Abstract
QuEChERS methodology is a new alternative for polyphenol analysis in foods and beverages. This extractive and clean up method includes several steps that should be optimized to accomplish a fast and efficient extraction. In this work, chemometrics tools were applied to optimize QuEChERS parameters for polyphenol extraction from beers. By means of D-optimal screening design, the most influential extraction parameters were defined, i.e., acetonitrile volume, acidity, PSA, and C18 amount. These parameters were optimized applying a central composite design with desirability function, establishing the following optimal conditions: 2.5 mL of acetonitrile as extraction volume, 0.5% v/v of formic acid for sample acidification, 40 mg PSA for d-SPE step, and 175 mg of C18. Method validation was carried out according to International Conference on Harmonization recommendations. Data calibration curves (0.10–10.00 mg L−1) fitted a linear regression model with determination coefficients (R2) ≥ 0.992. Repeatability (relative standard deviation, RSD) and intermediate precision (RSD) showed values ≤ 4.81% (n = 6) and ≤ 6.71% (n = 3), respectively. Recovery (n = 3) at three levels ranged from 93.98 to 119.92% (RDS ≤ 4.40%) and quantification limits ranged from 0.009 to 0.118 μg mL−1. Applying the optimized and validated method, 10 beer samples were analyzed. The principal phenolic acids found were t-ferulic acid, caffeic acid, p-coumaric acid, and p-hydroxybenzoic acid. Individually, t-ferulic acid showed the highest concentration in all samples presenting a content ranged from 0.01 ± 0.01 to 2.25 ± 0.02 μg mL−1. The proposed methodology proved to be fast, reliable, and efficient for the determination of polyphenols in beer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Beer is an alcoholic beverage with antioxidant activity provided by compounds like polyphenols, sulfites, and Maillard reaction products. Polyphenols play an important role in brewing process preventing the oxidation of compounds responsible of beer flavor. Conversely, at high concentration, they can negatively influence beer physical stability producing the effect known as haze, which is the formation of insoluble compounds (Fernández de Córdova and Medina 2014). Besides these positive and negative effects, polyphenols are very interesting bioactive molecules extensively studied due to its beneficial impact on human health. Its routinely consumption has been associated with protection against the risk of development certain type of cancers, and cardiovascular and neurodegenerative diseases (Pandey and Rizvi 2009). Regarding daily intake, polyphenol content in beers is lower than in wines, but beer consumption is much higher due to its low alcohol content and lower cost. Thus, in some countries like the USA, beers may provide two times more polyphenols intake per day than wine (Vinson et al. 2003). Polyphenol analysis is mostly performed by liquid chromatography, which commonly requires a sample preparation step, e.g., liquid-liquid extraction (LLE) with solvents like diethyl ether, ethyl acetate (Mitić et al. 2013), toluene and cyclohexane (Molina-García et al. 2011), and solid-phase extraction (SPE) using C18 (Alonso García et al. 2004) or anionic exchange (Quifer-Rada et al. 2015) columns. QuEChERS (Quick, Easy, Cheap, Effective, Rugged and Safe), originally proposed for pesticides residues analysis (Anastassiades et al. 2003), is an extractive methodology with adequate extraction performance, flexible and modifiable according to analyte properties, matrix composition, equipment availability, etc. (Lehotay et al. 2010). This methodology has been adapted and applied to determine drug residues (Bourdat-Deschamps et al. 2014), mycotoxins (Fernandes et al. 2013) and polyphenols in vegetables (Silva et al. 2012), legumes (Delgado-Zamarreño et al. 2012), and wines (Fontana and Bottini 2014). The objective of the present work was to apply QuEChERS methodology for polyphenol extraction from beers. In previous works, appropriate QuEChERS parameters/conditions for polyphenol extraction have been defined through univariate experimental design (one factor at a time) (Delgado-Zamarreño et al. 2012; Fontana and Bottini 2014; Silva et al. 2012), but this approach has some disadvantages, i.e., high number of experiments, factor interactions are not evaluated, and only a small part of the experimental domain is examined. In situations where the relationships among several factors can affect the response, design of experimental (DOE) is the most appropriate option. To the best of our knowledge, this is the first report about QuEChERS conditions optimization by DOE for polyphenol extraction from beers (Dejaegher and Vander Heyden 2011). Since QuEChERS parameters are numerous, D-optimal design is an adequate alternative to identify the most influencing/significant factors. This kind of DOE is an efficient fractional factorial design that covers the maximum of the experimental region with a minimal number of experiments (Eriksson et al. 2008). After defining the most important factors by D-optimal design, a response surface methodology (RSM) is a natural choice for a multi-response optimization. Central composite design (CCD) with multi-response optimization by desirability function is one of the most employed chemometric technique because shows a high efficiency with a reduced number of experiments (Ferreira et al. 2007). The responses studied were the chromatographic peak area of each (eight) polyphenols. Quantification was carried out applying a novel chromatographic method capable of selectively detecting polyphenols in beers. This method was validated according to International Conference on Harmonization (ICH) recommendations.
