Abstract
Dengue fever is a mosquito-borne disease that claims the lives of millions of people around the world. A number of factors like disease’s non-specific symptoms, increased viral mutation, growing antiviral drug resistance due to reduced susceptibility, unavailability of an effective vaccine for dengue, weak immunity against the virus, and many more are involved. Dengue belongs to the Flaviviridae family of viruses. The two species of the vector transmitting dengue are Aedes aegypti and Aedes albopictus, with the former one being dominant. Serotypes 2 of dengue fever are spread to the human body and cause severe illness. Recently, dengue has imposed an aggressive effect synergistically with the COVID-19 pandemic. As a result, we concentrated our efforts on finding a potential therapeutic. For this, we chose natural compounds to fight dengue fever, which is currently regarded as successful among many drug therapies. Following this, we started the in silico experiment with 922 plant extracts as lead compounds to fight serotype 2. In this study, we used SwissADME for analyzing ligand drug-likeness, pkCSM for designing an ADMET profile, Autodock vina 4.2 and Swissdock tools for molecular docking, and finally Desmond for molecular dynamics simulation. Ultimately 45 were found effective against the 2'O methyltransferase protein of serotype 2. CHEMBL376820 was found as possible therapeutic candidates for inhibiting methyltransferase protein in this thorough analysis. Nevertheless, more in vitro and in vivo research are required to substantiate their potential therapeutic efficacy.
Similar content being viewed by others

Avoid common mistakes on your manuscript.
Introduction
The outbreak of dengue fever has spread all over the world, affecting millions of people. It is a mosquito-borne illness spread by infected Aedes aegypti and Aedes albopictus mosquitos, with the latter contributing secondarily to Aedes aegypti mosquitos [1]. The primary and more prevalent symptom of the dengue virus injection is fever. When the infection remains unaddressed or overlooked, it can develop from silent or benign to devastating dengue shock syndrome [2]. According to the World Health Organization (WHO), dengue fever has become prevalent throughout America and South-East Asia, and Western Pacific regions are the most seriously with Asia representing ~ 70% of the global burden of disease [3], including India [4], Nepal [5], China [6], Taiwan [7], and others. Each year, dengue fever causes between 100 and 400 million new illnesses. The amount of new cases and deaths from dengue fever has been steadily growing. According to World Health Organization (WHO), the global incidence of dengue has increased by 30-fold in the past 50 years and estimated some 50 to 100 million new infections occurred annually, with approximately 20,000 deaths [8]. However, there is no particular treatment for this disease. It is almost uncontrollable due to a lack of safe and effective vaccine and appropriate antiviral therapies. Although few ways have been tried to lessen it, they are not working sufficiently. Hence, effective therapies need to be discovered to prevent worldwide dengue severity.
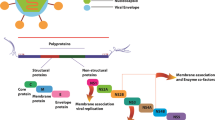
There are five different serotypes transmitted to humans known as dengue virus type 1 to 5 or DENV 1 to 5 [9, 10]. Among them, Serotype 2 of the Dengue Virus is one of four viral serotypes that are antigenically unique but closely connected. It can be divided into phylogenetic clusters or genotypes with distinct epidemiologic correlations that provide a potential source to cause infection. This genotype emerged independently of viral strains that cause human illness epidemics [11]. It is responsible for rapid disease transmission throughout Asia, Africa, and America [12]. In DENV2, the cap shape is critical for mRNA splicing, intracellular RNA transport, RNA stability and degradation, and translation initiation factor 4E (eIF4E) identification for effective translation [13]. Both (guanine-N7)-methyltransferase and nucleoside-2′-O methyltransferase activities are necessary for successive methylation of the cap structure located at the 5′ end of the RNA genome by the Flavivirus NS5 protein [14]. For the development of type 1 (m7GpppNm) caps structure and internal RNA methylation in serotype 2, 2′-O methyltransferases (MTases) proteins are necessary. The 2′-O methyltransferase is essential for the creation of a particular framework at the 5′ end of the RNA template, which safeguard the pathogen RNA against the human innate immune system. This structure is known as RNA cap, mimics the host cell’s original mRNA and facilitate RNA expression [15]. Furthermore, the 2′-O methyl part in Cap-1 is necessary to shield pathogen RNA against interferon-driven reaction [16]. Therefore, for the survival of the dengue virus, 2′-O methyltransferases play a crucial role which can be targeted as a possible option for drug development [17].
Despite the severity of serotype 2 of the dengue virus, there is no specific therapy to cure yet [18]. A recent study suggested that enzymes involved in viral capping could be used as antiviral targets [19]. Recently, an antimalarial drug, quinine has shown modest antiviral effects when administered to dengue fever patients. When treated with quinine, the length of the infection, and the severity of fever, were not substantially different but treatment showed a significant reduction in pain [20]. Therefore, as it did not alter the duration of the disease or the intensity and days of fever, further studies are crucial to ensure the clinical effects of quinine in dengue. Vaccine development was another method of preventing dengue fever. Sanofi Pasteur introduced to science a vaccine called Dengvaxia® (CYD-TDV) [21]. The establishment of a dengue vaccine has been hindered by the distinct and intricate immunopathology of the dengue virus. The development of dengue vaccines has also been hampered by crucial concerns, such as a shortage of animal models for the disease, a lack of appropriate protective immunity markers, and the presence of four distinct dengue serogroups [22]. As there is no particular targeted therapy for the dengue virus, our research is critical for finding a new drug candidate. Natural compounds with antiviral properties might be one alternative for drug development to combat the dengue virus.
