Abstract
Adrenocortical carcinoma (ACC) is an end‐stage hormonal syndrome. Although profound attempts have been made to illuminate the pathogenesis, the molecular mechanisms of ACC remain to be clarified. To identify the important genes in the progression of ACC, microarray datasets GSE19775 was downloaded from the gene expression omnibus database. The differentially-expressed genes (DEGs) were identified, and pathway and GO enrichment analyses were performed. The protein–protein interaction (PPI) network was constructed and the module analysis was performed using the protein interaction network analysis and Cytoscape. Also constructed target genes–miRNA regulatory network and target genes–TF regulatory network. Correlation of the hub genes were analyzed in The Cancer Genome Atlas. The prognostic values of hub genes were further validated by online tool UALCAN. Mutation analysis was done by online tool CBio Cancer Genomics Portal. A total of 884 DEGs were identified, with 441 in up regulation and 443 in down regulation. Pathways in catecholamine biosynthesis, aldosterone synthesis and secretion, pyrimidine deoxyribonucleosides salvage and systemic lupus erythematosus were the most significantly enriched for DEGs (up and down regulated). Blood vessel morphogenesis and cell cycle phase transition were the most significantly enriched term in biological processes, while extracellular matrix and chromosome, centromeric region were in cellular component and heparin binding and protein dimerization activity were in molecular function. Among the PPI networks and its module, target genes–miRNA regulatory network and target genes–TF regulatory network, hub genes were YWHAZ, FN1, GRK5, VCAM1, GATA6, TXNIP, HSPA1A, and F11R. Hub genes such as YWHAZ, STAT1, ICAM1, SH3BP5, CD83, FN1, TK1, HIST1H1C, CABLES1, and MCM3 were associated with poor overall survival, while hub genes such as STAT1, ICAM1, CD83, FN1, TK1, HIST1H1C, and MCM3 were highly expressed in stage 4. In conclusion, DEGs and hub genes diagnosed in this study may deepen our understanding of molecular mechanisms underlying the progression of ACC, and provide important targets for diagnosis and treatment of ACC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Adrenocortical carcinoma (ACC) is rare tumor cell form in the outer layer of the adrenal gland. 0.02–0.2% ACC related deaths were reported each year in the world [1]. 5-year survival rate of ACC patients was 16–47% [2]. Due to the inadequate diagnostic methods and tools, ACC is often diagnosed at an advanced stage. Despite the key improvements in surgical treatment, the 5-year survival rate for ACC patients diagnosed with metastatic is still low at approximately 33% [3]. Thus, it’s urgently needed for compassionate the molecular mechanisms of ACC progression and recognition of novel biomarkers and pathways are used for the early detection and prognosis assessment of ACC.
Over the past decades, advance of molecular biology have increased our compassionate of the pathogenesis of ACC. Earlier researches have indicated that ACC is a genetic disease, which depends on modification of several of oncogenes and tumor suppressor genes [4]. A growing number of genes and their coding proteins related to ACC have been examined. Earlier studies reported that they play key roles in a large number of physiological and pathological processes including cell proliferation [5], differentiation [6], apoptosis [7], and metastasis [8]. However, the precise molecular mechanisms of ACC are still far from being deep understood. Newly, several studies have identified a group of ACC related candidate biomarkers and pathways by microarray analysis [9]. Genes such as PDL1 [10], IGF and IGFBP [11], TARBP2 [12], IGFII and MIB1 [13], and TOP2A [14] were responsible for pathogenesis of ACC. Pathways such as Wnt/beta-catenin pathway [15], TGF-β pathway [16], and mTOR pathway [17] were important for progression of ACC. However, few studies have assessed the key genes associated in ACC using a bioinformatics approach. Accordingly, it was crucial to explore the molecular mechanisms in ACC and to detect novel and specific biomarkers, pathways, and targets.
In this study, we have downloaded GSE19775 from gene expression omnibus (GEO) database and utilized limma R bioconductor tool to identify the DEGs between ACC and normal whole adrenal gland. Additionally, other approaches including pathway enrichment analysis from different pathway databases, Gene Ontology (GO) enrichment analysis, protein–protein interaction (PPI) network construction, sub-modules analysis, target genes–miRNA, interaction construction, target genes–TF interaction construction, survival analysis, and validation of hub genes. The aims of our study were to identify novel biomarkers, pathways, and potential therapeutic targets for ACC.
Materials and methods
Data collection
The gene expression profile (GSE19775) included 15 ACC samples and 15 normal whole adrenal gland samples based on platform of GPL887Agilent-012097 Human 1A Microarray (V2) G4110B (Feature Number version) was downloaded from the GEO (http://www.ncbi.nlm.nih.gov/geo/) database, which was deposited by Demeure et al. [18].
Identification of DEGs
We utilized the “limma” R bioconductor package (R Software version 3.5.2 and R Packages limma 3.38.2) [19] to screen the DEGs between ACC samples and normal whole adrenal gland samples. The FDR (false discovery rate) P < 0.05 and |log2fold change (FC)| > 1.335 for up regulated genes and |log2fold change (FC)| < − 0.891 for down regulated genes were chosen as the cut-off criteria. FDR (false discovery rate) was applied to help correct false-positives.
Pathway enrichment analysis
BIOCYC (https://biocyc.org/) [20], Kyoto Encyclopedia of Genes and Genomes (KEGG) (http://www.genome.jp/kegg/pathway.html) [21], Pathway Interaction Database (PID) (https://wiki.nci.nih.gov/pages/viewpage.action?pageId=315491760) [22], REACTOME (https://reactome.org/) [23], GenMAPP (http://www.genmapp.org/) [24], MSigDB C2 BIOCARTA (v6.0) (http://software.broadinstitute.org/gsea/msigdb/collections.jsp) [25], PantherDB (http://www.pantherdb.org/) [26], Pathway Ontology (http://www.obofoundry.org/ontology/pw.html) [27], and Small Molecule Pathway Database (SMPDB) (http://smpdb.ca/) [28] were reference knowledge bases involving systems information, genomic information, and chemical information. To further explore the biological significance of DEGs, we used a online tool ToppCluster ToppCluster (https://toppcluster.cchmc.org/) [29] with the ability to analyze and visualize data to perform enrichment analysis of pathways. P value < 0.05 was considered a significant pathway enrichment.
Gene ontology (GO) enrichment analysis
Gene ontology (GO) (http://www.geneontology.org/) [30] is frequently used to annotate genes from high-throughput genome or transcriptome data. ToppCluster (https://toppcluster.cchmc.org/) [29] is an online functional annotation tool for multiple genes or proteins. P value < 0.05 was considered a significant GO enrichment.
PPI network and sub-modules analysis
Protein interaction network analysis (PINA) (http://omics.bjcancer.org/pina/home.do) [31] is an online software that contains comprehensive information of multiple proteins and is mainly used to evaluate the PPI information and this software integrates different PPI databases such as IntAct (https://www.ebi.ac.uk/intact/) [32], Molecular INTeraction Database (MINT, https://mint.bio.uniroma2.it/) [33], Biological General Repository for Interaction Datasets (BioGRID, https://thebiogrid.org/) [34], Database of Interacting Proteins (DIP, https://dip.doe-mbi.ucla.edu/dip/Main.cgi) [35], and Human Protein Reference Database (HPRD, http://www.hprd.org/) [36]. Cytoscape (http://www.cytoscape.org/) [37] version (3.7.1) was used to visualize the PPI networks of both up and down regulated genes. Confidence score ≥ 0.4 and maximum number of interactors = 0 were selected as the cut-off criteria. Five topological properties such as node degree [38], betweenness [39], stress [40], closeness [41], and clustering coefficient [42] were calculated in PPI networks. PEWCC1 [43] java plug-in of Cytoscape was utilized to illuminate the biological significance of gene modules in colon cancer with degree cut-off = 2, node score cut-off = 0.2, k-core = 2, and max. depth = 100.
Construction of target genes–miRNA regulatory network
Regulatory relationship between target genes (up and down regulated) and miRNA using online tool miRNet (https://www.mirnet.ca/) [44] which integrates 10 miRNA databases such as TarBase (http://diana.imis.athena-innovation.gr/DianaTools/index.php?r=tarbase/index) [45], miRTarBase (http://mirtarbase.mbc.nctu.edu.tw/php/download.php) [46], miRecords (http://miRecords.umn.edu/miRecords) [47], miR2Disease (http://www.mir2disease.org/) [48], HMDD (http://www.cuilab.cn/hmdd) [49], PhenomiR (http://mips.helmholtz-muenchen.de/phenomir/) [50], SM2miR (http://bioinfo.hrbmu.edu.cn/SM2miR/) [51], PharmacomiR (http://www.pharmaco-mir.org/) [52], EpimiR (http://bioinfo.hrbmu.edu.cn/EpimiR/) [53], and starBase (http://starbase.sysu.edu.cn/) [54]. Target genes–miRNA regulatory network was visualized by Cytoscape (http://www.cytoscape.org/) [37].
Construction of target genes–TF regulatory network
Regulatory relationship between target genes (up and down regulated) and TF using online tool NetworkAnalyst (https://www.networkanalyst.ca/) [55] online tool which integrates one TFs database ENCODE (https://www.encodeproject.org/pipelines/ENCPL138KID/) [56]. Target genes–TF regulatory network was visualized by Cytoscape (http://www.cytoscape.org/) [37].
Survival analysis and stage-related expression analysis of hub genes
UALCAN (http://ualcan.path.uab.edu/cgi-bin/ualcan-res.pl) [57] is an online tool for analyzing the gene expression profiles of different tumor stages and survival analysis from the Cancer Genome Atlas (TCGA). Thus, we could verify the expression levels of hub genes in all stages of ACC as well as survival analysis based on the high and low expression of hub genes in ACC. P value < 0.05 was considered as having statistical significance.
Genetical modification of hub genes
Online tool CBio Cancer Genomics Portal (http://www.cbioportal.org) [58] is an open strong platform that is use full for visualization, analysis, and downloads of large-scale cancer genomic dataset of ACC. This online tool is use full for researchers to explore and compare genetic alterations across samples.
Results
Identification of DEGs
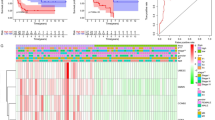
GSE19775 dataset (15 ACC samples and 15 normal whole adrenal gland samples) were first downloaded and preprocessed (normalized) is shown in Fig. 1a and b. After preprocessing and removing batch effects, we analyzed the DEGs of GSE19775 using the limma package, using FDR (false discovery rate) P < 0.05 and |log2fold change (FC)| > 1.335 for up regulated genes and |log2fold change (FC)| < − 0.891 for down regulated genes as the cut-off criterion. We screened 884 DEGs, including 441 up regulated genes and 443 down regulated genes in ACC samples compared to normal whole adrenal gland samples was shown in volcano plot (Fig. 2 and Table 1). The results of the cluster analysis of DEGs (up and down regulated) come to know that significant differences between the ACC samples and normal whole adrenal gland samples (Figs. 3 and 4).
