCTGGCTCAG-3′) and pH (5′-AAGGAGGTGATCCAGCCGCA-3′). The PCR amplification conditions were carried as described by Yadav et al. (2016). 16S rRNA gene sequencing (Xcelris Labs, Ahmedabad, India) was to identify species of selected bacterium isolate. Consensus sequences were aligned using ClustalW. Basic local alignment search tool (BLAST) of the National Center for Biotechnology Information (NCBI) was used to study the homology of sequences. Neighbor joining method was used to construct phylogenetic tree by the aid of Molecular Evolutionary Genetics Analysis (MEGA) software version 6.06. Under accession numbers KY992944, 16S rRNA gene sequences of isolate SCA1007 were submitted to GenBank. Isolate SCA1007 was deposited at culture collection facility of ICAR-National Bureau of Agriculturally Important Microorganisms (NBAIM), Mau Uttar Pradesh, India.
Decolorization assay
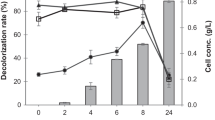
Decolorization was studied in Erlenmeyer flasks (250 mL) having MSM (100 mL) amended with reactive orange 16 dye. Four percent of freshly grown Bacillus stratosphericus SCA1007 was inoculated in each flask. Absorbance of cells was recorded at λ = 600. Inoculum size was maintained (1.50 × 106 CFU/mL) and was left for incubation at 35 °C for 24 h. Reactive orange 16 dye was examined for absorbance (A) maximum (λmax) (493 nm) using spectrophotometer UV-1800 Shimadzu. Two milliliters of reaction medium were removed and centrifuged at consistent intervals of 0 h, 24 h, 48 h, and 72 h respectively. Absorbance of supernatant was recorded at 493 nm. Medium without dye and inoculum was used as blank. All experiments were carried out in triplicate and standard deviation was calculated. The bacterial isolates which showed maximum decolorizing ability were used for subsequent investigations (Karim et al. 2018).
The decolorization capacity showed by different isolates was measured in terms of percentage decolorization (Dos et al. 2007) and calculated as follows:
where,
A = Absorbance of media before incubation
B = Absorbance of media after decolorization
Optimization of culture conditions for efficient decolorization
One variable at a time method was used to check the impact of different factors on decolorization of reactive orange 16 by Bacillus stratosphericus SCA1007 was studied. Experiments were performed at different concentration of dye ranging from (25–400 mg/L), pH (4–11), temperature (5–55 °C), NaCl (1–7%), and shaking/static. Buffers like sodium hydroxide (1 mol/L) and hydrochloric acid (1 mol/L) was used to maintain lower and higher pH. Data represent are means of three replicates.
Degradation analysis
Degraded metabolites obtained after 10 h of incubation was extracted by mixing the product with equal volume of ethyl acetate. The extracted product was dried over anhydrous sodium sulfate and evaporated to dryness in a rotary vacuum evaporator. Fourier transform infrared (FTIR) analysis of the extracted product was done using PerkinElmer 783 spectrophotometer in the mid IR region of 500–4500 cm−1 (Gao et al. 2020).
Liquid chromatography-mass spectrometry (LC–MS/MS) analysis
Monitoring of UV–vis spectra and FTIR gave information about change in the structures of parent dyes. To identify dye metabolites, analysis was performed using LC–MS/MS. Extracted metabolites were re-suspended in HPLC grade methanol. The mobile phase comprised of acetonitrile, ammonium formate (2 mM), and formic acid (0.1%) with a flow rate of 0.4 mL/min. The purity was checked by a UV detector at 450 nm. Elution was executed in an isocratic mode and with 15 min run time. LCMS was carried out by the Thermo Scientific, USA, equipped with C18 column. Mass spectra was obtained using an ion trap mass spectrometer fitted with an electronic spray (ESI, Thermo Scientific, USA) interface operated in a positive ionization mode (100–1000 m/z) with a spray voltage of 5 kV, capillary voltage 12 V, capillary temperature 275 °C, sheath gas at 30 AU, and auxiliary gas at 5 AU. The instrument was HPLC coupled with MS (LTQ XL system (linear ion trap)). Data was recorded using Xcalibur software (Asses et al. 2018).
