Abstract
Pseudomonas aeruginosa, a Gram-negative, rod-shaped bacterium causes widespread diseases in humans. This bacterium is frequently related to nosocomial infections such as pneumonia, urinary tract infections (UTIs) and bacteriaemia especially in immunocompromised patients. The current review focuses on the recent perspectives on biofilms formation by these bacteria. Biofilms are communities of microorganisms in which cells stick to each other and often adhere to a surface. These adherent cells are usually embedded within a self-produced matrix of extracellular polymeric substance (EPS). Pel, psl and alg operons present in P. aeruginosa are responsible for the biosynthesis of extracellular polysaccharide which plays an important role in cell surface interactions during biofilm formation. Recent studies suggested that cAMP signalling pathway, quorum-sensing pathway, Gac/Rsm pathway and c-di-GMP signalling pathway are the main mechanism that leads to the biofilm formation. Understanding the bacterial virulence depends on a number of cell-associated and extracellular factors and is very essential for the development of potential drug targets. Thus, the review focuses on the major genes involved in the biofilm formation, the state of art update on the biofilm treatment and the dispersal approaches such as targeting adhesion and maturation, targeting virulence factors and other strategies such as small molecule-based inhibitors, phytochemicals, bacteriophage therapy, photodynamic therapy, antimicrobial peptides and natural therapies and vaccines to curtail the biofilm formation by P. aeruginosa.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pseudomonas aeruginosa is a Gram-negative, aerobic rod-shaped bacterium which belongs to the bacterial family Pseudomonadaceae, a member of γ-proteobacteria. This family is classified into eight groups. P. aeruginosa is one of the subtypes among 12 member groups (Todar 2008). P. aeruginosa is a free-living organism found ubiquitously in diverse environmental settings. It has earned the name of an opportunistic pathogen that forms biofilm and is responsible for 10–20% of infections in hospitals (Fazeli et al. 2012). It is especially prevalent among patients who are already immunocompromised. P. aeruginosa have been identified as nosocomial contaminants and epidemics have been identified in the hospital environment (Bodey et al. 1983).
The ability of these bacteria to survive with basic minimal nutrient requirements and their tolerance to numerous physical conditions has allowed persisting in both urban and natural settings. In the natural environment and during infection of hosts, many bacteria including P. aeruginosa grow as populations or groups entrenched in a matrix, the extracellular polymeric substances (EPS). These groups of bacteria encased in a matrix are known as bacterial biofilms. These bacteria in the biofilm display different features from planktons, such as high resistance or opposition to the immune system, therapeutics and physiology (Wei and Ma 2013). Due to this, the formation of biofilms possesses a major health concern in chronic infections (Laverty et al. 2014). P. aeruginosa is very strenuous to suppress immunity when forming dense antibiotic-resistant biofilms, lowering the proficiency of immune systems in patients as organisms that exist in group can endure some period of time (Laverty et al. 2014).
The biofilms provide basic mechanism of resistance to not only antibiotics, but also bacteriophages, disinfectants and other host defense systems (Dheilly et al. 2010) by constituting multi-layered protection mechanisms (Stewart 2002). Furthermore, the microbes in biofilms develop flexible stress responses resulting in higher number of premature deaths and increased morbidity rates. This makes P. aeruginosa a significant model organism for learning about the development of bacterial biofilm and resistance to various antibacterial agents (Habash et al. 2014).
According to the recent reports by the World Health Organization, P. aeruginosa is considered as one of the priority pathogens which became extremely drug resistant to most of the current generation antibacterial and create high threat and mortality rate to public health. Hence, it is very essential to study the recent advances in the biofilm formation by the bacteria which contribute the virulence and antibiotic resistance. Although there are many previous reviews which focused on the mechanism of biofilm formation by P. aeruginosa, the detailed molecular basis of biofilm formation by P. aeruginosa, various approaches used for the treatment and biofilm dispersal have profound scope in applied microbiology. Thus, the current review tried to provide cutting-edge knowledge and up-to-date perspectives on the molecular mechanism of biofilm formation by P. aeruginosa. The review focuses on the role of biofilm in bacterial pathogenesis and the mechanism of antibiotic resistance. The review further discusses the biochemical mechanism of biofilm formation, especially major aspects of attachment, biofilm maturation especially the role of bis-(3′-5′)-cyclic-dimeric-guanosine monophosphate (c-di-GMP)-dependent polysaccharides, two-component regulatory systems and quorum-sensing and quorum-quenching systems. The review finally illuminates the recent perspectives and approaches used for the treatment and dispersal of biofilm formation by P. aeruginosa.
Epidemiology
P. aeruginosa is a predominant pathogen, triggering various infections, mainly in patients who are unable to develop a normal immune response. An enduring complication regarding the treatment of infections caused by these bacteria is to form biofilms on indwelling and entrenched devices (Costerton et al. 2003). Most of the infections are connected with urinary tract infections, which are the foremost nosocomial infections, followed by cathetherisation and intubation infections (Nicolle 2014; Xu et al. 2015). The infections are commonly observed in chronic obstructive pulmonary disorder (COPD) and cystic fibrosis (CF) patients (Mulcahy et al. 2014).
Biofilm-related infections where the significance of biofilm is highlighted include infections on medical devices such as cardiac devices, prosthetic joints, intravascular catheters, shunts and prosthetic vascular grafts. In patients with CF, the widespread of pseudomonal pneumonia encompasses from 21 to 80% in the age groups between 1 and 19 years. Participation of the gastrointestinal tract usually is common in patients with hematologic malignancies; neutropenia arisen from chemotherapy is also observed in infants (Friedrich 2015). It is also one of the customary causes of health care-associated pneumonia (HCAP), ventilator-associated pneumonia (VAP) and hospital-acquired pneumonia (HAP) and overreached in prevalence only by Staphylococcus aureus (Driscoll et al. 2007).
P. aeruginosa causes 17% of nosocomial pneumonia, 7% urinary tract infection, 8% quotidian cause of surgical-site infection and 9% infection in general from all sites (Friedrich 2015). It has a rate of 36 infections per 10,000 releases. The epidemiology is related to their capability to survive in tiny ecological niches and emerge as a nosocomial pathogen with the emergence of antimicrobial resistance (Morrison and Wenzel 1984).
Most of these infections have hostile consequences in patients who undergo supplementary surgeries and lengthy subjections to universal treatments which are uncertain to resolve the infection. The most common worry is in periprosthetic infection, where the pathogenic bacteria can gain entrance by two major routes with related time scales (Green 2011). The first is the perioperative period, usually via the surgical cut itself. The second is from hematogenous spread, which normally arises during the postoperative period. Infections can be existent in the patients acutely and/or can persist chronically, and the analysis of infection and the type of biofilm growth remains a hard task for clinicians often necessitating highly dedicated research equipment and time-consuming techniques (McConoughey et al. 2014).
Biochemistry of biofilm
Confocal scanning laser microscopy revealed the 3-D structure of the biofilm to be composed of microcolonies (clusters of cells) of different species of microbial cells (around15% by volume) and of matrix material (85%) (Kokare et al. 2009). The spatial arrangement of these microcolonies plays significant role in the determination of the biofilm complex and has a profound implication on their function. Extracellular polymeric substance (EPS) or exopolysaccharide is a sticky organic matter that forms an integral part of the structural organization of biofilm (Davey and O’Toole 2000). It comprises primarily of polysaccharides and an undefined proportion of protein and nucleic acids such as extracellular DNA. P. aeruginosa produces three major types of polysaccharides: pel, psl and alginate, each differing in their chemical structures (Schurr 2013). Alginate was the first P. aeruginosa exopolysaccharide to be discovered. It was identified in mucoid strains of P. aeruginosa recovered from CF patients (Doggett 1969). The various components of EPS interact in a coordinated manner, which is vital for the effective colonisation of the bacteria which form biofilm (Flemming et al. 2016).
Based on the composition of the EPS matrix, biofilm may assume a number of differentiated forms during development such as mushroom-like microcolonies and filamentous streamers (Chew et al. 2014). The vertical structures of biofilm are separated by interstitial spaces. The biofilm acquire nutrients easily and rapidly from the surrounding medium and move the toxic by-products away by the interstitial spaces (Secinti et al. 2011). The shape and the mechanical stability of the biofilm are determined by the physical properties of the matrix (Chew et al. 2014).
In order to form and maintain biofilm communities, pseudomonads have an inherent capacity to generate a manifold of biofilm-linked macro- and micro-molecules that constitute the matrix. There are five distinct stages involved in the exopolysaccharide biosynthesis. First, the precursor substrate, a nucleotide-activated sugar, is manufactured in the cytoplasm. The ancestor substrate is then polymerised onto the growing polysaccharide (Varki et al. 2009). Next, the polysaccharide is shipped across the internal membrane to the periplasm and undergoes enzymatic modification. It is consequently exported through the external membrane (Colvin et al. 2013).
Pel and psl polysaccharide
Pel, a cationic exopolysaccharide, was identified while screening mutant libraries of the strain PA14 for the lack of pellicles (Franklin et al. 2011). The products of the pel gene operon (PA3058-PA3064) control its synthesis (Friedman and Kolter 2004a, b). The establishment of solid surface-associated biofilms is greatly dependent on the presence of pel (Wei and Ma 2013). Currently, the structure of pel has not been resolved, but it is suggested to be a glucose-abundant polysaccharide (Friedman and Kolter 2004a, b).
Strains of P. aeruginosa that produces insufficient amount of alginate such as nonmucoid strains use pel and/or psl as the primary structural scaffold (Jennings et al. 2015). Pel also enhances the specific resistance of biofilms against antibiotics such as aminoglycosides and play protective role in P. aeruginosa biofilms (Colvin et al. 2013).
Reverse genetics was employed to identify psl in P. aeruginosa PAO1. The psl gene cluster consists of 15 genes: pslA to pslO (PA2231-2245 gene cluster) out of which 11 are necessary for psl synthesis (Matsukawa and Greenberg 2004; Byrd et al. 2009).
Psl is composed of L-rhamnose, D-glucose and D-mannose repeats. The pentasaccharide is expressed in two forms: a cell-related, high molecular weight form and a small, soluble form (Byrd et al. 2009; Mann and Wozniak 2012). It distributes helically around the cell surface and maintains the biofilm organization by facilitating interactions between cells and the cell surface (Ma et al. 2009). The presence of psl enhances the cross-linking and elasticity of the matrix. This reinforces the scaffold and promotes the establishment of microcolonies. Psl is exceedingly important during the primary attachment of sessile cells to biotic and abiotic substrates (Byrd et al. 2010). It was found that in the absence of psl, the viscosity of the matrix increases, which in turn promotes the biofilm diffusion (Chew et al. 2014). Although psl is intrinsically expressed in planktonic cells, the expression is confined to the centre of growing biofilm implying that psl has a part in biofilm differentiation (Overhage et al. 2005). Mature biofilm on the other hand reveals the peripheral presence of psl (Ryder et al. 2007).
Alginate
Alginate is a high molecular weight linear anionic polysaccharide, comprised of β-1-4 glycosidic linked α-L-guluronic acid and β-D-mannuronic acid (Hay et al. 2009). It guards the bacteria from environmental adversities and enhances surface adhesion. Alginate biosynthetic genes are transcribed upon attachment to surfaces, resulting in the development of biofilms. A 12-gene cluster encodes the regulatory and biosynthetic machineries for alginate (PA3540–PA3551). It is initially assembled as D-mannuronic acid homopolymer, which is transformed in the periplasm by selective O-acetyl substitution (Franklin et al. 2011; Wiens et al. 2014).
Within the biofilm, various hydrolytic enzymes and alginate are secreted at different quantities by various strains of P. aeruginosa (Tielen et al. 2013). They shield the pathogen from antibiotics and the host’s immune attack contributing to drug resistance. Initially, the CF lungs are infected with nonmucoid strains, but as it persists, mucoid forms begin to rise and turn into the dominant lung pathogen. The conversion from nonmucoid to mucoid phenotype is due to the production of alginate (Hay et al. 2009). It initiates the mobilisation of inflammatory cells to the locus of infection, which liberates reactive oxygen species causing extensive tissue damage. Alginate scavenges the free radicals liberated by macrophages and protects P. aeruginosa from phagocytic clearance (Ryder et al. 2007). Further, biofilm produced by alginate overproduction present well-ordered structures, providing higher resistance to antibiotics such as tobramycin (Hentzer et al. 2001). It also facilitates coexistence with S. aureus as observed during CF infections. Alginate overproduction reduces the synthesis of key compounds such as siderophores and rhamnolipids required to kill S. aureus and affect the antimicrobial potential (Limoli et al. 2017).
