Abstract
The p53-family member, p73, plays a key role in the development of the central nervous system (CNS), in senescence, and in tumor formation. The role of p73 in neuronal differentiation is complex and involves several downstream pathways. Indeed, in the last few years, we have learnt that TAp73 directly or indirectly regulates several genes involved in neural biology. In particular, TAp73 is involved in the maintenance of neural stem/progenitor cell self-renewal and differentiation throughout the regulation of SOX-2, Hey-2, TRIM32 and Notch. In addition, TAp73 is also implicated in the regulation of the differentiation and function of postmitotic neurons by regulating the expression of p75NTR and GLS2 (glutamine metabolism). Further still, the regulation of miR-34a by TAp73 indicates that microRNAs can also participate in this multifunctional role of p73 in adult brain physiology. However, contradictory results still exist in the relationship between p73 and brain disorders, and this remains an important area for further investigation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The transcription factor p73 belongs to the p53 family [1], involved in several complex biological processes including cell cycle arrest, DNA repair, apoptosis, metabolism, autophagy, and senescence [2–10]. Despite of evident functional overlapping functions (extensively reviewed in [1, 11]), the three p53 relatives also have distinct roles and functions, with p73 clearly involved in neurological developmental abnormalities [12].
The role of p53 in suppressing cancer and promoting cell death and senescence [6, 13, 14] is strongly supported by its frequent mutations in human cancers, mostly showing a gain of function able to promote tumorigenesis [15–19]. Conversely, p63 is a master regulator of epidermal development [20–23], including skin annexes such as breast, prostate, vibrissae, and teeth [24, 25]; it is also involved in the development of the heart [26, 27]. As a consequence of its role in the development of pluri-stratified epithelia, mutations in the Tp63 gene cause several human genetic syndromes including EEC syndrome (ectrodactyly, ectodermal dysplasia, and cleft lip/palate), SHFM syndrome (nonsyndromic split hand–split foot malformation), and AEC syndrome (ankyloblepharon, ectodermal defects, cleft lip/palate) [28–30].
p73 regulates cell survival and genomic stability, thus affecting cancer development [31–34]. Accordingly, over 70 % of the TAp73 selective knockout mice (TAp73−/−) show an increased susceptibility to both spontaneous and induced carcinogenesis [34]. In addition, TAp73−/− mice are infertile and exhibit hippocampal dysgenesis, indicating a role for the TAp73 isoform in the regulation of reproduction and in neuronal development [35–38].
The Tp73 gene is expressed as two main isoforms, TAp73 (transcribed from the P1 promoter) or ΔNp73 (transcribed from the P2 promoter), containing or not an N-terminal transactivation (TA) domain, codified by the first three exons [39, 40]. Like p53, TAp73 can transactivate target genes that regulate apoptosis and senescence [41, 42]. Through a direct competition for the promoter or by formation of inactive hetero-oligomeric complexes, the ΔNp73 protein acts as a dominant negative for TAp73 (and p53) and therefore shows an antiapoptotic effect [43].
In addition to the N-terminal isoforms, seven different alternative splicing C-terminal isoforms of p73 are expressed at the RNA level (α, β, γ, ζ, δ, ϵ, η), although it remains unclear if all these isoforms are expressed as proteins and their differential biological importance; indeed, no mouse model exists for their study [44–46].
All the transgenic mouse models for the various p73 isoforms show neurological defects (see Fig. 1), demonstrating the importance of p73 for neuronal development.
p73 Causes Neuronal Development Defects
The p73 knockout mouse (p73−/−), generated by the McKeon’s group in 2000, immediately revealed the importance of p73 in neuronal development [35]. These p73−/− mice displayed mild hydrocephalus at birth and hippocampal dysgenesis, characterized by an unusual organization of regions CA1 and CA3 and the dentate gyrus (DG). In particular, the DG lacks the infrapyramidal blade and the suprapyramidal blade is hypertrophied and extended. In addition, p73 expression was found only in Cajal-Retzius (CR) neurons that are distributed along the marginal zone of the cortex and in the molecular layer of the DG. Moreover, the expression of reelin, a glycoprotein involved in neuronal migration and a marker of CR neurons, was lost in the cortical and hippocampal marginal zone of the p73−/− mice, suggesting that loss of p73 leads to the disappearance of this cell type [47]. This could, in part, explain the observed hippocampal phenotype.
However, there are, of course, alternative explanations: The Kaplan group have found p73 to be essential for the survival of young postnatal and adult cortical neurons [48]. Indeed, the number of cortical neurons in p73−/− mice is normal at birth but decreases by postnatal day 14 (P14) as a consequence of enhanced cortical apoptosis peaking between P4 and P6 [48]. The same group has also observed that deletion of p73 leads to increased death of sympathetic neurons in the developing superior cervical ganglia [49].
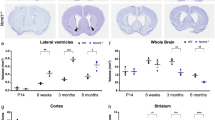
Because the strategy used to generate the first p73−/− mouse targeted the DNA-binding domain of p73, resulting in the loss of all isoforms, both TAp73 and ΔNp73 forms of each, it was impossible to discriminate the contribution of each major isoform type to the phenotype. An important step forward in our understanding of p73 in neurobiology came when, in collaboration with the Tak Mak lab, we generated the TA isoform selective p73 knockout mouse (TAp73−/−) [34]. Histological analysis of the TAp73−/− brain again revealed an abnormal hippocampal formation. In particular, the lower blade of the DG was missing or truncated. Notably, this abnormality occurs postnatally, between P6 and P14, suggesting that the TAp73 isoform is necessary for postnatal neurogenesis. In contrast, the size of the lateral ventricles and the thickness of the cortex were not affected by the loss of TAp73, indicating that TAp73 regulates hippocampal morphology while ΔNp73 isoforms, which are still expressed in this line, appear sufficient to prevent the loss of cortical neurons.
A direct antiapoptotic role for ΔNp73 in cortical neurons was subsequently demonstrated by Tissir and colleagues in a ΔNp73−/− mouse line [50]. Here, the selective inactivation of all ΔNp73 isoforms resulted in an increase in cell death in specific brain regions including the preoptic area and the vomeronasal organ and a reduction of CR and gonadotropin-releasing hormone (GnRH) neurons. This antiapoptotic effect was confirmed in a second ΔNp73 selective knockout mouse line. This ΔNp73−/− line also displayed some signs of neurodegeneration and a small reduction in cortical thickness and neuron number in older mice [31]. Although these studies have elegantly shown distinct roles for the p73 isoforms in the CNS, it should be noted that the phenotypes displayed by the isoform selective knockouts are milder than that observed in the total knockout lacking both TA and ΔN isoforms.
The hippocampus plays a central role in many aspects of memory [51], and the above studies indicate that the TAp73 isoform plays an important role in the morphogenesis of this structure, at least: Does such abnormal hippocampal anatomy have behavioral consequences? The p73−/− mice show a general reduction in performance across several behavioral tests. Performance in the Barnes maze, a test of spatial learning and memory formation, is markedly impaired in p73−/− mice [52]. The p73−/− mice also have impaired reflex and neuromuscular function and sensorimotor coordination and increased anxiety. Many of these behavioral abnormalities observed in the p73−/− mice were also found in the TAp73−/− mice [36]. Hippocampal dysfunction is specifically associated with a reduction in burrowing and open field performance [53, 54], and the TAp73−/− mice also exhibit a reduction in burrowing and in speed and rearing time in open field tests. Moreover, the degree of impairment in both increased with age [36]. Importantly, these behavioral deficits manifest with accompanying electrophysiological abnormalities.
In contrast, the ΔNp73−/− mice, in particular older animals, show only a slight deficit in the open field test. However, the limited number of observations and a lack of any electrophysiological data do not yet give a full picture of the role of ΔNp73 in the CNS, a line of enquiry worthy of further investigation.
How Does p73 Regulate so Many Aspect of Neural Development?
