Abstract
The differentiation from multipotent hematopoietic stem cells (HSC) to mature and functional blood cells requires the finely tuned regulation of gene expression at each stage of development. Specific transcription factors play a key role in this process as they modulate the expression of their target genes in an exquisitely lineage-specific manner. A large number of important transcriptional regulators have been identified which establish and maintain specific gene expression patterns during hematopoietic development. Hematopoiesis is therefore a paradigm for investigating how transcription factors function in mammalian cells, thanks also to the evolution of genome-wide and the next-generation sequencing technologies. In this review, we focus on the current knowledge of the biological and functional properties of the hematopoietic master regulator RUNX1 (also known as AML1, CBFA2, PEBP2aB) transcription factor and its main downstream target PU.1. We will outline their relationship in determining the fate of the myeloid lineage during normal stem cell development and under conditions when hematopoietic development is subverted by leukemic transformation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hematopoiesis is the dynamic process by which all the blood lineages originate from pluripotent hematopoietic stem cells. Lineage-specific transcription factors are at the heart of this process, a fact that is impressively highlighted by the finding that the development of specific lineages cannot take place in the absence of specific factors (Fig. 1). In this review we will highlight the specific role of two of these factors: RUNX1, which is part of the core-binding factor complex (CBF), and PU.1. In the absence of RUNX1, HSCs are absent, whereas in the absence of PU.1 myeloid cells and B-cells are not formed. However, in spite of their different positions in the hematopoietic hierarchy RUNX1 and PU.1 are in the same pathway within the hematopoietic transcriptional network. As mutations in this pathway lay at the heart of many cases of acute myeloid leukemia (AML) we will also discuss the role of the RUNX1/PU.1 axis in AML where myeloid differentiation is blocked [1].
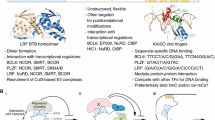
Hematopoietic hierarchy, representing the development of all blood lineages arising from totipotent cells from the inner cell mass. HSC hematopoietic stem cell, MPP multipotent progenitor, LMPP lymphoid-primed multipotent progenitor, CMP common myeloid progenitor, CLP common lymphoid progenitor, GMP granulocyte–macrophage progenitor, MEP megakaryocyte–erythrocyte progenitor. Lineage-determining transcription factors are depicted at the position where their action in critical for further blood cell development
Identification and cloning of RUNX1 and SPI1 (PU.1)
The core-binding factor (CBF) is also known as SL3-3 enhancer factor 1 [2], polyomavirus enhancer-binding factor 2 (PEBP2) [3] and SL3 and AKV core-binding factor [4] and was first purified in 1992 by Wang and Speck from calf thymus nuclei by combination of selective pH denaturation and chromatographic procedures [5]. The same investigator team described the isolation of cDNA clones encoding different CBF polypeptides from a mouse thymus cDNA library, a DNA-binding form named CBFα and a non-DNA-binding subunit, CBFβ. These subunits exist as a heterodimer prior to forming a protein–DNA complex [6]. Although the CBFα subunit is able to bind to DNA as a monomer in vitro, its affinity increases dramatically when it binds CBFβ, and their association results in the decrease of the rate of dissociation of the CBF–DNA complex [6, 7]. HPLC assays of tryptic peptides of the CBFα polypeptide reveal that one of these fragment shares 70 % sequence identity with the protein encoded by the Drosophila runt gene. Most importantly, the link between RUNX1 and blood has been established by the finding that CBFα has 100 % homology with a protein encoded by the human proto-oncogene acute myeloid leukemia 1 (AML1), which is localized on chromosome 21 and whose cDNA had been isolated and sequenced by Miyoshi et al. in 1991 as a partner in an AML causing translocation event [6, 8]. The AML1 protein is a member of a family of transcription factors that directly bind the enhancer core DNA sequence TGT/cGGT which is present in several viral and cellular promoters and enhancers [9–11]. AML1 binds DNA with a central RUNT homology domain (RHD) which constitutes 128 amino acids and is also relevant for protein–protein interactions [11, 12].
During hematopoiesis, the function of RUNX1 is strictly regulated. Mechanisms for finely controlling RUNX1 activity include alternative splicing, transcriptional control by two different promoters [13], translational control and the presence of posttranslational modifications, such as acetylation, methylation, phosphorylation, which promote transcriptional activity of RUNX1, and phosphorylation and ubiquitination, which regulate RUNX1 protein stability, reviewed in [14]. RUNX1 targets multiple genes, many of which are also pivotal transcription regulators involved in the formation of all hematopoietic lineages, including the hematopoietic-specific member of E-twenty-six (ETS) family, PU.1 [15, 16]. The gene encoding PU.1, also known as Sfpi1 (SFFV proviral integration, the human gene is named SPI1), was first identified as a novel oncogene isolated from a murine erythroleukemia induced by the acute leukemogenic retrovirus spleen focus forming virus (SFFV) [17]. PU.1 cDNA was cloned in 1990 and PU.1 protein has been shown to be a transcriptional activator in macrophages and B-cells [18]. The PU.1 protein has an 85 amino acid-long DNA-binding domain in the C-terminus which is highly conserved among all Ets family members and recognizes a purine-rich DNA sequence containing the core sequence 5′-GGAA-3′ [19]. Structural analysis of this domain revealed that PU.1 binds to DNA with its winged helix–turn–helix domain [20]. Other relevant sequences within the PU.1 activation domain include three acidic subdomains and one glutamine-rich subdomain towards the N-terminal half of the protein that are required for its transactivation function [21]. Phosphorylation of a central serine residue in position 148 has been found to be responsible for protein–protein interactions, which are also decisive in the control of transcriptional activity [22].
Targeted mutagenesis of RUNX1 and PU.1 genes
Loss-of-function animal models in conjunction with cutting-edge genome-wide technologies represent a powerful strategy to understand how cell-specific transcription factors regulate biological processes [23]. The importance of RUNX1 in embryonic hematopoiesis was shown in mice where both alleles of RUNX1 were disrupted. The absence of RUNX1 results in embryonic lethality at day E11.5–E12.5, caused by perivascular edema, hemorrhaging in the central nervous system, and a block of fetal liver hematopoiesis [24]. A similar phenotype is observed in CBFβ-knock-out mice, whereby CBFβ deficient ESCs are able to contribute to nonhematopoietic organs but not to hematopoietic tissues, i.e., peripheral blood and thymus [25]. However, Runx1 can work independently from its partner CBFβ, as suggested in a recent study, where it was shown that RUNX1 is able to drive the formation of nascent HSCs in zebrafish embryo model in the absence of CBFβ [26]. Another independent study found that murine AML1 −/− ESCs are able to give rise primitive erythroid progenitors in vitro but no myeloid or erythroid cells are produced in either fetal livers or yolk sac of AML1 −/− embryos [27]. To assess the role of RUNX1 in the development of adult blood cells, conditional RUNX1 knockout mice were generated and analyzed by several independent groups. These studies overall show that loss of RUNX1 in adult stages does not totally compromise hematopoiesis in general (i.e., HSCs are still present) [28–30], but causes expansion of a lineage negative Sca1+ Kit+ (LSK) population and the myeloid progenitor compartment in bone marrow [30, 31]. Moreover, in vivo long-term HSC activity is negatively regulated by RUNX1 [32].