Under certain conditions, P. aeruginosa expresses an alginate lyase (algL) that divides the alginate polymer into short oligosaccharides eliminating the anchoring ability, resulting in the separation of bacteria from the surface. This enhances biofilm dispersal and allows the microorganisms to colonise new sites (Boyd and Chakrabarty 1995). Studies have demonstrated that alginate lyase shows potential for treating mucoid P. aeruginosa infections as they have shown to lower culture viscosity in clinical isolates and in CF sputum, striping biofilms from abiotic surfaces, enhancing phagocytosis and the destruction of P. aeruginosa by human immune system and improving the effectiveness of anti-pseudomonal antibiotics (Lamppaa and Griswold 2013).
eDNA
The importance of eDNA as a structural component of biofilm was first demonstrated in P. aeruginosa. eDNA fortifies the biofilm, functions as a source of nutrient during period of starvation, provides resistance to the antibiotics and helps in biofilm expansion (Wang et al. 2015). It further helps to connect microcolonies in the biofilm (Wei and Ma 2013). Several hypotheses have been proposed for the generation of eDNA such as the release of small membrane vesicles, lysis of subpopulation and direct secretion (Wei and Ma 2013). A biofilm development regulator, BfmR, has been found to correspond with DNA liberation and localised cell death. It is expressed during situations that result in the membrane disruption. ΔBfmR mutant biofilm displayed enhanced eDNA liberation and cell lysis. This indicates that BfmR does not eliminate, but suppresses, the processes (Petrova et al. 2011). eDNA release is also regulated by the expression of phenazine molecules such as pyocyanin. Mutants of P. aeruginosa with defective phenazine synthesis produce comparatively less eDNA than phenazine producing strains (Das and Manefield 2012, 2013). The interaction of pyocyanin with eDNA modulates cell surface hydrophobicity which in turn influences the bacterial surface energy components and nonspecific interactions. eDNA removal reduced the biofilm thickness by ∼ 40% and phenazine deficiency reduced the biofilm thickness and biomass by ∼ 40 and ∼ 80%, respectively (Das et al. 2016).
In P. aeruginosa, eDNA is commonly released through autolysis and may be produced by two separate pathways. A pathway not coupled to quorum sensing is responsible for a base-level production of eDNA and the late log phase associated with abundant eDNA release is generated by quorum-sensing-dependent mechanisms (Montanaro et al. 2011).
Two major elements of the P. aeruginosa biofilm, psl and eDNA, interact to build the mesh of psl-eDNA fibres. It provides a scaffold that permits bacterial adherence and growth (Wang et al. 2015). The negatively charged eDNA binds to Ca2+ which stabilises and promotes cross-bridging among bacterial cells, leading to the biofilm formation (Das et al. 2014). Similarly, eDNA binds to other positively charged antimicrobials such as aminoglycosides and antimicrobial peptides which were demonstrated in a study by Chiang et al. (2013). eDNA is also responsible for the decrease in pH and acidification of biofilm. This induces the PhoPQ and PmrAB two-component systems regulating virulence and aminoglycosides resistance (Wilton et al. 2015).
eDNA has been found to play a role in the regulation of type VI secretion system (T6SS) found in Gram-negative bacteria to translocate proteins and virulence factors. Wilton et al. (2016) suggested that eDNA stimulates H1-T6SS through chelation of outer membrane bound cations. This enables P. aeruginosa to attack other species in the vicinity in an indiscriminate manner. Due to the widespread availability of eDNA in P. aeruginosa inhabitants, they gain a competitive edge in poly-microbial communities (Wilton et al. 2016).
Matrix proteins
Cell elements or adhesins, chiefly flagella and type IV pili (cell surface appendages), assist in biofilm formation (Cai et al. 2015). Different adhesins bind to glycosylated receptor molecules located in the apical and basal region on the epithelial surface of the cells. Type IV pili interact with apical region, whereas flagella associate with the basal region surface proteoglycan heparin sulphate chains (Bucior et al. 2012). Type IV pili (T4P) are protein fibers produced on the bacterial cell surface. Pilins are small structural proteins with a protein interaction domain and a transmembrane domain (Giltner et al. 2012). They are built from pilA, a protein subunit that is transported outside the cell by pilQ, a secretin, to produce a fimbrial strand (Laverty et al. 2014). Type IV pili are involved in various processes such as colonisation during infection, twitching motility (bacterial translocation), biofilm formation, bacteriophage infection, DNA uptake and natural transformation. A specialised feature of pilin is the ability to reversibly produce polymeric fibers and aggregate to form ordered bundles (Hazes et al. 2000).
The pilus assembly system comprises of inner and outer membrane complexes and connecting subcomplexes, composed of pilMNOP. A study by McCallum et al. (2016) showed that when pilM binds to pilN at the N-terminus, there are significant structural changes in pilM causing monomerization (McCallum et al. 2016).
Flagella are required for cell attachment, swimming locomotion and biofilm development. The flagellum is primarily composed of flagellin which can be grouped into a and b serotypes (Campodónico et al. 2010). It is the globular protein flagellin that is identified by the innate immune system (Bucior et al. 2012). The flagellar genes such as fleQ, fleR, fleS, fliA, flgM and fleN encode proteins that participate in the regulation of the flagellar transcriptional circuit and are grouped into three distinct regions in the chromosome (Dasgupta et al. 2003). Many animal studies have demonstrated flagella as an important virulence factor in P. aeruginosa and have certified as target antigens for vaccination (Campodónico et al. 2010). Bivalent type a and b flagellar vaccine have shown promising results in terms of preventing P. aeruginosa infection in CF patients (Doring et al. 2007).
Although the gene encoding for chemotaxes in P. aeruginosa are homologous with other model organisms displaying chemotaxes such as Escherichia coli, it was shown that chemotaxes occurs by consistently extending and reducing the periods of both clockwise and counterclockwise flagellar rotations when swimming up and down the chemoattractant gradient, respectively (Cai et al. 2016).
Biofilm and bacterial pathogenesis
P. aeruginosa infections are complex in the pathogenesis and are dependent on various virulence factors such as secretory factors, elastase, phospholipase C, alkaline protease, pyocyanin, hydrogen cyanide, pyoverdine and rhamnolipids (van ‘t Wout et al. 2015). The lipopolysaccharide, pili, flagella and cell associated factors that allow the bacteria to form biofilm when attached to biotic or abiotic surfaces (Pier and Ramphal 2005). The effects of P. aeruginosa infections are due to bacterial virulence factors. The main factors are proteases and exotoxins that cause severe host tissue damages by distorting cytoskeletal structures, cleaving the immunoglobulin A and G and depolymerising the actin filaments. The exoenzymes of P. aeruginosa are responsible for the host tissue damages, thus facilitating dissemination, invasion and development of chronic and acute diseases (Sadikot et al. 2005). The virulence factors such as pili, LecA and LecB help the bacteria to attach to their host; siderophores allow the bacterial multiplication in limited iron environments and alginate gives protection from certain immune attacks by breaking down oxygen species, by hindering complement factors and also by limiting polymorphonuclear chemotaxis (Chatterjee et al. 2016).
Additionally, the biological function of oxylipins have shown their roles in bacterial biofilm pathogenesis and are studied in plants, animals and algae but are largely unidentified in prokaryotes. P. aeruginosa shows a diol synthase activity which transforms various mono-unsaturated fatty acids into mono- and di-hydroxylated derivatives. The oxylipins derived from this activity restrict flagellum-driven motility and increase type IV twitching motility of P. aeruginosa. These oxylipins increase bacterial organization and thereby increase the ability of P. aeruginosa to form biofilms (Martinez and Gomez 2016).
Pseudomonas infections can be involved in the bloodstream (bacteremia), GI tract (diarrhoea, enteritis), enterocolitis respiratory tract (pneumonia), CNS (meningitis, brain abscess), heart (endocarditis), eye (bacterial keratitis, endophthalmitis), ear (otitis externa and media), bones and joints (osteomyelitis) and urinary tract and skin (ecthymagangrenosum) (Friedrich 2015). An arrangement of major virulence factors of P. aeruginosa discussed earlier offset the host defenses and forms the basis of host tissue damages or increases the competitiveness of the bacteria (Gellatly and Hancock 2013).
An insight on P. aeruginosa infections with respect to CF patients is one of most perused researches. CF is a disorder resulting from the mutations in the CFTR gene that code for an ion-transport protein. Defects in this gene result in various physiological defects most importantly, the accumulation of alveolar fluid and mucus in lungs in human. This leads to severe colonisation by P. aeruginosa which eventually leads to fatal infections and pneumonia (Rajan and Saiman 2002; Lovewell et al. 2014).
Mechanism of antibiotic resistance and tolerance of biofilm
The bacteria in biofilm display distinctive abilities to survive to antibiotic treatments that are accountable directly for a substantial number of complications in the clinical atmosphere (Lebeaux et al. 2014). Resistance to the antibiotics in bacterial biofilms is due to an amalgamation of various factors which act organised to give rise to a resistance level. These distinct factors include persister (cells) formation, initiation of lipid-modified operons by eDNA, slow release of antibiotics in the biofilms, mutations in chromosomes and gaining of genes responsible for resistance (Poole 2011). Acidic pH and acidification from eDNA produces a signal that is recognised by P. aeruginosa to enhance the expression of genes regulated by the PhoPQ and PmrAB two-component systems (Zhang et al. 2013). P. aeruginosa cultured under acidic conditions give around eightfold increases in aminoglycoside resistance. This resistance requires the production of spermidine and also aminoarabinose modification of lipid A (Wilton et al. 2016). Changes in mutations affecting resistance to treatments include an increase in the rate of multidrug efflux mechanisms, an enhancement of antimicrobial expulsion, a modification to antimicrobial targets and a depression and alteration of AmpC leading to a broad-spectrum β-lactamase which increases the enzyme specificity to the substrates (Poole et al. 2011).
Polysaccharides are the major constituents in the matrix of the biofilms, but their association with the resistance characteristics of biofilm is largely unknown. Certain continuous flow and static biofilm experiments have shown that psl imparts a nonspecific first line of defense towards treatments with antibiotics, with various possessions during the inceptive stages of development of biofilm (Billings et al. 2013). AlgC promotes to the synthesis of psl, promoting resistance to polysorbate-80 (PS80) (biofilm inhibitor), a non-ionic detergent and surfactant that prevent biofilm formation at lower strengths and is tolerated well by human tissues (Zegans et al. 2012). While multiple factors contribute to tolerance of biofilm towards antimicrobial, only sparse information about the time at which induction of biofilm tolerance occurs is known. The tolerance of P. aeruginosa biofilm is coupled to the ‘two component system SagS’ which controls the formation of biofilm with conversion to an irreversible stage of attachment. It also reveals the time when biofilm shift to high levels of tolerance. Inactivation of this hybrid system resulted in biofilms but not the planktonic cells sensitive to norfloxacin, tobramycin and hydrogen peroxide (Gupta et al. 2013). Blr equivalently gave rise to tobramycin and colistin resistance in PAO1 strain of P. aeruginosa. Clinical patients with an increase in expression of blr gave rise to an increased tobramycin resistance and displayed minimum inhibitory concentration. However, the minimum inhibitory concentration of colistin deteriorated and increased the susceptibility of colistin (Chambers and Sauer 2013). The major resistance mechanism exhibited by P. aeruginosa is shown in Fig. 1.
Mechanism of biofilm formation
Biofilm formation depends on the bacterial ability to produce the extracellular matrix components that enable them adhere to each other (Bucior et al. 2012). During the course of biofilm maturation, the inhabited bacteria get embedded and secured in the matrix. In Gram-negative bacteria, biofilm formation is regulated by three main processes: attachment, maturation and dispersion (Laverty et al. 2014). The major events in the biofilm formation are shown in Fig. 2.
Schematic showing the various stages of biofilm development by P. aeruginosa which characterised by initiation, maturation and dispersal. The SEM images show the biofilm formation initiation and biofilm maturation. The images have reproduced with permission (Rendueles and Ghigo 2012)
Attachment/reversible adhesion
Gram-negative bacteria can attach to surfaces either through flagella, pili or fimbriae (Laverty et al. 2014). This process is controlled by a number of factors that include environmental aspects, bacterial species, surface composition and essential gene products (Dunne 2002). The cells can be transported by the physical forces of the media or by flagella and other appendages. Some of the cells reversibly adsorb onto the surface upon contact. Physical forces include electrostatic interactions, steric interactions and the van der Waals forces, known as the DVLO (Derjaguin, Verwey, Landau and Overbeek) forces (Garrett et al. 2008).
The following stages are involved in bacterial adhesion: first, a reversible docking phase (primary stage) involving the initial interaction between the microorganism and a surface that has been modified or conditioned. Conditioning enhances the surface properties for bacterial adhesion (Garrett et al. 2008). This phase is influenced by physiochemical variables involved in the interaction. Next, the locking phase (secondary stage) which involves the binding between specific adhesins and the surface brought about by molecular interactions. This is followed by biofilm development and maturation (Dunne 2002).
Type IV pili of P. aeruginosa facilitates the initial adhesion through a combination with two forms A and B of O-polysaccharide chain (Makin and Beveridge 1996). P. aeruginosa is capable of altering the lipopolysaccharide composition for adherence, therefore supporting the survival and formation of biofilm on various surfaces. Lipopolysaccharide-A production is aided to increase hydrophobicity of the cell surface enabling the adherence to hydrophobic surfaces, while lipopolysaccharide-B induces hydrophilicity which enables attachment to hydrophilic surfaces such as glass (Makin and Beveridge 1996).
