Abstract
In the present study, bacterial isolates were screened for arsenic resistance efficiency. Environmental isolates were isolated from arsenic-rich soil samples (i.e., from Rajnandgaon district of Chhattisgarh state, India). Amplification and sequencing of 16S rRNA gene revealed that the isolates were of Bacillus firmus RSN1, Brevibacterium senegalense RSN2, Enterobacter cloacae RSN3, Stenotrophomonas pavanii RSN6, Achromobacter mucicolens RSN7, and Ochrobactrum intermedium RSN10. Arsenite efflux gene (arsB) was successfully amplified in E. cloacae RSN3. Atomic absorption spectroscopy (AAS) analysis showed an absorption of 32.22% arsenic by the RSN3 strain. Furthermore, results of scanning electron microscopy (SEM) for morphological variations revealed an initial increase in the cell size at 1 mM sodium arsenate; however, it was decreased at 10 mM concentration in comparison to control. This change of the cell size in different metal concentrations was due to the uptake and expulsion of the metal from the cell, which also confirmed the arsenite efflux system.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Heavy metal contamination in the environment is a buzzing topic nowadays. Extensive growth in the industrial activities has led to the production of several heavy metal pollutants that contaminate soil and water, thereby harming our agroecosystem (Agrawal et al. 2015). Several novel physical and chemical strategies have been adopted for solving this problem, which ultimately proved to be costly and ineffective (Ayangbenro and Babalola 2017). Over few years, different new avenues of microbial bioremediation, i.e., bioaccumulation, biosorption, bioprecipitation, bioleaching, biotransformation, biosurfactants, and siderophore formation, have emerged as a new hope (Agrawal et al. 2018; Mosa et al. 2016; Kang et al. 2016). These eco-friendly methods are quite successful in addressing the issues lacking in their predecessors. It has reported that bioremediation can save up to 50–65% of the total cost as compared to conventional approaches (Ojuederie and Babalola 2017). Due to their widespread abundance, easy culturability, and varied mechanisms towards heavy metal tolerance, bacteria can be used as a suitable tool for heavy metal remediation.
A typical ars operon is generally transcribed as a single unit (Rosen 1999) containing either three (arsRBC) or five (arsRDABC) genes, which can be either plasmid or chromosome borne (Diorio et al. 1995). Sequential events and their associated genes involved in detoxification of arsenic are arsC, encoding As5+ reductase responsible for the reduction of As5+ to As3+ in the cytoplasm (Ji and Silver 1992); arsB, encoding a membrane protein that functions as an As3+ pump using the membrane potential force (Tisa and Rosen 1990); and arsR, as a regulator of the arsRBC operon by repressing the transcription of this operon in the absence of As3+ (Wu and Rosen 1993).
Thus, in this way, arsB gene indirectly provides resistance to arsenate by converting it into arsenite, which is further detoxified to form monomethylarsenite (MMAs(III)), dimethylarsenite (DMAs(III)), and trimethylarsine (TMAs) (Shen et al. 2013; Kruger et al. 2013). It has reported that bacteria contain arsenite oxidizing gene and arsenite transporter gene or any one of them (Sanyal et al. 2016). In the present study, six arsenic-resistant bacterial isolates were screened from different arsenic-contaminated sites of Chhattisgarh. A molecular approach was employed to decipher the arsenic resistance mechanism and the bioremediation potential of these native isolates.
Materials and methods
Collection of soil sample and isolation of As-resistant bacteria
Surface soil samples (0–15 cm) were collected from Rajnandgaon district of Chhattisgarh, a state in central India. This state was earlier reported for heavy arsenic content in the soil. All samples (250 g) were collected in sterile plastic bags and refrigerated until further use. The soil samples were serially diluted up to 10−4 plated on nutrient agar (NA) plates containing 1 mM concentrations of sodium arsenite (NaAsO2). The plates were incubated at 37 °C for 48 h to 72 h until colonies developed. Colonies with distinct morphology were further screened for sodium arsenate tolerance.