Materials and Methods
Standards, Solvents, and Samples
Gallic acid (97.5–102.5%), p-hydroxybenzoic acid (Bzc, ≥ 99.0%), caffeic acid (Caf, ≥ 98.0%), trans-ferulic acid (Fer, 99.0%), vanillic acid (Van, ≥ 97.0%), p-coumaric acid (Cou, ≥ 98.0%), trans-resveratrol (Res, ≥ 99.0%), (±)-catechin (Cat, ≥ 96.0%), (−)-epicatechin (Epi, ≥ 97%), anhydrous magnesium sulfate (MgSO4, ≥ 97.0%), and ammonium formate (≥ 99.0%) were obtained from Sigma-Aldrich (St. Louis. MO, USA). Bulk sorbent for d-SPE including primary secondary amines (PSA) and octadecylsilane (C18) was purchased from United Chemical Technologies (UCT, Bristol PA, USA). Acetonitrile (ACN) and methanol, both HPLC grade, acetic acid glacial, formic acid (FA, 98–100%) and sodium chloride (NaCl) were purchased from Merck (Darmstadt, Germany). Ultrapure water (18.2 MΩ cm) was produced by means of a Millipore Simplicity system (Bedford, MA, USA). All beer samples correspond to malted barley beers, considering the type of fermentation (yeast), five samples were lager (bottom fermentation) and five ale (top fermentation). According to the kind of brewery, lager beers were produced by macrobreweries or industrial brewery, and ale beers by microbreweries or craft brewery. Regarding to the beer style, two were stout, five blonde, and three amber. All beer samples were purchased from Chilean market during 2016, stored at 4 °C and analyzed immediately after uncapping.
QuEChERS Extraction
Beer samples were degassed in an ultrasonic bath for 30 min at 20 °C. Ten milliliters of degassed sample was transferred to 50-mL centrifuge tube and acidified with 50 μL of formic acid. Then, 2.5 mL of acetonitrile was added and the solution was vigorous mixed in Thermolyne (Dubuque, USA) vortex for 2 min. For salting-out step, 4 g of MgSO4 and 2 g of NaCl were added, vortex mixed for 1 min and centrifuged (Centrifuge Hettich Universal 32R, Tuttlingen, Germany) at 3992×g for 10 min at 20 °C. For d-SPE step, 1 mL of supernatant was transferred into a 2-mL microcentrifuge tube containing 40 mg of PSA, 175 mg C18 sorbent, and 150 mg MgSO4; the mixture was vortex mixed and centrifuged (Centrifuge Sigma 1-14, Osterode am Harz, Germany) at 10625×g for 2 min at room temperature. An aliquot of 0.5 mL was evaporated to dryness under a gently stream of nitrogen and dissolved in 0.5 mL of water:methanol (4:1 v/v). The sample was filtered through 13 mm PVDF syringe filter (0.22-μm pore size) before injection into chromatography system.
Chromatography
Polyphenol quantification was performed using a Shimadzu (Kyoto, Japan) Prominence HPLC system consisted of LC-20AT pump, DGU-20A5R degassing unit, CTO-20AC column oven and SPD 20AV UV/VIS detector. Data were acquired and recorded by means of Shimadzu LabSolutions software version 5.51. Chromatography was carried on Phenomenex (Torrence, USA) Kinetex XB-C18 100A (150 × 4.6 mm; 5 μm) column connected to guard-column C18 (4.6 id), both set at 30 °C, using a mobile phase composed of acetonitrile (A) and 10 mM ammonium formate pH 3 (B). The following gradient program was used at flow rate of 0.5 mL min−1: 0–3 min, 18–20% (A); 3–4 min, 20–30% (A); 4–6 min, 30% (A) (isocratic step); 6–10 min, 30–80% (A); 10–14 min, 80% (A) (isocratic step); 14–16 min, 80–18% (A); and 16–20 min 18–18% (A) (column conditioning). Detection was performed by UV absorption at 280 nm. Mass spectrometry analyses were carried out using Shimadzu UHPLC-DAD-ESI-MS system composed of Nexera X2 UHPLC coupled online with diode array detector (DAD) and ESI interface of LCMS 8030 triple quadrupole. Chromatography was performed using the same conditions described above, and mass spectrometer was operated in scan mode using the following conditions: electrospray ionization (ESI) in negative mode with a voltage of 4.5 kV; nebulizer gas (N2) 3 L min−1, desolvation gas (N2) 15 L min−1; desolvation line temperature 250 °C and heat block temperature 400 °C.
