Abstract
In Arabidopsis thaliana (Arabidopsis) treated with the harpin protein HrpNEa, resistance to the green peach aphid Myzus persicae, a generalist phloem-feeding insect, develops with induced expression of the AtMYB44 gene. Special GLUCAN SYNTHESIS-LIKE (GSL) genes and β-1,3-glucan callose play an important role in plant defence responses to attacks by phloem-feeding insects. Here we report that AtGLS5 and AtMYB44 are both required for HrpNEa-induced repression of M. persicae feeding from the phloem of Arabidopsis leaves. In 24 h successive surveys on large-scale aphid populations, the proportion of feeding aphids was much smaller in HrpNEa-treated plants than in control plants, and aphids preferred to feed from the 37 tested atgsl mutants rather than the wild-type plant. The atgsl mutants were generated previously by mutagenesis in 12 identified AtGSL genes (AtGSL1 through AtGSL12); in the 24 h survey, both atgsl5 and atgsl6 tolerated aphid feeding, and atgsl5 was the most tolerant. Consistently, atgsl5 was also most inhibitive to the deterrent effect of HrpNEa on the phloem-feeding activity of aphids as monitored by the electrical penetration graph technique. These results suggested an important role of the AtGSL5 gene in the effect of HrpNEa. In response to HrpNEa, AtGSL5 expression and callose deposition were induced in the wild-type plant but not in atgsl5. In response to HrpNEa, moreover, the AtMYB44 gene known to be required for repression of aphid reproduction on the plant was also required for repression of the phloem-feeding activity. Small amounts of the AtGSL5 transcript and callose deposition were detected in the atmyb44 mutant, as in atgsl5. Both mutants performed similarly in tolerating the phloem-feeding activity and impairing the deterrent effect of HrpNEa, suggesting that AtGSL5 and AtMYB44 both contributed to the effect.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Harpins are multifunctional proteins produced by Gram-negative plant pathogenic bacteria (Kim and Beer 2000; Peng et al. 2004; Liu et al. 2006; Chen et al. 2008). HrpNEa is the first-characterized (Wei et al. 1992), well-studied harpin (Dong et al. 1999, 2004, 2005a; Kim and Beer 2000; Peng et al. 2003; Zhang et al. 2007; Ren et al. 2008; Liu et al. 2010a,b; Sun et al. 2010) secreted by Erwinia amylovora, the bacterial pathogen that causes fire blight disease in rosaceous plants (van der Zwet and Beer 1999). Harpins’ multiple functions, especially in eliciting plant defence responses, were also elucidated initially by studies using HrpNEa as a paradigm (Wei et al. 1992; Dong et al. 1999; Kim and Beer 2000). Early studies demonstrated that the external application of HrpNEa was able to induce resistance in a variety of plant species (Dong et al. 1999, 2004, 2005a; Peng et al. 2003; Ren et al. 2008), and that the induced resistance effectively protected plants from attacks by herbivorous insects (Zitter and Beer 1998; Dong et al. 2005a; Gou et al. 2009; Liu et al. 2010a). HrpNEa-induced resistance to insects was first suggested on the basis of observations of field-grown peppers. Plants that had been treated with HrpNEa incurred fewer injuries from the European corn borer Ostrinia nubilalis Hubner than comparable untreated plants (Kim and Beer 2000). A deterrent effect on the striped cucumber beetle Acalymma vittatum Fabricius was observed in HrpNEa-treated cucumber; striped cucumber beetles preferred to colonize untreated control plants rather than HrpNEa-treated plants (Zitter and Beer 1998). HrpNEa-induced resistance was also effective in impeding infestations of aphids, which represent typical phloem-feeding herbivores (Dong et al. 2005a; Gou et al. 2009; Liu et al. 2010a). In cucumbers grown in environmentally controlled conditions, the plant treatment with HrpNEa had a deterrent effect on colonization by the muskmelon aphid Aphis gossypii Glover (Gou et al. 2009). In Arabidopsis, HrpNEa-induced resistance was shown to repress multiplication of the green peach aphid Myzus persicae Sulzer (Dong et al. 2004), a generalist phloem-feeding insect (Pollard 1972).
Phloem-feeding insects are highly specialized in their mode of feeding and present a unique stress on plant fitness (Douglas 2006). They use their slender stylets to feed from a single-cell type, the phloem sieve element; the feeding process can be monitored by the electrical penetration graph (EPG) technique (Tjallingii 1987, 2006). Of pivotal importance, a stylet puncturing of the host plant cell, shown as a probe in the EPG, may lead to uptake of the phloem sap. In order to prevent protein clogging inside the sieve element, ejection of watery saliva is essential in feeding from the phloem. This ejection is detected in the EPG as E1 salivation and always precedes phloem sap ingestion. During ingestion from the sieve element, the watery E2 salivation occurs, and this E2 saliva is added to the ingested sap, and is thought to prevent phloem proteins from clogging inside the capillary food canal. Therefore, salivation is a crucial event during the phloem-feeding process for insects to overcome a number of phloem-related plant properties and reactions (Klingler et al. 2005; Pegadaraju et al. 2007; Mutti et al. 2008; Will and van Bel 2008; De Vos and Jander 2009; Louis et al. 2010).
In response to the phloem-feeding stress, plants defend themselves by using the phloem-based defense mechanism (Kehr 2006; Will and van Bel 2006, 2008). This mechanism involves the biosynthesis of β-1,3-glucan callose and subsequent closure of sieve pores and coagulation on sieve plates (Stone and Clarke 1992; Willats and Knox 2003; Kuśnierczyk et al. 2008; Saheed et al. 2009; Villada et al. 2009). Biochemical evidence and molecular studies in several plant species indicate that callose is synthesized by a class of enzymes, termed callose synthases or glucan synthases (Hong et al. 2001; Verma and Hong 2001). Twelve genes encoding putative glucan synthase have been identified and designated as AtGSL1 through AtGSL12 in Arabidopsis (for review, see Chen and Kim 2009). Based on analyses of mutations made by knocking out the individual GSL genes, it is found that the genes exhibit partially redundant roles in plant growth and development; a single GSL gene can also have diverse functions (Chen and Kim 2009). For example, GSL5 is responsible for the synthesis of wound-inducible and pathogen-inducible callose in leaf tissue (Jacobs et al. 2003; Nishimura et al. 2003; Wawrzynska et al. 2010); it also plays an important role in exine formation and pollen wall patterning (Dong et al. 2005b; Enns et al. 2005). However, genetic evidence is incomplete regarding functions of the different GSL genes; so far, evidence is available only for AtGSL1, 2, 5, 6, 8 and 10 (Kaliff et al. 2007; Dong et al. 2008; Huang et al. 2008; Töller et al. 2008; Chen et al. 2009a; Consonni et al. 2009; Xie et al. 2009).
