Abstract
Attention deficit/hyperactivity disorder (ADHD) is one of the most highly heritable psychiatric disorders in childhood. The risk gene mutation accounts for about 60 to 90 % cases. Synaptosomal-associated protein of 25 kDa (SNAP-25) is a presynaptic plasma membrane protein which is expressed highly and specifically in the neuronal cells. A number of evidences have suggested the role of SNAP-25 in the etiology of ADHD. Notably, the animal model of coloboma mouse mutant bears a ∼2-cM deletion encompassing genes including SNAP25 and displays spontaneous hyperkinetic behavior. Previous investigators have reported association between SNPs in SNAP25 and ADHD, and controversial results were observed. In this study, we analyzed the possible association between six polymorphisms (rs3746544, rs363006, rs1051312, rs8636, rs362549, and rs362998) of SNAP25 and ADHD in a pooled sample of ten family-based studies and four case–control studies by using meta-analysis. The combined analysis results were significant only for rs3746544 (P = 0.010) with mild association (odds ratio (OR) = 1.14). And, the meta-analysis data for rs8636, rs362549, and rs362998 are the first time to be reported; however, no positive association was detected. In conclusion, we report some evidence supporting the association of SNAP25 to ADHD. Future research should emphasize genome-wide association studies in more specific subgroups and larger independent samples.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Attention deficit/hyperactivity disorder (ADHD [MIM143465]) is a common psychiatric disorder [1, 2] in childhood worldwide that affects 6–7 % of school age children when diagnosed via the Diagnostic and Statistical Manual IV (DSM-IV) criteria [3]. Boys are diagnosed approximately threefold greater incidence than girls [4, 5]. It is typically characterized by significant problems of attention, hyperactivity, or acting impulsively that are not appropriate for a person’s age [6]. Three different subtypes are recognized according to DSM-IV as follows: primarily inattentive (ADHD-I), primarily hyperactive/impulsive (ADHD-HI), and combined (ADHD-C), among them, the inattentive and combined subtypes are the most prevalent [7]. ADHD children are at high risk of negative long-term outcomes including academic underachievement and high accident rates and difficulty sustaining stable social relationships [8]. And, if they were left untreated, the effects tend to persist in adulthood [9, 10] with approximately 2.5 % of adults meeting diagnostic criteria for ADHD [11]. ADHD adults are at high risk of car accidents, divorce, substance misuse, and frequent job changes [12–14]. It is believed that both genetic factors and environmental factors and its interactions can contribute to this disorder [15]. Family, twin and adoption studies have indicated a strong genetic component in the susceptibility to ADHD, with an average estimated heritability (the proportion of phenotypic variance explained by genetic factors) of 0.76 [8].
Pathophysiology of ADHD
Animal and human studies have implicated the dysregulation of prefrontal cortical areas [8], basal ganglia, cerebellum, and temporal and parietal cortex [8, 16] in the pathophysiology of ADHD. And, this condition is complex and likely associated with functional impairments in some of the brain’s neurotransmitter systems, particularly those involving dopamine and norepinephrine. To date, a large number of studies on different candidate genes for ADHD have been published (dopaminergic neurotransmission system: DRD4, DAT1/SLC6A3, DRD5, COMT, and DBH; noradrenergic neurotransmission system: NET1/SLC6A2, ADRA2A, and ADRA2C; serotonergic: 5-HTT/SLC6A4, HTR1B, HTR2A, and TPH2; and central nervous system development pathway: SNAP25 and BDNF) in the etiology of ADHD [17–20]. Alterations in the dopamine and norepinephrine pathways can impair the prefrontal cortex (PFC) function which is responsible for integrating cortical and subcortical inputs to execute essential cognitive functions such as attention, motivation, working memory planning, and decision making [21–24]. There may additionally be abnormalities of this disorder in serotoninergic and cholinergic pathways [25, 26]. And, the synaptosomal-associated protein 25 (SNAP-25) is interesting for its involvement in a number of processes including axonal growth, synaptic plasticity, and the vesicular release of neurotransmitters [27, 28].
In addition, the environmental factors such as prenatal nicotine [29], alcohol [30] or lead exposure, and low birth weighting [31] are believed to play a lesser role in the pathogenesis of ADHD.
Structure of SNAP-25
Synaptosomal-associated protein 25 gene (SNAP25 [MIM 600322]) is located at chromosome 20p11.2 [32] in humans. Differential splicing of SNAP25 results in the expression of two transcripts, SNAP25a and SNAP25b [33, 34]. These splice variants differ by only 9 out of 206 amino acids, a result of differential usage of two alternative exon 5 sequences (exon 5a/5b). SNAP-25 has been identified in contributing two α-helix motifs to the N-ethylmalemide-sensitive factor attachment protein receptor (SNARE) complex [35], SN1 and SN2, as revealed from the crystal structure of the four-α-helix domain complex [36]. The C-terminus of SNAP-25 SN2 is known to be the target of botulinum neurotoxins A and E (BoNT/A and BoNT/E), which block the release of neurotransmitters in vivo [37–39]. Biochemical analyses have shown that SNAP-25 amino acids (AA) 7–83 and 141–204 are essential motifs that are spontaneously assembled into helical SNARE complexes with Syntaxin1 and synaptobrevin 2 motifs [40]. SNAP-25 mutations introduced to the C terminal of the protein at AA positions 78, 81, and 202 resulted in a near elimination of exocytosis [40].
Functions of SNAP-25
Regulating Neural Signal Transmission: Role as a SNARE Protein (Fig. 1)
SNAP-25 is a presynaptic membrane bound protein which is anchored to the cytosolic surface of membranes via palmitoyl side chains located in the central region of the molecule [41, 42]. Together with synaptobrevin/VAMP and syntaxin/HPC-1, SNAP-25 constitutes the soluble SNARE protein core complex [43, 44] which is essential for docking and holding synaptic vesicles at the presynaptic membrane in preparation for Ca2+-triggered neurotransmitter exocytosis [42, 45–47]. SNAP-25 has been identified in contributing two α-helices to the SNARE complex, one located around the center and another at the C-terminal end of the SNARE bundle [48, 49]. At the presynaptic plasma membrane site wherein SNAP-25 located primarily, on contact, SNARE protein complex is initiated amino-terminally and proceeds toward the C terminus in a zipper-like fashion, thus pulling the synaptic vesicle and the presynaptic membranes together [40, 50]. Recently, it has been discovered that the interaction with the central SNARE motifs of SNAP-25 is essential for vesicle docking, priming, and fast fusion-triggering exocytosis. As to the C-terminal binding interface, it only plays a subsidiary role in triggering but is required for the full size of the readily releasable pool [49].