From in vivo studies, it emerges that either the deletion of both major p73 isoforms or the selective deletion of just the N-terminal isoforms leads to a complex neural phenotype. Initially, this phenotype was ascribed to the pro-survival role of ΔNp73 [48], supported by the observation that cortical neurons from p73−/− mice are more susceptible to glutamate-induced cell death [55]. However, when cortical neurons derived from the three mouse models, p73−/−, TAp73−/−, and ΔNp73−/−, were cultured in vitro, no signs of cell death were observed. Furthermore, no differences were seen in cortical neurons from these lines when challenged with DNA-damaging agents [56, 36]. Therefore, the pro-survival role of ΔNp73 can only partially explain the neural phenotypes of the various p73 knockout models. Further histological analysis of the p73−/− hippocampus shows a disorganized distribution, with cells lacking correct basal-apical orientation. In particular, p73−/− hippocampal neurons have a reduced number of branches and shorter dendrites than do WT cells and impaired morphology is observed in hippocampal neurons in CA3 and DG. This aberrant morphology suggests that p73, in particular the TAp73 isoforms, are playing a role in hippocampal neurogenesis. Indeed, earlier work, albeit only in vitro, had indicated a possible role for p73 in neuronal differentiation [57, 58].
Neurogenesis is a complex multistep process through which nerve cells are generated and integrated into existing neuronal circuits [59, 60]. Under normal conditions, adult neurogenesis takes place in two different regions of the brain, the subventricular zone (SVZ) of the lateral ventricle and the subgranular zone (SGZ) of the DG [60, 61]. Neural stem cells (NSC) of course play a key role in neurogenesis and have the ability to differentiate into different brain cell types (neurons, astrocytes, and oligodendrocytes) while also retaining the capacity to produce identical NSC progeny (self-renewal) [62]. Other p53 family members, p53 itself and ΔNp63, have already been implicated in the regulating NSC behavior [63–65].
p73 and Stemness
Numerous studies have demonstrated that p73 is also a positive regulator of embryonic and adult NSCs [66, 52, 67–69]. Indeed, neurospheres derived from p73−/− mice are smaller and grow more slowly than do WT. This phenotype is due to an impairment of cell proliferation with a reduced number of cells in S-phase and is associated with an increase in the senescent population [52]. Interestingly, Talos et al. found no differences in apoptosis between p73−/− and control NSCs indicating that apoptosis does not play a part in the reduced neurosphere size. The same phenotype has been observed in TAp73−/− NSCs, indicating that it is the TA isoform that is responsible for NSC maintenance [67]. In support of this, TAp73 is the main isoform expressed in embryonic NSCs and the endogenous expression of TAp73 increases during NSC differentiation [66]. As a consequence, both p73−/− and TAp73−/− mice have significantly depleted stem cell compartments in SGZ and SVZ.
The potential downstream candidates responsible for this phenotype are genes involved in the regulation of proliferation and/or self-renewal [70]. The loss of p73 leads to the transcriptional dysregulation of SOX-2, SOX-3, NANOG, NOTCH-1, NOTCH-2, HES-5, JAG2, HEY-2, and DELTEX (Fig. 2a). Although additional studies are required to address how p73 physiologically regulates these factors, so far only Hey-2 has been shown to be a direct transcriptional target of TAp73 [67]. Of possible relevance here, given the HES/HEY family are transcriptional targets of the Notch-1 intracellular domain (N1ICD), we have shown that TAp73, but not ∆Np73, isoforms are able to directly bind the N1ICD and antagonize its transcriptional activity [71]. Of particular note, a CNS-specific conditional Sox-2 knockout mouse line closely phenocopies the p73 and TAp73 null mice [72], manifesting with reduced cortical mass, hydrocephalus, and progressive loss of the lower blade of the DG, while neurospheres generated from this line also show a gradual reduction in stem cell number.
TAp73 play a critical role in regulating NSC biology. a TAp73 regulates the maintenance of both embryonic and adult NSC throughout the expression regulation of several genes involved in the regulation of self-renewal and proliferation. In particular, Hey-2 is a transcriptional target of TAp73. The question mark indicates that it is not known yet whether TAp73 directly or indirectly regulates Sox-2 gene. b TAp73 regulates differentiation of NSC into neurons by regulating the expression of miR-34a and TRIM32. The downstream target of miR-34a is still an open question (indicate with a question mark)
p73 and Neuronal Stem Cell Differentiation
In addition to the above, there is experimental evidence that TAp73 also regulates the differentiation of NSCs. Indeed, it has been shown that neurons derived from p73−/− NSCs do not differentiate fully and exhibit dendritic arborization defects and reduced synaptic connectivity [52]. p73 has been implicated in oligodendrocyte development [58], and oligodendrocytes derived from p73−/− NSCs are fewer in number and of “poorer quality” than those derived from WT cells. The role of TAp73 in the regulation of NSC differentiation has been further confirmed using mouse embryonic stem cells (mESCs) committed to neuronal differentiation. Inhibition of TAp73 expression results in a reduction of the number of differentiated neurons together with reduced neurite connectivity. Although the molecular mechanisms underlying TAp73 function in NSC differentiation are not fully characterized, a possible explanation has recently emerged. Firstly, TAp73 drives the expression of miR-34a in mESCs [73], while miR-34a modulates the appearance of neurons and neurite outgrowth, by a mechanism that involves, at least in part, SIRT-1 (Fig. 2b) [74]. Importantly, miR-34a levels are reduced in the hippocampus of p73−/− mice, suggesting that a TAp73/miR-34a axis exists in vivo. And, in addition, miR-34a knockout mice show a significant reduction of proliferating precursor cells (i.e., Ki-67-positive cells) in the DG SGZ, reminiscent of the p73−/− mouse phenotype. As for TAp73, the molecular mechanisms underlying the role of miR-34a in mESC neuronal differentiation are not yet fully characterized, although the inverse relationship between miR-34a and SIRT-1 is suggestive of SIRT-1 being a miR-34a target [75]. Furthermore, Wnt signalling is involved in the regulation of the self-renewal [76] and Wnt-1 is downregulated during ESC differentiation [77]. Recently, Wnt-1 has also been shown to be a miR-34a target [78]; thus, miR-34a may affect the differentiation of ESCs by acting on Wnt-1 signalling. Notch signalling also plays a central role in neuronal differentiation, and the inhibitory effect of TAp73 on Notch we allude to above may also be at work here. Moreover, the Notch and Wnt signalling pathways are known to interact at several levels and to exert mutually antagonistic effects on each other [79]. Indeed, the TAp73β the isoform we found to be most antagonistic towards Notch has been shown to enhance canonical Wnt signalling [80], indicating that p73 may be another hub through which the Notch and Wnt pathways interact.
Another plausible mechanism by which TAp73 could regulate NSC differentiation is via the ubiquitin/proteasome pathway, specifically through the E3 ubiquitin ligase tripartite motif protein 32 (TRIM32), which has been found necessary for the correct induction of neuronal differentiation of NSCs [81, 82]. TAp73 directly binds the TRIM32 promoter to drive its expression (Fig. 2b); TAp73 and TRIM32 levels increase in parallel during NSC differentiation, and TRIM32 steady-state expression is reduced in p73−/− NSCs and in the SVZ of the p73−/− mouse [83].
p73 and Multipotency
Multipotency is the ability of NSC to differentiate into the three neural lineages, neurons, astrocytes, and oligodendrocytes. Overall, the loss of p73 does not affect the multipoptency of NSCs as dissociated p73−/− NSCs maintain their ability to differentiate along each lineage [52].
