Abstract
N6-methyladenosine (m6A) is the most abundant—internal modification of eukaryotic mRNA. m6A can be installed and removed by specific enzymes. The “writer,” “eraser,” and “reader” of m6A modification have been reported. These discoveries facilitate our understanding of the functional significance of m6A. m6A plays an essential role in diverse biological processes by recruiting the corresponding YTH domain-containing proteins, as well as recruiting additional translation initiation factors. Here, we provide an update on the various aspects of YTH domain-containing proteins, including an introduction to the YTH domain, the categories, distribution in cells, and biological roles of YTH proteins. Then we focus on the mechanisms that YTH proteins recognize m6A and mediate the fate of methylated-RNAs in eukaryotic cells.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
N6-methyladenosine (m6A) is the most abundant and widely conserved internal modification of message RNA [1, 2], which was discovered in a large-scale of RNAs in 1970s [3,4,5]. However, due to the technical limitations, the function of m6A has not been intensively studied. In 2011, the discovery of the m6A demethylase FTO defines m6A modification as a dynamic process [6]. It revives the interest of mRNA/lncRNA methylation field. In 2012, the novel m6A profiling analysis, which is based on m6A specific antibody-mediated capture and next-generation sequencing, advanced our understandings of the methylated transcriptome in mammals [7, 8]. These results reignited research into the mechanisms of m6A-mediated functions. Subsequent studies show that m6A contributes to diverse aspects of gene regulation [9], cell renewal and differentiation [10], and gene stability maintenance [11] by recruiting specific proteins [1]. The m6A modification performs diverse functions through binding to specific proteins. It is feasible that m6A mRNA modification executes its functions through two major methods: destabilize local RNA structure to block or induce protein–RNA interactions, or directly recruit specific proteins to mRNA therefore induce subsequent reactions. m6A recruits translation initiation factor eIF3 [12] and YTH domain-containing proteins which include YTHDF1, YTHDF2, YTHDF3, YTHDC1, YTHDC2 [13, 14]. Studies reveal that YTH domain-containing proteins combine to m6A transcript and determine its subsequent preference. Here, we review the various capabilities of YTH domain-containing proteins in regulating m6A in transcriptome.
m6A is a Dynamic and Reversible Modification in RNA
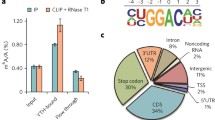
m6A methylation was firstly reported in 1974 [15]. It tends to be regularly distributed over the coding sequences (CDS), 3′-untranslated region (3′ UTR), stop codons, and the long exon regions of the mRNA [7, 8, 13, 16]. These modified regions have a high degree of conservation in human and mice [7, 8, 13]. Early studies used a liquid chromatography technique to reveal the m6A content, which m6A accounted for 0.2–0.6% of the total adenosines content in the mRNA [17]. It was also found that the modified regions tended to contain “RRACH” or “DRACH” (with D = G, A, or U; R = G or A; and H = C, A, or U) consensus motif [4, 18]. As other epigenetic modifications, m6A also has a dynamic and reversible mechanism, which could be established by the METTL3–METTL14–WTAP methyltransferase complex (writers of methylation) [18, 19] and removed by demethylases (erasers) including FTO [6] and ALKBH5 (Fig. 1) [20]. A multiprotein methyltransferase complex methylates m6A on transcriptome [21,22,23]. METTL3 is firstly identified as a S-adenosyl-l-methionine (SAM)-binding component and processes catalytic functions of the methyltransferase complex [23]. Although METTL14 is a homologue gene of METTL3 [17], only METTL3 can bind to SAM, whereas METTL14 binds to the RNA substrate and regulates the activity of METTL3 [18, 24, 25]. Deficiency of METTL3 or METTL14 both lead to depletion of m6A on mRNA [18, 26, 27]. WTAP also cooperates with METTL3–METTL14 core complex [18, 19] and facilitates complex translocation to nuclear speckles [19]. FTO and ALKBH5 are identified as demethylases and they both belong to the α-ketoglutarate-dependent dioxygenase family [6, 20, 28]. The activity of FTO and ALKBH5 demethylases is likely transcript-specific. Only certain transcripts are actively demethylated by these enzymes [29, 30]. Deficiency of FTO in HeLa and 293FT cells enhances total m6A levels in transcriptome, and overexpression of FTO reduces m6A levels on RNA [6]. Equally, knockdown of ALKBH5 on human cell lines turns to increased total m6A levels.
The regulation process of the m6A writer proteins (m6A methyltransferase complex METTL3, METTL14, WTAP RBM15), eraser proteins (ALKBH5, FTO), and reader proteins (YTH domain-containing proteins). METTL3 methyltransferase like 3, METTL14 methyltransferase like 14, WTAP Wilms’ tumor 1 associated protein, ALKBH5 AlkB homolog 5 RNA demethylase, FTO fat mass and obesity-associated protein, YTH YT521-B homology
In the last few years, numerous literatures showed the significant roles of m6A in biological processes. It impacts the pluripotency of embryonic stem cells (ESCs) [26, 27, 31], cancer [32, 33], circadian rhythm [34,35,36], virus replication [37, 38], immunity [39, 40], and sex determination [41, 42]. m6A modification influences the fate of mRNA in cells by affecting RNA stability, mRNA translation, and post-transcriptional processing [9]. However, the underlying mechanisms of m6A functions still are a riddle. Besides, m6A can destabilize local RNA structure [43] and promote proteins to binding sites adjacent to m6A. Another influential effect of m6A is to recruit precise binding proteins to transcriptome. These proteins recognize m6A through a distinct YTH domain. Here, we review the various functions of YTH domain-containing proteins about mediating the results of m6A in mRNA.