The adhesin genes such as cdrAB were first observed in P. aeruginosa as a bi-cistronic operon that was induced under influence of high c-di-GMP. These genes belonged to a two-component secretion system known as type Vb secretion system. CdrB gene forms barrel structures that form pores in the outer membrane functioning as the channel through which adhesin cdrA is released. CdrA harbors two cysteine residues in the C-terminal region that have been concerned in other TPSS adhesins in joining to the outer membrane. The interaction of adhesin cdrA with the matrix exopolysaccharide psl is specific. This interaction is the probable reason for stabilising matrix integrity, with mutant strains of cdrA exhibiting aggregates with freely connected psl and depicting biofilms with compromised integrity. It is suggested that cdrA was approximately 220-kDa protein that was originated to be cell-linked and also as a processed form (150 kDa) in the bacterial cultures overexpressing it. This system was induced at high c-di-GMP content with the concurrent P. aeruginosa-induced biofilm formation through the concurrent expression of matrix-binding adhesions and matrix polysaccharides (Parsek 2015).
Studies suggested that the mechanism of biofilm formation is varied in various strains such as PAO1 and PA14 based on the polysaccharide components (Colvin et al. 2011, 2012). The PAO1 and PA14 are that two major laboratory strains that vary in the polysaccharide used as major structural component for the biofilm development. PAO1 mainly uses psl and PA14 uses pel (Colvin et al. 2011). Colvin et al. (2012) validated this observation using two distinct biofilm culturing formats. A microtiter dish assay was measured biomass in the biofilm that forms at the air-liquid interface after incubation and confocal laser scanning microscopy (CLSM) was used to monitor live biofilms growing in the medium using a flow-cell reactor. PAO1 and PA14 and their analogous pel, psl and pel psl mutant strains were compared to study the attachment and formation of biofilm in the microtiter dish assay. The study suggested that PA14 pel, PAO1 pel and PAO1 psl mutants all lack surface attachment (Colvin et al. 2012). However, pel demonstrated significant impact on later stages of biofilm formation. A pel mutant of PA14 developed significant reduction in biofilm biomass in comparison with PA14, while a PAO1 pel biofilm was identical to PAO1. Further, the PAO1 psl and PAO1 pel psl double mutants were impaired for biofilm development.
Glycosylation of mucin in the patients have been implicated in the adhesion of P. aeruginosa (Venkatakrishnan et al. 2013). P. aeruginosa also adheres to cell surface-associated mucin, mucin-1 (MUC1), and other mucin carbohydrates via adhesins that are distinct from pilin. Several classes of adhesins displayed on the bacterial surface of P. aeruginosa are involved in the colonisation and infection of the human respiratory tract (Ramphal et al. 1991). During infection, P. aeruginosa were associated with ciliated cells trapped at the extremities of cilia (Crabbé et al. 2014). The epithelial layer along with mucus-secreting cells layering the luminal surface forms the mucosal barrier (Enge and Eran 2011). Fracturing of the epithelial barrier by disruption or injury of narrow junctions renders the basolateral surface accessible to P. aeruginosa. Basement membrane proteins of the extracellular matrix such as fibronectin, collagens and laminin facilitate P. aeruginosa adhesion and can influence dissemination and colonisation (Crabbé et al. 2014).
A polyamine, norspermidine, has been reported to disrupt the process of biofilm formation and sometimes prevent it in the case of Gram-negative bacteria. The inhibitory effects of norspermidine is due to its ability to prevent the attachment of cells to surfaces, downregulating quorum-sensing (QS) genes and inhibiting swimming motility by reducing the available eDNA and exopolysaccharides (Qu et al. 2016; Si et al. 2016). A study conducted by Qu et al. (2016) showed that the degree of cell-surface attachment was found to vary proportionally with the concentration of norspermidine. Exposure to norspermidine at concentrations of 4 mmol/L decreased bacterial attachment to 71.22% (Qu et al. 2016). Norspermidine was encapsulated into polyelectrolyte multilayer coatings to provide a surface-mediated releasing approach. This arrangement was found to have a high inhibitory effect on the biofilm attachment and formation against pure strains of P. aeruginosa and on mixed cultures (Si et al. 2016).
Biofilm maturation
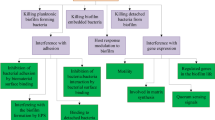
The differences of microcolonies into true biofilms are regulated by a variety of bacterial machinery, such as the quorum-sensing (QS) systems, the RetS/LadS and GacS/GacA two-component regulatory systems and the c-di-GMP-mediated polysaccharide regulation (Rasamiravaka et al. 2015b). The role of selected examples of the regulatory cascades in the biofilm formation is shown in Fig. 3.
The molecular events governed in the formation of biofilm by P. aeruginosa. The figure shows only a subset of the regulatory connections. The surface colonisation, biofilm formation and development provide various advantages to microorganisms. The main factors relevant in the biofilm formation by P. aeruginosa include cAMP/Vfr signalling, quorum-sensing (QS) systems, Gac/Rsm pathway and c-di-GMP signalling. cAMP/ Vfr signalling regulates the transcription of various gene encoding virulence factors, including the type 2 and type 3 secretion systems and their associated toxins, type IV pili and flagella. It is suggested that the increase of cAMP inhibits the attachment phase of biofilm development. The QS systems are set hierarchically with the las system positively regulating the rhl and PQS systems. These three QS systems are responsible for virulence factor production, biofilm maturation and motility phenotypes. GacS/GacA (two-component system) facilitates the expression of RsmY and RsmZ, which sequester RsmA (translational repressor). RsmA induces the production of sessile and biofilm determinants. c-di-GMP is a ubiquitous bacterial second messenger; the formation is assisted by diguanylate cyclases (DGC) and phosphodiesterases (PDE). Increased concentrations of c-di-GMP induce the formation of biofilm matrix, while decreased concentration of c-di-GMP promotes bacterial motility and shift into the planktonic growth. This figure revealed that WspB/D and methyltransferase WspC also play an important role; rhlR is regulated by LasR, which is an important regulatory pathway in P. aeruginosa
Quorum-sensing systems
Formation of biofilm is a dynamic process, and for many microorganisms, it is dependent on small chemical signal molecules called autoinducers that are self-produced. This process is termed as quorum sensing (QS). Once the bacterial colony attains quorum level, the autoinducers couple with their corresponding transcription regulators and repress or activate the target genes. Thus, quorum sensing displays a coordinated response (Annous et al. 2009). This also facilitates same-species and interspecies communication (Miller and Bassler 2001). In many bacteria, quorum sensing depicts the fundamental mechanism to regulate social activities, allowing bacteria to gain benefits that would otherwise be unattainable to them as individual cells (Li and Tian 2012a, b).
QS regulates the social behavior of bacteria by various interconnected signalling pathways (LaSarre and Federle 2013). It permits bacterial communities to control various biological processes which are essential for bacterial adaptation and endurance. This event relies on regulating the expression of specific genes in response to a critical threshold of signalling molecules (autoinducers). QS mediates the population density-dependent responses and is therefore favorable for community adaptation and survival. QS plays vital role for survival and colonisation by organising phenotypic changes at initial stages of infection (González and Keshavan 2006). The acute to chronic progression of infection is significantly enhanced by QS-dependent gene expression. QS can regulate more than 10% of genes in P. aeruginosa. These genes are primarily implicated in the production of virulence factors, biofilm formation, antibiotic resistance mortality and the amendment of metabolic pathways for stress responses (Moradali et al. 2017).
The relevance of QS in biofilm formation and maturation
The biofilm inhabitants exploit QS systems for spatio-temporal regulation of specific gene expression and cell-to-cell communication. Majority of the colonising population was expected to lose QS due to hypermutation and other phenotypic alterations during chronic infection. Studies have suggested that in P. aeruginosa, the genes involved in the progress of biofilm maturation and persistence, are positively regulated by QS. The QS-deficient mutants of P. aeruginosa (ΔlasIΔrhlI and ΔlasRΔrhlR) formed thin and less developed biofilms which were sensitive to antibiotic treatment and suppression (Nelson et al. 2009). Further, the studies suggested that at a part of QS pathways such as rhl encoded system and the formation of C4-HSL signals was preserved in mucoid population latterly of chronic stages match with overproduction of biosurfactants (rhamnolipids) and alginates (Bjarnsholt et al. 2010). It has been expected that rhamnolipids plays an important role in safeguarding of the architecture of biofilm by contributing to the development of interior cavities within the mature biofilm, permitting suitable flow of nutrients and water (Chrzanowski et al. 2012). In addition, QS-mediated production of pyocyanin is a vital component for biofilm maturation. Pyocyanin can promote the release of eDNA by inducing the cell lysis of the bacteria. Pyocyanin interact with eDNA enhancing its solution viscosity which accelerate the physicochemical interactions of the biofilm matrix and facilitates the aggregation of the cells (Jennings et al. 2015; Das et al. 2015). Such kind of cellular and molecular interactions along with additional polymeric substances resulted in the establishment of vigorous and mature biofilm (Moradali et al. 2017).
Studies have revealed that complicated QS network hierarchy is present in P. aeruginosa, which include groups of connected systems, such as las, pqs, iqs and rhl. Among these, las and rhl are the most important QS systems (Lee and Zhang 2015) (Fig. 3).
las and rhl quorum-sensing systems
The QS systems based on las and rhl are associated with N-Acyl-homoserine lactone (AHL). They regulate the expression of virulence genes in P. aeruginosa. The las system comprises of LasR, the transcriptional activator protein which is associated with AHL synthase LasI that regulates the synthesis of N-(3-oxododecanoyl)-l-homoserine lactone, the autoinducer (Kievit 2009). Similarly, the Rhl system consists of the transcriptional activator RhlR with its associated AHL, N-butyryl-l-homoserine lactone, which is synthesised by RhlI (Latifi et al. 1996).
A number of transcription factors regulate the rhlI-rhlR and the lasI-lasR quorum-sensing systems by direct or indirect mechanisms in order to adopt their expression to environmental variations (Kievit 2009). RhlI and lasI control the synthesis of the major autoinducers C4-HSL and 3-oxo-C12-HSL, respectively. A third signalling molecule called Pseudomonas quinolone signal (PQS), 2-heptyl-3-hydroxy-4(1H)-quinolone, is also produced by P. aeruginosa (Kievit 2009). The various cellular and secreted virulence factors that the las and rhl systems regulate include lectins, alkaline protease, lipase, elastase, exotoxin A, phospholipase C, LasA protease, pyocyanin and rhamnolipids (Duan and Surette 2007).
The lasR gene is regulated by two different control systems: the las quorum-sensing system (controlling lasI and lasR expression) and the global regulators Vfr and GacA. The las quorum-sensing system creates an autoinduction feedback loop. In a similar manner, the expression of rhlR is regulated by GacA, and the las system controls the rhlR and rhlI genes to some degree (Kievit and Iglewski 2000).
c-di-GMP-dependent polysaccharides biosynthesis
Bacteria use various small molecules to aid signalling pathways in order to modify their internal physiological conditions in response to extracellular environmental cues. Bis-(3′-5′)-cyclic-dimeric-guanosine monophosphate (c-di-GMP) is a ubiquitous intracellular secondary messenger in bacteria that is found in many of these pathways (D’Argenio and Miller 2004). c-di-GMP was first recognised in Gluconacetobacter xylinus as an allosteric effector of cellulose synthase (Ross et al. 1987). The role of c-di-GMP-dependent pathway in the biofilm formation of P. aeruginosa is shown in Fig. 3.
The pelA-G of c-di-GMP system is activated by the C4-HSL molecule produced by the rhl system associated with AHL (Rasamiravaka et al. 2015b). A majority of the c-di-GMP-dependent signalling pathways govern the interaction between the bacteria and biotic (eukaryotic and bacterial cells) or abiotic surfaces (Römling et al. 2013). Transformation from the motile to the sessile state occurs at high levels of c-di-GMP which promotes the biosynthesis of polysaccharides (alginate through algA-44-X gene and Pel through pelA-G gene) and conversely, lower levels of c-di-GMP enhances bacterial motility by promoting flagellar development and bacterial dispersion (Merighi et al. 2007).
The c-di-GMP-dependent pathways also aid the transition from the virulence phase as observed in acute infections to a more resilient and less virulence phase during chronic infections. Transmutation of these bacterial pathways could provide an alternative approach towards controlling biofilm formation and dispersal. As it is recognised by the mammalian immune systems, it is considered as a promising vaccine adjuvant (Römling et al. 2013).
P. aeruginosa undertakes an extensive adaptation to the lung and form persistent, low-virulence state and phenotypically distinct morphological form called small colony variants (SCVs). These are small, autoaggregative isolates that demonstrate superior biofilm formation, deep attachment to cell surfaces and increased production of exopolysaccharides. This often stabilises and contributes resistance to multiple antibiotics. The presence of SCVs in the patients is associated with a worse clinical situation (Evans 2015). The current mechanism for the generation of the SCV remains unclear. However, studies have suggested that the formation of SCVs is linked with overproduction of c-di-GMP which is probably responsible for the SCV phenotype in various isolates. Hence, the phenotypic variant to contribute to bacterial adaptation and survival of bacterial pathogens within the lung of patients and contributes to the pulmonary damage (Malone 2015).