Overall, TAp73 is required for the maintenance of NSCs and for the proper differentiation of these cells into neurons and oligodendrocytes. Is TAp73 also required for the commitment in NSCs from the neuroectoderm, a process that takes place in the early phase of CNS development? Recent studies have provided at least a partial answer to this question [84]. Mouse embryonic fibroblasts isolated from p73−/− mice can be normally reprogrammed into induced pluripotent stem cells (iPSC), indicating that p73 deficiency does not affect iPSC generation, self-maintenance, or pluripotency. Moreover, iPSC from p73−/− mice are able to differentiate normally into NSCs, clearly suggesting that p73 is dispensable for NSC formation.
p73 and Terminal Neuronal Differentiation
Once neuronal progenitors exit from the cell cycle, the immature postmitotic neurons engage in a series of developmental processes including migration, axonal and dendritic growth, synapse formation, and integration in the preexisting neuronal circuitry [85, 60]. Extrinsic factors including brain-derived neurotrophic factor (BDNF), neurotrophin 3, and nerve growth factor (NGF) play key roles in axon growth and dendritic morphology in cortical neurons [86]. The p75 neurotrophin receptor (p75NTR) is a member of the tumor necrosis factor receptor family that transduces signals from pro- and mature neurotrophins, including NGF [87–89]. p75NTR has multiple functions within the nervous system, ranging from neurite outgrowth and survival to apoptosis; these multiple functions are a reflection of the variety of ligands as well as the ability of p75NTR to interact with other receptors such as tyrosine kinase receptor B (TrkB) and sortilin [90–93]. p75NTR promotes neurite outgrowth, elongation, and branching via activation of a ceramide, Ras/ERK pathway [94–96]. Furthermore, p75NTR facilitates cell survival through PI3-K-mediated Akt activation and NF-κB, an antiapoptotic signalling factor [97–100].
p75NTR expression is reduced in both the cortex and hippocampus of TAp73−/− mice. Moreover, TAp73−/− cortical neurons show reduction in total neurite length and branch points after NGF treatment, resulting in a reduction in the complexity of the dendritic arbor and a reduction in network connectivity, as confirmed by electrophysiology [36]. At the molecular level, the same authors demonstrated that TAp73 binds the p75NTR promoter and regulates its expression (Fig. 3a).
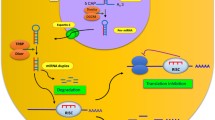
TAp73 regulates neuronal terminal differentiation. TAp73 expression increases during the developmental stage from immature to mature neurons. Loss of TAp73 results in a neuron with less connectivity and electrophysiological defects. This complex phenotype results from the direct regulation of several genes by TAp73. a TAp73 binds p75NTR promoter and regulates its expression, which in turn regulates neurite outgrowth, elongation, and branching for activation of ceramide, Ras/ERK pathway, and cell survival through PI3-K and NF-κB. b miR-34a is a direct target of TAp73. miR-34a inhibits the expression of synaptotagmin-1 (Syt-1) and sintaxin-1A (Stx-1A), resulting in a modulation of synaptogenesis. c Glutaminase-2 regulates the conversion of glutamine into glutamate. Modulation of GLS2 expression affects neurite outgrowth. GLS2 is a transcriptional target of TAp73
Several observations indicate that p75NTR also plays an important role in the peripheral nervous system (PNS) where it exerts a positive effect on myelination [101–103]. TAp73−/− mice show similar PNS defects to those observed in p75NTR knockout mice [36]. In addition, sciatic nerves from TAp73−/− mice contain fewer axons which also have a reduction in the diameter of the myelin sheath, while these mice manifest an associated thermal sensitivity defect.
During development, a number of miRs show distinct expression patterns during maturation of the central nervous system [104, 105]. miR-34a, discussed above, is highly expressed in the brain, and its ectopic expression in neuroblastoma cell lines modulates neuron-specific genes [106], suggesting a possible developmental role of miR-34a in the CNS. The expression of miR-34a increases during postnatal development of the brain and cerebellum, when synaptogenesis takes place, and miR-34a expression is modulated during in vitro differentiation of cortical neurons. TAp73 is an important factor modulating miR-34a expression during neuronal development, during in vitro differentiation of neuroblastoma cells and cortical neurons [56].
An in silico search for targets that might explain the role of miR-34a in this system revealed that several synaptic proteins (synapsin-II, synaptotagmin-I and synaptotagmin-IV, sintaxin-1A, synaptobrevin-2) contained putative miR-34a consensus sequences within their 3′UTRs (Fig. 3b). Of these, synaptotagmin-I and sintaxin-1A were validated as direct targets of the miR-34 family. Ectopic expression of miR-34a in primary cortical neurons reduced neurite arborization, and a reduction in synaptotagmin-1 and sintaxin-1A expression, while the reduced neurite complexity due to miR-34a expression was partially rescued by ectopic expression of synaptotagmin-1. In parallel, inhibition of miR-34a expression with an antagomir resulted in increases in neurite outgrowth length and branch number. These phenotypic changes resulting from modulation of miR-34a expression were also associated with changes in neurite spinal morphology and with electrophysiological abnormalities, which are consistent with miR-34a acting at the level of inhibitory synapses (Fig. 3b).
A recent report has suggested that TAp73 regulates the expression of glutaminase 2 (GLS2), an enzyme that mediates the conversion of glutamine into glutamate during the neuronal differentiation of neuroblastoma cells (Fig. 3c) [107]. Moreover, direct manipulation of GLS2 expression itself modulates neuroblastoma differentiation, and glutamine deprivation influences the differentiation of cortical neurons in vitro, suggesting that the neuronal effects of TAp73 are partly due an effect on metabolism. Although TAp73 is not essential for the in vivo regulation of GLS2 expression, metabolic profiling performed on TAp73- and ΔNp73-deficient cortical neurons suggests that TAp73 loss does affect glutamate metabolism. Indeed, cortical neurons derived from TAp73−/− mice show a reduction in the levels of the neurotransmitters, glutamate, and GABA, without any significant changes in aspartate and N-acetylaspartylglutamate. On the contrary, N-acetylaspartylglutamate and glycine are reduced in ΔNp73−/− neurons, suggesting isoform-specific metabolic functions of p73.
p73 and Alzheimer’s Disease
Alzheimer’s disease (AD) is confirmed in the postmortem brain by an abundant presence of senile plaques (extracellular deposits of the β-amyloid peptide) and neurofibrillary tangles (intraneuronal aggregates of hyperphosphorylated forms of the microtubule-associated protein, tau) in the entorhinal cortex, hippocampal formation, and temporal and frontal cortices. In its initial formulation, the “amyloid cascade hypothesis” of AD neuropathology proposed that an increase in the extracellular deposition of β-amyloid leads, in some yet to be determined way, to an affect on tau leading to tangle formation which in turn resulted in cognitive impairment and neurodegeneration. Almost 25 years on the hypothesis still holds but is coming under considerable criticism largely due to the lack of success of a number of large clinical trials evaluating therapeutics aimed at targeting β-amyloid [108–111].
Some argue that it is now time to reject the amyloid cascade hypothesis outright [112]. While others point out that although considerable evidence demonstrate that amyloid at any stage of aggregation is not alone sufficient to cause AD, it is at the very least necessary for AD to manifest [113]. Indeed, few would argue β-amyloid does not play an important role in the etiology and pathology of the disease, and the general consensus now is that it is not the amyloid aggregates themselves but rather the oligomeric, soluble forms of β-amyloid that are the toxic species, exerting effects on a multitude of cellular processes including inflammation, autophagy, oxidative stress, calcium homeostasis, mitochondrial function, synaptic function, excitotoxicity, and neuronal cell death [114, 115].