YTH Proteins Recognize m6A Through YTH Domain
Initially, yeast two-hybrid screening technique has been used for exploring functions of RA301 which is an RNA-binding protein. Furthermore, researchers cloned a new RNA-binding protein, designated YT521-B, for RA301 [44]. YT521-B is the first discovered YTH protein which acts as a novel candidate for RNA splicing-related proteins [45]. Studies show that YT521-B homology is typical and abundant in the eukaryotes and plants, termed YT homology (YTH) domain. Studies reveal that YTH domains are formed with six β strands and four or five α helices [46]. These six β strands form a β barrel, with the α helices wrapped against the β strands to stabilize the hydrophobic activation core [46]. The approximately 140 amino acid domain shows remarkable conservation across the species which includes 14 invariant and 19 highly conserved residues. The aromatic residues in the β-sheets of the YTH domain are homologous with the RNA-binding RRM (RNA recognition motif) domain, indicating that the YTH domain probably serves as the m6A-binding module. The RRM β-sheet domain has a crucial function on binding to RNA [47,48,49]. To recognize the binding mechanism of YTH domain, subsequent studies have explored the molecular basis of YTH domain. The aromatic cage of YTH domain is pivotal in recognizing m6A modification. The residues of tyrosine (Y260) and tryptophan (W200) in the protein sandwich the m6A group. The methyl group is pointed toward tryptophan (W254) which forms the base of the cage. N6 amino group utilizes its remaining hydrogen atom to bond the Ser201 main-chain carbonyl oxygen. The interactions between m6A and aromatic cage can make a combination of m6A and YTH domain-containing proteins and simultaneously prevent YTH domain from binding double methylated adenines [50, 51]. The amino acids of the cage are highly conserved in YTH proteins across diverse organisms. The affinity of YTH domain with m6A is controversial. Some studies showed the affinity is in the range of 100 and 300 nM [46, 50, 52], while other studies showed an affinity of between 1 and 3 µM [51, 53]. According to the non-redundant protein–RNA-binding benchmark dataset, unlike the specific RNA-binding interactions, the non-methylated RNA generally perform a weaker binding affinity about fivefold to tenfold, which is in a low nanomolar range [54]. It suggests that although the YTH domain binds to RNA primarily through m6A site, it also can bind RNA through the presence of other modifications. Is it possible for the YTH domain to interact with other methylation modifications of adenosine, for example, N6,2′-O-dimethyladenosine (m6Am), which is also one of the prevalent modifications? m6Am is a reversible modification that influences cellular mRNAs fate of stability. It sits at the 5′ end of mRNAs, the first encoded nucleotide adjacent to the 7-methylguanosine cap [55, 56]. YTHDF1, YTHDF2, and YTHDF3 could show a specific recognition of the 2′-OH with a side chain asparagine, and YTHDC1 mediates this function through a distinct asparagine. Thus, these YTH domains unlikely bind to m6Am [46, 50, 53]. The structural and CLIP studies suggest that the YTH domain does not bind to other modified adenosines, such as m1A, or m6′6A [57]. Recent studies employ a quantitative proteomics method, identifying YTHDF1-3 and YTHDC1, besides YTHDC2, directly bind to m1A in RNA [58, 59]. This is consistent with previously published data, which reveal transcriptome-wide colocalization of YTH domain-containing proteins and the m1A sites in HeLa cells [59, 60]. Researchers have found that Trp432 which is conserved in the hydrophobic pocket of the YTH domain in YTHDF2 is necessary for recognition of m1A [58].
Diversity of m6A-binding proteins
The YTH domain exists in 174 different proteins and is conserved in the eukaryotic species. YTH domain-containing proteins have three categories: YTHDF family proteins, YTHDC1 and YTHDC2 protein. The YTHDF family proteins have more similarities in amino acids identity, whereas YTHDC1 and YTHDC2 are not homologous to YTHDF family except for the YTH domain and family name [41, 42]. YTHDF family proteins include three classic proteins which share high gene homology and amino acids composition upon their entire structure: YTHDF1, YTHDF2, and YTHDF3. The YTHDF family proteins contain a C-terminal YTH domain in the gene structure and a simple region of approximately 350 amino acids, which lacks specific modular protein domain and includes some P/Q/N-rich patches [61]. Recent studies have also examined YTHDC2 homologs across species [61]. YTHDC2 includes R3H, DEXDc, ankyrin repeats (ANK), HELICc, HA2, and OB-fold domains beside a YTH domain. YTH proteins show different distribution over cells, among which YTHDF family proteins and YTHDC2 are widely distributed in the cytoplasm, while YTHDC1 is abundant in nucleus. YTH proteins have a function in enhancing the translation efficiency and promoting mRNA degradation [13, 62,63,64]. YTHDC1 mainly distributes in the nucleus, and it relates RNA post-transcriptional splicing and chromosome modification [9, 57].
With the exception of YTH domain-containing proteins, eukaryotic initiation factor 3 (eIF3) is also binding to the m6A [65]. Study shows that eIF3 facilitates eIF4-independent translation initiation through a m6A-mediated cap-independent manner in the 5′ UTRs [65]. Besides, HNRNPA2B1, a RBP containing RRM domain, could also bind to m6A-rich sites in the mRNA [66,67,68]. The recent study reveals that insulin-like growth factor 2 mRNA-binding proteins (IGF2BPs; including IGF2BP1/2/3) could target mRNA transcripts through recognizing the m6A site and acts as a m6A-binding protein [69].
Several YTH domain-containing proteins are expressed in plants [70]. The YTH domain-containing proteins, ECT proteins, part of EVOLUTIONARILY CONSERVED C-TERMINAL REGION1-11 (ECT1-11) family proteins in Arabidopsis, are revealed as m6A reader proteins. The mRNA-protein interactome study indicates that YTH domain of several ECT family proteins has the capacity to bind to m6A site of mRNA in vivo [71]. The homology modeling strongly indicates the existence of intact m6A binding sites in ECT2 and ECT3 [72]. Researchers have developed the formaldehyde cross-linking and immunoprecipitation method to recognize that the 3′ UTRs of target genes are enriching ECT2 binding sites and ECT2 leads to the recognition of a plant-specific m6A motif, URUAY (R = G > A, Y = U > A, where the majority [over 90%] is UGUAY) [71]. Studies suggest that the location of ECT2 causes dual roles in modulating 3′ UTR processing in the nucleus and thereby facilitating mRNA stability in the cytoplasm sequently [71, 72].
Biological Functions of YTH Proteins
The YTH proteins, as the “reader” of m6A, target to a specific m6A site via a methylation-dependent manner. By recruiting diverse “readers” to bind m6A sites, the YTH domain-containing proteins, no less than other RBPs, contribute to biological process in many aspects.