PU.1 mutant embryos display a variable onset of anemia between days E14.5 and E17.5 due to defects in multiple hematopoietic lineages, and no viable embryos are found after day E18 [33, 34]. Further experiments conducted in another strain of PU.1 null mice demonstrate multiple hematopoietic abnormalities, leading to septicemia within 24 h and to death by 48 h after birth [34]. In PU.1 null mice, macrophages are totally absent and neutrophil development is severely impaired. Abnormalities are also detected in B cell compartment, which is blocked at a very early stage of maturation [34]. PU.1 has been shown to play a role also in early committing blood precursor stages. TALE-effector-mediated PU.1 repression in a murine ESC differentiation confirmed the presence of PU.1 at sites of mouse definitive hematopoiesis [35]. However, using the same experimental model, Lancrin et al. [36] investigated the capability of the major downstream targets of RUNX1, i.e., GFI1, GFI1B and PU.1, to rescue the defective phenotype of Runx1 −/− FLK1+ cells in cell differentiation assays and find that the retroviral overexpression of PU.1 in Runx1 −/− FLK1+ cells is not able to rescue their impairment in forming blast colonies, confirming its role downstream of RUNX1.
Developmental-stage-specific RUNX1 and PU.1 function
During embryogenesis the hematopoietic system originates from the mesoderm and consists of two distinct developmental waves: primitive and definitive hematopoiesis, which take place in the yolk sac and dorsal aorta, respectively [37]. As opposed to cells derived from primitive hematopoiesis, definitive hematopoiesis generates true HSCs which give rise to all blood cell compartments throughout the entire lifespan of an individual. HSCs emerge from a specialized endothelium inside the dorsal aorta, i.e., the hemogenic endothelium (HE), from which they detach and form intra-aortic cell clusters that eventually take part in blood circulation before homing to the fetal liver and the bone marrow [38]. The process that comprises the loss of endothelial specificity and the emergence of round, free-moving cells is referred to as the endothelial-to-hematopoietic transition (EHT) [39–41]. The upregulation of RUNX1 during the EHT is responsible for the upregulation of Pu.1/Sfpi1, with Runx1 mRNA being detected as early as E7.5 and Pu.1/Sfpi1 mRNA at E8.5 in the HE in the mouse [40, 42, 43].
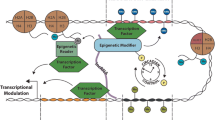
RUNX1 was shown to be essential for the formation of intra-aortic clusters, HSCs and hematopoietic progenitor formation [31], via facilitated expression of critical regulators of the EHT. These include GFI1 and GFIB, which trigger down-regulation of the endothelial markers Tek/TIE2, VE-Cadherin/CDH5 and KIT, even in the absence of RUNX1 [36]. Genome-wide studies have shown that hematopoietic genes such as PU.1 are actually primed in the HE by occupancy of TAL1/SCL and FLI1 to their regulatory regions and that a RUNX1-mediated reorganization of these factors is critical during EHT [44]. PU.1, on the other hand, contributes towards HSC maintenance by balancing the expression of cell-cycle regulators [45]. However, above all, PU.1 is a master regulator of later hematopoiesis and is required for commitment to the myeloid lineage since together with C/EBPα it controls the expression of the receptors for GM-CSF [46], and of CSF-1 which are critical cytokines for myelopoiesis [47, 48]. In addition, high levels of CSF-1 result in higher PU.1 expression [49]. C/EBPα which is required for the production of granulocyte–macrophage progenitors (GMPs) and beyond upregulates Spi1 (PU.1) but PU.1 also upregulates the expression of CEBPA [50–53]. Besides being part of such a feed-forward loop driving differentiation, high PU.1 levels and thus commitment towards the myeloid lineages are achieved via lengthening of the cell cycle and ensuing protein accumulation [54]. As the end-product of these developmentally controlled activities, PU.1 and RUNX1 function synergistically by forming a complex that excludes co-repressors [55] in myeloid cells, ultimately resulting in the regulation of lineage-specific genes such as the CSF-1 receptor gene (Csf1r) [56] and Ig-like transcripts [57]. Finally, RUNX1 is also capable of recruiting Polycomb repressive complex 1 (PRC1) to regulatory elements of myeloid genes [58]. However, in the case of RUNX1 deficiencies, this property is conferred to PU.1, although the loss of RUNX1 function prevents later switches from co-repressor to co-activator complexes, resulting in blocked differentiation [59].
PU.1 is not only a crucial factor for HSCs and myeloid progenitors but also plays an important role as a regulator of macrophage function. Macrophages are a highly heterogeneous cell type expressing high levels of this factor. In the last years, genome-wide studies have provided an explanation for its crucial role in the terminally differentiated state and made some headway into understanding its function in setting the stage for macrophage heterogeneity. PU.1 binds to more than 40,000 sites in macrophages thus opening up chromatin for rapid binding of a variety of incoming signal-inducible factors in the context of an inflammatory response [60–62] and cooperating with different types of factors in different macrophage types [63–65]. Macrophages are therefore primed to respond to a multitude of signals with the expression of a large number of different genes. In summary, the RUNX1/PU.1 axis is critical for the entire process of myeloid differentiation.
RUNX1 and PU.1 in T and B-cells
Besides its important role in myelopoiesis, RUNX1 is also a master regulator of T-cell differentiation, due to its binding to regulatory elements of T-cell-specific genes [66]. In CD4+ single positive (SP) and CD4+/CD8+ double positive (DP) T-cells, genome-wide studies have conferred a key role of RUNX1 in the modulation of the expression of these genes via distal elements [67, 68]. In CD4−/CD8− double negative (DN) and DP thymocytes, RUNX1 enhances the expression of critical genes such as the TCRα and β loci by binding to their respective enhancers [69, 70]. Furthermore, RUNX1 represses CD4 at the DN and CD8+ SP stages by binding to its silencer element [71, 72]. This repression, however, requires cooperative binding with RUNX3, the latter driving T-cell differentiation towards the CD8+ lineages [73–75], reviewed in [76]. At later CD4+ SP, stages, RUNX1 favors Th1 differentiation by inhibiting the expression of GATA3 and IL4 [77, 78], reviewed in [79]. RUNX1 also suppresses the emergence of regulatory T-cells (Treg) by inhibiting the expression of FOXP3 [80, 81]. In B-cells, the expression of RUNX1 and its binding to enhancer regions of critical pre-B cell transition genes are required for the production and survival of early B-cell progenitors [82]. Early B-cell development is also facilitated by enhanced expression of Ebf1 via the RUNX1–CBFβ interaction [83].
PU.1 is expressed in early thymocytes up to the DN2 stage, after which its down-regulation is required for normal T-cell development [84–86]. However, T-cell differentiation is severely impaired in PU.1-deficient mice [34], indicating a critical role for this transcription factor at the onset of thymopoiesis. This was later confirmed by genome-wide studies whereby binding of PU.1 revealed factor-specific epigenetic marking of crucial early T-cell differentiation genes [87]. In contrast, low levels of PU.1 expression are specifically required for normal B-cell development, since higher levels drive precursors into myeloid differentiation [88, 89]. Similar to T-cells, PU.1 expression is not required for the expression of very early B-cell genes; however, it is essential for the formation of progenitors, as it also regulates the expression of Ebf1 [90]. Likewise, PU.1 triggers chromatin remodeling in early B-cells, followed by deposition of activation-specific histone marks in distal regions later bound by B-cell-specific transcription factors [60].