Determination of minimum inhibitory concentration
Flasks containing minimal broth (Dextrose, 1 g/L; dipotassium phosphate 7 g/L; monopotassiumphosphate, 2 g/L; sodium citrate, 0.5 g/L; magnesium sulphate, 0.1 g/L; ammonium sulphate, 1 g/L) were amended with different concentrations of heavy metal (HM) salt; sodium arsenate (20, 40, 50, 100, 200, 400, 600 and 700 mM). Single colony (CFU) from a pre-streaked plate of each isolate was inoculated with different concentrations of respective HM and incubated at 37 °C for 72 h. The growth of each isolate was determined spectrophotometrically by measuring the optical density (O.D.) at 600 nm.
Isolation of genomic DNA
Six bacterial isolates (RSN1, RSN2, RSN3, RSN6, RSN7, and RSN10) were revived in nutrient agar medium in the presence of 1-mM sodium arsenate (Na3AsO4) by the streak plate method. This concentration (1 mM) was previously by many researchers for initial screening of arsenic resistance bacteria (Banerjee et al. 2011; Selvi et al. 2014). Briefly, 100 μL of inoculum (0.6 OD overnight grown culture) was inoculated in nutrient broth (with 1 mM Na3AsO4) and incubated at 37 °C in a BOD incubator with constant shaking at 100 rpm for 12 h after that genomic DNA was isolated using Genomic DNA Isolation Kit (Himedia Laboratories Pvt. Ltd., Mumbai, India).
Amplification of 16S rRNA gene and in silico analysis
The isolated genomic DNA of all bacterial isolates were amplified for 16S rRNA gene using a set of universal primers U1 (5′-CCAGCAGCCGCGGTAATACG-3′) and U2 (5′ATCGGCTACCTTGTTACGACTTC-3′) originally designed by Lu et al. (2000). The PCR mixture (25 μL) was composed of 1 μm of each primer, 2.5 mM of dNTPs, 1 U Taq polymerase, PCR buffer with 2.5 mM MgCl2, and 1 μL of template DNA. The PCR reaction was carried out in a T100TM thermal cycler (Bio-Rad Laboratories, Inc.) with an initial denaturing temperature of 94 °C for 10 min followed by denaturing temperature of 94 °C for 1 min, annealing temperature of 55 °C for 1 min, extension temperature of 72 °C for 2 min and with having a final extension temperature of 72 °C for 10 min. The amplified products were partially sequenced by Sanger sequencing technique, and the gene sequences were deposited in NCBI GenBank. Furthermore, the BLAST algorithm (NCBI BLAST) was used to compare the gene sequences with the known ones to find their level of identity. The sequences were aligned using ClustalW, and their phylogenetic relationship was established by constructing a phylogram through neighbor-joining method in MEGA X.
Antibiotic susceptibility test and silver nitrate assay
All the cultures were studied for their co-tolerance towards heavy metals and antibiotics. The cultures were first grown in nutrient broth with 1 mM Na3AsO4 overnight for 16 h and after that 100 μL culture was evenly spread over nutrient agar plates having the same concentration metal along with HiPer Antibiotic Sensitivity discs (HiMedia Laboratories Pvt. Ltd., India) consisting of eight antibiotics. Silver nitrate assay was performed for arsenic-resistant bacterial strains to check the As(III) to As(V) and As(V) to As(III) transformation abilities of the isolates. The entire experiment was performed in duplicate (Simeonova et al. 2004; Selvi et al. 2014).
PCR amplification of arsB gene
The genomic DNA of all bacterial isolates were used for amplification of arsenite efflux gene arsB through a set of primers, i.e., darsB1F (5′-GGTGTGGAACATCGTCTGGAAYGCNAC3′) and darsB1R (5′-CAGGCCGTACACCACCAGRTACATNCC-3′), originally designed by Achour et al. (2007). The polymerase chain reaction protocol for arsB was performed in a 25 μL mixture composed of 1 μm of each primer, 2.5 mM of dNTPs, 1 U Taq polymerase, PCR buffer with 2.5 mM MgCl2, and 1 μL of template for 35 cycles (denaturation for 45 s at 94 °C, annealing of 50 °C for 45 s, extension of 72 °C for 50 s, and final extension of 72 °C for 10 min after completion of an initial denaturation of 94 °C for 5 min).
