Abstract
Aponogeton microphyllus, previously placed under the synonymy of A. undulatus, is recognized here as a distinct species based on morphology, chromosome number, and molecular phylogenetics (nuclear ribosomal internal transcribed (ITS) spacer region). Observations on the type and live specimens revealed morphological differences between the two species. Aponogeton microphyllus flowered regularly and set seeds. Aponogeton undulatus flowered rarely, did not set seeds, but showed formation of young plantlets on the inflorescence axis. Similarly, different chromosome numbers were recorded in Aponogeton microphyllus and the two forms of A. undulatus, viz., AF1 and AF2, which occur in distinct populations. Aponogeton microphyllus exhibited polysomaty with root-tip cells showing 2n=40, 42, and 44 chromosomes. The two forms of A. undulatus, AF1 and AF2, showed 2n=84 and 86 chromosomes, respectively. Based on the ITS data, both species occupied two separate clades. Plastid trnK intron region indicated a close relationship between both species. Our study suggests the need for comprehensive phylogenetic analyses of A. undulatus across its distribution range based on more advanced techniques such as high-throughput sequencing data to understand the A. undulatus species complex and to detect natural hybrids of this species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
The Cape pondweed family (Aponogetonaceae Planch.) is a monogeneric family comprising 60 species. These tuberous hydrogeophytes occur mainly in the tropical and subtropical regions of the Old World (POWO 2023). The species of this genus usually bear a simple or two-branched spike (fragrant biforked inflorescence in A. distachyos L.f.). Some species are also known for their unique leaves (fenestrate leaves in A. madagascariensis (Mirb.) H. Bruggen). The most important utility of the genus is its use as an ornamental freshwater aquarium plant (van Bruggen 1985).
The species of this genus are mostly concentrated in Africa, Madagascar, Asia, and Australia. In India, Yadav and Gaikwad (2003) carried out a taxonomic revision of the genus, describing seven species. Since then two new species, viz. A. nateshii S.R.Yadav (Yadav et al. 2015) and A. wolfgangianus S.R.Yadav (Yadav 2017), have been described. Very recently, Dey et al. (2021) re-collected A. lakhonensis A. Camus from the Poba Reserve Forest, Dhemaji District, Assam, after a gap of 123 years. At present, nine species occur in India, of which five are endemic (Yadav et al. 2015; Yadav 2017). Taxonomic delimitation has always been tricky in this genus on account of polymorphism and frequent hybridization (van Bruggen 1985). Consequently, species complexes are usually observed in nature. Hybridization has been described as ‘common’ in this genus, which includes mainly self-compatible species, several of which are also interfertile (Les and Philbrick 1993). Both natural and artificial hybrids have been reported. The natural hybrids are A. appendiculatus x A. undulatus, A. crispus x A. undulatus, and A. natans x A. undulatus (Les and Philbrick 1993; Gaikwad et al. 1998). Gaikwad et al. (1998) reported natural hybrids (A. appendiculatus x A. undulatus) from two places, Padubidri, Udupi Taluk, Karnataka, and Khodungullor, Thrissur District, Kerala. Both populations show characters intermediate between A. undulatus and A. appendiculatus. The natural hybrids from Padubidri produce many inflorescences, but fail to develop fruit and seeds because of the absence of stamens in flowers, and the plants propagate vegetatively by forming young plants on inflorescence-like axes as in A. undulatus and have submerged leaves as in A. appendiculatus (Gaikwad et al. 1998). The hybrids from Khodungullor also produce many inflorescences with normal flowers, but this population does not set seeds because of high pollen sterility and, hence, propagate vegetatively by formation of propagules on inflorescence-like axes (Gaikwad et al. 1998). Cytogenetical studies of the hybrids revealed diploid numbers of 2n=74 and 84 for the Khodungullor and Padubidri populations, respectively. Artificial hybrids (A. decaryi x A. satarensis, A. boivinianus x A. ulvaceus, and A. madagascariensis x A. ulvaceus) have also been generated and are found to be vigorous and flowering abundantly (van Bruggen 1985; Yadav 1995).
As a part of taxonomic studies on Indian Aponogeton, we collected some specimens from Bankati, on the way to Dudhwa Wildlife Sanctuary, Lakhimpur, Uttar Pradesh. After critical scrutiny of the relevant literature and examination of fresh material, we found that the specimens tallied with Roxburgh’s species A. microphyllus Roxb. van Bruggen (1985) reduced this species to a synonym of A. undulatus. Therefore, the aims of the present study were (1) to determine whether A. undulatus and A. microphyllus are distinct species, using morphology, karyology, and molecular phylogenetics and (2) to determine whether A. undulatus/A. microphyllus are part of a species complex.