Two-component regulatory systems
Two-component system (TCS) signalling pathways are vital signalling mechanisms in bacteria, archaea, simple eukaryotes and higher plants (Wolanin et al. 2002; Goodman et al. 2009). Classical TCS pathways possess a preserved central framework consisting of a sensor, a homo-dimerising histidine kinase protein domain and its response regulator, a cognate receiver domain, mechanistically linked by histidine-aspartic acid phosphorelay (Goodman et al. 2009). Changes in the environment are sensed by TCS in order to exhibit virulence factors that induce both acute and chronic infections (Okkotsu et al. 2014). Among the 60 such systems found in the genome of P. aeruginosa, the GacS/GacA system acts as a super-regulator of the QS system and plays important role in the formation of biofilm and multiple virulence factors (Rasamiravaka et al. 2015b).
GacS/GacA and RetS/LadS
The two-component regulatory system involves GacS, the sensor kinase, and GacA, its response regulator found in many γ-proteobacteria. It is involved in the regulation of quorum sensing, secondary metabolism, virulence, biofilm formation, motility and multiple features of bacterial physiology (Byrd et al. 2009).
GacA/GacS TCS regulates the expression of the psl and pel genes in P. aeruginosa. Once activated, GacA controls the transcription of two small regulatory RNAs (sRNAs), RsmZ and RsmY, that bind to repressor proteins, RsmA and RsmE, resulting in decrease or increase in the translation of the pel or psl operon (Wei and Ma 2013). The role of Gac/Ras pathway in biofilm formation is shown in Fig. 3.
The whole complex pathway involving GacS/A, RsmA and RsmY/Z is further directed by two additional hybrid sensor kinases, RetS and LadS (Dötsch et al. 2012). RetS is an activator of type III secretion (TTSS) and inhibits biofilm formation, contributing to acute infections. LadS, on the other hand, has been shown to up regulate biofilm formation, contributing to chronic infections. RetS/LadS interact with the GacS/GacA system by modulating the GacS phosphorylation state, which accordingly inhibits or promotes the phosphorylation and activation of GacA (Workentine et al. 2009). The GacS/GacA system regulates the AHL system as it inactivates freely available RsmA which is involved in negative control of C4-HSL and 3-oxo-C12-HSL synthesis, thereby monitoring extracellular virulence factors controlled by Las and Rhl systems (Rasamiravaka et al. 2015b).
Quorum-quenching systems
The virulence of P. aeruginosa PAO1 is governed by an N-Acyl-homoserine lactone (AHL)-dependent quorum-sensing system. The acylase gene present in the P. aeruginosa, PA2385 gene in PAO1 genome, was depicted to be encoded an acylase that eliminates the fatty acid side chain from the homoserine lactone (HSL) nucleus of AHL quorum-sensing signal molecules. Studies demonstrated that the posttranslational modification of acylase and hydrolysis are analogous to those of β-lactam acylases, suggesting that PA2385 gene product is associated with N-terminal nucleophile hydrolase superfamily. The purified acylase was demonstrated to degrade AHLs with side chains from 11 to 14 carbons. The substituent at the 3′ end of the side chain was not affected by the activity and showed high AHL quorum-quenching activity. The AHL signal molecule such as 3-oxo-C12-HSL is degraded by the acylase enzyme. The AHL acylase enable P. aeruginosa PAO1 to amend its own quorum-sensing-dependent pathogenic capabilities and suggest possibilities for novel antibacterial therapies (Si et al. 2016).
Biofilm treatment
Biofilms are described as niches of bacteria enclosed in a grid of protective extracellular matrix comprised of polymers. Bacteria in these biofilms reveal distinct features such as different morphology, physiology and high resistivity to antibiotic treatments and immune system. Biofilms have displayed high antagonism against therapeutic treatments thus allowing the bacteria to develop immunity. Finding an alternative mode of treatment of the infections caused by biofilms is highly laborious and strenuous task in the process of antimicrobial drug development (Wei and Ma 2013). The various approaches used for the biofilm treatment of P. aeruginosa are targeting adhesion and maturation, dispersal and virulence factors. These approaches are considered as the best promising strategies to eradicate biofilm formation.
Targeting adhesion and maturation
The lectins such as LecA and LecB are considered as putative drug targets for P. aerugenosa. LecA and LecB are soluble proteins binding to galactose and fucose, respectively. These proteins have pivotal role in the attachment of the bacteria to human cells, creating epithelial tissue damage and play important role in the development of P. aeruginosa biofilms, hence, acting as major virulence factors. LecA and LecB are small proteins with a size of 121 and 115 amino acid residues, respectively. As lectins play major role in the cell adhesion and biofilm formation, targeting these proteins can be a therapeutic remedy (Grishin et al. 2015). LecA has low affinity binding with D-galactose and N-acetyl-D-galactosamine. L-fucose showed high affinity binding to LecB. Both the lectins have same quaternary structures and form homotetrameric complexes where each monomer has its own binding site. Studies suggested that Lectin A binds with compounds such as α-D-galactose, p-amino-phenyl-galacto-pyranoside, phenyl-galacto-pyranoside, p-nitro-phenyl-galacto-pyranoside, p-tolyl-galacto-pyranoside and naphthyl-galacto-pyranoside, and the binding of these compounds can provide an inhibitory activities of the bacterial adhesion (Grishin et al. 2015).
Biofilm dispersal
Biofilm must disperse and liberate differentiated cells in order to inhabit new locations. Biofilm dispersal is responsible for the transfer of bacteria from environmental sources to human hosts, especially for many pathogenic bacteria (Kaplan 2010). It also promotes the bacterial survival by aiding in escaping hostile environments or those of declining suitability and successfully colonising more advantageous and unpopulated habitat patches (Berne et al. 2010).
Dispersion involves a decrease in bacterial adhesiveness and modulation or breakdown of the biofilm matrix (Harmsen et al. 2010). Dispersal takes place by three simultaneous and interdependent processes occurring within mature biofilms: (i) production of nitrosative or oxidative stress-inducing molecules (ii) prophage induction and (iii) cell disruption (Barraud et al. 2006).
By using flow chambers irrigated with Luria-Bertani (LB) medium, local dispersion of P. aeruginosa biofilm was observed as a cavity-like formation (Purevdorj-Gage et al. 2005). Initially, a barrier establishing a group of nonmotile cells formed the exterior component of the biofilm, and a swiftly moving motile group of cells are found within. The motile population would eventually find its way out of the biofilm, resulting in the formation of a central void. The biofilm must reach critical size in order for this phenomenon to occur (Harmsen et al. 2010).
Sauer et al. (2004) demonstrated that the dispersal of P. aeruginosa PAO1 biofilms is influenced by the presence of carbon substrates. A sudden increase in the availability of glucose, glutamate and succinate greatly facilitates the biofilm dispersion. The phenotypes displayed by single dispersed cells are different from those within the biofilm (Sauer et al. 2002). Biofilm dispersion induced by nutrients demonstrated reduced expression of pilus genes in dispersed cells and a corresponding increase in the flagellar expression (Sauer et al. 2004).
Studies revealed that the use of gaseous nitric oxide prevents treatment of the biofilm produced by methicillin-resistant Staphylococcus aureus, P. aeruginosa and Acinetobacter baumannii (Sulemankhil et al. 2012). In particular, nitric oxide (NO), a signalling molecule in P. aeruginosa, was observed to initiate biofilm dispersal at depressed and nontoxic concentration. It was also found to increase susceptibility to antibiotics (Barraud et al. 2006). Decreased c-di-GMP levels are related to dispersal and hence to planktonic bacteria while higher levels are seen in biofilm cells. Diguanylate cyclase (DGCs) regulates the biosynthesis of c-di-GMP and phosphodiesterase (PDE) regulates its degradation (Iyer et al. 2003; Barraud et al. 2009). Studies suggested that the decrease in the intracellular levels of c-di-GMP is induced by NO in turn stimulates PDE, consequently decreasing c-di-GMP levels in P. aeruginosa facilitating dispersion (Barraud et al. 2009).
Endogenous nitric oxide (NO) released by macrophages plays an important role in host defense, and NO is a therapeutic agent in preventing the formation of biofilms in vitro. By making use of a novel and controlled electrochemical NO-releasing catheters, the physiological levels of NO released has combined effects of killing bacteria and dispersing biofilm. Moreover, NO greatly increases the efficacy of therapeutic agents, reduces the c-di- GMP levels and thus helps in destroying the biofilm and its detached cells (Ren et al. 2016).
Recent studies suggested that the detachment processes of biofilm such as erosion or sloughing are considered as passive processes. These are the processes in which dispersal of biofilm occurs as a result of complex spatial differentiation and various molecular events in the bacterial cells in response to environmental factors. There are various biological factors that force bacterial cells to disperse from the biofilm (Kim and Lee 2016).
Targeting virulence factors
The major virulence factors that associated with P. aeruginosa during biofilm development are pili, T2SS such as exotoxin A, LasA, LasB, Type IV protease, alkaline protease, protease IV and phospholipase H. The major T3SS systems included ExoS, ExoT, ExoU, ExoY, T5SS and lectins such as LecA and LecB, siderophores, pyocyanin and quorum sensing. The major lipopolysaccharide involved as virulence factors are lipid A and O-specific polysaccharide. Inhibition of the functions of these virulence targets probably contributes newer insight in the development of novel therapeutic strategies against P. aeruginosa biofilm development.
Other advances in biofilm treatment
Some of the earlier approaches and strategies include a variety of chemical, biochemical and enzymatic techniques that can be visualised for disruption of the biofilms (Sharma et al. 2011; Landini et al. 2010; Bjarnsholt et al. 2013; Kostakioti et al. 2013). A direct approach is the chemical attack on the biofilm matrix which may be due to the success of halogens such as free chlorine and other biocides present in the therapeutics used against biofilms (Davison et al. 2010). Enzymatic cleavage of the biofilm extracellular matrix and other approaches using dispersin B (that degrades linear polymers of N-acetyl glucosamine, a common biofilm polysaccharide) are also used for cessation of biofilms. Exposure to this enzyme can remove biofilms of mixed species of bacteria (Stewart 2014).
The elimination of biofilms by commercially attainable therapeutics is inefficacious since biofilms are highly resistant. Tobramycin was effective to an extent, but it decreased significantly when employed to treat P. aeruginosa biofilms, although it caused cell death in mature biofilms (Reffuveille et al. 2014). A novel benzimidazole named anti-biofilm compound 1 (ABC-1), identified in a small-molecule screen, was found to prevent the initiation step in biofilm formation in many Gram-positive and Gram-negative pathogens including P. aeruginosa. This molecule is active at nanomolar concentrations. Further, layering the surface of biofilms with ABC-1 decreases the formation of biofilms (Sambanthamoorthy et al. 2011).
A different approach in the treatment of biofilms is by the use of therapeutic enzymes that breaks down the biofilm matrix. Dornase alfa (deoxyribonuclease I) is the enzyme in clinical use that destroys P. aeruginosa biofilm. This potent enzyme acts by hydrolysing eDNA in the extracellular matrix (Baker et al. 2016). Glycoside hydrolase therapy has the ability to concentrate large biofilms from many Gram-positive and Gram-negative bacteria which are relevant to both health care and industry. Glycoside hydrolase degrades the exopolysaccharide component of the biofilm matrix. PelA and pslG inhibit formation of biofilms over a 24-h period with an effective half-maximal concentration of 69.3 ± 1.2 and 4.1 ± 1.1 nM, respectively, and thus, they are capable of inhibiting pre-existing biofilms. This treatment reduced the biofilms by 58 to 94% (Baker et al. 2016). Biofilm formation on devices such as shunts, prosthetic vascular grafts, cardiac devices, intravascular catheters and prosthetic joints may be prevented by surface charges or by insertion of antimicrobials in the device that prevents formation of biofilms (Joo and Otto 2012).
Many naturally occurring compounds have played important role in P. aeruginosa to reduce biofilm production as they affect the QS-controlled gene expression. These included 6-gingerol (stifling oil from ginger), ajoene, eugenol prepared from cloves, some flavonoids and S-phenyl-L-cysteine sulfoxide. The bifurcation of Dalbergia trichocarpa bark has given rise to the purification and recognition of an aldehyde, coumarate which is oleanolic in nature, as a new therapeutic for biofilms (Rasamiravaka et al. 2015b). Cranberry proanthocyanidins (PACs) have properties against P. aeruginosa biofilms, especially A-type proanthocyanidins. These reduced the motility of various strains of P. aeruginosa. Proteomic analysis showed that different proteins were expressed with the treatment of PACs. These potentiate the activity of gentamycin, thus, inhibiting biofilms in vitro and in vivo (Ulrey et al. 2014).
An amalgamation of various medicinal plant extracts (Ayurvedic medicines), in which most of their mechanisms of action are known, are commonly used in India to treat bacterial colonisation that leads to biofilm formation. Some of them are Hareetaki Churna, Triphala Churna and Dashmula Churna. These exhibit properties against P. aeruginosa under in vitro conditions. Certain phytochemicals can also initiate the activity of antibacterial in combinations. The combination of sulfamethoxazole with gallic acid or ellagic acid and tetracycline with gallic acid showed effective killing of biofilms by synergistic mode of interaction (Chatterjee et al. 2016).