One of the main purposes of this study was to elucidate if AtGSL and callose play a role in HrpNEa-induced resistance to M. persicae in Arabidopsis. We present evidence that AtGSL5 is required for HrpNEa-induced repression in aphid feeding activities. Recently we showed that AtMYB44 is the most HrpNEa-inducible of the 37 transcription factor genes tested and that AtMYB44 is required for the induction of resistance to M. persicae in Arabidopsis (Liu et al. 2010a). In this study, we sought to verify if AtMYB44 also affects the repression of aphid feeding activities.
2 Materials and methods
2.1 Plant growth and treatment
Arabidopsis genotypes used in this study included the wild type (WT) of the ecotype Col-0 and both atmyb44 and atgsl mutants previously generated under the background of Col-0 (table 1). Plants were grown in 9 cm pots (5 plants/pot) and incubated in an environmentally controlled chamber (22°C; 14 h illumination at 250 μE/m2/s) for 40 days before use. HrpNEa was prepared and purified as described (Dong et al. 1999). The protein was used in a 10 μg/ml aqueous solution and applied in the presence of surfactant Silwet-77 (0.02%, v/v) by spraying plant tops with a low-pressure atomizer. Similar treatment with pure water in the presence of Silwet-77 (0.02%, v/v) was used as a control.
2.2 Aphid culture
A single isolate of M. persicae was collected from the field-grown radish (Raphanus sativus L.) near Nanjing in China. A clone of apterous (wingless) agamic females was obtained by acclimatization in WT Arabidopsis grown in the chamber (22°C; 14 h illumination at 250 μE/m2/s). The colony was maintained in nursery WT Arabidopsis seedlings and was transferred to fresh plants every 2 weeks. Uniform 10-day-old apterous agamic females were used in this study and were transferred to experimental plants with a fine Chinese writing brush.
2.3 Plant colonization
Plants were treated with pure water and purified HrpNEa separately. Water-treated and HrpNEa-treated plants were distributed alternately in 45 × 30 cm plastic trays. Four days after treatment, plants were fed by root irrigation with a 2 μM aqueous solution of hexyl rhodamine B (HRB), a rose fluorescence dye with the chemical character of water solubility and plant-permissible diffusion (Boevink et al. 1996; Zheng et al. 2005). Twenty-four hours later, HRB-fed plants were colonized with uniform 10-day-old aphids by placing the aphids on lower sides of the top two expanded leaves of the plants (10 aphids per leaf). A total of 1200 aphids were monitored in four repetitions of the experiments for each single recombination of a treatment and a plant genotype. In each experimental repetition, 300 aphids were placed on 30 leaves of 15 plants treated specifically. The leaves and aphids were observed at intervals in 24 h using a fluorescence microscope of low magnification. Aphid feeding activities were judged on the basis of HRB fluorescence from their esophagus globes. The number of feeding aphids in a leaf colony was scored.
2.4 Aphid feeding behaviour
Activities of aphids placed on leaves of HRB-fed plants were observed with the aid of HRB fluorescence, either from wounds caused by the stylet puncturing of leaves or from esophagus globes of feeding aphids. HRB fluorescence spots on leaves were regarded as a result of spillage of the plant sap. The term ‘feeding aphids’ was used to indicate aphid individuals that had accomplished a feeding process and had ingested the plant sap, seen as strong signals of HRB fluorescence from esophagus globes of the insects. The effect of HrpNEa on feeding activities was quantified as a percentage decrease in the number of feeding aphids using the formula:
Here, x and y refer to proportions of feeding aphids in plants treated with water and HrpNEa, respectively.
Aphid feeding activities were also observed by the EPG technique using the Giga Amplifier system (Laboratory of Entomology, Wageningen Agricultural University, Wageningen, The Netherlands; http://www.epgsystems.eu/systems.htm). Uniform 10-day-old aphids were placed on the upper sides of the top first-expanded leaves of plants. For each genotype of the plant or each combination of a genotype and treatment (with EVP or HrpNEa), 20 aphids placed on 20 plants were monitored in five repetitions of the experiments. Immediately after the aphids were placed on the leaves, a 20-mm-diameter gold wire was attached to the dorsal surface of each aphid’s abdomen using the silver conductive paint. The other end of the wire was connected to a four-channel Giga-4 direct-current amplifier with 109 Ω input resistance in an electrical circuit that was also connected to the plant via an electrode placed in the soil. The behaviour of individual aphids was monitored for 4 h. Voltage waveforms were digitized at 100 Hz with an A/D converter USB device. Waveform patterns were identified according to previously described categories (Tjallingii and Esch 1993). Waveform recordings were dissected every 5 s with the EPG analysis software STYLET 2.5 installed in a computer connected to Giga-4 direct-current amplifier.
2.5 AtGSL5 expression and callose deposition
The gene expression in HrpNEa-treated plants and water-treated control plants was determined immediately and 3 days after treatment. Total RNA isolated from plant tops and the top two leaves was subjected to reverse transcriptase–polymerase chain reaction (RT-PCR) performed using superscript II RNAse Hˉ Reverse Transcriptase (Invitrogen) and using the EF1α gene as a reference (Peng et al. 2003). Specific primers were synthesized as per reported sequences of AtGSL5 and EF1α (Arabidopsis Genome Initiative Numbers AT4G03550 and AT1G07930). Primers specific to AtGSL5 were 5′-CCGCCACCGCACCGTCCCGCCGCAA-3′ and 5′-CGTCGTAGTTCCTCCACTTACAATG-3′, yielding a 752 bp product. Primers specific to EF1α were 5′-CCCCTTCGTCTCCCACTTCAGGATGTCTA-3′ and 5′-GTTGTCACCTGGAAGTGCCTCAAGAAG-3′, yielding a 189 bp product. RT-PCR products were confirmed by sequencing and sequence comparison. They were resolved by electrophoresis and visualized by staining with ethidium bromide.
Callose deposition in HrpNEa-treated plants and water-treated control plants was determined 3 days after treatment. Callose deposition in leaves was visualized as a violet colour by staining with aniline blue (Reuber et al. 1998; Zhang et al. 2009). The top two leaves were infiltrated with 5 ml of a solution made of phenol, glycerol, lactic acid, water and 95% ethanol (1:1:1:1:2, v/v). Leaves in solution were incubated in a 65°C bath until they were judged clear and then stained with aniline blue. The staining reaction was held in the dark for 4 h. Samples were observed by microscopy under ultraviolet field. Leaf photos were resolved by Adobe Photoshop to determine relative levels of callose in leaves. The size of a leaf and the size of callose-deposited areas were determined using the Magic Wand and Edit Tools of Adobe Photoshop as described (Kwack et al. 2005; Sun et al. 2010). The relative level of callose in a leaf was qualified as the ratio of violet callose-deposited area vs. leaf size.
