Abstract
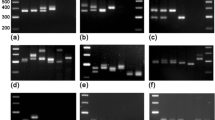
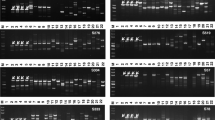
Dwarf bunt of wheat, caused by Tilletia controversa Kühn, is a destructive disease on wheat as well as an important internationally quarantined disease in many countries. The primer ISSR818 generated a polymorphic pattern displaying a 867-bp DNA fragment specific for T. controversa. The marker was converted into a sequence characterized amplified region (SCAR), and specific primers (TCKSF3/TCKSR3) designed for use in PCR detection assays; they amplified a unique DNA fragment in all isolates of T. controversa but not in the related pathogens. The detection limit with the primer set (TCKSF3/TCKSR3) was 5 ng of DNA which could be obtained from 5.5 μg of teliospores in a 25-μL PCR reaction mixture.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Abbreviations
- AFLP:
-
amplified fragment length polymorphism
- CAPS:
-
cleavage amplified polymorphic sequence
- ISSR:
-
inter-simple sequence repeat
- ITS:
-
internal transcribed spacer
- PCR:
-
polymerase chain reaction
- RAPD:
-
random amplification of polymorphic DNA
- RFLP:
-
restriction fragment length polymorphism
- SCAR:
-
sequence characterized amplified region
- SSR:
-
simple sequence repeat
References
Agarwal M., Shrivastava N., Padh H.: Advances in molecular marker techniques and their applications in plant sciences. Plant Cell Rep.27, 617–631 (2008).
Bakkeren G., Kronstad J.W.: Linkage of mating-type loci distinguishes bipolar from tetrapolar mating in basidiomycetous smut fungi. Proc.Nat.Acad.Sci.USA91, 7085–7089 (1994).
Bauer R., Oberwinkler F., Vánky K.: Ultrastructural markers and systematics in smut fungi and allied taxa. Can.J.Bot.75, 1273–1314 (1997).
Bautista R., Crespillo R., Cánovas F.M., Claros M.G.: Identification of olive tree cultivars with SCAR markers. Euphytica129, 33–41 (2003).
Bornet B., Branchard M.: Nonanchored inter simple sequence repeat (ISSR) markers: reproducible and specific tools for genome fingerprinting. Plant Mol.Biol.Reprod.19, 209–215 (2001).
Burgess T., Wingfield M.J., Wingfield B.W.: Simple sequence repeat markers distinguish among morphotypes of Sphaeropsis sapinea. Appl.Environ.Microbiol.67, 354–362 (2001).
Charlesworth B., Sniegowski P., Stephan W.: The evolution dynamics of repetitive DNA in eukaryotes. Nature371, 215–220 (1994).
Dongre A., Parkhi V.: Identification of cotton hybrid through the combination of PCR based RAPD, ISSR and microsatellite markers. J.Plant Biochem.Biotechnol.14, 53–55 (2005).
Duran R., Fischer G.W.: The Genus Tilletia. Washington State University Press, Pullman (WA, USA) 1961.
Eibel P., Wolf G.A., Koch E.: Detection of Tilletia caries, causal agent of common bunt of wheat, by ELISA and PCR. J.Phytopathol.153, 297–306 (2005).
ELENA ESTRADA M., CAMACHO M.V., BENITO C.: The molecular diversity of different isolates of Beauveria bassiana (BALS) VUILL. as assessed using intermicrosatellites (ISSRs). Cell Mol.Biol.Lett.12, 240–252 (2007).
Errampalli D., Saunders J., Cullen D.: A PCR-based method for detection of potato pathogen Helminthosoprium solani, in silver scurf infected tuber tissue and soils. J.Microbiol.Meth.44, 59–68 (2001).
Frederick R.D., Synder K.E., Tooley P.W., Berthier-Schaad Y., Peterson G.L., Bonde M.R., Schaad N.W., Knorr D.A: An improved PCR method utilizing TagMan for the detection and differentiation of Tilletia indica, the causal organism of Karnal bunt of wheat, and a related ryegrass smut. Phytopathology88, S29 (1998).