Plants produce natural compounds that are small molecules and can be extracted in trace amounts. They include the plant’s primary and secondary metabolites. These isolated sections may contain unique and complex elements with a wide variety of pharmacological actions that can be utilized in the search for new drugs. Natural products offering chemical diversity and bio-reactivity are readily available and accessible and have a significantly high level of public acceptance due to their long history of use [23]. These phytochemicals derived from plants are far less harmful and safer than synthesized chemical substances [24]. Hence, there is a possibility that different phytochemicals isolated from a natural source may show protection against dengue virus infection [25]. Therefore, we examined the efficacy of the natural compounds as a drug candidate against the 2′-O methyltransferase (1R6A) protein of dengue virus serotype 2.
In this study, we retrieved 922 distinct plant-derived compounds from ChEMBL Database and examined their drug-like activities to select potent drug candidates against serotype 2 of the dengue virus (DENV-2) 2'O methyltransferase (1R6A) protein. Initially, these compounds were chosen for drug-likeness, pharmacokinetics, and toxicity analyses employing SwissADME and pkCSM tools, respectively. To assess the best small molecules with greater binding affinity, 45 compounds were chosen for molecular docking purposes utilizing Swissdock and Autodock vina tools. This evaluation showed CHEMBL376820 as a potential candidate for targeting the 2'O methyltransferase (1R6A) receptor. Finally, Molecular Dynamics Simulation was done to evaluate the stability of the lead drug-like substances complexed with the respective binding receptors.
Materials and Methods
Assessment of Drug-Likeness of the Ligands (SwissADME Analysis)
ADME properties stand for absorption, distribution, metabolism, and excretion. ADME features were analyzed to evaluate the physicochemical properties of drug-like compounds. The SwissADME server (http://www.swissadme.ch/index.php) was utilized in this study to determine the major physicochemical, pharmacokinetics, drug-like, and associated properties of the 922 small molecules. To predict the properties, this tool exploited Lipinski’s five-criterion rule [26], Ghose’s rule [27], Veber’s rule [28], Muegge’s rule [29], TPSA, and the number of rotatable bonds. SMILE format of the ligands was obtained from the ChEMBL database (https://www.ebi.ac.uk/chembl/) as input structures for the SwissADME server.
pkCSM: Predicting Small-Molecule Pharmacokinetic and Toxicity Prediction
pkCSM is a program that measures ADMET (absorption, distribution, metabolism, excretion, and toxicity) properties of drug-like molecules. ADMET characteristics that are important for drug development. The pkCSM (http://structure.bioc.cam.ac.uk/pkcsm) server [30] was used to explore an ADMET profile for 220 ligands. Using the canonical smiles of the ligands retrieved from the ChEMBL database, pharmacokinetic, and toxicity properties were calculated.
Molecular Docking
Protein Preparation
The three-dimensional structure of the receptor 2'O methyltransferase in complex with S-adenosyl homocysteine and ribavirin 5' triphosphate (1R6A) was obtained using the RCSB Protein Data Bank (https://www.rcsb.org/search). The PDB protein was prepared for molecular docking study by removing the water molecules from the structure using the PyMOL [31]. Then, they were refined and optimized using Swiss-PdbViewer that utilizes GROMOS 43B1 force field [29].
Ligand Preparation
For molecular docking analysis, we utilized the ChEMBL database (https://www.ebi.ac.uk/chembl/) to fetch the 3D-SDF structures of the drug-like molecules. Further, we energy minimized those ligands using Avogadro, employing MMFF94 force field [32]. Using Avogadro software, 3D-SDF structures were converted to Mol2 format.
Molecular Docking
SwissDock
SwissDock server (http://www.swissdock.ch/docking) [33] allows simulating protein–ligand docking with a simple and user-friendly interface. The structures of the target protein and ligands were prepared before docking. Similarly, a total of 45 ligands were reutilized for molecular docking on the SwissDock website.
Autodock vina
The binding conformations and affinity of all the ligands with the viral proteins were evaluated using molecular docking, a crucial aspect of rational drug development. A total of 45 ligands were docked with 2'O methyltransferase (1R6A) using Autodock vina 4.2 [34, 35]. Using the Lamarckian Genetic Algorithm, this program calculates the binding energy between phytochemicals and target protein. Rigid docking between proteins and chosen molecules were conducted in Autodock vina, with a target region of 27,000 m3 and an exhaustiveness of 10. Receptors were maintained rigid, whereas ligands were remained flexible [36]. Utilizing Lamarckian Genetic model, Autodock vina estimate the interacting energy of target complexes and adjusts their binding poses [37]. After the protein and ligands were created, the Autodock tool was used to convert PDB structures into vina-compliant PDBQT format. Grid boxes with the desired dimensions were created using the Grid box preparation module. The position of the grid box for 2'O methyltransferase (PDB ID: 1R6A) was set at center_x = 11.44, center_y = − 46.682, and Center_z = 3.4 and size_x = 76, size_y = 98, size_z = 58, and exhaustion = 10 [12]. The CASTp server was used to find ligand-binding sites on the viral protein [38].