2.6 Data treatment
The experiments were repeated three to five times with similar results. The Student’s t-test was used to compare data obtained from HrpNEa-treated plants with those obtained from water-treated control plants, and to compare data obtained from the WT and mutant plants. Quantitative data were also analysed using the ANOVA test to compare differences among plants of the WT and different mutants.
3 Results
3.1 Plant treatment with HrpN Ea reduces proportion of feeding aphids
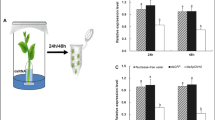
We devised a luminescence assay using HRB, a fluorescence dye with the chemical character of water solubility and plant-permissible diffusion (Boevink et al. 1996), to monitor M. persicae behaviours after colonization of Arabidopsis leaves. After the plants were fed by root irrigation with an aqueous HRB solution, leaves were stained rose, whereas feeding sites appeared as orange spots when uniform apterous females of the insect were placed on the leaves (10 aphids/leaf) of HRB-fed plants (figure 1A, section a). The stylet puncturing of leaves occurred in 1 h (figure 1A, section b); puncturing sites appearing as orange fluorescence spots was thought to indicate spillages of the plant sap from wounds caused by the stylet puncturing of leaves (figure 1A, sections b–c). Subsequently, the number of puncturing sites and the number of plant sap spillages increased with time during the course of 24 h monitoring experiments (figure 1A, sections b–c). At the 24th hour, strong signals of HRB fluorescence were found in esophagus globes of aphids on leaves of water-treated control plants but not HrpNEa-treated plants (figure 1B), suggesting that feeding activities were repressed by the plant treatment with HrpNEa. The luminescence assay method was used to investigate 24 h feeding activities of a large-scale aphid population of 1200 aphids placed on HrpNEa-treated plants and water-treated control plants, respectively. As shown in figure 1C, HrpNEa-treated plants had much smaller number of feeding aphids than water-treated control plants (Student’s t-test, P < 0.01) at each time point from 1 h after colonization. The largest degree of difference in the number of feeding aphids between HrpNEa-treated and control plants was found at the 4th hour. Similar degrees of difference were observed subsequently. The proportion of feeding aphids remained smaller in HrpNEa-treated plants than in control plants (figure 1C), suggesting that the HrpNEa treatment induced a deterrent effect on M. persicae feeding on the plant.
Hexyl rhodamine B (HRB) fluorescence assays to observe the green peach aphid Myzus persicae feeding on leaves of wild-type (WT) Arabidopsis plants. (A–C) Plants were treated with pure water (control) and purified HrpNEa separately. Four days later, treated plants were fed by root irrigation with an aqueous HRB solution. Twenty-four hours later, the top two expanded leaves of HRB-fed plants were colonized by uniform aphids with 10 adults placed on the lower side of the top two expanded leaves. Leaves and aphids were observed with a fluorescence microscope. (A) Leaves of HrpNEa-treated plants photographed immediately (section a) and at the indicated time points (sections b–c) after colonization. (B) Aphids on leaves photographed at the 24th hour after colonization. (C) A 24 h survey on proportions of feeding aphids monitored at a large-scale population (1200 aphids/treatment). The curves represent mean ± standard deviation (SD) of the results obtained from four repetitions of experiments (15 plants/repetition; 20 aphids/plant).
3.2 Tests of 37 atgsl alleles identify atgsl5 and atgsl6 as compromised in HrpN Ea -induced repression of aphid feeding activities
To determine if any of the 12 AtGSL genes already identified in Arabidopsis (Hong et al. 2001; Verma and Hong 2001; Chen and Kim 2009) plays a role in HrpNEa-induced deterrent effect on M. persicae feeding activities, we investigated Arabidopsis mutants generated previously by T-DNA insertion into the individual AtGSL genes (Alonso et al. 2003; Jacobs et al. 2003; Nishimura et al. 2003; Enns et al. 2005; Huang et al. 2008; Töller et al. 2008; http://www.arabidopsis.org). A total of 37 mutants were selected as representative of mutagenesis alleles of the individual AtGSL genes, including 3 and 4 mutation alleles of AtGSL5 and AtGSL6, respectively (table 1). The 37 mutants were compared with the WT plant in terms of response to HrpNEa and the effects on M. persicae feeding. Feeding activities were visualized by the HRB fluorescence assay applied to 1200 aphids in each of the plant genotypes treated with HrpNEa or water. As shown in figure 2, different genotypes of the plant varied greatly in regard to aphid feeding activities and the effect of HrpNEa. The proportion of feeding aphids was decreased by HrpNEa treatment compared with water treatment control (Student’s t-test, P < 0.01) in the WT plant, and also in 30 atgsl mutants rather than atgsl5 and atgsl6. In atgsl5 and atgsl6 mutants, especially atgsl5-1 and atgsl6-2, HrpNEa treatment did not reduce the proportion of feeding aphids to a significant extent. These observations suggested that atgsl5 and atgsl6 were both inhibitive to the deterrent effect of HrpNEa on aphid feeding. Based on multiple comparisons (ANOVA test, P < 0.01), the three atgsl5 alleles were found to be more inhibitive than the four atgsl6 alleles, and atgsl5-1 was the strongest inhibitor of the HrpNEa-induced deterrent effect on M. persicae feeding activities. So, both AtGSL5 and AtGSL6 genes were important for the effect of HrpNEa with AtGSL5 being more critical than AtGSL6.
Quantification of proportions of aphids feeding on wild-type (WT) Arabidopsis and the atgsl mutants that represent different mutation alleles of 12 AtGSL genes. Plants were treated with pure water (control) and purified HrpNEa separately. Four days later, treated plants were fed by root irrigation with an aqueous hexyl rhodamine B (HRB) solution. Twenty-four hours later, the top two expanded leaves of HRB-fed plants were colonized by uniform aphids with 10 adults placed on the lower side of the top two expanded leaves. Leaves and aphids were observed with a fluorescence microscope. The number of feeding aphids that had the HRB fluorescence in their esophagus globes was scored. Histograms represent mean ± SD of the results obtained from four repetitions of experiments (15 plants/repetition; 20 aphids/plant).
3.3 Deterrent effect of HrpN Ea on aphid feeding occurs mainly in phloem phase
To further elucidate the effects of HrpNEa and AtGSL genes on M. persicae feeding from Arabidopsis, feeding activities and the phloem-feeding behaviour in particular were studied by the EPG technique. The technique was used to first characterize the feeding behaviour and the effect of HrpNEa treatment. As shown in figure 3A, aphid feeding activities were detected in the EPG as different waveform patters recognized according to the standard previously established (Tjallingii and Esch 1993) and widely used (Tjallingii 1987, 2006; Klingler et al. 2005; Mutti et al. 2008). Based on a 4 h EPG record of aphid feeding activities, it was observed that all the 20 aphids tested in five repetitions of the experiments for HrpNEa-treated plants and water-treated control plants, respectively, accomplished major steps of the feeding process, but aphid activities varied greatly depending on the feeding process (figure 4; table 2) and varied greatly with different genotypes of the plant (figures 3 and 4).