Overexpression of YTHDF1 could enhance the HIV-1 replication [37]. Scholars precisely describe the m6A editing site map on the HIV-1 genome showing that the m6A sites cluster in the HIV 3′-untranslated region (3′ UTR). YTHDF proteins bind to m6A sites to enhance the HIV-1 replication [73]. YTHDF family protein-mediated inhibition of HIV-1 infection is due to the negative regulation on HIV-1 post-entry infection. YTHDF family proteins and HIV-1 Gag make a complex with RNA in cells. YTHDF1-3 blocks the viral reverse transcription by degrading the HIV-1 Gag RNA in infected cells [73]. YTHDF family proteins affect the HIV virus self-replication and reverse transcription process, thus affecting the HIV virus infection [37]. Recent studies show that YTHDF1 is associated with various tumor behaviors and cancer stages. YTHDF2 is also thought to play roles in acute myeloid leukemia [74]. Researchers utilized 3′ rapid amplification of cDNA ends method and panhandle polymerase chain reaction to identify YTHDF2 as a novel RUNX1 translocation partner gene in the three acute myeloid leukemia (AML) patients [74]. The functions of YTHDF2 in the AML disease need to be further researched. Heat shock stress is an important issue, which related to many aspects of animal health, breeding and development, and production performance. The nuclear YTHDF2 preserves 5′ UTR methylation upon heat shock stress-induced transcripts by limiting the m6A demethylase FTO from demethylation [75]. The increased m6A content promotes cap-independent translational initiation and provides a mechanism underlying the selective mRNA translation under the heat shock response situation [75]. In our group, we find that YTHDF1 plays important roles in adipogenesis. Our study shows that the ratio of m6A/A in the layer of backfat (LB) is prominently higher in Landrace (lean models) than that in Jinhua (fat models). Patatin-like phospholipase domain containing 2 (PNPLA2) is one of the unique m6A peak genes in Landrace LB. The result shows that overexpression of YTHDF1 increases PNPLA2 expression [76], further inhibiting the adipogenesis. YTHDF family proteins also affect the cell cycle. We find EGCG could increase the expression of YTHDF2 and lead to the down-regulation of protein levels of CCNA2 and CDK2. It means that YTHDF2 plays an important role in cell cycle [77]. In addition, YTHDF2 accelerates the clearance of over one-third of maternal-to-zygotic transition (MZT) related transcript during the early life of embryos [78]. Deficiency of YTHDF2 in zebrafish embryos hinders zygotic genome expression which fails to initiate timely MZT, cell cycle, and delays the development of larval life [78]. These results highlight the vital roles of m6A readers in animal development.
The recent accompanying features find that several ECT family proteins are the m6A readers of Arabidopsis thaliana and the m6A binding function of ECT family proteins is required for the development of Arabidopsis morphology [72, 79, 80]. Studies indicate that one of the ECT family proteins, ECT2, is a m6A reader protein [79]. ECT2 is highly expressed in rapidly developing tissues and has a function on trichome morphogenesis [79]. Consistent research demonstrates that ECT2, ECT3, and ECT4 are strongly expressed at leaf formation sites in the young seedlings shoot apex and the division zone of developing leave [72]. Mutation of ECT4 enhances the delayed leaf emergence and mutation of ECT2 and ECT3 causes the defection of leaf morphology [72], In addition, ECT2 is required for normal branching of trichome through controlling ploidy levels. These indicate the m6A-YTH regulatory module play a vital role in the plant organogenesis.
YTHDC2 also influences the development of mammals. Study shows that YTHDC2 knockout mice have significantly smaller testes and ovaries than other littermates, which means YTHDC2 plays an important role in motivating early spermatogenesis [81]. In addition, YTHDC2 also relates to cell proliferation. The target genes of YTHDC2 are involved in meiosis-related classifications [81]. The MEIOC might interact with YTHDC2 for controlling the germline transition into meiosis [82]. In addition, YTHDC1 also acts on the herpes simplex virus (HSV), by affecting its infection process [83]. The results show that YTHDC1 responds to HSV-1 infection and the modulation depends on the expression of the viral E3 ubiquitin-ligase ICP0 [83]. Removing an arginine-rich region at the C-terminus of the YTHDC1 impacts its RNA-binding ability. Some researchers have figured out that the truncation of YTHDC1 is a potential candidate cancer driver mutation in Jurkat cells [84]. YTHDC2 is a member of the DExD/H-box family which belongs to RNA helicases. Knockdown of YTHDC2 reduces protein expression hypoxia-inducible factor-1 alpha (HIF-1α) and other metastasis genes, which inhibit metastasis of colon tumor cells in turn [64]. It suggests that YTHDC2 is a potential diagnostic marker and a target gene of colon cancer. It conduces to colon tumor metastasis by promoting translation of HIF-1α [64].
The recent study shows that YTH domain-containing protein, Mmi1, could sequestrate meiotic-related transcripts from the translation machinery in the fission yeast Schizosaccharomyces pombe [85]. Mmi1 tethers the specific mRNA to the nuclear foci during vegetative proliferation in order to prevent mistimed expression of meiotic proteins [85].
YTH Proteins Mediate Intracellular Biological Functions
YTH domain-containing proteins participate in extensive post-transcriptional regulation through directly binding to m6A site. As we know, YTHDF family proteins are mainly distributed among cytoplasm, suggesting that its function may be related to RNA in the cytoplasm, as they play an important role in translation and degradation of RNA [14, 55, 75]. Some researchers have found that YTHDF1 could specifically recognize and bind to m6A among 3′ UTR. It could recruit the 43S pre-initiation complex to begin translation processes [12, 65]. eIF3 is a multiprotein complex which could stimulate the eukaryotic translational initiation. The study reveals that YTHDF1 could promote ribosome binding on its target mRNAs and directly interact with initiation factors to promote translation, such like eIF3 [13]. YTHDF1 could be spatially close to translational initiation sites through eIF4G which looks like a bridge. During the classic YTHDF1-mediated translation initiation, eIF4G binds both poly (A)-binding protein and cap-binding protein eIF4E to form a “closed-loop” structure [13]. If eIF4G-mediated loop structure is interrupted to form, the stimulation of YTHDF1 on mRNAs translation would be weaker. These integrated results propose a mechanism of how YTHDF1 promotes m6A-dependent translation of mRNA. Recently, YTHDF3 is also discovered to promote translation [37]. YTHDF3 facilitates the translation of methylated mRNA through interacting with 40S and 60S ribosome subunits by cooperating with YTHDF1 [12, 86]. But we still need a clear mechanism of YTHDF1 and YTHDF3 in translational enhancement process.
