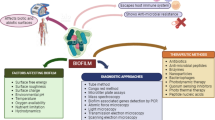
Abstract
Biofilms are considered as a severe problem in the treatment of bacterial infections; their development causes some noticeable resistance to antibacterial agents. Biofilms are responsible for at least two-thirds of all infections, displaying promoted resistance to classical antibiotic treatments. Therefore, finding new alternative therapeutic approaches is essential for the treatment and inhibition of biofilm-related infections. Therefore, this review aims to describe the potential therapeutic strategies that can inhibit bacterial biofilm development; these include the usage of antiadhesion agents, AMPs, bacteriophages, QSIs, aptamers, NPs and PNAs, which can prevent or eradicate the formation of biofilms. These antibiofilm agents represent a promising therapeutic target in the treatment of biofilm infections and development of a strong capability to interfere with different phases of the biofilm development, including adherence, polysaccharide intercellular adhesion (PIA), quorum sensing molecules and cell-to-cell connection, bacterial aggregation, planktonic bacteria killing and host-immune response modulation. In addition, these components, in combination with antibiotics, can lead to the development of some kind of powerful combined therapy against bacterial biofilm-related infections.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Most nosocomial infections have been caused by opportunistic pathogens in the recent decades; these are associated with various infections including bacteremia, urinary tract infections (UTIs), wound, meningitis and biofilm-associated infections [1]. Bacteria that are firmly attached to artificial medical devices cause biofilm-associated infections. Treatment of these infections is difficult because of the increased antibiotic resistance in the biofilm, which is commonly due to multidrug-resistant strains [2, 3]. Nowadays, biofilm infections have become a serious concern and a main global healthcare problem [1]. The identification of novel therapeutic targets to fight biofilm-related infections, thus, signifies one of the primary problems in the field of antibiotic therapy [4]. Bacteria are present in the environment in two forms: free-living or biofilm. Bacteria inside a biofilm can demonstrate low levels of sensitivity to some antimicrobial compounds including biocides and antibiotics [2, 3]. Therefore, antibiofilm compounds can be regarded as one of the most encouraging options to combat biofilm-related infections. These compounds have shown a broad range of biological functions including antibacterial properties on free-living cells, as well as antibiofilm properties [5].
Biofilm is known as a bacterial population in which cells adhere to biotic or abiotic surfaces by extracellular polymeric substances (EPS). Biofilm could be considered as a strategy some bacteria could adopt to withstand adverse conditions such as desiccation, host defense system and antibacterial agents [6].
The stages involved in the formation of biofilm in bacteria comply with a generic model consisting of three stages. The features of each stage are characteristics in different species, but the general traits are all alike [7]. The main stages in the biofilm formation are attachment, accumulation and dispersal [8]. In the first stage, the attachment to the surface is reversible and not very strong. Next, a loose bond is established between cells, and the extracellular matrix begins to secrete. In the later stages, the biofilm structure gains shape and other bacteria still in the environment attach themselves to this structure. Following the evolution of the biofilm structure, the cells are released back into the environment, causing new biofilm foci elsewhere [9].
In the biofilm phase, bacteria are surrounded in a self-produced extracellular matrix principally made of EPS [10]. EPS comprises over 90% of the dry weight of the biofilm, facilitating bonding to surfaces, which can cause microcolony formation and resistance to antimicrobial agents [11]. EPS is made up of exopolysaccharides, proteins and extracellular DNA (eDNA) [10]. EPS matrix reduces the permeation of antibiotics to spread bacteria within the biofilm by dispersion restriction or neutralization of antibacterial agents with extracellular polysaccharides [12]. Bacterial biofilms are significant in causing infectious diseases, especially chronic infections, in the host [13, 14]. According to some estimations previously carried out, approximately 80% of bacterial infections in the human body are associated with the formation of biofilms, which can increase the mortality rate in the hospitalized patients [2, 14]. Therefore, managing biofilm-related infections is challenging because of the problems inherent in inhibiting and treating them [3].
Given the specific conditions in the biofilm environment, treating and employing antibiotics to tackle this problem can be regarded as one of the important challenges the medical sciences face. Biofilm-associated infections cannot be treated by classical antibiotics, and they are a challenge worldwide. This has led to a set of studies aiming to find modern treatments for biofilm infections [15]. Therefore, new antibacterial or antibiofilm strategies with various mechanisms of action are urgently needed. Among these strategies, development of new classes of antimicrobial peptides (AMPs) [16], bacteriophages [17], quorum sensing inhibitors (QSIs) [18], aptamers [19], nanoparticles (NPs) [20] and peptide nucleic acids (PNAs) [21] can be considered as the most feasible solution.
In the recent years, several innovative antibiofilm agents have been developed to limit bacterial adhesion to abiotic and biotic surfaces; these are intended to target bacterial signals for removing grown biofilms or displacing cells from the established biofilms [4]. Therefore, this review aims to review and describe the potential therapeutic strategies that could be applied to prevent bacterial biofilm development; these include the usage of antiadhesion agents, AMPs, bacteriophages, QSIs, aptamers, NPs and PNAs, which can prevent or eradicate the biofilm formation.
Properties of Desirable Antibiofilm Agents
Biofilms are bacterial populations demonstrating exclusive properties in comparison with their free-living forms [22]. These properties should be correctly considered when assessing the potential of biofilm inhibition mechanisms. The EPS matrix in the biofilm development plays a significant and critical role in determining antibiotic resistance mechanisms of biofilm [4]. It basically establishes a dispersion barrier which can prevent the interaction of antibacterial agents with bacterial cells [23]. Restricting or preventing EPS accumulation and having the capability to permeate EPS could be regarded as characteristics of an ideal antibiofilm agent [24]. Therefore, characteristics of an ideal antibiofilm substance include antibacterial activities, easy penetration into the cell, unique structure, interference with the machinery of bacterial cells communication and synergism with other antibacterial agents [4]. Many of such properties can be observed in natural and synthetic antibiofilm substances [25,26,27,28]. Inhibition of the biofilm development by these compounds may simplify the treatment of biofilm-related infections.
Strategies for the Inhibition and Disruption of Bacterial Biofilms
Biofilms are highly resistant to conventional antibiotics; therefore, new alternative therapeutic approaches are needed to treat biofilm-related infections [3, 29]. There are several strategies to inhibit and eradicate biofilm development; these include antiadhesion agents, AMPs, bacteriophages, QSIs, aptamers, NPs and PNAs.
Antiadhesion Agents
The introduction of bacteria to a surface is influenced by the stochastic process, which is driven by gravitational forces, the surrounding hydrodynamic forces and Brownian motion [30, 31]. To overcome repulsive and hydrodynamic forces, motile bacteria utilize flagella, which can play an important role in the initial attachment in the case of several pathogens such as Escherichia coli, Pseudomonas aeruginosa, Listeria monocytogenes and Vibrio cholera [32,33,34,35]. The roles of chemotaxis in directing attachment and biofilm formation in response to nutrient composition have been demonstrated. Schmidt et al. [36], for instance, showed that the CheR1 methyltransferase mutations of P. aeruginosa could alter the amino acid response and inhibit biofilm maturation through impairing attachment. In addition, biofilm defects have been revealed in the tar gene (encodes methyl-accepting chemotaxis protein II) in uropathogenic E. coli (UPEC) [37]. Type 1 pili is another type of adhesion in UPEC and other E. coli that can play a role in the initial attachment, maintaining a steadfast grip on the surface and shear forces [38, 39]. These pili, which are assembled by the chaperon usher pathway (CUP), are multi-subunit adhesive [40]. CUP pili systems facilitate adherence in a niche-specific manner [37, 40, 41]. Adherence in the type 1 pili is mediated by the FimH adhesion, which recognizes mannose-rich regions [42]. Antigen 43 and curli fibers are other types of adhesion mediating interbacterial interactions and attachment on biotic and abiotic surfaces [43]. There are other attachment organelles in bacteria, including type IV pili and numerous CUP fimbriae (such as CupA) in P. aeruginosa [44], Ace, Esp and Ebp in Enterococcus faecalis [45], and SagA and Acm in Enterococcus faecium [46]. Therefore, several studies have focused on the development of compounds interacting and interfering with the first step of attachment, leading to the biofilm formation. Mannosides are molecules that competitively inhibit FimH mannose binding [47]. Among mannosylated proteins, the highest affinity of the monovalent ligands has been displayed in long chain aryl mannosides and alkyl mannosides, which could be due to the increased interactions with Tyr-48, Tyr-137 and Ile-52 of the binding pocket [48]. The development of monomeric biphenyl mannosides could lead to FimH inhibitors [49], which can prevent in vitro UPEC biofilm formation and interfere with the in vivo adherence and invasion of UPEC [49]. Cusumano et al. [49] also demonstrated that the combination of mannosides with TMP-SMZ (trimethoprim-sulfamethoxazole) could enhance antimicrobial treatment in clinical settings. In addition, the bladder bacterial load of UPEC was reduced in orally mannoside treated mice within 6 h, thus serving as an efficient therapeutic strategy for chronic UTIs [49].
In parallel, pilicides have been introduced to inhibit CUPs and type 1 pili, which are components derived from peptodomimetic scaffolds such as C-2 substituted thiazolo and dihydrothiazolo ring-fused 2-pyridones, as well as the bromomethyl substituted scaffold that can interfere and interact with the exportation of the subunits corresponding to the pili structure [50, 51]. Furthermore, curlicides (components derived from ring-fused 2-pyridones, such as FN075 and BibC6) have been introduced to inhibit the curli synthesis followed by the inhibition of the biofilm formation [43]. According to some studies, bacteria utilize a variety of receptors and adhesion types during their adherence process, facilitating adhesion in a niche-specific manner; therefore, interactions of multiple molecules may need to be inhibited to remove a biofilm from surfaces.
Antimicrobial Peptides
Antimicrobial peptides (AMPs) are small cationic molecules displaying a broad range of activity against microorganisms [52]. AMPs are divided into natural and synthetic groups. Table 1 shows the list of AMPs and their properties. Natural AMPs are produced by cellular tissues in a wide range of organisms; they can be a source for synthetic AMPs [53, 54]. AMPs are suggested as a potential method to cope with the biofilm formation. AMPs as antibiofilm agents have received much attention. Many AMPs have a positive charge, enabling them to interact with the phosphate groups of lipopolysaccharides as negatively charged components of the cell membrane in Gram-negative bacteria or lipoteichoic acids in Gram-positive bacteria [55, 56]. Most AMPs exhibit strong antibiofilm properties against antibiotic-resistant bacteria, which can be effective with different mechanisms at various phases of the biofilm development and on different molecular targets [16]. Some peptides can kill bacteria through membrane distraction and/or pore formation, as well as inhibition of the bacterial cell division; further, they can be effective on bacteria through preventing the adherence of the bacterial cells to the substrate surface, downregulating QS signals and removing the pre-formed biofilm [16]. Downregulation of genes involved in the motility and inhibition of a series of cellular biological procedures, such as the synthesis of cell walls, DNA, RNA and proteins, are other instances of AMP-antibiofilm mechanisms of function [57,58,59,60]. Figure 1 shows multiple mechanisms of antimicrobial peptides on the bacterial biofilm formation.