Toxicity Study
The phytotoxicity as well as microbial toxicity was performed. The phytotoxicity test was performed by petri dish method. The test was carried out with 150 mg/L concentration of reactive orange 16 dye and its degraded metabolites by dissolving in 10 ml of distilled water. The sterile petri dish containing a double layered of Whatman no. 3 filter papers were prepared. Three dishes were prepared: dish soaked with water (control), dish soaked with dye solution and dish soaked with degraded metabolites. The healthy seeds of Vigna radiata were selected and surface sterilized with 1.2% sodium hypochlorite solution then washed thoroughly with deionized water. The sterilized seeds were soaked in respective solution for 1 h and then transferred to soaked petri dish (10 seeds on each petri dish). To maintain the moisture, seeds were water very alternate day. Length of root, shoot and germination rate (%) were calculated after 7 days of incubation (Shahi and Sapkota 2018; Haque et al. 2021). Experimental setup contained distilled water as control, dye, and its degraded metabolite. Growth inhibition zone of Escherichia coli was measured after 24 h of incubation at 35 °C.
Results and discussion
Isolation and screening of dye decolorizing bacteria
Nineteen bacterial isolates were obtained, because of their ability to grow on and decolorize reactive orange 16 dye. Of these, fourteen were Gram negative and five were Gram positive in nature. The Gram-negative bacteria resists harsh environment due to changes in the outer membrane such as mutation in porins, modified hydrophobic properties etc. (Breijeh et al. 2020). Based on the clear zone formation in plate assay containing reactive orange 16 dye (25 mg/L), six isolates were screened which showed decolorization potential. Efficiency of decolorization by each selected isolate was checked by preliminary batch experiments. Bacterial isolate was inoculated to media and incubated for 24 h, 48 h, and 72 h at 35 °C. Absorbance (A) was measured at 493 nm. Among the isolates, maximum decolorization efficiency was observed in case of SCA1007 (Fig. 1).
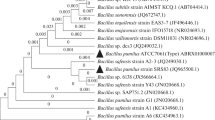
The bacterial isolate SCA1007 was initially identified according to Bergey’s Manual of Determinative Bacteriology (Table 1). Further identification of the strain was done based on 16S rRNA gene sequencing. Consensus sequence was aligned using clusterW. Basic local alignment search tool (BLAST) of the National Center for Biotechnology Information (NCBI) was used to study the homology of sequences. These sequences were selected based on maximum identity score and their alignment was done using clustalW. A phylogenetic tree was constructed using MEGA 6.06 software. (Fig. 2). Analysis of 16S rRNA gene sequence showed closest similarity with Bacillus stratosphericus. The 16S rRNA gene sequence of SCA1007 has been submitted GenBank under the accession number KY992944 (https://www.ncbi.nlm.nih.gov/nuccore/KY992944).
Decolorization analysis
UV–visible spectroscopy was used to confirm the decolorization of reactive orange 16. If color removal is through biodegradation, two things could happen: disappearance of visible light absorbance peak or appearance of new peak (Chen et al. 2003; Patil et al. 2010). The absorbance peak (at 0 h) of dye reduced within 10 h of incubation (Fig. 3). Culture, which was utilized for decolorization of reactive orange 16 was pelleted through centrifugation (at 10,000 rpm for 15 min). Absence of color in the pellet of Bacillus stratosphericus SCA1007 showed that decolorization was due to degradation of compound, not through adsorption. Laccase enzyme been produced by Bacillus stratosphericus SCA1007 which is confirmed by appearance of reddish-brown zone around bacteria. We believe laccase enzyme may attribute to bioconversion of reactive orange 16 (Kiiskinen et al. 2004; López et al. 2006).