On the contrary, m6A is selectively recognized by the YTHDF2 protein, which promotes mRNA degradation. The YTH domain position in C-terminal of YTHDF2 selectively binds to m6A-containing mRNAs. Whereas the N-terminal domain of YTHDF2 is intended to localize the YTHDF2-mRNA complex to P-body, which is the RNA decay site in cells [14, 87]. Other researchers suggest that YTHDF2 mediates mRNA degradation by generating mRNA deadenylation subsequently translocating to P-bodies [57]. The N-terminus of YTHDF2 recruits CCR4-NOT transcription complex subunit 1 (CNOT1), a component of the CCR4-NOT deadenylation complex, promoting deadenylation of mRNA [57]. In addition, researchers find that YTHDF family proteins recruit CCR4–NOT and enhance the deadenylation of mRNA [57]. In contrast, other researches find that that YTHDF1 has effects on mRNA stability [13]. In consideration of the different functions of YTHDF1 and YTHDF2, the researchers have found that YTHDF1 binds to RNA earlier than YTHDF2 during the mRNA life cycle, suggesting that two proteins act a cooperation to mediate translation efficiency of transcripts which are requiring a transient and dynamic control [86]. In this regulating model, YTHDF3 also plays a vital role, helping regulate the mRNAs fate by manipulating YTHDF1 and YTHDF2 [86]. Knockdown of YTHDF3 reduces the RNA-binding specificity of both YTHDF1 and YTHDF2 [86].
YTHDF family proteins have a dynamic interaction function in the cytosol: when m6A-contained mRNAs are exported to the cytoplasm from cell nucleus, it is immediately recognized by the YTHDF3 or YTHDF3-YTHDF1 complex in order to enhance the translation efficiency and then eventually bound by YTHDF2 to decay site. YTHDF3 influences the functions of YTHDF1 and YTHDF2 and mRNA translation efficiency or mRNA stability [86, 88]. Translation and degradation are two reverse fates of mRNA, suggesting a complicated dynamic network of the overall functions of m6A on mRNAs. This machinery achieves a fast response with adequate gene expression and could keep a steady protein quantity meanwhile reduce the noise of gene expression process. Recent studies show that neither of the YTHDF family proteins performs only single function, and all the three proteins have similar properties in terms of translation and degradation of mRNAs. A model that accurately characterizes the respective functions and interaction network of the three proteins is needed.
Recent studies demonstrate the subcellular localization of YTH domain-containing protein ECT2 in Arabidopsis. ECT is present in both the nucleus and cytoplasm which indicates that ECT plays dual roles on pre-mRNA processing in nucleus and mRNA metabolism in cytoplasm [79]. Researchers have found that ECT2 may specifically bind to m6A-containing poly(A), subsequently recruiting the polyadenylation machinery to process alternative polyadenylation and 3′ UTR processing [79]. Besides, ECT2 may facilitate m6A-mediated mRNA stability in the cytoplasm [79].
YTHDC1 was firstly discovered as a splicing tool for exon inclusion [89]. It is in the nucleus and the binding site is almost entirely coincident with the m6A site, indicating that it functions on nuclear mRNA processing and nuclear-localized RNAs. YTHDC1 regulates mRNA splicing through recruiting pre-mRNA splicing factors SRSF3 (SRp20) and repressing SRSF10 binding site to promote the exon inclusion [9]. The expression of SRSF3 is 30-fold higher than SRSF10 in endogenous condition, which means the exon inclusion measure by YTHDC1 and SRSF3 is probably predominant [9]. The binding ability is based on existence of m6A locus [9]. However, the evidence of m6A splicing regulation is unclear. The first in-depth analysis of altered splicing finds that knockdown METTL3 induced < 100 m6A contained-exons has been alternatively spliced [7]. Further experiments reveal that METTL3 might be involved in the splicing of mRNAs and 0.5% of the exons has been found alternatively spliced in METTL3−/− mouse embryonic stem cells [7, 27, 90].
Besides, YTHDC1 also mediates the nuclear non-coding RNA X-inactive specific transcript (XIST). XIST mediates the transcriptional silencing of genes on the X chromosome by recruiting chromatin-modifying factors and coating the X chromosome [91]. XIST is highly methylated with almost 78 m6A in human cells [57]. The highly enriched m6A throughout XIST offers the possibility about the recruitment of the nuclear m6A-binding protein YTHDC1. XIST interacts with gene-silencing proteins to achieve the silencing of X chromosome gene transcription during development of female mammalian. XIST binds to YTHDC1 therefore activates gene-silencing mechanisms [57, 91]. YTHDC1 recognizes m6A residues of XIST preferentially and is probably required for XIST functions. But how YTHDC1 binds to XIST resulting in gene silencing remains indistinct. Loss of m6A or YTHDC1 equally rescues XIST-mediated gene silencing [57].
In contrast to the other YTH domain-containing proteins, YTHDC2 has several undefined domains [92]. Recent studies reveal that apart from YTH domain, the ankyrin repeats of YTHDC2 regulate an RNA-independent interaction with the 5′–3′ exoribonuclease XRN1. Researchers have uncovered that YTHDC2 cooperates with the small ribosomal subunit in the close proximity sites of entrance and exitance of mRNA [92].
A major problem is whether these proteins have independent functions or their functions overlap with each other. Several studies report the debatable conclusions, and find that YTHDF1, YTHDF2, and YTHDF3 maybe in a similar category. Their similar sequence is an important reason to prove that three proteins have similar functions: YTHDF1, YTHDF2, and YTHDF3 do not contain diverse modules which might grant unique property to each protein. In the meantime, the m6A-mediate functions of YTHDC2 have not been completely and clearly found either. By CLIP data, YTHDC2 does not exhibit clear binding to m6A [57], thus, YTHDC2 does not show single binding mode toward m6A. YTHDC2 might bind m6A under unique condition or in certain cell type. Tumor necrosis factor-alpha induces YTHDC2 to regulate cell type [82, 93], and increase the translation efficiency of interaction to hypoxia-inducible factor-lα (HIF1-α) by spreading its 5′-UTR region through the YTHDC2 RNA helicase domain [64]. Whether these processes are mediated by m6A is unclear.
Remark and Outlook
In the past few years, although > 100 RNA modifications have been discovered in organisms, their distinct roles need further exploration. m6A mediates the extensive functions of eukaryotic RNAs. The YTH-contained proteins are relying on the YTH domain to recognize and bind m6A, thereby regulate the translational efficiency or the other fates of mRNAs. From the current research, the crystal structure and the protein conformation of YTH domain have been studied clearly. Few studies focus on the functions of YTH domain-containing proteins. It is controversial that the specific function of the YTHDF proteins is unitary or not. YTH proteins regulate the fate of gene expression by the specific control methods, which still need further studies to prove. YTHDF2 has higher expression than YTHDF1 or YTHDF3, which means using YTH proteins knockdown experiment is hampered to exhibit a significant effect as the compensation of loss YTHDF2 do. It indicates that knockout all three YTHDF proteins from cells and construct one of YTHDF proteins exogenous expression at the same time may offer the ability on finding out definite function of these proteins.