Antibiofilm peptides (ABPs) are a type of AMPs with the antibiofilm activity; they prevent the biofilm development at concentrations much lower than those of common antibiotics. The minimum inhibitory concentration (MIC) is, in fact, greater than the minimum biofilm inhibitory concentration (MBIC) [71]. ABPs can be linked to ppGpp molecules (second messenger nucleotide), disrupting them; they are involved in several metabolic pathways like colonization, attachment and aggression in bacteria. Therefore, ABPs prevent the increase of these molecules in the cell and inhibit the biofilm development in bacteria [72, 73]. ABPs commonly interact with signal molecules to apply control or distraction effects on the biofilms development; therefore, they can help other antibacterial agents to cope with the bacterial cells. For instance, Ribeiro et al. [74] described that ABPs promoted the sensitivity of carbapenemase-producing Klebsiella pneumoniae to carbapenems.
It has been demonstrated that most AMPs can be in combined with antibiotics to enhance their ability to prevent biofilm development and eliminate mature biofilms. Synergism of AMPs with other antibacterial agents is promised in the cleaning of biofilm-related infections, as they can increase the antibiofilm effect and reduce the drug dose [68]. Tong et al. [75] also reported the synergistic interaction between nisin (a natural AMP) and penicillin or chloramphenicol against the E. faecalis biofilm. The synergistic efficacy of nisin with nafcillin on biofilm in Streptococcus mutans was evaluated by Tong et al. [76]. Their results showed that nisin in combination with antibiotics considerably decreased the biofilm development. Therefore, AMPs could be regarded as potential antibiofilm agents through different mechanisms including functional inhibition of proteins, cell-membrane-disrupting action, detoxification of lipoteichoic acid and lipopolysaccharide, and binding with DNA.
Bacteriophages
The viruses that can infect and kill bacteria are known as bacteriophages (phages); they are not able to infect human or animal cells [77]. Therefore, phages are known as the key hunters of bacteria in the environment, and phage therapy has always been of interest for medical goals. Phage therapy emphasizes on lytic phages because they kill their host bacterial cells. In addition, lytic phages lack integrases and/or genes that participate in the horizontal gene transfer. Lytic phages are capable of lysing their host bacterial cells; as well, they can be amplified inside the host bacterial cells. Lysis of the host bacterial cells destroys the bacteria, as well as releasing the progeny phages for the re-infection of more bacteria. Phages are species-specific; thus, they can be used to target special pathogenic bacteria, without any effect on the commensal bacteria [17].
Phage therapy has several advantages in comparison to common antibiotics; these include specificity of function, a narrow range of action, greater safety, greater tolerability, an effect limited to the site of infection and cost effectiveness [78]. The use of phages to inhibit bacterial infections is being followed as an alternative to antibiotics [79]. According to some recent studies, the role of phages in the eradication and/or prevention of bacteria biofilms has been highlighted. Phages are capable of penetrating into the structure of bacterial biofilms and removing them [80, 81]. There are several water channels in the biofilm structure, allowing the phages to easily penetrate into the biofilms' inner structure [82]. Furthermore, most of the phages generate depolymerases, which are capable of hydrolysing the EPS of the bacterial biofilms [83]. For instance, phage phi 15 produces the polysaccharide depolymerase enzyme, which hydrolyses the EPS of Pseudomonas putida and inhibits its biofilm formation [84]. As well, it has been observed that T4-like and Φ29-like phages from the family of Myoviridae and Podoviridae, respectively, suppress the biofilm of Staphylococcus aureus [85].
Phage-derived enzymes like lysin enzymes are known as bactericidal components that hydrolase peptidoglycan, the key composition of the cell-wall of both Gram-negative and Gram-positive bacteria. Destruction of peptidoglycan by lysine could induce lysis and degrade the bacterial cell-wall and biofilm structure [86, 87]. Current studies have shown that phage-derived lysin enzymes can potentially act as antibiofilm development and antibacterial components [81, 88, 89]. It has also been reported that osmotic lysis, independent of bacterial metabolism, can occur upon applying Art-175 lysin against P. aeruginosa biofilms. This phenomenon is often important for the elimination of the bacterial biofilms; owing to lysin, even at low metabolic levels, persistent bacteria inside biofilms can be killed [90].
Some phages cannot produce specific enzymes to permeate and diffuse into the EPS matrix for the biofilms inhibition [91]. Nevertheless, phages are often genetically modified to produce enzymes that can damage the EPS matrix and ease the elimination of biofilms. Lu et al. [80] demonstrated that a modified phage T7 of E. coli intracellularly produced a hydrolase during infection, which could increase the biofilm removal by being released into the extracellular matrix. An eradication rate more than 99% was observed by testing on E. coli biofilms, thus confirming the advantage of using the modified phages.
In phage therapy, individual or cocktail (mixture) phages can be used. The usage of cocktails versus individual phages could considerably increase the host spectrum and decrease the generation of phage-resistant types. The cocktails of phages have been confirmed to be effective in the prevention of the biofilm development and biofilm removal. Antibacterial components such as disinfectants and antibiotics may be applied with phages to increase the efficiency of the biofilm eradication [81].
The combination therapy of phages and antibiotics can increase the treatment effectiveness, as well as inhibiting the resistance to phages without enhancing the toxicity of the antibiotics [79]. A combined usage of phages with antibiotics increases the synergistic activation of these factors in improving the biofilm disruption. Combined therapy of phage T4 and tobramycin on E. coli biofilms has also been done; this could drastically decrease tobramycin-resistant E. coli. The same experiment was reported against P. aeruginosa biofilms by phage PB-1 [92]. Also, the synergistic efficacy of the phage with amoxicillin on the biofilm of Klebsiella pneumoniae B5055 was demonstrated by Bedi et al. [79]. The combined therapy of amoxicillin and phages significantly increased the elimination of the biofilm development in the case K. pneumoniae, as compared to each of the agents alone [79]. Henriksen et al. [93] also demonstrated that sub-MIC concentrations combined ciprofloxacin and phages have a synergic effect on P. aeruginosa biofilm, showing a decrease of about 6 log in the biofilm formation. Depolymerized enzymes in combination with antibiotics could promote the antibacterial efficacy by facilitating the entrance of antibiotics to the biofilm inside. These enzymes could diminish the adherence of EPS matrix and bacteria, thus favoring the function of antibiotics [94].
However, a main limitation of the combined therapy of phages and antibiotics is that antibiotic-resistant biofilms may be increased because phages could favorably infect antibiotic-sensitive bacteria [17]. Furthermore, the interference with bacterial metabolism is needed for the DNA replication and protein synthesis of phages. Since phage therapy is strictly associated to the growth condition of its bacterial host and common antibiotics potentially act on the log phase of bacterial growth, which have the maximum metabolism, it can be regarded as one of the obstacles in combining antibiotics and phage therapy [81, 95, 96]. Therefore, to prevent incompatibility, the potential adverse effects of the combination therapy of antibiotics should be considered. As some pathogenic bacteria prefer to remove competitors, the use of the combination therapy of antibiotics and phage cocktails is especially interesting for the treatment of mixed infections.
Quorum Sensing Inhibitors
Bacterial cell-to-cell communication is known as Quorum sensing (QS); it directly acts in the biofilm development of various bacterial species. This system can control the expression of various pathogenic and virulence genes in the biofilm phase [97, 98]. In this system, small molecules, known as autoinducers, are responsible for the communication of bacteria with each other. The gene expression levels may display major changes when the bacterial density reaches the concentration threshold of autoinducers [99]. Changes in the gene expression levels could affect (induction or suppression) various virulence factors in bacteria; these also include the biofilm production. The changes in the environment surrounding the microorganism can modify the planktonic state to become a biofilm. The gene expression in the planktonic state undergoes many changes during the transition to the biofilm state. Cell surface molecules, specific metabolic pathways, and the production of various factors can all contribute to the bacterial survival under biofilm conditions [44]. Quorum quenchings (QQ) or quorum sensing inhibitors (QSIs) are described as molecules produced by eukaryotes and/or prokaryotes with the ability to prevent the QS systems, which can lead to decreasing the expression of efflux pump genes and the disruption of bacterial biofilms [99]. Several methods have been applied to disturb QS, such as blocking the production of acyl-homoserine lactones (AHLs), diminishing the activity of the AHL synthase, disturbing and inactivating AHLs, and utilizing numerous competitors compounds as the signaling molecules antagonists [100,101,102]. Given that QS controls different phases of the biofilm development, including initial colonization/adhesion, bacterial aggregation, biofilm maturation and cell dispersion, inhibiting QS will prevent the biofilm formation [103]. QSIs are, therefore, applied as a therapeutic agent in treating biofilm infection to inhibit the biofilm formation [104]. Antibiotics, synthetic compounds and natural products may influence the QSIs function [99]. Some synthetic and natural compounds that act as QSIs and reduce the biofilm formation in bacteria are shown in Table 2. Drugs like aspirin, piroxicam and meloxicam, which are in the category of nonsteroidal anti-inflammatory drugs, may be applied as potential inhibitors to control the QS signaling molecules of P. aeruginosa, as well as biofilm development [105,106,107]. Antibiotics such as azithromycin, erythromycin, ciprofloxacin, ceftazidime, gentamicin, tobramycin, piperacillin, spectinomycin and streptomycin display good levels of the QSI activity [108, 109]. Bacteria could be resistant to a single synthetic and natural compound, leading to decreasing the effective activity of them. Therefore, a combined therapy of antibiotics and QSIs is recommended. Consequently, this combined therapy can improve the efficiency of treatment without increasing the antibiotics toxicity and inhibiting the resistance to a single QSI [110, 111]. The combination of Aminoglycoside antibiotics with resveratrol significantly diminishes the biofilm production, as compared with each of the compounds alone [112]. The synergistic effectiveness of curcumin with ceftazidime, ciprofloxacin, gentamicin and azithromycin on the P. aeruginosa QS signaling molecule showed that the sub-MIC of each of the compounds, both alone and in combination, could significantly decline the biofilm development [28, 113]. Zinc oxide (ZnO) nanoparticles, Chitosan and Chitosan-ZnO nanocomposite, in combination with gentamicin, significantly decreased the biofilm formation of both S. aureus and P. aeruginosa, when they were treated with MIC and 1/4 MIC of the compounds [114]. Therefore, targeting QS by various anti-QS agents could be a potential application in the treatment of antibiofilm infections.