Electrical penetration graph (EPG) analyses of aphid feeding on Arabidopsis leaves. (A) The EPG waveforms obtained from untreated wild-type (WT) plants [Probe, the stylet puncturing of leaf; np, no probe (nonpuncturing); Path, pathway; Xylem, the xylem phase; Phloem, the phloem phase]. (B) The 2nd hour EPG patterns of aphid feeding from the WT plant and AtGSL5-deficient mutant atgsl5-1. Plants were treated with pure water (control) and purified HrpNEa separately. Five days later, four uniform aphids were placed on the top first-expanded leaves of four plants; feeding behaviours were monitored by a four-channel Giga Amplifier system (Laboratory of Entomology, Wageningen Agricultural University, Wageningen, The Netherlands). Each EPG picture represents 20 aphids tested in five repetitions of the experiments.
Key parameters of the EPG analyses of aphid feeding from the wild-type (WT) plant and representative atgsl mutants. (A–C) Plants were treated with pure water (control) and purified HrpNEa separately. Five days later, four uniform aphids were placed on the top first-expanded leaves of four plants; feeding behaviours were monitored by the Giga Amplifier system (Laboratory of Entomology, Wageningen Agricultural University, Wageningen, The Netherlands). The total durations of the three phases were scored from 4 h EPG records. (D) Relative effect of HrpNEa on the phloem-feeding activity. The effect in each genotype of the plant was quantified as a percentage reduction in total duration of the phloem phase under the condition of HrpNEa treatment in contrast to water treatment control. Each histogram in (A–D) represents 20 aphids monitored in five repetitions of the EPG study.
In the WT plant, the proportion of time that aphids spent outside the cuticle (nonpenetration; figure 3A, np) until the first stylet puncturing of leaf cells (figure 3A, probe) under the condition of HrpNEa treatment was similar to that under the condition of water treatment control (table 2). Under both conditions, equivalent proportion of time was used to penetrate between cells en route to the vascular tissue, referred to as the pathway phase in the EPG (figure 3A, path; table 2). The pathway phase represents insect’s efforts in navigating the phloem and preparing to ingest sap from sieve elements (Klingler et al. 2005; Tjallingii 2006). Subsequently, aphid stylets may proceed to the phloem phase (figure 3A, phloem) in which ingestion of the phloem sap may occur (Tjallingii 2006). The pathway phase may be also connected with the xylem phase (figure 3A, Xylem), indicating stylet penetration of the xylem in the vascular tissue (Tjallingii 2006). The EPG records indicated that aphid activities did not change evidently in HrpNEa-treated plants compared with control plants until the stylet entry of vascular tissues (table 2). Proportions of time in the pathway phase and in the first phloem phase were much longer in HrpNEa-treated plants than in control plants, suggesting a HrpNEa-induced impediment to aphids in locating the ingestion site within the vascular tissue. Moreover, on HrpNEa-treated plants, fewer penetrations of the phloem were observed after the first stylet entry of phloem phase (table 2). These results suggest that phloem properties of HrpNEa-treated plants were unfavourable for feeding. Consistently, the total duration of the phloem phase was much shorter in HrpNEa-treated plants than in control plants (Student’s t-test, P < 0.01) during the course of the 4 h EPG monitoring (table 2). In particular, durations of the phloem phase in the 2nd hour of EPG monitoring were 32 and 15 min on average in control plants and HrpNEa-treated plants (Student’s t-test, P < 0.01), respectively, further supporting the deterrent effect of HrpNEa treatment on the phloem-feeding activity of M. persicae (figure 4B).
3.4 atgsl5-1 impairs deterrent effect of HrpN Ea on phloem feeding of aphids
The effects of AtGSL genes on the phloem-feeding behaviour of M. persicae were elucidated by investigating 12 atgsl mutants using the EPG technique. The 12 mutants were selected because they represented highly effective alleles of atgsl1 through atgsl12 in suppressing the HrpNEa-induced deterrent effect on aphid feeding activities (figure 2). In the EPG experiments, the total duration of the nonpuncturing phase in all the plants was close under conditions of water treatment control and HrpNEa treatment, suggesting that the atgsl alleles had little effects on the stylet puncturing of leaf cells (figure 4A). However, feeding behaviours subsequent to the stylet entry of leaf tissues changed greatly due to the HrpNEa treatment in WT and 11 atgsl mutants but not in atgsl5-1. In WT and the 11 mutants, the total duration of the pathway phase was longer in HrpNEa treatment than in control (Student’s t-test, P < 0.01), suggesting that aphids incurred frustrations in navigating the phloem owing to the effect of HrpNEa (figure 4B). In atgsl5-1, however, the total duration of the pathway phase in HrpNEa treatment was as long as in the control (figure 4B). Moreover, in atgsl5-1, the total duration of the phloem phase were also close, suggesting that aphids fed from the phloem equally well, under HrpNEa treatment and control conditions (figures 3B and 4C). On the contrary, the other 11 mutants performed as did WT in supporting the deterrent effect of HrpNEa on the phloem-feeding activity. In these plants, total durations of the phloem phase were decreased significantly by the HrpNEa treatment compared with the control (figure 5C; Student’s t-test, P < 0.01).
Determinations of AtGSL5 expression and callose deposition in WT Arabidopsis and the atgsl5-1 and atmyb44 mutants that have defects in AtGSL5 and AtMYB44 genes. (A) Reverse transcriptase–polymerase chain reaction analysis of AtGSL5 expression in plants treated with pure water (control) and purified HrpNEa, respectively [dpt, day(s) post-treatment; EF1α, a reference gene]. (B) Leaf callose deposition visualized as a violet colour by staining the leaves with aniline blue at 3 hpt (hours post-treatment) (scale bar, 1 mm). (C) Relative content of callose in leaves. Relative levels of callose were given as the violet callose-deposited area/leaf size ratio. Data are presented as mean ± SD of the results obtained from three repetitions of experiments (15 plants/repetition).
The relative degree of the HrpNEa-induced deterrent effect on the phloem-feeding activity in each genotype of the plant was quantified as the rate of decrease in total duration of the phloem phase under the HrpNEa treatment condition in contrast to the control (figure 4D). Based on this criterion, only 3 of the 12 mutants were found to be inhibitive to the effect of HrpNEa. The three mutants were atgsl5-1, atgsl8-2, and atgsl9-3 – all inhibited the effect of HrpNEa to significant extents (ANOVA test, P < 0.01). Comparatively, atgsl5-1 proved to be most inhibitive to the HrpNEa-induced deterrent effect on phloem feeding by M. persicae, confirming the importance of AtGSL5.