Regulating Neural Signal Transmission: Role as a Cellular Calcium Modulator (Fig. 1)
Calcium channel activity, as a kind of second messenger, has an immediate impact on synaptic activity [51]. SNAP-25 interacts with different types of voltage-gated calcium channels (VGCCs) [52], including N-type [53], P/Q-type [54, 55], and L-type [56, 57], inhibiting their function and thus reducing neuronal calcium responsiveness to depolarization [58–60]. It is notable that the different neuropsychiatric alterations where SNAP-25 has been involved are characterized by a dysregulation of calcium homeostasis [61, 62]. Different levels of SNAP-25 expression in excitatory versus inhibitory neurons may profoundly modulate neuronal responses to synaptic stimuli in a dose-dependent manner, and therefore, SNAP-25 is involved in the regulation of neuronal excitability [58]. SNAP-25 has been discovered to be a target of protein kinase C (PKC) on its residue serine in position 187 (Ser187), and PKC phosphorylation of SNAP-25 at Ser187 was found to be crucial for the negative regulation of VGCCs [59]. For the reason that Ser187 phosphorylation is transiently induced by neuronal activity [59], it is suggested that SNAP-25 provides a negative feedback mechanism for controlling neuronal excitability. It is also possible that the effects of reducing endogenous SNAP-25 expression have a greater impact on VGCC regulation than on the function of the protein as a SNARE [63, 64].
Axonal Growth and Synaptic Plasticity
Evidence derived from organisms ranging from reptiles such as geckos to humans indicates that SNAP-25 can promote outgrowth and elongation of neuritis [65, 66]. A high level of SNAP-25 expression in the adult brain was found to contribute to nerve terminal plasticity [43]. Upregulating and downregulating SNAP-25 by combined lentiviral packaging and siRNA, the result that SNAP-25 is specific for neural remodeling has been obtained [67]. Findings that selective inhibition of SNAP-25 expression imposed restrictions on neurite outgrowth have been reported [27, 68–70]. In addition, SNAP25 is associated with neuronal maturation and synaptogenesis during development [71] and also affects the expression of receptors like NMDARs in the plasma membrane [72, 73]. These findings indicate that SNAP25 may be involved in a mechanism relevant to axonal growth and synaptic plasticity.
SNAP-25 and ADHD
According to the functions of SNAP-25, it is possible that any variation in SNAP-25 which located primarily and specifically in axons and nerve terminals [67, 68] might interfere in the susceptibility of ADHD by influencing the release of neurotransmitters [74–76] and establishing neural circuits during central nervous system (CNS) development [77].
SNAP25 and ADHD Mouse Model
The coloboma mouse mutant, heterozygous for a ∼2-cM deletion of chromosome 2 (Cm) that encompasses the SNAP-25 gene and therefore exhibiting 50 % reduction of SNAP-25 expression [78], is considered an animal model of ADHD, as it displays certain hallmarks of ADHD including mainly locomotor hyperactivity [79] as well as inattention and impulsivity [80]. Furthermore, the hyperactive phenotype of coloboma mouse has been shown to be ameliorated by the psychostimulant d-amphetamine or with a transgenic insertion [81–83]. Animal studies have also shown involvements of SNAP-25 in neurotransmitter systems like dopamine and norepinephrine pathways [84, 85]. A significant reduction of Ca2+-dependent dopamine release from the dorsal striatum region but not ventral striatum [82, 86] and an increase of up to 40 % in noradrenaline within the striatum and the nucleus accumbens [87] of the coloboma (Cm/+) mouse have been reported. Additionally, an enhanced calcium response through L-type channels has been described in the SHR model of ADHD, where a reduction of SNAP-25 also occurs [88]. Consequently, SNAP-25 mediates neurotransmitter release like acetylcholine [89, 90] and plays essential roles in neurotransmitter release at different steps. These findings represent therefore that the gene SNAP25 could play a part in several heritable neurocognitive and behavioral abnormalities [91].
The Genetic Variation of SNAP25 and ADHD
Using a transmission disequilibrium test (TDT), Barr et al. [92] found a trend of excess transmission of the C allele of rs1051312 in Canadian and Brophy et al. [93] reported preferential transmission of allele T of rs1051312 in Irish ADHD cases. Significant association between rs3746544 (1065 T>G) and ADHD was reported in Chinese [94] and Colombian [95] populations in case–control studies, while negative results were reported in Irish [21, 93], Indian [96], Canadian [92, 97], US Caucasian [97], and UK Caucasian [98] populations. Feng et al. [97] examined 12 SNPs in two independent samples of ADHD families and found significant over-transmission of the rs66039806-C, rs362549-A, rs362987-A, and the rs362998-C alleles which located in introns 2, 4 and 4 and exon 6, respectively, in the Canadian sample, but not in the southern California sample. When they tested Canadian sample with quantitative analysis for hyperactivity and inattention ADHD subtypes, they found associations for both of behavioral ADHD subtypes with SNAP25. Mill [98] and Kim [99] reported significant association with additional SNPs rs363006 (intron 7) and rs3787283 (intron 7), respectively. Nevertheless, several studies [100, 101] have yielded negative results for the association of ADHD with the polymorphisms mentioned above.
In our study, we carry out a comprehensive meta-analysis to summarize the associations of the reported polymorphisms of SNAP-25 gene (rs3746544, rs363006, rs1051312, rs8636, rs362549, and rs362998) (Fig. 2) with childhood ADHD.