Aptamers
Aptamers are peptides or single-stranded oligonucleotide molecules produced in an in vitro procedure called Systematic Evolution of Ligands by Exponential Enrichment (SELEX) [141]. Due to their 3-dimensional constructions, they can be linked with great affinity and specificity to select target molecules including small molecules, proteins, drugs, metal ions, and even whole cells [142, 143]. Such properties of aptamers lead to a broad range of activities as antibiofilm and antibacterial agents [144]. Bacterial cell-wall depolarization may be due to the antibacterial effects of aptamer [145]. Although the available research is limited, several studies have suggested that aptamers could be used as an alternative strategy to inhibit the development of biofilms [144, 146, 147]. Figure 2 shows multiple mechanisms of aptamers for the suppression of the bacterial biofilm formation. The aim of these researches has been selection of aptamers against targets involved in the biofilm development. Thevendran et al. [148] developed a quickly growing field of research by scientists from various scientific branches. The flexibility of aptamers as agents of both diagnostics and therapy has introduced them as good candidates for a broad range of uses. Bacterial flagella are responsible for the initial attachment and motility, which are necessary for the biofilm development. Therefore, inhibiting flagella represents an efficient potential approach to prevent the biofilm development. Ning et al. [142] demonstrated that a specific aptamer targeting the flagella of Salmonella choleraesuis could develop the antibiofilms effect on the inhibition of the bacterial biofilms. Yu et al. [149] also screened the aptamers targeted QS signal molecule that could interfere with the virulence gene expression of P. aeruginosa. Furthermore, Mao et al. [146] reported that aptamer-graphene oxide conjugates displayed excellent antibiofilm properties against Salmonella typhimurium and could serve as a long-term approach to control the bacterial biofilm formation. In addition, Shatila et al. [144] demonstrated that the DNA aptamer (Apt17) targeted invasion protein A of Salmonella enteritidis (SidA) could reduce the biofilm formation, either alone or in combination with ampicillin. Further, Wang et al. [150] revealed that the PA-ap1 aptamer in combination with SWNTs (Single-walled carbon nanotubes) could reduce the biofilm development of P. aeruginosa about 36%, in comparison to the SWNTs alone. In addition, the combination of ciprofloxacin with PA-ap1 aptamer–SWNTs had a greater antibiofilm efficacy than any compound alone. As bacteria accumulation represents a significant phase at the initial step of the biofilm development, aptamer-SWNTs could spontaneously catch and link to bacterial cells, quickly accumulating a greater effective concentration of aptamer-SWNTs around bacterial cells that could interfere with the bacteria accumulation. In addition, it was revealed that aptamer–SWNTs significantly decreased the biofilm formation, in comparison to the SWNTs alone, thus implying the possibility of using the targeted effective aptamers in the removal of the biofilm. Aptamers, as special targeting agents, can be used for the treatment of bacterial infections, which could increase the efficient concentration of antibiotics and decrease the off-target effects. It was reported that the combination of aptamer with antibiotics could lead to attacking more bacterial cells, in comparison to the use of antibiotics alone [142]. Wang et al. [150] also demonstrated that C4-HSL aptamers could efficiently decrease the biofilm development of P. aeruginosa with inhibiting QS, in comparison to that of the untreated groups. In enteropathogenic E. coli (EPEC), the biofilm formation is upregulated using some genes such as cell interaction (lsrA), motility (motB) and curli gene (csgA). Aptamer SELEX 10 Colony 5 could reduce the mRNA level of csgA, lsrA and motB gene, consequently decreasing the biofilm formation of the treated group [151]. Sengupta et al. [152] also reported that aptamer-DNA templated Ag-NC (silver-nanocluster) could act as a sensor, thus creating a novel possibility to detect planktonic and biofilm forms. In addition, Ag-NC aptamer could be potentially effective in inhibiting the biofilm development in P. aeruginosa. Therefore, aptamer and aptamer coupled with excellent agents could provide high target specificity to inhibit the bacterial growth and biofilm development; thus, they could be regarded as ideal strategies for the development of antibacterial and antibiofilm agents.
Nanoparticles
Recently, nanoparticles have received much attention due to their antibacterial and antibiofilm characteristics. Therefore, NPs can be presented as an alternative therapeutic strategy to inhibit the development of the bacterial biofilm [20]. Figure 3 shows different physicochemical interactions between bacterial biofilms and nanoparticles. One of the advantages of NPs is the high surface-area to volume ratio, which can provide a platform for the development of materials with a wide range of chemical, mechanical, magnetic and electrical characteristics [153]. NPs can easily interact with the bacterial cell because of their shape, small size, surface charge, hydrophobicity and high surface-to-volume ratio [154, 155].
NPs with a positive charge prefer to interact with the negatively charged components of the cell walls and plasma membranes of bacteria (polysaccharide, extracellular DNA (eDNA) and proteins). Furthermore, NPs with a negative charge prefer to interact with the positively charged components of the cell-wall, enhancing the bacterial membrane permeability and flowing out of the cytoplasm contents, which can cause the bacterial cell death. In addition, there are several water channels in the bacterial biofilm structure, which can enable bacteria to transport nutrients through these pores. Therefore, NPs can have an antibacterial effect on the biofilm through spreading from these channels [156].
The inhibition of the efflux pumps of bacteria by NPs is another potential mechanism for the antibiofilm activity [157]. Gupta et al. [157] indicated that efflux pumps could play effective roles in the bacterial biofilm formation; as well, regulation of their expression could directly control the antibiotic resistance and development of biofilms. In addition, Padwal et al. [158] showed the combination of antibiotics with NPs could inhibit the activity of efflux pumps, thus suggesting that these compounds could be applied as efflux pump inhibitors. They proposed that NPs could distract efflux kinetics through linking to the active site of efflux pumps and decreasing their activities, thus leading to inhibition of the extrusion of antibiotics outside cells. Similar results have revealed the ZnO nanoparticles could have an inhibitory effect on such efflux pumps as MexAM-OPrM of P. aeruginosa and NorA of S. aureus [159, 160]. Therefore, the downregulation or inhibition of efflux pumps with NPs could be regarded as a potential therapeutic approach to inhibit and/or decrease the bacterial biofilm development.
EPS produced by bacteria play critical roles in the primary attachment of the bacterial cells to the host cell surface and the development of a complex biofilm structure, which could protect biofilms against the classical antibiotics and host-immune system [161, 162]. Production of EPS is one of the critical strategies to protect the bacterial biofilm against the antibiotic activity. Therefore, one of the mains restrictive strategies for the biofilm development is also EPS reduction [25]. Inhibition of the EPS development can be one of the antibiofilm mechanisms of NPs; it has been proved that NPs could reduce the biofilm development by the disruption of the EPS production [155, 163]. In S. aureus biofilm, the chemical compounds of EPS contain polysaccharide, proteins and eDNA. One of the major compounds of the biofilm matrix is Polysaccharide Intercellular Adhesin (PIA), which is a compound with a positive charge. Cytoplasmic proteins and eDNA are positively and negatively charged, respectively [164]. NPs like chitosan NPs, due to having a positive charge, can bind to eDNA with a negative charge, disrupting the biofilm formation with their penetration into the biofilm structure [163]. In contrast, NPs with a negative charge like ZnO NPs can interact with positively charged PIA and modify the permeability of the biofilm structure; then they can easily penetrate into the biofilm matrix, inhibiting the biofilm development by inducing ROS [164,165,166,167]. The biofilm matrix of P. aeruginosa consists of exopolysaccharides and eDNA. In P. aeruginosa, there are three types of exopolysaccharides including alginate, Pel (a glucose-rich polysaccharide polymer encoded by pel operon including pelA-F genes), and Psl (composed of a repeating pentamer containing d-glucose, l-rhamnose, and d-mannose, which is encoded by the psl operon including pslA-L); they perform different roles in the biofilm development [168, 169]. Alginate is a major compound of the biofilm matrix accounting for the cell surfaces binding and biofilm sustainability. Alginate and Pel are negatively and positively charged polysaccharides, respectively [170]. The positive and negative NPs could be charged by binding to alginate and Pel, respectively, which can prevent the attachment of the bacteria to cell surfaces; this is the early stage of biofilm development. These nanoparticles can disrupt the development of biofilm by preventing the bacterial attachment to the cellular surfaces [163, 170, 171]. Messiaen et al. [172] also demonstrated that the negatively charged components of tobramycin-loaded liposome were placed near the bacterial cell clusters of Burkholderia cepacia complex biofilms, while fiber-like structures such as the extracellular DNA became immobilized by the interactions with the positively charged particles of nanospheres in the biofilm matrix.
In addition, owing to the antibacterial activity of NPs, they can reduce the bacterial attachment to surfaces and to themselves, as well as the replication and development of biofilm [154]. Table 3 shows nanoparticles inhibiting the bacterial biofilm formation and development. According to these studies, NPs could be regarded as potential antibiofilm agents as a function of their anti-adhesive activity, bactericidal activity and delivery capability. However, NPs may be toxic, but they could be modified to reduce their toxicity, making them useful for biomedical applications. Therefore, further studies should be directed toward changing the oxidative state and charge density, by applying surface coatings and altering the ability to aggregate. Clinical and nonclinical studies are also needed to determine the safety and tolerance of NPs formulating the potential commercial applications.
Peptide Nucleic Acids
Peptide nucleic acids (PNAs) are structurally similar to DNA or RNA; they are the synthetic analogs of nucleic acid. Their structure is the repetition of N-(2-aminoethyl)-glycine units in nucleotide bases instead of sugar phosphate back bones connected by unnatural pseudodipeptide bonds [182]. Properties of PNAs are a combination of both nucleic acids and peptides, which are chemically similar to protein and structurally similar to nucleic acid, which are relatively stable against enzymes degrading proteins and nucleic acids [182, 183]. Antimicrobial and antibiofilm effects of PNAs are related to their small size and the targeted nucleotide sequences of genes involved in the biofilm formation, in the absence or presence of a linker between the peptide conjugated to PNAs and its position in the structure of peptide-PNA [21]. The major property of PNA is its great affinity and specificity binding. Therefore, it has a good potential to synthesize PNA-based specific antibacterials for particular genes involved in the biofilm formation of the selected bacteria. PNAs demonstrate an enormous potential to inhibit the increase of the resistant bacteria [184]. However, one of the most widespread negative properties of PNAs is the hydrophobicity of its structure, which could be due to its insolubility in the aqueous solution. The impermeability of bacterial membranes to PNA is another important limitation preventing the development of the antibacterial PNA due to the difficulty in finding the effective carriage and carriers of PNA to the bacterial cells [183]. The lipid bilayer, lipopolysaccharide and peptidoglycan of bacteria serve as main barriers to the entrance of PNAs [185].
Different mechanisms have been suggested to increase the transport of PNAs to bacterial cells: chemical changes of the PNAs structure can increase their hydrophilicity. In addition, covalent conjugation of various cell penetrating peptides (CPPs) with PNA [186], such as photochemical internalization (PCI), chloroquine, cationic lipids and HIV-TAT [187], and the use of the tetrahedral DNA nanostructure (TDN) can act a complementary base pair between PNAs and DNA [188].
PNAs can prevent or overcome the bacterial biofilm development. PNAs treated at the efaA gene have a negative influence on the biofilm development in Enterococcus faecalis; this gene serves an important role in the bacterial attachment to surfaces [21]. In addition, Otsuka et al. [189] showed that the acpP gene targeted by PNA-peptides caused antibacterial and antibiofilm effects on both planktonic and biofilm states of Haemophilus influenzae. Binding PNAs to their targeted genes could prevent and interfere with the gene expression. The motA gene could contribute to the downstream adhesion events of attachment in the development of biofilm in P. aeruginosa, playing a critical role in the initial state of the biofilm formation, which encodes the element of the flagellar motor complex [190]. Xia et al. [191] also revealed that PNA-peptides targeting the motA gene could decrease the motA expression and inhibit the biofilm development. PNAs have also been used in combination with classical antibiotics to increase their antibacterial and antibiofilm activity. Castillo et al. [192] showed the synergistic activity of mRNA targeted PNAs in E. coli O157:H7. The synergistic effect of trimethoprim and polymyxin B antibiotics in combination with anti-acpP PNA (encoding acyl carrier protein (AcpP) was determined using the checkerboard titration method. Their results showed that acpP mRNA targeted PNAs could improve the antibacterial activity of antibiotics by suppressing the acpP expression, which is functionally related with antibiotic resistance. In general, designing PNAs targeting specific genes involved in the development and formation of biofilm, as well as targeting critical genes involved in the living and surviving bacteria, could enhance the susceptibility of bacteria and their biofilm.
