Abstract
Background
Lung cancer in never smokers presents predominately as adenocarcinoma and in females. MicroRNA-183 (miR-183) has various expression patterns in types of human cancers. In the present study, we evaluated the expression of miR-183-3p in female lung adenocarcinoma and adjacent noncancerous tissues and explored its relationship with clinicopathological characteristics and prognosis.
Methods
In the present study, a hundred female nonsmoking patients who were newly diagnosed and histologically confirmed as lung adenocarcinoma at Tianjin Medical University Cancer Hospital were included. miR-183-3p expression of surgically removed NSCLC tissues and their corresponding normal lung tissues was measured by qRT-PCR assay. Associations of miR-183-3p expression with clinicopathological features were determined using the Student’s t test. Log-rank test, and Cox proportional hazards model were used for survival analysis.
Results
At first, miR-183-3p was up-regulated in lung cancer tissues when compared with the corresponding noncancerous lung tissues. Moreover, the expression of miR-183-3p in tumor tissue was found to be associated with lymph node metastasis (P = 0.043), clinical stage (P = 0.015), and EGFR mutation (P = 0.003). At last, high miR-183-3p expression was also associated with both poor overall survival and progression-free survival of women with lung adenocarcinoma (P = 0.005 and P = 0.010, respectively).
Conclusion
This study suggested that miR-183-3p expression might be involved in lung cancer pathogenesis and progression, and could be used as a potential prognostic biomarker of female lung adenocarcinoma.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nonsmoking lung cancer consists of approximately 20–25 % of all lung cancer cases [1, 2]. Evidence from the epidemiological and pathological studies has shown that lung cancers are quite different between smokers and never smokers [3, 4]. In recent years, the incidence of lung adenocarcinoma has been continuously rising, which is especially true for Asian women, who have a high percentage of nonsmokers and are more likely to respond to the target therapy [5]. New prognostic and predictive biomarkers are urgently needed for these patients.
MiRNAs are endogenous noncoding RNAs of 19–24 nucleotides in length. Abnormal expression of miRNAs has been associated with numerous diseases, especially cancer, and several studies have demonstrated the diagnostic or prognostic significance of miRNA expression in cancer [6, 7]. For example, we have reported that circulating level of miR-195 and miR-122 may have prognostic values in predicting the overall survival and EGFR mutation status in nonsmoking female patients with lung adenocarcinoma in our previous study [8]. However, few reports involve the clinical significance of miRNA tissue expression in lung adenocarcinoma of female nonsmokers. In the present study, we explored the role of tissue miR-183-3p in lung adenocarcinoma of female nonsmokers.
MiR-183 is located in q32.2 of chromosome 7 and it is a member of the miR-183 family, which consists of miR-96, miR-182, and miR-183. In addition, miR-183 has been identified as a potential oncogene in types of cancer, including NSCLC [9]. To understand better the clinical significance of miR-183 in NSCLC, we used real-time PCR to analyze the expression of miR-183-3p in tissue samples of nonsmoking women with lung adenocarcinoma in the present study, and our results showed that tissue miR-183-3p expression may be a potential biomarker for predicting the prognosis of lung adenocarcinoma.
Materials and methods
Patients and tissue samples
This study was endorsed by the Research Ethics Committee of Tianjin Medical University Cancer Hospital, China. Written informed consent was obtained from all patients. All specimens were handled and made anonymous according to the ethical and legal standards. All recruited patients underwent pulmonary surgery between 2007 and 2009 at Tianjin Medical University Cancer Institute and Hospital. Finally, a hundred female nonsmoking patients, who were newly diagnosed and histologically confirmed as lung adenocarcinoma patients were enrolled with a median follow-up of 34 months (range 4–72 months), and the mean age of patients at diagnosis was 58.56 ± 9.94 years. The clinicopathological features of all patients are summarized in Table 1.
The specimens were snap-frozen in liquid nitrogen immediately after resection and stored at −80 °C until the extraction of RNA. The normal adjacent tissues (NAT) were obtained from tumor-free margins, and were at least 5 cm away from the tumor site. All patients were followed after surgery through clinical visit and regular telephone contact. The final date of follow-up was August 20 in 2013. At the last follow-up, 64 patients were still alive, whereas 36 had died. Overall survival time was calculated from the date of the initial surgical operation to death. Progression-free survival was calculated from the date of the initial surgical operation to the date of tumor recurrence or distant metastases.