3.5 atgsl5-1 does not accumulate AtGSL5 transcript and callose
The role of ATGSL5 in repression of aphid feeding activities was correlated with induced expression of the gene in Arabidopsis. Three days after treatment with HrpNEa, the expression of AtGSL5 was induced conspicuously in WT but not in atgsl5-1 (figure 5A). Moreover, in response to HrpNEa, an abundant amount of callose was induced in WT rather than atgsl5-1 (figure 5B). In WT, callose had a greater level when induced by the HrpNEa treatment compared with control; in atgsl5-1, however, the relative level of callose was lower and did not change evidently in response to HrpNEa (figure 5C).
3.6 AtGSL5 expression and callose deposition are arrested in atmyb44
Recently we showed that AtMYB44 was the most HrpNEa-responsive of the 37 transcription factor genes tested and that AtMYB44 was required for the induction of resistance to M. persicae in Arabidopsis (Liu et al. 2010a). In this study, we found that AtMYB44 was required for the induction of AtGSL5 expression and callose deposition. Both events were induced in WT but not atmyb44, an Arabidopsis mutant generated previously through T-DNA insertion at the promoter region of AtMYB44 (Liu et al. 2010b; http://www.arabidopsis.org). The expression of AtGSL5 was induced in WT but not in atmyb44 following treatment with HrpNEa (figure 5A). In contrast to the absence of AtGSL5 expression in atmyb44 and in water-treated control plants of WT, the gene was highly expressed in HrpNEa-treated WT plants tested 3 days after treatment. Moreover, in response to HrpNEa, an abundant amount of callose was induced in WT rather than atmyb44 (figure 5B). In WT, callose had a greater level as induced by the HrpNEa treatment compared with control; in atmyb44, however, the level of callose was lower and did not change evidently in response to HrpNEa (figure 5C).
3.7 atmyb44 is more susceptible than atgsl5-1 to phloem feeding by aphids
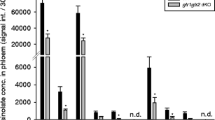
Comparison of data shown in figure 5 revealed that the amount of callose was smaller in atmyb44 than in atgsl5-1. This difference was found to be consistent with the extents by which atmyb44 and atgsl5-1 compromised the HrpNEa-induced deterrent effect on aphid feeding activities. In parallel experiments, WT, atmyb44 and atgsl5-1 treated with water or HrpNEa were colonized each with 1200 and 20 aphids, respectively, for monitoring by the HRB fluorescence and EPG assays. In comparison with WT, both atmyb44 and atgsl5-1 allowed greater proportions of feeding aphids (figure 6A) and longer durations of the phloem phase (figure 6B). However, atmyb44 differed from atgsl5-1 in both assays. More aphids failed to feed from atgsl5-1 than from atmyb44 in cases of both control and HrpNEa treatment (figure 6A). In atgsl5-1, a smaller proportion of feeding aphids was scored in the case of HrpNEa treatment compared with control (Student’s t-test, P < 0.01). In atmyb44, however, the proportion of feeding aphids in the case of HrpNEa was similar to that in control. Based on 4 h EPG records, the total duration of the phloem phase was longer in atmyb44 than in atgsl5-1 and was reduced by HrpNEa treatment to a greater extent in atgsl5-1 than in atmyb44 (figure 6B). These analyses suggested that atmyb44 was more susceptible than atgsl5-1 while both mutants were more susceptible than WT to phloem feeding by aphids. Moreover, atmyb44 was more inhibitive than atgsl5-1 to the deterrent effect of HrpNEa on aphid feeding activities, especially the phloem-feeding behaviour (figure 6C).
Comparison of WT, atgsl5-1 and atmyb44 in terms of effects on aphid feeding activities. (A) Proportions of feeding aphids. Plants were treated with pure water (control) and purified HrpNEa separately. Four days later, treated plants were fed by root irrigation with an aqueous hexyl rhodamine B (HRB) solution. Twenty-four hours later, the top two expanded leaves of HRB-fed plants were colonized by uniform aphids with 10 adults placed on the lower side of the top two expanded leaves. Leaves and aphids were observed with a fluorescence microscope. Proportions of feeding aphids were scored (mean ± SD) with a large-scale population – 1200 aphids per treatment-plant genotype combination, in four repetitions of the experiments. (B) Plants were treated with pure water (control) and purified HrpNEa separately. Five days later, four uniform aphids were placed on the top first-expanded leaves of four plants; feeding behaviours were monitored by the Giga Amplifier system (Laboratory of Entomology, Wageningen Agricultural University, Wageningen, The Netherlands). The total duration of the phloem phase was scored from 4 h EPG records. (C) The effect of HrpNEa on aphid feeding activities. Relative levels of the effect was quantified as a percentage reduction in the number of feeding aphids, or quantified as a percentage reduction in total duration of the phloem phase, under the condition of HrpNEa treatment in contrast to water treatment control. Each histogram in (B) and (C) represents 20 aphids.
4 Discussion
One purpose of this study was to screen and identify GLS genes involved in the induction of resistance to M. persicae in Arabidopsis following treatment with HrpNEa, a bacterial harpin protein (Wei et al. 1992). Previously, the induced resistance was shown to inhibit colonization of Arabidopsis by M. persicae (Liu et al. 2010a) and to inhibit multiplication of the insect on the plant (Dong et al. 2004). HRB fluorescence assays suggest that the HrpNEa treatment causes a decrease in feeding population of aphids that colonizes leaves of treated Arabidopsis (figure 1). Use of the technique to investigate previously generated 37 atgsl alleles has identified atgstl5 and atgsl6 as mostly compromised in HrpNEa-induced repression of aphid feeding activities (figure 2). The 37 mutants represent different mutagenesis alleles of the 12 AtGSL genes (Jacobs et al. 2003; Nishimura et al. 2003; Enns et al. 2005; Huang et al. 2008; Töller et al. 2008; http://www.arabidopsis.org) previously identified in Arabidopsis (Hong et al. 2001; Verma and Hong 2001; Chen and Kim 2009). The different mutation alleles of individual AtGSL genes perform consistently in response to HrpNEa and in the effect on aphid feeding activities (figure 2), suggesting that the response and effect are a stable attribute of the genes under the conditions in this study. The atgsl alleles mostly inhibitive to the effect of HrpNEa include atgsl5-1 and atgsl6-2. Subsequent studies by the EPG technique suggest that the deterrent effect of HrpNEa on aphid feeding activities occurs mainly in the phloem phase (figures 3 and 4; table 2). It is atgsl5-1, rather than atgsl6-2, that markedly compromises, but does not eliminate, the deterrent effect (figure 4), suggesting that AtGSL5 is required for a significant proportion of the effect.