Methods
Study Sample Identification and Inclusion/Exclusion Criteria
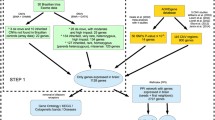
We used a three-stage approach to identify relevant studies for meta-analysis. Firstly, we conducted searches of four databases—PubMed, Web of Science, Elsevier, and Google Scholar—to identify an initial set of articles. These potentially relevant reports were published in a definite time from June 1996 to February 2016. The search terms used to query the databases included “ADHD” or “attention deficit hyperactivity disorder” and related terms such as “ADD,” “attention deficit,” “inattention,” and “hyperactivity.” Each of these terms was combined with “Synaptosomal-associated protein 25” or “SNAP25” to conduct searches. Then, we searched reference lists of relevant review and original published studies identified in the first stage to identify additional studies that might have been missed in stage 1. Finally, to determine which studies would be included in the meta-analysis for a given polymorphism, a series of inclusion and exclusion criteria were applied as follows: (1) studies evaluating the association between SNAP-25 gene and ADHD were included, and other pharmacological, functional, biochemical, or animal model studies were excluded; (2) family-based studies employing haplotype-based haplotype relative risk (HHRR) procedure or transmission disequilibrium test (TDT) and case–control studies were included; (3) studies including SNPs (rs3746544, rs1051312 et al.) of SNAP-25 gene and containing useful original data which was able to calculate the odds ratio (OR) were included; (4) each included study was required to report data from an independent sample; (5) only studies that used children samples (age range from 7 to 16) were included, and studies using adult samples were excluded from the present investigation; (6) all ADHD probands or cases used in each included study met DSM-IV/III criteria; and (7) all subjects used in each included study had an intelligence quotient (IQ) test score above 70 and were free of other neurological disorders.
Meta-Analytic Methods
Heterogeneity in effect sizes across studies was assessed using the Q-statistic [102], and its magnitude was quantified using I 2 [103], which is an index that describes the proportion of total variation in study effect size. The heterogeneity was considered significant when P < 0.10 for Q-statistic and qualified by I 2. Mild heterogeneity might account for less than 30 % of the variability in point estimates, notable heterogeneity more than 50 %, and moderate heterogeneity between them [103]. For TDT studies, ORs were estimated from the number of transmissions versus non-transmissions of the designated “high-risk” allele to ADHD cases from heterozygous parents. For case–control studies, ORs were estimated by contrasting the ratio of counts of the “high-risk” versus “low-risk” alleles in ADHD cases versus non-disordered controls. We calculated the pooled ORs with 95 % CIs and drew forest plots. A fixed effect model (FEM) was applied if the heterogeneity is not statistically significant (P < 0.05); otherwise, a random effect model (REM) was adopted [104]. Funnel plot, Egger’s linear regression model, and Begg’s rank correlation test were applied to evaluate the evidence for publication bias, and no publication bias exists when P > 0.05. Sensitivity analyses were performed if the heterogeneity was moderate and notable to estimate the sources of heterogeneity. The meta-analysis was performed by the metafor package (version 1.9-5; http://cran.r-project.org/web/packages/metafor/index.html) (Nicodemus KK 2008 [105]) in R (version 3.1.2; http://www.r-project.org/).
Results
A total of 14 studies were finally included after the meta-analysis literature selection, and the flow chart is shown in Supplementary Fig. 1. Ten were TDT designs, three were case–control designs, and one study adopted both methods. When one study employed two association analysis methods, data from the method that presented the largest dataset were included according to the finding that estimates of association were equivalent in aggregate across methods with observed differences most likely due to uncertainty in the estimates resulting from small sample sizes [106]. As a result, data from these 14 studies that analyzed six common variants within the SNAP25 gene was applied for meta-analytical procedures (Supplementary Fig. 2) and the descriptive characteristics of the studies are shown in Tables 1 and 2. Meta-analytic results for associations between candidate gene polymorphisms in the SNAP 25 gene and childhood ADHD and general information about the SNPs are shown in Table 3.
For the meta-analysis of the association between childhood ADHD and rs3746544, eight TDT studies and four case–control studies were identified. Moderate heterogeneity in effect sizes across studies was observed (Q-statistic χ 2 = 19.3528, P = 0.055, I 2 = 41.56 %). The pooled results (Fig. 3a) indicated a significant and modest association between ADHD and the “T” allele (fixed effects: OR = 1.14, 95 % CI 1.03–1.26, P = 0.010). For sensitivity analysis, three of the pooled ORs changed qualitatively after excluding one single study each time. It suggested that the results of this meta-analysis were not stable (Supplementary Table 1). The funnel plot was generally symmetrical, showing no evidence of publication bias (Supplementary Fig. 3a). And, Egger’s test (P = 0.109) and Begg’s test (P = 0.153) also suggested that publication bias was not significant.
Forest plot OR and pooled OR from the meta-analysis of ADHD and SNAP-25 SNPs. a The single study and pooled ORs for the association between ADHD and rs3746544 (pooled OR = 1.14, 95 % CI 1.03–1.26, P = 0.010; Q-statistic χ 2 = 19.3528, P = 0.055, I 2 = 41.56 %). b The single study and pooled ORs for the association between ADHD and rs1051312 (pooled OR = 0.96, 95 % CI 0.79–1.17, P = 0.688; Q-statistic χ 2 = 16.207, P = 0.040, I 2 = 50.78 %). c The single study and pooled ORs for the association between ADHD and rs363006 (pooled OR = 1.04, 95 % CI 0.87–1.25, P = 0.667; Q-statistic χ 2 = 5.937, P = 0.312, I 2 = 0.00 %). d The single study and pooled ORs for the association between ADHD and rs8636 (pooled OR = 1.09, 95 % CI 0.93–1.27, P = 0.289; Q-statistic χ 2 = 0.831, P = 0.842, I 2 = 0.00 %). e The single study and pooled ORs for the association between ADHD and rs362549 (pooled OR = 1.14, 95 % CI 0.97–1.34, P = 0.122; Q-statistic χ 2 = 5.849, P = 0.119, I 2 = 47.61 %). f The single study and pooled ORs for the association between ADHD and rs362998 (pooled OR = 1.31, 95 % CI 0.98–1.75, P = 0.069; Q-statistic χ 2 = 3.965, P = 0.265, I 2 = 0.00 %)
The meta-analysis for rs363006 (Fig. 3c) included six TDT studies. The heterogeneity in effect sizes across studies was mild (Q-statistic χ 2 = 5.9374, P = 0.3124, I 2 = 0.00 %). The pooled results were non-significant (fixed effects: OR = 1.04, 95 % CI 0.87–1.25, P = 0.667). And, no significant publication bias existed (Egger’s test: P = 0.514; Begg’s test: P = 0.272).