Box plots of the expression profiles. A: Before normalization (red) and B: after normalization (yellow). (A1,A2, A3, A4, A5, A6, A7, A8, A9,A10, A11, A12, A13, A14, A15 = adrenocortical carcinoma samples, B1,B2, B3, B4, B5,B6, B7, B8, B9,B10, B11, B12, B13, B14, B15 = normal whole adrenal gland samples)
Heat map of gene expression (Upregulated genes). The color key represents the logFC of DEGs. FC, fold change (A1,A2, A3, A4, A5, A6, A7, A8, A9,A10, A11, A12, A13, A14, A15 = adrenocortical carcinoma samples, B1,B2, B3, B4, B5,B6, B7, B8, B9,B10, B11, B12, B13, B14, B15 = normal whole adrenal gland samples)
Heat map of gene expression (Down-regulated genes). The color key represents the logFC of DEGs. FC, fold change (A1,A2, A3, A4, A5, A6, A7, A8, A9,A10, A11, A12, A13, A14, A15 = adrenocortical carcinoma samples, B1,B2, B3, B4, B5,B6, B7, B8, B9,B10, B11, B12, B13, B14, B15 = normal whole adrenal gland samples)
Pathway enrichment analysis
Pathway enrichment analysis from different pathway databases (BIOCYC, KEGG, PID, REACTOME, GenMAPP, MSigDB C2 BIOCARTA, PantherDB, and SMPDB) showed that the up regulated genes were mainly associated with catecholamine biosynthesis, glucocorticoid biosynthesis, aldosterone synthesis and secretion, tyrosine metabolism, IL6-mediated signaling events, IL23-mediated signaling events, metallothioneins bind metals, metabolism of steroid hormones, steroid hormone metabolism, tyrosine metabolism, ensemble of genes encoding extracellular matrix and extracellular matrix-associated proteins, biosynthesis of neurotransmitters, Dopamine receptor mediated signaling pathway, adrenaline and noradrenaline biosynthesis, leptin system, insulin-like growth factor signaling, catecholamine biosynthesis, and steroidogenesis are listed in Table 2, mean while down regulated genes were mainly associated with pyrimidine deoxyribonucleosides salvage, glutamate removal from folates, systemic lupus erythematosus, FOXM1 transcription factor network, E2F transcription factor network, SIRT1 negatively regulates rRNA Expression, epigenetic regulation of gene expression, methionine metabolism, pyrimidine metabolism, CDK Regulation of DNA Replication, ensemble of genes encoding core extracellular matrix including ECM glycoproteins, collagens and proteoglycans, integrin signaling pathway, serine glycine biosynthesis, mismatch repair pathway, nucleotide excision repair, and adenosine deaminase deficiency are listed in Table 3.
Gene ontology (GO) enrichment analysis
Gene ontology (GO) enrichment analysis from all three GO categories [biological process (BP), cellular component (CC), and molecular function (MF)] showed that the up regulated genes were mainly associated with blood vessel morphogenesis, response to other organism, extracellular matrix, extracellular space, heparin binding, and G protein-coupled receptor binding are listed in Table 4, mean while down regulated genes were mainly associated with cell cycle phase transition, negative regulation of chromosome segregation, chromosome, centromeric region, spindle, protein dimerization activity, and protein heterodimerization activity are listed in Table 5.
PPI network and sub-modules analysis
We constructed a PPI network to further explore the interaction between the common DEGs (up and down regulated) by using PINA database and Cytoscape. A total of 5050 nodes and 9534 edges were included in the PPI networks (up regulated) (Fig. 5). The hub genes with highest node degree in this PPI network such as YWHAZ (degree = 805), HSPA5 (degree = 394), STAT3 (degree = 213), APOE (degree = 183), and GRK5 (degree = 175) are listed in Table 6. A total of 5146 nodes and 11,499 edges were included in the PPI networks (down regulated) (Fig. 6). The hub genes with highest node degree in this PPI network such as FN1 (degree = 1012), VCAM1 (degree = 671), UBD (degree = 664), BARD1 (degree = 252), and TK1 (degree = 169) are listed in Table 6. R square = 0.761 and correlation coefficient = 0.967 for node degree (up regulated) (Fig. 7a), meanwhile R square = 0.761 and correlation coefficient = 0.978 for node degree (down regulated) (Fig. 7b). The hub genes with highest betweenness in PPI network for up regulated genes such as YWHAZ (betweenness = 0.26018915), HSPA5 (betweenness = 0.10211544), STAT3 (betweenness = 0.05179592), TH (betweenness = 0.05146886), and CD83 (betweenness = 0.04395976) are listed in Table 6. R square = 0.602 and correlation coefficient = 0.105 for betweenness (Fig. 8a). The hub genes with highest stress in PPI network for up regulated genes such as YWHAZ (stress = 110811588), HSPA5 (stress = 43035116), ICAM1 (stress = 27529814), HOXB2 (stress = 23,027,092), and STAT3 (stress = 22,678,172) are listed in Table 6. R square = 0.001 and correlation coefficient = 0.111 for stress (Fig. 8b). The hub genes with highest closeness in PPI network for up regulated genes such as YWHAZ (closeness = 0.40211725), HSPA5 (closeness = 0.38225204), NR4A1 (closeness = 0.36144752), IRS1 (closeness = 0.35719883), and MAP3K5 (closeness = 0.35502372) are listed in Table 6. R square = 0.186 and correlation coefficient = 0.338 for closeness (Fig. 8c). The hub genes with lowest clustering coefficient in PPI network for up regulated genes such as CRLF1 (clustering coefficient = 0), CRH (clustering coefficient = 0), ARHGAP18 (clustering coefficient = 0), FJX1 (clustering coefficient = 0), and FMO3 (clustering coefficient = 0) are listed in Table 6. R square = 0.447 and correlation coefficient = 0.686 for clustering coefficient (Fig. 8d). The hub genes with highest betweenness in PPI network for down regulated genes such as FN1 (betweenness = 0.24710836), UBD (betweenness = 0.14892195), VCAM1 (betweenness = 0.11467148), BARD1 (betweenness = 0.05118512), and TK1 (betweenness = 0.04488624) are listed in Table 6. R square = 0.574 and correlation coefficient = 0.109 for betweenness (Fig. 9a). The hub genes with highest stress in PPI network for down regulated genes such as FN1 (stress = 358,695,218), UBD (stress = 286,343,980), VCAM1 (stress = 222,971,648), SLPI (stress = 46,566,718), and BARD1 (stress = 43,410,948) are listed in Table 6. R square = 0.011 and correlation coefficient = 0.090 for stress (Fig. 9b). The hub genes with highest closeness in PPI network for down regulated genes such as FN1 (closeness = 0.42073523), VCAM1 (closeness = 0.39252408), MCM3 (closeness = 0.39092723), PHGDH (closeness = 0.38138386), and MCM6 (closeness = 0.3806143) are listed in Table 6. R square = 0.198 and correlation coefficient = 0.293 for closeness (Fig. 9c). The hub genes with lowest clustering coefficient in PPI network for down regulated genes such as CYP3A5 (clustering coefficient = 0), POPDC2 (clustering coefficient = 0), PNLIPRP2 (clustering coefficient = 0), KIAA1407 (clustering coefficient = 0), and CX3CR1 (clustering coefficient = 0) are listed in Table 6. R square = 0.663 and correlation coefficient = 0.810 for clustering coefficient (Fig. 9d).
Subsequently, we performed module analysis of the whole network by the PEWCC1 plug-in. A total of 773 modules were obtained from the PPI network (up regulated). The four most significant modules were selected in this PPI network such as module 6, module 30, module 37, and module 55 (Fig. 10). Module 6 had 48 nodes and 119 edges. Hub genes in this modules such as YWHAZ (degree = 805), GRK5 (degree = 175), and NOLC1 (degree = 99). Module 30 had 10 nodes and 19 edges. Hub genes in this modules such as ID2 (degree = 46), STAT3 (degree = 213), HES1 (degree = 43), NR4A1 (degree = 116), ZNF331 (degree = 11), and YWHAZ (degree = 805). Module 37 had 8 nodes and 22 edges. Hub genes in this modules such as APOE (degree = 183), PLAT (degree = 34), C1S (degree = 12), SERPING1 (degree = 137), HP (degree = 160), APOC1 (degree = 151), and SH3BP5 (degree = 142). Module 55 had 7 nodes and 12 edges. Hub genes in this modules such as STAT1 (degree = 174), IPO7 (degree = 47), and HSPA4L (degree = 40). Meanwhile, total of 923 modules were obtained from the PPI network (down regulated). The four most significant modules were selected in this PPI network such as module 2, module 8, module 10, and module 27 (Fig. 11). Module 2 had 69 nodes and 239 edges. Hub genes in this modules such as VCAM1 (degree = 671), FN1 (degree = 1012), MCM3 (degree = 126), MCM2 (degree = 110), and MCM6 (degree = 100). Module 8 had 36 nodes and 74 edges. Hub genes in this modules such as UBD (degree = 664), MAGED2 (degree = 37), HIST1H1C (degree = 144), MAD2L1 (degree = 60), and PHGDH (degree = 76). Module 10 had 29 nodes and 89 edges. Hub genes in this modules such as HSPB1 (degree = 129), HIST1H2BJ (degree = 23), VCAM1 (degree = 671), UHRF1 (degree = 35), MCM3 (degree = 126), NCOR2 (degree = 140), HIST1H1C (degree = 144), TOP2A (degree = 110), FN1 (degree = 1012), PBK (degree = 91), ASNS (degree = 43), MCM2 (degree = 110), TMPO (degree = 58), MCM6 (degree = 100), HMGB2 (degree = 39), HIST1H2BL (degree = 21), TYMS (degree = 39), CHAF1B (degree = 26), and PHGDH (degree = 76). Module 27 had 17 nodes and 47 edges. Hub genes in this modules such as CCNE1 (degree = 84), POLD1 (degree = 67), CDT1 (degree = 55), and PRC1 (degree = 27).
Construction of target genes–miRNA regulatory network
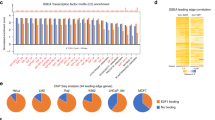
Target genes (up and down regulated) interacts with miRNA are shown Figs. 12 and 13. Top five up regulated targeted genes such as YWHAZ interacts with 212 miRNAs, GATA6 interacts with 207 miRNAs, LDLR interacts with 167 miRNAs, BZW1 interacts with 149 miRNAs, and IGFBP5 interacts with 144 miRNAs. Meanwhile, top five down regulated targeted genes such as TXNIP interacts with 228 miRNAs, MAPKAPK5 interacts with 179 miRNAs, PMAIP1 interacts with 147 miRNAs, RAD51 interacts with 147 miRNAs, and MICA interacts with 142 miRNAs (Table 7).
Construction of target genes–TF regulatory network
Target genes (up and down regulated) interacts with TF are shown Figs. 14 and 15. Top five up regulated targeted genes such as HSPA1A interacts with 67 TFs, PHOX2B interacts with 54 TFs, H19 interacts with 52 TFs, CEBPD interacts with 51 TFs, and DNAJC3 interacts with 50 TFs. Meanwhile, top five down regulated targeted genes such as F11R interacts with 91 TFs, CHAF1B interacts with 85 TFs, FANCG interacts with 82 TFs, CENPF interacts with 82 TFs, and XRCC1 interacts with 82 TFs (Table 8).