Apart from atgsl5-1, the other 11 atgsl alleles seem somewhat inhibitive to the effect of HrpNEa (figure 4), indicating that the AtGSL genes other than AtGSL5 play some roles in repressing the phloem-feeding activity. This result indicates possible redundancy in functions of GSL genes. Different GSL genes exhibit partially redundant roles in plant growth, development and defences (Chen and Kim 2009). It has been shown that the GSL5 enzyme is responsible for the synthesis of wound- and pathogen-inducible callose in leaf tissue (Jacobs et al. 2003; Nishimura et al. 2003; Wawrzynska et al. 2010). GSL5 also plays an important role in exine formation and pollen wall patterning (Dong et al. 2005b; Enns et al. 2005). When the AtGSL5 gene plays a dominant role in the HrpNEa-induced deterrent effect on the phloem-feeding activity of aphids, other AtGSL genes may be partially redundant in function with AtGSL5 and play additive roles in the effect. The representative atgsl1 though atgsl12 alleles were investigated (figure 2; table 1) because genetic evidence regarding functions of different GSL genes was available for several but not all of the genes (Kaliff et al. 2007; Dong et al. 2008; Huang et al. 2008; Töller et al. 2008; Chen et al. 2009b; Consonni et al. 2009; Xie et al. 2009). However, the present study does not provide information about contributions of the different individual AtGSL genes to the repression of aphid feeding activities except evidence of the major role of AtGSL5 in the HrpNEa-induced deterrent effect on the phloem-feeding behaviour.
A related issue is that evidence is lacking in terms of regulation of GSLs’ defensive function. In Arabidopsis, HrpNEa-induced resistance to M. persicae develops via the ethylene signalling pathway (Dong et al. 2004). The pathway uses different regulators to fulfill distinct functions (Guo and Ecker 2004; Chen et al. 2009a; Camehl et al. 2010). EIN2 is an integral membrane protein and an essential regulator of ethylene signalling, and therefore is indispensable for multiple processes, including defense responses (Alonso et al. 1999; Wang et al. 2002; Dong et al. 2004; Camehl et al. 2010). In HrpNEa-treated plants, induced EIN2 expression is critical for the induction of resistance to M. persicae (Dong et al. 2004). Induced EIN2 expression is regulated by AtMYB44 (Liu et al. 2010b), a multifunctional transcription factor (Kirik et al. 1998; Kranz et al. 1998; Jung et al. 2008, 2010; Pitzschke et al. 2009). Downstream events were unclear.
Our evidence suggests that AtMYB44 is required for HrpNEa-induced expression of AtGLS5 and the concomitant deposition of callose in Arabidopsis (figure 5). AtMYB44 is also required for the HrpNEa-induced deterrent effect on the phloem-feeding activity of M. persicae (figure 6). On comparison of the plant mutants atmyb44 and atgsl5-1, atmyb44 is found to be more susceptible to aphids and more inhibitive to the HrpNEa-induced deterrent effect on the phloem-feeding activity. These results indicate other components, besides AtGSL5 and callose, are involved in HrpNEa-induced resistance to M. persicae in the plant (Zhang et al. 2011), and especially support the function of AtMYB44 as a regulator of the induced resistance (Dong et al. 2004; Liu et al. 2010a,b). It would be great of interest to study in the future the physiological connection between AtMYB44 and AtGSL5 during induction of the defense.
Abbreviations
- EPG:
-
electrical penetration graph
- GSL :
-
GLUCAN SYNTHESIS-LIKE
- HRB:
-
hexyl rhodamine B
- RT-PCR:
-
reverse transcriptase–polymerase chain reaction
- WT:
-
wild type
References
Alonso JM, Hirayama T, Roman G, Nourizadeh S and Ecker JR 1999 EIN2, a bifunctional transducer of ethylene and stress responses in Arabidopsis. Science 284 2148–2152
Alonso JM, Stepanova AN, Leisse TJ, Kim CJ, Chen H, Shinn P, Stevenson DK, Zimmerman J, et al. 2003 Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301 653–657
Boevink P, Cruz S, Hawes C, Harris N and Oparka KJ 1996 Virus-mediated delivery of the green fluorescent protein to the endoplasmic reticulum of plant cells. Plant J. 10 935–941
Camehl I, Sherameti I, Venus Y, Bethke G, Varma A, Lee J and Oelmüller R 2010 Ethylene signalling and ethylene-targeted transcription factors are required to balance beneficial and nonbeneficial traits in the symbiosis between the endophytic fungus Piriformospora indica and Arabidopsis thaliana. New Phytol. 185 1062–1073
Chen H, Xue L, Chintamanani S, Germain H, Lin H, Cui H, Cai R, Zuo J, et al. 2009a ETHYLENE INSENSITIVE3 and ETHYLENE INSENSITIVE3-LIKE1 repress SALICYLIC ACID INDUCTION DEFICIENT2 expression to negatively regulate plant innate immunity in Arabidopsis. Plant Cell 21 2527–2540
Chen L, Qian J, Qu SP, Long JY, Yin Q, Zhang CL, Wu XJ, Sun F, et al. 2008 Identification of specific fragments of HpaGXooc, a harpin from Xanthomonas oryzae pv. oryzicola, that induce disease resistance and enhance growth in plants. Phytopathology 98 781–791
Chen XY, Liu L, Lee EK, Han X, Rim Y, Chu H, Kim SW, Sack F and Kim JY 2009b The Arabidopsis callose synthase gene, GSL8, is required for cytokinesis and cell patterning. Plant Physiol. 150 105–113
Chen XY and Kim JY 2009 Callose synthesis in higher plants. Plant Signal. Behav. 4 489–492
Consonni C, Bednarek P, Humphry M, Francocci F, Ferrari S, Harzen A, van Themaat EV and Panstruga R 2009 Tryptophan-derived metabolites are required for antifungal defence in the Arabidopsis thaliana mlo2 mutant. Plant Physiol. 152 1544–1561
De Vos M and Jander G 2009 Myzus persicae (green peach aphid) salivary components induce defence responses in Arabidopsis thaliana. Plant Cell Environ. 32 1548–1560
Dong HP, Peng JL, Bao ZL, Meng XD, Bonasera JM, Chen GY, Beer SV and Dong HS 2004 Downstream divergence of the ethylene signaling pathway for harpin-stimulated Arabidopsis growth and insect defense. Plant Physiol. 136 3628–3638
