Abstract
Introduction of more than one gene into crop plants simultaneously or sequentially, called transgene stacking, has been a more effective strategy for conferring higher and durable insect and disease resistance in transgenic plants than single-gene technology. Transgenes can be stacked against one or more pathogens or for traits such as herbicide tolerance or anthocyanin pigmentation. Polygenic agronomic traits can be improved by multiple gene transformation. The most widely engineered stacked traits are insect resistance and herbicide tolerance as these traits may lead to lesser use of pesticides, higher yield, and efficient control of weeds. In this review, we summarize transgene stacking of two or more transgenes into crops for different agronomic traits, potential applications of gene stacking, its limitations and future prospects.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In conventional transformation system, usually a single-target gene is transformed in a plant species [1, 2]. However, for improved and more effective disease resistance, more than one gene need to be pyramided (also called gene stacking) as the pathogens may overcome the single-gene resistance [3]. Gene stacking or gene pyramiding is the integration of multiple genes, each conferring resistance to single or a separate pest following their independent host pathways. Stacking multiple genes in a genotype is one of the promising tools for breeding higher and durable resistance, especially with the resistant genes that originate from the different gene clusters and represent different host resistance (HR) interactions between the resistance (R) genes and their Avr (effectors) proteins [4]. Employing traditional breeding for gene stacking may lead to linkage drag in gene stacking, while genetic engineering is an efficient and effective strategy to integrate multiple resistance genes into the existing variety. The area of genetically altered crops with stacked genes or traits is likely to rise in near future with combination of the new traits to fulfill needs of the consumers and the producers [5]. Pyramiding genes for disease resistance in crops has become possible because of availability of disease-resistant genes [6] with the improved gene manipulation tools [7].
This means that one may stack genes conferring resistance to insect pests, or one may also pyramid genes that may confer resistance to both the insects and weeds. For instance, transgenic cotton, Bollgard II, developed by Monsanto and registered in U.S. in 2002, expressed two Bt genes, Cry1A(c) and Cry2A(b)2 for conferring resistance to lepidopterans on cotton.
A number of efforts have been made to pyramid more than one transgene in plants for enhanced and effective disease resistance against one or more pathogens. Transgenic rice transformed with a polyprotein and two antimicrobial proteins, Dahlia merckii-antimicrobial protein (Dm-AMP1) and Raphanus sativus- antimicrobial protein (Rs-AFP2) exhibited higher protection to Rhizoctonia solani and Magnaporthe oryzae compared to non-transformed controls or the plants engineered with single-gene constructs [8]. Transgenic potatoes containing chitinase C gene (extracted from Streptomyces griseus) were re-transformed with wasabi defensin (WD) gene (extracted from Wasabia japonica). The transgenic plants stacked with the ChiC and the WD genes were found more resistant to Alternaria solani and Fusarium oxysporum than the wild-type and/or the transgenic plants expressing single gene (ChiC or WD) [9] (Table 1).
Transgene Stacking for Insect Resistance
Currently, commercial transgenic crops are usually stacked with insect-resistant and herbicide-tolerant genes as both traits are highly valuable in the production of major crop plants such as corn and cotton [10]. Bt gene-stacking strategy introduces different insecticidal genes into plant, and proved to be an efficient way of delaying insect resistance to Bt toxin [11]. Thus, the Bt genes with their different modes of action were usually stacked together in the newly developed transgenic crops. Multiple insect-resistant genes and herbicide-tolerant genes were stacked in newly developed commercial transgenic crops. For example, a total of eight genes for insect resistance or herbicide tolerance were stacked in the Genuity® SmartStax™ corn released by Monsanto (USA).