Bioremediation assay
The selected isolate was grown in nutrient broth (Dey et al. 2016) in the presence (1 mM Na3AsO4) and absence of the arsenic salt along with a control (uninoculated). It was continuously shaken (at 100 rpm) for 7 days, and its growth was periodically monitored in every 24 h at 600 nm in a Vis-spectrophotometer (Systronics India). Simultaneously, the environmental isolate was also grown in 100 ppm (0.5 mM) Na3AsO4 along with positive control (no metal), and samples were drawn on the 8th day (in decline phase) for AAS analysis. The sample was centrifuged at 6000 rpm for 5 min, and then the cell-free supernatant was processed for AAS analysis.
Scanning electron microscopy analysis
The best bacterial isolate was selected for cell size differentiation analysis using SEM. The culture was grown in minimal media with 1 mM and 10 mM metal salt concentrations; however, culture without metal salt was treated as control. The cells were fixed with 2.5% glutaraldehyde in 0.1-M phosphate buffer (pH 7.2) for 24 h at 4 °C and 2% aqueous osmium tetroxide post-fixed for 4 h. The processed samples were then examined under the scanning electron microscope (model: FEI Quanta 250) at 30 kV with 30,000X magnification for detection of any morphological changes in shape, size, density, and volume of cells (work done at Central Drug Research Institute, Lucknow, India). Cell size was measured by the manual method, i.e., “Old Way by Hand” method. The scale bar indicates that the actual distance on the specimen (for that bar) is 1 μm. The bar distance was the same for all three SEM micrographs. The length of scale bar was measured in mm, which was considered to be equivalent to 1 μm. Similarly, cell sizes (5 cells / micrograph) were measured in mm and converted them into μm.
Results and discussion
Isolation of As-resistant bacteria
Only 20 bacterial isolates were obtained in nutrient agar medium containing 10 ppm arsenite. Most of the arsenic-resistant bacteria were obtained from soil samples from Rajnandgaon and, thus, suggested the presence of As in soil. Microbial resistance to arsenate [As (V)] and arsenite [As (III)] was determined by visible growth after 72 h in minimal broth supplemented with varying concentrations of sodium arsenate and sodium arsenite. Six bacterial isolates (RSN1, RSN2, RSN3, RSN6, RSN7, and RSN10) have shown growth on both the arsenic salts. The highest MIC was shown by isolate RSN3, i.e., 600 mM grown in minimal salt medium with sodium arsenate. This tolerance capacity is better than some previously reported environmental isolates (Anderson and Cook 2004; Wolfe-Simon et al. 2011; Bagade et al. 2016; Firrincieli et al. 2019). However, based on maximum tolerance, RSN3 was finally selected for further bioremediation studies (Fig. 1a).
a Minimum inhibitory concentrations of selected isolates grown in minimal broth with variable concentrations of metal salt. b Growth curve of RSN3 isolate in the presence and absence of arsenic (Na3AsO4). Each point represents the mean of three independent experiments. Bars, standard error of the means
Molecular characterization
The amplification of the 16S rRNA gene resulted in yielding approx. 996 bp DNA amplicons as reported previously by Lu et al. (2000). These sequenced amplicons were compared with the BLAST algorithm (https://blast.ncbi.nlm.nih.gov/Blast.cgi) for homology (Fig. 2)). All the identified genera’s have already reported earlier showing resistance towards arsenic Achromobacter (Li et al. 2012; Bhosale et al. 2014), Bacillus (Dey et al. 2016), Enterobacter (Selvi et al. 2014), Brevibacterium (Sunitha et al. 2015), Ochrobactrum (Sarkar et al. 2014), and Stenotrophomonas (Bachate et al. 2009). The amplified products were partially sequenced, and the gene sequences were deposited in NCBI GenBank retrieving the following accession numbers, i.e., MG857853 (RSN1), MG857854 (RSN2), MG857855 (RSN3), MG857856 (RSN6), MG857857 (RSN7), and MG857858 (RSN10).