As to the meta-analysis for rs1051312 (Fig. 3b), seven TDT studies and two case–control studies were included. The pooled results did not support the association with ADHD (random effects: OR = 0.96, 95 % CI 0.79–1.17, P = 0.688). Notable heterogeneity in effect sizes across studies was observed (Q-statistic χ 2 = 16.207, P = 0.040, I 2 = 50.78 %). Subsequently, sensitivity analysis was performed and the pooled ORs did not change qualitatively after excluding one single study each time. It suggested that the results of this meta-analysis were stable (Supplementary Table 1). The funnel plot (Supplementary Fig. 3b) was substantially symmetrical, with Egger’s test (P = 0.763) and Begg’s test (P = 0.920) results shown no evidence of publication bias.
Each of the meta-analyses for rs8636 (Fig. 3d), rs362549 (Fig. 3e), and rs362998 (Fig. 3f) included four studies which is the low threshold for meta-analysis procedure. The pooled results did not indicate any association with ADHD (fixed effects: OR = 1.09, 95 % CI 0.93–1.27, P = 0.289; OR = 1.14, 95 % CI 0.97–1.34, P = 0.122; OR = 1.31, 95 % CI 0.98–1.75, P = 0.069, respectively). The heterogeneity (Q-statistic χ 2 = 0.831, P = 0.842, I 2 = 0.00 %; Q-statistic χ 2 = 5.849, P = 0.119, I 2 = 47.61 %; Q-statistic χ 2 = 3.965, P = 0.265, I 2 = 0.00 %, respectively) in effect sizes across studies was mild for rs8636 and rs362998 and moderate for rs362549. We did sensitivity analysis for rs362549, and the pooled ORs did not change qualitatively after excluding one single study each time, suggesting that the results of this meta-analysis were stable (Supplementary Table 1). Egger’s test (P = 0.100; P = 0.425; P = 0.728, respectively) and Begg’s test (P = 0.750; P = 0.750; P = 0.750, respectively) results did not show any evidence of publication bias.
Discussion
Imaging studies have suggested the contribution of a wider range of dysfunctions of neural networks to the diversity of ADHD symptoms [23]. Based on the effect of psychostimulants used in the pharmacological treatment of ADHD [107, 108], such as methylphenidate or amphetamines, dysfunctions in neuroplasticity mechanisms and synapses have been postulated to be involved in the pathogenesis of ADHD [109–111]. Specifically, the lowered expression of SNAP-25 in regions that are critical for attention and inhibition, such as inferior frontal gyrus (IFG), may ultimately decrease the efficiency of neurotransmitter release and synaptic function, impair behavior and cognition, and confer risk to ADHD [21]. Furthermore, the physiological functions of SNAP-25 in docking and fusion of synaptic vesicles in presynaptic neurons, as well as in axonal growth and synaptic plasticity, also make SNAP25 an important candidate gene for ADHD.
In the present study, we investigated the association of six SNPs within SNAP25 with childhood ADHD and did find some evidences to support the association. In a latest meta-analysis, Gizer and colleagues [20] evaluated four previously studied SNPs in the 3′-UTR and introns of SNAP25 in association with ADHD and found only one significant association (a pooled OR of 1.15 for the T allele of rs3746544 was found, including data from seven studies). When we extract data from previous studies for a pooled analysis, also only significant mild association between rs3746544 and childhood ADHD was observed. These results are in accordance with Gizer’s [20] and the previous meta-analyses [112, 113]. Although the association between 3′-UTR rs3746544 polymorphism and childhood ADHD existed moderate heterogeneity in effect sizes across studies in our meta-analysis, it still generated the even increased positive evidence for this association (OR 1.21, 95 % CI 1.08–1.35) when the study by Hawi et al. [21] according to the sensitivity analysis was removed.
In contrast to this positive result, non-significant combined results between the rest of five SNPs (rs1051312-T, rs363006-G, rs8636-T, rs362549-A, and rs362998-C) and childhood ADHD were observed, which are also uniform with the meta-analysis by Gizer [20]. And, to our knowledge, the meta-analysis results for rs8636, rs362549, and rs362998 are the first time to be reported. It is worthwhile to note that the association between rs1051312 and rs362549 polymorphisms and childhood ADHD existed notable and moderate heterogeneity in effect sizes across studies, respectively. However, when further sensitivity analyses were performed, none of the results changed qualitatively after excluding one single study each time. This indicated that our negative meta-analysis results were stable.
Heterogeneity in effect sizes across studies characterized several of the reviewed markers, including some that showed significant evidence for association and others that did not. This highlights the need for future studies that examine differences in methodological aspects and sample characteristics that can explain such heterogeneity and point to ways of maximizing associations.
All of the associations that we meta-analyzed and reviewed herein were for single polymorphisms in genes, but multiple polymorphisms and/or their interactions were also observed. For example, strong linkage disequilibriums (LDs) between rs3746544-rs8636 in subjects from USA and Canada [97], British [114], and Indian [96] as well as LD between rs1051312-rs8636 [96, 97] and rs3746544-rs1051312 [96] were reported. Other SNPs in SNAP25 which were not presented in our study like rs1889189 and rs362569 were also reported to have a LD in Dutch [115]. Such samples both from single site and multi-sites studies will facilitate the detection of replicable associations and be more accurate estimation of the magnitude of risk conferred [116]. Thus, it is possible that the SNPs that we estimated might have allele-dependent functional effects and probably in LD with genetic variations (located in protein-coding or regulatory regions) of functional relevance.
The sample included in our meta-analysis is obviously insufficient compared with the large samples in the genetic studies on other psychiatric diseases (the meta-analysis for schizophrenia is provided with approximately 3500 or even more participants on average [117]). This can be partly explained by the use of family-based approaches like the TDT adopted by most of our included studies, which have the inherent disadvantage of the effective samples being substantially smaller than the initial samples [118].