Multiple insect-resistant genes stacking in the transgenic Bt crops have been employed to confer resistance to the insects and herbicides. The first transgenic Bt crop (cotton) with stacked genes, Cry1Ac and Cry2Ab2, registered for use in the U.S. in 2002, was Bollgard II. These stacked genes in the transgenic cotton have been very effective against pink bollworms (Pectinophora gossypiella) Steffey et al. [12]. These genes (Cry1Ac and Cry1C), also stacked in transgenic Bt broccoli, had the potential to delay resistance to diamondback moth (Plutella xylostella) more effectively than the transgenic plants with single-Bt gene [11]. Roundup Ready Flex, a GM cotton also developed by Monsanto, contains three transgenes, Cry1A(c), Cry2A(b)2 and EPSPS genes conferring resistance to insects and herbicide, glyphosate. An altered form of EPSPS (5-enolpyruvylshikimate-3-phosphate synthase), an enzyme causing conversion of sugars to amino acids, was isolated from a soil bacterium which was not affected by glyphosate. Transgenic rice, transformed with Cry1Ac and Cry1Ig and altered glyphosate-tolerant EPSPS genes through single T-DNA, exhibited enhanced resistance to striped rice stem borer (Chilo suppressalis) and rice leafroller (Cnaphalocrocis medinalis), and to glyphosate as well [13]. The triple genes construct (Cry1Ac-Cry2Ab-EPSPS) was also expressed in Nicotiana benthamiana. The transgenic plants had higher resistance to armyworm (Spodoptera littoralis) and herbicides compared to non-transformed control [14]. Transgenic corn, with triple genes stacks, containing Cry3B(b)1 for protection against corn root worm (CRW), Cry1A(b) for imparting resistance to stalk-boring insect, and Roundup Ready® trait for herbicide tolerance were also produced (Table 1).
Transformation Strategies for Stacking Transgenes in Plants
Transformation with the Genes of Interest on a Plasmid or Separate Plasmid
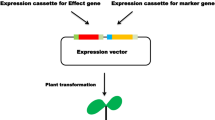
The transgenes with their appropriate promoters and terminators (transgenic cassettes) are placed on a single T-DNA and transformed into plants as a unit to a single locus. Jha and Chattoo [8] transformed rice with Agrobacterium tumefaciens harbouring Dm-AMP1, a linker peptide of the I. balsamina antimicrobial peptides (Ib-AMP) and Rs-AFP2 on a single plasmid, pFAJ3105 (Fig. 1d). The co-expression of the transgenes, Rs-AFP2 and Dm-AMP1 in the transformed plant provided more resistance to fungal pathogens than the singly transformed plants. Transgenic chickpea (Cicer arietinum) co-transformed with the two insecticidal genes, cry1Ab and cry1Ac, exhibited higher resistance to pod borer larvae of Helicoverpa armigera than the single-gene transgenic plants expressing one toxin [15].
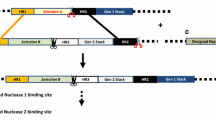
Schematic representation of Tranformation for gene stacking. A-C, Sequential transformation of two disease resistance genes. a and b Marker-free transgenic potato plants were produced by MAT vector system. c The Marker-free transgenic potato plants were re-transformed with wasabi defensin gene [28]. dDm-AMP1 and Rs-AFP2 genes were connected by linker peptide on same plasmid between left and right borders [8]. e Three genes, Rpi-sto1, Rpi-vnt1.1 and Rpi-blb3 were transformed simultaneously in potato [19]. f In GAANTRY system more than three genes could be transformed simultaneously in plants [45]. GUS, beta-glucuronidase. hpt, hygromycin phophotransferase. nptII, neomycin phophotransferase. LP, linker peptide region isolated from the seeds of Impatiens balsamina. Dm-AMP1, Antimicrobial proteins from Dahlia merckii. Rs-AFP2, Antimicrobial proteins from Raphanus sativus. Rpi-sto1, Resistance gene for Phytophthora infestans from Solanum stoloniferum. Rpi-vnt1.1, Rpi from S. venturii. Rpi-blb3, Rpi from S. bulbocastanum. TBS transformation booster sequence, MYB CsMybA, Bar bialaphos resistance, GFP enhanced green fluorescent, Luc firefly luciferase, Sul1 sulfadiazine resistance
Transgenic rice co-transformed, simultaneously, with three genes, the cry1Ac and cry2A genes and the lectin gene from snowdrop (Galanthus nivalis agglutinin; gna) exhibited higher resistance to the important insect pests of rice: rice leafroller (Cnaphalocrocis medinalis), brown planthopper (Nilaparvata lugens) and yellow stem borer (Scirpophaga incertulas) [16]. Walnut rootstock genotype was co-transformed to stack resistance genes for crown gall and nematodes infection using A. tumefaciens binary vector, pDE00.0201 carrying the iaaM (tryptophan 2-monooxygenase), ipt (isopentenyl transferase), GUS (β-glucuronidase), and nptII genes, and A. rhizogenes vector, pGR-Pv010 carrying Pv010 (from Pratylenchus vulnus) and GFP genes. The genes, iaaM, ipt and pv010 were silenced using the RNAi. Silencing of these genes in the transgenic lines caused complete suppression of the crown gall and 32% fewer nematodes than the control lines [17]. Transgenic potato were generated by transforming with a gene construct containing both the cisgenic late blight (Phytophthora infestans (Pi) R genes, the Rpi-vnt1.1 (Solanum venturii) and the Rpi-sto1 (S. stoloniferum), but with no selection marker gene, nptII, using Agrobacterium-mediated transformation. The transformed cells or events were screened by PCR analysis in the regenerated shoots. The transgenic marker-free potato exhibited broadspectrum and durable resistance to the late blight infection [18]. In another attempt, Zhu et al. [19] transformed potato susceptible to late blight by introducing the three genes, Rpi-sto1, Rpi-vnt1.1 and Rpi-blb3 (S. bulbocastanum). The transgenic plants stacked with the triple Rpi genes were found highly resistant to late blight (Table 1 and Fig. 1e).