Survival analysis and stage-related expression analysis of hub genes
TCGA data of ACC patients are used via the UALCAN data portal. Kaplan–Meier curve for overall survival of TCGA patients with ACC is obtained according to the low and high expression of each gene. The results showed that patients in the high mRNA expression group for YWHAZ had worse overall survival than those in the low expression group (P = 0.012) (Fig. 16a), the high mRNA expression group for STAT1 had worse overall survival than those in the low expression group (P = 0.024) (Fig. 16b), the high mRNA expression group for ICAM1 had worse overall survival than those in the low expression group (P = 0.008) (Fig. 16c, a), the high mRNA expression group for SH3BP5 had worse overall survival than those in the low expression group (P = 0.016) (Fig. 16d), the high mRNA expression group for CD83 had worse overall survival than those in the low expression group (P = 0.00016) (Fig. 16e), the high mRNA expression group for FN1 had worse overall survival than those in the low expression group (P = 0.0069) (Fig. 16f), the high mRNA expression group for TK1 had worse overall survival than those in the low expression group (P = 0.0001) (Fig. 16g), the high mRNA expression group for HIST1H1C had worse overall survival than those in the low expression group (P = 0.0057) (Fig. 16h), the high mRNA expression group for CABLES1 had worse overall survival than those in the low expression group (P = 0.017) (Fig. 16i), and the high mRNA expression group for MCM3 had worse overall survival than those in the low expression group (P = 0.00011) (Fig. 16j). TCGA data analysis showed that hub genes such as YWHAZ, STAT1, ICAM1, CD83, FN1, TK1, HIST1H1C, and MCM3 were highly expressed in stages 4 (Fig. 17), while hub genes such as SH3BP5 and CABLES1 were highly expressed in stages 2 and stages 1 (Fig. 18).
Genetical alteration of the five hub genes
Five hub genes’ alteration statuses in TCGA bladder cancer patients were analyzed using the CbioPortal database. The five hub genes such as YWHAZ, STAT1, SH3BP5, TK1, and HIST1H1C altered in 0% patients, while five hub genes such as ICAM1, CD83, FN1, CABLES1, and MCM3 altered in 3% (missence mutation and amplification) and 1.1% (missence mutation and amplification) patients and the frequency of alteration of each hub gene are shown in Fig. 19.
Discussion
In recent years, although the morbidity of ACC has less, it is still malignant tumors worldwide. Specific biomarkers and effective targets are still needed to be diagnosed.
In this study, a total of 884 DEGs were picked out from the profile of GSE19775, including 441 up regulated genes and 443 down regulated genes. Gene such as phenylethanolamine N-methyltransferase (PNMT) [59] and cocaine- and amphetamine-regulated transcript (CART) [60] were associated with development of adrenal pheochromocytoma, but these genes may be responsible for progression of ACC. Methylation inactivation of tumor suppressor gene proenkephalin (PENK) was responsible for development of colorectal cancer [61], but inactivation of this gene may be identified with growth of ACC. Chromogranin A (parathyroid secretory protein 1) (CHGA) was important for progression of gastroenteropancreatic neuroendocrine tumors [62], but this gene may be diagnosed with development of ACC. Genes such as FGF23 [63] and NEK3 [64] were liable for invasion of prostate cancer cells, but these genes may be associated with invasion of ACC cells. Single nucleotide polymorphism (SNP) in SLC2A4RG was linked with development of glioblastoma [65], but SNP in this gene may be liable for advancement of ACC. Alteration in CHTF18 was linked with development of endometrial cancer [66], but mutation in this gene may be responsible for pathogenesis of ACC. HMGB2 was identified with development chemoresistance in gastric cancer [67], but this gene may be responsible for drug resistance in ACC. EZH2 was reasonable for pathogenesis of ACC [68].
In pathway enrichment analysis, up regulated genes were mainly associated with several pathways from different pathway databases (BIOCYC, KEGG, PID, REACTOME, GenMAPP, MSigDB C2 BIOCARTA, PantherDB, and SMPDB) such as catecholamine biosynthesis, aldosterone synthesis and secretion, IL6-mediated signaling events, metallothioneins bind metals, steroid hormone metabolism, ensemble of genes encoding extracellular matrix and extracellular matrix-associated proteins, dopamine receptor mediated signaling pathway, and leptin system. SNP in genes such as CYP11A1 [69], HSD3B1 [70], SCARB1 [71], lumican (LUM) [72], and IRS1 [73] were liable for progression of different cancers such as colorectal cancer, prostate cancer, thyroid cancer and breast cancer, but SNP in these genes may be involved in development of ACC. Gene such as CYP11B2 [74], HSD3B2 [75], STAT3 [76], CYP17A1 [77], CCN3 [78], IGFBP2 [79], IGFBP5 [80], IGFBP6 [80], and PRKAR2B [81] were diagnosed with growth of ACC. Genes such as MC2R [82], CCAAT/enhancer binding protein (C/EBP), beta (CEBPB) [83], and ANGPTL4 [84] were involved in proliferation of different cancer cells such as prostate cancer, breast cancer, and colorectal cancer, but these genes may be associated with proliferation of ACC cells. Gene such as NR4A1 [85], NR4A2 [86], PRKD3 [87], ADAMTS1 [88], CCL13 [89], CRLF1 [90], CXCL12 [91], CXCL2 [92], decorin (DCN) [93], FGF9 [94], GDF15 [95], plasminogen activator, tissue (PLAT) [96], S100A8 [97], S100A9 [97], SEMA6A [98], SRPX2 [99], and tenascin XB (TNXB) [100] were important for invasion of different cancer cells such as breast cancer, colorectal cancer, prostate cancer, thyroid cancer, ovarian cancer, endometrial cancer, gastric cancer, glioblastoma, and mesothelioma, but these genes may be linked with invasion of ACC cells. IL6 was responsible for inflammation in breast cancer [101], but this gene may be linked with inflammation in ACC. High expression of jun B proto-oncogene (JUNB) [102], STAT1 [103], MT1E [104], FGF7 [105], fibromodulin (FMOD) [106], SPON2 [107], and THBS1 [108] were important for advancement of different cancers such as renal cancer, colon cancer, breast cancer, prostate cancer and esophageal squamous cell carcinoma, but high expression of these genes may be identified with growth of ACC. Methylation inactivation of tumor suppressor genes such as SOCS3 [109], MT1G [110], MT1H [111], ADAMTS9 [112], ANGPTL1 [113], FBLN1 [114], FCN2 [115], SEMA3B [116], and TIMP4 [117] were diagnosed with growth of different cancers such as prostate cancer, gastric cancer hepatocellular carcinoma, and lung cancer, but loss of these genes may be responsible for pathogenesis of ACC. Decrease expression of tumor suppressor gene MT1F was liable for progression of hepatocellular carcinoma [118], but less expression of this gene may be identified with growth of ACC. Genes such as CCL2 [119], CXCL1 [120], and EFEMP1 [121] were associated with angiogenesis in different cancers such as breast cancer and colorectal cancer, but these genes may be linked with angiogenesis in ACC. Tyrosine hydroxylase (TH) [122] and neuropeptide Y (NPY) [123] were responsible for pathogenesis of pheochromocytoma, but these genes may be associated with development of ACC. Dopa decarboxylase (aromatic l-amino acid decarboxylase) (DDC), CYP21A2, KCNK3, low density lipoprotein receptor (LDLR), CCAAT/enhancer binding protein (C/EBP), delta (CEBPD), MT1A, MT1B, MT1X, CTSD, FBLN5, FGF12, MFAP4, SERPING1, SPOCK3, dopamine beta-hydroxylase (dopamine beta-monooxygenase) (DBH) EPB41L1 and guanine nucleotide binding protein (G protein), alpha z polypeptide (GNAZ) were novel biomarkers for pathogenesis of ACC in these pathways. Meanwhile down regulated genes were mainly associated with several pathways from different pathway databases such as pyrimidine deoxyribonucleosides salvage, systemic lupus erythematosus, FOXM1 transcription factor network, SIRT1 negatively regulates rRNA expression, methionine metabolism, CDK regulation of DNA replication, integrin signaling pathway, mismatch repair pathway, and adenosine deaminase deficiency. High expression of genes such as TK1 [124], thymidylatesynthetase (TYMS) [125], CCNE1 [126], CDT1 [127], FN1 [128], NTN4 [129], and adenosine deaminase (ADA) [130] were associated with pathogenesis of different cancer such as breast cancer, colorectal cancer, ovarian cancer, and prostate cancer, but elevated expression of these genes may be liable for development of ACC. Genes such as HLA-DQB1 [131], MCM2 [132], and MSH2 [133] were involved in pathogenesis of ACC. Metylation inactivation of tumor suppressor gene DNMT1 was responsible for progression of colon cancer [134], but loss of this gene may be associated with pathogenesis of ACC. Genes such as MCM3 [135] and MCM6 [136] were associated with proliferation of different cancer cells such as ovarian cancer and lung cancer, but these genes may be liable for advancement of ACC. Genes such as COL1A1 [137], COL3A1 [138], LAMA3 [139], and LAMB1 [140] were linked with invasion of different cancer cells such as gastric cancer, bladder cancer, head and neck cancers, and colorectal cancer, but these genes may be diagnosed with growth of ACC. Mutation in genes such as EXO1 [141] and POLD1 [142] were associated with pathogenesis of colorectal cancer, but alteration in this gene may be identified with growth of ACC. SNP in LIG1 was involved in progression of lung cancer [143], but this gene may be responsible for development of ACC. HIST1H2BB, HIST1H2BC, HIST1H2BD, HIST1H2BE, HIST1H2BF, HIST1H2BH, HIST1H2BI, HIST1H2BJ, HIST1H2BL, HIST1H2BN, HIST1H2BO, HIST1H4F, HIST1H4H, HIST1H4J, HIST1H4K, HLA-DRB4, MAT1A, COL14A1, COL27A1, COL4A5, COL5A2, RFC4, and RFC5 were novel biomarkers for pathogenesis of ACC in these pathways.