Several of the papers reported associations in the stratified analysis. For example, a trend toward sex-dependent transmission of alleles from parents to ADHD probands has been reported earlier for the T alleles of rs3746544 [119] and rs1051312 [93], and in both of these investigations, the over-transmission was paternal. In addition, associations for both of behavioral ADHD subtypes with SNAP25 were reported in a Canadian sample [97]. Due to the lack of data, we did not do the further stratified analysis, but it is expected that future meta-analysis will do more detailed and stratified analysis.
Depending on the position and flanking sequence in the gene, SNPs may have varied functional effects on protein sequence, transcriptional regulation, RNA splicing, or microRNA (miRNA) binding. According to a web server SNPinfo (http://www.niehs.nih.gov/snpinfo) [120], rs3746544 located in 3′-UTR is to be predicted as a binding site of miRNAs (miRanda) which are ∼22-nucleotide-long endogenous non-coding RNA regulators of gene activity at the post-transcriptional level [121, 122]. This is also consistent with our meta-analysis and supports the association between SNAP25 and ADHD. However, the direct evidence is still missing.
In conclusion, we did provide modest support for one of the first reported markers (rs3746544, MnlI) of SNAP25 with ADHD, but we were not able to confirm the association of variants of SNAP25 with ADHD. In order to explore more effective and direct evidence, further extensive animal experiments and pharmacological studies and larger and more various and detailed genome-wide association studies are crucial and conclusive in light of the impact of SNAP25 on disease processes.
References
Kooij SJ, Bejerot S, Blackwell A et al (2010) European consensus statement on diagnosis and treatment of adult ADHD: the European Network Adult ADHD. BMC Psychiatry 10:67
Lange KW, Reichl S, Lange KM et al (2010) The history of attention deficit hyperactivity disorder. Atten Defic Hyperact Disord 2(4):241–255
Willcutt EG (2012) The prevalence of DSM-IV attention-deficit/hyperactivity disorder: a meta-analytic review. Neurotherapeutics 9(3):490–499
Emond V, Joyal C, Poissant H (2009) Structural and functional neuroanatomy of attention-deficit hyperactivity disorder (ADHD). Encéphale 35(2):107–114
Singh I (2008) Beyond polemics: science and ethics of ADHD. Nat Rev Neurosci 9(12):957–964
Childress AC, Berry SA (2012) Pharmacotherapy of attention-deficit hyperactivity disorder in adolescents. Drugs 72(3):309–325
Kenemans JL, Bekker EM, Lijffijt M et al (2005) Attention deficit and impulsivity: selecting, shifting, and stopping. Int J Psychophysiol 58(1):59–70
Biederman J, Faraone SV (2005) Attention-deficit hyperactivity disorder. Lancet 366(9481):237–248
Hall CL, Newell K, Taylor J et al (2013) ‘Mind the gap’—mapping services for young people with ADHD transitioning from child to adult mental health services. BMC Psychiatry 13:186
Swift KD, Hall CL, Marimuttu V et al (2013) Transition to adult mental health services for young people with attention deficit/hyperactivity disorder (ADHD): a qualitative analysis of their experiences. BMC Psychiatry 13:74
Simon V, Czobor P, Balint S et al (2009) Prevalence and correlates of adult attention-deficit hyperactivity disorder: meta-analysis. Br J Psychiatry 194(3):204–211
Faraone SV, Biederman J, Spencer T et al (2000) Attention-deficit/hyperactivity disorder in adults: an overview. Biol Psychiatry 48(1):9–20
Wilens TE, Faraone SV, Biederman J (2004) Attention-deficit/hyperactivity disorder in adults. JAMA 292(5):619–623
Advokat C, Martino L, Hill BD et al (2007) Continuous Performance Test (CPT) of college students with ADHD, psychiatric disorders, cognitive deficits, or no diagnosis. J Atten Disord 10(3):253–256
Thapar A, Cooper M, Eyre O et al (2013) What have we learnt about the causes of ADHD? J Child Psychol Psychiatry 54(1):3–16
Rhodes SM, Coghill DR, Matthews K (2004) Methylphenidate restores visual memory, but not working memory function in attention deficit-hyperkinetic disorder. Psychopharmacology (Berl) 175(3):319–330
Berry MD (2007) The potential of trace amines and their receptors for treating neurological and psychiatric diseases. Rev Recent Clin Trials 2(1):3–19
Sotnikova TD, Caron MG, Gainetdinov RR (2009) Trace amine-associated receptors as emerging therapeutic targets. Mol Pharmacol 76(2):229–235
Kebir O, Tabbane K, Sengupta S et al (2009) Candidate genes and neuropsychological phenotypes in children with ADHD: review of association studies. J Psychiatry Neurosci 34(2):88–101
Gizer IR, Ficks C, Waldman ID (2009) Candidate gene studies of ADHD: a meta-analytic review. Hum Genet 126(1):51–90
Hawi Z, Matthews N, Wagner J et al (2013) DNA variation in the SNAP25 gene confers risk to ADHD and is associated with reduced expression in prefrontal cortex. Plos One 8(4):e60274
Wallace TL, Bertrand D (2013) Importance of the nicotinic acetylcholine receptor system in the prefrontal cortex. Biochem Pharmacol 85(12):1713–1720
Castellanos FX, Proal E (2012) Large-scale brain systems in ADHD: beyond the prefrontal-striatal model. Trends Cogn Sci 16(1):17–26
Cortese S, Kelly C, Chabernaud C et al (2012) Toward systems neuroscience of ADHD: a meta-analysis of 55 fMRI studies. Am J Psychiatry 169(10):1038–1055
Bidwell LC, McClernon FJ, Kollins SH (2011) Cognitive enhancers for the treatment of ADHD. Pharmacol Biochem Behav 99(2):262–274