In this light, β-amyloid is regarded as a trigger of other downstream events that bring about neurodegeneration [113]. The effectors of those events may well include members of the p53 family. Only a few years after the amyloid cascade hypothesis was first put forward reports began to appear implicating the p53 family in AD [116], and their number has steadily grown over the intervening years. p53 in particular is closely linked with the three familial AD genes, the β-amyloid precursor protein (APP) [117, 118] and presenilin 1 and presenilin 2 (PSEN1 and PSEN2) [119]. The presenilins are necessary components of γ-secretase, the multimeric protein complex responsible of the final proteolytic cleavage of APP resulting in the generation of β-amyloid [119].
p73, the p53 family member preferentially expressed in the brain, has also been implicated in AD, although its role remains a matter of contention. In hippocampal pyramidal neurons of adult human brain, the p73 protein displays a cytoplasmic expression pattern. However, in the brains of AD sufferers, p73 shows a more nuclear localization pattern in these cells. Our own in vitro studies show that TAp73 induces an increase in tau phosphorylation at phosphoepitopes found to be increased in AD brain. Furthermore, brains from aged heterozygous p73+/− mice showed an age-dependent increase in tau phosphorylation levels and the formation of filamentous aggregates resembling neurofibrillary tangles. It was suggested that this effect on tau was predominantly due to a reduction in ∆Np73 isoform expression and was mediated through an effect on JNK [55]. The p73/miR-34a axis has also been implicated in AD, as high levels of TAp73 and miR-34a have been found in brains from both mouse models of AD and AD patients [120–122, 56].
Evidence placing p73 downstream of β-amyloid and upstream of tau appeared when the heterozygous p73 mouse was crossed with two mouse AD models resulting in a much earlier appearance of increased tau phosphorylation and filamentous aggregates and the activation of tau kinases [123]. However, in a later study, no change in tau phosphorylation was observed in the aged p73+/− mice or when the line was crossed with the same mouse AD model (TgCRND8) [124, 125]. The authors of this study also looked, but found no polymorphisms or change in the copy number of TP73 associated with AD [126]. The reasons for these disparities presently remain unclear. However, an issue is that the p73+/− line and the TgCRND8 lines used were not on an identical genetic background, a factor which is known to have strong effects on phenotype.
A potential area yet to be investigated in AD is the metabolic activities of p73, such as its regulation of glutamine metabolism, which could help clarify this controversy. The long history of research into the role of p53 in tumorigenesis has led to important advances in the treatment of cancer: it is hoped by those in the field that an improved understanding of the role of p53/p73 in AD will, similarly, result in therapeutic advances which are so urgently needed for this most devastating of diseases.
Perspectives and Conclusions
The p53 family member, p73, like the founder member is also a tumor suppressor protein. However, the phenotypes of the mice in which all or certain of different isoforms of p73 have been deleted are not principally those of enhanced tumor susceptibility. Rather, abnormalities of the nervous system, particularly the CNS, occur. Of note, the p73 null mouse is normal at birth and the neurological abnormalities begin to manifest around postnatal day 6. Given p73 expression increases during neuronal maturation, these features indicate that the appropriate expression levels of the p73 isoforms are required for proper CNS development and subsequent neuronal function. Although our model does not allow for discrimination between neuronal phenotypes caused by alterations of development or aberrant neuronal biology, it demonstrates that p73 is an important factor involved in the regulation of postnatal neurological function. However, the downstream mechanisms involved appear complex and remain to be fully worked out. It is clear that one molecular mechanism underlying the phenotype observed in the p73 null mouse is linked to its function as a transcription factor. Indeed, TAp73 either directly or indirectly regulates several genes known to be involved in neuronal biology, including SOX-2, Hey-2, TRIM32, and p75NTR. Moreover, the regulation of miR-34a by TAp73 indicates that microRNAs also participate in the multifunctional role of p73 in neurons. In addition, p73 regulates several metabolic enzymes such as GLS2, which plays a central role in the production of the neurotransmitter, GABA.
The loss of p73 isoforms could also have an impact at the level of neuronal circuits. An altered neurotransmitter profile has been documented in our p73 mouse models, which, together with an effect on dendritic arborization, could readily lead to the disruption of neuronal connectivity.
Although the data so far accumulated indicate that the complex in vivo phenotype observed in the genetically modified mice is mainly due to the functions of the TAp73 isoforms in neurons, one cannot exclude a contribution from the glial cell population. Indeed, TAp73 has been shown to be involved in the proper differentiation of oligodendrocyte precursor cells (OPCs). These cells normally dived several times then differentiate. However, the exogenous overexpression of TAp73 in OPCs, in vitro, induces OPCs to spontaneously differentiate into oligodendrocytes, while exogenous ΔNp73 completely inhibits this effect [58]. More recently, Talos et al. using NSC isolated from p73−/− mice confirmed the role of p73 in oligodendrocyte differentiation.
To date, no reports have appeared indicating a role of p73 in astrocytes, and astrocytes generated from p73−/− NSC appear normal [52], suggesting that this p53 family member plays little role in astrocyte biology. Even so, further investigations are needed to fully rule this out.
In the last few years, epidemiological studies and experimental findings have postulated a link between cancer and neurodegenerative disease [127]. Indeed, several genes that are involved in neurodegeneration are often deregulated or mutated in cancer. Among those genes, several are tumor suppressor genes or oncogenes. Therefore, we would like to speculate that p73 could join the club of those genes with overlapping function in both cancer and neurodegenerative disorders.
Abbreviations
- CNS:
-
Central nervous system
- NSC:
-
Neural stem cell
- (MBP):
-
Myelin basic protein
- NGF:
-
Nerve growth factor
- PNS:
-
Peripheral nervous system
- p75NTR :
-