In GO enrichment analysis, up regulated genes were mainly associated with all GO categories (BP, CC and MF) such as blood vessel morphogenesis, extracellular matrix, and heparin binding. Genes such as adrenomedullin (ADM) [144], apolipoprotein E (APOE) [145], GATA6 [146], and RARRES2 [147] were responsible for pathogenesis of ACC. Genes such as CDH2 [148], CITED1 [149], FZD4 [150], STARD13 [151], XBP1 [152], and latexin (LXN) [153] were associated with invasion of different cancer cells such as lung cancer, colorectal cancer, prostate cancer, breast cancer, and thyroid cancer, but these genes may be liable for invasion of ACC cells. Methylation inactivation of tumor suppressor genes such as DLL1 [154], HAND2 [155], and UNC5B [156] were identified with progression of different cancer such as lung cancer endometrial cancer and colorectal cancer, but loss of these genes may be involved in development of ACC. High expression of genes such as ENPP2 [157], GJA1 [158], HES1 [159], RAMP1 [160], SIRT1 [161], TFAP2B [162], TNFRSF11B [163], and SAA1 [164] were important for advancement of different cancer such as hepatocellular carcinoma, endometrial cancer, colon cancer, prostate cancer, lung cancer, and gastric cancer, but elevated expression of these genes may be diagnosed with ACC. Genes such as HOXA5 [165] and HOXA7 [166] were linked with cell cycle progression in different cancer such as breast cancer and ovarian cancer, but these genes may be involved in cell cycle progression in ACC. SNP in NRXN3 was important for development of breast cancer [167], but this polymorphic gene may be involved in development of ACC. Low expression of TGFBR3 was responsible for advancement of prostate cancer [168], but loss of this gene may associated with growth of ACC. Platelet-derived growth factor receptor, alpha polypeptide (PDGFRA), SCG2, TCDD-inducible poly(ADP-ribose) polymerase (TIPARP), and IL1RL1 were novel biomarkers for pathogenesis of ACC in these GO categories. Meanwhile, down regulated genes were mainly associated with all GO categories such as cell cycle phase transition, chromosome, centromeric region, and protein dimerization activity. High expression of genes such as anillin, actin binding protein (ANLN) [169], NEK2 [170], timeless homolog (TIMELESS) [171], TPX2 [172], TTK protein kinase (TTK) [173], CDCA8 [174], KIF18A [175], OIP5 [176], SPC25 [177], ZWINT [178], asparagine synthetase (ASNS) [179], E2F8 [180], LCN2 [181], and RAB11FIP4 [182] were important for progression of different cancers such as pancreatic cancer, breast cancer, lung cancer, colorectal cancer, gastric cancer, ovarian cancer, and hepatocellular carcinoma, but increased expression of these genes may be associated with development of ACC. Genes such as BIRC5 [183], BUB1 [184], CCNB1 [185], CCNB2 [186], CDCA5 [187], CDKN3 [188], centromere protein F (CENPF) [189], and SPAG5 [190] were associated with proliferation of different cancer cells such as breast cancer, gastric cancer, hepatocellular carcinoma, and ovarian cancer, but these genes may be linked with proliferation of ACC cells. Methylation inactivation of tumor suppressor genes such as CABLES1 [191], GADD45A [192], PLK2 [193], and NR3C1 [194] were involved in pathogenesis of different cancers such as ovarian cancer, breast cancer, and colorectal cancer, but loss these genes may be identified with development of ACC. SNP in genes such as hyaluronan-mediated motility receptor (HMMR) [195]. DNMT3B [196], aryl hydrocarbon receptor (AHR) [197] hydroxyprostaglandin dehydrogenase 15-(NAD) (HPGD) [198], and UGT1A6 [199] were liable for advancement of different cancer such as breast cancer, lung cancer, and colorectal cancer, but these polymorphic genes may be linked with development of ACC. Alteration in genes such as MAD2L1 [200] and BARD1 [201] were associated with pathogenesis of breast cancer, but mutation in this gene may be involved in development of ACC. Genes such as maternal embryonic leucine zipper kinase (MELK) [202], RRM2 [203], NR0B1 [204], and TOP2A [14] were important for pathogenesis of ACC. Alteration in tumor suppressor gene PLAGL1 was liable for growth of pheochromocytomas [205], but mutation in this gene may be associated with pathogenesis of ACC. CEP192, KNTC1, MCM8, KIF22, NUP107, STAG3, BTBD11, CD2, CD3D, DPP4, fibroblast activation protein, alpha (FAP) granzyme A (granzyme 1, cytotoxic T-lymphocyte-associated serine esterase 3) (GZMA), H2BFS, hematopoietically expressed homeobox (HHEX), HOMER1, v-kit Hardy-Zuckerman 4 feline sarcoma viral oncogene homolog (KIT), NCOA6, SRGAP2, and TCFL5 were novel biomarkers for pathogenesis of ACC in these GO categories.
In this study, we finally identify five hub genes (up regulated) were in PPI network with high node degree such as YWHAZ, HSPA5, STAT3, APOE, and GRK5. High expression of tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, zeta polypeptide (YWHAZ) was responsible for progression of head and neck cancer [206], but elevated expression of this gene may be identified with growth of ACC. Genes such as HSPA5 [207] and GRK5 [208] were associated with invasion of different cancer cells such as colorectal cancer and prostate cancer, but these genes may be liable for invasion of ACC cells. Five hub genes (up regulated) were in PPI network with high betweenness such as YWHAZ, HSPA5, STAT3, TH, and CD83. SNP in CD83 was diagnosed with growth of cervical cancer [209], but this polymorphic gene may be involved in development of ACC. Five hub genes (up regulated) were in PPI network with high stress such as YWHAZ, HSPA5, ICAM1, HOXB2, and STAT3. ICAM1 was linked with invasion of breast cancer cells [210], but this gene may be associated with invasion of ACC cells. High expression of HOXB2 was involved in pathogenesis of cervical cancer [211], but elevated expression of this gene may be responsible for progression of ACC. Five hub genes (up regulated) were in PPI network with high closeness such as YWHAZ, HSPA5, NR4A1, IRS1, and MAP3K5. MAP3K5 was important for invasion of prostate cancer cells [212], but this gene may be linked with invasion of ACC cells. Five hub genes (up regulated) were in PPI network with low clustering coefficient such as CRLF1, CRH, ARHGAP18, FJX1, and FMO3. Corticotropin releasing hormone (CRH) was liable for progression of pheochromocytomas [213], but this gene may be associated with advancement of ACC. ARHGAP18 was identified with invasion of breast cancer cells [214], but this gene may be involved in invasion of ACC cells. High expression of FJX1 was diagnosed with growth of nasopharyngeal carcinoma [215], but over expression of this gene may be identified with progression of ACC. SNP in FMO3 was responsible for advancement of colorectal cancer [216], but this polymorphic gene may be associated with development of ACC. Meanwhile, five hub genes (down regulated) were in PPI network with high node degree such as FN1, VCAM1, UBD, BARD1, and TK1. High expression of genes such as VCAM1 [217] and ubiquitin D (UBD) [218] were responsible for pathogenesis of different cancers such as gastric cancer and colon cancer, but elevated expression of these genes may be liable for growth of ACC. Five hub genes (down regulated) were in PPI network with high betweenness such as FN1, UBD, VCAM1, BARD1, and TK1. Five hub genes (down regulated) were in PPI network with high stress such as FN1, UBD, VCAM1, SLPI, and BARD1. Secretary leukocyte protease inhibitor (SLPI) was linked with invasion of ovarian cancer cells [219], but this gene may be involved in invasion of ACC cells. Five hub genes (down regulated) were in PPI network with high closeness such as FN1, VCAM1, MCM3, PHGDH, and MCM6. High expression phosphoglycerate dehydrogenase (PHGDH) was responsible for pathogenesis of melanoma [220], but over expression of this gene may be identified with growth of ACC. Five hub genes (down regulated) were in PPI network with low clustering coefficient such as CYP3A5, POPDC2, PNLIPRP2, KIAA1407, and CX3CR1. SNP in CYP3A5 was important for pathogenesis of breast cancer [221], but this polymorphic gene may associated with development of ACC. CX3CR1 was linked with invasion of prostate cancer cells [222], but this gene may be involved in invasion of ACC cells. POPDC2, PNLIPRP2, and KIAA1407 were novel biomarkers for pathogenesis of ACC in these PPI network.
Hub genes (up regulated) such as YWHAZ, GRK5, NOLC1, ID2, STAT3, HES1, NR4A1, ZNF331, APOE, PLAT, C1S, SERPING1, HP, APOC1, SH3BP5, STAT1, IPO7, and HSPA4L were in all four modules. NOLC1 was associated with proliferation of hepatocellular carcinoma cell [223], but this gene may be responsible for progression of ACC cells. High expression of genes such as ID2 [224], APOC1 [225], and HSPA4L [226] were involved in pathogenesis of different cancer such as pancreatic cancer, lung cancer, and hepatocellular carcinoma, but increase expression these genes may be linked with growth of ACC. Methylation inactivation of tumor suppressor gene ZNF331 was important for development of colorectal cancer [227], but loss of this gene may be liable for growth of ACC. SNP in haptoglobin (HP) was associated with pathogenesis of breast cancer [228], but this polymorphic gene may be liable for growth of ACC. IPO7 was involved in proliferation of prostate cancer cells [229], but this gene may be associated with proliferation of ACC cells. C1S and SH3BP5 were novel biomarkers for pathogenesis of ACC in these modules. Meanwhile, hub genes (down regulated) such as VCAM1, FN1, MCM3, MCM2, MCM6, UBD, MAGED2, HIST1H1C, MAD2L1, PHGDH, HSPB1, HIST1H2BJ, UHRF1, NCOR2, TOP2A, PBK, ASNS, TMPO, HMGB2, HIST1H2BL, TYMS, CHAF1B, CCNE1, POLD1, CDT1, and PRC1 were in all four modules. Methylation inactivation of tumor suppressor gene HSPB1 was responsible for progression of prostate cancer [230], but loss of this gene may be identified with pathogenesis of ACC. High expression of genes such as UHRF1 [231] and PDZ binding kinase (PBK) [232] were important for advancement of different cancer such as lung cancer and prostate cancer, but elevated expression of these genes may be associated with development of ACC. NCOR2 was responsible for proliferation of glioblastoma cells [233], but this gene may be linked with proliferation of ACC cells. MAGED2 HIST1H1C, thymopoietin (TMPO), CHAF1B, and PRC1 were novel biomarkers for pathogenesis of ACC in these modules.
Target genes (up regulated) such as YWHAZ, GATA6, LDLR, BZW1, and IGFBP5 were in target genes–miRNA regulatory network. BZW1 was novel biomarker for pathogenesis of ACC in this network. Meanwhile, target genes (down regulated) such as TXNIP, MAPKAPK5, PMAIP1, RAD51, and MICA were in target genes–miRNA regulatory network. Decrease expression of tumor suppressor genes such as thioredoxin interacting protein (TXNIP) [234] and PMAIP1 [235] was diagnosed with growth of different cancer types such as thyroid cancer and pancreatic cancer, but loss of expression of these genes may liable for pathogenesis of ACC. High expression of RAD51 was involved in pathogenesis of breast cancer [236], but elevated expression of this gene may be linked with growth of ACC. MAPKAPK5 and MHC class I polypeptide-related sequence A (MICA) were novel biomarkers for pathogenesis of ACC in this network.
Target genes (up regulated) such as HSPA1A, PHOX2B, H19, CEBPD, and DNAJC3 were in target genes–TF regulatory network. HSPA1A was associated with pathogenesis of colorectal cancer [237], but this gene may be important for development of ACC. Inactivation of H19 was involved in pathogenesis of ACC [238]. PHOX2B and DNAJC3 were novel biomarkers for pathogenesis of ACC in this network. Meanwhile, target genes (down regulated) such as F11R, CHAF1B, FANCG, CENPF, and XRCC1 were in target genes–TF regulatory network. F11R was linked with advancement of glioma [239], but this gene may be associated with development of ACC. SNP in genes such as fanconianemia, complementation group G (FANCG) [240], and XRCC1 [241] were liable for progression of different cancers such as pancreatic cancer and breast cancer, but these genes may be liable for growth of ACC.
Survival analysis revealed that genes such as YWHAZ, STAT1, ICAM1, SH3BP5, CD83, FN1, TK1, HIST1H1C, CABLES1, and MCM3 were predicting shorter survival of ACC. Expression levels revealed that hub genes such as STAT1, ICAM1, CD83, FN1, TK1, HIST1H1C, and MCM3 were highly expressed in ACC (stages 4), meanwhile those such as SH3BP5 and CABLES1 were highly expressed in ACC (stage 2 and stage 1).