RNA extraction and QRT-PCR
Total RNA was isolated from surgical resection tissue samples using the TRizol reagent (Applied Biosystems, Foster City, CA, USA). RNA concentration and purity were measured using the Nanodrop ND-1000 (Agilent Biosystems TM, Santa Clara, CA). Only the samples with the OD A260/A280 ratio close to value of 2.0, which indicates that the RNA is pure, were eventually analyzed. All RNA samples were diluted into 50 ng/μL before our experiment.
MiR-183-3p and the small nuclear RNAU6B (Applied Biosystems, Foster City, CA, USA) expression in lung cancer and adjacent normal lung tissues were analyzed in triplicates by real-time quantitative PCR assay. RNA was processed by the TaqMan MicroRNA Reverse Transcription Kit (Applied Biosystems, Foster City, CA, USA) and the real-time RT-PCR was performed following the protocol described in the Ref. [10]. Following the RT, quantitative real-time PCR was performed in ABI 7900 Real-Time PCR System (Applied Biosystems, Foster City, CA, USA) at 95 °C for 10 min, followed by 40 cycles of 95 °C for 15 s and 60 °C for 1 min. The quantification cycle (Cq) was calculated using SDS 2.4 software (Applied Biosystems, Foster City, CA, USA). In addition, the expression of miR-183-3p in tissues was normalized with RNU6B using the 2−ΔΔCq method.
Statistical analyses
Statistical analyses were performed using the SPSS statistical software package, version 16.0 (SPSS Inc., Chicago, IL, USA) and Graphpad Prism 5.0 statistical software (Graphpad Software, La Jolla, CA). The value for miRNA expression levels were calculated based on the formula of \(\text {log}_{10^{2- \Delta \Delta {\text{Cq}}}}\). Data were expressed as mean ± SD. Differences in miR-183 levels between lung adenocarcinoma tissue and paired NAT samples were compared using a paired Student’s t test. MiR-183 levels between groups (inter-group differences) were compared using Independent two-sample Student’s t test. The Kaplan–Meier method was used for survival analysis, and differences in survival were estimated using the log-rank test. The Cox proportional hazards model for multivariate survival analysis was used to assess predictors related to survival. All tests were two tailed, and the significance level was set at P < 0.05.
Results
Overexpression of miR-183-3p in lung adenocarcinoma tissue
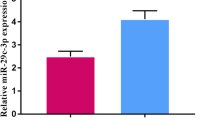
The qRT-PCR was performed to analyze the expression of the miR-183-3p in 24 pairs of lung adenocarcinoma and the corresponding noncancerous lung tissues. We showed that when compared with the adjacent normal lung tissues, the level of the miR-183 was significantly increased in lung cancer tissues (P = 0.036, Fig. 1).
MiR-183-3p expression in 24 pairs of clinical NSCLC and nontumorous lung tissues were respectively detected by real-time quantitative PCR assay. After normalization to RNU6B expression levels, the expression level of miR-183-3p in lung adenocarcinoma tissues (mean ± SD: 8.3 ± 0.4) was significantly higher than that in nontumorous tissues (mean ± SD: 7.9 ± 0.8, P = 0.036)
Correlation between miR-183-3p and clinicopathological features of lung adenocarcinoma
The associations of miR-183-3p expression with various clinicopathological parameters of patients with lung adenocarcinoma are summarized in Table 1. The lung adenocarcinoma patients with high miR-183-3p expression more frequently had advanced clinical stage (P = 0.015, Fig. 2a), positive lymph node metastasis (P = 0.043, Fig. 2b), and EGFR mutation (P = 0.003, Fig. 2c). However, the expression of miR-183-3p was not correlated with other factors of patients, including age, family history of cancer, history of previous lung disease, and tumor size (all P > 0.05).
Overexpression of miR-183-3p was associated with poor survival
The correlation of miR-183-3p expression with prognosis in lung adenocarcinoma patients was also investigated. As shown in Fig. 3, the overall survival of lung adenocarcinoma patients with high miR-183-3p expression was significantly poorer than those with low expression (P = 0.005, Fig. 3a), which was similar to the prognostic value of miR-183-3p in progression-free survival of lung adenocarcinoma patients (P = 0.010, Fig. 3b).