Experimental Design
D-optimal screening design was focused in eight y-responses corresponding to chromatographic peak area of the eight polyphenols under study with the purpose of obtaining an efficient extraction. Considering these responses and the previous works carried out by Delgado-Zamarreño et al. (2012), Fontana and Bottini (2014), Silva et al. (2012), and Herrero et al. (2013), seven x-factors were selected: acetonitrile volume (x1, ACN), sample acidity (x2, %FA), MgSO4 amount (x3, MgSO4), NaCl amount (x4, NaCl), time of vortex mixing (x5, Vortex), PSA amount (x6, PSA), and C18 amount (x7, C18). The design was generated by MODDE 7.0 software (Umetrics, Sweden) and the model adjusted by mean of PLS regression, statistically validated through an analysis of variance (ANOVA) with a significance level (α) of 0.05. Optimization (maximization) of polyphenol extraction was carried by response surface methodology applying a face-centered central composite design (CCD) with desirability function, which was generated with Statgraphics Centurion XV version 15.1.02 software (Rockville, MD, USA). The factors optimized were % FA, PSA, and C18 selected according to the results obtained with D-optimal design. The response evaluated was recovery of each polyphenols.
Results and Discussion
D-optimal Design Evaluation and PLS Model Interpretation
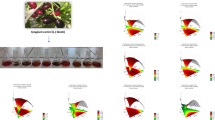
For screening design, the following ranges were established for each factor: acetonitrile volume (2.0–5.0 mL), sample acidity (0–2.5% v/v FA), MgSO4 (2.0–4.0 g), NaCl (0–3.0 g), time of vortex mixing (1.0–3.0 min), PSA (25–200 mg), and C18 (0–200 mg). The experimental design consisted of 40 experiments including 4 central points. All experiments were randomly conducted in order to minimize the effects of uncontrolled factors using beer samples spiked with 5 μg mL−1 of each polyphenol. Results showed a G-efficiency of 71.9% and condition number of 1.18, both values close to the recommended ones, 60–70% and < 3, respectively (Eriksson et al. 2008). MODDE software allows to fit experimental data applying multiple linear regression (MLR) o partial least squares (PLS). For this research, MLR was considered not efficient because separate regression models are fitted for each response. PLS regression also named projection to latent structures is a multivariate calibration that relates two data matrices, a factor/variable matrix X and response matrix Y, and models their relationships and structures by X and Y decomposition into latent structures. PLS regression can model simultaneously the variation of all responses to the variation of all variables providing an overall representation of their relationships instead of a separate model for each response (Campos-Requena et al. 2015; Eriksson et al. 2008). This fitted model has been proposed to evaluate screening experiments when several responses are measured, finding the most important experimental variables in loading projections (Carlson et al. 1991; Eriksson et al. 1998). For these reasons, PLS regression was the model of choice to fit experimental data. In MODDE software, the number of significant PLS component is determined by automatic cross-validation, after that prediction residual sum of square (PRESS) is computed for each model dimensions. Since PRESS is the square difference between observed Y and predicted Y when the observations were kept out from the model estimation, the predictive power of a PLS model is evaluated by Q2 (goodness of prediction), which is the fraction of the total response variation that can be predicted by the model and it is computed based on PRESS. MODDE software computes an overall Q2 for all PLS components, for all responses and for individual responses. Large Q2 or higher than 0.5 indicates that the model has good predictive ability with small prediction errors. Also, PLS model fitness evaluation can by determined by the explained variation, which is expressed as R2 (goodness of the model) and it corresponds to the response variation explained by the model (Campos-Requena et al. 2014; Eriksson et al. 2008). Considering the information obtained from coefficient and residual plots, the original model was refined removing non-significant (p < 0.05) terms and outliers data. Then, the model was fitted again and four x-factors and four interactions were established as significant: ACN volume, %FA, PSA, and C18 amounts, and