Optimization of culture conditions
In order to examine the potential of Bacillus stratosphericus SCA1007 to decolorize dye, it was exposed to varying dye concentrations (25–400 mg/L). At concentrations above 150 mg/L, the decolorization percentage was inversely proportional to dye concentration (Fig. 4a). This is because, on increasing dye concentration, toxicity of dyes increases. This leads to restriction of metabolic activity which in turn inhibits cell growth (Shobana and Hangam 2012). Thus, dye concentration plays an important role in the choice of microbes utilized for removal of dyes from effluent, as high concentration can decrease the decolorization capacity due to the toxic nature of dyes (Khehra et al. 2006a).
The effect of pH on dye decolorization efficiency of Bacillus stratosphericus SCA1007 was checked, over a wide range of pH 4–11. Increase in decolorization percentage from pH 4 to 7 was observed. The optimum pH for decolorization of reactive orange 16 was found to be pH 7, at which complete decolorization of dye was recorded within 10 h of incubation (Fig. 4b). Mostly, azo dye decolorizing microbes have been reported to degrade dye at pH near neutral. Alcaligenes aquatilis showed maximum decolorization of Synozol Red at pH 7 (Ajaz et al. 2019). Reduced pH has a negative impact on decolorization efficiency of bacteria. Negative impact is due to formation of protonated azo dye at low pH. This alteration in chemical structure of dye hinders with bacterial decolorization (Deng et al. 2008).
To study the impact of temperature on dye decolorization, a range of temperatures from 4 to 55 °C was investigated. In this experiment, the initial dye concentration and the pH were fixed at 150 mg/L and 7 respectively. Results showed that the impact of temperature on decolorization was prominent over the examined range. The dye decolorization increased in proportion to the increase in temperature up to a maximum of 35 °C. On raising the temperature to 40 °C and above, there was drastic decrease in decolorizing potency of microorganisms. Thus, 35 °C was found to be optimal temperature for decolorization of reactive orange 16 (Fig. 4c). The result obtained is in accordance with a previous study which showed that 35 °C is the optimum temperature for maximum decolorization of reactive orange 16 by Pseudomonas sp. (Patel et al. 2013). Denaturation of catabolic enzymes might be the cause of decrease of metabolic activity at higher temperature (Chang and Lin 2001). In another study by a Masarbo et al. 2018, Bacillus sp. strain AK1, Lysinibacillus sp. strain AK2, and Kerstersia sp. strain VKY1 were able to decolorize sulfonated azo dye (Methyl Orange) at 35 °C within 18 h of incubation. Ambient temperature for maximum decolorization of reactive red 198 by consortia consisting of Enterococcus faecalis and Klebsiella variicola was found to be 37 °C. On further increasing the temperature to 40 °C, there was decline in decolorization percentage (Eslami et al. 2019).
The dye effluents contain high amounts of salt dissolved in it (Cui et al. 2014). The effect of salinity on the decolorization capacity of bacterial isolate was checked. The experiment was conducted using different concentrations of NaCl in a range of 0.1–7%. It was found that SCA1007 was tolerant to a wide range of salinity. Maximum decolorization of dyes was observed for up to 4% of salt concentration after 10 h of incubation (Fig. 4d), followed by decrease in decolorization. Similar results have been reported, where Bacillus fusiformis strain KMK5 could decolorize Acid orange 10 effectively within 48 h in 0.5–3% salt concentrations (Kolekar et al. 2008). In another study, the percentage decolorization using Klebsiella sp. was maintained above 80% at 1–4% salt concentrations at 48 h (Cui et al. 2014). A decline in decolorizing efficiency by microorganisms was probable due to increased salt concentration, which causes plasmolysis of cells which ultimately leads to decline in cellular activity (Wood 2015). Bacillus stratosphericus SCA1007 could efficiently decolorize reactive orange 16 over a wide range (1–4%), unlike in case of Comamonas acidovorans MTCC 3364 which showed tolerance up to 2% of sodium chloride (Rudakiya and Pawar 2013).