Cortese S (2012) The neurobiology and genetics of attention-deficit/hyperactivity disorder (ADHD): what every clinician should know. Eur J Paediatr Neurol 16(5):422–433
Osen-Sand A, Catsicas M, Staple JK et al (1993) Inhibition of axonal growth by SNAP-25 antisense oligonucleotides in vitro and in vivo. Nature 364(6436):445–448
Kovacs-Nagy R, Hu J, Ronai Z et al (2009) SNAP-25: a novel candidate gene in psychiatric genetics. Neuropsychopharmacol Hung 11(2):89–94
Abbott LC, Winzer-Serhan UH (2012) Smoking during pregnancy: lessons learned from epidemiological studies and experimental studies using animal models. Crit Rev Toxicol 42(4):279–303
Burger PH, Goecke TW, Fasching PA et al (2011) How does maternal alcohol consumption during pregnancy affect the development of attention deficit/hyperactivity syndrome in the child. Fortschr Neurol Psychiatr 79(9):500–506
Thapar A, Cooper M, Jefferies R et al (2012) What causes attention deficit hyperactivity disorder? Arch Dis Child 97(3):260–265
Maglott DR, Feldblyum TV, Durkin AS et al (1996) Radiation hybrid mapping of SNAP, PCSK2, and THBD (human chromosome 20p). Mamm Genome 7(5):400–401
Bark IC, Wilson MC (1994) Human cDNA clones encoding two different isoforms of the nerve terminal protein SNAP-25. Gene 139(2):291–292
Bark IC (1993) Structure of the chicken gene for SNAP-25 reveals duplicated exon encoding distinct isoforms of the protein. J Mol Biol 233(1):67–76
Pevsner J, Hsu SC, Braun JE et al (1994) Specificity and regulation of a synaptic vesicle docking complex. Neuron 13(2):353–361
Sutton RB, Fasshauer D, Jahn R et al (1998) Crystal structure of a SNARE complex involved in synaptic exocytosis at 2.4 A resolution. Nature 395(6700):347–353
Binz T, Blasi J, Yamasaki S et al (1994) Proteolysis of SNAP-25 by types E and A botulinal neurotoxins. J Biol Chem 269(3):1617–1620
Blasi J, Chapman ER, Link E et al (1993) Botulinum neurotoxin A selectively cleaves the synaptic protein SNAP-25. Nature 365(6442):160–163
Schiavo G, Rossetto O, Catsicas S et al (1993) Identification of the nerve terminal targets of botulinum neurotoxin serotypes A, D, and E. J Biol Chem 268(32):23784–23787
Stein A, Weber G, Wahl MC et al (2009) Helical extension of the neuronal SNARE complex into the membrane. Nature 460(7254):525–528
Hess DT, Slater TM, Wilson MC et al (1992) The 25 kDa synaptosomal-associated protein SNAP-25 is the major methionine-rich polypeptide in rapid axonal transport and a major substrate for palmitoylation in adult CNS. J Neurosci 12(12):4634–4641
Jahn R, Scheller RH (2006) SNAREs—engines for membrane fusion. Nat Rev Mol Cell Biol 7(9):631–643
Oyler GA, Higgins GA, Hart RA et al (1989) The identification of a novel synaptosomal-associated protein, SNAP-25, differentially expressed by neuronal subpopulations. J Cell Biol 109(6 Pt 1):3039–3052
Sudhof TC, Rothman JE (2009) Membrane fusion: grappling with SNARE and SM proteins. Science 323(5913):474–477
Chen YA, Scheller RH (2001) SNARE-mediated membrane fusion. Nat Rev Mol Cell Biol 2(2):98–106
Rizo J, Sudhof TC (2002) Snares and Munc18 in synaptic vesicle fusion. Nat Rev Neurosci 3(8):641–653
Jahn R, Lang T, Sudhof TC (2003) Membrane fusion. Cell 112(4):519–533
Chua JJ, Kindler S, Boyken J et al (2010) The architecture of an excitatory synapse. J Cell Sci 123(Pt 6):819–823
Mohrmann R, de Wit H, Connell E et al (2013) Synaptotagmin interaction with SNAP-25 governs vesicle docking, priming, and fusion triggering. J Neurosci 33(36):14417–14430
Jahn R, Fasshauer D (2012) Molecular machines governing exocytosis of synaptic vesicles. Nature 490(7419):201–207
Zamponi GW (2003) Regulation of presynaptic calcium channels by synaptic proteins. J Pharmacol Sci 92(2):79–83
Catterall WA, Few AP (2008) Calcium channel regulation and presynaptic plasticity. Neuron 59(6):882–901
Sheng ZH, Rettig J, Cook T et al (1996) Calcium-dependent interaction of N-type calcium channels with the synaptic core complex. Nature 379(6564):451–454
Rettig J, Sheng ZH, Kim DK et al (1996) Isoform-specific interaction of the alpha1A subunits of brain Ca2+ channels with the presynaptic proteins syntaxin and SNAP-25. Proc Natl Acad Sci U S A 93(14):7363–7368
Martin-Moutot N, Charvin N, Leveque C et al (1996) Interaction of SNARE complexes with P/Q-type calcium channels in rat cerebellar synaptosomes. J Biol Chem 271(12):6567–6570
Wiser O, Bennett MK, Atlas D (1996) Functional interaction of syntaxin and SNAP-25 with voltage-sensitive L- and N-type Ca2+ channels. Embo J 15(16):4100–4110
Wiser O, Trus M, Hernandez A et al (1999) The voltage sensitive Lc-type Ca2+ channel is functionally coupled to the exocytotic machinery. Proc Natl Acad Sci U S A 96(1):248–253
Verderio C, Pozzi D, Pravettoni E et al (2004) SNAP-25 modulation of calcium dynamics underlies differences in GABAergic and glutamatergic responsiveness to depolarization. Neuron 41(4):599–610
Pozzi D, Condliffe S, Bozzi Y et al (2008) Activity-dependent phosphorylation of Ser187 is required for SNAP-25-negative modulation of neuronal voltage-gated calcium channels. Proc Natl Acad Sci U S A 105(1):323–328
Condliffe SB, Corradini I, Pozzi D et al (2010) Endogenous SNAP-25 regulates native voltage-gated calcium channels in glutamatergic neurons. J Biol Chem 285(32):24968–24976
Lidow MS (2003) Calcium signaling dysfunction in schizophrenia: a unifying approach. Brain Res Brain Res Rev 43(1):70–84