the interactions: ACN*FA, ACN*PSA, FA*PSA, and PSA*C18. As can be observed in Fig. 1a, four PLS components were required for overall regression model showing R2 = 0.777 and Q2 = 0.596 (p ≤ 0.05). These values suggested a good correlation considering that eight different responses were modeled at the same time. Individual R2 and Q2 values for each response ranged from 0.741 to 0.819 and from 0.557 to 0.669, respectively (p ≤ 0.05, Fig. 1b). The estimation of PLS model significance was determined with ANOVA with a significance level (α) of 0.05. Values showed a satisfactory significance of the regression model (p ≤ 0.05) for each y-response without lack of fit (p ≥ 0.05). Score plots shown that PLS component had an adequate correlation (Fig. 2), and even when PLS component showed a higher dispersion, the correlation structure was reasonably strong. The two first PLS components indicated that changes on factors were influenced for the monitored responses and the most relevant information can be inferred from these PLS components. The third and fourth PLS components indicated a more scattered correlation (data not shown). As can be observed in loading plot (Fig. 3), ACN volume and % FA factors showed a strong impact on the measured responses. Contrary, PSA and C18 had closeness to plot origin showing a lower impact. Four interactions were significant (p ≤ 0.05), being FA*PSA the most relevant. In general terms, the closeness of %FA and Bzc, Caf, Cou, Van, and Fer responses can be interpreted as a correlation between them. Figure 3a, b shows that ACN and %FA*PSA are the most relevant factors for polyphenols (Res) extraction, followed by PSA and %FA. In particular case of Res, ACN, %FA, and PSA factors intercept the baseline on the opposite side of the plot origin, showing therefore a negatively correlation, contrarily, FA*PSA showed a positive effect on this polyphenol extraction (Fig. 3a). Different correlations were observed depending on the polyphenol evaluated, for example %FA showed a positive correlation with Van extraction (Fig. 3b). The new model equation (Eq. 1) obtained by PLS regression was
where b corresponds to regression coefficients and their values are detailed in Table 1. According to the coefficient plots (Fig. 4), %FA had the highest impact on polyphenol extraction, mainly on Bzc, Caf, Cou, Van, and Fer. These phenolic acids are weak acids with pKa ranged from 4.0 to 4.5 (Beltrán et al. 2003) and therefore are partially ionized in beers (pH ca. 4.1). Thus, the FA addition favors the equilibrium towards neutral molecules increasing the partition to acetonitrile (Fontana and Bottini (2014). In the case of very weak acids with higher pKa (8.7–9.2), phenolic acids like Cat, Epi, and Res showed different behaviors. Cat was positively and Epi and Res negatively affected by FA addition but with a minor impact (Fig. 4). These results are not in agree with the reported by Fontana and Bottini (2014), who reported a non-significant difference between these three polyphenols. The use of PSA in d-SPE showed important effects but of lower impact than %FA. Since PSA sorbent is used to eliminate acid compounds such as pigments, fatty acid, and some sugars, it is also capable of removing phenolic acids. Therefore, PSA effect should be evaluated together with %FA to avoid misleading conclusions because the use of inadequate FA volumes could lead to polyphenols removal by PSA. Thus, in this work, the %FA-PSA ratio was also evaluated. Fontana and Bottini (2014) reported similar conclusions regarding the importance of PSA and %FA, but without described a simultaneous evaluation of both factors. C18 sorbent had a lower impact on extraction efficiency, but its evaluation or incorporation should not be discarded because it has an impact on matrix cleaning, decreasing the background observed in UV-chromatograms. In comparison with previous works, the applied screening model allowed to have an overview of relationships between all factors and all responses simultaneously, and to evaluate the impact of each factor over the response. The model interpretation in terms of loading is a powerful tool, which may be supplemented with other more traditional tools such as the coefficient overview plot (Eriksson et al. 2008). Thereby, the applied chemometric tools were very useful for establishing the more relevant QuEChERS factors/conditions that affect polyphenol extraction from beers.