Although YTHDC1 and YTHDC2 have similar names, their genetic structures are quite different. YTHDC2 functions are still not clear and need to be further explored. YTHDC1 is the only m6A-binding protein localized in the nucleus which means it plays an important role in the regulation of RNA fate in the nucleus. Recent studies investigate several parts of complicated modules of YTHDC1 and reveal that these modules modulate the interaction between mRNA with YTHDC1 [92]. But the remaining modules which functions are still waiting to be explored. In addition, whether the functions of YTHDC2 are mediated by m6A-dependent process is completely unknown.
While studying the function of YTH proteins, it is important to investigate whether m6A has other reader-mediated mechanisms in cells and verify the function of YTH domain exerted through m6A. The corresponding effect of m6A can be studied by YTH proteins knockout cells.
And more remarkably, the biological functions of YTH domain-containing proteins are becoming a hotspot. In the past several years, researchers have focused on the capacity of YTH domain-containing protein on animal growth, reproduction, and disease. And recent studies explore the novel function of YTH domain-containing proteins on regulating the development of plants. This will provide the new thinking on prospective researches. In addition, researchers have revealed that m6A modification influences the reprogram of Naïve T cells for proliferation and differentiation [39]. Recent studies indicate the potential relationship between m6A with systemic lupus erythematosus [40]. These studies demonstrate that m6A modification acts as a critical regulator of immune cell homeostasis through mediating the gene expression of immune-related signaling pathway. However, whether YTH domain-containing proteins participate in the immune processes is still unknown. These findings suggest that YTH domain-containing proteins might act as regulators of immune-related signaling pathway through a m6A-dependent manner.
YTHDF family proteins are regulated diversely by particular signals need to be further explored. Phosphoproteomic experiments show existing mass phosphorylation sites among the P/Q/N-rich regions and adjacent to the YTH domain of the YTHDF family proteins [94]. Phosphorylation could add phosphate (PO4) to a non-polar R group of amino acid residue and change hydrophobic portion into a polar hydrophilic portion in order to alter the structure of the protein which could prevent proteins from granules [95]. Besides phosphorylation, YTHDF proteins have found to be myristoylated [96]. It suggests that post-translational modifications are likely to regulate YTHDF family proteins clustering. At present, there are no research findings that signaling pathways could modulate m6A. Therefore, whether YTH domain is regulated by some signaling pathways would advance our learning of m6A-mediated regulation and processes.
Numerous studies show that m6A plays important roles in various diseases including viral infections and cancers [37, 56, 63, 73, 97,98,99,100,101,102]. It attracts attention to the study of m6A readers [64, 83, 84]. Future study can focus on screening out the target genes of YTH proteins, to explore which are related to YTH proteins. This provides an advance for further studies of the biological functions of YTH proteins.
References
Fu, Y., Dominissini, D., Rechavi, G., & He, C. (2014). Gene expression regulation mediated through reversible m6A RNA methylation. Nature Reviews Genetics, 15, 293–306.
Wu, R., Jiang, D., Wang, Y., & Wang, X. (2016). N6-methyladenosine m6A methylation in mRNA with a dynamic and reversible epigenetic modification. Molecular Biotechnology, 58, 450–459.
Adams, J. M., & Cory, S. (1975). Modified nucleosides and bizarre 5′-termini in mouse myeloma messenger-RNA. Nature, 255, 28–33.
Wei, C. M., & Moss, B. (1977). Nucleotide-sequences at N6-methyladenosine sites of HeLa-cell messenger ribonucleic-acid. Biochemistry-US, 16, 1672–1676.
Krug, R. M., Morgan, M. A., & Shatkin, A. J. (1976). Influenza viral mRNA contains internal N6-methyladenosine and 5′-terminal 7-methylguanosine in cap structures. Journal of Virology, 20, 45–53.
Jia, G. F., Fu, Y., Zhao, X., Dai, Q., et al. (2011). N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nature Chemical Biology, 7, 885–887.
Dominissini, D., Moshitch-Moshkovitz, S., Schwartz, S., Salmon-Divon, M., et al. (2012). Topology of the human and mouse m6A RNA methylomes revealed by m6A-sEq. Nature, 485, 201–206.
Meyer, K. D., Saletore, Y., Zumbo, P., Elemento, O., et al. (2012). Comprehensive analysis of mRNA methylation reveals enrichment in 3’ UTRs and near stop codons. Cell, 149, 1635–1646.
Roundtree, I. A., & He, C. (2016). Nuclear m6A reader YTHDC1 regulates mRNA splicing. Trends Genetics, 32, 320–321.
Cui, Q., Shi, H. L., Ye, P., Li, L., et al. (2017). m6A RNA methylation regulates the self-renewal and tumorigenesis of glioblastoma stem cells. Cell Reports, 18, 2622–2634.
Xiang, Y., Laurent, B., Hsu, C. H., Nachtergaele, S., et al. (2017). RNA m6A methylation regulates the ultraviolet-induced DNA damage response. Nature, 543, 573–576.
Lee, A. S., Kranzusch, P. J., & Cate, J. H. (2015). eIF3 targets cell-proliferation messenger RNAs for translational activation or repression. Nature, 522, 111–114.
Wang, X., Zhao, B. S., Roundtree, I. A., Lu, Z., et al. (2015). N6-methyladenosine modulates messenger RNA translation efficiency. Cell, 161, 1388–1399.
Wang, X., Lu, Z., Gomez, A., Hon, G. C., et al. (2014). N6-methyladenosine-dependent regulation of messenger RNA stability. Nature, 505, 117–120.
Desrosiers, R., Friderici, K., & Rottman, F. (1974). Identification of methylated nucleosides in messenger-RNA from Novikoff hepatoma-cells. Proceedings of the National Academy of Sciences USA, 71, 3971–3975.
Batista, P. J., Molinie, B., Wang, J., Qu, K., et al. (2014). m6A RNA modification controls cell fate transition in mammalian embryonic stem cells. Cell Stem Cell, 15, 707–719.
Molinie, B., Wang, J., Lim, K. S., Hillebrand, R., et al. (2016). m6A-LAIC-seq reveals the census and complexity of the m6A epitranscriptome. Nature Methods, 13, 692–698.
Liu, J., Yue, Y., Han, D., Wang, X., et al. (2014). A METTL3-METTL14 complex mediates mammalian nuclear RNA N6-adenosine methylation. Nature Chemical Biology, 10, 93–95.