Phylogenetic tree of arsenic-resistant bacterial isolates constructed by the neighbor-joining method using MEGA X. The scale bar indicates 2% nucleotide sequence substitution. The optimal tree with the sum of branch length = 0.73389547 is shown. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (1000 replicates) is shown next to the branches
Antibiotics resistance and silver nitrate assay
The results obtained through antibiotic susceptibility test (Fig. 3) showed that all the isolates were resistant to the penicillin and rifamycin. Rifamycin (rifampin like an antibiotic) resistance occurred in various species of bacteria may be due to the modification in targeting this antibiotic. The rpoB gene codes for β-subunit of RNA polymerase (Padayachee and Klugman 1999). Among these six isolates, Enterobacter cloacae RSN3 and Ochrobactrum intermedium RSN10 had shown maximum tolerance for penicillin as it was the first generation antibiotic. However, E. cloacae was not earlier reported for these antibiotics resistance, but O. intermedium was previously reported as an extensive antibiotic-resistant against penicillin and rifampin (Johnning et al. 2013; Poszytek et al. 2018). Results exhibited tolerance developed by bacterial strain for older antibiotics. However, an increase in environmental pollutants may also be responsible for resistance towards antibiotics (Samanta et al. 2012). It has reported that heavy metals act as co-selecting agents in metal-resistant bacterial strains for the propagation of resistance in bacteria against antibiotics (Nguyen et al. 2019; Eduardo-Correia et al. 2020). Although the co-occurrence of heavy metal resistance and antibiotic resistance in the contaminated area was not clearly understood, the selection of either of the properties may characterize the genes for other.
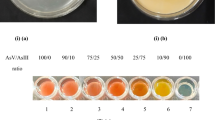
Silver nitrate test resulted in the formation of a light yellow-colored precipitate, which revealed that all bacterial isolates were arsenate reducing bacteria. These results were in agreement with previous reports of Govarthanan et al. 2015 and Selvankumar et al. 2017.
PCR amplification of arsB gene
The arsB gene was successfully amplified from RSN3 bacterial isolate, i.e., E. cloacae RSN3 out of six isolates. The amplicon was of approx. 750 bp in size as shown in Fig. 4, and it was the expected size according to Achour et al. 2007. The arsB is mainly code for an arsenite efflux gene that can either work independently or in association with arsA gene (Dey and Rosen 1995). We were able to amplify this gene from chromosomal DNA of the bacteria; however, this gene was found both in chromosome and plasmid (Butcher et al. 2000). Further, ArsB is an integral membrane protein able to evict arsenite from the cell cytoplasm, thus diminishing arsenite accumulation in the cell (Yang et al. 2012). The earlier experiments conducted in other bacteria explained that ArsB functions more efficiently when it works in association with ArsA (Rosen 2002). Interestingly, arsBC gene pair was common in the chromosomes of Gram-negative bacteria and chromosomes and plasmids of Gram-positive bacteria, but in the latter case, no arsA gene was present. As RSN3 is also a Gram-negative bacterium, and this information also confirmed the presence of arsA gene in RSN3 (Ben Fekih et al. 2018). These findings also revealed that arsB gene in E. cloacae works independently without its association with arsA. It means that the presence of arsB gene gives some insight about the probable mechanism of arsenic resistance in E. cloacae RSN3. On the other hand, arsenic resistance genes were characterized by several Enterobacteriaceae family members, but we found very few publications related to the amplification arsB gene from E. cloacae (Saltikov and Olson 2002).
Growth in the presence of As
Results exhibited that exponential phase of isolate RSN3 was very short (i.e., of only 24 h) when allowed to grow without heavy metal salt, but after 24 h it moved towards declining phase (Fig. 1b). In contrast, it has extended log phase of 3 days with a short stationary phase in the presence of arsenic (As) after that the declining phase starts from the 5th day onwards. The similar finding was also reported by Abbas et al. (2014) and reported that the stationary phase in non-stressed condition was longer than stressed condition (in presence of As). Results of AAS samples analysis revealed that RSN3 was able to absorb 32.22% arsenic (i.e., around 1/3 reduction of arsenic from control), which was quite good, and thus, it may be effectively used as a bioremediation tool in future.