The use of bacteriophages to destroy biofilms has become a popular source of interest. T7 bacteriophage which encodes a lactonase enzyme was developed. These T7 bacteriophages exhibited Aii A lactonase to breakdown acyl homoserine lactones from various bacterial strains. These T7 bacteriophages when introduced in P. aeruginosa biofilms cause inhibition of biofilm formation (Pei and Lamas-Samanamud, 2014). Phages and drugs usually have minor effects in killing the bacteria. Phages are capable of limiting the bacteria to which minor populations of resistant bacteria ascend. However, some phage-drug combinations have significantly reduced bacteria to a great extent. In some cases, bacterial densities were reduced to levels even below the expected levels of synergy. Phages are also capable of limiting the extent to which minority populations of bacteria resistant to the treatment of antibiotics ascend (Chaudhry et al. 2017).
There is an urgent need for new methodologies to deal with biofilm-associated microorganisms, and antimicrobial photodynamic therapy (aPDT) may be a promising approach. A PDT includes the amalgamation of a nontoxic dye and low visible light which, in manifestation of oxygen, produces cytotoxic reactive oxygen species. This method has recognised the fact that many biofilms are gullible to aPDT (Melo et al. 2013).
There has been no record of well-defined methods accessible for practitioners to approve the occurrence of biofilms in injury. A diversity of microscopic and histological approaches to practical to biopsies of tissues is the recent enlightening techniques accessible for representing the occurrence of biofilms and the consequences they have in indorsing tenderness and downregulation of functions at the cellular level (Percival et al. 2015).
The available vaccines towards the biofilm by P. aeruginosa became limited. The increased comprehension of pathogenesis of P. aeruginosa facilitated in the recognition of several immunogens which could be effectively used for the development of novel vaccines. In spite of these efforts, a notable and an effective vaccine in opposition to P. aeruginosa has not been accomplished (Sharma et al. 2011). Different techniques were introduced to improve the life of patients and to improve quality. One of the major approaches is to develop the methods for inhibition of alginate in bacterial cells (Krylov et al. 2016). Raffinose, a galactotrisaccharide, shows successful biofilm inhibition of P. aeruginosa by binding with LecA protein responsible for biofilm inhibition. Further, raffinose decreased the concentration of c-di-GMP, by increasing the activity of a c-di-GMP-specific phosphodiesterase. Thus, the ability of raffinose to inhibit biofilm formation opens new possibilities for pharmacological and industrial applications (Kim et al. 2016).
Proteomics and gene expression studies of bacteria suggested that the recent advancement in these areas aid to analyse the genome of the bacteria and functional genes which are significant in upholding the infection and provide novel insight for the development of therapeutic agents (Mulcahy et al. 2014).
It was lately suggested that the biofilm display greater degree of resistance to antibiotics when compared to that of planktonic bacteria as they have an extracellular matrix that acts as a physical protection (Hall-Stoodley et al. 2004). A high-dose norspermidine has bactericidal effect on P. aeruginosa biofilms as norspermidine inhibits biofilm formation by inhibiting swimming motility, preventing cell surface attachment, and also by downregulating QS-related gene expression. Therefore, studies have suggested that norspermidine could be a potent inhibitior for P. aeruginosa biofilms (Ou et al. 2016).
Many bacteria comprise certain functional amyloid fibers on their cell surface that are responsible for biofilm formation and other community behaviors. Curli are extracellular functional amyloid fibers formed by many enterobacteriaceae members. Curli and type 1 pili displayed important roles in promoting the biofilms formation in many bacteria. Two analogs of FN075 and BibC6 of ring-fused 2-pyridones inhibit curli and type 1 pili and provide anti-biofilm and anti-virulence properties. Further, two molecules such as AA-861 (a benzoquinone derivative) and parthenolide (a sesquiterpene lactone) act as potential inhibitors against TasA protein of amyloid-like fibers and open therapeutic insight towards many biofilm forming bacteria including P. aeruginosa.
A vital polyphenolic compound is found in green tea which is detected to be at levels of 0.7 g/L which is about 1.5 mM in concentration; this compound is epigallocatechinn-3-gallate (EGCG) (Yang and Wang 1993). EGCG has wide variety of antibacterial effects (Steinmann et al. 2013); it acts by interfering with the quorum-sensing process and when it is exposed to P. aeruginosa PAO1, it inhibits swarming and downregulates the Las and PQS of the quorum-sensing process (Yang et al. 2010). EGCG plays an important role in binding with various proteins that are responsible for the formation of biofilm (Ishii et al. 2008; Ishii et al. 2011; Ehrnhoefer et al. 2008; Lorenzen et al. 2014; Jankun et al. 1997; Liang et al. 1997). One such approach is by blocking the outer membrane porins and production of hydrogen peroxide (Cui et al. 2012; Nakayama et al. 2013). EGCG can rearrange the previously formed amyloid fibrils to unstructured aggregates through the hydrophobic sites on the fibrils (Palhano et al. 2013); it inhibits fibrillation of amyloidogenic proteins such as Sup35 prion (Roberts et al. 2009), α-synuclein and β-amyloid (Ehrnhoefer et al. 2008; Bieschke et al. 2010), losozyme (He et al. 2009) and transthyretin (Ferreira et al. 2009).
The differential expansion of various imaging methods endures to push novel results and increase the knowledge on biofilm structure, formation and activity. The arrival and use of scanning laser confocal microscopy and optical sectioning have reformed the interpretation of biofilms, lashing their intangible changeover from one-dimensional to complex, three-dimensional structures. Furthermore, by the intent use of confocal video microscopy and particle-tracking velocity, variations in fluid flow around microcolonies, 3-D outlines of growth, development of arrangements of biofilm morphologies, and antimicrobial killing patterns can be observed (Haussler and Fuqua 2013).
Conclusions
Although reports are available on the biofilm formation of many Gram-negative bacteria, limited reports are available on the molecular basis and major virulence factors for biofilm development by P. aeruginosa which recently became extremely resistant to most of the conventional antibacterial agents. The current review provides a comprehensive, state-of-the-art information and recent aspects on the molecular basis of biofilm formation by P. aeruginosa. This review also provides insight on the relevance of quorum-sensing pathway, Gac/Rsm pathway and c-di-GMP signalling pathway in biofilm development. The present review suggest that both quorum-sensing and c-di-GMP pathways are probably the best pathways implicated in the biofilm formation by P. aeruginosa and targeting the virulence factors in these pathways could be one of the promising approaches for developing anti-biofilm agents. Thus, the review enlightens with the recent developments in the biofilm dispersal and treatment that can provide therapeutic intervention towards many strains of multidrug-resistant P. aeruginosa.
References
Annous BA, Fratamico PM, Smith JL (2009) Quorum sensing in biofilms: why bacteria behave the way they do. J Food Sci 74(1):R24–R34. https://doi.org/10.1111/j.1750-3841.2008.01022.x
Baker P, Hill PJ, Snarr BD, Alnabelseya N, Pestrak MJ, Lee MJ, Jennings LK, Tam J, Melnyk RA, Parsek MR, Sheppard DC, Wozniak DJ, Howell PL (2016) Exopolysaccharide biosynthetic glycoside hydrolases can be utilized to disrupt and prevent Pseudomonas aeruginosa biofilms. Sci Adv 2(5):e1501632. https://doi.org/10.1126/sciadv.1501632
Barraud N, Hassett DJ, Hwang SH, Rice SA, Kjelleberg S, Webb JS (2006) Involvement of nitric oxide in biofilm dispersal of Pseudomonas aeruginosa. J Bacteriol 188(21):7344–7353. https://doi.org/10.1128/JB.00779-06
Barraud N, Schleheck D, Webb JS, Hassett DJ, Rice SA, Kjelleberg S (2009) Nitric oxide signaling in Pseudomonas aeruginosa biofilms mediates phosphodiesterase activity, decreased cyclic di-GMP levels, and enhanced dispersal. J Bacteriol 191(23):7333–7342. https://doi.org/10.1128/JB.00975-09
Berne C, Kysela DT, Brun YV (2010) A bacterial extracellular DNA inhibits settling of motile progeny cells within a biofilm. Mol Microbiol 77(4):815–829. https://doi.org/10.1111/j.1365-2958.2010.07267.x
Bieschke J, Russ J, Friedrich RP, Ehrnhoefer DE, Wobst H, Neugebauer K, Wanker EE (2010) EGCG remodels mature alpha-synuclein and amyloid-beta fibrils and reduces cellular toxicity. Proc Natl Acad Sci U S A 107(17):7710–7715. https://doi.org/10.1073/pnas.0910723107
Billings N, Millan MR, Caldara M, Rusconi R, Tarasova Y, Stocker R, Ribbeck K (2013) The extracellular matrix component psl provides fast-acting antibiotic defense in Pseudomonas aeruginosa biofilms. PLoS Pathog 9(8):e1003526. https://doi.org/10.1371/journal.ppat.1003526
Bjarnsholt T, Jensen PØ, Jakobsen TH, Phipps R, Nielsen AK, Rybtke MT, Tolker-Nielsen T, Givskov M, Høiby N, Ciofu O, the Scandinavian Cystic Fibrosis Study Consortium (2010) Quorum sensing and virulence of Pseudomonas aeruginosa during lung infection of cystic fibrosis patients. PLoS One 5(4):e10115. https://doi.org/10.1371/journal.pone.0010115
Bjarnsholt T, Alhede M, Eickhardt-Sorensen SR, Moser C, Kühl M, Jensen PO, Hoiby N (2013) The in vivo biofilm trends. Microbiology 21:466–474
Bodey GP, Bolivar R, Fainstein V, Jadeja L (1983) Infections caused by Pseudomonas aeruginosa. Rev Infect Dis 5(2):279–313. https://doi.org/10.1093/clinids/5.2.279
Boyd A, Chakrabarty AM (1995) Pseudomonas aeruginosa biofilms: role of the alginate exopolysaccharide. J Ind Microbiol 15(3):162–168. https://doi.org/10.1007/BF01569821
Bucior I, Pielage JF, Engel JN (2012) Pseudomonas aeruginosa pili and flagella mediate distinct binding and signaling events at the apical and basolateral surface of airway epithelium. PLoS Pathog 8(4):e1002616. https://doi.org/10.1371/journal.ppat.1002616
Byrd MS, Sadovskaya I, Vinogradov E, Lu H, Sprinkle AB, Richardson SH, Ma L, Ralston B, Parsek MR, Anderson EM, Lam JS, Wozniak DJ (2009) Genetic and biochemical analyses of the Pseudomonas aeruginosa psl exopolysaccharide reveal overlapping roles for polysaccharide synthesis enzymes in psl and LPS production. Mol Microbiol 73(4):622–638. https://doi.org/10.1111/j.1365-2958.2009.06795.x
Byrd MS, Pang B, Mishra M, Swords WE, Wozniak DJ (2010) The Pseudomonas aeruginosa exopolysaccharide Psl facilitates surface adherence and NF-kappaB activation in A549 cells. MBio 1. doi:https://doi.org/10.1128/mBio.00140-10
Cai Z, Liu Y, Chen Y, Yam JKH, Chew SC, Chua SL, Wang K, Givskov M, Yang L (2015) RpoN regulates virulence factors of Pseudomonas aeruginosa via modulating the PqsR quorum sensing regulator. Int J Mol Sci 16(12):28311–28319. https://doi.org/10.3390/ijms161226103