2 Materials and methods
2.1 Taxonomy
Aponogeton microphyllus was collected from Bankati, near Dudhwa Wildlife Sanctuary, Lakhimpur, Uttar Pradesh. Aponogeton undulatus Form 1 (AF1) was collected from Kittur, Belgaum District, Karnataka, and A. undulatus Form 2 (AF2) from Thrissur District, Kerala. Flowering was not observed in AF1, whereas AF2 flowered regularly. Both the forms propagated vegetatively by formation of young plants on an inflorescence-like axis. Morphological analysis and description are based on observations made in the field and examination of live specimens. The identity of the specimen was confirmed by consulting protologs and types. Descriptions were made following the terminology of Hickey and King (2001).
2.2 Cytogenetics and pollen viability
Materials for cytogenetical studies were taken from the plants maintained in the Botanical Garden, Shivaji University, Kolhapur, Maharashtra, India. The voucher specimens of the cytogenetically examined plants were deposited in the Herbarium of the Department of Botany, Shivaji University, Kolhapur, India (table 1).
For mitotic studies, the root tips obtained from tubers were used after pre-treatment with an aqueous saturated solution of para-dichlorobenzene at 8–10°C for 4 to 5 h. Root tips were fixed in freshly prepared modified Carnoy’s fluid (3:1 of ethanol and propionic acid, respectively) for 24 h. Fixed roots were washed thoroughly in distilled water, hydrolyzed in 1 N HCl, and stained and squashed with 2% propionic-orcein. For meiotic studies, floral buds of appropriate size were fixed in Carnoy’s fluid. Anther smears were prepared in 2% propionic-orcein. Meiotic and pollen viability studies for AF1 could not be carried out as these plants do not flower. Suitable somatic and meiotic plates from freshly prepared slides were photographed with a LEICA DM 750 microscope.
Pollen viability was estimated through stainability using glyceroacetocarmine (1:1). A total of 2500 pollen were analyzed. Stained pollen were taken as fertile or viable, while shriveled and unstained pollen were scored as sterile or non-viable. Slides were photographed with a LEICA DM 750 microscope.
2.3 Molecular phylogeny
A total of eight Aponogeton species were collected from different regions in India. The plant material was identified, and voucher specimens deposited in the herbarium, Department of Botany, Shivaji University, Kolhapur, India (table 1).
DNA was extracted from either live plants or silica gel-dried plants by the CTAB method (Doyle and Doyle 1987). The nuclear ribosomal DNA internal transcribed spacers (ITS) and the plastid tRNA-Lys (trnK) intron along with partial maturase K (matK) gene were used for molecular phylogenetics analysis as in Chen et al. (2015). The ITS region was amplified using ITS4 and ITS5 primers (White et al. 1990) and the plastid (cp) trnK intron region was amplified using M-apo-f1 and M-apo-r1 primers (Chen et al. 2015).
PCR amplification was done using a 2X PCR master mix (APS Labs, Pune, India). Each 25 μL reaction consisted of 12.5 μL 2X master mix, 1 μL each of the forward and reverse primers (10 pmol/μL), and 1 μL of template DNA extracted by the CTAB method which was approximately 100 ng. The PCR cycle was as follows: 95°C for 3 min, followed by 35 cycles of 95°C for 30 s, 58°C for 30 s, and 72°C for 45 s, followed by a final extension for 5 min. The PCR products were visualized in 1% agarose gels stained with 0.5 μg/mL ethidium bromide. PCR product purification and Sanger sequencing were performed by Barcode Biosciences, Bengaluru, India. For one specimen of A. undulatus, the ITS PCR products were cloned into a TA-cloning vector and plasmids were isolated from 10 colonies and sequenced from both forward and reverse directions. PCR product cloning and sequencing of the cloned plasmid DNA was outsourced to Barcode Biosciences, Bengaluru, India.
Contigs from both forward and reverse strands were assembled using pregap4 and gap4 modules of the STADEN package ver. 2.0.0b11 (Bonfield and Whitwham 2010). The assembled contigs were separately aligned with sequences of Aponogeton available in the NCBI GenBank database (table 2) and aligned using the MUSCLE algorithm in Aliview ver. 1.28 (Larsson 2014). Alignments were checked and refined manually in Aliview.
The congruence of the two datasets was tested using the partition homogeneity test as implemented in PAUP ver. 4.0b10 (Swofford 2003), and the null hypothesis that the datasets are congruent was rejected (p=0.01). As p=0.01, we analyzed the two genes separately. The best model of sequence evolution was checked using jModelTest ver. 2.1.10 (Darriba et al. 2012). Bayesian phylogenetics analysis was performed using MrBayes ver. 3.2.7a (Ronquist et al. 2012) using the CIPRES portal (www.phylo.org) (Miller et al. 2010). The Bayesian phylogenetic (BP) analysis consisted of two independent runs of 4 chains of Metropolis-coupled Monte Carlo simulations run for 2 million generations sampling trees every 1000 generations. The convergence of the runs was monitored with effective sample sizes (ESS) which were greater than 200. The first 20% of the sampled trees were discarded as the burn-in and the remaining trees were summarized. Trees were visualized in FigTree ver. 1.4.4. Maximum likelihood (ML) phylogenetic analysis was performed using RAxML ver. 8.2.12 (Stamatakis 2014). ML bootstrap analysis was performed using the GTRGAMMA model under the non-parametric bootstrapping method with 100 replicates in RAxML.