Future Directions
In the post-antibiotic era, development of novel antibiofilm strategies and alternative therapeutic agents has been of interest to combat the threat of bacterial biofilm infections. In this era, we need more complex biological approaches to investigate the interactions between single- and multi-species bacterial biofilms and their environments in chronic infections. Some of the introduced strategies could only be used to treat single-species biofilms and may not be applicable to mixed-species biofilms. In addition, it is possible for bacteria to protect themselves during the use of these agents; this is a great challenge that should be addressed in regard to combating bacteria and their biofilm formation. Therefore, combination of these agents and flexibility in the use of them could be great promises for future biofilm infection treatments. The applicability of these strategies could be summarized in three steps including entry, delivery and release into the biofilm matrix. Recruitment of alternative and promising therapeutic approaches or strategies could increase the local concentration of antibiotics or antibiofilm agents improve their local accumulation and decrease their systematic toxicity. Nanomedicine, as the most versatile and flexible strategy, can be combined with other novel therapeutic strategies as the state-of-art concepts. There are several tenuous antibiofilm agents and a few of them could be used in clinical relevant models. Therefore, great efforts should be devoted to the evaluation of the biosafety and efficacy of strategies; effective routs for the entry, delivery and release of the agents to the biofilm matrix should be developed; further, the novel agents-biofilm interactions should be studies, and the complexity of prescription and cost of fabrication should be reduced.
Conclusions
Antibiofilm agents such as antiadhesion agents, antimicrobial peptides, aptamers, nanoparticles and peptide nucleic acids represent a promising therapeutic target to treat biofilm infections and develop a strong capability to interfere with different phases of the biofilm development, including adherence, polysaccharide intercellular adhesion (PIA), quorum sensing molecules and cell-to-cell connection, bacterial aggregation, planktonic bacteria killing and host-immune response modulation. Therefore, rational and experimental design approaches allow the synthesis of new antibiofilm compounds with improved and varied biological functions. In addition, these novel strategies can be promising methods against bacterial biofilm due to their improved permeation and targeted delivery of antibacterials inside the biofilm. As well, these components in combination with antibiotics can lead to developing a powerful approach for the treatment of bacterial biofilm-related infections.
References
Høiby, N., Bjarnsholt, T., Moser, C., Bassi, G., Coenye, T., Donelli, G., Hall-Stoodley, L., Holá, V., Imbert, C., & Kirketerp-Møller, K. (2015). ESCMID guideline for the diagnosis and treatment of biofilm infections 2014. Clinical Microbiology & Infection, 21, S1–S25.
Evans, J. J., & Bolz, D. D. (2019). Regulation of virulence and antibiotic resistance in Gram-positive microbes in response to cell wall-active antibiotics. Current Opinion in Infectious Diseases, 32, 217–222.
Khatoon, Z., McTiernan, C. D., Suuronen, E. J., Mah, T.-F., & Alarcon, E. I. (2018). Bacterial biofilm formation on implantable devices and approaches to its treatment and prevention. Heliyon, 4, e01067.
Batoni, G., Maisetta, G., & Esin, S. (2016). Antimicrobial peptides and their interaction with biofilms of medically relevant bacteria. Biochimica et Biophysica Acta (BBA)-Biomembranes, 1858, 1044–1060.
Lynch, A. S., & Robertson, G. T. (2008). Bacterial and fungal biofilm infections. Annual Review of Medicine, 59, 415–428.
Hall-Stoodley, L., Costerton, J. W., & Stoodley, P. (2004). Bacterial biofilms: From the natural environment to infectious diseases. Nature Reviews Microbiology, 2, 95–108.
Costerton, J. W., Cheng, K., Geesey, G. G., Ladd, T. I., Nickel, J. C., Dasgupta, M., & Marrie, T. J. (1987). Bacterial biofilms in nature and disease. Annual Reviews in Microbiology, 41, 435–464.
Lei, M. G., Gupta, R. K., & Lee, C. Y. (2017). Proteomics of Staphylococcus aureus biofilm matrix in a rat model of orthopedic implant-associated infection. PLoS ONE, 12, e0187981.
Watnick, P., & Kolter, R. (2000). Biofilm, city of microbes. Journal of Bacteriology, 182, 2675–2679.
Galdiero, E., Lombardi, L., Falanga, A., Libralato, G., Guida, M., & Carotenuto, R. (2019). Biofilms: Novel strategies based on antimicrobial peptides. Pharmaceutics, 11, 322.
Staudt, C., Horn, H., Hempel, D., & Neu, T. (2004). Volumetric measurements of bacterial cells and extracellular polymeric substance glycoconjugates in biofilms. Biotechnology and Bioengineering, 88, 585–592.
Dibdin, G. H., Assinder, S. J., Nichols, W. W., & Lambert, P. A. (1996). Mathematical model of β-lactam penetration into a biofilm of Pseudomonas aeruginosa while undergoing simultaneous inactivation by released β-lactamases. Journal of Antimicrobial Chemotherapy, 38, 757–769.
Bhagirath, A. Y., Li, Y., Somayajula, D., Dadashi, M., Badr, S., & Duan, K. (2016). Cystic fibrosis lung environment and Pseudomonas aeruginosa infection. BMC Pulmonary Medicine, 16, 1–22.
Ebbensgaard, A., Mordhorst, H., Overgaard, M. T., Nielsen, C. G., Aarestrup, F. M., & Hansen, E. B. (2015). Comparative evaluation of the antimicrobial activity of different antimicrobial peptides against a range of pathogenic bacteria. PLoS ONE, 10, e0144611.
Kåhrström, C. T. (2013). Entering a post-antibiotic era? Nature Reviews Microbiology, 11, 146–146.
Di Somma, A., Moretta, A., Canè, C., Cirillo, A., & Duilio, A. (2020). Antimicrobial and antibiofilm peptides. Biomolecules, 10, 652.
Ferriol-González, C., & Domingo-Calap, P. (2020). Phages for biofilm removal. Antibiotics, 9, 268.
Haque, S., Ahmad, F., Dar, S. A., Jawed, A., Mandal, R. K., Wahid, M., Lohani, M., Khan, S., Singh, V., & Akhter, N. (2018). Developments in strategies for Quorum Sensing virulence factor inhibition to combat bacterial drug resistance. Microbial Pathogenesis, 121, 293–302.
Shatila, F., Yaşa, İ, & Yalçın, H. (2020). Inhibition of Salmonella enteritidis biofilms by Salmonella invasion protein-targeting aptamer. Biotechnology Letters. https://doi.org/10.1007/s10529-020-02920-2
Fulaz, S., Vitale, S., Quinn, L., & Casey, E. (2019). Nanoparticle–biofilm interactions: The role of the EPS matrix. Trends in Microbiology, 27, 915–926.
Narenji, H., Teymournejad, O., Rezaee, M. A., Taghizadeh, S., Mehramuz, B., Aghazadeh, M., Asgharzadeh, M., Madhi, M., Gholizadeh, P., Ganbarov, K., Yousefi, M., Pakravan, A., Dal, T., Ahmadi, R., & Samadi Kafil, H. (2020). Antisense peptide nucleic acids againstftsZ andefaA genes inhibit growth and biofilm formation of Enterococcus faecalis. Microbial Pathogenesis, 139, 103907.
Arias, C. A., & Murray, B. E. (2009). Antibiotic-resistant bugs in the 21st century—A clinical super-challenge. New England Journal of Medicine, 360, 439–443.
Wood, L. F., Leech, A. J., & Ohman, D. E. (2006). Cell wall-inhibitory antibiotics activate the alginate biosynthesis operon in Pseudomonas aeruginosa: roles of σ22 (AlgT) and the AlgW and Prc proteases. Molecular Microbiology, 62, 412–426.
Wiens, J. R., Vasil, A. I., Schurr, M. J., & Vasil, M. L. (2014). Iron-regulated expression of alginate production, mucoid phenotype, and biofilm formation by Pseudomonas aeruginosa. MBio. https://doi.org/10.1128/mBio.01010-13
Bhattacharyya, P., Agarwal, B., Goswami, M., Maiti, D., Baruah, S., & Tribedi, P. (2018). Zinc oxide nanoparticle inhibits the biofilm formation of Streptococcus pneumoniae. Antonie van Leeuwenhoek, 111, 89–99.
Ali, S. G., Ansari, M. A., Alzohairy, M. A., Alomary, M. N., AlYahya, S., Jalal, M., Khan, H. M., Asiri, S. M. M., Ahmad, W., & Mahdi, A. A. (2020). Biogenic gold nanoparticles as potent antibacterial and antibiofilm nano-antibiotics against Pseudomonas aeruginosa. Antibiotics, 9, 100.
Kim, H.-S., Lee, S.-H., Byun, Y., & Park, H.-D. (2015). 6-Gingerol reduces Pseudomonas aeruginosa biofilm formation and virulence via quorum sensing inhibition. Science and Reports, 5, 8656.
Bahari, S., Zeighami, H., Mirshahabi, H., Roudashti, S., & Haghi, F. (2017). Inhibition of Pseudomonas aeruginosa quorum sensing by subinhibitory concentrations of curcumin with gentamicin and azithromycin. Journal of Global Antimicrobial Resistance, 10, 21–28.
De la Fuente-Núñez, C., Reffuveille, F., Fernández, L., & Hancock, R. E. (2013). Bacterial biofilm development as a multicellular adaptation: Antibiotic resistance and new therapeutic strategies. Current Opinion in Microbiology, 16, 580–589.
Donlan, R. M. (2002). Biofilms: Microbial life on surfaces. Emerging Infectious Diseases, 8, 881.
Beloin, C., Valle, J., Latour-Lambert, P., Faure, P., Kzreminski, M., Balestrino, D., Haagensen, J. A., Molin, S., Prensier, G., & Arbeille, B. (2004). Global impact of mature biofilm lifestyle on Escherichia coli K-12 gene expression. Molecular Microbiology, 51, 659–674.
Klausen, M., Aaes-Jørgensen, A., Molin, S., & Tolker-Nielsen, T. (2003). Involvement of bacterial migration in the development of complex multicellular structures in Pseudomonas aeruginosa biofilms. Molecular Microbiology, 50, 61–68.
Klausen, M., Heydorn, A., Ragas, P., Lambertsen, L., Aaes-Jørgensen, A., Molin, S., & Tolker-Nielsen, T. (2003). Biofilm formation by Pseudomonas aeruginosa wild type, flagella and type IV pili mutants. Molecular Microbiology, 48, 1511–1524.