Consequently, due to their critical role in hematopoiesis, both RUNX1 and PU.1 exhibit oncogenic potential in B- and T-cell lineages. In t [12, 21] B-cell acute lymphoblastic leukemia (ALL), the most common mutation in child leukemia, the RUNX1–ETV6 fusion protein causes aberrant upregulation of proliferation genes [82, reviewed in 91]. On the other hand, RUNX1 is downregulated or mutated in TLX1 T-ALL, as its normal expression entails tumor-suppressing effects [92]. Conversely, the role of PU.1 in ALL is less established, although it is expressed in most B-ALLs, and that its deletion together with SpiB systematically results in leukemia [93, 94]. In T-ALL recent studies have reported expression of PU.1; however, the mechanisms in which it could contribute towards leukemogenesis remain unclear [93, 95].
Developmental regulation of RUNX1 and PU.1: one regulates the other and both regulate themselves
Both RUNX1 and PU.1 contain distal regulatory elements harboring RUNX and ETS motifs [16, 96], indicating that both genes are subject to autoregulation [44, 45, 89, 97]. Using an inducible system, Lichtinger et al. [44] showed that the induction of Runx1 leads to binding of RUNX1 to the proximal promoter and to a strong upregulation of its own expression. Only one transcriptional start site (TSS) is known to initiate PU.1 mRNA transcription. However, an antisense RNA was described to regulate PU.1 protein levels originating from a promoter within the gene which also is regulated by the −14 kb upstream regulatory element (URE) [98]. Early in development prior to hematopoietic specification, low levels of RUNX1 prime Pu.1/Sfpi1 by binding to its −14 kb URE, an event that is accompanied by remodeling of the chromatin of cis-elements in the Pu.1 locus [99]. In adult and embryonic mouse hematopoietic progenitors RUNX1 expressed at high levels then recruits the methyltransferase mixed lineage leukemia (MLL) to the -14 Kb URE to further upregulate expression [15, 44]. Moreover, RUNX1-binding sites within the URE are crucial for PU.1 expression and the interaction between the URE and its promoter [100]. Autoregulation of the Pu.1/Sfpi1 locus is achieved by different mechanisms, depending on the cell type. Intermediate levels of PU.1 expression in B-cells are achieved by a cooperation of PU.1 with E2A and FOXO1 which bind to the -14 Kb URE. In myeloid cells, C/EBP factors activate additional myeloid-specific enhancers at -12 and -10 kb which leads to an upregulation of gene expression through the cooperation of PU.1 and C/EBP factors [89].
The deletion of the URE in mice causes the development of AML, indicating that high levels of PU.1 are required to maintain a healthy balance between proliferation and differentiation [101]. Aberrant down-regulation of PU.1 thus occurs in AML and MDS patients with frame-shift and missense RUNX1 mutations which impact its interaction with MLL [102]. These phenotypes are explained by RUNX1 binding being an absolute requirement for Pu.1/Sfpi1 expression [103], since its loss causes a block in myeloid and B-cell differentiation as well as increased platelet and T-cell development due to the tissue-specific nature of Pu.1/Sfpi1 expression [15]. In T-cells, Pu.1/Sfpi1 expression is abrogated by specific TCF/LEF binding in the absence of Wnt signaling [104].
The Runx1 locus contains two temporally modulated promoters: a proximal one (P2) which transcribes a shorter isoform, expressed in early embryonic hematopoiesis, as well as a distal one (P1) which corresponds to isoforms expressed from hematopoietic progenitors onwards [105, 106]. P1 is bound and repressed by RUNX1 itself [44, 105–107], while in T-cells P2 is activated by NFAT binding [108]. To date, an enhancer has been characterized at +23 kb in the Runx1 locus, conferring hematopoietic specificity to both isoforms [109]. Runx1 expression is controlled by TAL1/SCL via binding at the +23 kb enhancer [96, 110]. Further, GATA1 as well as ETS factors, including PU.1, bind sites required for enhancer activity at the +23 kb element, suggesting that Runx1 is a PU.1 target [96]. Finally, it was shown that RUNX1 protein expression is downregulated in AML with MLL fusion proteins, which, along with the loss of correct MLL function [111] and interaction with RUNX1 may contribute to the leukemic phenotype via incorrect regulation of RUNX1 and consequently PU.1. In summary, these studies demonstrate a profound interdependence of these two factors in driving correct hematopoietic development.
RUNX1 and PU.1 in AML
The genes encoding RUNX1 and CBFβ, its heterodimeric partner, are hot spots for chromosomal rearrangements and are the most frequent mutations associated with AML [8]. In a clinical study conducted in patients with normal karyotype, most RUNX1 mutations were detected in AML with an immature phenotype characterized by French–American–British classification subtype M0, less in M1 and M2 [112]. Genetic aberrations that can affect these genes include reciprocal chromosomal translocations, which give rise to chimeric fusion protein, such as RUNX1–ETO and CBFβ–MYH11, partial tandem duplication, intragenic loss-of-function mutations and loss of heterozygosity [112–115] (Fig. 2). Point mutations in RUNX1 have been found in both de novo and secondary AML, following myelodysplastic syndrome or chemotherapy [114, 116] and are capable of driving these diseases when introduced into the mouse germ line [117]. The majority of acquired RUNX1-associated point mutations occurs in the RHD domain or the TAD domain and confers a very poor prognosis [112, 116, 118].
Different types of genetic aberrations occur in RUNX1 locus in AML and other myeloproliferative disorders. The most frequent are point mutations, which can affect the main domains of the protein, often resulting in loss of function (a); reciprocal chromosomal translocations involving human chr21q22 locus (b); loss of heterozygosity, in which the loss of the wild-type allele results in the aberrant expression of the non-functioning allele (c). RHD RUNT homology domain, TAD transactivation domain, LOH loss of heterozygosity
The best characterized of these fusion proteins is RUNX1–ETO and it has for many years served as a paradigm for studying the molecular basis of AML [119]. In the t[8, 21] chromosomal aberration, part of the N-terminal portion of the RUNX1 protein, including its RHD, is fused to the repressor protein ETO, whose gene is localized on chromosome 8, thus producing a RUNX1–ETO chimeric protein [8, 10, 120]. RUNX1–ETO retains the ability to interact with the enhancer core DNA sequence and has been shown to interfere with RUNX1-dependent transactivation [11, 121–123] and to alter the transcriptional regulation of normal RUNX1 target genes [27, 122]. The fusion protein forms a tetrameric complex that interacts with other important hematopoietic regulators, namely the bridging factors LMO2 and LDB1, the E-Box binding factor HEB and the ETS family members FLI1 and ERG [123–126]. Genome-wide studies demonstrated that the majority of RUNX1–ETO binding sites overlap with binding sites for RUNX1 [122, 125] and it was shown later that RUNX1 and RUNX1–ETO form complexes with the same transcriptional regulators and compete for the same binding sites. However, RUNX1–ETO and RUNX1 have distinct preferences for co-repressors and co-activators, with RUNX1–ETO being mostly associated with repressed genes [123]. These data show that RUNX1–ETO and RUNX1 are in balance with each other, and additional studies have shown that each regulate distinct sets of genes which are both required to keep AML cells alive [127]. This dependency on the wild-type copy of RUNX1 has also been demonstrated for other RUNX1 and CBF translocations [127, 128]. Genome-wide binding studies have shown that RUNX1–ETO binds to thousands of genes in chromatin [122, 125]. The nature of these binding sites is dictated by the nature of the RUNX1–ETO complex binding to sequences with RUNX, E-Box and ETS binding motifs whereas RUNX1 has a much larger choice of interaction partners and also binds to additional sequences [122]. Using a combination of chromatin immunoprecipitation studies and knock-down approaches, Ptasinska et al. [123] identified and characterized the core t[8, 21] specific transcriptional network and demonstrated that the binding of RUNX1–ETO occurs at chromatin regions that are destined for differentiation-drive transcription factor exchange during myeloid differentiation. At such sites, C/EBPα is the main driver of this exchange, together with RUNX1, binding to thousands of new sites once RUNX1–ETO is depleted. This exchange does not occur in t[8, 21] AML since RUNX1–ETO represses CEBPA, differentiation is thus blocked [122, 129]. Besides providing a system-wide molecular explanation of the block in differentiation, this work therefore also provided the molecular explanation for the crucial role of C/EBPα for myelopoiesis [52].