Interpretation example of PLS loading plots with the first and second PLS component for two responses. a Relationship of resveratrol (open square) with the most important individual factors and interactions among factors (dotted circle). b Relationship of vanillic acid with the most important individual factors and interactions among factors
Optimization of QuEChERS Extraction
From D-optimal results, four factors (%FA, PSA, C18, ACN volume) were chosen to carry out the optimization of QuEChERS extraction conditions. The first factor, ACN volume, was evaluated individually resulting in an optimal volume of 2.5 mL, while the other three factors, %FA (0–1.5%), PSA (0–100 mg), and C18 (100–200 mg), were optimized applying a face-centered CCD resulting an experimental plan with 17 runs including three central points. All experiments were carried out in duplicate using a beer sample spiked with 1 μg mL−1 of each polyphenol. Van analysis was excluded due to the presence of matrix interference peaks, including instead gallic acid. Data fitted a quadratic response surface model, which was validated through ANOVA (p < 0.05) showing determination coefficients (R2) from 79.57 to 93.30. Optimal conditions for eight responses were calculated by multiple response optimization using desirability function with an overall desirability of 0.82. Thus, QuEChERS optimal conditions were established as follows: 2.5 mL of ACN (calculated individually), 0.35% FA, 175 mg C18, and 0.00 mg PSA. However, PSA sorbent showed an important role removing sample interferences; therefore, its use was considered necessary, and the optimal values adjusted accordingly. As can be observed in response surface plot (Fig. 5), using 40 mg of PSA, the overall desirability was close to 0.80 with 0.5% of FA; these values are within the optimal zone, hence, they could be used to accomplish optimal polyphenol recoveries. These optimal conditions were experimentally evaluated resulting in adequate recoveries and chromatograms with lower matrix interference (Fig. 6). Applying optimal conditions, 10 mL of acidified beer with 0.5% of FA reached a pH ca. 2.6, which is lower than studied polyphenols pKa, finding therefore in non-ionized form facilitating the extraction with ACN. Regarding the PSA amount selected, Fontana and Bottini (2014) reported a decrease in polyphenol extraction using 100 mg PSA in d-SPE step, but in the same experiments using 50 mg PSA, recoveries showed values from 75 to 119%, which is concordant with the PSA amount proposed in the present work. Contrarily, Burin et al. (2014) using the same amount of PSA (50 mg) reported low recoveries from grapes (57 to 79%), which evidence the importance of studying factors behavior for each matrix. In the case of polyphenol determination in vegetables, Silva et al. (2012), using 25 mg of PSA and C18 reported recoveries values from 78.4 to 99.8%. These reports are in agreement with our results showing that higher PSA amount negatively affect polyphenol extraction; however, it is possible to use lower amount without significantly change on polyphenol extraction but obtaining chromatograms with low background or matrix interference.
Validation
The proposed methodology was validated following ICH recommendations (ICH 2005). Calibrations with and without matrix were established to evaluate a possible matrix effect. Since slopes were statistically different (p < 0.05) calibrations were established in beer matrix at seven levels from 0.10 to 10.00 μg mL−1. Calibration data fitted a linear regression model with determination coefficients (R2) higher than 0.992. Method precision was evaluated through repeatability and intermediate precision using a beer sample spiked with 1 μg mL−1 of each polyphenol. Repeatability (n = 6) showed RSDs values from 0.70 to 4.81% and intermediate precision was assayed in three different days (n = 3), showing RSDs values ≤ 6.71%. Method accuracy was determined through recovery evaluation with three different levels defined according to the calibration range and reported values. Beer samples were spiked with 0.25, 0.50, and 1.00 μg mL−1 of each polyphenol. Recovery values expressed as percentage showed a ranged from 93.98 to 119.92% with RSD values ≤ 4.40%. Detection and quantification limits were calculated using signal-to-noise ratios (S/N) of 3 and 10, respectively. Considering an injection volume of 20 μL, the detection and quantification limits ranged from 0.003 to 0.036 μg mL−1, and from 0.009 to 0.118 μg mL−1, respectively. Selectivity was evaluated in two steps: first, the identification of polyphenols peaks was carried out comparing retention time (tR) with pure standard and standard addition method. Then, selectivity was evaluated determining peak purity by UV absorption using a diode array detector, showing r > 0.99 for all polyphenols. Additionality, by mass spectrometry analysis a full scan spectra were acquired from m/z 50 to 1000, showing unique and the corresponding m/z value of each polyphenol. Comparatively with other methods, recovery values (94.0–119.9%) were higher than those obtained with laborious methodologies using solid phase extraction, e.g., Alonso García et al. (2004) (61.0–100.0%) and Dvořáková et al. (2007) (63.7–102.7%). Comparing the proposed method with reports that used QuEChERS for polyphenol extraction, recoveries were slightly higher than those reported by Silva et al. (2012) (78.4–99.9%) and Fontana and Bottini (2014) (75.0–119.6%). LOD values (0.003–0.036 mg L−1) were slightly lower than the ranges presented by Silva et al. (2012) (LOD 0.005–0.62 mg L−1) and similar than those reported by Fontana and Bottini (2014) (LOD 0.004–0.079 mg L−1). Even when MS quantification provides lower LOD (0.0007–0.0015 mg L−1), the reproducibility (RSD ≤ 25.8%) (Delgado-Zamarreño et al. 2012) is inferior compared to the present method (RSD ≤ 6.71). Thus, the proposed methodology proved to be faster with an adequate detection capacity (fit for propose) for polyphenol determination in beers.