Gang D.R., Weber D.J.: Preparation of genetic DNA for RAPD analysis from thick-walled dormant teliospores of Tilletia species. Biotechniques19, 93–97 (1995).
Garg A., Singh A.K., Parbhu K.V., Mohapatra T., Tyagi N.K., Nandakumar N., Singh R., Zaman F.U.: Utility of a fertility restorer gene linked marker for testing genetic purity of hybrid seeds in rice (Oryza sativa L.). Seed Sci.Technol.34, 9–18 (2006).
Goates J.B.: Common bunt and dwarf bunt, pp. 12–25 in Bunt and Smut Diseases of Wheat: Concept and methods of disease management (R.D. Wilcoxson, E.E. Saari, D.F. Mexico, Eds). Centro Internacional de Mejoramiento de Maíz y Trigo (CIMMYT) 1996.
Goldwin I.D., Aitken A.B., Smith L.W.: Application of inter simple sequence repeats (ISSR) markers to plant genomics. Electrophoresis18, 1524–1528 (1997).
Hansen M., Hallden C., Sall T.: Error rates and polymorphism frequencies for three RAPD protocols. Plant Mol.Biol.Rep.16, 139–146 (1998).
Hantula J., Dusabenyagasani M., Hamelin R.C.: Random amplified microsatellites (RAMS) — a novel method for characterizing genetic variation within fungi. Eur.J.Forest Pathol.26, 159–166 (1996).
Josefsen L., Christiansen K.: PCR as a tool for the early detection and diagnosis of common bunt in wheat, caused by Tilletia tritici. Mycol.Res.106, 1287–1292 (2002).
Karp A., Kresovich S., Bhat K.V., Ayada W.G., Hodgkin T.: Molecular tools in plant genetic resources conservation: a guide to the technologies. IPGRI Technical Bulletin no. 2. International Plant Genetic Resources Institute, Rome 1997.
Kauserud H.: Widespread vegetative compatibility groups in the dry-rot fungus Serpula lacrymans. Mycologia96, 232–239 (2003).
Kauserud H., Schumacher T.: Regional and local population structure of the pioneer wood-decay fungus Trichaptum abietinum. Mycologia95, 416–425 (2003).
Levy L., Castlebury L.A., Carris L.M., Meyer R.J., Pimentel G.: Internal transcribed spacer sequence-based phylogeny and polymerase chain reaction-restriction fragment length polymorphism differentiation of Tilletia walkeri and T. indica. Phytopathology91, 935–940 (2001).
Liang H., Peng Y.L., Zhang G.Z., Chen W.Q., Liu T.G.: Amplification and sequence analysis of the rDNA intergenic spacer (rDNA 2 IGS) from three Tilletia species. (In Chinese) Acta Phytopathol.Sinica36, 407–412 (2006).
Liu J.H., Gao L., Liu T.G., Chen W.Q.: Development of a SCAR marker for diagnosis of dwarf bunt of wheat and detection of Tilletia controversa KÜHN. Lett.Appl.Microbiol.49, 235–240 (2009).
Mathre D.E.: Dwarf bunt: politics, identification, and biology. Ann.Rev.Phytopathol.34, 67–85 (1996).
Mcdonald J.G., Wong E., White G.P.: Differentiation of Tilletia species by rep-PCR genomic fingerprinting. Plant Dis.84, 1121–1125 (2000).
Meng X., Chen W.: Applications of AFLP and ISSR techniques in detecting genetic diversity in the soybean brown stem rot pathogen Phialophora gregata. Mycol.Res.105, 936–940 (2001).
Menzies J.G., Bakkeren G., Matheson F., Procunier J.D., Woods S.: Use of inter-simple sequence repeats and amplified fragment length polymorphisms to analyze genetic relationships among small grain infecting species of Ustilago. Phytopathology93, 167–175 (2003).
Mishra P.K., Tewari J.P., Clear R.M., Turkington T.K.: Molecular genetic variation and geographical structuring in Fusarium graminearum. Ann.Appl.Biol.145, 299–307 (2004).