Although PU.1 mutations occurring in the DNA-binding domain and the transactivation domain have been found in AML patients [130], they are not common [131]. However, PU.1 expression is required for leukemia development in a RUNX1/Eto9a-dependent leukemia model as a reduction of PU.1 levels results in a delayed onset of the AML phenotype [100]. In addition, although PU.1 expression is not directly controlled by chimeric MLL proteins, it is required for initiating and maintaining the leukemic phenotype induced by the MLL fusions, via cooperation with HOXA9 and MEIS1 proteins, which play a central role in the MLL-dependent leukemogenicity. Indeed, reduced expression of PU.1 in MLL-AF9 cells also results in a delay in the onset of leukemia [132]. Similarly, the upregulation of the CSF1R by the cooperation between PU.1 and the fusion protein MOZ-TIF2 is essential in the establishment and maintenance of an AML phenotype [47]. As outlined above, reduction in PU.1 expression induces a premalignant state that is associated with a high frequency of chromosomal rearrangements, leading to the AML development [101]. In addition, mutations within the URE and loss of heterozygosity in the PU.1 locus, detected in AML patients, strongly suggest that these rearrangements could be associated with AML [133, 134]. Taken together, these data highlight the importance of the RUNX1–PU.1 axis for both normal and leukemia stem cells.
Although in the last few years many details about the mechanisms by which RUNX1 and PU.1 cooperate have been elucidated, many relevant insights remain still elusive. In particular, our knowledge of the molecular details of how these two factors program the epigenome and alter transcriptional networks is still incomplete and this holds true for both normal and abnormal hematopoiesis. However, such understanding is vital if we want to revert malignant phenotypes and design more efficient therapeutic treatments for leukemias and other blood disorders. There is still a large amount of fundamental science to do.
References
Bonifer C, Bowen DT. Epigenetic mechanisms regulating normal and malignant haematopoiesis: new therapeutic targets for clinical medicine. Expert Rev Mol Med. 2010;12:e6.
Thornell A, Hallberg B, Grundstrom T. Differential protein binding in lymphocytes to a sequence in the enhancer of the mouse retrovirus SL3-3. Mol Cell Biol. 1988;8:1625–37.
Kamachi Y, Ogawa E, Asano M, Ishida S, Murakami Y, Satake M, Ito Y, Shigesada K. Purification of a mouse nuclear factor that binds to both the A and B cores of the polyomavirus enhancer. J Virol. 1990;64:4808–19.
Boral AL, Okenquist SA, Lenz J. Identification of the SL3-3 virus enhancer core as a T-lymphoma cell-specific element. J Virol. 1989;63:76–84.
Wang SW, Speck NA. Purification of core-binding factor, a protein that binds the conserved core site in murine leukemia virus enhancers. Mol Cell Biol. 1992;12:89–102.
Wang S, Wang Q, Crute BE, Melnikova IN, Keller SR, Speck NA. Cloning and characterization of subunits of the T-cell receptor and murine leukemia virus enhancer core-binding factor. Mol Cell Biol. 1993;13:3324–39.
Ogawa E, Inuzuka M, Maruyama M, Satake M, Naito-Fujimoto M, Ito Y, Shigesada K. Molecular cloning and characterization of PEBP2 beta, the heterodimeric partner of a novel Drosophila runt-related DNA binding protein PEBP2 alpha. Virology. 1993;194:314–31.
Miyoshi H, Shimizu K, Kozu T, Maseki N, Kaneko Y, Ohki M. t[8, 21] breakpoints on chromosome 21 in acute myeloid leukemia are clustered within a limited region of a single gene, AML1. Proc Natl Acad Sci USA. 1991;88:10431–4.
Daga A, Tighe JE, Calabi F. Leukaemia/Drosophila homology. Nature. 1992;356:484.
Erickson P, Gao J, Chang KS, Look T, Whisenant E, Raimondi S, Lasher R, Trujillo J, Rowley J, Drabkin H. Identification of breakpoints in t[8, 21] acute myelogenous leukemia and isolation of a fusion transcript, AML1/ETO, with similarity to Drosophila segmentation gene, runt. Blood. 1992;80:1825–31.
Meyers S, Downing JR, Hiebert SW. Identification of AML-1 and the [8, 21] translocation protein (AML-1/ETO) as sequence-specific DNA-binding proteins: the runt homology domain is required for DNA binding and protein-protein interactions. Mol Cell Biol. 1993;13:6336–45.
Levanon D, Negreanu V, Bernstein Y, Bar-Am I, Avivi L, Groner Y. AML1, AML2, and AML3, the human members of the runt domain gene-family: cDNA structure, expression, and chromosomal localization. Genomics. 1994;23:425–32.
Ghozi MC, Bernstein Y, Negreanu V, Levanon D, Groner Y. Expression of the human acute myeloid leukemia gene AML1 is regulated by two promoter regions. Proc Natl Acad Sci USA. 1996;93:1935–40.
Goyama S, Huang G, Kurokawa M, Mulloy JC. Posttranslational modifications of RUNX1 as potential anticancer targets. Oncogene. 2014. doi:10.1038/onc.2014.305.
Huang G, Zhang P, Hirai H, Elf S, Yan X, Chen Z, Koschmieder S, Okuno Y, Dayaram T, Growney JD, et al. PU.1 is a major downstream target of AML1 (RUNX1) in adult mouse hematopoiesis. Nat Genet. 2008;40:51–60.
Hoogenkamp M, Krysinska H, Ingram R, Huang G, Barlow R, Clarke D, Ebralidze A, Zhang P, Tagoh H, Cockerill PN, et al. The PU.1 locus is differentially regulated at the level of chromatin structure and noncoding transcription by alternate mechanisms at distinct developmental stages of hematopoiesis. Mol Cell Biol. 2007;27:7425–38.
Moreau-Gachelin F, Tavitian A, Tambourin P. Spi-1 is a putative oncogene in virally induced murine erythroleukaemias. Nature. 1988;331:277–80.
Klemsz MJ, McKercher SR, Celada A, Van Beveren C, Maki RA. The macrophage and B cell-specific transcription factor PU.1 is related to the ETS oncogene. Cell. 1990;61:113–24.
Karim FD, Urness LD, Thummel CS, Klemsz MJ, McKercher SR, Celada A, Van Beveren C, Maki RA, Gunther CV, Nye JA, et al. The ETS-domain: a new DNA-binding motif that recognizes a purine-rich core DNA sequence. Genes Dev. 1990;4:1451–3.
Kodandapani R, Pio F, Ni CZ, Piccialli G, Klemsz M, McKercher S, Maki RA, Ely KR. A new pattern for helix-turn-helix recognition revealed by the PU.1 ETS-domain-DNA complex. Nature. 1996;380:456–60.
Klemsz MJ, Maki RA. Activation of transcription by PU.1 requires both acidic and glutamine domains. Mol Cell Biol. 1996;16:390–7.
Pongubala JM, Van Beveren C, Nagulapalli S, Klemsz MJ, McKercher SR, Maki RA, Atchison ML. Effect of PU.1 phosphorylation on interaction with NF-EM5 and transcriptional activation. Science. 1993;259:1622–5.