Scanning electron microscopy
The bacterial cells recovered after growth curve experiment were observed at higher magnification (i.e., at × 30,000) for any morphological changes (Fig. 5, 6, and 7). Results showed a significant increase in the cell size (2.01 μm) as compared to control (1.24 μm) having no metal at 1-mM concentration. This suggested that there was an accumulation of arsenic inside the bacterial cell while growing in the presence of metal (Table 1). Interestingly, with a further rise in the metal concentration up to 10 mM, a noticeable decrease (1.51 μm) in the cell volume was observed. It was already confirmed from PCR amplification that the bacterium has arsB gene, which was responsible efflux of arsenic. ArsB mediated a primary arsenite pump in E. coli also (Yoon 2005). Similarly, morphological changes like an increase in cell size and shape were reported in Acidocella bacterium as a defense mechanism to reduce the heavy metal stress (Chakravarty et al. 2007). It has reported that some bacteria also form a chain like an arrangement in the presence of metal (Dey et al. 2016) and sometimes they decrease their surface area (Mohamed and Farag 2015). Further, our SEM finding also revealed that the change of cell length was due to the arsenic stress in E. cloacae.
Conclusion
The results obtained from PCR amplification and scanning electron microscopic studies indicated the presence of arsenite efflux system in E. cloacae RSN3 through which reduced arsenic extruded from the cell through arsB pump. Moreover, atomic absorption spectrophotometer analysis showed absorption of 32.22% of arsenic from the culture medium by this bacterium, which was quite significant, and therefore its arsenic resistance property can be further exploited for bioremediation purpose. Meanwhile, antibiotic susceptibility test revealed that this bacterium was resistant towards most of the antibiotics (particularly older generation antibiotics) besides showing resistance towards heavy metal (arsenic), therefore displaying co-resistance and cross-resistance towards these two components, i.e., antibiotics and heavy metals.
References
Abbas SZ, Riaz M, Ramzan N, Zahid MT, Shakoori FR, Rafatullah M (2014) Isolation and characterization of arsenic resistant bacteria from wastewater. Braz J Microbiol 45:1309–1315
Achour AR, Bauda P, Billard P (2007) Diversity of arsenite transporter genes from arsenic-resistant soil bacteria. Res Microbiol 158:128–137
Agrawal R, Satlewal A, Varma A (2015) Characterization of plant growth-promoting rhizobacteria (pgpr): a perspective of conventional versus recent techniques. In: Sherameti I, Varma A (eds) Heavy metal contamination of soils. Soil biology, vol 44. Springer, Cham
Agrawal R, Verma A, Satlewal A (2018) Bioprospecting PGPR microflora by novel immuno based techniques. In: Prasad R, Gill SS, Tuteja N (eds) New and future developments in microbial biotechnology and bioengineering: crop improvement through microbial biotechnology. Elsevier,pp 465–478. https://doi.org/10.1016/B978-0-444-63987-5.00024-4
Anderson CR, Cook GM (2004) Isolation and characterization of arsenate-reducing bacteria from arsenic-contaminated sites in New Zealand. Curr Microbiol 48:341–347
Ayangbenro AS, Babalola OO (2017) A new strategy for heavy metal polluted environments: a review of microbial biosorbents. Int J Environ Res Public Health 14:1–16
Bachate SP, Cavalca L, Andreoni V (2009) Arsenic-resistant bacteria isolated from agricultural soils of Bangladesh and characterization of arsenate-reducing strains. J Appl Microbiol 107:145–156
Bagade AV, Bachate SP, Dholakia BB, Giri AP, Kodam KM (2016) Characterization of Roseomonas and Nocardioides spp. for arsenic transformation. J Hazard Mater 318:742–750
Banerjee S, Datta S, Chattyopadhyay D, Sarkar P (2011) Arsenic accumulating and transforming bacteria isolated from contaminated soil for potential use in bioremediation. J Environ Sci Health A 46:1736–1747