Braunewell KH (2005) The darker side of Ca2+ signaling by neuronal Ca2+-sensor proteins: from Alzheimer’s disease to cancer. Trends Pharmacol Sci 26(7):345–351
Bronk P, Deak F, Wilson MC et al (2007) Differential effects of SNAP-25 deletion on Ca2+ -dependent and Ca2+ -independent neurotransmission. J Neurophysiol 98(2):794–806
Delgado-Martinez I, Nehring RB, Sorensen JB (2007) Differential abilities of SNAP-25 homologs to support neuronal function. J Neurosci 27(35):9380–9391
Wang Y, Dong Y, Song H et al (2012) Involvement of gecko SNAP25b in spinal cord regeneration by promoting outgrowth and elongation of neurites. Int J Biochem Cell Biol 44(12):2288–2298
Martinez-Arca S, Coco S, Mainguy G et al (2001) A common exocytotic mechanism mediates axonal and dendritic outgrowth. J Neurosci 21(11):3830–3838
Wang W, Wang F, Liu J et al (2014) SNAP25 ameliorates sensory deficit in rats with spinal cord transection. Mol Neurobiol 50(2):290–304
Aikawa Y, Xia X, Martin TF (2006) SNAP25, but not syntaxin 1A, recycles via an ARF6-regulated pathway in neuroendocrine cells. Mol Biol Cell 17(2):711–722
Osen-Sand A, Staple JK, Naldi E et al (1996) Common and distinct fusion proteins in axonal growth and transmitter release. J Comp Neurol 367(2):222–234
Wu CS, Lin JT, Chien CL et al (2011) Type VI adenylyl cyclase regulates neurite extension by binding to Snapin and Snap25. Mol Cell Biol 31(24):4874–4886
Catsicas S, Larhammar D, Blomqvist A et al (1991) Expression of a conserved cell-type-specific protein in nerve terminals coincides with synaptogenesis. Proc Natl Acad Sci U S A 88(3):785–789
Selak S, Paternain AV, Aller MI et al (2009) A role for SNAP25 in internalization of kainate receptors and synaptic plasticity. Neuron 63(3):357–371
Lau CG, Takayasu Y, Rodenas-Ruano A et al (2010) SNAP-25 is a target of protein kinase C phosphorylation critical to NMDA receptor trafficking. J Neurosci 30(1):242–254
Thapar A, O’Donovan M, Owen MJ (2005) The genetics of attention deficit hyperactivity disorder. Hum Mol Genet 14(2):R275–R282
Rizo J, Sudhof TC (2012) The membrane fusion enigma: SNAREs, Sec1/Munc18 proteins, and their accomplices—guilty as charged? Annu Rev Cell Dev Biol 28:279–308
Washbourne P, Thompson PM, Carta M et al (2002) Genetic ablation of the t-SNARE SNAP-25 distinguishes mechanisms of neuroexocytosis. Nat Neurosci 5(1):19–26
Bark C, Bellinger FP, Kaushal A et al (2004) Developmentally regulated switch in alternatively spliced SNAP-25 isoforms alters facilitation of synaptic transmission. J Neurosci 24(40):8796–8805
Hess EJ, Jinnah HA, Kozak CA et al (1992) Spontaneous locomotor hyperactivity in a mouse mutant with a deletion including the Snap gene on chromosome 2. J Neurosci 12(7):2865–2874
Steffensen SC, Wilson MC, Henriksen SJ (1996) Coloboma contiguous gene deletion encompassing Snap alters hippocampal plasticity. Synapse 22(3):281–289
Bruno KJ, Freet CS, Twining RC et al (2007) Abnormal latent inhibition and impulsivity in coloboma mice, a model of ADHD. Neurobiol Dis 25(1):206–216
Hess EJ, Collins KA, Wilson MC (1996) Mouse model of hyperkinesis implicates SNAP-25 in behavioral regulation. J Neurosci 16(9):3104–3111
Wilson MC (2000) Coloboma mouse mutant as an animal model of hyperkinesis and attention deficit hyperactivity disorder. Neurosci Biobehav Rev 24(1):51–57
Steffensen SC, Henriksen SJ, Wilson MC (1999) Transgenic rescue of SNAP-25 restores dopamine-modulated synaptic transmission in the coloboma mutant. Brain Res 847(2):186–195
Jones MD, Williams ME, Hess EJ (2001) Abnormal presynaptic catecholamine regulation in a hyperactive SNAP-25-deficient mouse mutant. Pharmacol Biochem Behav 68(4):669–676
Fortin GD, Desrosiers CC, Yamaguchi N et al (2006) Basal somatodendritic dopamine release requires snare proteins. J Neurochem 96(6):1740–1749
Raber J, Mehta PP, Kreifeldt M et al (1997) Coloboma hyperactive mutant mice exhibit regional and transmitter-specific deficits in neurotransmission. J Neurochem 68(1):176–186
Jones MD, Williams ME, Hess EJ (2001) Expression of catecholaminergic mRNAs in the hyperactive mouse mutant coloboma. Brain Res Mol Brain Res 96(1-2):114–121
Li Q, Wong JH, Lu G et al (2009) Gene expression of synaptosomal-associated protein 25 (SNAP-25) in the prefrontal cortex of the spontaneously hypertensive rat (SHR). Biochim Biophys Acta 1792(8):766–776
Nagy G, Milosevic I, Fasshauer D et al (2005) Alternative splicing of SNAP-25 regulates secretion through nonconservative substitutions in the SNARE domain. Mol Biol Cell 16(12):5675–5685
Chapman ER (2002) Synaptotagmin: a Ca(2+) sensor that triggers exocytosis? Nat Rev Mol Cell Biol 3(7):498–508
Corradini I, Donzelli A, Antonucci F et al (2014) Epileptiform activity and cognitive deficits in SNAP-25(+/-) mice are normalized by antiepileptic drugs. Cereb Cortex 24(2):364–376
Barr CL, Feng Y, Wigg K et al (2000) Identification of DNA variants in the SNAP-25 gene and linkage study of these polymorphisms and attention-deficit hyperactivity disorder. Mol Psychiatry 5(4):405–409
Brophy K, Hawi Z, Kirley A et al (2002) Synaptosomal-associated protein 25 (SNAP-25) and attention deficit hyperactivity disorder (ADHD): evidence of linkage and association in the Irish population. Mol Psychiatry 7(8):913–917
Gao XP, Su LY, Zhao AL et al (2009) Association of 14 polymorphisms in the five candidate genes and attention deficit hyperactivity disorder. Zhongguo Dang Dai Er Ke Za Zhi 11(8):617–622