DeVilbiss AW, Sanalkumar R, Johnson KD, Keles S, Bresnick EH. Hematopoietic transcriptional mechanisms: from locus-specific to genome-wide vantage points. Exp Hematol. 2014;42:618–29.
Wang Q, Stacy T, Binder M, Marin-Padilla M, Sharpe AH, Speck NA. Disruption of the CBFA2 gene causes necrosis and hemorrhaging in the central nervous system and blocks definitive hematopoiesis. Proc Natl Acad Sci USA. 1996;93:3444–9.
Wang Q, Stacy T, Miller JD, Lewis AF, Gu TL, Huang X, Bushweller JH, Bories JC, Alt FW, Ryan G, et al. The CBFbeta subunit is essential for CBFalpha2 (AML1) function in vivo. Cell. 1996;87:697–708.
Bresciani E, Carrington B, Wincovitch S, Jones M, Gore AV, Weinstein BM, Sood R, Liu PP. CBFbeta and RUNX1 are required at 2 different steps during the development of hematopoietic stem cells in zebrafish. Blood. 2014;124:70–8.
Okuda T, van Deursen J, Hiebert SW, Grosveld G, Downing JR. AML1, the target of multiple chromosomal translocations in human leukemia, is essential for normal fetal liver hematopoiesis. Cell. 1996;84:321–30.
Ichikawa M, Asai T, Saito T, Seo S, Yamazaki I, Yamagata T, Mitani K, Chiba S, Ogawa S, Kurokawa M, et al. AML-1 is required for megakaryocytic maturation and lymphocytic differentiation, but not for maintenance of hematopoietic stem cells in adult hematopoiesis. Nat Med. 2004;10:299–304.
Putz G, Rosner A, Nuesslein I, Schmitz N, Buchholz F. AML1 deletion in adult mice causes splenomegaly and lymphomas. Oncogene. 2006;25:929–39.
Growney JD, Shigematsu H, Li Z, Lee BH, Adelsperger J, Rowan R, Curley DP, Kutok JL, Akashi K, Williams IR, et al. Loss of Runx1 perturbs adult hematopoiesis and is associated with a myeloproliferative phenotype. Blood. 2005;106:494–504.
Chen MJ, Yokomizo T, Zeigler BM, Dzierzak E, Speck NA. Runx1 is required for the endothelial to haematopoietic cell transition but not thereafter. Nature. 2009;457:887–91.
Ichikawa M, Goyama S, Asai T, Kawazu M, Nakagawa M, Takeshita M, Chiba S, Ogawa S, Kurokawa M. AML1/Runx1 negatively regulates quiescent hematopoietic stem cells in adult hematopoiesis. J Immunol. 2008;180:4402–8.
Scott EW, Simon MC, Anastasi J, Singh H. Requirement of transcription factor PU.1 in the development of multiple hematopoietic lineages. Science. 1994;265:1573–7.
McKercher SR, Torbett BE, Anderson KL, Henkel GW, Vestal DJ, Baribault H, Klemsz M, Feeney AJ, Wu GE, Paige CJ, et al. Targeted disruption of the PU.1 gene results in multiple hematopoietic abnormalities. EMBO J. 1996;15:5647–58.
Wilkinson AC, Kawata VK, Schutte J, Gao X, Antoniou S, Baumann C, Woodhouse S, Hannah R, Tanaka Y, Swiers G, et al. Single-cell analyses of regulatory network perturbations using enhancer-targeting TALEs suggest novel roles for PU.1 during haematopoietic specification. Development. 2014;141:4018–30.
Lancrin C, Mazan M, Stefanska M, Patel R, Lichtinger M, Costa G, Vargel O, Wilson NK, Moroy T, Bonifer C, et al. GFI1 and GFI1B control the loss of endothelial identity of hemogenic endothelium during hematopoietic commitment. Blood. 2012;120:314–22.
Medvinsky A, Rybtsov S, Taoudi S. Embryonic origin of the adult hematopoietic system: advances and questions. Development. 2011;138:1017–31.
Medvinsky A, Dzierzak E. Definitive hematopoiesis is autonomously initiated by the AGM region. Cell. 1996;86:897–906.
Kissa K, Herbomel P. Blood stem cells emerge from aortic endothelium by a novel type of cell transition. Nature. 2010;464:112–5.
Lancrin C, Sroczynska P, Stephenson C, Allen T, Kouskoff V, Lacaud G. The haemangioblast generates haematopoietic cells through a haemogenic endothelium stage. Nature. 2009;457:892–5.
Eilken HM, Nishikawa S, Schroeder T. Continuous single-cell imaging of blood generation from haemogenic endothelium. Nature. 2009;457:896–900.
Moignard V, Woodhouse S, Fisher J, Gottgens B. Transcriptional hierarchies regulating early blood cell development. Blood Cells Mol Dis. 2013;51:239–47.
Swiers G, Baumann C, O’Rourke J, Giannoulatou E, Taylor S, Joshi A, Moignard V, Pina C, Bee T, Kokkaliaris KD, et al. Early dynamic fate changes in haemogenic endothelium characterized at the single-cell level. Nat Commun. 2013;4:2924.
Lichtinger M, Ingram R, Hannah R, Muller D, Clarke D, Assi SA, Lie ALM, Noailles L, Vijayabaskar MS, Wu M, et al. RUNX1 reshapes the epigenetic landscape at the onset of haematopoiesis. EMBO J. 2012;31:4318–33.
Staber PB, Zhang P, Ye M, Welner RS, Nombela-Arrieta C, Bach C, Kerenyi M, Bartholdy BA, Zhang H, Alberich-Jorda M, et al. Sustained PU.1 levels balance cell-cycle regulators to prevent exhaustion of adult hematopoietic stem cells. Mol Cell. 2013;49:934–46.
Hohaus S, Petrovick MS, Voso MT, Sun Z, Zhang DE, Tenen DG. PU.1 (Spi-1) and C/EBP alpha regulate expression of the granulocyte-macrophage colony-stimulating factor receptor alpha gene. Mol Cell Biol. 1995;15:5830–45.
Aikawa, Y., Katsumoto, T., Zhang, P., Shima, H., Shino, M., Terui, K., Ito, E., Ohno, H., Stanley, E.R., Singh, H. et al. (2010) PU.1-mediated upregulation of CSF1R is crucial for leukemia stem cell potential induced by MOZ-TIF2. Nat Med, 16, 580-585, 581p following 585.
Rieger MA, Hoppe PS, Smejkal BM, Eitelhuber AC, Schroeder T. Hematopoietic cytokines can instruct lineage choice. Science. 2009;325:217–8.
Mossadegh-Keller N, Sarrazin S, Kandalla PK, Espinosa L, Stanley ER, Nutt SL, Moore J, Sieweke MH. M-CSF instructs myeloid lineage fate in single haematopoietic stem cells. Nature. 2013;497:239–43.
Heath V, Suh HC, Holman M, Renn K, Gooya JM, Parkin S, Klarmann KD, Ortiz M, Johnson P, Keller J. C/EBPalpha deficiency results in hyperproliferation of hematopoietic progenitor cells and disrupts macrophage development in vitro and in vivo. Blood. 2004;104:1639–47.
Suh HC, Gooya J, Renn K, Friedman AD, Johnson PF, Keller JR. C/EBPalpha determines hematopoietic cell fate in multipotential progenitor cells by inhibiting erythroid differentiation and inducing myeloid differentiation. Blood. 2006;107:4308–16.