Toutain, C. M., Caizza, N. C., Zegans, M. E., & O’Toole, G. A. (2007). Roles for flagellar stators in biofilm formation by Pseudomonas aeruginosa. Research in Microbiology, 158, 471–477.
Watnick, P. I., & Kolter, R. (1999). Steps in the development of a Vibrio cholerae El Tor biofilm. Molecular Microbiology, 34, 586–595.
Schmidt, J., Müsken, M., Becker, T., Magnowska, Z., Bertinetti, D., Möller, S., Zimmermann, B., Herberg, F. W., Jänsch, L., & Häussler, S. (2011). The Pseudomonas aeruginosa chemotaxis methyltransferase CheR1 impacts on bacterial surface sampling. PLoS ONE, 6, e18184.
Hadjifrangiskou, M., Gu, A. P., Pinkner, J. S., Kostakioti, M., Zhang, E. W., Greene, S. E., & Hultgren, S. J. (2012). Transposon mutagenesis identifies uropathogenic Escherichia coli biofilm factors. Journal of Bacteriology, 194, 6195–6205.
Beloin, C., Roux, A., & Ghigo, J.-M. (2008). Escherichia coli biofilms. In Bacterial biofilms. (pp. 249–289). Springer.
Anderson, G. G., Palermo, J. J., Schilling, J. D., Roth, R., Heuser, J., & Hultgren, S. J. (2003). Intracellular bacterial biofilm-like pods in urinary tract infections. Science, 301, 105–107.
Waksman, G., & Hultgren, S. J. (2009). Structural biology of the chaperone–usher pathway of pilus biogenesis. Nature Reviews Microbiology, 7, 765–774.
Spurbeck, R. R., Stapleton, A. E., Johnson, J. R., Walk, S. T., Hooton, T. M., & Mobley, H. L. (2011). Fimbrial profiles predict virulence of uropathogenic Escherichia coli strains: contribution of ygi and yad fimbriae. Infection and Immunity, 79, 4753–4763.
Thumbikat, P., Berry, R. E., Zhou, G., Billips, B. K., Yaggie, R. E., Zaichuk, T., Sun, T.-T., Schaeffer, A. J., & Klumpp, D. J. (2009). Bacteria-induced uroplakin signaling mediates bladder response to infection. PLoS Pathogens, 5, e1000415.
Cegelski, L., Pinkner, J. S., Hammer, N. D., Cusumano, C. K., Hung, C. S., Chorell, E., Åberg, V., Walker, J. N., Seed, P. C., & Almqvist, F. (2009). Small-molecule inhibitors target Escherichia coli amyloid biogenesis and biofilm formation. Nature Chemical Biology, 5, 913–919.
Klebensberger, J., Birkenmaier, A., Geffers, R., Kjelleberg, S., & Philipp, B. (2009). SiaA and SiaD are essential for inducing autoaggregation as a specific response to detergent stress in Pseudomonas aeruginosa. Environmental Microbiology, 11, 3073–3086.
Najafi, K., Ganbarov, K., Gholizadeh, P., Tanomand, A., Rezaee, M. A., Mahmood, S. S., Asgharzadeh, M., & Kafil, H. S. (2020). Oral cavity infection by Enterococcus faecalis: Virulence factors and pathogenesis. Reviews in Medical Microbiology, 31, 51–60.
Sillanpää, J., Nallapareddy, S. R., Prakash, V. P., Qin, X., Hook, M., Weinstock, G. M., & Murray, B. E. (2008). Identification and phenotypic characterization of a second collagen adhesin, Scm, and genome-based identification and analysis of 13 other predicted MSCRAMMs, including four distinct pilus loci, in Enterococcus faecium. Microbiology (Reading, England), 154, 3199.
Han, Z., Pinkner, J. S., Ford, B., Obermann, R., Nolan, W., Wildman, S. A., Hobbs, D., Ellenberger, T., Cusumano, C. K., & Hultgren, S. J. (2010). Structure-based drug design and optimization of mannoside bacterial FimH antagonists. Journal of Medicinal Chemistry, 53, 4779–4792.
Bouckaert, J., Berglund, J., Schembri, M., De Genst, E., Cools, L., Wuhrer, M., Hung, C. S., Pinkner, J., Slättegård, R., & Zavialov, A. (2005). Receptor binding studies disclose a novel class of high-affinity inhibitors of the Escherichia coli FimH adhesin. Molecular Microbiology, 55, 441–455.
Cusumano, C. K., Pinkner, J. S., Han, Z., Greene, S. E., Ford, B. A., Crowley, J. R., Henderson, J. P., Janetka, J. W., & Hultgren, S. J. (2011). Treatment and prevention of urinary tract infection with orally active FimH inhibitors. Science Translational Medicine, 3, 109ra115.
Chorell, E., Pinkner, J. S., Phan, G., Edvinsson, S., Buelens, F., Remaut, H., Waksman, G., Hultgren, S. J., & Almqvist, F. (2010). Design and synthesis of C-2 substituted thiazolo and dihydrothiazolo ring-fused 2-pyridones: Pilicides with increased antivirulence activity. Journal of Medicinal Chemistry, 53, 5690–5695.
Chorell, E., Bengtsson, C., Banchelin, T.S.-L., Das, P., Uvell, H., Sinha, A. K., Pinkner, J. S., Hultgren, S. J., & Almqvist, F. (2011). Synthesis and application of a bromomethyl substituted scaffold to be used for efficient optimization of anti-virulence activity. European Journal of Medicinal Chemistry, 46, 1103–1116.
Yazici, A., Ortucu, S., Taskin, M., & Marinelli, L. (2018). Natural-based antibiofilm and antimicrobial peptides from micro-organisms. Current Topics in Medicinal Chemistry, 18, 2102–2107.
Ageitos, J., Sánchez-Pérez, A., Calo-Mata, P., & Villa, T. (2017). Antimicrobial peptides (AMPs): Ancient compounds that represent novel weapons in the fight against bacteria. Biochemical Pharmacology, 133, 117–138.
Zhang, L., & Falla, T. J. (2006). Antimicrobial peptides: Therapeutic potential. Expert Opinion on Pharmacotherapy, 7, 653–663.
Yeaman, M. R., & Yount, N. Y. (2003). Mechanisms of antimicrobial peptide action and resistance. Pharmacological Reviews, 55, 27–55.
Jenssen, H., Hamill, P., & Hancock, R. E. (2006). Peptide antimicrobial agents. Clinical Microbiology Reviews, 19, 491–511.
Brogden, K. A. (2005). Antimicrobial peptides: Pore formers or metabolic inhibitors in bacteria? Nature Reviews Microbiology, 3, 238–250.
Cassone, M., Frith, N., Vogiatzi, P., Wade, J. D., & Otvos, L. (2009). Induced resistance to the designer proline-rich antimicrobial peptide A3-APO does not involve changes in the intracellular target DnaK. International Journal of Peptide Research and Therapeutics, 15, 121–128.
Shah, P., Hsiao, F. S. H., Ho, Y. H., & Chen, C. S. (2016). The proteome targets of intracellular targeting antimicrobial peptides. Proteomics, 16, 1225–1237.
Graf, M., & Wilson, D. N. (2019). Intracellular antimicrobial peptides targeting the protein synthesis machinery. In Antimicrobial peptides. (pp. 73–89). Springer.
Okuda, K.-I., Zendo, T., Sugimoto, S., Iwase, T., Tajima, A., Yamada, S., Sonomoto, K., & Mizunoe, Y. (2013). Effects of bacteriocins on methicillin-resistant Staphylococcus aureus biofilm. Antimicrobial Agents and Chemotherapy, 57, 5572–5579.
Overhage, J., Campisano, A., Bains, M., Torfs, E. C., Rehm, B. H., & Hancock, R. E. (2008). Human host defense peptide LL-37 prevents bacterial biofilm formation. Infection and Immunity, 76, 4176–4182.
Brancatisano, F. L., Maisetta, G., Di Luca, M., Esin, S., Bottai, D., Bizzarri, R., Campa, M., & Batoni, G. (2014). Inhibitory effect of the human liver-derived antimicrobial peptide hepcidin 20 on biofilms of polysaccharide intercellular adhesin (PIA)-positive and PIA-negative strains of Staphylococcus epidermidis. Biofouling, 30, 435–446.
Blower, R. J., Barksdale, S. M., & van Hoek, M. L. (2015). Snake cathelicidin NA-CATH and smaller helical antimicrobial peptides are effective against Burkholderia thailandensis. PLoS Neglected Tropical Diseases, 9, e0003862.
Anunthawan, T., De La Fuente-Núñez, C., Hancock, R. E., & Klaynongsruang, S. (2015). Cationic amphipathic peptides KT2 and RT2 are taken up into bacterial cells and kill planktonic and biofilm bacteria. Biochimica et Biophysica Acta (BBA)-Biomembranes, 1848, 1352–1358.
De Brucker, K., Delattin, N., Robijns, S., Steenackers, H., Verstraeten, N., Landuyt, B., Luyten, W., Schoofs, L., Dovgan, B., & Fröhlich, M. (2014). Derivatives of the mouse cathelicidin-related antimicrobial peptide (CRAMP) inhibit fungal and bacterial biofilm formation. Antimicrobial Agents and Chemotherapy, 58, 5395–5404.
Mataraci, E., & Dosler, S. (2012). In vitro activities of antibiotics and antimicrobial cationic peptides alone and in combination against methicillin-resistant Staphylococcus aureus biofilms. Antimicrobial Agents and Chemotherapy, 56, 6366–6371.
Cao, Y., Yin, H., Wang, W., Pei, P., Wang, Y., Wang, X., Jiang, J., Luo, S.-Z., & Chen, L. (2020). Killing Streptococcus mutans in mature biofilm with a combination of antimicrobial and antibiofilm peptides. Amino Acids, 52, 1–14.
Gopal, R., Lee, J. H., Kim, Y. G., Kim, M.-S., Seo, C. H., & Park, Y. (2013). Anti-microbial, anti-biofilm activities and cell selectivity of the NRC-16 peptide derived from witch flounder, Glyptocephalus cynoglossus. Marine Drugs, 11, 1836–1852.
Chen, L., Jia, L., Zhang, Q., Zhou, X., Liu, Z., Li, B., Zhu, Z., Wang, F., Yu, C., & Zhang, Q. (2017). A novel antimicrobial peptide against dental-caries-associated bacteria. Anaerobe, 47, 165–172.
Cooper, V. S., Carlson, W. A., & LiPuma, J. J. (2009). Susceptibility of Caenorhabditis elegans to Burkholderia infection depends on prior diet and secreted bacterial attractants. PLoS ONE, 4, e7961.
Peng, X., Zhang, Y., Bai, G., Zhou, X., & Wu, H. (2016). Cyclic di-AMP mediates biofilm formation. Molecular Microbiology, 99, 945–959.
Pletzer, D., Coleman, S. R., & Hancock, R. E. (2016). Anti-biofilm peptides as a new weapon in antimicrobial warfare. Current Opinion in Microbiology, 33, 35–40.