Molecular Dynamics (MD) Simulation
The dynamic properties and conformational flexibility or stability of docked CHEMBL376820 into the dengue virus protein 2'O methyltransferase (PDB ID: 1R6A) were investigated using MD simulation. The Desmond module of the Schrödinger 2021-1 suite installed on Ubuntu 18.04 (HP Z2 G2 TOWER workstation [with an NVIDIA Quadro 6000 4 GB graphics processing unit (GPU)] was used to conduct MD studies [39]. The Autodock vina-generated ligand–protein (CHEMBL376820-2'O methyltransferase and quinine-2'O methyltransferase) complex was imported into the Schrodinger’s Maestro interface. The ligand–protein complex structure was refined with the help of the “protein preparation wizard,” which assigned bond order, adjusted formal charges, and corrected side and backbone chains [40]. The CHEMBL376820-2'O methyltransferase complex system was solvated in an orthorhombic box (x = 9.721 Å, y = 9.821 Å, z = 6.810 Å) with at least 10 Å distance from the box’s edge with SPC water molecules (simple point-charge). Counter ions (11 Na+ and 25 Cl− ions) were incorporated to neutralize charges, followed by 0.15 M NaCl salt concentrations to mimic human physiological conditions. The energy of the entire system was then minimized using the OPLS3e force field with 2000 iterations and a convergence criterion of 1 kcal/mol [41, 42]. The MD simulation was run for 100 ns at an NPT (isothermal–isobaric) ensemble, and temperature and pressure were kept constant at 300 K and 1.01325 bar, respectively [43, 44]. Every 100 ps, the structure’s coordinates where saved in order to generate trajectories of 1000 frames. Following the completion of the production run, MD simulation trajectories were used for a variety of dynamics analyses, including root mean square deviation (RMSD), root mean square fluctuation (RMSF), and protein contact mapping using the Schrödinger “simulation interaction diagram (SID) panel.”
Results
Drug-Likeness
The drug-likeness analysis explores a compound’s structural or physicochemical characteristics to estimate its likelihood of becoming the best drug candidate. The SwissADME uses five alternative rules (Lipinski, Ghose, Veber, Egan, and Muegge) and utilizes different algorithms to identify phytochemicals with desired characteristics. Molecular weight (optimal range: < 500), amount of hydrogen bond donors (optimal range: ≤ 5), amount of hydrogen bond acceptors (optimal range: ≤ 10), lipophilicity (expressed as LogP, Standard range: < 5), and molar refractivity (optimal range: 40–130) were all refined initially using Lipinski’s rule of five filter standards. Only ligands that fell within these parameters were selected. They were re-examined using the Ghose, Veber, Egan, and Muegge rules. The number of bioavailability scores, rotatable bonds, topological polar surface area (Standard range: 20–130 Å), and log S parameters comprises these rules. Finally, 220 compounds were chosen for further investigation. In Supplementary Table 1, the drug-likeness properties of the selected ligands were reported.
ADMET (Absorption, Distribution, Metabolism, Excretion, Toxicity) Analysis
pkCSM is a valuable tool for predicting the ADMET properties of different ligands. The site utilizes smile format as the input data for evaluating drug-like molecules. Supplementary Table 2 contains the ADMET test results, including the probability score. Almost all ligands have a high intestine absorption rate, oral bioavailability, and positive CaCo-2 permeability in the absorption section. However, not all of them are permeable to the blood–brain barrier in terms of distribution (BBB). Besides, none of the ligands block any CYP450 enzymes in terms of distribution and excretion, even though some of them act as substrates for CYP3A4, CYP2C9, and CYP2D6, while others have no function as a substrate for any of the CYP450 enzymes. Finally, the carcinogenicity and Ames mutagenicity tests revealed that all of the tested ligands were negative. They also have no hepatotoxic or toxic effects when taken orally. Following ADMET analysis, a total of 45 drug-like compounds were chosen. In Table 1, ADMET properties of the selected ligands are provided.
Molecular Docking Analysis
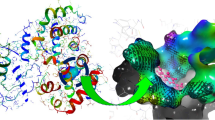
A total of 45 ligands were screened in the Swissdock server and AutoDock vina software, with the most promising molecules for therapeutic development being filtered out. Suitable ligands were chosen based on the bound conformations and the binding affinity of all ligands with the proteins. Because lower binding energy indicates a better affinity, ligands with the lowest binding energy were considered to be the best therapeutic candidates. Binding affinity and the number of hydrogen bonds between ligand and protein were evaluated for identifying the best ligand hit. Based on nonbonded interaction and binding affinity from both docking tool, CHEMBL376820 had the highest binding affinity and estimated ΔG values, accompanied by three hydrogen binding and two hydrophobic interactions. CHEMBL376820 showed better binding affinity on both docking tools. Results of the Swissdock server and AutoDock vina are shown in Table 2. Schematic representation of protein–ligand complexes is displayed in Fig. 1A and B. Complete methodology of this study is shown in Fig. 2.