In conclusion, we diagnosed several hub genes and pathways that were closely related to the initiation and progression of ACC using a series of bioinformatics analyses of DEGs in cancer and normal samples. Identification of the associated genes, YWHAZ, FN1, GRK5, VCAM1, GATA6, TXNIP, HSPA1A, and F11R, provided greater insight into the specific molecular mechanisms underlying ACC occurrence and development, especially in terms of the pathways involved in catecholamine biosynthesis, aldosterone synthesis and secretion, pyrimidine deoxyribonucleosides salvage, and systemic lupus erythematosus. Upon further experimental validation of these results, these genes may serve as potential research targets for therapy and thereby contribute to greater understanding of the molecular mechanisms underlying the progression of ACC.
Availability of data and materials
The datasets supporting the conclusions of this article are available in the GEO (Gene Expression Omnibus) (https://www.ncbi.nlm.nih.gov/geo/) repository. [(GSE19775) (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc= GSE19775)].
References
Calissendorff J, Calissendorff F, Falhammar H. Adrenocortical cancer: mortality, hormone secretion, proliferation and urine steroids—experience from a single centre spanning three decades. BMC Endocr Disord. 2016;16:15. https://doi.org/10.1186/s12902-016-0095-9.
Wajchenberg BL, Albergaria Pereira MA, Medonca BB, Latronico AC, Campos Carneiro P, Alves VA, Zerbini MC, Liberman B, Carlos Gomes G, et al. Adrenocortical carcinoma: clinical and laboratory observations. Cancer. 2000;88(4):711–36.
Icard P, Louvel A, Chapuis Y. Survival rates and prognostic factors in adrenocortical carcinoma. World J Surg. 1992;16(4):753–8.
Gara SK, Lack J, Zhang L, Harris E, Cam M, Kebebew E. Metastatic adrenocortical carcinoma displays higher mutation rate and tumor heterogeneity than primary tumors. Nat Commun. 2018;9(1):4172. https://doi.org/10.1038/s41467-018-06366-z.
Doghman M, Cazareth J, Lalli E. The T cell factor/beta-catenin antagonist PKF115-584 inhibits proliferation of adrenocortical carcinoma cells. J Clin Endocrinol Metab. 2008;93(8):3222–5. https://doi.org/10.1210/jc.2008-0247.
Miettinen M. Neuroendocrine differentiation in adrenocortical carcinoma. New immunohistochemical findings supported by electron microscopy. Lab Investig. 1992;66(2):169–74.
Bernini GP, Moretti A, Viacava P, Bonadio AG, Iacconi P, Miccoli P, Salvetti A. Apoptosis control and proliferation marker in human normal and neoplastic adrenocortical tissues. Br J Cancer. 2002;86(10):1561–5. https://doi.org/10.1038/sj.bjc.6600287.
Assié G, Antoni G, Tissier F, Caillou B, Abiven G, Gicquel C, Leboulleux S, Travagli JP, Dromain C, Bertagna X, et al. Prognostic parameters of metastatic adrenocortical carcinoma. J Clin Endocrinol Metab. 2007;92(1):148–54. https://doi.org/10.1210/jc.2006-0706.
Fernandez-Ranvier GG, Weng J, Yeh RF, Khanafshar E, Suh I, Barker C, Duh QY, Clark OH, Kebebew E. Identification of biomarkers of adrenocortical carcinoma using genomewide gene expression profiling. Arch Surg. 2008;143(9):841–6. https://doi.org/10.1001/archsurg.143.9.841.
Fay AP, Signoretti S, Callea M, Telό GH, McKay RR, Song J, Carvo I, Lampron ME, Kaymakcalan MD, Poli-de-Figueiredo CE, et al. Programmed death ligand-1 expression in adrenocortical carcinoma: an exploratory biomarker study. J Immunother Cancer. 2015;3:3. https://doi.org/10.1186/s40425-015-0047-3.
Patel D, Ellis R, Howard B, Boufraqech M, Gara SK, Zhang L, Quezado MM, Nilubol N, Kebebew E. Analysis of IGF and IGFBP as prognostic serum biomarkers for adrenocortical carcinoma. Ann Surg Oncol. 2014;21(11):3541–7. https://doi.org/10.1245/s10434-014-3768-5.
Caramuta S, Lee L, Ozata DM, Akçakaya P, Xie H, Höög A, Zedenius J, Bäckdahl M, Larsson C, Lui WO. Clinical and functional impact of TARBP2 over-expression in adrenocortical carcinoma. Endocr Relat Cancer. 2013;20(4):551–64. https://doi.org/10.1530/ERC-13-0098.
Schmitt A, Saremaslani P, Schmid S, Rousson V, Montani M, Schmid DM, Heitz PU, Komminoth P, Perren A. IGFII and MIB1 immunohistochemistry is helpful for the differentiation of benign from malignant adrenocortical tumours. Histopathology. 2006;49(3):298–307. https://doi.org/10.1111/j.1365-2559.2006.02505.x.
Jain M, Zhang L, He M, Zhang YQ, Shen M, Kebebew E. TOP2A is overexpressed and is a therapeutic target for adrenocortical carcinoma. Endocr Relat Cancer. 2013;20(3):361–70. https://doi.org/10.1530/ERC-12-0403.
El Wakil A, Lalli E. The Wnt/beta-catenin pathway in adrenocortical development and cancer. Mol Cell Endocrinol. 2011;332(1–2):32–7. https://doi.org/10.1016/j.mce.2010.11.014.
Parviainen H, Schrade A, Kiiveri S, Prunskaite-Hyyryläinen R, Haglund C, Vainio S, Wilson DB, Arola J, Heikinheimo M, et al. Expression of Wnt and TGF-β pathway components and key adrenal transcription factors in adrenocortical tumors: association to carcinoma aggressiveness. Pathol Res Pract. 2013;209(8):503–9. https://doi.org/10.1016/j.prp.2013.06.002.
De Martino MC, van Koetsveld PM, Pivonello R, Hofland LJ. Role of the mTOR pathway in normal and tumoral adrenal cells. Neuroendocrinology. 2010;92(Suppl 1):28–34. https://doi.org/10.1159/000314280.
Demeure MJ, Coan KE, Grant CS, Komorowski RA, Stephan E, Sinari S, Mount D, Bussey KJ. PTTG1 overexpression in adrenocortical cancer is associated with poor survival and represents a potential therapeutic target. Surgery. 2013;154(6):1405–16. https://doi.org/10.1016/j.surg.2013.06.058.
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43(7):e47. https://doi.org/10.1093/nar/gkv007.
Caspi R, Billington R, Ferrer L, Foerster H, Fulcher CA, Keseler IM, Kothari A, Krummenacker M, Latendresse M, Mueller LA, et al. The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of pathway/genome databases. Nucleic Acids Res. 2016;44(D1):D471–80. https://doi.org/10.1093/nar/gkv1164.
Tanabe M, Kanehisa M. Using the KEGG database resource, Chapter 1: Unit 1.12. Curr Protoc Bioinform. 2012. https://doi.org/10.1002/0471250953.bi0112s38.
Schaefer CF, Anthony K, Krupa S, Buchoff J, Day M, Hannay T, Buetow KH. PID: the pathway interaction database. Nucleic Acids Res. 2009;37(Database issue):D674–9. https://doi.org/10.1093/nar/gkn653.
Sidiropoulos K, Viteri G, Sevilla C, Jupe S, Webber M, Orlic-Milacic M, Jassal B, May B, Shamovsky V, Duenas C, et al. Reactome enhanced pathway visualization. Bioinformatics. 2017;33(21):3461–7. https://doi.org/10.1093/bioinformatics/btx441.
Dahlquist KD, Salomonis N, Vranizan K, Lawlor SC, Conklin BR. GenMAPP, a new tool for viewing and analyzing microarray data on biological pathways. Nat Genet. 2002;31(1):19–20. https://doi.org/10.1038/ng0502-19.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102(43):15545–50. https://doi.org/10.1073/pnas.0506580102.
Mi H, Huang X, Muruganujan A, Tang H, Mills C, Kang D, Thomas PD. PANTHER version 11: expanded annotation data from Gene Ontology and Reactome pathways, and data analysis tool enhancements. Nucleic Acids Res. 2017;45(D1):D183–9. https://doi.org/10.1093/nar/gkw1138.
Petri V, Jayaraman P, Tutaj M, Hayman GT, Smith JR, De Pons J, Laulederkind SJ, Lowry TF, Nigam R, Wang SJ, et al. The pathway ontology—updates and applications. J Biomed Semant. 2014;5(1):7. https://doi.org/10.1186/2041-1480-5-7.
Jewison T, Su Y, Disfany FM, Liang Y, Knox C, Maciejewski A, Poelzer J, Huynh J, Zhou Y, Arndt D, et al. SMPDB 2.0: big improvements to the Small Molecule Pathway Database. Nucleic Acids Res. 2014;42(Database issue):D478–84. https://doi.org/10.1093/nar/gkt1067.
Kaimal V, Bardes EE, Tabar SC, Jegga AG, Aronow BJ. ToppCluster: a multiple gene list feature analyzer for comparative enrichment clustering and network-based dissection of biological systems. Nucleic Acids Res. 2010;38(Web Server issue):W96–102. https://doi.org/10.1093/nar/gkq418.
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet. 2000;25(1):25–9. https://doi.org/10.1038/75556.
Cowley MJ, Pinese M, Kassahn KS, Waddell N, Pearson JV, Grimmond SM, Biankin AV, Hautaniemi S, Wu J. PINA v2.0: mining interactome modules. Nucleic Acids Res. 2012;40(Database issue):D862–5. https://doi.org/10.1093/nar/gkr967.
Orchard S, Ammari M, Aranda B, Breuza L, Briganti L, Broackes-Carter F, Campbell NH, Chavali G, Chen C, del-Toro N, et al. The MIntAct project–IntAct as a common curation platform for 11 molecular interaction databases. Nucleic Acids Res. 2014;42(Database issue):D358–63. https://doi.org/10.1093/nar/gkt1115.
Licata L, Briganti L, Peluso D, Perfetto L, Iannuccelli M, Galeota E, Sacco F, Palma A, Nardozza AP, Santonico E, et al. MINT, the molecular interaction database: 2012 update. Nucleic Acids Res. 2012;40(Database issue):D857–61. https://doi.org/10.1093/nar/gkr930.
Oughtred R, Stark C, Breitkreutz BJ, Rust J, Boucher L, Chang C, Kolas N, O’Donnell L, Leung G, McAdam R, et al. The BioGRID interaction database: 2019 update. Nucleic Acids Res. 2019;47(D1):D529–41. https://doi.org/10.1093/nar/gky1079.
Salwinski L, Miller CS, Smith AJ, Pettit FK, Bowie JU, Eisenberg D. The Database of Interacting Proteins: 2004 update. Nucleic Acids Res. 2004;32(Database issue):D449–51. https://doi.org/10.1093/nar/gkh086.