Re-transformation
Re-transformation of a transgenic plant can be employed to stack or pyramid transgenes into one plant line. For example, Singla-Pareek et al. [20] re-transformed a transgenic tobacco containing glyoxalase I (gly-I) with the glyoxalase II (gly-II) gene which showed enhanced salinity tolerance. [21] re-transformed transgenic potato containing dermaseptin (from Phyllomedusa sauvagii) with double gene construct, the AP24 osmotin (from Nicotiana tabacum) and the lysozyme (from Gallus gallus). Increased level of resistance to Erwinia carotovora was found in the transgenic lines expressing the dermaseptin and lysozyme sequences. These transgenes also exhibited enhanced resistance against F. solani, Phytophthora infestans and Rhizoctonia solani, depicting that stacking of transgenes is an effective strategy to get higher resistance to the fungal and bacterial pathogens.
The disadvantages of using the consecutive transformation strategy include the need for unique selection agents and markers, which are limited and the multiple genomic locations of transgene insertion. It is difficult to obtain offspring with transgenes all localized together in the progeny following genetic segregation. After each transformation, transgenic lines have to be screened for position effects, which render this method less than optimal and practical. Marker-free transformation in which the selection marker gene is excised from the transgenic plants can be an alternate and environment-friendly option to re-transform the transgenic plants with another gene. Site-specific recombination systems (Cre-Lox, FLP-FRT and R-RS recombination) have been used for deletion and integration of DNA sequence at specific sites within genome [22,23,24,25]. When selection marker-free transgenic tobacco containing the ChiC gene was re-transformed with WD gene, the transgenic plants co-expressing both the genes were found significantly with higher resistance to F. oxysporum f.sp. nicotianae (Fon) than the corresponding isogenic lines expressing single gene [26]. Previously, using the multi-auto-transformation (MAT) vector system [27], we produced transgenic potato (free of the selection marker) containing transgene, ChiC, [28]. The marker-free potato was re-transformed with WD gene to stack the two antifungal genes, ChiC and WD (Fig. 1a–c). The transgenic plants expressing both the transgenes were found more resistant to F. oxysporum and A. solani than the wild-type and the single-gene transgenic lines [9].
The marker-free transgenic rice containing the rice chi11 gene was re-transformed with AP24 (tobacco osmotin) using the A. tumefaciens harbouring cointegrate vector, pGV2260::pSSJ1 (a single-copy) and the binary vector, pBin19DnptII-ap24 (a multi-copy) in the same cell. The transgenic plants expressing the stacked genes were found highly resistant to Rhizoctonia solani [29].
Stacked Genes with Linker Peptide
Uncoordinated expression is considered a major constraint in the co-expression of the transgenes, even if the transgenes are linked physically [30]. Multiple copies of transgenes in genome of the transgenic plants may also lead to silencing of the transgenes [31]. To cover these limitations, gene sequences for different proteins can be introduced in an open reading frame using the short linkers. The linker peptides, subsequently, are cleaved in protein units by proteinase from the host cell when passing through the endomembrane system [32].
Jha and Chatto prepared a gene construct, consisting of two antimicrobial proteins, Rs-AFP2 and Dm-AMP1 linked by Ib-AMP linker peptide (16 amino acids) extracted from seeds of Impatiens balsamina (Fig. 1d). Transgenic rice was produced with single-protein gene and the cleavable chimeric polyprotein gene constructs using Agrobacterium-mediated transformation. It was found that the transgenic rice showed increased resistance to rice blast fungus (90% higher) and Rhizoctonia bacteria (79% higher) than the wild-type rice [8].