A Schematic representation of CHEMBL376820-2'O methyltransferase complex; ligands were in magenta color, parts of protein in blue color, and interacting amino acids were shown with green color. B Schematic representation of quinine-2'O methyltransferase complex; ligands were in magenta color, parts of protein in blue color, and interacting amino acids were shown with green color
Molecular Dynamics (MD) Simulation
Molecular docking results have indicated a strong binding affinity of CHEMBL376820 with 2'O methyltransferase protein. Thus, further MD study was conducted to check the stability of CHEMBL376820 in the active site of 2'O methyltransferase protein. The results of molecular dynamics were analyzed for RMSD, RMSF, and protein–ligand contact mapping. The deviation in the structure of a protein or protein–ligand complex from its initial pose is measured as the RMSD. The measurement of RMSD in molecular dynamics simulations gives an estimate of the protein–ligand complex’s stability, dynamic nature, and conformational perturbations that occur in the backbone of the protein during the simulation time scale [45, 46]. Figure 3A represents an analysis of RMSD of CHEMBL376820 and 2'O methyltransferase protein. When lead compound CHEMBL376820 was bound to 2'O methyltransferase protein, Cα atom showed RMSD values between 1.25 and 2.0 Å with an average value of 1.54 Å. Changes in the protein RMSD values when considering the Cα atoms were on the order of 1–3 Å in CHEMBL376820-2'O methyltransferase complex, which is typical for small, globular proteins. The results have shown higher fluctuation of the ligand initially due to equilibration, but after it becomes stable up to 100 ns, the major fluctuation is observed at 83 ns with a 4.44 Å RMSD as shown in Fig. 3A. When the control drug quinine was attached to the 2'O methyltransferase protein, the Cα amino acid had RMSD values ranging from 1 to 2.4, with an average of 1.80. This result has suggested that protein and ligand have robust conformational interactions.
MD simulation analysis of CHEMBL376820 in complex with dengue virus protein 2'O methyltransferase (PDB ID: 1R6A) A RMSD (Protein RMSD is shown in gray, while RMSD of compound CHEMBL376820 are shown in red), B Protein RMSF, C 2d interaction diagram, and D Protein–ligand contact analysis of MD trajectory
The RMSF value represents a protein structure’s mobility and flexibility. The RMSF of protein residues was measured in order to better characterize the protein’s local changes. The β-strand and αhelical domains are emphasized with red and blue backgrounds, respectively. These areas are defined by helices or strands that last for 70% of the simulation. While the loop region is highlighted by a white background [47, 48]. As depicted in Fig. 3B, the fluctuations of the simulated compound had no substantial changes. With the exception of the N terminus and loop regions, the RMSF values of most residues were less than 1.5 Å, indicating that the residue conformation is relatively stable during the simulation. In case of quinine and 2'O methyltransferase construct, it had slightly higher RMSF value ranging from 0.5 to 2.0 Å.
Discussion
A new phase of pharmaceutical research has started since the development of computer-assisted drug design. Due to high toxicity, inadequate antiviral activity, and a significant risk of viral resistance, therapy for the dengue virus is limited [49]. As a result, bioactive compounds with antiviral properties for the dengue virus are needed immediately to combat this life-threatening virus (Fig. 4).
MD simulation analysis of quinine in complex with dengue virus protein 2'O methyltransferase (PDB ID: 1R6A) A RMSD (Protein RMSD is shown in gray, while RMSD of compound CHEMBL376820 are shown in red), B1/B2 Protein RMSF, C 2d interaction diagram, and D Protein–ligand contact analysis of MD trajectory
Nonstructural proteins enable the viral pathogen to bypass the host’s innate immunity and remodel the host–cell underlying structure, in addition to their involvement in RNA replication. Among flavivirus proteins, nonstructural protein 5 (NS5) is by far the most conserved. More specifically, methyltransferase (MTase) activity is in charge of successively covering developing genomic RNA with S-adenosylmethionine by successive methylation on the N7 atoms of the cap guanine and the 2'O atoms of the genome’s ribose of adenine. As a result, the dengue virus can evade the innate immune response of the host [50, 51]. Our work was evolved around finding inhibitors of the methylation process of methyltransferase protein. Recent drug development initiatives to target 2′-O methyltransferase of SARS COV-2 were carried out by Sulimov et al. and Eldehnab et al. [52, 53].