Samples Analysis
Malt is the main source of beer polyphenols contributing near to 80% of total polyphenol content, the rest is provided by hop (Quifer-Rada et al. 2015). Using the optimized and validated method, ten beer samples were analyzed. In agreement with previous results (Table 2), the principal phenolic acids found were t-ferulic acid, caffeic acid, p-coumaric acid, and p-hydroxybenzoic acid, which were present in all samples. Individually, t-ferulic acid was the major polyphenol found with a mean value of 1.15 ± 0.60 mg L−1. Analyzing the individual content of each polyphenols, p-hydroxybenzoic acid levels have been informed in just a few reports, Callemien and Collin (2010) described similar values (0.02 to 3.09 mg L−1), but Floridi et al. (2003) reported values at least 25 times higher. Gallic acid concentrations in beers are usually lower than 2 mg L−1; the values observed in Chilean samples were lower than the ones reported elsewhere. The values detected for hydroxycinnamic acids, p-coumaric acid (0.10–0.76 mg L−1), and t-ferulic acid (0.01–2.25 mg L−1) were concordat with those reported by Alonso García et al. (2004), Piazzon et al. (2010), and Mitić et al. (2013). Caffeic acid (0.18–1.61 mg L−1) values were similar to those reported by Dvořáková et al. (2007) and Zhao et al. (2010), but slightly higher to other reports (Table 2). According to the results, hydroxycinnamic acids were the main responsible of total polyphenol content. Other type of phenolic structures could affect the physical characteristics of beer; it has been described a haze-forming activity with high (+)-catechin and (−)-epicatechin contents due to their possible interaction with proteins. In case of (+)-catechin, the values found (0.06–0.31 mg L−1) were lower than those previously reported (Table 2), while (−)-epicatechin values (nd −2.22 mg L−1) were similar to those published by Dvořáková et al. (2007) and Callemien and Collin (2010) (0.80 to 1.90 mg L−1), but relatively higher than the ones reported by Alonso García et al. (2004) and Zhao et al. (2010). However, no haze-forming activity was visually observed. The stilbene resveratrol was not detected in any sample; however, some previous articles have reported resveratrol presence in beers at levels lower than 0.07 mg L−1 (Chiva-Blanch et al. 2011; Molina-García et al. 2011).
Conclusions
The present work reports for the first time the use of experimental designs to optimize QuEChERS conditions for polyphenol extraction from beers. D-optimal, as screening chemometric tools, allowed the selection of four factors that significantly affected polyphenol extraction. These factors were optimized using a face-centered CCD, where acidity (%FA) was one of the critical factor, affecting also the inclusion of PSA and its optimal amount. According to optimization results, only three QuEChERS factors were important for polyphenol extraction from beers. Considering validation results and sample analysis, the method was appropriate, reliable, and sensitive for all polyphenols studied. This methodology was applied to carried out for the first time a preliminary study of polyphenol content in beers commercialized in Chilean market.