p75 neurotrophin receptor
- WT:
-
Wild type
- TAp73−/−:
-
TAp73 knockout
- p73−/−:
-
p73 knockout mice
- DG:
-
Dentate gyrus
- AD:
-
Alzheimer’s disease
- Aβ:
-
β-amyloid
- NFTs:
-
Neurofibrillary tangles
References
Dotsch V, Bernassola F, Coutandin D, Candi E, Melino G (2010) p63 and p73, the ancestors of p53. Cold Spring Harb Perspect Biol 2(9):a004887. doi:10.1101/cshperspect.a004887
Montero J, Dutta C, van Bodegom D, Weinstock D, Letai A (2013) p53 regulates a non-apoptotic death induced by ROS. Cell Death Differ 20(11):1465–1474. doi:10.1038/cdd.2013.52
Fan YH, Cheng J, Vasudevan SA, Dou J, Zhang H, Patel RH, Ma IT, Rojas Y et al (2013) USP7 inhibitor P22077 inhibits neuroblastoma growth via inducing p53-mediated apoptosis. Cell Death Dis 4:e867. doi:10.1038/cddis.2013.400
Valentino T, Palmieri D, Vitiello M, Pierantoni GM, Fusco A, Fedele M (2013) PATZ1 interacts with p53 and regulates expression of p53-target genes enhancing apoptosis or cell survival based on the cellular context. Cell Death Dis 4:e963. doi:10.1038/cddis.2013.500
Kenzelmann Broz D, Spano Mello S, Bieging KT, Jiang D, Dusek RL, Brady CA, Sidow A, Attardi LD (2013) Global genomic profiling reveals an extensive p53-regulated autophagy program contributing to key p53 responses. Genes Dev 27(9):1016–1031. doi:10.1101/gad.212282.112
Qian Y, Chen X (2013) Senescence regulation by the p53 protein family. Methods Mol Biol 965:37–61. doi:10.1007/978-1-62703-239-1_3
Levine AJ, Tomasini R, McKeon FD, Mak TW, Melino G (2011) The p53 family: guardians of maternal reproduction. Nat Rev Mol Cell Biol 12(4):259–265. doi:10.1038/nrm3086
He Z, Liu H, Agostini M, Yousefi S, Perren A, Tschan MP, Mak TW, Melino G et al (2013) p73 regulates autophagy and hepatocellular lipid metabolism through a transcriptional activation of the ATG5 gene. Cell Death Differ 20(10):1415–1424. doi:10.1038/cdd.2013.104
Alexandrova EM, Petrenko O, Nemajerova A, Romano RA, Sinha S, Moll UM (2013) DeltaNp63 regulates select routes of reprogramming via multiple mechanisms. Cell Death Differ 20(12):1698–1708. doi:10.1038/cdd.2013.122
Molchadsky A, Ezra O, Amendola PG, Krantz D, Kogan-Sakin I, Buganim Y, Rivlin N, Goldfinger N et al (2013) p53 is required for brown adipogenic differentiation and has a protective role against diet-induced obesity. Cell Death Differ 20(5):774–783. doi:10.1038/cdd.2013.9
Candi E, Agostini M, Melino G, Bernassola F (2014) How the TP53 family proteins TP63 and TP73 contribute to tumorigenesis: regulators and effectors. Hum Mutat 35(6):702–714. doi:10.1002/humu.22523
Levrero M, De Laurenzi V, Costanzo A, Gong J, Wang JY, Melino G (2000) The p53/p63/p73 family of transcription factors: overlapping and distinct functions. J Cell Sci 113(Pt 10):1661–1670
Rai TS, Adams PD (2013) Lessons from senescence: chromatin maintenance in non-proliferating cells. Biochim Biophys Acta 1819(3–4):322–331
Wiman KG (2013) p53 talks to PARP: the increasing complexity of p53-induced cell death. Cell Death Differ 20(11):1438–1439. doi:10.1038/cdd.2013.111
Huang Q, Yu L, Levine AJ, Nussinov R, Ma B (2014) Dipeptide analysis of p53 mutations and evolution of p53 family proteins. Biochim Biophys Acta 1844(1 Pt B):198–206. doi:10.1016/j.bbapap.2013.04.002
Mello SS, Attardi LD (2013) Not all p53 gain-of-function mutants are created equal. Cell Death Differ 20(7):855–857. doi:10.1038/cdd.2013.53
Hanel W, Marchenko N, Xu S, Yu SX, Weng W, Moll U (2013) Two hot spot mutant p53 mouse models display differential gain of function in tumorigenesis. Cell Death Differ 20(7):898–909. doi:10.1038/cdd.2013.17
Wang W, Cheng B, Miao L, Mei Y, Wu M (2013) Mutant p53-R273H gains new function in sustained activation of EGFR signaling via suppressing miR-27a expression. Cell Death Dis 4:e574. doi:10.1038/cddis.2013.97
Chen YC, Chan JY, Chiu YL, Liu ST, Lozano G, Wang SL, Ho CL, Huang SM (2013) Grail as a molecular determinant for the functions of the tumor suppressor p53 in tumorigenesis. Cell Death Differ 20(5):732–743. doi:10.1038/cdd.2013.1
Vanbokhoven H, Melino G, Candi E, Declercq W (2011) p63, a story of mice and men. J Investig Dermatol 131(6):1196–1207. doi:10.1038/jid.2011.84
Candi E, Cipollone R, di Val R, Cervo P, Gonfloni S, Melino G, Knight R (2008) p63 in epithelial development. Cell Mol Life Sci CMLS 65(20):3126–3133. doi:10.1007/s00018-008-8119-x
Chari NS, Romano RA, Koster MI, Jaks V, Roop D, Flores ER, Teglund S, Sinha S et al (2013) Interaction between the TP63 and SHH pathways is an important determinant of epidermal homeostasis. Cell Death Differ 20(8):1080–1088. doi:10.1038/cdd.2013.41
Masse I, Barbollat-Boutrand L, Molina M, Berthier-Vergnes O, Joly-Tonetti N, Martin MT, Caron de Fromentel C, Kanitakis J et al (2012) Functional interplay between p63 and p53 controls RUNX1 function in the transition from proliferation to differentiation in human keratinocytes. Cell Death Dis 3:e318. doi:10.1038/cddis.2012.62
Rufini A, Barlattani A, Docimo R, Velletri T, Niklison-Chirou MV, Agostini M, Melino G (2011) p63 in tooth development. Biochem Pharmacol 82(10):1256–1261. doi:10.1016/j.bcp.2011.07.068
Paradis MR, Raj MT, Boughner JC (2013) Jaw growth in the absence of teeth: the developmental morphology of edentulous mandibles using the p63 mouse mutant. Evol Dev 15(4):268–279. doi:10.1111/ede.12026
Rouleau M, Medawar A, Hamon L, Shivtiel S, Wolchinsky Z, Zhou H, De Rosa L, Candi E et al (2011) TAp63 is important for cardiac differentiation of embryonic stem cells and heart development. Stem Cells 29(11):1672–1683. doi:10.1002/stem.723
Paris M, Rouleau M, Puceat M, Aberdam D (2012) Regulation of skin aging and heart development by TAp63. Cell Death Differ 19(2):186–193. doi:10.1038/cdd.2011.181
Barrow LL, van Bokhoven H, Daack-Hirsch S, Andersen T, van Beersum SE, Gorlin R, Murray JC (2002) Analysis of the p63 gene in classical EEC syndrome, related syndromes, and non-syndromic orofacial clefts. J Med Genet 39(8):559–566
Ferone G, Mollo MR, Thomason HA, Antonini D, Zhou H, Ambrosio R, De Rosa L, Salvatore D et al (2013) p63 control of desmosome gene expression and adhesion is compromised in AEC syndrome. Hum Mol Genet 22(3):531–543. doi:10.1093/hmg/dds464
Ferone G, Thomason HA, Antonini D, De Rosa L, Hu B, Gemei M, Zhou H, Ambrosio R et al (2012) Mutant p63 causes defective expansion of ectodermal progenitor cells and impaired FGF signalling in AEC syndrome. EMBO Mol Med 4(3):192–205. doi:10.1002/emmm.201100199
Wilhelm MT, Rufini A, Wetzel MK, Tsuchihara K, Inoue S, Tomasini R, Itie-Youten A, Wakeham A et al (2010) Isoform-specific p73 knockout mice reveal a novel role for delta Np73 in the DNA damage response pathway. Genes Dev 24(6):549–560. doi:10.1101/gad.1873910