Francoise and his co-workers expressed a chimeric polyprotein, consisting of two AMPs linked by the LP4, in transgenic Arabidopsis. The chimeric proteins conferred antifungal activity under in vitro condition [32]. 2A, a linker peptide, isolated from the virus causing foot-and-mouth disease, has widely been used in potato, tobacco, tomato, and other crops for gene fusions [33,34,35]. Researchers have also used 2A for transgene stacking in staple food crops [36, 37]. 2A has also been used as linker peptide between carotene desaturase gene (isolated from Pantoea) and phytoene synthase gene (isolated from Capsicum) to make a fusion vector construct and was introduced into rice for high carotenoid contents in “Golden Rice” [38].
Advantages of Stacking Transgenes
Pyramiding more than one transgene in crops may offer broader and more effective disease resistance and other agronomic characters that farmers need for higher yield and quality products. Gene stacking has the potential to pyramid transgenes for control of insect pests, fungal, bacterial and viral pathogens, weeds and abiotic stresses. The Bt gene technology has demonstrated well the multi-gene insect resistance for stronger and durable resistance against different types of insect pests as it is likely that the pest may not overcome the multiple insecticidal proteins [39]. Similarly, transgene stacking for conferring resistance to the commonly used herbicides has also been reported for the different herbicidal mode of action [40]. For example, the glyphosate-resistant gene, epsps was stacked with the pat (phosphinothricin N-acetyltransferase) gene for increased resistance to the herbicide, glufosinate, and/or with the dmo (bacterial dicamba monooxygenase) gene for higher resistance to the herbicide, dicamba [40].
The pyramided transgenic plants, as reported, exhibited stronger activation of the PR genes in response to pathogen infection than the single-gene-expressed plants [41]. The higher and stronger expression of the endogenous PR genes could be the result of synergistic effect of the stacked genes. Along with the PR genes, expression of other genes, such as allene oxide synthase (AOS), phenylalanine ammonia-lyase (OsPAL), and genes for jasmonic acid (JA) and SA (salicylic acid)-dependent signaling pathways, chitin-induced phytoalexins encoding gene from rice (OsMAPK6), and a rice homolog of the Arabidopsis NPRl (OsNHI), was also found stronger in the pyramided transgenic rice compared to the single-gene transgenics [41].
Gene stacking has been one of the effective approaches for metabolic engineering of the plants as most of the metabolic processes, the biochemical pathways and complex traits involve several interacting genes [42]. For example, all the signaling pathway for biosynthesis of the provitamin A (β-carotene) was genetically engineered in rice endosperm by stacking three β-carotene biosynthesis genes; the phytoene synthase (psy) isolated from daffodil (Narcissus pseudonarcissus, the phytoene desaturase (crtI) originated from Erwinia uredovora and the lycopene beta-cyclase (from N. pseudonarcissus) into rice [43]. Biosynthesis of provitamin-A was found improved in the endosperm of transgenic rice with the stacked transgenes. A modified flower color was developed in roses (biotech rose) by pyramiding two genes in pathway of the anthocyanin biosynthesis that altered pigmentation of the flower, imparting the biotech roses novel shades of blue coloration [44]. Genetic manipulation for down or up-regulation of flavonoid and anthocyanin pathway has lead to the development of changed color varieties in roses and other cut-flower plants [44].
Limitations
Stacking genetically modified traits may offer durable and effective multiple insect pests or pathogens resistance or multiple metabolic engineering for improving nutritional food quality and quantity. However, the advancement of the GM traits is still difficult because of some major hurdles. Some of the traits like yield, nutritional value, or quality of yield products need several genes to alter the several interconnected pathways regulating the complex traits. Very few genetically modified crops transformed with three or more stacked genes have yet obtained the regulatory approval such as multiple virus resistance in the squash. In addition, re- or co-transformation of the multiple transgenes driven by same promoter may result in transgene silencing.