3 Results
3.1 Cytogenetics and pollen viability
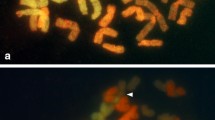
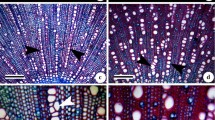
Aponogeton microphyllus exhibited polysomaty. Three counts with 2n=40, 42, and 44 chromosomes were observed (figure 1a–c). The chromosome number 2n=40 was the most frequent and observed in 47.62% of the nuclei, whereas 2n=42 and 44 was recorded in 30.16% and 22.22% of the nuclei, respectively. In addition to numerical alterations, structural alterations in chromosomes were observed. In one of them, i.e., 2n=40, two chromosomes with secondary constrictions were noted (figure 1a). Aponogeton undulatus AF1 and AF2 showed 2n=84 and 2n=86 chromosomes, respectively (figure 1e, f). Meiosis was normal in A. microphyllus and n=20 bivalents were observed at diakinesis (figure 1d). Aponogeton undulatus AF2 showed n=43 bivalents at diakinesis (figure 1g) with meiotic anomalies such as bridge formation, precocious separation, and laggards (figure 1h–l).
Aponogeton microphyllus exhibited high (98%) pollen viability (figure 2a) and seed set (figure 3g). AF2 had very low (30%) pollen viability (figure 2b) with no seed set. These plants propagated vegetatively by the formation of propagules (figure 3m, n).
Karyotypes in two species of Aponogeton (a, b, c) Mitotic metaphases showing 2n=40, 42 and 44 chromosomes, respectively (A. microphyllus), note secondary constrictions (arrowheads); (d) pollen mother cell (PMC) at diakinesis showing n=20 bivalents (A. microphyllus); (e) Mitotic metaphase showing 2n=84 (A. undulatus AF1); (f) Mitotic metaphase showing 2n=86 chromosomes (A. undulatus AF2); meiotic stages in A. undulatus AF2 (g–l); (g) PMC at diakinesis showing n=43 bivalents; (h, i) PMCs at metaphase I showing precocious separation (arrowheads); (j) PMC at anaphase I showing a bridge (arrowhead); (k) PMC at metaphase II showing precocious separations (arrowheads); (l) PMC at telophase II showing bridge and laggards (arrowheads). Scale bars: 5 μm.
Comparison of morphological features of A. microphyllus and A. undulatus (AF2). (a–h) A. microphyllus. (a) Inflorescence; (b) tepals; (c) stamen; (d) infructescence; (e) gynoecium; (f) fruit; (g) seed; (h) embryo; (i–n) AF2; (i) inflorescence; (j) tepals; (k) stamen; (l) gynoecium; (m) inflorescence with a young plantlet (propagule); (n) the young plantlet (propagule) forming a new individual.
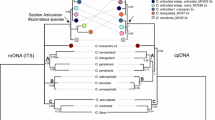
Bayesian phylogenetic tree based on the ITS region. Numbers above branches are posterior probabilities and maximum-likelihood bootstrap value, respectively. Sequences generated for this study are indicated in bold text. GenBank accession numbers are indicated beside taxon names. Scale indicates the number of substitutions along the branch lengths. Branches are colour-coded according to their biogeographic distribution. (Orange: S/S.E. Asia, Pink: Tropical Australasia, Green: Madagascar, Blue: Continental and E. Africa, Red: S.W. Africa.)
Bayesian phylogenetic tree based on the plastid trnK intron. Numbers above branches are posterior probabilities and maximum-likelihood bootstrap values, respectively. Sequences generated for this study are indicated in bold text. GenBank accession numbers are indicated beside taxon names. Scale indicates number of substitutions along the branch lengths. Branches are colour-coded according to their biogeography. (Orange: S/S.E. Asia, Pink: Tropical Australasia, Green: Madagascar, Blue: Continental and E. Africa, Red: S.W. Africa.)
3.2 Molecular phylogeny
The aligned ITS dataset consisted of 111 accessions and 837 nucleotides. There were 564 variable sites, of which 462 were parsimony-informative. The cp trnK intron dataset consisted of 86 accessions and 1010 nucleotides of which 233 were variable and 176 were parsimony-informative. The BP and ML trees were similar in topology (ML tree not shown). The ITS tree (figure 4) was better resolved than the plastid trnK intron tree (figure 5). The best-fit model of sequence evolution for the ITS dataset and cpDNA datasets was GTR+G and SYM+G, respectively.