Cai Q, Lia Z, Ouyanga Q, Luoa C, Gordonb VD (2016) Singly flagellated Pseudomonas aeruginosa chemotaxes efficiently by unbiased motor regulation. MBio 7(2):e00013-16. https://doi.org/10.1128/mBio.00013-16
Campodónico VL, Llosa NJ, Grout M, Döring G, Maira-Litrán T, Pier GB (2010) Evaluation of flagella and flagellin of Pseudomonas aeruginosa as vaccines. Infect Immun 78(2):746–755. https://doi.org/10.1128/IAI.00806-09
Chambers JR, Sauer K (2013) The MerR-like regulator BrlR impairs Pseudomonas aeruginosa biofilm tolerance to colistin by repressing PhoPQ. J Bacteriol 195(20):4678–4688. https://doi.org/10.1128/JB.00834-13
Chatterjee M, Anju CP, Biswas L, Anil Kumar V, Gopi Mohan C, Biswas R (2016) Antibiotic resistance in Pseudomonas aeruginosa and alternative therapeutic options. Int J Med Microbiol 306(1):48–58. https://doi.org/10.1016/j.ijmm.2015.11.004
Chaudhry WN, Acevedo JC, Park T, Andleeb S, Bull JJ, Levin BR (2017) Synergy and order effects of antibiotics and phages in killing Pseudomonas aeruginosa biofilms. PLoS One 12(1):e0168615. https://doi.org/10.1371/journal.pone.0168615
Chew SC, Kundukad B, Seviour T, van der Maarel JRC, Yang L, Rice SA, Doyle P, Kjelleberg S (2014) Dynamic remodeling of microbial biofilms by functionally distinct exopolysaccharides. MBio 5(4):e01536–e01514. https://doi.org/10.1128/mBio.01536-14
Chiang WC, Nilsson M, Jensen PO, Hoiby N, Nielsen TE, Givskov M, Tolker-Nielsen T (2013) Extracellular DNA shields against aminoglycosides in Pseudomonas aeruginosa biofilms. Antimicrob Agents Chemother 57(5):2352–2361. https://doi.org/10.1128/AAC.00001-13
Chrzanowski L, Lawniczak L, Czaczyk K (2012) Why do microorganisms produce rhamnolipids? World J Microb Biotechnol 28(2):401–419. https://doi.org/10.1007/s11274-011-0854-8
Colvin KM, Gordon VD, Murakami K, Borlee BR, Wozniak DJ, Wong GC, Parsek MR (2011) The pel polysaccharide can serve a structural and protective role in the biofilm matrix of Pseudomonas aeruginosa. PLoS Pathog 7(1):e1001264. https://doi.org/10.1371/journal.ppat.1001264
Colvin KM, Irie Y, Tart CS, Urbano R, Whitney JC, Ryder C, Howell PL, Wozniak DJ, Parsek MR (2012) The pel and psl polysaccharides provide Pseudomonas aeruginosa structural redundancy within the biofilm matrix. Environ Microbiol 14(8):1913–1928. https://doi.org/10.1111/j.1462-2920.2011.02657.x
Colvin KM, Alnabelseya N, Baker P, Whitney JC, Howell PL, Parsek MR (2013) PelA deacetylase activity is required for Pel polysaccharide synthesis in Pseudomonas aeruginosa. J Bacteriol 195(10):2329–2339. https://doi.org/10.1128/JB.02150-12
Costerton W, Veeh R, Shirtliff M, Pasmore M, Post C, Ehrlich G (2003) The application of biofilm science to the study and control of chronic bacterial infections. J Clin Invest 112(10):1466–1477. https://doi.org/10.1172/JCI200320365
Crabbé A, Ledesma MA, Nickerson CA (2014) Mimicking the host and its microenvironment in vitro for studying mucosal infections by Pseudomonas aeruginosa. Pathog Dis 71(1):1–19. https://doi.org/10.1111/2049-632X.12180
Cui Y, Oh YJ, Lim J, Youn M, Lee I, Pak HK, Park W, Jo W, Park S (2012) AFM study of the differential inhibitory effects of the green tea polyphenol (−)-epigallocatechin-3- gallate (EGCG) against Gram-positive and Gram-negative bacteria. Food Microbiol 29(1):80–87. https://doi.org/10.1016/j.fm.2011.08.019
D’Argenio DA, Miller SI (2004) Cyclic di-GMP as a bacterial second messenger. Microbiol Res 150:2497–2502
Das T, Manefield M (2012) Pyocyanin promotes extracellular DNA release in Pseudomonas aeruginosa. PLoS One 7(10):e46718. https://doi.org/10.1371/journal.pone.0046718
Das T, Manefield M (2013) Phenazine production enhances extracellular DNA release via hydrogen peroxide generation in Pseudomonas aeruginosa. Commun Integr Biol 6(3):e23570. https://doi.org/10.4161/cib.23570
Das T, Sehar S, Koop L, Wong YK, Ahmed S, Siddiqui KS, Manefield M (2014) Influence of calcium in extracellular DNA mediated bacterial aggregation and biofilm formation. PLoS One 9(3):e91935. https://doi.org/10.1371/journal.pone.0091935
Das T, Kutty SK, Tavallaie R, Ibugo AI, Panchompoo J, Sehar S, Aldous L, Yeung AWS, Thomas SR, Kumar N, Gooding JJ, Manefield M (2015) Phenazine virulence factor binding to extracellular DNA is important for Pseudomonas aeruginosa biofilm formation. Sci Rep 5(1):8398. https://doi.org/10.1038/srep08398
Das T, Ibugo AI, Klare W, Manefield M (2016) Role of pyocyanin and extracellular DNA in facilitating Pseudomonas aeruginosa biofilm formation, microbial biofilms - importance and applications, Dr. Dharumadurai Dhanasekaran (Ed.). InTech. https://doi.org/10.5772/63497
Dasgupta N, Wolfgang MC, Goodman AL, Arora SK, Jyot J, Lory S, Ramphal R (2003) A four-tiered transcriptional regulatory circuit controls flagellar biogenesis in Pseudomonas aeruginosa. Mol Microbiol 50(3):809–824. https://doi.org/10.1046/j.1365-2958.2003.03740.x
Davey ME, O’Toole GA (2000) Microbial biofilms: from ecology to molecular genetics. Microbiol Mol Biol Rev 64(4):847–867. https://doi.org/10.1128/MMBR.64.4.847-867.2000
Davison WM, Pitts B, Stewart PS (2010) Spatial and temporal patterns of biocide action against Staphylococcus epidermidis biofilms. Antimicrob Agents Chemother 54(7):2920–2927. https://doi.org/10.1128/AAC.01734-09
Dheilly A, Soum-Soutéra E, Klein GL, Bazire A, Compère C, Haras D, Dufour A (2010) Antibiofilm activity of the marine bacterium Pseudoalteromonas sp. strain 3J6. Appl Environ Microbiol 76(11):3452–3461. https://doi.org/10.1128/AEM.02632-09
Doggett RG (1969) Incidence of mucoid Pseudomonasaeruginosa from clinical sources. Appl Microbiol 18(5):936–937
Döring G, Meisner C, Stern M (2007) A double-blind randomized placebo-controlled phase III study of a Pseudomonas aeruginosa flagella vaccine in cystic fibrosis patients. Proc Natl Acad Sci U S A 104(26):11020–11025. https://doi.org/10.1073/pnas.0702403104
Dötsch A, Eckweiler D, Schniederjans M, Zimmermann A, Jensen V, Scharfe M, Geffers R, Häussler S (2012) The Pseudomonas aeruginosa transcriptome in planktonic cultures and static biofilms using RNA sequencing. PLoS One 7(2):e31092. https://doi.org/10.1371/journal.pone.0031092
Driscoll JA, Brody SL, Kollef MH (2007) The epidemiology, pathogenesis and treatment of Psuedomonas. Drugs 67(3):351–368. https://doi.org/10.2165/00003495-200767030-00003
Duan K, Surette MG (2007) Environmental regulation of Pseudomonas aeruginosa PAO1 Las and Rhl quorum-sensing systems. J Bacteriol 189(13):4827–4836. https://doi.org/10.1128/JB.00043-07
Dunne WM Jr (2002) Bacterial adhesion: seen any good biofilms lately? Clin Microbiol Rev 15(2):155–166. https://doi.org/10.1128/CMR.15.2.155-166.2002
Ehrnhoefer DE, Bieschke J, Boeddrich A, Herbst M, Masino L, Lurz R, Engemann S, Pastore A, Wanker EE (2008) EGCG redirects amyloidogenic polypeptides into unstructured, off-pathway oligomers. Nat Struct Mol Biol 15(6):558–566. https://doi.org/10.1038/nsmb.1437
Enge J, Eran Y (2011) Subversion of mucosal barrier polarity by Pseudomonas Aeruginosa. Front Microbiol 2:114
Evans TJ (2015) Small colony variants of Pseudomonas aeruginosa in chronic bacterial infection of the lung in cystic fibrosis. Future Microbiol 10(2):231–239. https://doi.org/10.2217/fmb.14.107
Fazeli H, Akbari R, Moghim S, Narimani T, Arabestani MR, Ghoddousi AR (2012) Pseudomonas aeruginosa infections in patients, hospital means, and personnel’s specimens. J Res Med Sci 17(4):332–337
Ferreira N, Cardoso I, Domingues MR, Vitorino R, Bastos M, Bai G, Saraiva MJ, Almeida MR (2009) Binding of epigallocatechin-3-gallate to transthyretin modulates its amyloidogenicity. FEBS Lett 583(22):3569–3576. https://doi.org/10.1016/j.febslet.2009.10.062
Flemming H, Neu T, Wingender J (2016) The perfect slime: microbial extracellular polymeric substances (EPS). IWA publishing
Franklin MJ, Nivens DE, Weadge JT, Howell PL (2011) Biosynthesis of the Pseudomonas aeruginosa extracellular polysaccharides, alginate, pel, and psl. Front Microbiol 2:167
Friedman L, Kolter R (2004a) Genes involved in matrix formation in Pseudomonas aeruginosa PA14 biofilms. Mol Microbiol 51(3):675–690
Friedman L, Kolter R (2004b) Two genetic loci produce distinct carbohydrate-rich structural components of the Pseudomonas aeruginosa biofilm matrix. J Bacteriol 186(14):4457–4465. https://doi.org/10.1128/JB.186.14.4457-4465.2004
Friedrich M (2015) Pseudomonasaeruginosa infections. http://emedicine.medscape.com/article/226748-overview#showall
Garrett TR, Bhakoob M, Zhang Z (2008) Bacterial adhesion and biofilms on surfaces. Prog Nat Sci 18(9):1049–1056. https://doi.org/10.1016/j.pnsc.2008.04.001
Gellatly SL, Hancock REW (2013) New insights into pathogenesis and host defenses. Pathog Dis 67(3):159–173. https://doi.org/10.1111/2049-632X.12033
Giltner CL, Nguyen Y, Burrows LL (2012) Type IV pilin proteins: versatile molecular modules. Microbiol Mol Biol Rev 76(4):740–772. https://doi.org/10.1128/MMBR.00035-12
González JE, Keshavan ND (2006) Messing with bacterial quorum sensing. Microbiol Mol Biol Rev 70(4):859–875. https://doi.org/10.1128/MMBR.00002-06
Goodman AL, Merighi M, Hyodo M, Ventre I, Filloux A, Lory S (2009) Direct interaction between sensor kinase proteins mediates acute and chronic disease phenotypes in a bacterial pathogen. Genes Dev 23(2):249–259. https://doi.org/10.1101/gad.1739009
Green LR (2011) Guide to the elimination of orthopedic surgery surgical site infections: an executive summary of the association for professionals in infection control and epidemiology elimination guide. Am J Infect Control 40:384–396
Grishin AV, Krivozubov MS, Karyagina AS, Gintsburg AL (2015) Pseudomonas aeruginosa lectins as targets for ovel antibacterials. Acta Nat 7(2):29–41
Gupta K, Marques CNH, Petrova OE, Sauer K (2013) Antimicrobial tolerance of Pseudomonas aeruginosa biofilms is activated during an early developmental stage and requires the two-component hybrid SagS. J Bacteriol 195(21):4975–4987. https://doi.org/10.1128/JB.00732-13
Habash MB, Park AJ, Vis EC, Harris RJ, Khursigara CM (2014) Synergy of silver nanoparticles and aztreonam against Pseudomonas aeruginosa PAO1 biofilms. Antimicrob Agents Chemother 58(10):5818–5830. https://doi.org/10.1128/AAC.03170-14
Hall-Stoodley L, Costerton JW, Stoodley P (2004) Bacterial biofilms: from the natural environment to infectious diseases. Nat Rev Microbiol 2(2):95–108. https://doi.org/10.1038/nrmicro821
Harmsen M, Yang L, Pamp SJ, Nielsen TT (2010) An update on Pseudomonas aeruginosa biofilm formation, tolerance, and dispersal. FEMS Immunol Med Microbiol 59(3):253–268. https://doi.org/10.1111/j.1574-695X.2010.00690.x
Haussler S, Fuqua C (2013) Biofilms 2012: new discoveries and significant wrinkles in a dynamic field. J Bacteriol 195(13):2947–2958. https://doi.org/10.1128/JB.00239-13
Hay ID, Remminghorst U, Rehm BHA (2009) MucR, a novel membrane-associated regulator of alginate biosynthesis in Pseudomonasaeruginosa. Appl Environ Microbiol 75(4):1110–1120. https://doi.org/10.1128/AEM.02416-08