The decolorization of reactive orange 16 in static and shaking culture conditions was studied. The decolorization potential of SCA1007 was adversely affected in shaking condition. Highest decolorization was achieved under static culture condition within 10 h of incubation under optimized culture condition (Fig. 5). Similarly, Nocardiopsis alba showed decolorization of reactive orange 16 when incubated under static conditions. On the other hand, under shaking condition, culture grew well with almost negligible decolorization (Shobana and Hangam 2012). Alcaligenes aquatilis showed maximum decolorization of Synozol Red 6HBN at 37 °C, pH 7 under static culture condition (Ajaz et al. 2019). This may be because of the competition between oxygen and azo compounds for the reduced electron carriers under aerobic condition (Mabrouk and Yusef 2008). Other authors have also reported that for efficient color reduction, shaking, and vigorous aeration should be avoided (Chen et al., 1999; Khehra et al., 2006a, b). Aerobic process inhibits dye reduction process. This is because the electron released due to oxidation of electron donors by the cells are preferred to reduce oxygen instead of azo dye (Yoo et al. 2001).
Degradation analysis
The FTIR spectral of control dye (reactive orange 16) and its degraded products by Bacillus stratosphericus SCA1007 were compared. The spectral of reactive orange 16 dye exhibited peak at 624 cm−1 for C–C stretching. Bending of C-N cm−1 was indicated at 664 cm−1. The peak at 837 cm−1 represents S = O stretching. Bending of C-CH3 was displayed at 1002 cm−1. A peak at 1049 cm−1 represents C = O stretching. A peak at 1411 cm−1 showed stretching vibration of the benzene ring. N = N stretching was displayed by 1670 cm−1. Peak at 2924 cm−1 indicates presence of CH3 and CH2 groups and peak at 3406 cm−1 indicates N–H stretching. A slight change in IR spectra of peaks of degraded products as compared to control dye was observed. The peak at 757 cm−1 showed C-N stretching. Bending of C-H was indicated by 1103 cm−1. Stretching of C–OH and N = O was indicated by 1302 cm−1 and 1452 cm−1, respectively. The peak at 2959 cm−1 indicates C-H stretching. Similar FTIR results were observed for reactive orange 16 dyes degradation by bacteria (Telke et al. 2009) (Fig. 6).
Liquid chromatography- mass spectrometry analysis (LC–MS)
It was observed from the LC–MS/MS spectra of both dye and its biodegraded metabolites that there has been change in the pattern. The several compounds obtained after dyes decomposition at different m/z ratios confirmed the biodegradation of dyes by Bacillus stratosphericus SCA1007 (Fig. 7). Reactive orange 16 having molar mass 617 g/mol was biodegraded by Bacillus stratosphericus SCA1007. During the process of degradation, acetyl group having m/z 43 was removed which leads to the formation of disodium;2-[4-[(2-amino-8-hydroxy-6-oxidoperoxysulfanylnaphthalen-1-yl)diazenyl]phenyl]sulfonylethylsulfate (A) 575 (m/z). At this point desulfonation takes place which gives rise to sodium (E)-6-amino-4-hydroxy-3-((4-(methylamino)phenyl)diazenyl)naphthalene-2-sulfonate (B) 405 (m/z). Further reaction was followed by two processes (i) removal of methyl group which gives an intermediate sodium (E)-6-amino-3-((4-aminophenyl)diazenyl)4-hydroxynaphthalene-2-sulfonate (C) 380 (m/z) and (ii) desulfonation which leads to formation of 7-amino-2-(p-tolyldiazenyl)naphthalen-1-ol compound with methyl-l2-azane (1:1) (D) 309 (m/z). After this step removal of aniline group, addition of sulfur and sodium and deamination reaction takes place which gives an intermediate sodium 3-acetamido-5-hydroxybenzenesulfonate (E) 253 (m/z). This step is followed by removal of ammonia and acetyl group by formation of sodium 3 hydroxy 5 acetlylbenzene sulphonate (F) 196 (m/z). Further removal of -OH from (F) 196 (m/z) leads to formation of benzene (G) 180 (m/z) (Fig. 8). Similar degradation pattern has been observed by Soni et al. (2015).