Ping, X. L., Sun, B. F., Wang, L., Xiao, W., et al. (2014). Mammalian WTAP is a regulatory subunit of the RNA N6-methyladenosine methyltransferase. Cell Research, 24, 177–189.
Zheng, G. Q., Dahl, J. A., Niu, Y. M., Fedorcsak, P., et al. (2013). ALKBH5 Is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Molecular Cell, 49, 18–29.
Narayan, P., & Rottman, F. M. (1988). An in vitro system for accurate methylation of internal adenosine residues in messenger RNA. Science, 242, 1159–1162.
Bokar, J. A., Rath-Shambaugh, M. E., Ludwiczak, R., Narayan, P., et al. (1994). Characterization and partial purification of mRNA N6-adenosine methyltransferase from HeLa cell nuclei. Internal mRNA methylation requires a multisubunit complex. Journal of Biological Chemistry, 269, 17697–17704.
Bokar, J. A., Shambaugh, M. E., Polayes, D., Matera, A. G., et al. (1997). Purification and cDNA cloning of the AdoMet-binding subunit of the human mRNA (N6-adenosine)-methyltransferase. RNA, 3, 1233–1247.
Wang, X., Feng, J., Xue, Y., Guan, Z., et al. (2017). Corrigendum: Structural basis of N6-adenosine methylation by the METTL3-METTL14 complex. Nature, 542, 260.
Wang, P., Doxtader, K. A., & Nam, Y. (2016). Basis for Cooperative function of Mettl3 and Mettl14 methyltransferases. Molecular Cell, 63, 306–317. Structural.
Wang, Y., Li, Y., Toth, J. I., Petroski, M. D., et al. (2014). N6-methyladenosine modification destabilizes developmental regulators in embryonic stem cells. Nature Cell Biology, 16, 191–198.
Geula, S., Moshitch-Moshkovitz, S., Dominissini, D., Mansour, A. A., et al. (2015). m6A mRNA methylation facilitates resolution of naive pluripotency toward differentiation. Science, 347, 1002–1006.
Gerken, T., Girard, C. A., Tung, Y. C., Webby, C. J., et al. (2007). The obesity-associated FTO gene encodes a 2-oxoglutarate-dependent nucleic acid demethylase. Science, 318, 1469–1472.
Frayling, T. M., Timpson, N. J., Weedon, M. N., Zeggini, E., et al. (2007). A common variant in the FTO gene is associated with body mass index and predisposes to childhood and adult obesity. Science, 316, 889–894.
Scuteri, A., Sanna, S., Chen, W. M., Uda, M., et al. (2010). Genome-wide association scan shows genetic variants in the FTO gene are associated with obesity-related traits. PLoS Genetics. https://doi.org/10.1371/journal.pgen.0030115.
Thomson, J. A., Itskovitz-Eldor, J., Shapiro, S. S., Waknitz, M. A., et al. (1998). Embryonic stem cell lines derived from human blastocysts. Science, 282, 1145–1147.
Jaffrey, S. R., & Kharas, M. G. (2017). Emerging links between m6A and misregulated mRNA methylation in cancer. Genome Medicine, 9, 2.
Wang, S. W., Sun, C. X., Li, J. H., Zhang, E. B., et al. (2017). Roles of RNA methylation by means of N6-methyladenosine in human cancers. Cancer Letters, 408, 112–120.
Fustin, J. M., Doi, M., Yamaguchi, Y., Hida, H., et al. (2013). RNA-methylation-dependent RNA processing controls the speed of the circadian clock. Cell, 155, 793–806.
Akhtar, R. A., Reddy, A. B., Maywood, E. S., Clayton, J. D., et al. (2002). Circadian cycling of the mouse liver transcriptome, as revealed by cDNA microarray, is driven by the suprachiasmatic nucleus. Current Biology, 12, 540–550.
Koike, N., Yoo, S. H., Huang, H. C., Kumar, V., et al. (2012). Transcriptional architecture and chromatin landscape of the core circadian clock in mammals. Science, 338, 349–354.
Kennedy, E. M., Bogerd, H. P., Kornepati, A. V. R., Kang, D., et al. (2017). Posttranscriptional m6A editing of HIV-1 mRNAs enhances viral gene expression (vol 19, pg 675, 2016). Cell Host & Microbe, 22, 830–830.
Brocard, M., Ruggieri, A., & Locker, N. (2017). m6A RNA methylation, a new hallmark in virus-host interactions. Journal of General Virology, 98, 2207–2214.
Li, H. B., Tong, J. Y., Zhu, S., Batista, P. J., et al. (2017). m6A mRNA methylation controls T cell homeostasis by targeting the IL-7/STAT5/SOCS pathways. Nature, 548, 338–342.
Li, L. J., Fan, Y. G., Leng, R. X., Pan, H. F., et al. (2018). Potential link between m6A modification and systemic lupus erythematosus. Molecular Immunology, 93, 55–63.
Kan, L., Grozhik, A. V., Vedanayagam, J., Patil, D. P., et al. (2017). The m6A pathway facilitates sex determination in Drosophila. Nature Communications, 8, 15737.
Haussmann, I. U., Bodi, Z., Sanchez-Moran, E., Mongan, N. P., et al. (2016). m6A potentiates Sxl alternative pre-mRNA splicing for robust Drosophila sex determination. Nature, 540, 301–304.
Roost, C., Lynch, S. R., Batista, P. J., Qu, K., et al. (2015). Correction to “structure and thermodynamics of N6-methyladenosine in RNA: A spring-loaded base modification”. Journal of the American Chemical Society, 137, 8308.
Imai, Y., Matsuo, N., Ogawa, S., Tohyama, M., et al. (1998). Cloning of a gene, YT521, for a novel RNA splicing-related protein induced by hypoxia/reoxygenation. Molecular Brain Research, 53, 33–40.
Hartmann, A. M., Nayler, O., Schwaiger, F. W., Obermeier, A., et al. (1999). The interaction and colocalization of Sam68 with the splicing-associated factor YT521-B in nuclear dots is regulated by the Src family kinase p59(fyn). Molecular Biology of the Cell, 10, 3909–3926.
Xu, C., Wang, X., Liu, K., Roundtree, I. A., et al. (2015). Structural basis for selective binding of m6A RNA by the YTHDC1 YTH domain (vol 10, pg 927, 2014). Nature Chemical Biology, 11, 815.
Altschul, S. F., Madden, T. L., Schaffer, A. A., Zhang, J., et al. (1997). Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Research, 25, 3389–3402.