Hazes B, Sastry PA, Hayakawa K, Read RJ, Irvin RT (2000) Crystal structure of Pseudomonas aeruginosa PAK pilin suggests a main-chain-dominated mode of receptor binding. J Mol Biol 299(4):1005–1017. https://doi.org/10.1006/jmbi.2000.3801
He J, Xing YF, Huang B, Zhang YZ, Zeng CM (2009) Tea catechins induce the conversion of preformed lysozyme amyloid fibrils to amorphous aggregates. J Agric Food Chem 57(23):11391–11396. https://doi.org/10.1021/jf902664f
Hentzer M, Teitzel GM, Balzer GJ, Heydorn A, Molin S, Givskov M, Parsek MR (2001) Alginate overproduction affects Pseudomonas aeruginosa biofilm structure and function. J Bacteriol 183(18):5395–5401. https://doi.org/10.1128/JB.183.18.5395-5401.2001
Ishii T, Mori T, Tanaka T, Mizuno D, Yamaji R, Kumazawa S, Nakayama T, Akagawa M (2008) Covalent modification of proteins by green tea polyphenol (−)-epigallocatechin-3- gallate through autoxidation. Free Radic Biol Med 45(10):1384–1394. https://doi.org/10.1016/j.freeradbiomed.2008.07.023
Ishii T, Ichikawa T, Minoda K, Kusaka K, Ito S, Suzuki Y, Akagawa M, Mochizuki K, Goda T, Nakayama T (2011) Human serum albumin as an antioxidant in the oxidation of (−)- epigallocatechin gallate: participation of reversible covalent binding for interaction and stabilization. Biosci Biotechnol Biochem 75(1):100–106. https://doi.org/10.1271/bbb.100600
Iyer LM, Anantharaman V, Aravind L (2003) Ancient conserved domains shared by animal soluble guanylylcyclases and bacterial signaling proteins. BMC Genomics 4(1):5–12. https://doi.org/10.1186/1471-2164-4-5
Jankun J, Selman SH, Swiercz R, Skrzypczak-Jankun E (1997) Why drinking green tea could prevent cancer. Nature 387(6633):561. https://doi.org/10.1038/42381
Jennings LK, Storek KM, Ledvina HE, Coulon C, Marmont LS, Sadovskaya I, Secor PR, Tseng BS, Scian M, Filloux A, Daniel J, Wozniak DJ, Howell PL, Parsek MR (2015) Pel is a cationic exopolysaccharide that cross-links extracellular DNA in the Pseudomonas aeruginosa biofilm matrix. Proc Natl Acad Sci U S A 112(36):11353–11358. https://doi.org/10.1073/pnas.1503058112
Joo HS, Otto M (2012) Molecular basis of in vivo biofilm formation by bacterial pathogens. Chem Biol 19(12):1503–1513. https://doi.org/10.1016/j.chembiol.2012.10.022
Kaplan JB (2010) Biofilm dispersal: mechanisms, clinical implications, and potential therapeutic uses. J Dent Res 89(3):205–218. https://doi.org/10.1177/0022034509359403
Kievit TR (2009) Quorum sensing in Pseudomonas aeruginosa biofilms. Environ Microbiol 11(2):279–288. https://doi.org/10.1111/j.1462-2920.2008.01792.x
Kievit TR, Iglewski BH (2000) Bacterial quorum sensing in pathogenic relationships. Infect Immun 68(9):4839–4849. https://doi.org/10.1128/IAI.68.9.4839-4849.2000
Kim SK, Lee JH (2016) Biofilm dispersion in Pseudomonas aeruginosa. J Microbiol 54(2):71–85. https://doi.org/10.1007/s12275-016-5528-7
Kim HS, Cha E, Kim Y, Jeon YH, Olson BH, Byun Y, Park HD (2016) Raffinose, a plant galactoside, inhibits Pseudomonas aeruginosa biofilm formation via binding to LecA and decreasing cellular cyclic diguanylate levels. Sci Rep 6(1):25318. https://doi.org/10.1038/srep25318
Kokare CR, Chakraborty S, Khopade AN, Mahadik KR (2009) Biofilm: importance and applications. IJBT 8:159–168
Kostakioti M, Hadjifrangiskou M, Hultgren SJ (2013) Bacterial biofilms: development, dispersal, and therapeutic strategies in the dawn of the post antibiotic era. Cold Spring Harb Perspect Med 3(4):a010306. https://doi.org/10.1101/cshperspect.a010306
Krylov V, Shaburova O, Pleteneva E, Bourkaltseva M, Krylov S, Kaplan A, Chesnokova E, Kulakov L, Magill D, Polygach O (2016) Modular approach to select bacteriophages targeting Pseudomonas aeruginosa for their application to children suffering with cystic fibrosis. Front Microbiol 7:1631
Lamppaa JW, Griswold KE (2013) Alginate lyase exhibits catalysis-independent biofilm dispersion and antibiotic synergy. Antimicrob Agents Chemother 57(1):137–145. https://doi.org/10.1128/AAC.01789-12
Landini P, Antoniani D, Burgess JG, Nijland R (2010) Molecular mechanisms of compounds affecting bacterial biofilm formation and dispersal. Appl Microbiol Biotechnol 86(3):813–823. https://doi.org/10.1007/s00253-010-2468-8
LaSarre B, Federle MJ (2013) Exploiting quorum sensing to confuse bacterial pathogens. Microbiol Mol Biol Rev 77(1):73–111. https://doi.org/10.1128/MMBR.00046-12
Latifi A, Foglino M, Tanaka K, Williams P, Lazdunski A (1996) A hierarchical quorum-sensing cascade in Pseudomonas aeruginosa links the transcriptional activators LasR and RhIR (VsmR) to expression of the stationary-phase sigma factor RpoS. Mol Microbiol 21(6):1137–1146. https://doi.org/10.1046/j.1365-2958.1996.00063.x
Laverty G, Gorman SP, Gilmore BF (2014) Biomolecular mechanisms of Pseudomonas aeruginosa and Escherichiacoli biofilm formation. Pathogens 3(3):596–632. https://doi.org/10.3390/pathogens3030596
Lebeaux D, Ghigo JM, Beloin C (2014) Biofilm-related infections: bridging the gap between clinical management and fundamental aspects of recalcitrance toward antibiotics. Microbiol Mol Biol Rev 78(3):510–543. https://doi.org/10.1128/MMBR.00013-14
Lee J, Zhang L (2015) The hierarchy quorum sensing network in Pseudomonas aeruginosa. Protein Cell 6(1):26–41. https://doi.org/10.1007/s13238-014-0100-x
Li YH, Tian X (2012a) Quorum sensing and bacterial social interactions in biofilms. Sensors (Basel) 12(12):2519–2538. https://doi.org/10.3390/s120302519
Li Y-H, Tian X (2012b) Quorum sensing and bacterial social interactions in biofilms. Sensors 12(12):2519–2538. https://doi.org/10.3390/s120302519
Liang YC, Lin-shiau SY, Chen CF, Lin JK (1997) Suppression of extracellular signals and cell proliferation through EGF receptor binding by (−)-epigallocatechin gallate in human A431 epidermoid carcinoma cells. J Cell Biochem 67(1):55–65. https://doi.org/10.1002/(SICI)1097-4644(19971001)67:1<55::AID-JCB6>3.0.CO;2-V
Limoli DH, Whitefield GB, Kitao T, Ivey ML, Davis MR, Grahl N, Hogan D, Rahme L, Howell LP, O’Toole G, Goldberg J (2017) Pseudomonas aeruginosa alginate overproduction promotes coexistence with Staphylococcus aureus in a model of cystic fibrosis respiratory infection. MBio 8(2):e00186-17. https://doi.org/10.1128/mBio.00186-17
Lorenzen N, Nielsen SB, Yoshimura Y, Vad BS, Andersen CB, Betzer C, Kaspersen JD, Christiansen G, Pedersen JS, Jensen PH, Mulder FA, Otzen DE (2014) How epigallocatechin gallate can inhibit alpha-synuclein oligomer toxicity in vitro. J Biol Chem 289(31):21299–21310. https://doi.org/10.1074/jbc.M114.554667
Lovewell RR, Patankar YR, Berwin B (2014) Mechanisms of phagocytosis and host clearance of Pseudomonas aeruginosa. Am J Phys Lung Cell Mol Phys 306(7):L591–L603. https://doi.org/10.1152/ajplung.00335.2013
Ma L, Conover M, Lu H, Parsek MR, Bayles K, Wozniak DJ (2009) Assembly and development of the Pseudomonas aeruginosa biofilm matrix. PLoS Pathog 5(3):e1000354. https://doi.org/10.1371/journal.ppat.1000354
Makin SA, Beveridge TJ (1996) The influence of A-band and B-band lipopolysaccharide on the surface characteristics and adhesion of Pseudomonas aeruginosa to surfaces. Microbiology 142(2):299–307. https://doi.org/10.1099/13500872-142-2-299
Malone JG (2015) Role of small colony variants in persistence of Pseudomonas aeruginosa infections in cystic fibrosis lungs. Infect Drug Resist 8:237–247. https://doi.org/10.2147/IDR.S68214
Mann EM, Wozniak DJ (2012) Pseudomonas biofilm matrix composition and niche biology. FEMS Microbiol Rev 36(4):893–916. https://doi.org/10.1111/j.1574-6976.2011.00322.x
Martinez E, Gomez JC (2016) Oxylipins produced by Pseudomonas aeruginosa promote biofilm formation and virulence. Nat Commun 7:13823. https://doi.org/10.1038/ncomms13823
Matsukawa M, Greenberg EP (2004) Putative exopolysaccharide synthesis genes influence Pseudomonas aeruginosa biofilm development. J Bacteriol 186(14):4449–4456. https://doi.org/10.1128/JB.186.14.4449-4456.2004
McCallum M, Tammam S, Little DJ, Robinson H, Koo J, Shah M, Calmettes C, Moraes TF, Burrows LL, Howell PL (2016) PilN binding modulates the structure and binding partners of the Pseudomonas aeruginosa type IVaPilus protein PilM. J Biol Chem 291(21):11003–11015. https://doi.org/10.1074/jbc.M116.718353
McConoughey SJ, Howlin R, Granger JF, Manring MM, Calhoun JH, Shirtlif M, Kathju S, Stoodley P (2014) Biofilms in periprosthetic orthopedic infections. Future Microbiol 9(8):987–1007. https://doi.org/10.2217/fmb.14.64
Melo WC, Avci P, Oliveira MN, Gupta A, Vecchio D, Sadasivam M, Chandran R, Huang YY, Yin R, Perussi LR, Tegos GP, Perussi JR, Dai T, Hamblin MR (2013) Photodynamic inactivation of biofilm: taking a lightly colored approach to stubborn infection. Expert Rev Anti-Infect Ther 11(7):669–693. https://doi.org/10.1586/14787210.2013.811861
Merighi M, Lee VT, Hyodo M, Hayakawa Y, Lory S (2007) The second messenger bis-(3′-5′)-cyclic-GMP and its PilZ domain-containing receptor Alg44 are required for alginate biosynthesis in Pseudomonas aeruginosa. Mol Microbiol 65(4):876–895. https://doi.org/10.1111/j.1365-2958.2007.05817.x
Miller MB, Bassler BL (2001) Quorum sensing in bacteria. Annu Rev Microbiol 55(1):165–199. https://doi.org/10.1146/annurev.micro.55.1.165
Montanaro L, Poggi A, Visai L, Ravaioli S, Campoccia D, Speziale P, Arciola CR (2011) Extracellular DNA in biofilms. Int J Artif Organs 34(9):824–831. https://doi.org/10.5301/ijao.5000051
Moradali MF, Ghods S, Rehm BH (2017) Pseudomonas aeruginosa lifestyle: a paradigm for adaptation, survival, and persistence. Front Cell Infect Microbiol 7:39. https://doi.org/10.3389/fcimb.2017.00039 eCollection 2017
Morrison AJ Jr, Wenzel RP (1984) Epidemiology of infections due to Pseudomonas aeruginosa. Rev Infect Dis 6(Supplement_3):S627–S642. https://doi.org/10.1093/clinids/6.Supplement_3.S627
Mulcahy LR, Isabella VM, Lewis K (2014) Pseudomonas aeruginosa biofilms in disease. Microb Ecol 68(1):1–12. https://doi.org/10.1007/s00248-013-0297-x
Nakayama M, Shimatani K, Ozawa T, Shigemune N, Tsugukuni T, Tomiyama D, Kurahachi M, Nonaka A, Miyamoto T (2013) A study of the antibacterial mechanism of catechins: isolation and identification of Escherichia coli cell surface proteins that interact with epigallocatechin gallate. Food Control 33(2):433–439. https://doi.org/10.1016/j.foodcont.2013.03.016
Nelson LK, D’Amours GH, Sproule-Willoughby KM, Morck DW, Ceri H (2009) Pseudomonas aeruginosa las and rhl quorum-sensing systems are important for infection and inflammation in a rat prostatitis model. Microbiology 155(Pt 8):2612–2619. https://doi.org/10.1099/mic.0.028464-0
Nicolle LE (2014) Catheter associated urinary tract infections. Antimicrob Resist Infect Control 25(3):23. https://doi.org/10.1186/2047-2994-3-23 eCollection 2014
Okkotsu Y, Little AS, Schurr MJ (2014) The Pseudomonas aeruginosa AlgZR two-component system coordinates multiple phenotypes. Front Cell Infect Microbiol 4:82
Ou L, She P, Wang Y, Liu F, Zhang D, Chen L, Luo Z, Xu H, Qi Y, Wu Y (2016) Effects of norspermidine on Pseudomonas aeruginosa biofilm formation and eradication. Microbiol Open 5:402–412
Overhage J, Schemionek M, Webb JS, Rehm BH (2005) Expression of the psl operon in Pseudomonas aeruginosa PAO1 biofilms: PslA performs an essential function in biofilm formation. Appl Environ Microbiol 71(8):4407–4413. https://doi.org/10.1128/AEM.71.8.4407-4413.2005