Rufini A, Agostini M, Grespi F, Tomasini R, Sayan BS, Niklison-Chirou MV, Conforti F, Velletri T et al (2011) p73 in cancer. Genes Cancer 2(4):491–502. doi:10.1177/1947601911408890
Liu T, Roh SE, Woo JA, Ryu H, Kang DE (2013) Cooperative role of RanBP9 and P73 in mitochondria-mediated apoptosis. Cell Death Dis 4:e476. doi:10.1038/cddis.2012.203
Tomasini R, Tsuchihara K, Wilhelm M, Fujitani M, Rufini A, Cheung CC, Khan F, Itie-Youten A et al (2008) TAp73 knockout shows genomic instability with infertility and tumor suppressor functions. Genes Dev 22(19):2677–2691. doi:10.1101/gad.1695308
Yang A, Walker N, Bronson R, Kaghad M, Oosterwegel M, Bonnin J, Vagner C, Bonnet H et al (2000) p73-deficient mice have neurological, pheromonal and inflammatory defects but lack spontaneous tumours. Nature 404(6773):99–103. doi:10.1038/35003607
Niklison-Chirou MV, Steinert JR, Agostini M, Knight RA, Dinsdale D, Cattaneo A, Mak TW, Melino G (2013) TAp73 knockout mice show morphological and functional nervous system defects associated with loss of p75 neurotrophin receptor. Proc Natl Acad Sci U S A 110(47):18952–18957. doi:10.1073/pnas.1221172110
Killick R, Niklison-Chirou M, Tomasini R, Bano D, Rufini A, Grespi F, Velletri T, Tucci P et al (2011) p73: a multifunctional protein in neurobiology. Mol Neurobiol 43(2):139–146. doi:10.1007/s12035-011-8172-6
Inoue S, Tomasini R, Rufini A, Elia AJ, Agostini M, Amelio I, Cescon D, Dinsdale D et al (2014) TAp73 is required for spermatogenesis and the maintenance of male fertility. Proc Natl Acad Sci U S A 111(5):1843–1848. doi:10.1073/pnas.1323416111
Seeliger MA, Moll UM (2013) p73—constitutively open for business. Cell Death Differ 20(8):972–973. doi:10.1038/cdd.2013.56
Luh LM, Kehrloesser S, Deutsch GB, Gebel J, Coutandin D, Schafer B, Agostini M, Melino G et al (2013) Analysis of the oligomeric state and transactivation potential of TAp73alpha. Cell Death Differ 20(8):1008–1016. doi:10.1038/cdd.2013.23
Flores ER, Lozano G (2012) The p53 family grows old. Genes Dev 26(18):1997–2000. doi:10.1101/gad.202648.112
Fricker M, Papadia S, Hardingham GE, Tolkovsky AM (2010) Implication of TAp73 in the p53-independent pathway of Puma induction and Puma-dependent apoptosis in primary cortical neurons. J Neurochem 114(3):772–783. doi:10.1111/j.1471-4159.2010.06804.x
Buhlmann S, Putzer BM (2008) DNp73 a matter of cancer: mechanisms and clinical implications. Biochim Biophys Acta 1785(2):207–216. doi:10.1016/j.bbcan.2008.01.002
Grespi F, Amelio I, Tucci P, Annicchiarico-Petruzzelli M, Melino G (2012) Tissue-specific expression of p73 C-terminal isoforms in mice. Cell Cycle 11(23):4474–4483. doi:10.4161/cc.22787
Tomasini R, Secq V, Pouyet L, Thakur AK, Wilhelm M, Nigri J, Vasseur S, Berthezene P et al (2013) TAp73 is required for macrophage-mediated innate immunity and the resolution of inflammatory responses. Cell Death Differ 20(2):293–301. doi:10.1038/cdd.2012.123
Marcel V, Petit I, Murray-Zmijewski F, Goullet de Rugy T, Fernandes K, Meuray V, Diot A, Lane DP et al (2012) Diverse p63 and p73 isoforms regulate Delta133p53 expression through modulation of the internal TP53 promoter activity. Cell Death Differ 19(5):816–826. doi:10.1038/cdd.2011.152
Meyer G, Perez-Garcia CG, Abraham H, Caput D (2002) Expression of p73 and reelin in the developing human cortex. J Neurosci Off J Soc Neurosci 22(12):4973–4986
Pozniak CD, Barnabe-Heider F, Rymar VV, Lee AF, Sadikot AF, Miller FD (2002) p73 is required for survival and maintenance of CNS neurons. J Neurosci Off J Soc Neurosci 22(22):9800–9809
Pozniak CD, Radinovic S, Yang A, McKeon F, Kaplan DR, Miller FD (2000) An anti-apoptotic role for the p53 family member, p73, during developmental neuron death. Science 289(5477):304–306
Tissir F, Ravni A, Achouri Y, Riethmacher D, Meyer G, Goffinet AM (2009) DeltaNp73 regulates neuronal survival in vivo. Proc Natl Acad Sci U S A 106(39):16871–16876. doi:10.1073/pnas.0903191106
Thompson CL, Pathak SD, Jeromin A, Ng LL, MacPherson CR, Mortrud MT, Cusick A, Riley ZL et al (2008) Genomic anatomy of the hippocampus. Neuron 60(6):1010–1021. doi:10.1016/j.neuron.2008.12.008
Talos F, Abraham A, Vaseva AV, Holembowski L, Tsirka SE, Scheel A, Bode D, Dobbelstein M et al (2010) p73 is an essential regulator of neural stem cell maintenance in embryonal and adult CNS neurogenesis. Cell Death Differ 17(12):1816–1829. doi:10.1038/cdd.2010.131
Deacon RM (2006) Burrowing in rodents: a sensitive method for detecting behavioral dysfunction. Nat Protoc 1(1):118–121. doi:10.1038/nprot.2006.19
Elder GA, Ragnauth A, Dorr N, Franciosi S, Schmeidler J, Haroutunian V, Buxbaum JD (2008) Increased locomotor activity in mice lacking the low-density lipoprotein receptor. Behav Brain Res 191(2):256–265. doi:10.1016/j.bbr.2008.03.036
Wetzel MK, Naska S, Laliberte CL, Rymar VV, Fujitani M, Biernaskie JA, Cole CJ, Lerch JP et al (2008) p73 regulates neurodegeneration and phospho-tau accumulation during aging and Alzheimer’s disease. Neuron 59(5):708–721. doi:10.1016/j.neuron.2008.07.021
Agostini M, Tucci P, Killick R, Candi E, Sayan BS, di Val R, Cervo P, Nicotera P et al (2011) Neuronal differentiation by TAp73 is mediated by microRNA-34a regulation of synaptic protein targets. Proc Natl Acad Sci U S A 108(52):21093–21098. doi:10.1073/pnas.1112061109
De Laurenzi V, Raschella G, Barcaroli D, Annicchiarico-Petruzzelli M, Ranalli M, Catani MV, Tanno B, Costanzo A et al (2000) Induction of neuronal differentiation by p73 in a neuroblastoma cell line. J Biol Chem 275(20):15226–15231
Billon N, Terrinoni A, Jolicoeur C, McCarthy A, Richardson WD, Melino G, Raff M (2004) Roles for p53 and p73 during oligodendrocyte development. Development 131(6):1211–1220. doi:10.1242/dev.01035
Kole AJ, Annis RP, Deshmukh M (2013) Mature neurons: equipped for survival. Cell Death Dis 4:e689. doi:10.1038/cddis.2013.220
Ming GL, Song H (2005) Adult neurogenesis in the mammalian central nervous system. Annu Rev Neurosci 28:223–250. doi:10.1146/annurev.neuro.28.051804.101459
Zhao C, Deng W, Gage FH (2008) Mechanisms and functional implications of adult neurogenesis. Cell 132(4):645–660. doi:10.1016/j.cell.2008.01.033
Yadirgi G, Marino S (2009) Adult neural stem cells and their role in brain pathology. J Pathol 217(2):242–253. doi:10.1002/path.2480
Meletis K, Wirta V, Hede SM, Nister M, Lundeberg J, Frisen J (2006) p53 suppresses the self-renewal of adult neural stem cells. Development 133(2):363–369. doi:10.1242/dev.02208
Dugani CB, Paquin A, Fujitani M, Kaplan DR, Miller FD (2009) p63 antagonizes p53 to promote the survival of embryonic neural precursor cells. J Neurosci Off J Soc Neurosci 29(20):6710–6721. doi:10.1523/JNEUROSCI.5878-08.2009