Stoilov, P., Rafalska, I., & Stamm, S. (2002). YTH: A new domain in nuclear proteins. Trends in Biochemical Sciences, 27, 495–497.
Hoffman, D. W., Query, C. C., Golden, B. L., White, S. W., et al. (1991). RNA-binding domain of the A protein component of the U1 small nuclear ribonucleoprotein analyzed by NMR spectroscopy is structurally similar to ribosomal proteins. Proceedings of the National Academy of Sciences USA, 88, 2495–2499.
Luo, S. K., & Tong, L. (2014). Molecular basis for the recognition of methylated adenines in RNA by the eukaryotic YTH domain. Proceedings of the National Academy of Sciences USA, 111, 13834–13839.
Zhu, T., Roundtree, I. A., Wang, P., Wang, X., et al. (2014). Crystal structure of the YTH domain of YTHDF2 reveals mechanism for recognition of N6-methyladenosine. Cell Research, 24, 1493–1496.
Theler, D., Dominguez, C., Blatter, M., Boudet, J., et al. (2014). Solution structure of the YTH domain in complex with N6-methyladenosine RNA: A reader of methylated RNA. Nucleic Acids Research, 42, 13911–13919.
Li, F., Zhao, D., Wu, J., & Shi, Y. (2014). Structure of the YTH domain of human YTHDF2 in complex with an m6A mononucleotide reveals an aromatic cage for m6A recognition. Cell Research, 24, 1490–1492.
Yang, X., Li, H., Huang, Y., & Liu, S. (2013). The dataset for protein-RNA binding affinity. Protein Sci, 22, 1808–1811.
Mauer, J., Luo, X., Blanjoie, A., Jiao, X., et al. (2017). Reversible methylation of m6Am in the 5′ cap controls mRNA stability. Nature, 541, 371–375.
Wei, C. M., & Moss, B. (1975). Methylated nucleotides block 5′-terminus of vaccinia virus messenger RNA. Proceedings of the National Academy of Sciences USA, 72, 318–322.
Patil, D. P., Chen, C. K., Pickering, B. F., Chow, A., et al. (2016). m6A RNA methylation promotes XIST-mediated transcriptional repression. Nature, 537, 369–373.
Dai, X. X., Wang, T. L., Gonzalez, G., & Wang, Y. S. (2018). Identification of YTH domain-containing proteins as the readers for N1-methyladenosine in RNA. Analytical Chemistry, 90, 6380–6384.
Safra, M., Sas-Chen, A., Nir, R., Winkler, R., et al. (2017). The m1A landscape on cytosolic and mitochondrial mRNA at single-base resolution. Nature, 551, 251.
Dominissini, D., Nachtergaele, S., Moshitch-Moshkovitz, S., Peer, E., et al. (2016). The dynamic N1-methyladenosine methylome in eukaryotic messenger RNA. Nature, 530, 441.
Oates, M. E., Romero, P., Ishida, T., Ghalwash, M., et al. (2013). (DP2)-P-2: Database of disordered protein predictions. Nucleic Acids Research, 41, D508–D516.
Du, H., Zhao, Y., He, J. Q., Zhang, Y., et al. (2016). YTHDF2 destabilizes m6A-containing RNA through direct recruitment of the CCR4-NOT deadenylase complex. Nature Communications, 7, 12626.
Lichinchi, G., Gao, S., Saletore, Y., Gonzalez, G. M., et al. (2016). Dynamics of the human and viral m6A RNA methylomes during HIV-1 infection of T cells. Nature Microbiology, 1, 16011.
Tanabe, A., Tanikawa, K., Tsunetomi, M., Takai, K., et al. (2016). RNA helicase YTHDC2 promotes cancer metastasis via the enhancement of the efficiency by which HIF-1 alpha mRNA is translated. Cancer Letter, 376, 34–42.
Meyer, K. D., Patil, D. P., Zhou, J., Zinoviev, A., et al. (2015). 5′ UTR m6A promotes cap-independent translation. Cell, 163, 999–1010.
Alarcon, C. R., Goodarzi, H., Lee, H., Liu, X. H., et al. (2015). HNRNPA2B1 is a mediator of m6A-dependent nuclear RNA processing events. Cell, 162, 1299–1308.
Chen, K., Lu, Z., Wang, X., Fu, Y., et al. (2015). High-resolution N6-methyladenosine map using photo-crosslinking-assisted m6A sequencing. Angewandte Chemie International Edition, 54, 1587–1590.
Wu, B. X., Su, S. C., Patil, D. P., Liu, H. H., et al. (2018). Molecular basis for the specific and multivariant recognitions of RNA substrates by human hnRNP A2/B1. Nature Communications, 9, 420
Huang, H., Weng, H., Sun, W., Qin, X., et al., (2018). Author Correction: Recognition of RNA N6-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nature Cell Biology, 20, 285–295.
Li, D. Y., Zhang, H. J., Hong, Y. B., Huang, L., et al. (2014). Genome-wide identification, biochemical characterization, and expression analyses of the YTH domain-containing RNA-binding protein family in Arabidopsis and rice. Plant Molecular Biology Reporter, 32, 1169–1186.
Reichel, M., Liao, Y., Rettel, M., Ragan, C., et al. (2016). In planta determination of the mRNA-binding proteome of Arabidopsis etiolated seedlings. The Plant Cell, 28, 2435–2452.
Arribas-Hernandez, L., Bressendorff, S., Hansen, M. H., Poulsen, C., et al. (2018). An m6A-YTH module controls developmental timing and morphogenesis in Arabidopsis. The Plant Cell, 30, 952–967.
Tirumuru, N., Zhao, B. S., Lu, W. X., Lu, Z. K., et al., (2016). N6-methyladenosine of HIV-1 RNA regulates viral infection and HIV-1 Gag protein expression. Elife, 5, e15528.
Nguyen, T. T., Ma, L. N., Slovak, M. L., Bangs, C. D., et al. (2006). Identification of novel Runx1 (AML1) translocation partner genes SH3D19, YTHDF2, and ZNF687 in acute myeloid leukemia. Genes Chromosomes Cancer, 45, 918–932.
Zhou, J., Wan, J., Gao, X., Zhang, X., et al. (2015). Dynamic m6A mRNA methylation directs translational control of heat shock response. Nature, 526, 591–594.
Wang, X., Sun, B., Jiang, Q. (2018). mRNA m6A plays opposite role in regulating UCP2 and PNPLA2 protein expression in adipocytes. International Journal of Obesity, 42, 1912–1924.