A recently introduced system for multi-gene transformation is reported by [45] in Arabidopsis. The GAANTRY (Gene Assembly in Agrobacterium by Nucleic acid Transfer using Recombinase technologY) system can be used for flexible and in vivo stacking of multifaceted genes within a T-DNA of Agrobacterium plasmid. They evaluated the system in Arabidopsis by introducing 10 genes-stack T-DNA consisting of eight transcriptional units; sul1 (sulfadiazine resistance), luc (firefly luciferase), eGFP (enhanced Green Fluorescent Protein), bar (bialaphos resistance), GUS (b-glucuronidase), CsMybA (Citrus sinensis anthocyanin-promoting Myb genes), tdTomato (Tandem dimeric orange fluorescent protein) and nptII genes (Fig. 1f). Many of the transgenic lines expressed all eight of the transgenic traits with varying level of expression. The GAANTRY system could be further evaluated in other plant species for multi-gene stacking and stable transformation.
Conclusions and Future Prospects
Transgene stacking has been used as one of the effective strategies of conferring disease resistance by incorporating more than one gene in transgenic crops. Introduction of traits controlled by single gene such as insect resistance or the herbicide tolerance has been proved well in agriculture, improvement of the multi-gene traits such as yield, nutritional quality and stress resistance will need integration of several genes and more sophisticated techniques. The recently introduced system, the GAANTRY system may enable the researchers to address some of these challenging tasks. In addition, therapeutic proteins can be expressed in transgenic plants as edible vaccines using multigene transformation technology.
References
Ronald, P. C. (1997). The molecular basis of disease resistance in rice. Plant Molecular Biology,35, 179–186.
Zhang, S., Song, W. Y., Chen, L., Ruan, D., Taylor, N., Ronald, P. C., et al. (1998). Transgenic elite indica rice varieties, resistant to Xanthomonas oryzae pv.oryzae. Molecular Breeding,4, 551–558.
Mew, T. W., Vera Cruz, C. M., & Medalla, E. S. (1992). Changes in the race frequency of Xanthomonas oryzae pv. oryzae in response to the planting of rice cultivars in the Philippines. Plant Disease,76, 1029–1032.
Douglas, E., & Halpin, C. (2010). Gene stacking. Molecular techniques in crop improvement (2nd ed., pp. 613–629). The Netherlands: Springer.
James, C. (2007). Global status of commercialized biotech/GM crops in 2007. New York: ISAAA.
Takakura, Y., Ito, T., Saito, H., Inoue, T., Komari, T., & Kuwata, S. (2000). Flower predominant expression of a gene encoding a novel class I chitinase in rice (Oryza sativa L.). Plant Molecular Biology,42, 883–897.
Kim, J. K., Jang, I. C., Wu, R., Zuo, W. N., Boston, R. S., Lee, Y. H., et al. (2003). Coexpression of a modified maize ribosome-inactivating protein and a rice basic chitinase gene in transgenic rice plants confers enhanced resistance to sheath blight. Transgenic Research,12, 475–484.
Jha, S., & Chattoo, B. B. (2009). Transgene stacking and coordinated expression of plant defensins confer fungal resistance in rice. Rice,2, 143–154.
Khan, R. S., Darwish, N. A., Khattak, B., Ntui, V., Kong, K., Shimomae, K., et al. (2014). Retransformation of marker-free potato for enhanced resistance against fungal pathogens by pyramiding Chitinase and Wasabi defensin genes. Molecular Biotechnol,56, 814–823.
James, C. (2013). Global status of commercialized biotech/GM crops: 2013 ISAAA brief no. 46 (p. 315). Ithaca, NY: International Service for the Acquisition of Agri-biotech Applications (ISAAA).
Zhao, J.-Z., et al. (2003). Transgenic plants expressing two Bacillus thuringiensis toxins delay insect resistance evolution. Nature Biotechnology,21, 1493–1497.
Steffey, K., Gray, M., & Estes, R. (2009). Traits for insect control with transgenic Bt Corn: What, why, and how now and in the future. In The proceedings of the 2009 University of Illinois Corn & Soybean Classics.
Zhao, Q. C., Liu, M. H., Zhang, X. W., Lin, C. Y., Zhang, Q., & Shen, Z. C. (2015). Generation of insect- resistant and glyphosate-tolerant rice by introduction of a T-DNA containing two Bt insecticidal genes and an EPSPS gene. Journal of Zhejiang University (Science B),16, 824–831.