This study investigated a total of 922 ligands, with quinine serving as a control drug [20]. To follow the structure-based drug design method, drug-likeness all of the ligands were evaluated. The concept of drug-likeness, which is based on interpretations of the physicochemical and molecular properties of small drug-like molecules, has been extensively used in drug development to screen out compounds that do not have properties that are compatible with established pharmacokinetic values [54]. The pharmaceutical companies suggested employing five different ruled-based filters successively to all of the analyzed ligands to screen out the optimum oral drug options [55]. The Lipinski filter is built on certain fundamental physicochemical characteristics, and it counts all nitrogen and oxygen atoms as H bond receivers as well as all nitrogen and oxygen with at least one hydrogen as H bond suppliers [56]. Greater molecular mass is associated with a poorer permeability rate in lipid bilayer membranes, and LogP under Five is about 90% likely to be orally soluble [57]. When a drug-like molecule satisfies all five fundamental principles, it shows greater pharmacokinetic properties and bioavailability. To establish a drug-like characteristic, all molecules need to satisfy the Ghose, Veber, and Muegge criteria [35]. Within that study, we selected the ligands that violated none of the five requirements. Before actually choosing the compounds, topological polar surface area (TPSA), bioavailability scores, and the number of rotatable bonds were all considered. 220 ligands were chosen for further investigation after the completion of SwissADME analyses.
The ADMET (Absorption, Distribution, Metabolism, Elimination, and Toxicity) features of a drug-like molecule should be studied before a drug is discovered to guarantee that it can remain in the body, perform its job, and be removed successfully [58]. The uptake of a medicinal ingredient via the intestine cell lining is a key factor in determining a drug’s bioavailability following systemic dosing [59]. CaCo-2 permeability rate and human intestinal absorption (HIA) assessment should be included in the ADMET profile. Our chosen compounds have a strong HIA ratio, and they all permeate the Caco-2 cell lines. Because P-glycoprotein is a crucial transport protein that aids in the transportation of most drugs, it is critical to determine if the substance is a P-glycoprotein substrate or inhibitor [59]. A large percentage of therapeutics that work in the central nervous system (CNS) need BBB permeability [60]. The drug-like molecules should be metabolized by CYP enzymes once it has been absorbed. The main subtypes of mammalian CYP associated with metabolism are CYP1A2, CYP2A6, CYP2C9, CYP2C19, CYP2D6, CYP2E1, and CYP3A4. Its catalytic activity varies for various substances, which is important in adjusting bioavailability and drug reactions. Inside this study, we targeted 5 of such CYPs: CYP1A2, CYP3A4, CYP2C9, CYP2C19, and CYP2D6 [61]. Multiple toxicity tests, such as hepatotoxicity, acute oral toxicity, and Ames mutagenesis, are performed while developing a therapeutic [62]. The Ames mutagenicity test determines if a drug possesses the mutagenic capacity, or the ability to cause gene aberration through genetic damage [63]. Several parameters (AMES toxicity, Max. tolerated dosage (human) hERG I inhibitor, hERG II inhibitor, Oral Rat Acute Toxicity (LD50) Hepatotoxicity) were used to filter out the ligands. Depending on certain ADMET characteristics, 45 drug-like compounds were selected from a pool of 220 compounds. All of the small molecules, had high human intestinal absorption rate. Interestingly, the selected hit CHEMBL376820 functions as a P-glycoprotein substrate. Additionally, the selected 45 ligands were metabolized by CYP3A4 cytochrome P450 isomer, while they did not inhibit CYP2D6 isomer. Furthermore, some of the ligands inhibited CYP1A2, CYP2C19, CYP2C9 isomers. CHEMBL376820 did not inhibit any of the isomers. Results indicated that CHEMBL376820 may have favorable metabolism and excretion activity in human body. In case of toxicity, all the ligands including CHEMBL376820 showed no hepatotoxicity and also were negative in AMES toxicity model. CHEMBL376820 had maximum tolerated dose of − 0.438 mg/kg/day. Moreover, CHEMBL376820 showed no hERG I inhibitor and hERG II inhibitor. The prediction model from pKCSM also suggested that CHEMBL376820 may have favorable pharmacokinetic activity.
Molecular docking has evolved into an essential component of computer-aided drug development [64]. This approach can help with therapeutic development and pharmaceutical science by revealing the understanding of structure-based recognition. We utilized two molecular docking suits (Swissdock and Autodock vina) to maintain the precision and accuracy of our methodology. Additionally, quinine was treated as a control drug during docking analysis. Previously filtered, 45 drug-like molecules were docked with the methyltransferase (1R6A) protein. CHEMBL376820 outperformed both the control drug and filtered ligands. We identified the active site of the 2'O methyltransferase (PDB ID: 1R6A) protein from CASTp webserver. CHEMBL376820 interacted with ILE147, LYS105 amino acids (hydrophobic bonds) and ARG84, VAL130, and GLY81 amino acids (hydrogen bonds) (Fig. 5). These amino acids are situated on the binding groove of the 2'O methyltransferase. So the ligand–protein interaction could diminish methyltransferase activity. The interaction of CHEMBL376820-protein complex along with the control-protein complex was further validated by molecular dynamics simulation.