Kaplan–Meier curves for survival time in patients with lung adenocarcinoma patients divided according to miR-183-3p expression. The overall survival and progression-free survival of patients with high versus low miR-183-3p expression levels are shown: a overall survival rate (P = 0.005); b progression-free survival rate (P = 0.010). Asterisk indicates statistically significant
Bootstrap Cox proportional hazards regression analysis, at the univariate level, clinical stage, lymph node metastasis, and high miR-183-3p expression were significantly correlated with poor overall survival (P = 0.000, 0.001, and 0.011, respectively; Table 2) and progression-free survival (P = 0.000, 0.000, and 0.012, respectively; Table 2) of lung adenocarcinoma patients. Multivariate analysis using the Cox proportional hazard model indicated that high miR-183-3p expression was an independent prognostic factor for both overall survival (P = 0.016, Table 3) and progression-free survival of NSCLC patients (P = 0.010, Table 3).
Discussion
According to the recent studies, miRNAs may offer a new regulatory model of gene expression, and miRNA expression signatures correlate well with specific clinical characteristics of cancer, so that they can be used to classify normal and cancerous tissues, as well as for prognosis [11, 12].
MiR-183 expression in cancer is controversial, because recent studies have indicated that it is up-regulated or down-regulated in different cancer tissues as compared with normal tissues. For example, miR-183 expression is up-regulated in several cancer types, including epithelial carcinoma [13], breast cancer [14], bladder cancer [15], colorectal cancer [16], prostate cancer [17], gastric cancer [18], ovarian cancer [19], medullary thyroid cancer [20], as well as urothelial carcinoma [21], but it is down-regulated in osteosarcoma [22]. For lung cancer, up to now, only Zhu et al. has reported that the expression levels of miR-183 in lung cancer tumor and sera were higher than that of their normal comparisons. Moreover, they also found miR-183 expression level was correlated with histological classification, invasion to lung membrane and overall survival in patients with lung cancer [9]. Similarly, our results showed that miR-183-3p was significantly up-regulated in female lung adenocarcinoma cancer tissues than adjacent noncancerous tissues and overexpression of miR-183-3p was related with advanced clinical stage, lymph node metastasis, poor overall, and progression-free survival of lung adenocarcinoma. It thus suggested that miR-183-3p is a significant predictor of advanced progression and poor prognosis for lung adenocarcinoma patients. However, precise molecular mechanisms for the altered expression of miR-183 in NSCLC remain unclear thus far.
Several tumor-related genes have been identified to be the targets of miR-183 such as FOXO1, FGF9, CPEB1, VIL2, Dkk-3, and SMAD4. FOXO1 has been identified as the target of miR-183 in endometrial cancer, glioma, and classical Hodgkin lymphoma (cHL). FOXO1 was down-regulated and induced apoptosis in cHL cell lines and blocked proliferation, accompanied with cell cycle arrest in the G0/G1 phase [23]. Lam et al. has identified several miRs (including miR-183) overexpressed in endometrial cancer that function in concert to repress FOXO1 expression, resulting in de-regulated cell cycle control and impaired apoptotic responses, and this may be central to endometrial tumorigenesis [13]. The findings of Tang et al. showed that inhibition of the miR-183/96/182 cluster induced ROS-mediated AKT/survival independent of three target genes FGF9, CPEB1, and FOXO1, and inhibition of the miRNA cluster induced p53/apoptosis signaling, which was dependent on these same genes [24]. Dkk-3 and SMAD4 have been identified as potential target genes of miR-183 and miR-183 knockdown decreased cell growth and motility in PC cells and significantly decreased prostate tumor growth in in vivo nude mice experiments [25]. Miller et al. induced overexpression of miR-183 inhibited migration of breast cancer cells by targeting the VIL2 gene [26], and VIL2 gene has also been identified as the target of miR-183 in ovarian cancer, as well as lung cancer [27]. Therefore, the biological results of miRNA are dependent on the combined outcome of different target genes, which may help in explaining our results on the role of miR-183-3p in nonsmoking female lung adenocarcinoma.