Naqvi, R. Z., Asif, M., Saeed, M., Asad, S., Khatoon, A., Amin, I., et al. (2017). Development of a triple gene Cry1Ac-Cry2Ab-EPSPS construct and its expression in Nicotiana benthamiana for insect resistance and herbicide tolerance in plants. Frontiers in Plant Science,8, 55.
Mehrotra, M., Singh, A. K., Sanyal, I., Altosaar, I., & Amla, D. V. (2011). Pyramiding of modified cry1Ab and cry1Ac genes of Bacillus thuringiensis in transgenic chickpea (Cicer arietinum L.) for improved resistance to pod borer insect Helicoverpa armigera. Euphytica,182, 87–102.
Maqbool, S. B., Riazuddin, S., Loc, N. T., Gatehouse, A. M. R., Gatehouse, J. A., & Christou, P. (2001). Expression of multiple insecticidal genes confers broad resistance against a range of different rice pests. Molecular Breeding,7, 85–93.
Walawage, S. L., Britton, M. T., Leslie, C. A., & Uratsu, S. L. (2013). Stacking resistance to crown gall and nematodes in walnut rootstocks. BMC Genomics,14, 668. https://doi.org/10.1186/1471-2164-14-668.
Jo, K. R., Kim, C. J., Kim, S. J., Kim, T. J., Bergervoet-van Deelen, J. E. M., Jongsma, M. A., et al. (2014). Development of late blight resistant potatoes by cisgenic stacking. BMC Biotechnology,14, 1472–6750.
Zhu, S., Li, Y., Vossen, J. H., Visser, R. G. F., & Jacobsen, E. (2012). Functional stacking of three resistance genes against Phytophthora infestans in potato. Transgenic Research,21, 89–99.
Singla-Pareek, S. L., Reddy, M. K., & Sopory, S. K. (2003). Genetic engineering of the glyoxlase pathway in tobacco leads to enhanced sa-linity tolerance. Proceedings of the National Academy of Sciences USA,100(25), 14672–14677.
Rivero, M., Furman, N., Mencaccia, N., Picca, P., Toum, L., Lentz, E., et al. (2012). Stacking of antimicrobial genes in potato transgenic plants confers increased resistance to bacterial and fungal pathogens. Journal of Biotechnology,157, 334–343.
Chakraborti, D., Sarkar, A., Hossain, A., Mondal, H. A., Schuer-mann, D., Hohn, B., et al. (2008). Cre/lox system to develop selectable marker-free transgenic tobacco plants conferring resistance against sap sucking homopteran insect. Plant Cell Reports,27(10), 1623–1633.
Hou, L., Yau, Y. Y., Wei, J., Han, Z., Dong, Z., & Ow, D. W. (2014). An open-source system for in planta gene stacking by Bxb1 and Cre recombinases. Molecular Plant,7, 1756–1765.
Khan, R. S., Nakamura, I., & Mii, M. (2010). Production and selection of marker-free transgenic plants of Petunia hybrida using sitespecific recombination. Biologia Plantarum,54, 265–271.
Sauer, B. (1987). Functional expression of the Cre-lox site-specific recombination system in the yeast Saccharomyces cerevisiae. Molecular and Cellular Biology,7, 2087–2096.
Ntui, V. O., Azadi, P., Thirukkumaran, G., Khan, R. S., Chin, D. P., Nakamura, I., et al. (2011). Increased resistance to fusarium wilt in transgenic tobacco lines co-expressing chitinase and wasabi defensin genes. Plant Pathology,60, 221–231.
Ebinuma, H., & Komamine, A. (2001). Mat (Multi-Auto-Transformation) vector system. The oncogenes of Agrobacterium as positive markers for regeneration and selection of marker-free transgenic plants. In Vitro Cellular & Developmental Biology-Plant,37, 103–113.
Khan, R. S., Ntui, V. O., Chin, D. P., Nakamura, I., & Mii, M. (2011). Production of marker-free disease-resistant potato using isopentenyl transferase gene as a positive selection marker. Plant Cell Reports,30, 587–597.
Ramana Rao, M., Parameswari, C., Sripriya, R., & Veluthambi, K. (2011). Transgene stacking and marker elimination in transgenic rice by sequential Agrobacterium-mediated co-transformation with the same selectable marker gene. Plant Cell Reports,30, 1241–1252.
Maqbool, S. B., & Christou, P. (1995). Multiple traits of agronomic importance in transgenic indica rice plants: Analysis of transgene integration patterns, expression levels and stability. Molecular Breeding,5, 471–480.