Keshava Prasad TS, Goel R, Kandasamy K, Keerthikumar S, Kumar S, Mathivanan S, Telikicherla D, Raju R, Shafreen B, Venugopal A, Balakrishnan L, et al. Human Protein Reference Database–2009 update. Nucleic Acids Res. 2009;37(Database issue):D767–72. https://doi.org/10.1093/nar/gkn892.
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;3:2498–504. https://doi.org/10.1101/gr.1239303.
Aittokallio T, Schwikowski B. Graph-based methods for analysing networks in cell biology. Brief Bioinform. 2006;7(3):243–55. https://doi.org/10.1093/bib/bbl022.
Ashtiani M, Salehzadeh-Yazdi A, Razaghi-Moghadam Z, Hennig H, Wolkenhauer O, Mirzaie M, Jafari M. A systematic survey of centrality measures for protein-protein interaction networks. BMC Syst Biol. 2018;12(1):80. https://doi.org/10.1186/s12918-018-0598-2.
Shi Z, Zhang B. Fast network centrality analysis using GPUs. BMC Bioinform. 2011;12:149. https://doi.org/10.1186/1471-2105-12-149.
Dopazo J. Erten C Graph-theoretical comparison of normal and tumor networks in identifying BRCA genes. BMC Syst Biol. 2017;11(1):110. https://doi.org/10.1186/s12918-017-0495-0.
Li M, Lu Y, Niu Z, Wu FX. United complex centrality for identification of essential proteins from PPI networks. IEEE/ACM Trans Comput Biol Bioinform. 2017;14(2):370–80. https://doi.org/10.1109/TCBB.2015.2394487.
Zaki N, Efimov D, Berengueres J. Protein complex detection using interaction reliability assessment and weighted clustering coefficient. BMC. Bioinform. 2013;14:163. https://doi.org/10.1186/1471-2105-14-163.
Fan Y, Xia J. miRNet-functional analysis and visual exploration of miRNA-target interactions in a network context. Methods Mol Biol. 2018;1819:215–33. https://doi.org/10.1007/978-1-4939-8618-7_10.
Vlachos IS, Paraskevopoulou MD, Karagkouni D, Georgakilas G, Vergoulis T, Kanellos I, Anastasopoulos IL, Maniou S, Karathanou K, Kalfakakou D, et al. DIANA-TarBase v7.0: indexing more than half a million experimentally supported miRNA:mRNA interactions. Nucleic Acids Res. 2015;43(Database issue):D153–9. https://doi.org/10.1093/nar/gku1215.
Chou CH, Shrestha S, Yang CD, Chang NW, Lin YL, Liao KW, Huang WC, Sun TH, Tu SJ, Lee WH, et al. miRTarBase update 2018: a resource for experimentally validated microRNA-target interactions. Nucleic Acids Res. 2018;46(D1):D296–302. https://doi.org/10.1093/nar/gkx1067.
Xiao F, Zuo Z, Cai G, Kang S, Gao X, Li T. miRecords: an integrated resource for microRNA-target interactions. Nucleic Acids Res. 2009;37(Database issue):D105–10. https://doi.org/10.1093/nar/gkn851.
Jiang Q, Wang Y, Hao Y, Juan L, Teng M, Zhang X, Li M, Wang G, Liu Y. miR2Disease: a manually curated database for microRNA deregulation in human disease. Nucleic Acids Res. 2009;37(Database issue):D98–104. https://doi.org/10.1093/nar/gkn714.
Huang Z, Shi J, Gao Y, Cui C, Zhang S, Li J, Zhou Y, Cui Q. HMDD v3.0: a database for experimentally supported human microRNA-disease associations. Nucleic Acids Res. 2019;47(D1):D1013–7. https://doi.org/10.1093/nar/gky1010z.
Ruepp A, Kowarsch A, Schmidl D, Buggenthin F, Brauner B, Dunger I, Fobo G, Frishman G, Montrone C, Theis FJ. PhenomiR: a knowledgebase for microRNA expression in diseases and biological processes. Genome Biol. 2010;11(1):R6. https://doi.org/10.1186/gb-2010-11-1-r6.
Liu X, Wang S, Meng F, Wang J, Zhang Y, Dai E, Yu X, Li X, Jiang W. SM2miR: a database of the experimentally validated small molecules’ effects on microRNA expression. Bioinformatics. 2013;29(3):409–11. https://doi.org/10.1093/bioinformatics/bts698.
Rukov JL, Wilentzik R, Jaffe I, Vinther J, Shomron N. Pharmaco-miR: linking microRNAs and drug effects. Brief Bioinform. 2014;15(4):648–59. https://doi.org/10.1093/bib/bbs082.
Dai E, Yu X, Zhang Y, Meng F, Wang S, Liu X, Liu D, Wang J, Li X, Jiang W. EpimiR: a database of curated mutual regulation between miRNAs and epigenetic modifications. Database. 2014;2014:bau023. https://doi.org/10.1093/database/bau023.
Li JH, Liu S, Zhou H, Qu LH, Yang JH. starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res. 2014;42(Database issue):D92–7. https://doi.org/10.1093/nar/gkt1248.
Zhou G, Soufan O, Ewald J, Hancock REW, Basu N, Xia J. NetworkAnalyst 3.0: a visual analytics platform for comprehensive gene expression profiling and meta-analysis. Nucleic Acids Res. 2019. https://doi.org/10.1093/nar/gkz240.
Davis CA, Hitz BC, Sloan CA, Chan ET, Davidson JM, Gabdank I, Hilton JA, Jain K, Baymuradov UK, Narayanan AK, et al. The Encyclopedia of DNA elements (ENCODE): data portal update. Nucleic Acids Res. 2018;46(D1):D794–801. https://doi.org/10.1093/nar/gkx1081.
Chandrashekar DS, Bashel B, Balasubramanya SAH, Creighton CJ, Ponce-Rodriguez I, Chakravarthi BVSK, Varambally S. UALCAN: a portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia. 2017;19(8):649–58. https://doi.org/10.1016/j.neo.2017.05.002.
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, Sun Y, Jacobsen A, Sinha R, Larsson E, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013;6(269):pl1. https://doi.org/10.1126/scisignal.2004088.
Funahashi H, Imai T, Tanaka Y, Tobinaga J, Wada M, Matsuyama T, Tsukamura K, Yamada F, Takagi H, Narita T, et al. Discrepancy between PNMT presence and relative lack of adrenaline production in extra-adrenal pheochromocytoma. J Surg Oncol. 1994;57(3):196–200.
Maletínská L, Maixnerová J, Matysková R, Haugvicová R, Sloncová E, Elbert T, Slaninová J, Zelezná B. Cocaine- and amphetamine-regulated transcript (CART) peptide specific binding in pheochromocytoma cells PC12. Eur J Pharmacol. 2007;559(2–3):109–14. https://doi.org/10.1016/j.ejphar.2006.12.014.
Roperch JP, Incitti R, Forbin S, Bard F, Mansour H, Mesli F, Baumgaertner I, Brunetti F, Sobhani I. Aberrant methylation of NPY, PENK, and WIF1 as a promising marker for blood-based diagnosis of colorectal cancer. BMC Cancer. 2013;13:566. https://doi.org/10.1186/1471-2407-13-566.
Lawrence B, Gustafsson BI, Kidd M, Pavel M, Svejda B, Modlin IM. The clinical relevance of chromogranin A as a biomarker for gastroenteropancreatic neuroendocrine tumors. Endocrinol Metab Clin N Am. 2011;40(1):111–334. https://doi.org/10.1016/j.ecl.2010.12.001.
Feng S, Wang J, Zhang Y, Creighton CJ, Ittmann M. FGF23 promotes prostate cancer progression. Oncotarget. 2015;6(19):17291–301. https://doi.org/10.18632/oncotarget.4174.
Miller SL, Antico G, Raghunath PN, Tomaszewski JE, Clevenger CV. Nek3 kinase regulates prolactin-mediated cytoskeletal reorganization and motility of breast cancer cells. Oncogene. 2007;26(32):4668–78. https://doi.org/10.1038/sj.onc.1210264.
Zhao Y, Yun D, Zou X, Jiang T, Li G, Hu L, Chen J, Xu J, Mao Y, Chen H, et al. Whole exome-wide association study identifies a missense variant in SLC2A4RG associated with glioblastoma risk. Am J Cancer Res. 2017;7(9):1937–47.
Price JC, Pollock LM, Rudd ML, Fogoros SK, Mohamed H, Hanigan CL, Le Gallo M. Sequencing of candidate chromosome instability genes in endometrial cancers reveals somatic mutations in ESCO1, CHTF18, and MRE11A. PLoS ONE. 2013;8(6):e63313. https://doi.org/10.1371/journal.pone.0063313.
An Y, Zhang Z, Shang Y, Jiang X, Dong J, Yu P, Nie Y, Zhao Q. miR-23b-3p regulates the chemoresistance of gastric cancer cells by targeting ATG12 and HMGB2. Cell Death Dis. 2015;6:e1766. https://doi.org/10.1038/cddis.2015.123.
Drelon C, Berthon A, Mathieu M, Ragazzon B, Kuick R, Tabbal H, Septier A, Rodriguez S, Batisse-Lignier M, Sahut-Barnola I, et al. EZH2 is overexpressed in adrenocortical carcinoma and is associated with disease progression. Hum Mol Genet. 2016;25(13):2789–800. https://doi.org/10.1093/hmg/ddw136.
Bethke L, Webb E, Sellick G, Rudd M, Penegar S, Withey L, Qureshi M, Houlston R. Polymorphisms in the cytochrome P450 genes CYP1A2, CYP1B1, CYP3A4, CYP3A5, CYP11A1, CYP17A1, CYP19A1 and colorectal cancer risk. BMC Cancer. 2007;7:123. https://doi.org/10.1186/1471-2407-7-123.
Chang BL, Zheng SL, Hawkins GA, Isaacs SD, Wiley KE, Turner A, Carpten JD, Bleecker ER, Walsh PC, Trent JM, et al. Joint effect of HSD3B1 and HSD3B2 genes is associated with hereditary and sporadic prostate cancer susceptibility. Cancer Res. 2002;62(6):1784–9.
Sigurdson AJ, Brenner AV, Roach JA, Goudeva L, Müller JA, Nerlich K, Reiners C, Schwab R, Pfeiffer L, Waldenberger M, et al. Selected single-nucleotide polymorphisms in FOXE1, SERPINA5, FTO, EVPL, TICAM1 and SCARB1 are associated with papillary and follicular thyroid cancer risk: replication study in a German population. Carcinogenesis. 2016;37(7):677–84. https://doi.org/10.1093/carcin/bgw047.
Kelemen LE, Couch FJ, Ahmed S, Dunning AM, Pharoah PD, Easton DF, Fredericksen ZS, Vierkant RA, Pankratz VS, Goode EL, et al. Genetic variation in stromal proteins decorin and lumican with breast cancer: investigations in two case-control studies. Breast Cancer Res. 2008;10(6):R98. https://doi.org/10.1186/bcr2201.
Slattery ML, Samowitz W, Curtin K, Ma KN, Hoffman M, Caan B, Neuhausen S. Associations among IRS1, IRS2, IGF1, and IGFBP3 genetic polymorphisms and colorectal cancer. Cancer Epidemiol Biomark Prev. 2004;13(7):1206–14.