Toxicity study
The release of effluent from textile industry in the form of treated and untreated wastewater affects the soil fertility, hence agricultural productivity. It is necessary to evaluate the toxicity of the dye before and after degradation because environmental health criteria require for both the removal and detoxification of pollutants. Since dyes are harmful to environment, it is important to check the microbial as well as phytotoxicity of dye and its degraded products. Phytotoxicity study was performed on Vigna radiata towards dyes and their degraded products by calculating seed germination (%). Germination of seeds was more with degraded metabolite and distilled water, as compared to dyes, indicating a less toxic effect of degradation products as compared to dyes. The present study revealed that biodegradation of reactive orange 16 by Bacillus strastosphericus SCA1007 led to detoxification and production of less toxic metabolites, thus suggesting bio-treated wastewater containing dye could be used for agricultural purpose (Fig. 9) (Table 2). For analysis of microbial toxicity, the inhibition zone was found to be (4 mm) with control dyes (reactive orange 16), while its degraded metabolite displayed no growth inhibition.
Conclusion
The study concludes that Bacillus stratosphericus SCA1007 can be utilized effectively for complete decolorization of reactive orange 16 at a dye concentration of 150 mg/L within 10 h at pH 7 incubated at 35 °C. The bacterial isolate could also tolerate salinity as high as 4%. It was also observed that dye decolorization by the isolated strain was by biological degradation. This is the first report of reactive orange 16 dye degradation using Bacillus strastosphericus SCA1007. Dye degradation property of Bacillus strastosphericus SCA1007 can be further used as a basic tool for bioremediation of various dyes in textile effluents.
Availability of data and materials
Not applicable.
Code availability
Not applicable.
References
Ajaz M, Rehman A, Khan Z et al (2019) Degradation of azo dyes by Alcaligenes aquatilis 3c and its potential use in the wastewater treatment. AMB Express 9:64. https://doi.org/10.1186/s13568-019-0788-3
Ali H (2010) Biodegradation of synthetic dyes—a review. Water Air Soil Pollut 213:251–273. https://doi.org/10.1007/s11270-010-0382-4
Asses N, Ayed L, Hkiri N, Hamdi M (2018) Congo Red Decolorization and detoxification by Aspergillus niger : removal mechanisms and dye degradation pathway. Biomed Res Int 2018:1–9. https://doi.org/10.1155/2018/3049686
Bhatia D, Sharma NR, Singh J, Kanwar RS (2017) Biological methods for textile dye removal from wastewater: a review. Crit Rev Environ Sci Technol 47:1836–1876. https://doi.org/10.1080/10643389.2017.1393263
Breed RS, Murray EGD, Smith NR (1957) Bergey’s manual of systematic bacteriology
Breijyeh Z, Jubeh B, Rafi K (2020) Resistance of gram negative bacteria to current antibacterial agent and approaches to resolve it. Molecules 25:1340. https://doi.org/10.3390/molecules25061340