Ben Fekih I, Zhang C, Li YP, Zhao Y, Alwathnani HA, Saquib Q, Rensing C, Cervantes C (2018) Distribution of arsenic resistance genes in prokaryotes. Front Microbiol 9:2473
Bhosale GP, Bachate SP, Kale SC (2014) Isolation and characterization of arsenate reducing bacteria from the waste water of an electroplating industry. Int J Curr Microbiol App Sci 3:444–452
Butcher BG, Deane SM, Rawlings DE (2000) The chromosomal arsenic resistance genes of thiobacillus ferrooxidans have an unusual arrangement and confer increased arsenic and antimony resistance to Escherichia coli. Appl Environ Microbiol 66:1826–1833
Chakravarty R, Manna S, Ghosh AK, Banerjee PC (2007) Morphological changes in an Acidocella strain in response to heavy metal stress. Res J Microbiol 2:742–748
Dey S, Rosen BP (1995) Dual mode of energy coupling by the oxyanion translocating ArsB protein. J Bacteriol 177:385–389
Dey U, Chatterjee S, Mondal NK (2016) Isolation and characterization of arsenic-resistant bacteria and possible application in bioremediation. Biotechnol Rep 10:1–7
Diorio C, Cai J, Marmor J, Shinder R, DuBow MS (1995) An Escherichia coli chromosomal ars operon homolog is functional in arsenic detoxification and is conserved in gram-negative bacteria. J Bacteriol 177(8):2050–2056
Eduardo-Correia B, Morales-Filloy H, Abad JP (2020) Bacteria from the multi-contaminated Tinto river estuary (sw, Spain) show high multi-resistance to antibiotics and point to Paenibacillus spp. as antibiotic-resistance-dissemination players. Front Microbiol 10:3071
Firrincieli A, Presentato A, Favoino G, Marabottini R, Allevato E, Stazi SR, Scarascia MG, Harfouche A, Petruccioli M, Turner RJ, Zannoni D, Cappelletti M (2019) Identification of resistance genes and response to arsenic in Rhodococcus aetherivorans BCP1. Front Microbiol 10:888
Govarthanan M, Park SH, Park YJ, Myung H, Krishnamurthy RR, Lee SH, Lovanh N, Kamala-Kannan S, Oh BT (2015) Lead biotransformation potential of allochthonous Bacillus sp. SKK11 with sesame oil cake extract in mine soil. RSC Adv 5:54564
Ji G, Silver S (1992) Reduction of arsenate to arsenite by the ArsC protein of the arsenic resistance operon of Staphylococcus aureus plasmid pI258. Proc Natl Acad Sci 89 (20):9474–9478
Johnning A, Moore ER, Svensson-Stadler L, Shouche YS, Larsson DG, Kristiansson E (2013) Acquired genetic mechanisms of a multiresistant bacterium isolated from a treatment plant receiving wastewater from antibiotic production. Appl Environ Microbiol 79:7256–7263
Kang CH, Kwon YJ, So JS (2016) Bioremediation of heavy metals by using bacterial mixtures. Ecol Eng 89:64–69
Kruger MC, Bertin PN, Heipieper HJ, Arsène-Ploetze F (2013) Bacterial metabolism of environmental arsenic—mechanisms and biotechnological applications. Appl Microbiol Biotechnol 97:3827–3841
Li X, Hu Y, Gong J, Lin Y, Johnstone L, Rensing C, Wang G (2012) Genome sequence of the highly efficient arsenite-oxidizing bacterium Achromobacter arsenitoxydans SY8. J Bacteriol 194:1243–1244
Lu JJ, Perng CL, Lee SY, Wan CC (2000) Use of PCR with universal primers and restriction endonuclease digestions for detection and identification of common bacterial pathogens in cerebrospinal fluid. J Clin Microbiol 38:2076–2080
Mohamed EAH, Farag AG (2015) Arsenic removal from aqueous solutions by different Bacillus and Lysinibacillus Species. Bioremediat J 19(4):269–276. https://doi.org/10.1080/10889868.2014.995375
Mosa KA, Saadoun I, Kumar K, Helmy M, Dhankher OP (2016) Potential biotechnological strategies for the cleanup of heavy metals and metalloids. Front Plant Sci 7:303
Nguyen CC, Hugie CN, Kile ML, Navab-Daneshmand T (2019) Association between heavy metals and antibiotic-resistant human pathogens in environmental reservoirs: a review. Front Environ Sci Eng 13:46
Ojuederie OB, Babalola OO (2017) Microbial and plant-assisted bioremediation of heavy metal polluted environments: a review. Int J Environ Res Public Health 14:1–26