Dong HP, Yu HQ, Bao ZL, Guo XJ, Peng JL, Yao Z, Chen GY and Dong HS 2005a The ABI2-dependent abscisic acid signalling controls HrpN-induced drought tolerance in Arabidopsis. Planta 221 313–327
Dong HS, Delaney TP, Bauer DW and Beer SV 1999 Harpin induces disease resistance in Arabidopsis through the systemic acquired resistance pathway mediated by salicylic acid and the NIM1 gene. Plant J. 20 207–215
Dong XY, Hong ZL, Chatterjee J, Kim S and Verma DP 2008 Expression of callose synthase genes and its connection with Npr1 signaling pathway during pathogen infection. Planta 229 87–98
Dong X, Hong Z, Sivaramakrishnan M, Mahfouz M and Verma DP 2005b Callose synthase (CalS5) is required for exine formation during microgametogenesis and for pollen viability in Arabidopsis. Plant J. 42 315–328
Douglas AE 2006 Phloem-sap feeding by animals: problems and solutions. J. Exp. Bot. 57 747–754
Enns LC, Kanaoka MM, Torii KU, Comai L, Okada K and Cleland RE 2005 Two callose synthases, GSL1 and GSL5, play an essential and redundant role in plant and pollen development and in fertility. Plant Mol. Biol. 58 333–349
Gou ZH, Zhang SP and Dong HS 2009 Effects of HrpNEa on inducing Aphid repellency on Cucumis melo. Acta Agric. Boreali-Sinica 24 188–192
Guo H and Ecker JR 2004 The ethylene signaling pathway: new insights. Curr. Opin. Plant Biol. 7 40–49
Hong Z, Delauney AJ and Verma DP 2001 A cell plate–specific callose synthase and its interaction with phragmoplastin. Plant Cell 13 755–768
Huang LJ, Chen XY, Rim Y, Han X, Cho WK, Kim SW and Kim JY 2008 Arabidopsis glucan synthase-like 10 functions in male gametogenesis. J. Plant Physiol. 166 344–352
Jacobs AK, Lipka V, Burton RA, Panstruga R, Strizhov N, Schulze-Lefert P and Fincher GB 2003 An Arabidopsis callose synthase, GSL5, is required for wound and papillary callose formation. Plant Cell 15 2503–2513
Jung C, Seo JS, Han SW, Koo YJ, Kim CH, Song SI, Nahm BH, Choi YD and Cheong JJ 2008 Overexpression of AtMYB44 enhances stomatal closure to confer abiotic stress tolerance in transgenic Arabidopsis. Plant Physiol. 146 623–635
Jung C, Shim JS, Seo JS, Lee HY, Kim CH, Choi YD and Cheong JJ 2010 Non-specific phytohormonal induction of AtMYB44 and suppression of jasmonate-responsive gene activation in Arabidopsis thaliana. Mol. Cell 29 71–76
Kaliff M, Staal J, Myrenås M and Dixelius C 2007 ABA is required for Leptosphaeria maculans resistance via ABI1- and ABI4-dependent signalling. Mol. Plant-Microbe Interact. 20 335–345
Kehr J 2006 Phloem sap proteins: their identities and potential roles in the interaction between plants and phloem-feeding insects. J. Exp. Bot. 57 767–774
Kim JF and Beer SV 2000 hrp genes and harpins of Erwinia amylovora: a decade of discovery; in Fire blight and its causative agent, Erwinia amylovora (ed.) JL Vanneste (Wallingford: CAB International) pp 141–162
Kirik V, Kolle K, Misera S and Baumlein H 1998 Two novel MYB homologues with changed expression in late embryogenesis-defective Arabidopsis mutants. Plant Mol. Biol. 37 819–827
Klingler J, Creasy R, Gao L, Nair RM, Calix AS, Jacob HS, Edwards OR and Singh KB 2005 Aphid resistance in Medicago trunculata involves antixenosis and phloem-specific, inducible antibiosis, and maps to a single locus flanked by NBS-LRR resistance gene analogs. Plant Physiol. 137 1445–1455
Kranz HD, Denekamp M, Greco R, Jin H, Leyva A, Meissner RC, Petroni K, Urzainqui A, Bevan M and Martin C 1998 Towards functional characterisation of the members of the R2R3-MYB gene family from Arabidopsis thaliana. Plant J. 16 263–276
Kuśnierczyk A, Winge P, Jørstad TS, Troczyńska J, Rossiter JT and Bones AM 2008 Towards global understanding of plant defence against aphids – timing and dynamics of early Arabidopsis defence responses to cabbage aphid (Brevicoryne brassicae) attack. Plant Cell Environ. 31 1097–1115
Kwack MS, Eui Nam Kim EN, Lee H, Kim J-W, Chun S-C and Kim KD 2005 Digital image analysis to measure lesion area of cucumber anthracnose by Colletotrichum orbiculare. J. Gen. Plant Pathol. 71 418–421
Liu FQ, Liu HX, Jia Q, Wu XJ, Guo XJ, Zhang SJ, Song F and Dong HS 2006 The internal glycine-rich motif and cysteine suppress several effects of the HpaGXooc protein in plants. Phytopathology 96 1052–1059
Liu RX, LÜ BB, Wang XM, Zhang CL, Zhang SP, Qian J, Chen L, Shi HJ and Dong HS 2010a Thirty-seven transcription factor genes differentially respond to a harpin protein and affect resistance to the green peach aphid in Arabidopsis. J. Biosci. 35 435–450
Liu R, Chen L, Jia Z, Lü B, Shi H, Shao W and Dong H 2010b Transcription factor AtMYB44 regulates induced expression of the ETHYLENE INSENSITIVE2 gene in Arabidopsis responding to a harpin protein. Mol. Plant-Microbe Interact. 24 377–389
Louis J, Leung Q, Pegadaraju V, Reese J and Shah J 2010 PAD4-dependent antibiosis contributes to the ssi2-conferred hyper-resistance to the green peach aphid. Mol. Plant-Microbe Interact. 23 618–627
Mutti NS, Louis J, Pappan LK, Pappan K, Begum K, Chen M, Park Y, Dittmer N, Marshall J, Reese JC and Reeck GR 2008 A protein from the salivary glands of the pea aphid, Acyrthosiphon pisum, is essential in feeding on a host plant. Proc. Natl. Acad Sci. USA 105 9965–9969
Nishimura MT, Stein M, Hou B, Vogel JP, Edwards H and Somerville SC 2003 Loss of a callose synthase results in salicylic acid-dependent disease resistance. Science 301 969–972
Pegadaraju V, Louis J, Singh V, Reese JC, Bautor J, Feys BJ, Cook G, Parker JE and Shah J 2007 Phloem-based resistance to green peach aphid is controlled by Arabidopsis PHYTOALEXIN DEFICIENT4 without its signaling partner ENHANCED DISEASE SUSCEPTIBILITY1. Plant J. 52 332–341
Peng JL, Bao ZL, Dong HS, Ren HY and Wang JS 2004 Expression of harpinXoo in transgenic tobacco induces pathogen defense in the absence of hypersensitive cell death. Phytopathology 94 1048–1055
Peng JL, Dong HS, Dong HP, Delaney TP, Bonasera BM and Beer SV 2003 Harpin-elicited hypersensitive cell death and pathogen resistance requires the NDR1 and EDS1 genes. Physiol. Mol. Plant Pathol. 62 317–326