Zhang DE, Zhang P, Wang ND, Hetherington CJ, Darlington GJ, Tenen DG. Absence of granulocyte colony-stimulating factor signaling and neutrophil development in CCAAT enhancer binding protein alpha-deficient mice. Proc Natl Acad Sci USA. 1997;94:569–74.
Yeamans C, Wang D, Paz-Priel I, Torbett BE, Tenen DG, Friedman AD. C/EBPalpha binds and activates the PU.1 distal enhancer to induce monocyte lineage commitment. Blood. 2007;110:3136–42.
Kueh HY, Champhekar A, Nutt SL, Elowitz MB, Rothenberg EV. Positive feedback between PU.1 and the cell cycle controls myeloid differentiation. Science. 2013;341:670–3.
Hu Z, Gu X, Baraoidan K, Ibanez V, Sharma A, Kadkol S, Munker R, Ackerman S, Nucifora G, Saunthararajah Y. RUNX1 regulates corepressor interactions of PU.1. Blood. 2011;117:6498–508.
Zhang DE, Hetherington CJ, Meyers S, Rhoades KL, Larson CJ, Chen HM, Hiebert SW, Tenen DG. CCAAT enhancer-binding protein (C/EBP) and AML1 (CBF alpha2) synergistically activate the macrophage colony-stimulating factor receptor promoter. Mol Cell Biol. 1996;16:1231–40.
Nakajima H, Asai A, Okada A, Ping L, Hamajima F, Sata T, Isobe K. Transcriptional regulation of ILT family receptors. J Immunol. 2003;171:6611–20.
Yu M, Mazor T, Huang H, Huang HT, Kathrein KL, Woo AJ, Chouinard CR, Labadorf A, Akie TE, Moran TB, et al. Direct recruitment of polycomb repressive complex 1 to chromatin by core binding transcription factors. Mol Cell. 2012;45:330–43.
Gu X, Hu Z, Ebrahem Q, Crabb JS, Mahfouz RZ, Radivoyevitch T, Crabb JW, Saunthararajah Y. Runx1 regulation of PU.1 corepressor/coactivator exchange identifies specific molecular targets for leukemia differentiation therapy. J Biol Chem. 2014;289:14881–95.
Heinz S, Benner C, Spann N, Bertolino E, Lin YC, Laslo P, Cheng JX, Murre C, Singh H, Glass CK. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol Cell. 2010;38:576–89.
Ghisletti S, Barozzi I, Mietton F, Polletti S, De Santa F, Venturini E, Gregory L, Lonie L, Chew A, Wei CL, et al. Identification and characterization of enhancers controlling the inflammatory gene expression program in macrophages. Immunity. 2010;32:317–28.
Natoli G, Ghisletti S, Barozzi I. The genomic landscapes of inflammation. Genes Dev. 2011;25:101–6.
Lavin Y, Winter D, Blecher-Gonen R, David E, Keren-Shaul H, Merad M, Jung S, Amit I. Tissue-resident macrophage enhancer landscapes are shaped by the local microenvironment. Cell. 2014;159:1312–26.
Gosselin D, Link VM, Romanoski CE, Fonseca GJ, Eichenfield DZ, Spann NJ, Stender JD, Chun HB, Garner H, Geissmann F, et al. Environment drives selection and function of enhancers controlling tissue-specific macrophage identities. Cell. 2014;159:1327–40.
Gosselin D, Glass CK. Epigenomics of macrophages. Immunol Rev. 2014;262:96–112.
Egawa T, Tillman RE, Naoe Y, Taniuchi I, Littman DR. The role of the Runx transcription factors in thymocyte differentiation and in homeostasis of naive T cells. J Exp Med. 2007;204:1945–57.
Hollenhorst PC, Chandler KJ, Poulsen RL, Johnson WE, Speck NA, Graves BJ. DNA specificity determinants associate with distinct transcription factor functions. PLoS Genet. 2009;5:e1000778.
Lepoivre C, Belhocine M, Bergon A, Griffon A, Yammine M, Vanhille L, Zacarias-Cabeza J, Garibal MA, Koch F, Maqbool MA, et al. Divergent transcription is associated with promoters of transcriptional regulators. BMC Genom. 2013;14:914.
Sun W, Graves BJ, Speck NA. Transactivation of the Moloney murine leukemia virus and T-cell receptor beta-chain enhancers by cbf and ets requires intact binding sites for both proteins. J Virol. 1995;69:4941–9.
Wotton D, Ghysdael J, Wang S, Speck NA, Owen MJ. Cooperative binding of Ets-1 and core binding factor to DNA. Mol Cell Biol. 1994;14:840–50.
Hayashi K, Natsume W, Watanabe T, Abe N, Iwai N, Okada H, Ito Y, Asano M, Iwakura Y, Habu S, et al. Diminution of the AML1 transcription factor function causes differential effects on the fates of CD4 and CD8 single-positive T cells. J Immunol. 2000;165:6816–24.
Taniuchi I, Osato M, Egawa T, Sunshine MJ, Bae SC, Komori T, Ito Y, Littman DR. Differential requirements for Runx proteins in CD4 repression and epigenetic silencing during T lymphocyte development. Cell. 2002;111:621–33.
Woolf E, Xiao C, Fainaru O, Lotem J, Rosen D, Negreanu V, Bernstein Y, Goldenberg D, Brenner O, Berke G, et al. Runx3 and Runx1 are required for CD8 T cell development during thymopoiesis. Proc Natl Acad Sci USA. 2003;100:7731–6.
Durst KL, Hiebert SW. Role of RUNX family members in transcriptional repression and gene silencing. Oncogene. 2004;23:4220–4.
Kohu K, Sato T, Ohno S, Hayashi K, Uchino R, Abe N, Nakazato M, Yoshida N, Kikuchi T, Iwakura Y, et al. Overexpression of the Runx3 transcription factor increases the proportion of mature thymocytes of the CD8 single-positive lineage. J Immunol. 2005;174:2627–36.
Carpenter AC, Bosselut R. Decision checkpoints in the thymus. Nat Immunol. 2010;11:666–73.
Komine O, Hayashi K, Natsume W, Watanabe T, Seki Y, Seki N, Yagi R, Sukzuki W, Tamauchi H, Hozumi K, et al. The Runx1 transcription factor inhibits the differentiation of naive CD4+ T cells into the Th2 lineage by repressing GATA3 expression. J Exp Med. 2003;198:51–61.
Naoe Y, Setoguchi R, Akiyama K, Muroi S, Kuroda M, Hatam F, Littman DR, Taniuchi I. Repression of interleukin-4 in T helper type 1 cells by RUNX/CBF beta binding to the Il4 silencer. J Exp Med. 2007;204:1749–55.
Wong WF, Kohu K, Chiba T, Sato T, Satake M. Interplay of transcription factors in T-cell differentiation and function: the role of Runx. Immunology. 2011;132:157–64.
Ono M, Yaguchi H, Ohkura N, Kitabayashi I, Nagamura Y, Nomura T, Miyachi Y, Tsukada T, Sakaguchi S. FOXP3 controls regulatory T-cell function by interacting with AML1/RUNX1. Nature. 2007;446:685–9.
Kitoh A, Ono M, Naoe Y, Ohkura N, Yamaguchi T, Yaguchi H, Kitabayashi I, Tsukada T, Nomura T, Miyachi Y, et al. Indispensable role of the RUNX1-CBFbeta transcription complex for in vivo-suppressive function of FOXP3+ regulatory T cells. Immunity. 2009;31:609–20.