More interestingly, our results indicated that miR-183 was significantly up-regulated in EGFR-positive group when compared with the EGFR-negative group for the first time. Abnormal expression of miR-183 noted in EGFR mutated adenocarcinoma has never been reported in lung adenocarcinoma. Subramanian et al. have revealed de-regulation of a miRNA network composed of miR-183–EGR1–PTEN by knocking down miR-183 in synovial sarcoma, rhabdomyosarcoma (RMS), and colon cancer cell lines, miR-183 is an important contributor to cell migration in these tumor types and has a potential oncogenic role through the regulation of EGR1 and PTEN [28]. Because the tumor suppressor gene PTEN (phosphatase and tensin homolog) encoding a phosphoinositide phosphatase, is a downstream molecule of EGFR pathway [29, 30], it is probably that the survival of lung adenocarcinoma patients with EGFR mutation might be improved using TKI and a miR-183 blocker in the future.
In summary, this study provides evidence that tissue miR-183-3p expression is associated with clinical factors and prognosis of lung adenocarcinoma patients of female nonsmokers. High tissue miR-83-3p expression may be an independent poor prognostic factor for survival in NSCLC patients. However, the molecular mechanisms underlying the role of this miRNA in cancer need to be addressed in future studies.
References
Siegel R, Naishadham D, Jemal A. Cancer statistics, 2012. CA Cancer J Clin. 2012;62:10–29.
Ferlay J, Shin H, Bray F, Forman D, Mathers C, Parkin D. Esti-mates of worldwide burden of cancer in 2008: GLOBOCAN 2008. Int J Cancer. 2010;127:2893–917.
Sun S, Schiller JH, Gazdar AF. Lung cancer in never smokers—a different disease. Nat Rev Cancer. 2007;7:778–90.
Yano T, Miura N, Takenaka T, Haro A, Okazaki H, Ohba T, et al. Never-smoking nonsmall cell lung cancer as a separate entity. Cancer. 2008;113:1012–8.
Toh CK, Gao F, Lim WT, Leong SS, Fong KW, Yap SP, et al. Never-smokers with lung cancer: epidemiologic evidence of a distinct disease entity. J Clin Oncol. 2006;24:2245–51.
Esquela-Kerscher ASF. Oncomirs—microRNAs with a role in cancer. Nat Rev Cancer. 2006;6:259–69.
Calin GA, Croce CM. MicroRNA signatures in human cancers. Nat Rev Cancer. 2006;6:857–66.
Zhang H, Su Y, Xu F, Kong J, Yu H, Qian B. Circulating microRNAs in relation to EGFR status and survival of lung adenocarcinoma in female non-smokers. PLoS One. 2013;8:e81408.
Zhu W, Liu X, He J, Chen D, Hunag Y, Zhang YK. Overexpression of members of the microRNA-183 family is a risk factor for lung cancer: a case control study. BMC Cancer. 2011;11:393.
Mei Q, Li X, Meng Y, Wu Z, Guo M, Zhao Y, et al. A facile and specific assay for quantifying microRNA by an optimized RT-qPCR approach. PLoS One. 2012;7:e46890.
Wang S, Zhao X, Wang J, Wen Y, Zhang L, Wang D, et al. Upregulation of microRNA-203 is associated with advanced tumor progression and poor prognosis in epithelial ovarian cancer. Med Oncol. 2013;30:681.
Võsa U, Vooder T, Kolde R, Fischer K, Välk K, Tõnisson N, et al. Identification of miR-374a as a prognostic marker for survival in patients with early-stage nonsmall cell lung cancer. Genes Chromosom Cancer. 2011;50:812–22.
Myatt SS, Wang J, Monteiro LJ, Christian M, Ho KK, Fusi L, et al. Definition of microRNAs that repress expression of the tumor suppressor gene FOXO1 in endometrial cancer. Cancer Res. 2010;70:367–77.
Chen L, Li Y, Fu Y, Peng J, Mo MH, Stamatakos M, et al. Role of deregulated microRNAs in breast cancer progression using FFPE tissue. PLoS One. 2013;8:e54213.
Han Y, Chen J, Zhao X, Liang C, Wang Y, Sun L, et al. MicroRNA expression signatures of bladder cancer revealed by deep sequencing. PLoS One. 2011;6:e18286.