Mongkolporn O., Dokmaihom Y., Kanchana-Udomkan C., Pakdeevaraporn P.: Genetic purity test of F1 hybrid Capsicum using molecular analysis. J.Hort.Sci.Biotechnol.79, 449–451 (2004).
Mulholland V., Mcewan M.: PCR-based diagnostics of Microdochium nivale and Tilletia tritici infection of winter wheat seeds. Bulletin OEPP/EPPO30, 543–547 (2000).
Nagaoka T., Ogihara Y.: Applicability of inter-simple sequence repeat polymorphisms in wheat for use as DNA markers in comparison to RFLP and RAPD markers. Theor.Appl.Genet.94, 597–602 (1997).
Nghia N.A., Kadir J., Sunderasan E., Abdullah M.P., Malik A., Napis S.: Morphological and inter simple sequence repeat markers analysis of Corynespora cassiicola isolates from rubber plantations in Malaysia. Mycopathologia166, 189–201 (2008).
O’gorman D., Xue B., Hsiang T., Goodwin P.H.: Detection of Leptosphaeria karrae with the polymerase chain reaction and primers from the ribosomal internal transcribed spacers. Can.J.Bot.72, 342–346 (1994).
Pažoutová S.: Genetic variation of Phoma sorghina isolates from southern Africa and Texas. Folia Microbiol.54, 217–229 (2009).
Pimental G.: Genetic variation among natural populations of Tilletia controversa and T. bromi. Phytopathology90, 376–383 (2000).
Pradeep Redy M., Sarla N., Siddiq A.: Inter simple sequence repeat (ISSR) polymorphism and its application in plant breeding. Euphytica128, 9–17 (2002).
Salaza O., Julian M.C., Rubio V.: Primers based on specific rDNA-ITS sequences for PCR detection of Rhizoctonia solani, R-solani AG 2 subgroups and ecological types, and binucleate Rhizoctonia. Mycol.Res.104, 281–285 (2000).
Sambrook J., Fritsch E.F., Maniatis T.: Molecular Cloning. A Laboratory Manual, 2nd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor (USA) 1989.
Shi Y.L., Loomis P., Christian D., Carris L.M., Leung H.: Analysis of the genetic relationships among the wheat bunt fungi using RAPD and ribosomal DNA markers. Phytopathology86, 311–318 (1996).
Tautz D.: Hypervariability of simple sequences as a general source for polymorphic DNA markers. Nucl.Acids Res.17, 6463–6471 (1989).
Vánky K.: Spore morphology in the taxonomy of the Ustilaginales. Trans.Mycol.Soc.Jpn.32, 381–400 (1991).
Virk P.S., Zhu J., Newbury H.J., Bryan G.J., Jackson M.T., Ford-Iloyd B.V.: Effectiveness of different classes of molecular markers for classifying and revealing variations in rice (Oryza sativa) germplasm. Euphytica112, 275–284 (2000).
Vos P., Hogers R., Bleeker M., Van De Lee T., Homes M., Frijters A., Peleman J., Kuiper M., Zabeau M.: AFLP: a new technique for DNA fingerprinting. Nucl.Acids Res.23, 4407–4414 (1995).
Weber J., Díez J., Selosse M.A., Tagu D., Le Tacon F.: SCAR markers to detect mycorrhizas of an American Laccaria bicolor strain inoculated in European Douglas-fir plantations. Mycorrhiza12, 19–27 (2002).
Williams J.G.K., Kubelik A.R., Livak K.J., Rafalski J.A., Tingey S.V.: DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucl.Acids Res.18, 6531–6535 (1990).
Zietkiewicz E., Rafalski A., Labuda D.: Genome fingerprinting by simple sequence repeat (SSR)-anchored polymerase chain reaction amplification. Genomics20, 176–183 (1994).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Gao, L., Chen, W.Q. & Liu, T.G. Development of a SCAR marker by inter-simple sequence repeat for diagnosis of dwarf bunt of wheat and detection of Tilletia controversa Kühn . Folia Microbiol 55, 258–264 (2010). https://doi.org/10.1007/s12223-010-0038-1
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-010-0038-1