Matzke, A. J., & Matzke, M. A. (1998). Position effects and epigenetic silencing of plant transgenes. Current Opinion in Plant Biology,1, 14–148.
Francois, I. E., De Bolle, M. F., Dwyer, G., Goderis, I. J., Woutors, P. F., Verhaert, P. D., et al. (2002). Transgenic expression in Arabidopsis of a polyprotein construct leading to production of two different antimicrobial proteins. Plant Physiology,128, 1346–1358.
Kwon, H. B., Kwon, S. J., & Hwang, E. W. (2004). Genetic engineering of drought resistant potato plants by co-introduction of genes encoding trehalose-6-phosphate synthase and trehalose-6-phosphate phosphatase of Zygosaccharomyces rouxii. Korean Journal of Genetics,26, 199–206.
Ralley, L., Enfissi, E. M., Misawa, N., Schuch, W., Bramley, P. M., & Fraser, P. D. (2004). Metabolic engineering of ketocarotenoid formation in higher plants. The Plant Journal,39, 477–486.
Randall, J., Sutton, D., Ghoshroy, S., Bagga, S., & Kemp, J. D. (2004). Co-ordinate expression of beta- and delta-zeins in transgenic tobacco. Plant Science,167, 367–372.
Park, S., Kang, K., Kim, Y. S., & Back, K. (2009). Endosperm-specific expression of tyramine N- hydroxycinnamoyltransferase and tyrosine decarboxylase from a single self-processing polypeptide produces high levels of tyramine derivatives in rice seeds. Biotechnology Letters,31, 911–915.
Quilis, J., Lopez-Garcia, B., Meynard, D., Guiderdoni, E., & San Segundo, B. (2014). Inducible expression of a fusion gene encoding two proteinase inhibitors leads to insect and pathogen resistance in transgenic rice. Plant Biotechnology Journal,12, 367–377.
Ha, S. H., Liang, Y. S., Jung, H., Ahn, M. J., Suh, S. C., Kweon, S. J., et al. (2010). Application of two bicistronic systems involving 2A and IRES sequences to the biosynthesis of carotenoids in rice endosperm. Plant Biotechnology Journal,8, 928–938.
Storer, N. P., Thompson, G. D., & Head, G. P. (2012). Application of pyramided traits against Lepidoptera in insect resistance management for Bt crops. GM Crops and Food,3(3), 154–162.
International Survey of Herbicide Resistant Weeds. (2017). Herbicide-Resistant Weeds by Site of Action. Retrieved from http://www.weedscience.org/Summary/SOASummary.aspx.
Karmakar, S., Molla, K. A., Das, K., Sarkar, S. N., Datta, S. K., & Datta, K. (2017). Dual gene expression cassette is superior than single gene cassette for enhancing sheath blight tolerance in transgenic rice. Scientific Reports,7, 7900.
Naqvi, S., Farre, G., Sanahuja, G., Capell, T., Zhu, C., & Christou, P. (2009). When more is better: Multigene engineering in plants. Trends in Plant Science,15, 48–56.
Ye, X. D., Al Babili, S., Kloti, A., Zhang, J., Lucca, P., Beyer, P., et al. (2000). Engineering the provitamin A (beta-carotene) biosynthetic pathway into (carotenoid-free) rice endosperm. Science,287, 303–305.
Tanaka, Y., Brugliera, F., & Chandler, S. (2009). Recent progress of flower colour modification by biotechnology. International Journal of Molecular Sciences,10, 5350–5369.
Collier, R., Thomson, J. G., & Thilmony, R. (2018). A versatile and robust Agrobacterium-based gene stacking system generates high quality transgenic Arabidopsis plants. Plant Journal. https://doi.org/10.1111/tpj.13992.
Monsanto. (2009). Monsanto biotechnology trait acreage: Fiscal years 1996 to 2009.
Bakhsh, A., Dinc, T., Hussain, T., Demirel, U., Aasim, M., & Çalışkan, M. E. (2018). Development of transgenic tobacco lines with pyramided insect resistant genes. Turkish Journal of Biology,42, 174–186.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Shehryar, K., Khan, R.S., Iqbal, A. et al. Transgene Stacking as Effective Tool for Enhanced Disease Resistance in Plants. Mol Biotechnol 62, 1–7 (2020). https://doi.org/10.1007/s12033-019-00213-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-019-00213-2