Palhano FL, Lee J, Grimster NP, Kelly JW (2013) Toward the molecular mechanism(s) by which EGCG treatment remodels mature amyloid fibrils. J Am Chem Soc 135(20):7503–7510. https://doi.org/10.1021/ja3115696
Parsek MR (2015) Controlling the connections of cells to the biofilm matrix. J Bacteriol 198(1):12–14. https://doi.org/10.1128/JB.00865-15
Pei R, Lamas-Samanamud GR (2014) Inhibition of biofilm formation by T7 bacteriophages producing quorum-quenching enzymes. Appl Environ Microbiol 80(17):5340–5348. https://doi.org/10.1128/AEM.01434-14
Percival SL, Vuotto C, Donelli Z, Lipsky BA (2015) Biofilms and wounds: an identification algorithm and potential treatment options. Adv Wound Care (New Rochelle) 4(7):389–397. https://doi.org/10.1089/wound.2014.0574
Petrova OE, Schurr JR, Schurr MJ, Sauer K (2011) The novel Pseudomonas aeruginosa two-component regulator BfmR controls bacteriophage-mediated lysis and DNA release during biofilm development through PhdA. Mol Microbial 81(3):767–783. https://doi.org/10.1111/j.1365-2958.2011.07733.x
Pier GB, Ramphal R (2005) In Mandell GL, Bennett JE, Dolin R (eds) Mandell, Douglas, and Bennett’s principles and practice of infectious diseases, 6th ed. Elsevier Inc
Poole K (2011) Pseudomonas aeruginosa: resistance to the max. Front Microbiol 2:65
Purevdorj-Gage B, Costerton WJ, Stoodley P (2005) Phenotypic differentiation and seeding dispersal in non-mucoid and mucoid Pseudomonas aeruginosa biofilms. Microbiology 151(5):1569–1576. https://doi.org/10.1099/mic.0.27536-0
Qu L, She P, Wang Y, Liu F, Zhang D, Chen L, Luo Z, Xu H, Qi Y, Wu Y (2016) Effects of norspermidine on Pseudomonas aeruginosa biofilm formation and eradication. Microbiology 5(3):402–412
Rajan S, Saiman L (2002) Pulmonary infections in patients with cystic fibrosis. Semin Respir Infect 17(1):47–55. https://doi.org/10.1053/srin.2002.31690
Ramphal R, Koo L, Ishimoto KS, Totten PA, Lara JC, Lory S (1991) Adhesion of Pseudomonas aeruginosa pilin-deficient mutants to mucin. Infect Immun 59(4):1307–1311
Rasamiravaka T, Vandeputte OM, Pottier L, Huet J, Rabemanantsoa C, Kiendrebeogo M, Rasamindrakotroka A, Stévigny C, Duez P, Jaziri ME (2015a) Pseudomonas aeruginosa biofilm formation and persistence, along with the production of quorum sensing-dependent virulence factors, are disrupted by a triterpenoid coumarate ester isolated from Dalbergia trichocarpa, a tropical legume. PLoS One 10(7):e0132791. https://doi.org/10.1371/journal.pone.0132791
Rasamiravaka T, Labtani Q, Duez P, Jaziri ME (2015b) The formation of biofilms by Pseudomonas aeruginosa: a review of the natural and synthetic compounds interfering with control mechanisms. BioMed Res Int 17 pages
Reffuveille F, Fuente-Núñez CL, Mansour S, Hancock REW (2014) A broad-spectrum antibiofilm peptide enhances antibiotic action against bacterial biofilms. Antimicrob Agents Chemother 58(9):5363–5371. https://doi.org/10.1128/AAC.03163-14
Ren H, Wu J, Colletta A, Meyerhoff ME, Xi C (2016) Efficient eradication of mature Pseudomonas aeruginosa biofilm via controlled delivery of nitric oxide combined with antimicrobial peptide and antibiotics. Front Microbiol 7:1260
Rendueles O, Ghigo JM (2012) Multi-species biofilms: how to avoid unfriendly neighbors. FEMS Microbiol Rev 36(5):972–989. https://doi.org/10.1111/j.1574-6976.2012.00328.x
Roberts BE, Duennwald ML, Wang H, Chung C, Lopreiato NP, Sweeny EA, Knight MN, Shorter J (2009) A synergistic small-molecule combination directly eradicates diverse prion strain structures. Nat Chem Biol 5(12):936–946. https://doi.org/10.1038/nchembio.246
Römling U, Galperin MY, Gomelsky M (2013) Cyclic di-GMP: the first 25 years of a universal bacterial second messenger. Microbiol Mol Biol Rev 77(1):1–52. https://doi.org/10.1128/MMBR.00043-12
Ross P, Weinhouse H, Aloni Y (1987) Regulation of cellulose synthesis in Acetobacter xylinum by cyclic diguanylate. Nature 325(6101):279–281. https://doi.org/10.1038/325279a0
Ryder C, Byrd M, Wozniak DJ (2007) Role of polysaccharides in Pseudomonas aeruginosa biofilm development. Curr Opin Microbiol 10(6):644–648. https://doi.org/10.1016/j.mib.2007.09.010
Sadikot RT, Blackwell TS, Christman JW, Prince AS (2005) Pathogen-host interactions in Pseudomonas aeruginosa pneumonia. Am J Respir Crit Care Med 171(11):1209–1223. https://doi.org/10.1164/rccm.200408-1044SO
Sambanthamoorthy K, Gokhale AA, Lao W, Parashar V, Neiditch MB, Semmelhack MF, Lee I, Waters CM (2011) Identification of a novel benzimidazole that inhibits bacterial biofilm formation in a broad-spectrum manner. Antimicrob Agents Chemother 55(9):4369–4378. https://doi.org/10.1128/AAC.00583-11
Sauer K, Camper AK, Ehrlich GD, Costerton JW, Davies DG (2002) Pseudomonas aeruginosa displays multiple phenotypes during development as a biofilm. J Bacteriol 184(4):1140–1154. https://doi.org/10.1128/jb.184.4.1140-1154.2002
Sauer K, Cullen MC, Rickard AH, Zeef LAH, Davies DG, Gillbert P (2004) Characterization of nutrient-induced dispersion in Pseudomonas aeruginosa PAO1 biofilm. J Bacteriol 186(21):7312–7326. https://doi.org/10.1128/JB.186.21.7312-7326.2004
Schurr MJ (2013) Which bacterial biofilm exopolysaccharide is preferred, psl or alginate? J Bacteriol 195(8):1623–1626. https://doi.org/10.1128/JB.00173-13
Secinti DK, Özalp H, Attar A, Sargon FM (2011) Nanoparticle silver ion coatings inhibit biofilm formation on titanium implants. J Clin Neurosci 18(3):391–395. https://doi.org/10.1016/j.jocn.2010.06.022
Sharma A, Krause A, Worgall S (2011) Recent developments for Pseudomonas vaccines. Hum Vaccin 7(10):999–1011. https://doi.org/10.4161/hv.7.10.16369
Si X, Quan X, Wu Y (2016) A small-molecule norspermidine and norspermidine-hosting polyelectrolyte coatings inhibit biofilm formation by multi-species wastewater culture. Appl Microbiol Biotechnol 99:10861
Steinmann J, Buer J, Pietschmann T, Steinmann E (2013) Anti-infective properties of epigallocatechin-3-gallate (EGCG), a component of green tea. Br J Pharmacol 168(5):1059–1073. https://doi.org/10.1111/bph.12009
Stewart PS (2002) Mechanisms of antibiotic resistance in bacterial biofilms. Int J Med Microbiol 292(2):107–113. https://doi.org/10.1078/1438-4221-00196
Stewart PS (2014) Biophysics of biofilm infection. Pathog Dis 70(3):212–218. https://doi.org/10.1111/2049-632X.12118
Sulemankhil I, Ganopolsky JG, Dieni CA, Dan AF, Jones ML, Prakash S (2012) Prevention and treatment of virulence bacterial biofilms with an enzymatic nitric oxide-releasing dressing. Antimicrob Agents Chemother 56(12):6095–6103. https://doi.org/10.1128/AAC.01173-12
Tielen P, Kuhn H, Rosenau F, Jaeger K, Flemming H, Wingender J (2013) Interaction between extracellular lipase LipA and the polysaccharide alginate of Pseudomonas aeruginosa. BMC Microbiol 13(1):159. https://doi.org/10.1186/1471-2180-13-159
Todar K (2008) Todar’s online textbook of bacteriology. University of Wisconsin-Madison Department of Bacteriology. http://textbookofbacteriology.net/pseudomonas.html
Ulrey RK, Barksdale SM, Zhou W, Hoek ML (2014) Cranberry proanthocyanidins have anti-biofilm properties against Pseudomonas aeruginosa. BMC Complement Altern Med 14(1):499. https://doi.org/10.1186/1472-6882-14-499
van ‘t Wout EF, van Schadewijk A, van Boxtel R, Dalton LE, Clarke HJ, Tommassen J, Marciniak SJ, Hiemstra PS (2015) Virulence factors of Pseudomonas aeruginosa induce both the unfolded protein and integrated stress responses in airway epithelial cells. PLoS Pathog 11(6):e1004946. https://doi.org/10.1371/journal.ppat.1004946
Varki A, Cummings RD, Esko JD, Freeze HH, Stanley P, Bertozzi CR, Hart GW, Etzler ME (2009) Essentials of glycobiology. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Venkatakrishnan V, Packer NH, Thaysen-Andersen M (2013) Host mucin glycosylation plays a role in bacterial adhesion in lungs of individuals with cystic fibrosis. Expert Rev Respir Med 7(5):553–576. https://doi.org/10.1586/17476348.2013.837752
Wang S, Liu X, Liu H, Ma LZ (2015) The exopolysaccharide psl-eDNA interaction enables the formation of a biofilm skeleton in Pseudomonas aeruginosa: psl-eDNA interaction and biofilm skeleton. Environ Microbiol Rep 7(2):330–340. https://doi.org/10.1111/1758-2229.12252
Wei Q, Ma LZ (2013) Biofilm matrix and its regulation in Pseudomonas aeruginosa. Int J Mol Sci 14(10):20983–21005. https://doi.org/10.3390/ijms141020983
Wiens JR, Vasilb AI, Schurrb MJ, Vasil ML (2014) Iron-regulated expression of alginate production, mucoid phenotype, and biofilm formation by Pseudomonas aeruginosa. MBio 5(1):e01010-13. https://doi.org/10.1128/mBio.01010-13
Wilton M, Mazenod LC, Moore R, Lewenza S (2015) Extracellular DNA acidifies biofilms and induces aminoglycoside resistance in Pseudomonas aeruginosa. Antimicrob Agents Chemother 60(1):544–553. https://doi.org/10.1128/AAC.01650-15
Wilton M, Wong MJQ, Tang L, Liang X, Moore R, Parkins MD, Lewenza S, Dong TG (2016) Chelation of membrane-bound cations by extracellular DNA activates the T6SS in Pseudomonas aeruginosa. Infect Immun 84(8):2355–2361. https://doi.org/10.1128/IAI.00233-16
Wolanin PM, Thomason PA, Stock JB (2002) Histidine protein kinases: key signal transducers outside the animal kingdom. Genome Biol 3(10):reviews3013.1. https://doi.org/10.1186/gb-2002-3-10-reviews3013
Workentine ML, Chang L, Ceri H, Turner RJ (2009) The GacS–GacA two-component regulatory system of Pseudomonasfluorescens: a bacterial two-hybrid analysis. FEMS Microbiol Lett 292(1):50–56. https://doi.org/10.1111/j.1574-6968.2008.01445.x
Xu W, He L, Liu C, Rong J, Shi Y, Song W, Zhang T, Wang L (2015) The effect of infection control nurses on the occurrence of Pseudomonas aeruginosa healthcare-acquired infection and multidrug-resistant strains in critically ill children. PLoS One 10(12):e0143692. https://doi.org/10.1371/journal.pone.0143692
Yang CS, Wang ZY (1993) Tea and cancer. J Natl Cancer Inst 85(13):1038–1049. https://doi.org/10.1093/jnci/85.13.1038
Yang L, Liu Y, Sternberg C, Molin S (2010) Evaluation of enoyl-acyl carrier protein reductase inhibitors as Pseudomonas aeruginosa quorum-quenching reagents. Molecules 15(2):780–792. https://doi.org/10.3390/molecules15020780
Zegans ME, Wozniak D, Griffin E, Toutain-Kidd CM, Hammond JH, Garfoot A, Lamd JS (2012) Pseudomonas aeruginosa exopolysaccharide psl promotes resistance to the biofilm inhibitor polysorbate 80. Antimicrob Agents Chemother 56(8):4112–4122. https://doi.org/10.1128/AAC.00373-12
Zhang L, Fritsch M, Hammond L, Landreville R, Slatculescu C, Colavita A, Mah TF (2013) Identification of genes involved in Pseudomonas aeruginosa biofilm-specific resistance to antibiotics. PLoS One 8(4):e61625. https://doi.org/10.1371/journal.pone.0061625
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Skariyachan, S., Sridhar, V.S., Packirisamy, S. et al. Recent perspectives on the molecular basis of biofilm formation by Pseudomonas aeruginosa and approaches for treatment and biofilm dispersal. Folia Microbiol 63, 413–432 (2018). https://doi.org/10.1007/s12223-018-0585-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-018-0585-4