Bandres E, Cubedo E, Agirre X, Malumbres R, Zarate R, Ramirez N, et al. Identification by real-time PCR of 13 mature microRNAs differentially expressed in colorectal cancer and non-tumoral tissues. Mol Cancer. 2006;5:29.
Schaefer A, Jung M, Mollenkopf HJ, Wagner I, Stephan C, Jentzmik F, et al. Diagnostic and prognostic implications of microRNA profiling in prostate carcinoma. Int J Cancer. 2010;126:1166–76.
Zheng WW, Huang CM, Xie JW, Zheng CH, Li P, Wang JB, et al. Expression of microRNA-183 in stage II (gastric cancer and its association with Ezrin protein. Zhonghua Wei Chang Wai Ke Za Zhi. 2012;15:723–6.
Li J, Liang S, Jin H, Xu C, Ma D. X. L. Tiam1, negatively regulated by miR-22, miR-183 andmiR-31, is involved in migration, invasion and viability of ovarian cancer cells. Oncol Rep. 2012;27:1835–42.
Mian C, Pennelli G, Fassan M, Balistreri M, Barollo S, Cavedon E, et al. MicroRNA profiles in familial and sporadic medullary thyroid carcinoma: preliminary relationships with RET status and outcome. Thyroid. 2012;22:890–6.
Yamada Y, Enokida H, Kojima S, Kawakami K, Chiyomaru T, Tatarano S, et al. MiR-96 and miR-183 detection in urine serve as potential tumor markers of urothelial carcinoma: correlation with stage and grade, and comparison with urinary cytology. Cancer Sci. 2011;102:522–9.
Zhu J, Feng Y, Ke Z, Yang Z, Zhou J, Huang X, et al. Down-regulation of miR-183 promotes migration and invasion of osteosarcoma by targeting Ezrin. Am J Pathol. 2012;180:2440–51.
Xie L, Ushmorov A, Leithäuser F, Guan HSC, Färbinger J, Pelzer C, et al. FOXO1 is a tumor suppressor in classical Hodgkin lymphoma. Blood. 2012;195:3503–11.
Tang H, Bian Y, Tu C, Wang Z, Yu Z, Liu Q, et al. The miR-183/96/182 cluster regulates oxidative apoptosis and sensitizes cells to chemotherapy in gliomas. Curr Cancer Drug Targets. 2013;13:221–31.
Ueno K, Hirata H, Shahryari V, Deng G, Tanaka Y, Tabatabai ZL, et al. microRNA-183 is an oncogene targeting Dkk-3 and SMAD4 in prostate cancer. Br J Cancer. 2013;108:1659–67.
Lowery AJ, Miller N, Dwyer RM, Kerin MJ. Dysregulated miR-183 inhibits migration in breast cancer cells. BMC Cancer. 2010;10:502.
Wang G, Mao W, Zheng S. MicroRNA-183 regulates Ezrin expression in lung cancer cells. FEBS Lett. 2008;582:3663–8.
Sarver AL, Li L. S. S. MicroRNA miR-183 functions as an oncogene by targeting the transcription factor EGR1 and promoting tumor cell migration. Cancer Res. 2010;70:9570–80.
Salmena L, Carracedo A, Pandolfi PP. Tenets of PTEN tumor suppression. Cell. 2008;133:403–14.
McCubrey JA, Steelman LS, Kempf CR, Chappell WH, Abrams SL, Stivala F, et al. Therapeutic resistance resulting from mutations in Raf/MEK/ERK and PI3K/PTEN/Akt/mTOR signaling pathways. J Cell Physiol. 2011;226:2762–81.
Acknowledgments
Research support: this study was supported by National Natural Science Foundation of China (No. 81372229) and Natural Science Foundation of Tianjin (No. 13JCYBJC23100).
Conflict of interest
The authors declared that they have no conflicts of interest to this work.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Xu, F., Zhang, H., Su, Y. et al. Up-regulation of microRNA-183-3p is a potent prognostic marker for lung adenocarcinoma of female non-smokers. Clin Transl Oncol 16, 980–985 (2014). https://doi.org/10.1007/s12094-014-1183-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-014-1183-9