Ribeiro, S. M., De La Fuente-Núñez, C., Baquir, B., Faria-Junior, C., Franco, O. L., & Hancock, R. E. (2015). Antibiofilm peptides increase the susceptibility of carbapenemase-producing Klebsiella pneumoniae clinical isolates to β-lactam antibiotics. Antimicrobial Agents and Chemotherapy, 59, 3906–3912.
Tong, Z., Zhang, Y., Ling, J., Ma, J., Huang, L., & Zhang, L. (2014). An in vitro study on the effects of nisin on the antibacterial activities of 18 antibiotics against Enterococcus faecalis. PLoS ONE, 9, e89209.
Tong, Z., Zhou, L., Jiang, W., Kuang, R., Li, J., Tao, R., & Ni, L. (2011). An in vitro synergetic evaluation of the use of nisin and sodium fluoride or chlorhexidine against Streptococcus mutans. Peptides, 32, 2021–2026.
Domingo-Calap, P., & Delgado-Martínez, J. (2018). Bacteriophages: Protagonists of a post-antibiotic era. Antibiotics, 7, 66.
Principi, N., Silvestri, E., & Esposito, S. (2019). Advantages and limitations of bacteriophages for the treatment of bacterial infections. Frontiers in Pharmacology, 10, 513.
Bedi, M. S., Verma, V., & Chhibber, S. (2009). Amoxicillin and specific bacteriophage can be used together for eradication of biofilm of Klebsiella pneumoniae B5055. World Journal of Microbiology & Biotechnology, 25, 1145.
Lu, T. K., & Collins, J. J. (2007). Dispersing biofilms with engineered enzymatic bacteriophage. Proceedings of the National Academy of Sciences, 104, 11197–11202.
Łusiak-Szelachowska, M., Weber-Dąbrowska, B., & Górski, A. (2020). Bacteriophages and lysins in biofilm control. Virologica Sinica. https://doi.org/10.1007/s12250-019-00192-3
Wood, S., Kirkham, J., Marsh, P., Shore, R., Nattress, B., & Robinson, C. (2000). Architecture of intact natural human plaque biofilms studied by confocal laser scanning microscopy. Journal of Dental Research, 79, 21–27.
Pires, D. P., Oliveira, H., Melo, L. D., Sillankorva, S., & Azeredo, J. (2016). Bacteriophage-encoded depolymerases: Their diversity and biotechnological applications. Applied Microbiology and Biotechnology, 100, 2141–2151.
Cornelissen, A., Ceyssens, P. J., T’syen, J., Van Praet, H., Noben, J. P., Shaburova, O. V., Krylov, V. N., Volckaert, G., & Lavigne, R. (2011). The T7-related Pseudomonas putida phage φ15 displays virion-associated biofilm degradation properties. PLoS ONE, 6, e18597.
Yoon, S., Choi, Y., Lee, S. Y., Son, J., Jun, S., & Kang, S. (2013) Bacteriophage or lytic protein derived from the bacteriophage which effective for the treatment of Staphylococcus aureus biofilm, Google Patents.
Borysowski, J., Łobocka, M., Międzybrodzki, R., Weber-Dąbrowska, B., & Górski, A. (2011). Potential of bacteriophages and their lysins in the treatment of MRSA. BioDrugs, 25, 347–355.
Fischetti, V. A. (2018). Development of phage lysins as novel therapeutics: A historical perspective. Viruses, 10, 310.
Sharma, U., Vipra, A., & Channabasappa, S. (2018). Phage-derived lysins as potential agents for eradicating biofilms and persisters. Drug Discovery Today, 23, 848–856.
Gray, J. A., Chandry, P. S., Kaur, M., Kocharunchitt, C., Bowman, J. P., & Fox, E. M. (2018). Novel biocontrol methods for Listeria monocytogenes biofilms in food production facilities. Frontiers in Microbiology, 9, 605.
Briers, Y., Walmagh, M., Grymonprez, B., Biebl, M., Pirnay, J.-P., Defraine, V., Michiels, J., Cenens, W., Aertsen, A., & Miller, S. (2014). Art-175 is a highly efficient antibacterial against multidrug-resistant strains and persisters of Pseudomonas aeruginosa. Antimicrobial Agents and Chemotherapy, 58, 3774–3784.
Donlan, R. M. (2009). Preventing biofilms of clinically relevant organisms using bacteriophage. Trends in Microbiology, 17, 66–72.
Coulter, L. B., McLean, R. J., Rohde, R. E., & Aron, G. M. (2014). Effect of bacteriophage infection in combination with tobramycin on the emergence of resistance in Escherichia coli and Pseudomonas aeruginosa biofilms. Viruses, 6, 3778–3786.
Issa, R., Chanishvili, N., Caplin, J., Kakabadze, E., Bakuradze, N., Makalatia, K., & Cooper, I. (2019). Antibiofilm potential of purified environmental bacteriophage preparations against early stage Pseudomonas aeruginosa biofilms. Journal of Applied Microbiology, 126, 1657–1667.
Maciejewska, B., Olszak, T., & Drulis-Kawa, Z. (2018). Applications of bacteriophages versus phage enzymes to combat and cure bacterial infections: An ambitious and also a realistic application? Applied Microbiology and Biotechnology, 102, 2563–2581.
Abedon, S. T. (2019). Phage-antibiotic combination treatments: Antagonistic impacts of antibiotics on the pharmacodynamics of phage therapy? Antibiotics, 8, 182.
Tagliaferri, T. L., Jansen, M., & Horz, H.-P. (2019). Fighting pathogenic bacteria on two fronts: Phages and antibiotics as combined strategy. Frontiers in Cellular and Infection Microbiology, 9, 22.
Fuqua, C., Parsek, M. R., & Greenberg, E. P. (2001). Regulation of gene expression by cell-to-cell communication: Acyl-homoserine lactone quorum sensing. Annual Review of Genetics, 35, 439–468.
Dijkshoorn, L., Nemec, A., & Seifert, H. (2007). An increasing threat in hospitals: Multidrug-resistant Acinetobacter baumannii. Nature Reviews Microbiology, 5, 939–951.
Hemmati, F., Salehi, R., Ghotaslou, R., Kafil, H. S., Hasani, A., Gholizadeh, P., Nouri, R., & Rezaee, M. A. (2020). Quorum Quenching: A potential target for antipseudomonal therapy. Infection and Drug Resistance, 13, 2989–3005.
Kalia, V. C. (2013). Quorum sensing inhibitors: An overview. Biotechnology Advances, 31, 224–245.
Dembitsky, V. M., Al Quntar, A. A. A., & Srebnik, M. (2011). Natural and synthetic small boron-containing molecules as potential inhibitors of bacterial and fungal quorum sensing. Chemical Reviews, 111, 209–237.
Kaufmann, G. F., Sartorio, R., Lee, S.-H., Mee, J. M., Altobell, L. J., Kujawa, D. P., Jeffries, E., Clapham, B., Meijler, M. M., & Janda, K. D. (2006). Antibody interference with N-acyl homoserine lactone-mediated bacterial quorum sensing. Journal of the American Chemical Society, 128, 2802–2803.
Bassler, B. L. (2002). Small talk: Cell-to-cell communication in bacteria. Cell, 109, 421–424.
Taylor, P. K., Yeung, A. T., & Hancock, R. E. (2014). Antibiotic resistance in Pseudomonas aeruginosa biofilms: Towards the development of novel anti-biofilm therapies. Journal of Biotechnology, 191, 121–130.
de Almeida, F. A., Vargas, E. L. G., Carneiro, D. G., Pinto, U. M., & Vanetti, M. C. D. (2018). Virtual screening of plant compounds and nonsteroidal anti-inflammatory drugs for inhibition of quorum sensing and biofilm formation in Salmonella. Microbial Pathogenesis, 121, 369–388.
Yang, E., Lu, Y., Xu, Y., Liang, Q., Wang, C., Wang, H., & Shen, H. (2014). Recombinant BCG coexpressing Ag85B, ESAT-6 and Rv3620c elicits specific Th1 immune responses in C57BL/6 mice. Microbial Pathogenesis, 69, 53–59.
Soheili, V., Bazzaz, B. S. F., Abdollahpour, N., & Hadizadeh, F. (2015). Investigation of Pseudomonas aeruginosa quorum-sensing signaling system for identifying multiple inhibitors using molecular docking and structural analysis methodology. Microbial Pathogenesis, 89, 73–78.
Skindersoe, M. E., Alhede, M., Phipps, R., Yang, L., Jensen, P. O., Rasmussen, T. B., Bjarnsholt, T., Tolker-Nielsen, T., Høiby, N., & Givskov, M. (2008). Effects of antibiotics on quorum sensing in Pseudomonas aeruginosa. Antimicrobial Agents and Chemotherapy, 52, 3648–3663.
Sofer, D., Gilboa-Garber, N., Belz, A., & Garber, N. C. (1999). ‘Subinhibitory’erythromycin represses production of Pseudomonas aeruginosa lectins, autoinducer and virulence factors. Chemotherapy, 45, 335–341.
Dong, Y.-H., & Zhang, L.-H. (2005). Quorum sensing and quorum-quenching enzymes. The Journal of Microbiology, 43, 101–109.
Chanda, S., & Rakholiya, K. (2011). Combination therapy: Synergism between natural plant extracts and antibiotics against infectious diseases. Microbiology Book Series, 1, 520–529.
Zhou, J.-W., Chen, T.-T., Tan, X.-J., Sheng, J.-Y., & Jia, A.-Q. (2018). Can the quorum sensing inhibitor resveratrol function as an aminoglycoside antibiotic accelerant against Pseudomonas aeruginosa? International Journal of Antimicrobial Agents, 52, 35–41.
Roudashti, S., Zeighami, H., Mirshahabi, H., Bahari, S., Soltani, A., & Haghi, F. (2017). Synergistic activity of sub-inhibitory concentrations of curcumin with ceftazidime and ciprofloxacin against Pseudomonas aeruginosa quorum sensing related genes and virulence traits. World Journal of Microbiology & Biotechnology, 33, 50.
Hemmati, F., Salehi, R., Ghotaslou, R., Kafil, H. S., Hasani, A., Gholizadeh, P., & Rezaee, M. A. (2020). The assessment of antibiofilm activity of chitosan-zinc oxide-gentamicin nanocomposite on Pseudomonas aeruginosa and Staphylococcus aureus. International Journal of Biological Macromolecules, 163, 2248–2258.
de Nys, R., Wright, A. D., König, G. M., & Sticher, O. (1993). New halogenated furanones from the marine alga Delisea pulchra (cf. fimbriata). Tetrahedron, 49, 11213–11220.
De Nys, R., Givskov, M., Kumar, N., Kjelleberg, S., & Steinberg, P. (2006). Furanones. In Antifouling compounds. (pp. 55–86). Springer.
Rasmussen, T. B., Skindersoe, M. E., Bjarnsholt, T., Phipps, R. K., Christensen, K. B., Jensen, P. O., Andersen, J. B., Koch, B., Larsen, T. O., & Hentzer, M. (2005). Identity and effects of quorum-sensing inhibitors produced by Penicillium species. Microbiology, 151, 1325–1340.