In the ITS dataset tree, the three different accessions of A. microphyllus formed a well-supported clade. This clade also contained samples named A. stachyosporus de Wit which were reported in other studies (AY926306 and AY926303 from Les et al. 2005) (table 3). In the same clade, one sequence from a cloned ITS region obtained from A. undulatus (A3 specimen, clone no. 3) collected from Thrissur, Kerala, India, was recovered.
The other clones of the same specimen, A3 (RNC-269), collected as A. undulatus from Kerala, and whose chromosome number indicates high ploidy formed a clade along with A. stachyosporus (AY926305, AY926304) and A. undulatus (AY926302) from Les et al. (2005). Another specimen, 10, collected as A. undulatus from Wayanad, Kerala, formed a close relationship with A. crispus Thunb. specimens although not strongly statistically supported. The occurrence of one isoform of ITS from A3 specimen of A. undulatus Roxb. in the A. microphyllus clade indicates the hybrid nature of this specimen.
The plastid trnK dataset was not well resolved, and the posterior probability (PP) as well as ML bootstrap (BS) supports of the majority of the branches were not significantly high. This might be due to fewer parsimony-informative characters (figure 5). The samples of A. microphyllus nested within A. undulatus with shallow branch lengths but with good branch supports. Aponogeton wolfangangianus was also found closely associated with A. undulatus in the trnK data. This clade was the closest in relation to another clade consisting of A. womersleyi H. Bruggen and A. euryspermus Hellq. & S.W.L. Jacobs and A. lakhonensis A. Camus. This sister relationship between these two clades was not well supported. The sample A3 used for ITS cloning and sequencing was in the A. undulatus clade, proving the maternal lineage of the hybrid.
3.3 Morphological study
Aponogeton microphyllus Roxb., Fl. Ind. Ed. Carey (1832) 2:211 (figures 3a–h and 6)
Type Roxb. Icon No.1232
Description Tuberous, monoecious, perennial, aquatic herbs. Tubers globose to obovoid, 1–2 cm, roots fibrous. Leaves both submerged and floating; submerged leaves petiolate, petiole ca. 15 cm long, lamina ca. 10×4 cm, elliptic lanceolate, obtuse or rounded at apex, base rounded or rarely cordate, leaf margin undulate, main nerve with 6–7 parallel nerves; floating leaves petiolate, petiole ca. 20–25 cm long, lamina ca. 8×3 cm, elliptic lanceolate, obtuse apex, base rounded, main nerve with 5–6 parallel nerves. Inflorescence a single spike, spatheate, spathe persistant, peduncle ca. 22–25 cm long, cylindrical; spike simple, 4–8 cm long, flowers turned in all directions on axis. Flowers bisexual, tepals 2, 4–7×2 mm, oblong-obovate, obtuse, 1-nerved, caducous, white coloured. Stamens 6, exserted, filamentous; filaments 1.5×1.7 mm long, narrow above and widened towards base; anthers 2-celled, globose, basifixed, yellow coloured. Carpels 3, 0.8–1.5×0.5–0.8 mm, ovules 2, basal. Follicles 5–7×4 mm with short, terminal, curved beak. Seeds ca. 5×2 mm with simple testa, embryo ca. 4–5×1.5–2 mm, plumule attached at the base with young foliage leaves.
Phenology flowering and fruiting from May to September
Aponogeton undulatus Roxb., Fl. Ind. Ed. Carey (1832) 2:211 (figures 3i–n, 7)
Type Roxb. Icon No. 936
Description Tuberous, monoecious, perennial, aquatic herbs. Tubers globose to obovoid 1–4×1–2 cm, roots fibrous. Leaves both submerged and floating; submerged leaves petiolate, petiole ca. 15–30 cm long, lamina ca. 9–15×1.5–2.5 cm, linear-lanceolate, acute apex, base cuneate, leaf margin undulate, main nerve with 5–7 parallel nerves; floating leaves petiolate, petiole ca. 25–30 cm long, lamina ca. 10×4 cm, elliptic lanceolate, obtuse apex, base rounded, main nerve with 5–6 parallel nerves. Inflorescence a single spike, spatheate, spathe caducous, peduncle ca. 25–30 cm long, cylindrical; spike simple, 4–6 cm long, flowers turned in all directions on axis. Flowers bisexual, tepals 2, 2.5–3×2 mm, oblong-obovate, rounded, 1-nerved, caducous, pinkish-white coloured. Stamens 6, exserted, filamentous; filaments 1.4×1.5 mm long, narrow above and widened towards base; anthers 2-celled, globose, basifixed, dark bluish to black coloured. Carpels 3, 1.2×0.8 mm, free, unilocular.