The conformational stability of potential therapeutics following interactions with 2'O methyltransferase enzyme was evaluated using Molecular Dynamics Simulation. Throughout the 100-ns simulation, the RMSD of the ligands CHEMBL376820 and quinine fluctuated slightly. The ligands CHEMBL376820 did not trigger conformational destabilization inside protein because the observed RMSD shift following complex formation maintained the similar trend as the quinine-protein complex. Our lead therapeutic CHEMBL376820 exhibited tight interaction with their binding area, as revealed by their small fluctuation under 1.5 Å inside 2'O methyltransferase, as per a thorough assessment of the Root Mean Square Variation (RMSF) curve of Cα atoms. However, RMSF variation was noticeable due to loop segments on 2'O methyltransferase. Since loop regions often contain unstructured portions of the protein that fluctuate more than secondary structural components (e.g., alpha helices and beta strands) [65]. Gly86, Arg84, and Ser56 formed H bonds within CHEMBL376820-2'O methyltransferase complex. Alternatively, the amino acid residue Gly148 created hydrogen bonds with the molecule quinine. The protein–ligand binding of CHEMBL376820-2'O methyltransferase maintained relatively stable conformation during the 100-ns simulation, with small backbone shifts in the complex. Our computational data support that CHEMBL376820 could be a viable drug candidate for dengue virus. Previously, Ando et al. reported that CHEMBL376820 isolated from Peperomia duclouxii plant showed anti-cancer activity of 49.8, 5.3, and 13.2 µg/mL against WI-38, VA-13, and HepG2 cell lines [66]. Therefore, CHEMBL376820 could have both antiviral and anti-cancer activities.
Amid certain limitations in terms of the quality of the results obtained as well as contradictions and inconsistencies in the existing knowledge base, computational approaches might indeed rapidly predict a list of potential therapeutic for any microorganism. Therefore, drug repurposing utilizing computational techniques is crucial because it takes less time and resources.
Conclusion
Drug repurposing is a prominent safety strategy for the identification of new bioactive molecules, with the major goal of minimizing the complexity and expenses of experimental drugs in a later clinical trial. Dengue fever caused by dengue serotype 2 has become a worldwide concern, and finding an effective treatment appears to be a big challenge. Based on our analyses, a holistic method including pharmacophore characterization and molecular modeling approaches like molecular docking and molecular dynamics simulations might be used to uncover new therapeutic agents that specifically target the methyltransferase protein of DENV-2. The molecular docking results revealed binding affinity and hydrogen bond interactions of lead compounds CHEMBL376820, whereas the pharmacophore investigations revealed drug-likeness properties. According to molecular dynamics simulations, CHEMBL376820 was constant and equilibrated throughout the simulation at the binding territory of the methyltransferase protein. To sum up, all of the drug-like molecules tested might provide a complete picture for finding a novel antiviral drug for the Dengue Virus 2.
References
Malavige, G., et al. (2004). Dengue viral infections. Postgraduate Medical Journal, 80(948), 588–601.
Hunsperger, E., et al. (2010). Dengue: A continuing global threat. Nature Reviews Microbiology, 8, S7–S16.
WHO Dengue and severe dengue. Available Online: https://bit.ly/3Tl1jZn. Accessed 15 May 2022.
Prakash, O., et al. (2015). Observation on dengue cases from a virus diagnostic laboratory of a tertiary care hospital in North India. The Indian Journal of Medical Research, 142(Suppl 1), S7.
Khadka, S., et al. (2020). Wolbachia: A possible weapon for controlling dengue in Nepal. Tropical Medicine and Health, 48(1), 1–6.
Zhang, H., et al. (2020). Increase in cases of dengue in China, 2004–2016: A retrospective observational study. Travel Medicine and Infectious Disease, 37, 101674.
Lai, W.-T., et al. (2018). Recognizing spatial and temporal clustering patterns of dengue outbreaks in Taiwan. BMC Infectious Diseases, 18(1), 1–11.
WHO. (2012). Global strategy for dengue prevention and control 2012–2020. WHO Library Cataloguing-in-Publication Data, Switzerland.
Mustafa, M., et al. (2015). Discovery of fifth serotype of dengue virus (DENV-5): A new public health dilemma in dengue control. Medical Journal Armed Forces India, 71(1), 67–70.
Karoli, R., et al. (2012). Clinical profile of dengue infection at a teaching hospital in North India. The Journal of Infection in Developing Countries, 6(07), 551–554.
Armstrong, P. M., & Rico-Hesse, R. (2003). Efficiency of dengue serotype 2 virus strains to infect and disseminate in Aedes aegypti. The American Journal of Tropical Medicine and Hygiene, 68(5), 539–544.
Dhar-Chowdhury, P., et al. (2017). Dengue seroprevalence, seroconversion and risk factors in Dhaka, Bangladesh. PLoS Neglected Tropical Diseases, 11(3), e0005475.
Dong, H., et al. (2014). Flavivirus RNA methylation. Journal of General Virology, 95(4), 763–778.
Kroschewski, H., et al. (2008). Mutagenesis of the dengue virus type 2 NS5 methyltransferase domain. Journal of Biological Chemistry, 283(28), 19410–19421.
El Hassab, M. A., et al. (2021). In silico identification of potential SARS COV-2 2’-O-methyltransferase inhibitor: Fragment-based screening approach and MM-PBSA calculations. RSC Advances, 11(26), 16026–16033.