Hadjal Y, Hadadeh O, Yazidi CE, Barruet E, Binetruy B (2013) A p38MAPK-p53 cascade regulates mesodermal differentiation and neurogenesis of embryonic stem cells. Cell Death Dis 4:e737. doi:10.1038/cddis.2013.246
Agostini M, Tucci P, Chen H, Knight RA, Bano D, Nicotera P, McKeon F, Melino G (2010) p73 regulates maintenance of neural stem cell. Biochem Biophys Res Commun 403(1):13–17. doi:10.1016/j.bbrc.2010.10.087
Fujitani M, Cancino GI, Dugani CB, Weaver IC, Gauthier-Fisher A, Paquin A, Mak TW, Wojtowicz MJ et al (2010) TAp73 acts via the bHLH Hey2 to promote long-term maintenance of neural precursors. Curr Biol CB 20(22):2058–2065. doi:10.1016/j.cub.2010.10.029
Gonzalez-Cano L, Herreros-Villanueva M, Fernandez-Alonso R, Ayuso-Sacido A, Meyer G, Garcia-Verdugo JM, Silva A, Marques MM et al (2010) p73 deficiency results in impaired self renewal and premature neuronal differentiation of mouse neural progenitors independently of p53. Cell Death Dis 1:e109. doi:10.1038/cddis.2010.87
Fatt MP, Cancino GI, Miller FD, Kaplan DR (2014) p63 and p73 coordinate p53 function to determine the balance between survival, cell death, and senescence in adult neural precursor cells. Cell Death Differ 21(10):1546–1559. doi:10.1038/cdd.2014.61
Molofsky AV, Pardal R, Morrison SJ (2004) Diverse mechanisms regulate stem cell self-renewal. Curr Opin Cell Biol 16(6):700–707. doi:10.1016/j.ceb.2004.09.004
Hooper C, Tavassoli M, Chapple JP, Uwanogho D, Goodyear R, Melino G, Lovestone S, Killick R (2006) TAp73 isoforms antagonize Notch signalling in SH-SY5Y neuroblastomas and in primary neurones. J Neurochem 99(3):989–999. doi:10.1111/j.1471-4159.2006.04142.x
Favaro R, Valotta M, Ferri AL, Latorre E, Mariani J, Giachino C, Lancini C, Tosetti V et al (2009) Hippocampal development and neural stem cell maintenance require Sox2-dependent regulation of Shh. Nat Neurosci 12(10):1248–1256. doi:10.1038/nn.2397
Agostini M, Tucci P, Steinert JR, Shalom-Feuerstein R, Rouleau M, Aberdam D, Forsythe ID, Young KW et al (2011) microRNA-34a regulates neurite outgrowth, spinal morphology, and function. Proc Natl Acad Sci U S A 108(52):21099–21104. doi:10.1073/pnas.1112063108
Aranha MM, Santos DM, Sola S, Steer CJ, Rodrigues CM (2011) miR-34a regulates mouse neural stem cell differentiation. PLoS One 6(8):e21396. doi:10.1371/journal.pone.0021396
Tarantino C, Paolella G, Cozzuto L, Minopoli G, Pastore L, Parisi S, Russo T (2010) miRNA 34a, 100, and 137 modulate differentiation of mouse embryonic stem cells. FASEB J Off Publ Fed Am Soc Exp Biol 24(9):3255–3263. doi:10.1096/fj.09-152207
Miyabayashi T, Teo JL, Yamamoto M, McMillan M, Nguyen C, Kahn M (2007) Wnt/beta-catenin/CBP signaling maintains long-term murine embryonic stem cell pluripotency. Proc Natl Acad Sci U S A 104(13):5668–5673. doi:10.1073/pnas.0701331104
Sampath P, Pritchard DK, Pabon L, Reinecke H, Schwartz SM, Morris DR, Murry CE (2008) A hierarchical network controls protein translation during murine embryonic stem cell self-renewal and differentiation. Cell Stem Cell 2(5):448–460. doi:10.1016/j.stem.2008.03.013
Hashimi ST, Fulcher JA, Chang MH, Gov L, Wang S, Lee B (2009) MicroRNA profiling identifies miR-34a and miR-21 and their target genes JAG1 and WNT1 in the coordinate regulation of dendritic cell differentiation. Blood 114(2):404–414. doi:10.1182/blood-2008-09-179150
Collu GM, Hidalgo-Sastre A, Brennan K (2014) Wnt-Notch signalling crosstalk in development and disease. Cell Mol Life Sci CMLS 71(18):3553–3567. doi:10.1007/s00018-014-1644-x
Ueda Y, Hijikata M, Takagi S, Takada R, Takada S, Chiba T, Shimotohno K (2001) p73beta, a variant of p73, enhances Wnt/beta-catenin signaling in Saos-2 cells. Biochem Biophys Res Commun 283(2):327–333. doi:10.1006/bbrc.2001.4788
Hillje AL, Pavlou MA, Beckmann E, Worlitzer MM, Bahnassawy L, Lewejohann L, Palm T, Schwamborn JC (2013) TRIM32-dependent transcription in adult neural progenitor cells regulates neuronal differentiation. Cell Death Dis 4:e976. doi:10.1038/cddis.2013.487
Schwamborn JC, Berezikov E, Knoblich JA (2009) The TRIM-NHL protein TRIM32 activates microRNAs and prevents self-renewal in mouse neural progenitors. Cell 136(5):913–925. doi:10.1016/j.cell.2008.12.024
Gonzalez-Cano L, Hillje AL, Fuertes-Alvarez S, Marques MM, Blanch A, Ian RW, Irwin MS, Schwamborn JC et al (2013) Regulatory feedback loop between TP73 and TRIM32. Cell Death Dis 4:e704. doi:10.1038/cddis.2013.224
Alexandrova EM, Talos F, Moll UM (2013) p73 is dispensable for commitment to neural stem cell fate, but is essential for neural stem cell maintenance and for blocking premature differentiation. Cell Death Differ 20(2):368. doi:10.1038/cdd.2012.134
de la Torre-Ubieta L, Bonni A (2011) Transcriptional regulation of neuronal polarity and morphogenesis in the mammalian brain. Neuron 72(1):22–40. doi:10.1016/j.neuron.2011.09.018
Chao MV, Bothwell M (2002) Neurotrophins: to cleave or not to cleave. Neuron 33(1):9–12
Zampieri N, Chao MV (2004) Structural biology. The p75 NGF receptor exposed. Science 304(5672):833–834. doi:10.1126/science.1098110
Arevalo JC, Chao MV (2005) Axonal growth: where neurotrophins meet Wnts. Curr Opin Cell Biol 17(2):112–115. doi:10.1016/j.ceb.2005.01.004
Tiveron C, Fasulo L, Capsoni S, Malerba F, Marinelli S, Paoletti F, Piccinin S, Scardigli R et al (2013) ProNGF\NGF imbalance triggers learning and memory deficits, neurodegeneration and spontaneous epileptic-like discharges in transgenic mice. Cell Death Differ 20(8):1017–1030. doi:10.1038/cdd.2013.22
Chen Y, Zeng J, Cen L, Chen Y, Wang X, Yao G, Wang W, Qi W et al (2009) Multiple roles of the p75 neurotrophin receptor in the nervous system. J Int Med Res 37(2):281–288
Hempstead BL (2002) The many faces of p75NTR. Curr Opin Neurobiol 12(3):260–267
Brito V, Puigdellivol M, Giralt A, del Toro D, Alberch J, Gines S (2013) Imbalance of p75(NTR)/TrkB protein expression in Huntington’s disease: implication for neuroprotective therapies. Cell Death Dis 4:e595. doi:10.1038/cddis.2013.116
Hu Y, Lee X, Shao Z, Apicco D, Huang G, Gong BJ, Pepinsky RB, Mi S (2013) A DR6/p75(NTR) complex is responsible for beta-amyloid-induced cortical neuron death. Cell Death Dis 4:e579. doi:10.1038/cddis.2013.110
Blochl A, Blumenstein L, Ahmadian MR (2004) Inactivation and activation of Ras by the neurotrophin receptor p75. Eur J Neurosci 20(9):2321–2335. doi:10.1111/j.1460-9568.2004.03692.x
Song MS, Posse de Chaves EI (2003) Inhibition of rat sympathetic neuron apoptosis by ceramide. Role of p75NTR in ceramide generation. Neuropharmacology 45(8):1130–1150
Xue L, Murray JH, Tolkovsky AM (2000) The Ras/phosphatidylinositol 3-kinase and Ras/ERK pathways function as independent survival modules each of which inhibits a distinct apoptotic signaling pathway in sympathetic neurons. J Biol Chem 275(12):8817–8824