Chang J, Lin C (2001) Decolorization kinetics of a recombinant Escherichia coli strain harboring azo-dye-decolorizing determinants from Rhodococcus sp. Biotechnol Lett 23:631–636. https://doi.org/10.1023/A:1010306114286
Chen K-C, Huang W-T, Wu J-Y, Houng J-Y (1999) Microbial decolorization of azo dyes by Proteus mirabilis. J Ind Microbiol Biotechnol 23:686–690. https://doi.org/10.1038/sj.jim.2900689
Chen KC, Wu JY, Liou DJ, Hwang SCJ (2003) Decolorization of the textile dyes by newly isolated bacterial strains. J Biotechnol 101:57–68. https://doi.org/10.1016/S0168-1656(02)00303-6
Cui D, Li G, Zhao M, Han S (2014) Decolourization of azo dyes by a newly isolated Klebsiella sp. strain Y3, and effects of various factors on biodegradation. Biotechnol Biotechnol Equip 28:478–486. https://doi.org/10.1080/13102818.2014.926053
Deng D, Guo J, Zeng G, Sun G (2008) Decolorization of anthraquinone, triphenylmethane and azo dyes by a new isolated Bacillus cereus strain DC11. Int Biodeterior Biodegradation 62:263–269. https://doi.org/10.1016/j.ibiod.2008.01.017
Dos SA, Cervantes F, Van LJ (2007) Review paper on current technologies for decolourisation of textile wastewaters: perspectives for anaerobic biotechnology. Bioresour Technol 98:2369–2385. https://doi.org/10.1016/j.biortech.2006.11.013
Elango G, G R, Elango S, (2017) Physico-chemical parameters of textile dyeing effluent and its impacts with case study. Int J Res Chem Env 7:17–24
Gao T, Qin D, Zuo S et al (2020) Decolorization and detoxification of triphenylmethane dyes by isolated endophytic fungus, Bjerkandera adusta SWUSI4 under non-nutritive conditions. Bioresour Bioprocess 7:53. https://doi.org/10.1186/s40643-020-00340-8
Geetha K, Velmani N (2015) Diverse technology and methods for dye treatment. Asian J Chem 27:1177–1184. https://doi.org/10.14233/ajchem.2015.17804
Habib S, Ahmad SA, Johari WLW et al (2018) Evaluation of conventional and response surface level optimisation of n-dodecane (n-C12) mineralisation by psychrotolerant strains isolated from pristine soil at Southern Victoria Island. Antarctica Microb Cell Fact 17:44. https://doi.org/10.1186/s12934-018-0889-8
Eslami H, Shariatifar A, Rafiee E, Shiranian M, Salehi Sadat Saeede F, Hosseini S, Eslami G, Ghanbari R, Ebrahimi AA (2019) Decolorization and biodegradation of reactive Red 198 Azo dye by a new Enterococcus faecalis–Klebsiella variicola bacterial consortium isolated from textile wastewater sludge. World J Microbiol Biotechnol 35(3) https://doi.org/10.1007/s11274-019-2608-y
Haque M, Haque A, Mosharaf k, Marcus PK, (2021) Decolorization, degradation and detoxification of carcinogenic sulfonated azo dye methyl orange by newly developed biofilm consortia. Saudi J Biol Sci 28:793–804. https://doi.org/10.1016/j.sjbs.2020.11.012
Kalyani DC, Telke AA, Govindwar SP et al (2009) Biodegradation and detoxification of reactive textile dye by isolated Pseudomonas sp SUK1. Water Environ Res 81:298-307.