Müller-Vieira U, Angotti M, Hartmann RW. The adrenocortical tumor cell line NCI-H295R as an in vitro screening system for the evaluation of CYP11B2 (aldosterone synthase) and CYP11B1 (steroid-11beta-hydroxylase) inhibitors. J Steroid Biochem Mol Biol. 2005;96(3–4):259–70. https://doi.org/10.1016/j.jsbmb.2005.04.032.
Kroiss M, Reuss M, Kühner D, Johanssen S, Beyer M, Zink M, Hartmann MF, Dhir V, Wudy SA, Arlt W, et al. Sunitinib inhibits cell proliferation and alters steroidogenesis by down-regulation of HSD3B2 in adrenocortical carcinoma cells. Front Endocrinol. 2011;2:27. https://doi.org/10.3389/fendo.2011.00027.
Zhu Y, Xu Y, Chen D, Zhang C, Rui W, Zhao J, Zhu Q, Wu Y, Shen Z, Wang W, et al. Expression of STAT3 and IGF2 in adrenocortical carcinoma and its relationship with angiogenesis. Clin Transl Oncol. 2014;16(7):644–9. https://doi.org/10.1007/s12094-013-1130-1.
Shigematsu K, Nishida N, Sakai H, Igawa T, Toriyama K, Nakatani A, Takahara O, Kawai K. Synaptophysin immunoreactivity in adrenocortical adenomas: a correlation between synaptophysin and CYP17A1 expression. Eur J Endocrinol. 2009;161(6):939–45. https://doi.org/10.1530/EJE-09-0596.
Doghman M, Arhatte M, Thibout H, Rodrigues G, De Moura J, Grosso S, West AN, Laurent M, Mas JC, Bongain A, et al. Nephroblastoma overexpressed/cysteine-rich protein 61/connective tissue growth factor/nephroblastoma overexpressed gene-3 (NOV/CCN3), a selective adrenocortical cell proapoptotic factor, is down-regulated in childhood adrenocortical tumors. J Clin Endocrinol Metab. 2007;92(8):3253–60. https://doi.org/10.1210/jc.2007-0342.
Boulle N, Logié A, Gicquel C, Perin L, Le Bouc Y. Increased levels of insulin-like growth factor II (IGF-II) and IGF-binding protein-2 are associated with malignancy in sporadic adrenocortical tumors. J Clin Endocrinol Metab. 1998;83(5):1713–20. https://doi.org/10.1210/jcem.83.5.4816.
Ilvesmäki V, Liu J, Heikkilä P, Kahri AI, Voutilainen R. Expression of insulin-like growth factor binding protein 1-6 genes in adrenocortical tumors and pheochromocytomas. Horm Metab Res. 1998;30(10):619–23. https://doi.org/10.1055/s-2007-978945.
Mantovani G, Lania AG, Bondioni S, Peverelli E, Pedroni C, Ferrero S, Pellegrini C, Vicentini L, Arnaldi G, Bosari S, et al. Different expression of protein kinase A (PKA) regulatory subunits in cortisol-secreting adrenocortical tumors: relationship with cell proliferation. Exp Cell Res. 2008;314(1):123–30. https://doi.org/10.1016/j.yexcr.2007.08.024.
Hafiz S, Dennis JC, Schwartz D, Judd R, Tao YX, Khazal K, Akingbemi B, Mo XL, Abdel-Mageed AB, Morrison E, et al. Expression of melanocortin receptors in human prostate cancer cell lines: MC2R activation by ACTH increases prostate cancer cell proliferation. Int J Oncol. 2012;41(4):1373–80. https://doi.org/10.3892/ijo.2012.1574.
Zahnow CA. CCAAT/enhancer-binding protein beta: its role in breast cancer and associations with receptor tyrosine kinases. Expert Rev Mol Med. 2009;11:e12. https://doi.org/10.1017/S1462399409001033.
Kim SH, Park YY, Kim SW, Lee JS, Wang D, DuBois RN. ANGPTL4 induction by prostaglandin E2 under hypoxic conditions promotes colorectal cancer progression. Cancer Res. 2011;71(22):7010–20. https://doi.org/10.1158/0008-5472.CAN-11-1262.
Zhou F, Drabsch Y, Dekker TJ, Vinuesa AG, Li Y, Hawinkels LJ, Sheppard KA, Goumans MJ, Luwor RB, Vries CJ, Mesker WE, et al. Nuclear receptor NR4A1 promotes breast cancer invasion and metastasis by activating TGF-β signalling. Nat Commun. 2014;5:3388. https://doi.org/10.1038/ncomms4388.
Holla VR, Mann JR, Shi Q, DuBois RN. Prostaglandin E2 regulates the nuclear receptor NR4A2 in colorectal cancer. J Biol Chem. 2006;281(5):2676–82. https://doi.org/10.1074/jbc.M507752200.
He JH, Li BX, Han ZP, Zou MX, Wang L, Lv YB, Zhou JB, Cao MR, Li YG. Zhang JZ et al Snail-activated long non-coding RNA PCA3 up-regulates PRKD3 expression by miR-1261 sponging, thereby promotes invasion and migration of prostate cancer cells. Tumour Biol. 2016. https://doi.org/10.1007/s13277-016-5450-y.
Tyan SW, Hsu CH, Peng KL, Chen CC, Kuo WH, Lee EY, Shew JY, Chang KJ, Juan LJ, Lee WH. Breast cancer cells induce stromal fibroblasts to secrete ADAMTS1 for cancer invasion through an epigenetic change. PLoS ONE. 2012;7(4):e35128. https://doi.org/10.1371/journal.pone.0035128.
Fan L, Zhu Q, Liu L, Zhu C, Huang H, Lu S, Liu P. CXCL13 is androgen-responsive and involved in androgen induced prostate cancer cell migration and invasion. Oncotarget. 2017;8(32):53244–61. https://doi.org/10.18632/oncotarget.18387.
Yu ST, Zhong Q, Chen RH, Han P, Li SB, Zhang H, Yuan L, Xia TL, Zeng MS, Huang XM. CRLF1 promotes malignant phenotypes of papillary thyroid carcinoma by activating the MAPK/ERK and PI3 K/AKT pathways. Cell Death Dis. 2018;9(3):371. https://doi.org/10.1038/s41419-018-0352-0.
Scotton CJ, Wilson JL, Scott K, Stamp G, Wilbanks GD, Fricker S, Bridger G, Balkwill FR. Multiple actions of the chemokine CXCL12 on epithelial tumor cells in human ovarian cancer. Cancer Res. 2002;62(20):5930–8.
Kavandi L, Collier MA, Nguyen H, Syed V. Progesterone and calcitriol attenuate inflammatory cytokines CXCL1 and CXCL2 in ovarian and endometrial cancer cells. J Cell Biochem. 2012;113(10):3143–52. https://doi.org/10.1002/jcb.24191.
Reed CC, Waterhouse A, Kirby S, Kay P, Owens RT, McQuillan DJ, Iozzo RV. Decorin prevents metastatic spreading of breast cancer. Oncogene. 2005;24(6):1104–10. https://doi.org/10.1038/sj.onc.1208329.
Sun C, Fukui H, Hara K, Zhang X, Kitayama Y, Eda H, Tomita T, Oshima T, Kikuchi S, Watari J, et al. FGF9 from cancer-associated fibroblasts is a possible mediator of invasion and anti-apoptosis of gastric cancer cells. BMC Cancer. 2015;15:333.
Husaini Y, Qiu MR, Lockwood GP, Luo XW, Shang P, Kuffner T, Tsai VW, Jiang L, Russell PJ, Brown DA, et al. Macrophage inhibitory cytokine-1 (MIC-1/GDF15) slows cancer development but increases metastases in TRAMP prostate cancer prone mice. PLoS ONE. 2012;7(8):e43833. https://doi.org/10.1371/journal.pone.0043833.
Duffy MJ, O’Grady P, Devaney D, O’Siorain L, Fennelly JJ, Lijnen HJ. Urokinase-plasminogen activator, a marker for aggressive breast carcinomas. Prelim Rep Cancer. 1988;62(3):531–3.
Hermani A, Hess J, De Servi B, Medunjanin S, Grobholz R, Trojan L, Angel P, Mayer D. Calcium-binding proteins S100A8 and S100A9 as novel diagnostic markers in human prostate cancer. Clin Cancer Res. 2005;11(14):5146–52. https://doi.org/10.1158/1078-0432.CCR-05-0352.
Zhao J, Tang H, Zhao H, Che W, Zhang L, Liang P. SEMA6A is a prognostic biomarker in glioblastoma. Tumour Biol. 2015;36(11):8333–40. https://doi.org/10.1007/s13277-015-3584-y.
Tanaka K, Arao T, Maegawa M, Matsumoto K, Kaneda H, Kudo K, Fujita Y, Yokote H, Yanagihara K, Yamada Y, et al. SRPX2 is overexpressed in gastric cancer and promotes cellular migration and adhesion. Int J Cancer. 2009;124(5):1072–80. https://doi.org/10.1002/ijc.24065.
Yuan Y, Nymoen DA, Stavnes HT, Rosnes AK, Bjørang O, Wu C, Nesland JM, Davidson B. Tenascin-X is a novel diagnostic marker of malignant mesothelioma. Am J Surg Pathol. 2009;33(11):1673–82. https://doi.org/10.1097/PAS.0b013e3181b6bde3.
Iliopoulos D, Hirsch HA, Wang G, Struhl K. Inducible formation of breast cancer stem cells and their dynamic equilibrium with non-stem cancer cells via IL6 secretion. Proc Natl Acad Sci USA. 2011;108(4):1397–402. https://doi.org/10.1073/pnas.1018898108.
Koo AS, Chiu R, Soong J, Dekernion JB, Belldegrun A. The expression of C-jun and junB mRNA in renal cell cancer and in vitro regulation by transforming growth factor beta 1 and tumor necrosis factor alpha 1. J Urol. 1992;148(4):1314–8.
Gregersen LH, Jacobsen AB, Frankel LB, Wen J, Krogh A, Lund AH. MicroRNA-145 targets YES and STAT1 in colon cancer cells. PLoS ONE. 2010;5(1):e8836. https://doi.org/10.1371/journal.pone.0008836.
Friedline JA, Garrett SH, Somji S, Todd JH, Sens DA. Differential expression of the MT-1E gene in estrogen-receptor-positive and -negative human breast cancer cell lines. Am J Pathol. 1998;152(1):23–7.
Lyakhovich A, Aksenov N, Pennanen P, Miettinen S, Ahonen MH, Syvälä H, Ylikomi T, Tuohimaa P. Vitamin D induced up-regulation of keratinocyte growth factor (FGF-7/KGF) in MCF-7 human breast cancer cells. Biochem Biophys Res Commun. 2000;273(2):675–80. https://doi.org/10.1006/bbrc.2000.2998.
Bettin A, Reyes I, Reyes N. Gene expression profiling of prostate cancer-associated genes identifies fibromodulin as potential novel biomarker for prostate cancer. Int J Biol Mark. 2016;31(2):e153–62. https://doi.org/10.5301/jbm.5000184.