References
Alonso García A, Cancho Grande B, Simal Gándara J (2004) Development of a rapid method based on solid-phase extraction and liquid chromatography with ultraviolet absorbance detection for the determination of polyphenols in alcohol-free beers. J Chromatogr A 1054:175–180. https://doi.org/10.1016/j.chroma.2004.07.092
Anastassiades M, Lehotay SJ, Štajnbaher D, Schenck FJ (2003) Fast and easy multiresidue method employing acetonitrile extraction/partitioning and “dispersive solid-phase extraction” for the determination of pesticide residues in produce. J AOAC Int 86:412–431
Beltrán JL, Sanli N, Fonrodona G, Barrón D, Özkan G, Barbosa J (2003) Spectrophotometric, potentiometric and chromatographic pKa values of polyphenolic acids in water and acetonitrile–water media. Anal Chim Acta 484:253–264. https://doi.org/10.1016/S0003-2670(03)00334-9
Bourdat-Deschamps M, Leang S, Bernet N, Daudin J-J, Nélieu S (2014) Multi-residue analysis of pharmaceuticals in aqueous environmental samples by online solid-phase extraction–ultra-high-performance liquid chromatography-tandem mass spectrometry: optimisation and matrix effects reduction by quick, easy, cheap, effective, rugged and safe extraction. J Chromatogr A 1349:11–23. https://doi.org/10.1016/j.chroma.2014.05.006
Burin VM, Ferreira-Lima NE, Panceri CP, Bordignon-Luiz MT (2014) Bioactive compounds and antioxidant activity of Vitis vinifera and Vitis labrusca grapes: evaluation of different extraction methods. Microchem J 114:155–163. https://doi.org/10.1016/j.microc.2013.12.014
Callemien D, Collin S (2010) Structure, organoleptic properties, quantification methods, and stability of phenolic compounds in beer-a review. Food Rev Int 26:1–84
Campos-Requena V, Rivas B, Pérez M, Wilhelm M (2014) Application of design of experiments, response surface methodology and partial least squares regression on nanocomposites synthesis. Polym Bull 71:1961–1982. https://doi.org/10.1007/s00289-014-1166-6
Campos-Requena VH, Rivas BL, Pérez MA, Garrido-Miranda KA, Pereira ED (2015) Polymer/clay nanocomposite films as active packaging material: modeling of antimicrobial release. Eur Polym J 71:461–475. https://doi.org/10.1016/j.eurpolymj.2015.08.018
Carlson R, Nordahl Å, Barth T, Myklebust R (1991) An approach to evaluating screening experiments when several responses are measured. Chemom Intell Lab Syst 12:237–255. https://doi.org/10.1016/0169-7439(92)80004-N
Chiva-Blanch G, Urpi-Sarda M, Rotchés-Ribalta M, Zamora-Ros R, Llorach R, Lamuela-Raventós RM, Estruch R, Andrés-Lacueva C (2011) Determination of resveratrol and piceid in beer matrices by solid-phase extraction and liquid chromatography–tandem mass spectrometry. J Chromatogr A 1218:698–705. https://doi.org/10.1016/j.chroma.2010.12.012
Dejaegher B, Vander Heyden Y (2011) Experimental designs and their recent advances in set-up, data interpretation, and analytical applications. J Pharm Biomed Anal 56:141–158. https://doi.org/10.1016/j.jpba.2011.04.023
Delgado-Zamarreño MM, Pérez-Martín L, Bustamante-Rangel M, Carabias-Martínez R (2012) A modified QuEChERS method as sample treatment before the determination of isoflavones in foods by ultra-performance liquid chromatography–triple quadrupole mass spectrometry. Talanta 100:320–328. https://doi.org/10.1016/j.talanta.2012.07.070
Dvořáková M, Hulín P, Karabín M, Dostálek P (2007) Determination of polyphenols in beer by an effective method based on solid-phase extraction and high performance liquid chromatography with diode-array detection. Czech J Food Sci 25:182–188
Eriksson L, Johansson E, Wikström C (1998) Mixture design—design generation, PLS analysis, and model usage. Chemom Intell Lab Syst 43:1–24. https://doi.org/10.1016/S0169-7439(98)00126-9
Eriksson L, Johansson E, Kettaneh-Wold N, Wikstrom C, Wold S (2008) Design of experimental: principles and applications, 3rd edn. MKS Umetrics, Malmo
Fernandes PJ, Barros N, Câmara JS (2013) A survey of the occurrence of ochratoxin A in Madeira wines based on a modified QuEChERS extraction procedure combined with liquid chromatography–triple quadrupole tandem mass spectrometry. Food Res Int 54:293–301. https://doi.org/10.1016/j.foodres.2013.07.020
Fernández de Córdova ML, Medina AR (2014) Chapter 29 - analytical methods for determination of polyphenols in beer. In: Preedy V (ed) Processing and impact on antioxidants in beverages. Academic Press, San Diego, pp 289–299. https://doi.org/10.1016/B978-0-12-404738-9.00029-5
Ferreira SLC, Bruns RE, da Silva EGP, dos Santos WNL, Quintella CM, David JM, de Andrade JB, Breitkreitz MC, Jardim ICSF, Neto BB (2007) Statistical designs and response surface techniques for the optimization of chromatographic systems. J Chromatogr A 1158:2–14. https://doi.org/10.1016/j.chroma.2007.03.051