Phenology flowering from May to September, fruiting absent.
The differences between A. microphyllus and A. undulatus are listed in table 3.
4 Discussion
4.1 Cytogenetics and pollen viability
The chromosome count in Aponogeton varies from 2n=16 (A. distachyos L.f. and A. madagascariensis (Mirb.) H. Bruggen) to 2n=~126 (A. junceus) (van Bruggen 1985). Other chromosome counts recorded in this genus are 2n=24, 26, 32, 38, 40, 48, 50, 52, 56, 60, 68, 70, 74, 76, 78, 80, 92, and ~100 (van Bruggen 1985; Gaikwad et al. 2014).
Aponogeton undulatus shows tremendous variation in chromosome number with 2n=~70, ~74, and 86 (van Bruggen 1985; Gaikwad 2000; Rice et al. 2014). The count of 2n=32 cannot be verified as it lacks voucher details (see Rice et al. 2014) and this could be a wrong identification. Non-flowering populations of A. undulatus collected from Bhandara District, Maharashtra, and Dastikoppa, Dharwad District, Karnataka, had 2n=70 chromosomes, whereas flowering populations of A. undulatus from Kittur, Belgaum District, Karnataka, and Ghadbadya Lake, near Amgaon, Bhandara District, Maharashtra, had 2n=74 and 2n=86 chromosomes, respectively (Gaikwad 2000). In our studies, we observed 2n=84 chromosomes in AF1 and 2n=86 chromosomes in AF2. Meiotic anomalies such as chromosome clumping and laggards have been reported in A. undulatus flowering individuals (2n=86) (Gaikwad 2000). We also observed meiotic anomalies such as bridge formation, precocious separation, and laggards in AF2. Consequently, AF2 showed very low (38%) pollen viability.
Polysomaty, the occurrence of different chromosome numbers in the cells of the same root tip, has been reported in many monocots such as Lilium Tourn. ex L., Drimiopsis Lindl. & Paxton, Ornithogalum L., Urginea Steinh., Allium L., Cymbidium Sw., Bulbophyllum auricomum Lindl., Cymodocea nodosa (Ucria) Asch., and Dendrobium pierardii R.Br. In the majority of cases, aneuploidy was observed frequently (Sen 1973; Fukai et al. 2002; Than et al. 2011; Gargiulo et al. 2020). Recently, Gargiulo et al. (2020) reported aneusomaty and polysomaty in Cymodocea nodosa (Cymodoceaceae). Similarly, polysomaty and chromosome number variation has been observed in the freshwater plant Ceratophyllum demersum L. by Gargiulo et al. (2022), who reported euploid and aneuploid (2n=24, 48, 52, 62, 72, 78, 82, 86, 92, 98) chromosome numbers as well as nuclei with different amounts of DNA (2C, 4C, 6C, 8C, 10C) in a single plant. Gargiulo et al. (2020, 2022) suggested that both endoreduplication and regular or anomalous mitoses together were responsible for the formation of nuclei with diverse chromosome numbers and ploidy levels in somatic cells. In the present investigation, we observed polysomaty in the root-tip cells of A. microphyllus (2n=40, 42, and 44) along with structural changes involving secondary constrictions. It has been suggested that the nucleocytoplasmic ratio in such polysomaty cases is maintained in the tissue taken as a whole, that is, between the entire cytoplasm and the nuclear complement of the tissue, rather than of the individual cells (Sen 1973). The exact cause of polysomaty in A. microphyllus is not known as of now. The data on cytogenetics and pollen viability suggest that both species are genetically different. Aponogeton microphyllus has low chromosome number and high pollen viability and reproduces sexually. On the other hand, A. undulatus has high chromosome number, polymorphism, and low pollen viability, and propagates vegetatively, indicating its hybrid origin.
4.2 Molecular phylogeny
The molecular phylogeny of the genus was constructed based on one nuclear marker and one plastid DNA marker, ITS and trnK intron, respectively. A total of 45 species and 36 species was included in the nuclear ITS and cp trnK intron data, respectively.
According to the ITS dataset, A. undulatus and A. microphyllus fall into two separate clades. However, in cpDNA data, A. undulatus and A. microphyllus show a close sister relationship. The close affinity of A. undulatus and A. microphyllus might have led to natural hybridization, and some specimens collected for this study might be those hybrids. Specimen A3, collected as A. undulatus, shows at least two isoforms of the ITS gene sequence. Among the nine sequences obtained by cloning the PCR product, one (A3) was found nested in the A. microphyllus clade (figure 4). Similar natural hybrids have been reported in previous works but without molecular sequence data (Les et al. 2005). Hybridization followed by polyploidization as a source of speciation has been reported in several plants (Soltis and Kuzoff 1995; Ackerman and Wen 2003; Kim and Donoghue 2008; Surveswaran et al. 2018).