Roux PP, Bhakar AL, Kennedy TE, Barker PA (2001) The p75 neurotrophin receptor activates Akt (protein kinase B) through a phosphatidylinositol 3-kinase-dependent pathway. J Biol Chem 276(25):23097–23104. doi:10.1074/jbc.M011520200
Yeiser EC, Rutkoski NJ, Naito A, Inoue J, Carter BD (2004) Neurotrophin signaling through the p75 receptor is deficient in traf6−/− mice. J Neurosci Off J Soc Neurosci 24(46):10521–10529. doi:10.1523/JNEUROSCI.1390-04.2004
Harrington AW, Kim JY, Yoon SO (2002) Activation of Rac GTPase by p75 is necessary for c-jun N-terminal kinase-mediated apoptosis. J Neurosci Off J Soc Neurosci 22(1):156–166
Carter BD, Kaltschmidt C, Kaltschmidt B, Offenhauser N, Bohm-Matthaei R, Baeuerle PA, Barde YA (1996) Selective activation of NF-kappa B by nerve growth factor through the neurotrophin receptor p75. Science 272(5261):542–545
Cosgaya JM, Chan JR, Shooter EM (2002) The neurotrophin receptor p75NTR as a positive modulator of myelination. Science 298(5596):1245–1248. doi:10.1126/science.1076595
Lee KF, Li E, Huber LJ, Landis SC, Sharpe AH, Chao MV, Jaenisch R (1992) Targeted mutation of the gene encoding the low affinity NGF receptor p75 leads to deficits in the peripheral sensory nervous system. Cell 69(5):737–749
Uesugi N, Kimura Y, Yamashita T (2013) Suppression of the p75 receptor signal attenuates the effect of ephrin-B3 and promotes axonal regeneration of the injured optic nerve. Cell Death Dis 4:e557. doi:10.1038/cddis.2013.83
Kosik KS (2006) The neuronal microRNA system. Nat Rev Neurosci 7(12):911–920. doi:10.1038/nrn2037
Volvert ML, Rogister F, Moonen G, Malgrange B, Nguyen L (2012) MicroRNAs tune cerebral cortical neurogenesis. Cell Death Differ 19(10):1573–1581. doi:10.1038/cdd.2012.96
Wei JS, Song YK, Durinck S, Chen QR, Cheuk AT, Tsang P, Zhang Q, Thiele CJ et al (2008) The MYCN oncogene is a direct target of miR-34a. Oncogene 27(39):5204–5213. doi:10.1038/onc.2008.154
Velletri T, Romeo F, Tucci P, Peschiaroli A, Annicchiarico-Petruzzelli M, Niklison-Chirou MV, Amelio I, Knight RA, Mak TW, Melino G, Agostini M (2013) GLS2 is transcriptionally regulated by p73 and contributes to neuronal differentiation. Cell Cycle 12(22)
De Strooper B (2010) Proteases and proteolysis in Alzheimer disease: a multifactorial view on the disease process. Physiol Rev 90(2):465–494. doi:10.1152/physrev.00023.2009
Palavicini JP, Wang H, Bianchi E, Xu S, Rao JS, Kang DE, Lakshmana MK (2013) RanBP9 aggravates synaptic damage in the mouse brain and is inversely correlated to spinophilin levels in Alzheimer’s brain synaptosomes. Cell Death Dis 4:e667. doi:10.1038/cddis.2013.183
Grolla AA, Sim JA, Lim D, Rodriguez JJ, Genazzani AA, Verkhratsky A (2013) Amyloid-beta and Alzheimer’s disease type pathology differentially affects the calcium signalling toolkit in astrocytes from different brain regions. Cell Death Dis 4:e623. doi:10.1038/cddis.2013.145
Ittner LM, Gotz J (2011) Amyloid-beta and tau—a toxic pas de deux in Alzheimer’s disease. Nat Rev Neurosci 12(2):65–72. doi:10.1038/nrn2967
Herrup K (2015) The case for rejecting the amyloid cascade hypothesis. Nat Neurosci 18(6):794–799. doi:10.1038/nn.4017
Musiek ES, Holtzman DM (2015) Three dimensions of the amyloid hypothesis: time, space and ‘wingmen’. Nat Neurosci 18(6):800–806. doi:10.1038/nn.4018
Crews L, Masliah E (2010) Molecular mechanisms of neurodegeneration in Alzheimer’s disease. Hum Mol Genet 19(R1):R12–R20. doi:10.1093/hmg/ddq160
Carrillo-Mora P, Luna R, Colin-Barenque L (2014) Amyloid beta: multiple mechanisms of toxicity and only some protective effects? Oxidative Med Cell Longev 2014:795375. doi:10.1155/2014/795375
Kitamura Y, Shimohama S, Kamoshima W, Matsuoka Y, Nomura Y, Taniguchi T (1997) Changes of p53 in the brains of patients with Alzheimer’s disease. Biochem Biophys Res Commun 232(2):418–421. doi:10.1006/bbrc.1997.6301
Xu X, Yang D, Wyss-Coray T, Yan J, Gan L, Sun Y, Mucke L (1999) Wild-type but not Alzheimer-mutant amyloid precursor protein confers resistance against p53-mediated apoptosis. Proc Natl Acad Sci U S A 96(13):7547–7552
Cuesta A, Zambrano A, Royo M, Pascual A (2009) The tumour suppressor p53 regulates the expression of amyloid precursor protein (APP). Biochem J 418(3):643–650. doi:10.1042/BJ20081793
Checler F, Dunys J, Pardossi-Piquard R, Alves da Costa C (2010) p53 is regulated by and regulates members of the gamma-secretase complex. Neurodegener Dis 7(1–3):50–55. doi:10.1159/000283483
Kiko T, Nakagawa K, Tsuduki T, Furukawa K, Arai H, Miyazawa T (2014) MicroRNAs in plasma and cerebrospinal fluid as potential markers for Alzheimer’s disease. J Alzheimer’s Dis JAD 39(2):253–259. doi:10.3233/JAD-130932
Dickson JR, Kruse C, Montagna DR, Finsen B, Wolfe MS (2013) Alternative polyadenylation and miR-34 family members regulate tau expression. J Neurochem 127(6):739–749. doi:10.1111/jnc.12437
Wang X, Liu P, Zhu H, Xu Y, Ma C, Dai X, Huang L, Liu Y et al (2009) miR-34a, a microRNA up-regulated in a double transgenic mouse model of Alzheimer’s disease, inhibits bcl2 translation. Brain Res Bull 80(4–5):268–273. doi:10.1016/j.brainresbull.2009.08.006
Wilson C, Henry S, Smith MA, Bowser R (2004) The p53 homologue p73 accumulates in the nucleus and localizes to neurites and neurofibrillary tangles in Alzheimer disease brain. Neuropathol Appl Neurobiol 30(1):19–29
Vardarajan B, Vergote D, Tissir F, Logue M, Yang J, Daude N, Ando K, Rogaeva E et al (2013) Role of p73 in Alzheimer disease: lack of association in mouse models or in human cohorts. Mol Neurodegener 8:10. doi:10.1186/1750-1326-8-10
Cancino GI, Miller FD, Kaplan DR (2013) p73 haploinsufficiency causes tau hyperphosphorylation and tau kinase dysregulation in mouse models of aging and Alzheimer’s disease. Neurobiol Aging 34(2):387–399. doi:10.1016/j.neurobiolaging.2012.04.010
Hooper C, Killick R, Tavassoli M, Melino G, Lovestone S (2006) TAp73alpha induces tau phosphorylation in HEK293a cells via a transcription-dependent mechanism. Neurosci Lett 401(1–2):30–34. doi:10.1016/j.neulet.2006.02.082
Plun-Favreau H, Lewis PA, Hardy J, Martins LM, Wood NW (2010) Cancer and neurodegeneration: between the devil and the deep blue sea. PLoS Genet 6(12):e1001257. doi:10.1371/journal.pgen.1001257
Acknowledgments
We thank Prof Eleonora Candi and Dr Ivano Amelio for scientific discussion. This work has been supported by the Medical Research Council, UK.
Conflict of Interest
The authors declare that they have no competing interests.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Niklison-Chirou, M.V., Killick, R., Knight, R.A. et al. How Does p73 Cause Neuronal Defects?. Mol Neurobiol 53, 4509–4520 (2016). https://doi.org/10.1007/s12035-015-9381-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-015-9381-1