Pitzschke A, Djamei A, Teige M and Hirt H 2009 VIP1 response elements mediate mitogen-activated protein kinase 3-induced stress gene expression. Proc. Natl. Acad. Sci. USA 106 18414–18419
Pollard DG 1972 Plant penetration by feeding aphids (Hemiptera, Aphidoidea): A review. Bull. Entomol. Res. 62 631–714
Ren XY, Liu F, Bao ZL, Zhang CL, Wu XJ, Chen L, Liu RX and Dong HS 2008 Root growth of Arabidopsis thaliana is regulated by ethylene and abscisic acid signaling interaction in response to HrpNEa, a bacterial protein of harpin group. Plant Mol. Biol. Rep. 26 225–240
Reuber TL, Plotnikova JM, Dewdney J, Rogers EE, Wood W and Ausubel FM 1998 Correlation of defense gene induction defects with powdery mildew susceptibility in Arabidopsis enhanced disease susceptibility mutants. Plant J. 16 473–485
Saheed SA, Cierlik I, Larsson KA, Delp G, Bradley G, Jonsson LM and Botha CE 2009 Stronger induction of callose deposition in barley by Russian wheat aphid than bird cherry-oat aphid is not associated with differences in callose synthase or β-1,3-glucanase transcript abundance. Physiol. Plant 135 150–161
Stone BA and Clarke AE 1992 Chemistry and physiology of higher plant 1,3-β-glucans (callose); in Chemistry and biology of 1,3-β-glucans (eds) BA Stone and AE Clarke (Bundoora Australia: La Trobe University Press) pp 365–429
Sun LJ, Ren HY, Liu RX, Li BY, Wu TQ, Sun F, Liu HM, Wang XM and Dong HS 2010 An h-type thioredoxin functions in tobacco defense responses to two species of viruses and an abiotic oxidative stress. Mol. Plant-Microbe Interact. 23 1470–1485
Tjallingii WF 1987 Electrical recording of stylet penetration activities; in Aphids: Their Biology, Natural Enemies and Control (eds) AK Minks and P Harrewijn (Elsevier Amsterdam) pp 95–108
Tjallingii WF 2006 Salivary secretions by aphids interacting with proteins of phloem wound responses. J. Exp. Bot. 57 739–745
Tjallingii WF and Esch TH 1993 Fine-structure of aphid stylet routes in plant tissues in correlation with EPG signals. Physiol. Entomol. 18 317–328
Töller A, Brownfield L, Neu C, Twell D and Schulze-Lefert P 2008 Dual function of Arabidopsis Glucan Synthase-Like genes GSL8 and GSL10 in male gametophyte development and plant growth. Plant J. 54 911–923
van der Zwet T and Beer SV 1999 Fire Blight–Its Nature, Prevention and Control: A Practical Guide to Integrated Disease Management. Agriculture Information Bulletin (U.S. Department of Agriculture) No. 631
Verma DPS and Hong Z 2001 Plant callose synthase complexes. Plant Mol. Biol. 47 693–701
Villada ES, González EG, López-Sesé AI, Castiel AF and Gómez-Guillamón ML 2009 Hypersensitive response to Aphis gossypii Glover in melon genotypes carrying the Vat gene. J. Exp. Bot. 60 3269–3277
Wang KL, Li H and Ecker JR 2002 Ethylene biosynthesis and signaling networks. Plant Cell 14 S131–151
Wawrzynska A, Rodibaugh NL and Innes RW 2010 Synergistic activation of defense responses in Arabidopsis by simultaneous loss of the GSL5 callose synthase and the EDR1 protein kinase. Mol. Plant-Microbe Interact. 23 578–584
Wei ZM, Laby RJ, Zumoff CH, Bauer DW, He SY, Collmer A and Beer SV 1992 Harpin, elicitor of the hypersensitive response produced by the plant pathogen Erwinia amylovora. Science 257 85–88
Will T and van Bel AJE 2006 Physical and chemical interactions between aphids and plants. J. Exp. Bot. 57 729–737
Will T and van Bel AJ 2008 Induction as well as suppression: How aphid saliva may exert opposite effects on plant defense. Plant Signal. Behav. 3 427–430
Willats WGT and Knox JP 2003 Molecules in context: probes for cell wall analysis; in The Plant Cell Wall (ed.) JKC Rose (Oxford UK: CRC) pp 92–110
Xie B, Wang X and Hong Z 2009 Precocious pollen germination in Arabidopsis plants with altered callose deposition during microsporogenesis. Planta 231 809–823
Zhang CL, Bao Z, Liang Y, Yang X, Wu XJ, Hong XY and Dong HS 2007 Abscisic acid mediates Arabidopsis drought tolerance induced by HrpNEa in the absence of ethylene signalling. Plant Mol. Biol. Rep. 25 98–114
Zhang CL, Shi H, Chen L, Wang X, Lü B, Zhang S, Liang Y, Liu R, Qian J, Sun W, You Z and Dong HS 2011 Harpin-induced expression and transgenic overexpression of the phloem protein gene AtPP2-A1 in Arabidopsis repress phloem feeding of the green peach aphid Myzus persicae. BMC Plant Biol. 11 doi:10.1186/1471-2229-11-11
Zhang SJ, Yang X, Sun MW, Sun F, Deng S and Dong HS 2009 Riboflavin-induced priming for pathogen defense in Arabidopsis thaliana. J. Integr. Plant Biol. 51 167–174
Zheng H, Rowland O and Kunst L 2005 Disruptions of the Arabidopsis enoyl-CoA reductase gene reveal an essential role for very-long-chain fatty acid synthesis in cell expansion during plant morphogenesis. Plant Cell 17 1467–1481
Zitter TA and Beer SV 1998 Harpin for insect control. Phytopathology 88 S104–S105
Acknowledgements
We thank the graduate students studying in our lab for their assistance in aphid investigations. This study was supported by grants (2009ZX08002-004B and 2008ZX08002-001) from the National Novel Transgenic Organisms Breeding Project in China.
Author information
Authors and Affiliations
Corresponding author
Additional information
[Lü B Sun W, Zhang S, Zhang C, Qian J, Wang X, Gao R and Dong H 2010 HrpNEa-induced deterrent effect on phloem feeding of the green peach aphid Myzus persicae requires AtGSL5 and AtMYB44 genes in Arabidopsis thaliana. J. Biosci. 36 0000–0000] DOI
Beibei Lü Weiwei Sun and Shuping Zhang contributed equally to this work.
Rights and permissions
About this article
Cite this article
Lü, B., Sun, W., Zhang, S. et al. HrpNEa-induced deterrent effect on phloem feeding of the green peach aphid Myzus persicae requires AtGSL5 and AtMYB44 genes in Arabidopsis thaliana . J Biosci 36, 123–137 (2011). https://doi.org/10.1007/s12038-011-9016-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12038-011-9016-2