Although A. undulatus and A. microphyllus have been synonymized as A. undulatus by van Bruggen (1985), several morphological features, karyology, and molecular phylogenetics using nuclear ITS provide evidence that they are distinct species. However, due to their close evolutionary history, as seen in the maternally inherited trnK intron data, the species might be able to hybridize naturally.
Some specimens collected as A. stachyosporus (Les et al. 2005) fall under the A. undulatus clade as well as the A. microphyllus clade in our study (figure 4). However, in the cpDNA phylogeny (figure 5), A. stachyosporus is placed along with A. undulatus. Les et al. (2005) have similar findings. They found that cpDNA sequences were identical in A. undulatus and A. stachyosporus; however, ITS sequences of the species differed by seven substitutions, indicating their distinctness. This might be the reason why Chen et al. (2015) used the name A. stachyosporus in their phylogeny (based on the combined nuclear and plastid data) but indicated that the species is synonymous with A. undulatus. A perusal of the protolog of A. stachyosporus revealed that the species was collected from Johore (now Johor, a state in Malaysia) from the Malay Peninsula and exhibited vivipary (de Wit 1958). The plants produced a digitately branched inflorescence, giving rise to reduced sterile inflorescences and young plants (de Wit 1958). The morphology of A. stachyosporus matches completely with the AF2, A3, and Ma specimens in our study. Therefore, the results of the present study provide clear evidence that A. stachyosporus is nothing but A. undulatus, which is a hybrid species.
In the ITS phylogeny, the widespread A. crispus, A. rigidifolius, and A. wolfgangianus were found in a close relationship, and one specimen of A. nateshii was also found inside this clade. The relationships between these taxa are well resolved by both nuclear and plastid markers. Aponogeton natans was found closely related to A. brugennii in the ITS data. Another widespread species, A. lakhonensis, was found in a clade consisting of Australian and Asian species with good branch support. There are some incongruencies between the cpDNA and ITS datasets, as expected (Reisberg and Soltis 1991; Doyle 1992). It was also found that in the ITS tree, one of the forms, i.e., A. undulatus (form 4), is sister to A. crispus, which indicates that these species may interbreed to produce hybrids.
The South/Southeast Asian species form a clade in both nuclear ITS and cpDNA datasets which is weakly supported in the cpDNA dataset (0.79/67) and only well supported by Bayesian posterior probability in the ITS dataset. In the cpDNA data, Australian A. eurysperumus and A. womersleyi are included in the South/Southeast Asian clade and this clade has relatively low support (only ML bootstrap support). The paraphyletic A. undulatus nested with A. microphyllus and sometimes in A. wolfgangianus. The ITS/plastid lineages of the A. crispus hybrids are more closely related to A. natans than to other A. crispus accessions themselves.
The close association of the A. undulatus dataset with A. wolfgangianus in the cpDNA dataset indicates a phylogenetic close relationship of these species. To further understand reticulation events in these closely related species, much more sequence data are required. To cite an example, in Lachemilla (Focke) Rydb. (Rosaceae), to resolve the incongruence between nuclear and cpDNA markers, a dense sampling of 396 nuclear loci by target capture approach and nearly complete plastome sequences from 27 species were used to resolve the relationships among the major groups. The data also helped understand the multiple sources of conflict between gene trees and species trees inferred with a multitude of approaches (Morales-Briones et al. 2018). Such an approach should be applied in Aponogeton, after sampling the genus from all over the world. In any case, the specimens of A. undulatus and A. microphyllus presented a major challenge in obtaining the sequences for our study. This might be due to their hybrid nature and varying ploidy levels. Therefore, a complete sampling of the specimens in various locations as well as reduced representation genome sequencing followed by phylogeny analysis are very important for a future study.
4.3 Taxonomy
Roxburgh (1832) described two species, A. microphyllus (figure 6) and A. undulatus (figure 7), but the former was synonymized under A. undulatus by van Bruggen (1985). He could not find any specimen of A. microphyllus and concluded that A. microphyllus represents a poor specimen of A. undulatus. Subsequent workers (Yadav and Gaikwad 2003) also considered A. microphyllus conspecific to A. undulatus. Roxburgh (1832) described the flowers as blue and the spathe as caducous, but his drawing (figure 6) depicts white flowers and a persistent spathe. This was also pointed out by van Bruggen (1985). Our observations revealed that A. microphyllus has white flowers and a persistent spathe, whereas A. undulatus has pinkish-white flowers and a caducous spathe. van Bruggen (1985) observed that A. undulatus rarely flowers in cultivation. He also observed the formation of a forked inflorescence, proliferation, and malformations of the inflorescence in the A. undulatus. de Wit (1958) also recorded branched inflorescence and propagules in A. undulatus. Yadav and Gaikwad (2003), based on their collections of A. undulatus from different localities in India, made similar observations. They found that some of these populations produced 1–3 inflorescences on the same axis on which propagules had developed and flowered regularly; however, these plants failed to set seeds, whereas in some populations flowers were not observed (Yadav and Gaikwad 2003). Our observations are consistent with those of van Bruggen (1985) and Yadav and Gaikwad (2003). Aponogeton undulatus AF1 flowers rarely, and therefore, the species has adapted vegetative propagation by forming a propagule, whereas A. undulatus AF2 flowers regularly but fails to set seeds. Aponogeton undulatus AF2 sometimes produces branched inflorescences that develop propagules which, when detached, form a new plant (figure 3m, n). Aponogeton microphyllus flowers regularly and sets seeds (figure 3g).