Floridi S, Montanari L, Marconi O, Fantozzi P (2003) Determination of free phenolic acids in wort and beer by coulometric array detection. J Agric Food Chem 51:1548–1554. https://doi.org/10.1021/jf0260040
Fontana AR, Bottini R (2014) High-throughput method based on quick, easy, cheap, effective, rugged and safe followed by liquid chromatography-multi-wavelength detection for the quantification of multiclass polyphenols in wines. J Chromatogr A 1342:44–53. https://doi.org/10.1016/j.chroma.2014.03.044
Herrero A, Ortiz MC, Sarabia LA (2013) D-optimal experimental design coupled with parallel factor analysis 2 decomposition a useful tool in the determination of triazines in oranges by programmed temperature vaporization–gas chromatography–mass spectrometry when using dispersive-solid phase extraction. J Chromatogr A 1288:111–126. https://doi.org/10.1016/j.chroma.2013.02.088
International Conference on Harmonisation (ICH) (2005) Harmonised tripartite guideline: validation of analytical procedures: text and methodology Q2 (R1)
Lehotay SJ, Son KA, Kwon H, Koesukwiwat U, Fu W, Mastovska K, Hoh E, Leepipatpiboon N (2010) Comparison of QuEChERS sample preparation methods for the analysis of pesticide residues in fruits and vegetables. J Chromatogr A 1217:2548–2560. https://doi.org/10.1016/j.chroma.2010.01.044
Mitić SS, Paunović DĐ, Pavlović AN, Tošić SB, Stojković MB, Mitić MN (2013) Phenolic profiles and total antioxidant capacity of marketed beers in Serbia. Int J Food Prop 17:908–922. https://doi.org/10.1080/10942912.2012.680223
Molina-García L, Ruiz-Medina A, Fernández-de Córdova ML (2011) A novel multicommuted fluorimetric optosensor for determination of resveratrol in beer. Talanta 83:850–856. https://doi.org/10.1016/j.talanta.2010.10.033
Pandey KB, Rizvi SI (2009) Plant polyphenols as dietary antioxidants in human health and disease. Oxidative Med Cell Longev 2:270–278
Piazzon A, Forte M, Nardini M (2010) Characterization of phenolics content and antioxidant activity of different beer types. J Agric Food Chem 58:10677–10683. https://doi.org/10.1021/jf101975q
Quifer-Rada P, Vallverdú-Queralt A, Martínez-Huélamo M, Chiva-Blanch G, Jáuregui O, Estruch R, Lamuela-Raventós R (2015) A comprehensive characterisation of beer polyphenols by high resolution mass spectrometry (LC–ESI-LTQ-Orbitrap-MS). Food Chem 169:336–343. https://doi.org/10.1016/j.foodchem.2014.07.154
Silva CL, Haesen N, Câmara JS (2012) A new and improved strategy combining a dispersive-solid phase extraction-based multiclass method with ultra high pressure liquid chromatography for analysis of low molecular weight polyphenols in vegetables. J Chromatogr A 1260:154–163. https://doi.org/10.1016/j.chroma.2012.08.082
Vinson JA, Mandarano M, Hirst M, Trevithick JR, Bose P (2003) Phenol antioxidant quantity and quality in foods: beers and the effect of two types of beer on an animal model of atherosclerosis. J Agric Food Chem 51:5528–5533. https://doi.org/10.1021/jf034189k
Zhao H, Chen W, Lu J, Zhao M (2010) Phenolic profiles and antioxidant activities of commercial beers. Food Chem 119:1150–1158. https://doi.org/10.1016/j.foodchem.2009.08.028
Acknowledgments
This work is part of Oscar Galarce-Bustos thesis to obtain the degree of Doctor in Science and Analytical Technology from the University of Concepcion, Chile. Authors want to thank to the National Commission of Scientific and Technological Research (CONICYT) of the Chilean Government for the doctoral scholarship granted.
Funding
This study was funded by the National Fund for Scientific and Technological Development (FONDECYT) project No. 1171857, by the National Fund for Scientific and Technological Equipment (FONDEQUIP) project No. 130209, and by the University of Concepcion.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
Oscar Galarce-Bustos declares that he has no conflict of interest. Lissette Novoa declares that she has no conflict of interest. Jessy Pavon declares that she has no conflict of interest. Karem Henríquez-Aedo declares that she has no conflict of interest. Mario Aranda declares that he has no conflict of interest.
Ethical Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed Consent
Not applicable.
Rights and permissions
About this article
Cite this article
Galarce-Bustos, O., Novoa, L., Pavon-Perez, J. et al. Chemometric Optimization of QuEChERS Extraction Method for Polyphenol Determination in Beers by Liquid Chromatography with Ultraviolet Detection. Food Anal. Methods 12, 448–457 (2019). https://doi.org/10.1007/s12161-018-1376-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12161-018-1376-x