Wilamowski, M., et al. (2021). 2’-O methylation of RNA cap in SARS-CoV-2 captured by serial crystallography. Proceedings of the National academy of Sciences of the United States of America, 118(21), e2100170118.
Züst, R., et al. (2013). Rational design of a live attenuated dengue vaccine: 2′-O-methyltransferase mutants are highly attenuated and immunogenic in mice and macaques. PLoS Pathogens, 9(8), e1003521.
Siddiqui, A. A., et al. (2014). Role of natural products in drug discovery process. The International Journal of Drug Development and Research, 6(2), 172–204.
Ferron, F., et al. (2012). The viral RNA capping machinery as a target for antiviral drugs. Antiviral Research, 96(1), 21–31.
Malakar, S., et al. (2018). Drug repurposing of quinine as antiviral against dengue virus infection. Virus Research, 255, 171–178.
Vannice, K. S., Durbin, A., & Hombach, J. (2016). Status of vaccine research and development of vaccines for dengue. Vaccine, 34(26), 2934–2938.
Ghosh, A., & Dar, L. (2015). Dengue vaccines: Challenges, development, current status and prospects. Indian Journal of Medical Microbiology, 33(1), 3–15.
Beutler, J. A. (2019). Natural products as a foundation for drug discovery. Current Protocols in Pharmacology, 86(1), e67.
Shawan, M. M. A. K., Halder, S. K., & Hasan, M. A. (2021). Luteolin and abyssinone II as potential inhibitors of SARS-CoV-2: An in silico molecular modeling approach in battling the COVID-19 outbreak. Bulletin of the National Research Centre, 45(1), 1–21.
Bhatt, S., et al. (2013). The global distribution and burden of dengue. Nature, 496(7446), 504–507.
Lipinski, C. A., et al. (1997). Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Advanced Drug Delivery Reviews, 23(1), 3–25.
Ghose, A. K., Viswanadhan, V. N., & Wendoloski, J. J. (1999). A knowledge-based approach in designing combinatorial or medicinal chemistry libraries for drug discovery. 1. A qualitative and quantitative characterization of known drug databases. Journal of Combinatorial Chemistry, 1(1), 55–68.
Veber, D. F., et al. (2002). Molecular properties that influence the oral bioavailability of drug candidates. Journal of Medicinal Chemistry, 45(12), 2615–2623.
Guex, N., & Peitsch, M. C. (1997). SWISS-MODEL and the Swiss-Pdb viewer: An environment for comparative protein modeling. Electrophoresis, 18(15), 2714–2723.
Pires, D. E. V., Blundell, T. L., & Ascher, D. B. (2015). pkCSM: Predicting small-molecule pharmacokinetic and toxicity properties using graph-based signatures. Journal of Medicinal Chemistry, 58(9), 4066–4072.
DeLano, W. L. (2002). The PyMOL molecular graphics system, version 1.8. Schrödinger, LLC.
Hanwell, M. D., et al. (2012). Avogadro: An advanced semantic chemical editor, visualization, and analysis platform. Journal of Cheminformatics, 4(1), 17.
Grosdidier, A., Zoete, V., & Michielin, O. (2011). SwissDock, a protein-small molecule docking web service based on EADock DSS. Nucleic Acids Research, 39, W270–W277.
Morris, G. M., et al. (2009). AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. Journal of Computational Chemistry, 30(16), 2785–2791.
Halder, S. K., & Elma, F. (2021). In silico identification of novel chemical compounds with antituberculosis activity for the inhibition of InhA and EthR proteins from Mycobacterium tuberculosis. Journal of Clinical Tuberculosis and Other Mycobacterial Diseases, 24, 100246.
Traversi, G., et al. (2016). Resveratrol and its methoxy-derivatives as modulators of DNA damage induced by ionising radiation. Mutagenesis, 31(4), 433–441.
Morris, G. M., et al. (1998). Automated docking using a Lamarckian genetic algorithm and an empirical binding free energy function. Journal of Computational Chemistry, 19(14), 1639–1662.
Shawan, M. M. A. K., Halder, S. K., & Hasan, M. A. (2021). Luteolin and abyssinone II as potential inhibitors of SARS-CoV-2: An in silico molecular modeling approach in battling the COVID-19 outbreak. Bulletin of the National Research Centre, 45(1), 27–27.
Maestro-Release, D. (2017). Desmond molecular dynamics system, maestro-desmond interoperability tools. DE Shaw Research.
Patel, H., et al. (2022). Investigating the impact of different acrylamide (electrophilic warhead) on osimertinib’s pharmacological spectrum by molecular mechanic and quantum mechanic approach. Combinatorial Chemistry & High Throughput Screening, 25(1), 149–166.
Jorgensen, W. L., Maxwell, D. S., & Tirado-Rives, J. (1996). Development and testing of the OPLS all-atom force field on conformational energetics and properties of organic liquids. Journal of the American Chemical Society, 118(45), 11225–11236.
Ahmad, I., et al. (2021). Synthesis, molecular modelling study of the methaqualone analogues as anti-convulsant agent with improved cognition activity and minimized neurotoxicity. Journal of Molecular Structure, 1251, 131972.