Niebuhr B, Kriebitzsch N, Fischer M, Behrens K, Gunther T, Alawi M, Bergholz U, Muller U, Roscher S, Ziegler M, et al. Runx1 is essential at two stages of early murine B-cell development. Blood. 2013;122:413–23.
Seo W, Ikawa T, Kawamoto H, Taniuchi I. RUNX1-CBFbeta facilitates early B lymphocyte development by regulating expression of EBF1. J Exp Med. 2012;209:1255–62.
Anderson MK, Weiss AH, Hernandez-Hoyos G, Dionne CJ, Rothenberg EV. Constitutive expression of PU.1 in fetal hematopoietic progenitors blocks T cell development at the pro-T cell stage. Immunity. 2002;16:285–96.
Dionne CJ, Tse KY, Weiss AH, Franco CB, Wiest DL, Anderson MK, Rothenberg EV. Subversion of T lineage commitment by PU.1 in a clonal cell line system. Dev Biol. 2005;280:448–66.
Del Real MM, Rothenberg EV. Architecture of a lymphomyeloid developmental switch controlled by PU.1, Notch and GATA3. Development. 2013;140:1207–19.
Zhang JA, Mortazavi A, Williams BA, Wold BJ, Rothenberg EV. Dynamic transformations of genome-wide epigenetic marking and transcriptional control establish T cell identity. Cell. 2012;149:467–82.
DeKoter RP, Singh H. Regulation of B lymphocyte and macrophage development by graded expression of PU.1. Science. 2000;288:1439–41.
Leddin M, Perrod C, Hoogenkamp M, Ghani S, Assi S, Heinz S, Wilson NK, Follows G, Schonheit J, Vockentanz L, et al. Two distinct auto-regulatory loops operate at the PU.1 locus in B cells and myeloid cells. Blood. 2011;117:2827–38.
Medina KL, Pongubala JM, Reddy KL, Lancki DW, Dekoter R, Kieslinger M, Grosschedl R, Singh H. Assembling a gene regulatory network for specification of the B cell fate. Dev Cell. 2004;7:607–17.
Pui CH, Relling MV, Downing JR. Acute lymphoblastic leukemia. N Engl J Med. 2004;350:1535–48.
Della Gatta G, Palomero T, Perez-Garcia A, Ambesi-Impiombato A, Bansal M, Carpenter ZW, De Keersmaecker K, Sole X, Xu L, Paietta E, et al. Reverse engineering of TLX oncogenic transcriptional networks identifies RUNX1 as tumor suppressor in T-ALL. Nat Med. 2012;18:436–40.
Nasr MR, Rosenthal N, Syrbu S. Expression profiling of transcription factors in B- or T-acute lymphoblastic leukemia/lymphoma and burkitt lymphoma: usefulness of PAX5 immunostaining as pan-Pre-B-cell marker. Am J Clin Pathol. 2010;133:41–8.
Sokalski KM, Li SK, Welch I, Cadieux-Pitre HA, Gruca MR, DeKoter RP. Deletion of genes encoding PU.1 and Spi-B in B cells impairs differentiation and induces pre-B cell acute lymphoblastic leukemia. Blood. 2011;118:2801–8.
McCune RC, Syrbu SI, Vasef MA. Expression profiling of transcription factors Pax-5, Oct-1, Oct-2, BOB.1, and PU.1 in Hodgkin’s and non-Hodgkin’s lymphomas: a comparative study using high throughput tissue microarrays. Mod Pathol. 2006;19:1010–8.
Nottingham WT, Jarratt A, Burgess M, Speck CL, Cheng JF, Prabhakar S, Rubin EM, Li PS, Sloane-Stanley J, Kong ASJ, et al. Runx1-mediated hematopoietic stem-cell emergence is controlled by a GATA/ETS/SCL-regulated enhancer. Blood. 2007;110:4188–97.
Chen H, Ray-Gallet D, Zhang P, Hetherington CJ, Gonzalez DA, Zhang DE, Moreau-Gachelin F, Tenen DG. PU.1 (Spi-1) autoregulates its expression in myeloid cells. Oncogene. 1995;11:1549–60.
Ebralidze AK, Guibal FC, Steidl U, Zhang P, Lee S, Bartholdy B, Jorda MA, Petkova V, Rosenbauer F, Huang G, et al. PU.1 expression is modulated by the balance of functional sense and antisense RNAs regulated by a shared cis-regulatory element. Genes Dev. 2008;22:2085–92.
Hoogenkamp M, Lichtinger M, Krysinska H, Lancrin C, Clarke D, Williamson A, Mazzarella L, Ingram R, Jorgensen H, Fisher A, et al. Early chromatin unfolding by RUNX1: a molecular explanation for differential requirements during specification versus maintenance of the hematopoietic gene expression program. Blood. 2009;114:299–309.
Staber PB, Zhang P, Ye M, Welner RS, Levantini E, Di Ruscio A, Ebralidze AK, Bach C, Zhang H, Zhang J, et al. The Runx-PU.1 pathway preserves normal and AML/ETO9a leukemic stem cells. Blood. 2014;124:2391–9.
Rosenbauer F, Wagner K, Kutok JL, Iwasaki H, Le Beau MM, Okuno Y, Akashi K, Fiering S, Tenen DG. Acute myeloid leukemia induced by graded reduction of a lineage-specific transcription factor, PU.1. Nat Genet. 2004;36:624–30.
Huang G, Zhao X, Wang L, Elf S, Xu H, Sashida G, Zhang Y, Liu Y, Lee J, Menendez S, et al. The ability of MLL to bind RUNX1 and methylate H3K4 at PU.1 regulatory regions is impaired by MDS/AML-associated RUNX1/AML1 mutations. Blood. 2011;118:6544–52.
Okada H, Watanabe T, Niki M, Takano H, Chiba N, Yanai N, Tani K, Hibino H, Asano S, Mucenski ML, et al. AML1(−/−) embryos do not express certain hematopoiesis-related gene transcripts including those of the PU.1 gene. Oncogene. 1998;17:2287–93.
Rosenbauer F, Owens BM, Yu L, Tumang JR, Steidl U, Kutok JL, Clayton LK, Wagner K, Scheller M, Iwasaki H, et al. Lymphoid cell growth and transformation are suppressed by a key regulatory element of the gene encoding PU.1. Nat Genet. 2006;38:27–37.
Pozner A, Lotem J, Xiao C, Goldenberg D, Brenner O, Negreanu V, Levanon D, Groner Y. Developmentally regulated promoter-switch transcriptionally controls Runx1 function during embryonic hematopoiesis. BMC Dev Biol. 2007;7:84.
Sroczynska P, Lancrin C, Kouskoff V, Lacaud G. The differential activities of RUNX1 promoters define milestones during embryonic hematopoiesis. Blood. 2009;114:5279–89.
Wong WF, Kurokawa M, Satake M, Kohu K. Down-regulation of RUNX1 expression by TCR signal involves an autoregulatory mechanism and contributes to IL-2 production. J Biol Chem. 2011;286:11110–8.
Wong WF, Looi CY, Kon S, Movahed E, Funaki T, Chang LY, Satake M, Kohu K. T-cell receptor signaling induces proximal RUNX1 transactivation via a calcineurin-NFAT pathway. Eur J Immunol. 2014;44:894–904.
Bee T, Ashley EL, Bickley SR, Jarratt A, Li PS, Sloane-Stanley J, Gottgens B, de Bruijn MF. The mouse RUNX1 +23 hematopoietic stem cell enhancer confers hematopoietic specificity to both RUNX1 promoters. Blood. 2009;113:5121–4.