Musthafa, K. S., Ravi, A. V., Annapoorani, A., Packiavathy, I. S. V., & Pandian, S. K. (2010). Evaluation of anti-quorum-sensing activity of edible plants and fruits through inhibition of the N-acyl-homoserine lactone system in Chromobacterium violaceum and Pseudomonas aeruginosa. Chemotherapy, 56, 333–339.
Girennavar, B., Cepeda, M. L., Soni, K. A., Vikram, A., Jesudhasan, P., Jayaprakasha, G., Pillai, S. D., & Patil, B. S. (2008). Grapefruit juice and its furocoumarins inhibits autoinducer signaling and biofilm formation in bacteria. International Journal of Food Microbiology, 125, 204–208.
Adonizio, A., Kong, K.-F., & Mathee, K. (2008). Inhibition of quorum sensing-controlled virulence factor production in Pseudomonas aeruginosa by South Florida plant extracts. Antimicrobial Agents and Chemotherapy, 52, 198–203.
Vandeputte, O. M., Kiendrebeogo, M., Rajaonson, S., Diallo, B., Mol, A., El Jaziri, M., & Baucher, M. (2010). Identification of catechin as one of the flavonoids from Combretum albiflorum bark extract that reduces the production of quorum-sensing-controlled virulence factors in Pseudomonas aeruginosa PAO1. Applied and Environment Microbiology, 76, 243–253.
Zhou, L., Zheng, H., Tang, Y., Yu, W., & Gong, Q. (2013). Eugenol inhibits quorum sensing at sub-inhibitory concentrations. Biotechnology Letters, 35, 631–637.
Packiavathy, I. A. S. V., Priya, S., Pandian, S. K., & Ravi, A. V. (2014). Inhibition of biofilm development of uropathogens by curcumin—An anti-quorum sensing agent from Curcuma longa. Food Chemistry, 148, 453–460.
Burt, S. A., Ojo-Fakunle, V. T., Woertman, J., & Veldhuizen, E. J. (2014). The natural antimicrobial carvacrol inhibits quorum sensing in Chromobacterium violaceum and reduces bacterial biofilm formation at sub-lethal concentrations. PLoS ONE, 9, e93414.
Pejin, B., Ciric, A., Glamoclija, J., Nikolic, M., & Sokovic, M. (2015). In vitro anti-quorum sensing activity of phytol. Natural Product Research, 29, 374–377.
Husain, F. M., Ahmad, I., Khan, M. S., Ahmad, E., Tahseen, Q., Khan, M. S., & Alshabib, N. A. (2015). Sub-MICs of Mentha piperita essential oil and menthol inhibits AHL mediated quorum sensing and biofilm of Gram-negative bacteria. Frontiers in Microbiology, 6, 420.
Kumar, L., Chhibber, S., Kumar, R., Kumar, M., & Harjai, K. (2015). Zingerone silences quorum sensing and attenuates virulence of Pseudomonas aeruginosa. Fitoterapia, 102, 84–95.
Luo, J., Dong, B., Wang, K., Cai, S., Liu, T., Cheng, X., Lei, D., Chen, Y., Li, Y., & Kong, J. (2017). Baicalin inhibits biofilm formation, attenuates the quorum sensing-controlled virulence and enhances Pseudomonas aeruginosa clearance in a mouse peritoneal implant infection model. PLoS ONE, 12, e0176883.
Li, Y., Huang, J., Li, L., & Liu, L. (2017). Synergistic activity of berberine with azithromycin against Pseudomonas aeruginosa isolated from patients with cystic fibrosis of lung in vitro and in vivo. Cellular Physiology and Biochemistry, 42, 1657–1669.
Vasavi, H. S., Arun, A. B., & Rekha, P. D. (2014). Anti-quorum sensing activity of Psidium guajava L. flavonoids against Chromobacterium violaceum and Pseudomonas aeruginosa PAO1. Microbiology and Immunology, 58, 286–293.
Jakobsen, T. H., Warming, A. N., Vejborg, R. M., Moscoso, J. A., Stegger, M., Lorenzen, F., Rybtke, M., Andersen, J. B., Petersen, R., & Andersen, P. S. (2017). A broad range quorum sensing inhibitor working through sRNA inhibition. Science and Reports, 7, 1–12.
Ilic-Tomić, T., Soković, M., Vojnović, S., Ćirić, A. D., Veljić, M., Nikodinović-Runić, J., & Novaković, M. M. (2017). Diarylheptanoids from Alnus viridis ssp viridis and Alnus glutinosa: Modulation of quorum sensing activity in Pseudomonas aeruginosa. Planta Medica, 83, 117–125.
Nafee, N., Husari, A., Maurer, C. K., Lu, C., de Rossi, C., Steinbach, A., Hartmann, R. W., Lehr, C.-M., & Schneider, M. (2014). Antibiotic-free nanotherapeutics: Ultra-small, mucus-penetrating solid lipid nanoparticles enhance the pulmonary delivery and anti-virulence efficacy of novel quorum sensing inhibitors. Journal of Controlled Release, 192, 131–140.
Geske, G. D., Wezeman, R. J., Siegel, A. P., & Blackwell, H. E. (2005). Small molecule inhibitors of bacterial quorum sensing and biofilm formation. Journal of the American Chemical Society, 127, 12762–12763.
Hentzer, M., Riedel, K., Rasmussen, T. B., Heydorn, A., Andersen, J. B., Parsek, M. R., Rice, S. A., Eberl, L., Molin, S., & Høiby, N. (2002). Inhibition of quorum sensing in Pseudomonas aeruginosa biofilm bacteria by a halogenated furanone compound. Microbiology, 148, 87–102.
Lönn-Stensrud, J., Petersen, F., Benneche, T., & Scheie, A. A. (2007). Synthetic bromated furanone inhibits autoinducer-2-mediated communication and biofilm formation in oral streptococci. Oral Microbiology and Immunology, 22, 340–346.
O’Loughlin, C. T., Miller, L. C., Siryaporn, A., Drescher, K., Semmelhack, M. F., & Bassler, B. L. (2013). A quorum-sensing inhibitor blocks Pseudomonas aeruginosa virulence and biofilm formation. Proceedings of the National Academy of Sciences, 110, 17981–17986.
Ishii, S., Fukui, K., Yokoshima, S., Kumagai, K., Beniyama, Y., Kodama, T., Fukuyama, T., Okabe, T., Nagano, T., & Kojima, H. (2017). High-throughput screening of small molecule inhibitors of the Streptococcus quorum-sensing signal pathway. Science and Reports, 7, 1–10.
Heidari, A., Noshiranzadeh, N., Haghi, F., & Bikas, R. (2017). Inhibition of quorum sensing related virulence factors of Pseudomonas aeruginosa by pyridoxal lactohydrazone. Microbial Pathogenesis, 112, 103–110.
Heidari, A., Haghi, F., Noshiranzadeh, N., & Bikas, R. (2017). (S, E)-2-hydroxy-N-(2-hydroxy-5-nitrobenzylidene) propane hydrazide as a quorum sensing inhibitor of Pseudomonas aeruginosa. Medicinal Chemistry Research, 26, 1947–1955.
Alizadeh, N., Memar, M., Mehramuz, B., Abibiglou, S., Hemmati, F., & Samadi Kafil, H. (2018). Current advances in aptamer-assisted technologies for detecting bacterial and fungal toxins. Journal of Applied Microbiology, 124, 644–651.
Ning, Y., Cheng, L., Ling, M., Feng, X., Chen, L., Wu, M., & Deng, L. (2015). Efficient suppression of biofilm formation by a nucleic acid aptamer. Pathogens and Disease, 73, ftv034.
Nimjee, S. M., Rusconi, C. P., & Sullenger, B. A. (2005). Aptamers: An emerging class of therapeutics. Annual Review of Medicine, 56, 555–583.
Shatila, F., Yaşa, İ, & Yalçın, H. T. (2020). Inhibition of Salmonella enteritidis biofilms by Salmonella invasion protein-targeting aptamer. Biotechnology Letters. https://doi.org/10.1007/s10529-020-02920-2
Kolovskaya, O. S., Savitskaya, A. G., Zamay, T. N., Reshetneva, I. T., Zamay, G. S., Erkaev, E. N., Wang, X., Wehbe, M., Salmina, A. B., & Perianova, O. V. (2013). Development of bacteriostatic DNA aptamers for salmonella. Journal of Medicinal Chemistry, 56, 1564–1572.
Mao, B., Cheng, L., Wang, S., Zhou, J., & Deng, L. (2018). Combat biofilm by bacteriostatic aptamer-functionalized graphene oxide. Biotechnology and Applied Biochemistry, 65, 355–361.
Lijuan, C., Xing, Y., Minxi, W., Wenkai, L., & Le, D. (2017). Development of an aptamer-ampicillin conjugate for treating biofilms. Biochemical and Biophysical Research Communications, 483, 847–854.
Thevendran, R., Sarah, S., Tang, T.-H., & Citartan, M. (2020). Strategies to bioengineer aptamer-driven nanovehicles as exceptional molecular tools for targeted therapeutics: A review. Journal of Controlled Release. https://doi.org/10.1016/j.jconrel.2020.04.051
Yu, Y. M., Xu, B. Y., Yan, S. S., Xu, J. F., Liu, F., Li, G. M., Ding, Y. L., & Wu, S. Q. (2013). Screening and anti-virulent study of N-acyl homoserine lactones DNA aptamers against Pseudomonas aeruginosa quorum sensing. Biotechnology and Bioprocess Engineering, 18, 406–412.
Wang, S., Mao, B., Wu, M., Liang, J., & Deng, L. (2018). Influence of aptamer-targeted antibiofilm agents for treatment of Pseudomonas aeruginosa biofilms. Antonie van Leeuwenhoek, 111, 199–208.
Oroh, S. B., Mustopa, A. Z., Budiarti, S., & Budiarto, B. R. (2020). Inhibition of enteropathogenic Escherichia coli biofilm formation by DNA aptamer. Molecular Biology Reports. https://doi.org/10.1007/s11033-020-05822-8
Sengupta, B., Adhikari, P., Mallet, E., Havner, R., & Pradhan, P. (2020). Spectroscopic study on Pseudomonas aeruginosa biofilm in the presence of the Aptamer-DNA scaffolded silver nanoclusters. Molecules, 25, 3631.
Whitesides, G. M. (2005). Nanoscience, nanotechnology, and chemistry. Small (Weinheim an der Bergstrasse, Germany), 1, 172–179.
Chen, M., Yu, Q., & Sun, H. (2013). Novel strategies for the prevention and treatment of biofilm related infections. International Journal of Molecular Sciences, 14, 18488–18501.
Mahamuni-Badiger, P. P., Patil, P. M., Badiger, M. V., Patel, P. R., Thorat-Gadgil, B. S., Pandit, A., & Bohara, R. A. (2019). Biofilm formation to inhibition: Role of zinc oxide-based nanoparticles. Materials Science and Engineering: C, 108, 110319.
Shi, S.-F., Jia, J.-F., Guo, X.-K., Zhao, Y.-P., Chen, D.-S., Guo, Y.-Y., & Zhang, X.-L. (2016). Reduced Staphylococcus aureus biofilm formation in the presence of chitosan-coated iron oxide nanoparticles. International JOURNAL of Nanomedicine, 11, 6499.