Wu, R., Yao, Y., Jiang, Q., Cai, M., et al. (2018). Epigallocatechin gallate targets FTO and inhibits adipogenesis in an mRNA m6A-YTHDF2-dependent manner. International Journal of Obesity (London), 42, 1378–1388.
Zhao, B. S., Wang, X., Beadell, A. V., Lu, Z., et al. (2017). m6A-dependent maternal mRNA clearance facilitates zebrafish maternal-to-zygotic transition. Nature, 542, 475–478.
Wei, L. H., Song, P. Z., Wang, Y., Lu, Z. K., et al. (2018). The m6A reader ECT2 controls trichome morphology by affecting mRNA stability in Arabidopsis. The Plant Cell, 30, 968–985.
Scutenaire, J., Deragon, J. M., Jean, V., Benhamed, M., et al. (2018). The YTH domain protein ECT2 is an m6A reader required for normal trichome branching in Arabidopsis. The Plant Cell, 30, 986–1005.
Hsu, P. J., Zhu, Y. F., Ma, H. H., Guo, Y. H., et al. (2017). YTHDC2 is an N6-methyladenosine binding protein that regulates mammalian spermatogenesis. Cell Research, 27, 1115–1127.
Jain, D., Puno, M. R., Meydan, C., Lailler, N., et al. (2018). ketu mutant mice uncover an essential meiotic function for the ancient RNA helicase YTHDC2. Elife, 7, e30919.
Drayman, N., Karin, O., Mayo, A., Danon, T., et al. (2017). Dynamic proteomics of herpes simplex virus infection. MBio. https://doi.org/10.1128/mBio.01612-17.
Ma, S., Menon, R., Poulos, R. C., & Wong, J. W. H. (2017). Proteogenomic analysis prioritises functional single nucleotide variants in cancer samples. Oncotarget, 8, 95841–95852.
Shichino, Y., Otsubo, Y., Kimori, Y., Yamamoto, M., et al. (2018). YTH-RNA-binding protein prevents deleterious expression of meiotic proteins by tethering their mRNAs to nuclear foci. Elife. https://doi.org/10.7554/eLife.32155.001.
Shi, H. L., Wang, X., Lu, Z. K., Zhao, B. X. S., et al. (2017). YTHDF3 facilitates translation and decay of N6-methyladenosine-modified RNA. Cell Research, 27, 315–328.
Hubstenberger, A., Courel, M., Benard, M., Souquere, S., et al. (2017). P-body purification reveals the condensation of repressed mRNA regulons. Molecular Cell, 68, 144–157 e5.
Li, A., Chen, Y. S., Ping, X. L., Yang, X., et al. (2017). Cytoplasmic m6A reader YTHDF3 promotes mRNA translation. Cell Research, 27, 444–447.
Zhang, Z. Y., Theler, D., Kaminska, K. H., Hiller, M., et al. (2010). The YTH domain is a novel RNA binding domain. Journal of Biological Chemistry, 285, 14701–14710.
Ke, S. D., Pandya-Jones, A., Saito, Y., Fak, J. J., et al. (2017). m6A mRNA modifications are deposited in nascent pre-mRNA and are not required for splicing but do specify cytoplasmic turnover. Genes & Development, 31, 990–1006.
Engreitz, J. M., Pandya-Jones, A., McDonel, P., Shishkin, A., et al. (2013). The Xist lncRNA exploits three-dimensional genome architecture to spread across the X chromosome. Science, 341,1237973.
Kretschmer, J., Rao, H., Hackert, P., Sloan, K. E., et al. (2018). The m6A reader protein YTHDC2 interacts with the small ribosomal subunit and the 5′-3′ exoribonuclease XRN1. RNA, 24, 1339–1350.
Tanabe, A., Konno, J., Tanikawa, K., & Sahara, H. (2014). Transcriptional machinery of TNF-alpha-inducible YTH domain containing 2 (YTHDC2) gene. Gene, 535, 24–32.
Hornbeck, P. V., Zhang, B., Murray, B., Kornhauser, J. M., et al. (2015). PhosphoSitePlus, 2014: mutations, PTMs and recalibrations. Nucleic Acids Research, 43, D512–D520.
Kato, M., Han, T. N. W., Xie, S. H., Shi, K., et al. (2012). Cell-free formation of RNA granules: low complexity sequence domains form dynamic fibers within hydrogels. Cell, 149, 753–767.
Thinon, E., Serwa, R. A., Broncel, M., Brannigan, J. A., et al. (2014). Global profiling of co- and post-translationally N-myristoylated proteomes in human cells. Nature Communications, 5, 4919.
Lichinchi, G., Zhao, B. S., Wu, Y. A., Lu, Z. K., et al. (2016). Dynamics of human and viral RNA methylation during zika virus infection. Cell Host & Microbe, 20, 666–673.
Gokhale, N. S., McIntyre, A. B. R., McFadden, M. J., Roder, A. E., et al. (2016). N6-methyladenosine in flaviviridae viral RNA genomes regulates infection. Cell Host & Microbe, 20, 654–665.
Dixit, D., Xie, Q., Rich, J. N., & Zhao, J. C. (2017). Messenger rna methylation regulates glioblastoma tumorigenesis. Cancer Cell, 31, 474–475.
Zhang, S. C., Zhao, B. S., Zhou, A. D., Lin, K. Y., et al. (2017). m6A demethylase ALKBH5 maintains tumorigenicity of glioblastoma stem-like cells by sustaining FOXM1 expression and cell proliferation program. Cancer Cell, 31, 591–606.
Kwok, C. T., Marshall, A. D., Rasko, J. E. J., & Wong, J. J. L. (2017). Genetic alterations of m6A regulators predict poorer survival in acute myeloid leukemia (vol 10, 39, 2017). Journal of Hematology & Oncology, 10, 39.
Canaani, D., Kahana, C., Lavi, S., & Groner, Y. (1979). Identification and mapping of N6-methyladenosine containing sequences in Simian Virus-40 RNA. Nucleic Acids Research, 6, 2879–2899.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant No. 31572413), the Natural Science Foundation of Zhejiang Province (No. LZ17C170001), the State Key Program of National Natural Science Foundation of China (No. 3163000269), and the Special Fund for Cultivation and Breeding of New Transgenic Organism (No. 2014ZX0800949B).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zhao, Y.L., Liu, Y.H., Wu, R.F. et al. Understanding m6A Function Through Uncovering the Diversity Roles of YTH Domain-Containing Proteins. Mol Biotechnol 61, 355–364 (2019). https://doi.org/10.1007/s12033-018-00149-z
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-018-00149-z