Landry JR, Kinston S, Knezevic K, de Bruijn MF, Wilson N, Nottingham WT, Peitz M, Edenhofer F, Pimanda JE, Ottersbach K, et al. Runx genes are direct targets of SCL/TAL1 in the yolk sac and fetal liver. Blood. 2008;111:3005–14.
Zhao X, Chen A, Yan X, Zhang Y, He F, Hayashi Y, Dong Y, Rao Y, Li B, Conway RM, et al. Downregulation of RUNX1/CBFbeta by MLL fusion proteins enhances hematopoietic stem cell self-renewal. Blood. 2014;123:1729–38.
Schnittger S, Dicker F, Kern W, Wendland N, Sundermann J, Alpermann T, Haferlach C, Haferlach T. RUNX1 mutations are frequent in de novo AML with noncomplex karyotype and confer an unfavorable prognosis. Blood. 2011;117:2348–57.
Mangan JK, Speck NA. RUNX1 mutations in clonal myeloid disorders: from conventional cytogenetics to next generation sequencing, a story 40 years in the making. Crit Rev Oncog. 2011;16:77–91.
Gaidzik VI, Bullinger L, Schlenk RF, Zimmermann AS, Rock J, Paschka P, Corbacioglu A, Krauter J, Schlegelberger B, Ganser A, et al. RUNX1 mutations in acute myeloid leukemia: results from a comprehensive genetic and clinical analysis from the AML study group. J Clin Oncol. 2011;29:1364–72.
Silva FP, Morolli B, Storlazzi CT, Anelli L, Wessels H, Bezrookove V, Kluin-Nelemans HC, Giphart-Gassler M. Identification of RUNX1/AML1 as a classical tumor suppressor gene. Oncogene. 2003;22:538–47.
Osato M, Asou N, Abdalla E, Hoshino K, Yamasaki H, Okubo T, Suzushima H, Takatsuki K, Kanno T, Shigesada K, et al. Biallelic and heterozygous point mutations in the runt domain of the AML1/PEBP2alphaB gene associated with myeloblastic leukemias. Blood. 1999;93:1817–24.
Matheny CJ, Speck ME, Cushing PR, Zhou Y, Corpora T, Regan M, Newman M, Roudaia L, Speck CL, Gu TL, et al. Disease mutations in RUNX1 and RUNX2 create nonfunctional, dominant-negative, or hypomorphic alleles. EMBO J. 2007;26:1163–75.
Matsuno N, Osato M, Yamashita N, Yanagida M, Nanri T, Fukushima T, Motoji T, Kusumoto S, Towatari M, Suzuki R, et al. Dual mutations in the AML1 and FLT3 genes are associated with leukemogenesis in acute myeloblastic leukemia of the M0 subtype. Leukemia. 2003;17:2492–9.
Lam K, Zhang DE. RUNX1 and RUNX1-ETO: roles in hematopoiesis and leukemogenesis. Front Biosci (Landmark Ed). 2012;17:1120–39.
Nisson PE, Watkins PC, Sacchi N. Transcriptionally active chimeric gene derived from the fusion of the AML1 gene and a novel gene on chromosome 8 in t[8, 21] leukemic cells. Cancer Genet Cytogenet. 1992;63:81–8.
Meyers S, Lenny N, Hiebert SW. The t[8, 21] fusion protein interferes with AML-1B-dependent transcriptional activation. Mol Cell Biol. 1995;15:1974–82.
Ptasinska A, Assi SA, Mannari D, James SR, Williamson D, Dunne J, Hoogenkamp M, Wu M, Care M, McNeill H, et al. Depletion of RUNX1/ETO in t[8, 21] AML cells leads to genome-wide changes in chromatin structure and transcription factor binding. Leukemia. 2012;26:1829–41.
Ptasinska A, Assi SA, Martinez-Soria N, Imperato MR, Piper J, Cauchy P, Pickin A, James SR, Hoogenkamp M, Williamson D, et al. Identification of a dynamic core transcriptional network in t[8, 21] AML that regulates differentiation block and self-renewal. Cell Rep. 2014;8:1974–88.
Sun XJ, Wang Z, Wang L, Jiang Y, Kost N, Soong TD, Chen WY, Tang Z, Nakadai T, Elemento O, et al. A stable transcription factor complex nucleated by oligomeric AML1-ETO controls leukaemogenesis. Nature. 2013;500:93–7.
Martens JH, Mandoli A, Simmer F, Wierenga BJ, Saeed S, Singh AA, Altucci L, Vellenga E, Stunnenberg HG. ERG and FLI1 binding sites demarcate targets for aberrant epigenetic regulation by AML1-ETO in acute myeloid leukemia. Blood. 2012;120:4038–48.
Mandoli A, Singh AA, Jansen PW, Wierenga AT, Riahi H, Franci G, Prange K, Saeed S, Vellenga E, Vermeulen M, et al. CBFB-MYH11/RUNX1 together with a compendium of hematopoietic regulators, chromatin modifiers and basal transcription factors occupies self-renewal genes in inv[16] acute myeloid leukemia. Leukemia. 2014;28:770–8.
Ben-Ami O, Friedman D, Leshkowitz D, Goldenberg D, Orlovsky K, Pencovich N, Lotem J, Tanay A, Groner Y. Addiction of t[8, 21] and inv[16] acute myeloid leukemia to native RUNX1. Cell Rep. 2013;4:1131–43.
Goyama S, Schibler J, Cunningham L, Zhang Y, Rao Y, Nishimoto N, Nakagawa M, Olsson A, Wunderlich M, Link KA, et al. Transcription factor RUNX1 promotes survival of acute myeloid leukemia cells. J Clin Investig. 2013;123:3876–88.
Pabst T, Mueller BU, Harakawa N, Schoch C, Haferlach T, Behre G, Hiddemann W, Zhang DE, Tenen DG. AML1-ETO downregulates the granulocytic differentiation factor C/EBPalpha in t[8, 21] myeloid leukemia. Nat Med. 2001;7:444–51.
Mueller BU, Pabst T, Osato M, Asou N, Johansen LM, Minden MD, Behre G, Hiddemann W, Ito Y, Tenen DG. Heterozygous PU.1 mutations are associated with acute myeloid leukemia. Blood. 2002;100:998–1007.
The Cancer Genome Atlas Consortium. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N Engl J Med. 2013;368:2059–74.
Zhou J, Wu J, Li B, Liu D, Yu J, Yan X, Zheng S, Wang J, Zhang L, He F, et al. PU.1 is essential for MLL leukemia partially via crosstalk with the MEIS/HOX pathway. Leukemia. 2014;28:1436–48.
Steidl U, Steidl C, Ebralidze A, Chapuy B, Han HJ, Will B, Rosenbauer F, Becker A, Wagner K, Koschmieder S, et al. A distal single nucleotide polymorphism alters long-range regulation of the PU.1 gene in acute myeloid leukemia. J Clin Invest. 2007;117:2611–20.
Bonadies N, Pabst T, Mueller BU. Heterozygous deletion of the PU.1 locus in human AML. Blood. 2010;115:331–4.
Acknowledgments
Research in C.B.’s lab is supported by a Specialist Programme grant from Leukaemia and Lymphoma Research, by the Biotechnology and Biological Sciences Research Council (BBSRC), the Kay Kendall Leukemia Fund and the Medical Research Council, UK.
Author information
Authors and Affiliations
Corresponding author
About this article
Cite this article
Imperato, M.R., Cauchy, P., Obier, N. et al. The RUNX1–PU.1 axis in the control of hematopoiesis. Int J Hematol 101, 319–329 (2015). https://doi.org/10.1007/s12185-015-1762-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12185-015-1762-8