Gupta, D., Singh, A., & Khan, A. U. (2017). Nanoparticles as efflux pump and biofilm inhibitor to rejuvenate bactericidal effect of conventional antibiotics. Nanoscale Research Letters, 12, 1–6.
Padwal, P., Bandyopadhyaya, R., & Mehra, S. (2014). Polyacrylic acid-coated iron oxide nanoparticles for targeting drug resistance in mycobacteria. Langmuir, 30, 15266–15276.
Banoee, M., Seif, S., Nazari, Z. E., Jafari-Fesharaki, P., Shahverdi, H. R., Moballegh, A., Moghaddam, K. M., & Shahverdi, A. R. (2010). ZnO nanoparticles enhanced antibacterial activity of ciprofloxacin against Staphylococcus aureus and Escherichia coli. Journal of Biomedical Materials Research Part B: Applied Biomaterials, 93, 557–561.
Nallathamby, P. D., Lee, K. J., Desai, T., & Xu, X.-H.N. (2010). Study of the multidrug membrane transporter of single living Pseudomonas aeruginosa cells using size-dependent plasmonic nanoparticle optical probes. Biochemistry, 49, 5942–5953.
Pérez-Laguna, V., García-Luque, I., Ballesta, S., Pérez-Artiaga, L., Lampaya-Pérez, V., Samper, S., Soria-Lozano, P., Rezusta, A., & Gilaberte, Y. (2018). Antimicrobial photodynamic activity of Rose Bengal, alone or in combination with Gentamicin, against planktonic and biofilm Staphylococcus aureus. Photodiagnosis and Photodynamic Therapy, 21, 211–216.
Khan, F., Lee, J.-W., Manivasagan, P., Pham, D. T. N., Oh, J., & Kim, Y.-M. (2019). Synthesis and characterization of chitosan oligosaccharide-capped gold nanoparticles as an effective antibiofilm drug against the Pseudomonas aeruginosa PAO1. Microbial Pathogenesis, 135, 103623.
Wang, Z., Bai, H., Lu, C., Hou, C., Qiu, Y., Zhang, P., Duan, J., & Mu, H. (2019). Light controllable chitosan micelles with ROS generation and essential oil release for the treatment of bacterial biofilm. Carbohydrate Polymers, 205, 533–539.
Kavanaugh, J. S., Flack, C. E., Lister, J., Ricker, E. B., Ibberson, C. B., Jenul, C., Moormeier, D. E., Delmain, E. A., Bayles, K. W., & Horswill, A. R. (2019). Identification of extracellular DNA-binding proteins in the biofilm matrix. MBio, 10, e01137–e1119.
Xia, T., Kovochich, M., Liong, M., Madler, L., Gilbert, B., Shi, H., Yeh, J. I., Zink, J. I., & Nel, A. E. (2008). Comparison of the mechanism of toxicity of zinc oxide and cerium oxide nanoparticles based on dissolution and oxidative stress properties. ACS Nano, 2, 2121–2134.
Zhang, L., Jiang, Y., Ding, Y., Povey, M., & York, D. (2007). Investigation into the antibacterial behaviour of suspensions of ZnO nanoparticles (ZnO nanofluids). Journal of Nanoparticle Research, 9, 479–489.
Ebrahimzadeh, S., Bari, M. R., Hamishehkar, H., Kafil, H. S., & Lim, L.-T. (2021). Essential oils-loaded electrospun chitosan-poly(vinyl alcohol) nonwovens laminated on chitosan film as bilayer bioactive edible films. LWT, 144, 111217.
Ma, L., Conover, M., Lu, H., Parsek, M. R., Bayles, K., & Wozniak, D. J. (2009). Assembly and development of the Pseudomonas aeruginosa biofilm matrix. PLoS Pathogens, 5, e1000354.
Whitchurch, C. B., Tolker-Nielsen, T., Ragas, P. C., & Mattick, J. S. (2002). Extracellular DNA required for bacterial biofilm formation. Science, 295, 1487–1487.
Kovach, K., Fleming, D., Rumbaugh, K. P., & Gordon, V. (2019). Specific disruption of established P. aeruginosa biofilms using polymer-attacking enzymes. bioRxiv, 598979.
Zhang, L., Jiang, Y., Ding, Y., Daskalakis, N., Jeuken, L., Povey, M., O’Neill, A. J., & York, D. W. (2010). Mechanistic investigation into antibacterial behaviour of suspensions of ZnO nanoparticles against E. coli. Journal of Nanoparticle Research, 12, 1625–1636.
Messiaen, A.-S., Forier, K., Nelis, H., Braeckmans, K., & Coenye, T. (2013). Transport of nanoparticles and tobramycin-loaded liposomes in Burkholderia cepacia complex biofilms. PLoS ONE, 8, e79220.
Pham, D. T. N., Khan, F., Phan, T. T. V., Park, S.-K., Manivasagan, P., Oh, J., & Kim, Y.-M. (2019). Biofilm inhibition, modulation of virulence and motility properties by FeOOH nanoparticle in Pseudomonas aeruginosa. Brazilian Journal of Microbiology, 50, 791–805.
Loo, C. Y., Rohanizadeh, R., Young, P. M., Traini, D., Cavaliere, R., Whitchurch, C. B., & Lee, W. H. (2016). Combination of silver nanoparticles and curcumin nanoparticles for enhanced anti-biofilm activities. Journal of Agriculture and Food Chemistry, 64, 2513–2522.
Ghotaslou, R., Bahari, Z., Aliloo, A., Gholizadeh, P., & Eshlaghi, B. S. (2017). The in vitro effects of silver nanoparticles on bacterial biofilms. Journal of Microbiology, Biotechnology and Food Sciences, 6, 1077–1080.
Mu, H., Tang, J., Liu, Q., Sun, C., Wang, T., & Duan, J. (2016). Potent antibacterial nanoparticles against biofilm and intracellular bacteria. Science and Reports, 6, 1–9.
Lin, W.-T., Tan, H.-L., Duan, Z.-L., Yue, B., Ma, R., He, G., & Tang, T.-T. (2014). Inhibited bacterial biofilm formation and improved osteogenic activity on gentamicin-loaded titania nanotubes with various diameters. International Journal of Nanomedicine, 9, 1215.
Khan, F., Manivasagan, P., Lee, J.-W., Pham, D. T. N., Oh, J., & Kim, Y.-M. (2019). Fucoidan-stabilized gold nanoparticle-mediated biofilm inhibition, attenuation of virulence and motility properties in Pseudomonas aeruginosa PAO1. Marine Drugs, 17, 208.
Ali, S. G., Ansari, M. A., Khan, H. M., Jalal, M., Mahdi, A. A., & Cameotra, S. S. (2017). Crataeva nurvala nanoparticles inhibit virulence factors and biofilm formation in clinical isolates of Pseudomonas aeruginosa. Journal of Basic Microbiology, 57, 193–203.
Huang, J., Liu, Y., Yang, L., & Zhou, F. (2019). Synthesis of sulfonated chitosan and its antibiofilm formation activity against E. coli and S. aureus. International Journal of Biological Macromolecules, 129, 980–988.
Khan, S. T., Ahmad, J., Ahamed, M., Musarrat, J., & Al-Khedhairy, A. A. (2016). Zinc oxide and titanium dioxide nanoparticles induce oxidative stress, inhibit growth, and attenuate biofilm formation activity of Streptococcus mitis. JBIC Journal of Biological Inorganic Chemistry, 21, 295–303.
Li, X., Wong, C.-H., Ng, T.-W., Zhang, C.-F., Leung, K.C.-F., & Jin, L. (2016). The spherical nanoparticle-encapsulated chlorhexidine enhances anti-biofilm efficiency through an effective releasing mode and close microbial interactions. International Journal of Nanomedicine, 11, 2471.
Wojciechowska, M., Równicki, M., Mieczkowski, A., Miszkiewicz, J., & Trylska, J. (2020). Antibacterial peptide nucleic acids—Facts and perspectives. Molecules, 25, 559.
Lee, H. T., Kim, S. K., & Yoon, J. W. (2019). Antisense peptide nucleic acids as a potential anti-infective agent. Journal of Microbiology, 57, 423–430.
Kurupati, P., Tan, K. S. W., Kumarasinghe, G., & Poh, C. L. (2007). Inhibition of gene expression and growth by antisense peptide nucleic acids in a multiresistant β-lactamase-producing Klebsiella pneumoniae strain. Antimicrobial Agents and Chemotherapy, 51, 805–811.
Barkowsky, G., Lemster, A.-L., Pappesch, R., Jacob, A., Krüger, S., Schröder, A., Kreikemeyer, B., & Patenge, N. (2019). Influence of different cell-penetrating peptides on the antimicrobial efficiency of PNAs in Streptococcus pyogenes. Molecular Therapy-Nucleic Acids, 18, 444–454.
Narenji, H., Gholizadeh, P., Aghazadeh, M., Rezaee, M. A., Asgharzadeh, M., & Kafil, H. S. (2017). Peptide nucleic acids (PNAs): Currently potential bactericidal agents. Biomedicine & Pharmacotherapy, 93, 580–588.
Readman, J. B., Dickson, G., & Coldham, N. G. (2017). Tetrahedral DNA nanoparticle vector for intracellular delivery of targeted peptide nucleic acid antisense agents to restore antibiotic sensitivity in cefotaxime-resistant Escherichia coli. Nucleic Acid Therapeutics, 27, 176–181.
Otsuka, T., Brauer, A. L., Kirkham, C., Sully, E. K., Pettigrew, M. M., Kong, Y., Geller, B. L., & Murphy, T. F. (2016). Antimicrobial activity of antisense peptide–peptide nucleic acid conjugates against non-typeable Haemophilus influenzae in planktonic and biofilm forms. Journal of Antimicrobial Chemotherapy, 72, 137–144.
Doyle, T. B., Hawkins, A. C., & McCarter, L. L. (2004). The complex flagellar torque generator of Pseudomonas aeruginosa. Journal of Bacteriology, 186, 6341–6350.
Xia, Y., Xiong, Y., Li, X., & Su, X. (2011). Inhibition of biofilm formation by the antisense peptide nucleic acids targeted at the motA gene in Pseudomonas aeruginosa PAO1 strain. World Journal of Microbiology & Biotechnology, 27, 1981–1987.
Castillo, J. I., Równicki, M., Wojciechowska, M., & Trylska, J. (2018). Antimicrobial synergy between mRNA targeted peptide nucleic acid and antibiotics in E. coli. Bioorganic & Medicinal Chemistry Letters, 28, 3094–3098.
Author information
Authors and Affiliations
Contributions
FH: Conceptualization; Data curation; Formal analysis; Investigation; Methodology; Roles/Writing—original draft; Writing—review & editing. MAR, SE, LY, RN & HSK: Data curation; Roles/Writing—original draft; Writing—review & editing. PG: Project administration; Resources; Software; Supervision; Validation; Visualization; Writing—review & editing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Hemmati, F., Rezaee, M.A., Ebrahimzadeh, S. et al. Novel Strategies to Combat Bacterial Biofilms. Mol Biotechnol 63, 569–586 (2021). https://doi.org/10.1007/s12033-021-00325-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-021-00325-8