Kapdan IK, Oztekin R (2003) Decolorization of textile dyestuff reactive orange 16 in fed-batch reactor under anaerobic condition. Enzyme Microb Technol 33:231–235. https://doi.org/10.1016/S0141-0229(03)00128-5
Karim ME, Dhar K, Hossain MT (2018) Decolorization of textile reactive dyes by bacterial monoculture and Consortium screened from textile dyeing effluent. J Genet Eng Biotechnol 16:375–380. https://doi.org/10.1016/j.jgeb.2018.02.005
Khehra MS, Saini HS, Sharma DK et al (2006a) Biodegradation of azo dye C.I. Acid Red 88 by an anoxic–aerobic sequential bioreactor. Dye Pigment 70:1–7. https://doi.org/10.1016/j.dyepig.2004.12.021
Khehra MS, Saini HS, Sharma DK et al (2006b) Biodegradation of azo dye C.I. Acid Red 88 by an anoxic - Aerobic sequential bioreactor. Dye Pigment 70:1–7. https://doi.org/10.1016/j.dyepig.2004.12.021
Kiiskinen LL, Rättö M, Kruus K (2004) Screening for novel laccase-producing microbes. J Appl Microbiol 97:640-646
Kolekar YM, Pawar SP, Gawai KR et al (2008) Decolorization and degradation of disperse blue 79 and acid orange 10, by Bacillus fusiformis KMK5 isolated from the textile dye contaminated soil. Bioresour Technol 99:8999–9003. https://doi.org/10.1016/j.biortech.2008.04.073L
López MJ, Guisado G, Vargas-García MC et al (2006) Decolorization of industrial dyes by ligninolytic microorganisms isolated from composting environment. Enzyme Microb Technol 40:42–45. https://doi.org/10.1016/j.enzmictec.2005.10.035
Mabrouk ME, Yusef H (2008) Decolorization of fast red by Bacillus subtilis HM. J Applies Sci Res 4:262–269
Masarbo RS, Ismailsab M, Monisha TR et al (2018) Enhanced decolorization of sulfonated azo dye methyl orange by single and mixed bacterial strains AK1, AK2 and VKY1. Bioremediat J 22:136–146. https://doi.org/10.1080/10889868.2018.1516612
Megha V, Meenakshi S, Jpn R (2015) Optimization of different parameters on synthetic dye decolorization by free and immobilized Mucor hiemalis MV04 (KR078215). Res J Chem Sci Res J Chem Sci 5:2231–2606
Ojekunle OZ, Lateef S (2017) Environmental impact of abattoir waste discharge on the quality of surface water and ground water in Abeokuta. J Environ Anal Toxicol 07:1–10. https://doi.org/10.4172/2161-0525.1000509
Park C, Lee M, Lee B et al (2007) Biodegradation and biosorption for decolorization of synthetic dyes by Funalia trogii. Biochem Eng J 36:59–65. https://doi.org/10.1016/j.bej.2006.06.007
Patel K, Shah M, Darji A, Nair S (2013) Exploiting application of Pseudomonas spp. ETL-2013 in microbial degradation and decolorization of disperse orange 3. J Bioremediation Biodegrad 04:1–7. https://doi.org/10.4172/2155-6199.1000193
Patil PS, Phugare SS, Jadhav SB, Jadhav JP (2010) Communal action of microbial cultures for Red HE3B degradation. J Hazard Mater 181:263–270. https://doi.org/10.1016/j.jhazmat.2010.05.006
Pillai HPJS (2017) Optimization of process conditions for effective degradation of azo blue dye by streptomyces DJP15. J Pure Appl Microbiol 11:1757–1765. https://doi.org/10.22207/JPAM.11.4.14
Popli S, Patel UD (2015) Destruction of azo dyes by anaerobic–aerobic sequential biological treatment: a review. Int J Environ Sci Technol 12:405–420. https://doi.org/10.1007/s13762-014-0499-x
Pritchard PH, Costa CF (1991) EPA’s Alaska oil spill bioremediation project. Part 5. Environ Sci Technol 25:372–379. https://doi.org/10.1021/es00015a002
Rudakiya DM, Pawar KS (2013) Optimization of culture condition for enhanced decolorization of reactive orange 16 by Comamonas acidovorans MTCC 3364. Int J Microbiol Appl Sci 2:467–476