Kalibaeva, G., Ferrario, M., & Ciccotti, G. (2003). Constant pressure-constant temperature molecular dynamics: A correct constrained NPT ensemble using the molecular virial. Molecular Physics, 101(6), 765–778.
Martyna, G. J. (1994). Remarks on ‘“Constant-temperature molecular dynamics with momentum conservation.”’ Physical Review E, 50(4), 3234.
Pawara, R., et al. (2021). Computational identification of 2, 4-disubstituted amino-pyrimidines as L858R/T790M-EGFR double mutant inhibitors using pharmacophore mapping, molecular docking, binding free energy calculation, DFT study and molecular dynamic simulation. In Silico Pharmacology, 9(1), 1–22.
Ghosh, S., et al. (2021). In silico validation of anti-viral drugs obtained from marine sources as a potential target against SARS-CoV-2 Mpro. Journal of the Indian Chemical Society, 98(12), 100272.
Zrieq, R., et al. (2021). Tomatidine and Patchouli alcohol as inhibitors of SARS-CoV-2 enzymes (3CLpro, PLpro and NSP15) by molecular docking and molecular dynamics simulations. International Journal of Molecular Sciences, 22(19), 10693.
Ahmad, I., Kumar, D., & Patel, H. (2022). Computational investigation of phytochemicals from Withania somnifera (Indian ginseng/ashwagandha) as plausible inhibitors of GluN2B-containing NMDA receptors. Journal of Biomolecular Structure and Dynamics, 40(17), 7991–8003.
Troost, B., & Smit, J. M. (2020). Recent advances in antiviral drug development towards dengue virus. Current Opinion in Virology, 43, 9–21.
El Sahili, A., & Lescar, J. (2017). Dengue virus non-structural protein 5. Viruses, 9(4), 91.
Lim, S. P., Noble, C. G., & Shi, P. Y. (2015). The dengue virus NS5 protein as a target for drug discovery. Antiviral Research, 119, 57–67.
Sulimov, A., et al. (2022). Novel inhibitors of 2’-O-methyltransferase of the SARS-CoV-2 coronavirus. Molecules, 27(9), 2721.
El Hassab, M. A., et al. (2021). In silico identification of novel SARS-COV-2 2’-O-methyltransferase (nsp16) inhibitors: Structure-based virtual screening, molecular dynamics simulation and MM-PBSA approaches. Journal of Enzyme Inhibition and Medicinal Chemistry, 36(1), 727–736.
Schneider, G. (2013). Prediction of drug-like properties, in Madame Curie Bioscience Database [Internet]. Landes Bioscience.
Sliwoski, G., et al. (2013). Computational methods in drug discovery. Pharmacological Reviews, 66(1), 334–395.
Benet, L. Z., et al. (2016). BDDCS, the rule of 5 and drugability. Advanced Drug Delivery Reviews, 101, 89–98.
Chae, S. Y., Jang, M. K., & Nah, J. W. (2005). Influence of molecular weight on oral absorption of water soluble chitosans. Journal of Controlled Release, 102(2), 383–394.
Lin, J., et al. (2003). The role of absorption, distribution, metabolism, excretion and toxicity in drug discovery. Current Topics in Medicinal Chemistry, 3(10), 1125–1154.
Hua, S. (2019). Physiological and pharmaceutical considerations for rectal drug formulations. Frontiers in Pharmacology, 10, 1196–1196.
Pardridge, W. M. (2012). Drug transport across the blood-brain barrier. Journal of Cerebral Blood Flow and Metabolism: Official Journal of the International Society of Cerebral Blood Flow and Metabolism, 32(11), 1959–1972.
Bibi, Z. (2008). Role of cytochrome P450 in drug interactions. Nutrition & Metabolism, 5, 27–27.
Larrey, D., & Pageaux, G. P. (2005). Drug-induced acute liver failure. European Journal of Gastroenterology and Hepatology, 17(2), 141–143.
Mortelmans, K., & Zeiger, E. (2000). The Ames Salmonella/microsome mutagenicity assay. Mutation Research, 455(1–2), 29–60.
Jakhar, R., et al. (2020). Relevance of molecular docking studies in drug designing. Current Bioinformatics, 15(4), 270–278.
Chaudhari, B., et al. (2022). Optimizing the Sunitinib for cardio-toxicity and thyro-toxicity by scaffold hopping approach. In Silico Pharmacol, 10(1), 10.
Li, N., et al. (2007). Bioactive lignans from Peperomia duclouxii. Journal of Natural Products, 70(4), 544–548.
Funding
The research was carried out on a self-funded basis.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Md Sakib Hossen is the primary correspondent and Md Salman Shakil is the secondary correspondent.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Halder, S.K., Ahmad, I., Shathi, J.F. et al. A Comprehensive Study to Unleash the Putative Inhibitors of Serotype2 of Dengue Virus: Insights from an In Silico Structure-Based Drug Discovery. Mol Biotechnol 66, 612–625 (2024). https://doi.org/10.1007/s12033-022-00582-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-022-00582-1