In conclusion, based on morphological studies, chromosome numbers, and molecular phylogenetic analysis of the ITS region, our study confirms that A. microphyllus is not conspecific to A. undulatus. Our studies also suggest that A. undulatus has a hybrid origin. There is a possibility that four species, viz., A. appendiculatus, A. crispus, A. microphyllus, and A. undulatus, hybridize whenever their distribution ranges overlap. Overall, it can be said that the Indian specimens of Aponogeton would provide an interesting model to study hybrid speciation and karyology by employing tools of molecular biology and techniques such as flow cytometry.
References
Ackerfield J and Wen J 2003 Evolution of Hedera (the Ivy Genus, Araliaceae): Insights from chloroplast DNA data. Int. J. Plant. Sci. 164 593–602
Bonfield JK and Whitwham A 2010 Gap5—editing the billion-fragment sequence assembly. Bioinformatics 26 1699–1703
van Bruggen HWE 1985 Monograph of the genus Aponogeton (Aponogetonaceae). Bioblioth. Bot. 33 1–76
Chen LU, Grimm GW, Wang QF, et al. 2015 A phylogeny and biogeography analysis for the Cape-Pondweed family Aponogetonaceae (Alismatales). Mol. Phylogenet. Evol. 82 111–117
Darriba D, Taboada GL, Doallo R, et al. 2012 jModelTest 2: more models, new heuristics and parallel computing. Nat. Methods 9 772
Doyle JJ 1992 Gene trees and species trees: molecular systematics as one character taxonomy. Syst. Biol. 17 144–163
Doyle JJ and Doyle JL 1987 A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem. Bull. 19 11–15
de Wit HCD 1958 Aponogeton stachyosporous sp. nov. Meded. Bot. Tuinen Belmonte Arbor. Wageningen. 2 96
Dey D, Yadav SR and Devi N 2021 Rediscovery of Aponogeton lakhonensis A. Camus (Aponogetonaceae): a long-lost aquatic plant of India. J. Threat. Taxa 13 19632–19635
Fukai S, Hasegawa A and Goi M 2002 Polysomaty in Cymbidium. HortScience 37 1088–1091
Gaikwad SP 2000 Studies on biology of some endemic aquatic monocots. Ph.D. Thesis, Department of Botany, Shivaji University, Kolhapur, India
Gaikwad SP, Sardesai MM and Yadav SR 1998 Naturliche hybriden (?) zwischen Aponogeton appendiculatus HWE van Bruggen und A. undulatus Roxburgh und ihr Nutzen fur die Aquaristik. Aqua Planta 2 54–67
Gaikwad SP, Yadav SR, Gore RD, et al. 2014 Karyomorphological studies of Aponogeton appendiculatus Bruggen and Aponogeton crispus Thunb. J. Natn. Sci. Foundation Sri Lanka 42 163–167
Gargiulo GM, Vilardo I, Gemelli F, et al. 2020 Aneusomaty and polysomaty in Cymodocea nodosa (Ucria) Ascherson from Mediterranean Sea (Sicily, Italy). Aquat. Bot. 162 103206
Gargiulo GM, Balkkouri BEl, Crisafulli A, et al. 2022 Polysomaty and chromosome number variation in a population of Ceratophyllum demersum L. from Aquila Lake (Aspromonte Mountains, Calabria, Italy). Aquat. Bot. 180 103530
Hickey M and King C 2001 The Cambridge illustrated glossary of botanical terms (Cambridge: Cambridge University Press)
Kim S and Donoghue MJ 2008 Incongruence between cpDNA and nrITS trees indicates extensive hybridization within Eupersicaria (Polygonaceae). Am. J. Bot. 95 1122–1135
Larsson A 2014 AliView: a fast and lightweight alignment viewer and editor for large data sets. Bioinformatics 30 3276–3278
Les DH and Tippery NP 2013 In time and with water…the systematics of alismatid monocotyledons. in Early events in monocot evolution. Eds. P. Wilkinand S. J Mayo, (Cambridge, United Kingdom: Cambridge University Press). pp 118–164
Les DH and Philbrick CT 1993 Studies of hybridization and chromosome number variation in aquatic angiosperms: evolutionary implications. Aquat. Bot. 44 181–228
Les DH, Crawford DJ, Kimball RT, et al. 2003 Biogeography of discontinuously distributed hydrophytes: A molecular appraisal of intercontinental disjunctions. Int. J. Plant Sci. 164 917–932
Les DH, Moody ML and Jacobs SWL 2005 Phylogeny and systematics of Aponogeton (Aponogetonaceae): the Australian species. Syst. Bot. 30 503–519