Saha P, Rao KVB (2019) Biotransformation of Reactive Orange 16 by alkaliphilic bacterium Bacillus flexus VITSP6 and toxicity assessment of biotransformed metabolites. Int J of Environ Sci and Technol 5:1–16. https://doi.org/10.1007/s13762-019-02256-z
Shahi D, Sapkota R (2018) A comparative study on dye degradation by leaf and root extracts of Parthenium hysterophorus L. Int J Appl Sci Biotechnol 6:327–331. https://doi.org/10.3126/ijasbt.v6i4.22110
Sharma S, Roy S (2015) Biodegradation of dye reactive black - 5 by a novel bacterial endophyte. Int Res J Environ Sci 4:44–53
Shobana S, Hangam BT (2012) biodegradation and decolorization of reactive orange 16 by Nocardiopsis alba soil isolate. J Bioremediation Biodegrad 03:1–7. https://doi.org/10.4172/2155-6199.1000155
Soni RK, Acharya PB, Modi HA (2015) Elucidation of biodegradation mechanism of reactive red 35 by Pseudomonas aeruginosa ARSKS20. IOSR J Environ Sci Ver I 9:31–40. https://doi.org/10.9790/2402-09813140
Telke AA, Kalyani DC, Dawkar VV, Govindwar SP (2009) Influence of organic and inorganic compounds on oxidoreductive decolorization of sulfonated azo dye C.I. Reactive Orange 16. J Hazard Mater 172:298–309. https://doi.org/10.1016/j.jhazmat.2009.07.008
Tomczak WE, Gorecki L (2012) Azo dyes - Biological activity and synthetic strategy. Chemik 66:1298–1307
Wood JM (2015) Bacterial responses to osmotic challenges. J Gen Physiol 145:381–388. https://doi.org/10.1085/jgp.201411296
Yadav AN, Sachan SG, Verma P, Saxena AK (2016) Bioprospecting of plant growth promoting psychrotrophic bacilli from the cold desert of north western Indian Himalayas. Indian J Exp Biol 54:142–150
Yaseen DA, Scholz M (2018) Treatment of synthetic textile wastewater containing dye mixtures with microcosms. Environ Sci Pollut Res 25:1980–1997. https://doi.org/10.1007/s11356-017-0633-7
Yoo ES, Libra J, Adrian L (2001) Mechanism of decolorization of azo dyes in anaerobic mixed culture. J Environ Eng 127:844–849. https://doi.org/10.1061/(ASCE)0733-9372(2001)127:9(844)
Zhou W, Zimmermann W (1993) Decolorization of industrial effluents containing reactive dyes by actinomycetes. FEMS Microbiol Lett 107:157–161. https://doi.org/10.1111/j.1574-6968.1993.tb06023.x
Zin KM, Effendi Halmi MI, Abd Gani SS et al (2020) Microbial decolorization of triazo dye, direct blue 71: an optimization approach using response surface methodology (RSM) and artificial neural network (ANN). Biomed Res Int 2020:1–16. https://doi.org/10.1155/2020/2734135
Acknowledgements
The authors would like to thank the Central Instrumentation Facility and Department of Bioengineering and Biotechnology, BIT, Mesra, Ranchi for providing the infrastructure and laboratory facilities. The authors are also thankful to the Department of Chemistry, Indian Institute of Technology, Patna for providing the facilities for research work. A special thanks to Amar Nath sir for their valuable suggestions for MS analysis.
Funding
This study is funded by BIT, Mesra for seed money scheme (DSR/SMS-2015/2015–2016(3122)).
Author information
Authors and Affiliations
Contributions
Conceptualization, methodology, original draft preparation, and review and editing: Kriti Akansha. Liquid chromatography-mass spectrometry analysis (LC–MS): Manish Kumar. Figures: Ajar Nath Yadav. Supervision: Shashwati Ghosh Sachan and Debashis Chakraborty.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Akansha, K., Yadav, A.N., Kumar, M. et al. Decolorization and degradation of reactive orange 16 by Bacillus stratosphericus SCA1007. Folia Microbiol 67, 91–102 (2022). https://doi.org/10.1007/s12223-021-00914-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-021-00914-9