Padayachee T, Klugman KP (1999) Molecular basis of rifampin resistance in Streptococcus pneumoniae. J Antimicrob 43:2361–2365
Poszytek K, Karczewska-Golec J, Ciok A, Decewicz P, Dziurzynski M, Gorecki A, Jakusz G, Krucon T, Lomza P, Romaniuk K, Styczynski M, Yang Z, Drewniak L, Dziewit L (2018) Genome-guided characterization of Ochrobactrum sp. POC9 enhancing sewage sludge utilization-biotechnological potential and biosafety considerations. Int J Environ Res Public Health 15:1501
Rosen BP (1999) Families of arsenic transporters. Trends Microbiol 7(5):207–212
Rosen BP (2002) Biochemistry of arsenic detoxification. FEBS Lett 529(1):86–92
Saltikov CW, Olson BH (2002) Homology of Escherichia coli R773 arsA, arsB, and arsC genes in arsenic-resistant bacteria isolated from raw sewage and arsenic-enriched creek waters. Appl Environ Microbiol 68:280–288
Samanta A, Paramita B, Mahamuda K, Chandrima S, Pinaki P, Asif L, Anurup M (2012) An investigation on heavy metal tolerance and antibiotic resistance properties of bacterial strain Bacillus sp. isolated from municipal waste. J Microbiol Biotech Res 2:178–189
Sanyal SK, Mou TJ, Chakrabarty RP, Hoque S, Hossain MA, Sultana M (2016) Diversity of arsenite oxidase gene and arsenotrophic bacteria in arsenic affected Bangladesh soils. AMB Express 6:1–11
Sarkar A, Kazy SK, Sar P (2014) Studies on arsenic transforming groundwater bacteria and their role in arsenic release from subsurface sediment. Environ Sci Pollut Res 21:8645–8662
Selvankumar T, Radhika R, Mythili R, Arunprakash S, Srinivasan P, Govarthanan M, Kim H (2017) Isolation, identification and characterization of arsenic transforming exogenous endophytic Citrobacter sp. RPT from roots of Pterisvittata. 3 Biotech 7(4):264
Selvi MS, Sasikumar S, Gomathi S, Rajkumar P, Sasikumar P, Sadasivam SG (2014) Isolation and characterization of arsenic resistant bacteria from agricultural soil, and their potential for arsenic bioremediation. Int J Agric Pol Res 2(11):393–405
Shen Z, Han J, Wang Y, Sahin O, Zhang Q (2013) The contribution of ArsB to arsenic resistance in Campylobacter jejuni. PLoS One 8(3):e58894
Simeonova DD, Lièvremont D, Lagarde F, Muller DA, Groudeva VI, Lett MC (2004) Microplate screening assay for the detection of arsenite-oxidizing and arsenate-reducing bacteria. FEMS Microbiol Lett 237(2):249–253
Sunitha MS, Prashanth SR, Kishor PB (2015) Characterization of arsenic-resistant bacteria and their ars genotype for metal bioremediation. Int J Sci Eng Res 6:304–309
Tisa LS, Rosen BP (1990) Transport systems encoded by bacterial plasmids. J Bioenerg Biomembr 22(4):493–507
Wolfe-Simon F, Switzer Blum J, Kulp TR et al (2011) A bacterium that can grow by using arsenic instead of phosphorus. Science 332:1163–1166
Wu J, Rosen BP (1993) Metalloregulated expression of the ars operon. J Biol Chem 268(1):52–58
Yang HC, Fu HL, Lin YF, Rosen BP (2012) Pathways of arsenic uptake and efflux. Curr Top Membr 69:325–358
Yoon KP (2005) Stabilities of artificially transconjugated plasmids for the bioremediation of cocontaminated sites. J Microbiol 43:196–203
Acknowledgments
Author RS wants to acknowledge his university, i.e., Indira Gandhi Krishi Vishwavidyalaya, Raipur, India, for the financial assistance. We would also like to acknowledge CDRI, Lucknow, and Department of Soil Science, Indira Gandhi Krishi Vishwavidyalaya, Raipur, for providing SEM and AAS facilities, respectively. We are also showing our gratitude towards Probecell: Scientific Writing Services for proofreading and edit.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Dash, B., Sahu, N., Singh, A.K. et al. Arsenic efflux in Enterobacter cloacae RSN3 isolated from arsenic-rich soil. Folia Microbiol 66, 189–196 (2021). https://doi.org/10.1007/s12223-020-00832-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-020-00832-2