Galvez JM, Forero DA, Fonseca DJ et al (2014) Evidence of association between SNAP25 gene and attention deficit hyperactivity disorder in a Latin American sample. Atten Defic Hyperact Disord 6(1):19–23
Sarkar K, Bhaduri N, Ghosh P et al (2012) Role of SNAP25 explored in eastern Indian attention deficit hyperactivity disorder probands. Neurochem Res 37(2):349–357
Feng Y, Crosbie J, Wigg K et al (2005) The SNAP25 gene as a susceptibility gene contributing to attention-deficit hyperactivity disorder. Mol Psychiatry 10(11):998–1005, 973
Mill J, Richards S, Knight J et al (2004) Haplotype analysis of SNAP-25 suggests a role in the aetiology of ADHD. Mol Psychiatry 9(8):801–810
Kim JW, Biederman J, Arbeitman L et al (2007) Investigation of variation in SNAP-25 and ADHD and relationship to co-morbid major depressive disorder. Am J Med Genet B Neuropsychiatr Genet 144B(6):781–790
Ilott NE, Saudino KJ, Asherson P (2010) Genetic influences on attention deficit hyperactivity disorder symptoms from age 2 to 3: a quantitative and molecular genetic investigation. BMC Psychiatry 10:102
Renner TJ, Walitza S, Dempfle A et al (2008) Allelic variants of SNAP25 in a family-based sample of ADHD. J Neural Transm 115(2):317–321
Lau J, Ioannidis JP, Schmid CH (1997) Quantitative synthesis in systematic reviews. Ann Intern Med 127(9):820–826
Higgins JP, Thompson SG (2002) Quantifying heterogeneity in a meta-analysis. Stat Med 21(11):1539–1558
Schmidt FL, Oh IS, Hayes TL (2009) Fixed- versus random-effects models in meta-analysis: model properties and an empirical comparison of differences in results. Br J Math Stat Psychol 62(Pt 1):97–128
Nicodemus KK (2008) Catmap: case-control and TDT meta-analysis package. BMC Bioinf 9:130
Evangelou E, Trikalinos TA, Salanti G et al (2006) Family-based versus unrelated case-control designs for genetic associations. PLoS Genet 2(8):e123
Stevenson RD, Wolraich ML (1989) Stimulant medication therapy in the treatment of children with attention deficit hyperactivity disorder. Pediatr Clin North Am 36(5):1183–1197
Greenhill L, Beyer DH, Finkleson J et al (2002) Guidelines and algorithms for the use of methylphenidate in children with attention-deficit/hyperactivity disorder. J Atten Disord 6(Suppl 1):S89–S100
Jensen V, Rinholm JE, Johansen TJ et al (2009) N-methyl-D-aspartate receptor subunit dysfunction at hippocampal glutamatergic synapses in an animal model of attention-deficit/hyperactivity disorder. Neuroscience 158(1):353–364
Forero DA, Casadesus G, Perry G et al (2006) Synaptic dysfunction and oxidative stress in Alzheimer’s disease: emerging mechanisms. J Cell Mol Med 10(3):796–805
Ramocki MB, Zoghbi HY (2008) Failure of neuronal homeostasis results in common neuropsychiatric phenotypes. Nature 455(7215):912–918
Forero DA, Arboleda GH, Vasquez R et al (2009) Candidate genes involved in neural plasticity and the risk for attention-deficit hyperactivity disorder: a meta-analysis of 8 common variants. J Psychiatry Neurosci 34(5):361–366
Faraone SV, Perlis RH, Doyle AE et al (2005) Molecular genetics of attention-deficit/hyperactivity disorder. Biol Psychiatry 57(11):1313–1323
Carroll LS, Kendall K, O’Donovan MC et al (2009) Evidence that putative ADHD low risk alleles at SNAP25 may increase the risk of schizophrenia. Am J Med Genet B Neuropsychiatr Genet 150B(7):893–899
Gosso MF, de Geus EJ, van Belzen MJ et al (2006) The SNAP-25 gene is associated with cognitive ability: evidence from a family-based study in two independent Dutch cohorts. Mol Psychiatry 11(9):878–886
Xu X, Rakovski C, Xu X et al (2006) An efficient family-based association test using multiple markers. Genet Epidemiol 30(7):620–626
Allen NC, Bagade S, McQueen MB et al (2008) Systematic meta-analyses and field synopsis of genetic association studies in schizophrenia: the SzGene database. Nat Genet 40(7):827–834
Cordell HJ, Clayton DG (2005) Genetic association studies. Lancet 366(9491):1121–1131
Kustanovich V, Merriman B, McGough J et al (2003) Biased paternal transmission of SNAP-25 risk alleles in attention-deficit hyperactivity disorder. Mol Psychiatry 8(3):309–315
Xu Z, Taylor JA (2009) SNPinfo: integrating GWAS and candidate gene information into functional SNP selection for genetic association studies. Nucleic Acids Res 37:W600–W605, Web Server issue
Ambros V (2004) The functions of animal microRNAs. Nature 431(7006):350–355
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116(2):281–297
Acknowledgments
This study is supported partially by the National Natural Science Foundation of China (81361120245, 31571039, 81101016, 81400816), Top-Notch Young Talents Program of China of 2014 to Dr. Ling-Qiang Zhu, Program of Outstanding Youth of Hubei Province, China (2014CFA017), and Program for Changjiang Scholars and Innovative Research Team in University (No. IRT13016).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare that they have no conflicts of interest.
Additional information
Yun-Sheng Liu, Xuan Dai and Wei Wu contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 75 kb)
Rights and permissions
About this article
Cite this article
Liu, YS., Dai, X., Wu, W. et al. The Association of SNAP25 Gene Polymorphisms in Attention Deficit/Hyperactivity Disorder: a Systematic Review and Meta-Analysis. Mol Neurobiol 54, 2189–2200 (2017). https://doi.org/10.1007/s12035-016-9810-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-016-9810-9