POWO 2023 Plants of the World Online (http://www.plantsoftheworldonline.org)
Morales-Briones DF, Liston A and Tank DC 2018 Phylogenomic analyses reveal a deep history of hybridization and polyploidy in the Neotropical genus Lachemilla (Rosaceae). New Phytol. 218 1668–1684
Miller MA, Pfeiffer W and Schwartz T 2010 Creating the CIPRES Science Gateway for inference of large phylogenetic trees. 2010 Gateway Computing Environments Workshop (GCE) pp 1–8
Rieseberg LH and Soltis DE 1991 Phylogenetic consequences of cytoplasmic gene flow in plants. Evol. Trends Plants 5 65–83
Rice A, Glick L, Abadi S, et al. 2014 The Chromosome Counts Database (CCDB) – a community resource of plant chromosome numbers. New Phytol. 206 19–26
Ronquist F, Teslenko M, van der Mark P, et al. 2012 MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61 539–542
Roxburgh W 1832 Flora Indica; or, descriptions of Indian Plants 2 (Thacker & Co., Serampore, Calcutta and Allen & Co., Parbury, London) p 211
Swofford DL 2003 PAUP* Phylogenetic analysis using parsimony (*and other methods). Version 4 (Sinauer Associates, Sunderland, Massachusetts)
Sen S 1973 Polysomaty and its significance in Liliales. Cytologia 38 737–751
Stamatakis A 2014 RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30 1312–1313
Soltis DE and Kuzoff RK 1995 Discordance between nuclear and chloroplast phylogenies in the Heuchera Group (Saxifragaceae). Evolution 49 727–742
Surveswaran S, Gowda V and Sun M 2018 Using an integrated approach to identify cryptic species, divergence patterns and hybrid species in Asian ladies’ tresses orchids (Spiranthes, Orchidaceae). Mol. Phylogenet. Evol. 124 106–121
Than MMM, Pal A and Jha S 2011 Chromosome number and modal karyotype in a polysomatic endangered orchid, Bulbophyllum auricomum Lindl., the Royal Flower of Myanmar Plant Syst. Evol. 294 167–175
White TJ, Bruns T, Lee S and Taylor J 1990 Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics; in PCR protocols: A guide to methods and applications (Eds.) MA Innis, DH Gelfand, JJ Sninsky, et al. (Academic Press, San Diego) pp 315–322
Yadav SR 2017 A new species of Aponogeton (Aponogetonaceae) from India with critical notes on embryo morphology. Phytotaxa 328 83–89
Yadav SR and Gaikwad SP 2003 A revision of the Indian Aponogetonaceae. Bull. Bot. Surv. India. 45 39–76
Yadav SR, Patil VS, Gholave AR, et al. 2015 Aponogeton nateshii (Aponogetonaceae): a new species from India. Rheedea 25 9–13
Yadav SR 1995 Die hybrid zwishen Aponogeton decaryi Jumelle and Aponogeton satarensis Raghavan, Kulkami and Yadav. Aqua Planta 20 71–80
Acknowledgements
The authors are grateful to the Head, Department of Botany, Shivaji University, Kolhapur, for providing necessary research facilities. Thanks are due to Dr. Susanne S. Renner, Honorary Professor of Biology, Washington University in St. Louis, Missouri, for going through previous versions of the manuscript and improving its content. SS thanks Pooja Kumari Shukla for help in sequence data analysis. SRY thanks Mr. Pushpendra Katiyar, NBRI, Lucknow, for his help during the field tour. SRY is grateful to University Grants Commission (UGC), New Delhi, for the award of the BSR faculty fellowship. We thank the Director, Royal Botanic Gardens, Kew, for permitting us to publish the selected images.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Authors have no conflict of interest to declare.
Additional information
Corresponding editor: R Geeta
Rights and permissions
About this article
Cite this article
Chougule, R.N., Surveswaran, S., George, A. et al. Roxburgh was right: Aponogeton microphyllus and Aponogeton undulatus are distinct species. J Biosci 48, 53 (2023). https://doi.org/10.1007/s12038-023-00366-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12038-023-00366-y