Qian X, Li C, Pang B, Xue M, Wang J, Zhou J. Spondin-2 (SPON2), a more prostate-cancer-specific diagnostic biomarker. PLoS ONE. 2012;7(5):e37225. https://doi.org/10.1371/journal.pone.0037225.
Zhou ZQ, Cao WH, Xie JJ, Lin J, Shen ZY, Zhang QY, Shen JH, Xu LY, Li EM. Expression and prognostic significance of THBS1, Cyr61 and CTGF in esophageal squamous cell carcinoma. BMC Cancer. 2009;9:291. https://doi.org/10.1186/1471-2407-9-291.
Pierconti F, Martini M, Pinto F, Cenci T, Capodimonti S, Calarco A, Bassi PF, Larocca LM. Epigenetic silencing of SOCS3 identifies a subset of prostate cancer with an aggressive behavior. Prostate. 2011;71(3):318–25. https://doi.org/10.1002/pros.21245.
Henrique R, Jerónimo C, Hoque MO, Nomoto S, Carvalho AL, Costa VL, Oliveira J, Teixeira MR, Lopes C, Sidransky D. MT1G hypermethylation is associated with higher tumor stage in prostate cancer. Cancer Epidemiol Biomark Prev. 2005;14(5):1274–8. https://doi.org/10.1158/1055-9965.EPI-04-0659.
Han YC, Zheng ZL, Zuo ZH, Yu YP, Chen R, Tseng GC, Nelson JB, Luo JH. Metallothionein 1 h tumour suppressor activity in prostate cancer is mediated by euchromatin methyltransferase. J Pathol. 2013;230(2):184–93. https://doi.org/10.1002/path.4169.
Du W, Wang S, Zhou Q, Li X, Chu J, Chang Z, Tao Q, Ng EK, Fang J, Sung JJ, et al. ADAMTS9 is a functional tumor suppressor through inhibiting AKT/mTOR pathway and associated with poor survival in gastric cancer. Oncogene. 2013;32(28):3319–28. https://doi.org/10.1038/onc.2012.359.
Chen HA, Kuo TC, Tseng CF, Ma JT, Yang ST, Yen CJ, Yang CY, Sung SY, Su JL. Angiopoietin-like protein 1 antagonizes MET receptor activity to repress sorafenib resistance and cancer stemness in hepatocellular carcinoma. Hepatology. 2016;64(5):1637–51. https://doi.org/10.1002/hep.28773.
Cheng YY, Jin H, Liu X, Siu JM, Wong YP, Ng EK, Yu J, Leung WK, Sung JJ, Chan FK. Fibulin 1 is downregulated through promoter hypermethylation in gastric cancer. Br J Cancer. 2008;99(12):2083–7. https://doi.org/10.1038/sj.bjc.6604760.
Yang G, Liang Y, Zheng T, Song R, Wang J, Shi H, Sun B, Xie C, Li Y, Han J, et al. FCN2 inhibits epithelial-mesenchymal transition-induced metastasis of hepatocellular carcinoma via TGF-β/Smad signaling. Cancer Lett. 2016;378(2):80–6. https://doi.org/10.1016/j.canlet.2016.05.007.
Tomizawa Y, Sekido Y, Kondo M, Gao B, Yokota J, Roche J, Drabkin H, Lerman MI, Gazdar AF, Minna JD. Inhibition of lung cancer cell growth and induction of apoptosis after reexpression of 3p21.3 candidate tumor suppressor gene SEMA3B. Proc Natl Acad Sci USA. 2001;98(24):13954–9. https://doi.org/10.1073/pnas.231490898.
Azhikina T, Kozlova A, Skvortsov T, Sverdlov E. Heterogeneity and degree of TIMP4, GATA4, SOX18, and EGFL7 gene promoter methylation in non-small cell lung cancer and surrounding tissues. Cancer Genet. 2011;204(9):492–500. https://doi.org/10.1016/j.cancergen.2011.07.010.
Lu DD, Chen YC, Zhang XR, Cao XR, Jiang HY, Yao L. The relationship between metallothionein-1F (MT1F) gene and hepatocellular carcinoma. Yale J Biol Med. 2003;76(2):55–62.
Soria G, Ben-Baruch A. The inflammatory chemokines CCL2 and CCL5 in breast cancer. Cancer Lett. 2008;267(2):271–85. https://doi.org/10.1016/j.canlet.2008.03.018.
Wang D, Wang H, Brown J, Daikoku T, Ning W, Shi Q, Richmond A, Strieter R, Dey SK, DuBois RN. CXCL1 induced by prostaglandin E2 promotes angiogenesis in colorectal cancer. J Exp Med. 2006;203(4):941–51. https://doi.org/10.1084/jem.20052124.
Sadr-Nabavi A, Ramser J, Volkmann J, Naehrig J, Wiesmann F, Betz B, Hellebrand H, Engert S, Seitz S, Kreutzfeld R, et al. Decreased expression of angiogenesis antagonist EFEMP1 in sporadic breast cancer is caused by aberrant promoter methylation and points to an impact of EFEMP1 as molecular biomarker. Int J Cancer. 2009;124(7):1727–35. https://doi.org/10.1002/ijc.24108.
Tümer N, Brown JW, Carballeira A, Fishman LM. Tyrosine hydroxylase gene expression in varying forms of human pheochromocytoma. Life Sci. 1996;59(19):1659–65.
Lundberg JM, Hökfelt T, Hemsén A, Theodorsson-Norheim E, Pernow J, Hamberger B, Goldstein M. Neuropeptide Y-like immunoreactivity in adrenaline cells of adrenal medulla and in tumors and plasma of pheochromocytoma patients. Regul Pept. 1986;13(2):169–82.
He Q, Zou L, Zhang PA, Lui JX, Skog S, Fornander T. The clinical significance of thymidine kinase 1 measurement in serum of breast cancer patients using anti-TK1 antibody. Int J Biol Mark. 2000;15(2):139–46.
Watson RG, Muhale F, Thorne LB, Yu J, O’Neil BH, Hoskins JM, Meyers MO, Deal AM, Ibrahim JG, Hudson ML, et al. Amplification of thymidylate synthetase in metastatic colorectal cancer patients pretreated with 5-fluorouracil-based chemotherapy. Eur J Cancer. 2010;46(18):3358–64. https://doi.org/10.1016/j.ejca.2010.07.011.
Nakayama N, Nakayama K, Shamima Y, Ishikawa M, Katagiri A, Iida K, Miyazaki K. Gene amplification CCNE1 is related to poor survival and potential therapeutic target in ovarian cancer. Cancer. 2010;116(11):2621–34. https://doi.org/10.1002/cncr.24987.
Bravou V, Nishitani H, Song SY, Taraviras S, Varakis J. Expression of the licensing factors, Cdt1 and Geminin, in human colon cancer. Int J Oncol. 2005;27(6):1511–8.
Ifon ET, Pang AL, Johnson W, Cashman K, Zimmerman S, Muralidhar S, Chan WY, Casey J, Rosenthal LJ. U94 alters FN1 and ANGPTL4 gene expression and inhibits tumorigenesis of prostate cancer cell line PC3. Cancer Cell Int. 2005;5:19. https://doi.org/10.1186/1475-2867-5-19.
Esseghir S, Kennedy A, Seedhar P, Nerurkar A, Poulsom R, Reis-Filho JS, Isacke CM. Identification of NTN4, TRA1, and STC2 as prognostic markers in breast cancer in a screen for signal sequence encoding proteins. Clin Cancer Res. 2007;13(11):3164–73. https://doi.org/10.1158/1078-0432.CCR-07-0224.
Durak I, Biri H, Devrim E, Sözen S, Avci A. Aqueous extract of Urtica dioica makes significant inhibition on adenosine deaminase activity in prostate tissue from patients with prostate cancer. Cancer Biol Ther. 2004;3(9):855–7.
Wolkersdörfer GW, Marx C, Brown J, Schröder S, Füssel M, Rieber EP, Kuhlisch E, Ehninger G, Bornstein SR. Prevalence of HLA-DRB1 genotype and altered Fas/Fas ligand expression in adrenocortical carcinoma. J Clin Endocrinol Metab. 2005;90(3):1768–74. https://doi.org/10.1210/jc.2004-1406.
Szajerka A, Dziegiel P, Szajerka T, Zabel M, Winowski J, Grzebieniak Z. Immunohistochemical evaluation of metallothionein, Mcm-2 and Ki-67 antigen expression in tumors of the adrenal cortex. Anticancer Res. 2008;28(5B):2959–65.
Berends MJ, Cats A, Hollema H, Karrenbeld A, Beentjes JA, Sijmons RH, Mensink RG, Hofstra RM, Verschueren RC, Kleibeuker JH. Adrenocortical adenocarcinoma in an MSH2 carrier: coincidence or causal relation? Hum Pathol. 2000;31(12):1522–7. https://doi.org/10.1053/hupa.2000.20409.
Jin B, Yao B, Li JL, Fields CR, Delmas AL, Liu C, Robertson KD. DNMT1 and DNMT3B modulate distinct polycomb-mediated histone modifications in colon cancer. Cancer Res. 2009;69(18):7412–21. https://doi.org/10.1158/0008-5472.CAN-09-0116.
Kobierzycki C, Pula B, Skiba M, Jablonska K, Latkowski K, Zabel M, Nowak-Markwitz E, Spaczynski M, Kedzia W, Podhorska-Okolow M, et al. Comparison of minichromosome maintenance proteins (MCM-3, MCM-7) and metallothioneins (MT-I/II, MT-III) expression in relation to clinicopathological data in ovarian cancer. Anticancer Res. 2013;33(12):5375–83.
Vigouroux C, Casse JM, Battaglia-Hsu SF, Brochin L, Luc A, Paris C, Lacomme S, Gueant JL, Vignaud JM, Gauchotte G. Methyl(R217)HuR and MCM6 are inversely correlated and are prognostic markers in non small cell lung carcinoma. Lung Cancer. 2015;89(2):189–96. https://doi.org/10.1016/j.lungcan.2015.05.008.
Li J, Ding Y, Li A. Identification of COL1A1 and COL1A2 as candidate prognostic factors in gastric cancer. World J Surg Oncol. 2016;14(1):297. https://doi.org/10.1186/s12957-016-1056-5.
Yuan L, Shu B, Chen L, Qian K, Wang Y, Qian G, Zhu Y, Cao X, Xie C, Xiao Y, et al. Overexpression of COL3A1 confers a poor prognosis in human bladder cancer identified by co-expression analysis. Oncotarget. 2017;8(41):70508–20. https://doi.org/10.18632/oncotarget.19733.
Moller-Levet CS, Betts GN, Harris AL, Homer JJ, West CM, Miller CJ. Exon array analysis of head and neck cancers identifies a hypoxia related splice variant of LAMA3 associated with a poor prognosis. PLoS Comput Biol. 2009;5(11):e1000571. https://doi.org/10.1371/journal.pcbi.1000571.
Lin Q, Lim HS, Lin HL, Tan HT, Lim TK, Cheong WK, Cheah PY, Tang CL, Chow PK, Chung MC. Analysis of colorectal cancer glyco-secretome identifies laminin β-1 (LAMB1) as a potential serological biomarker for colorectal cancer. Proteomics. 2015;15(22):3905–20. https://doi.org/10.1002/pmic.201500236.
Wu Y, Berends MJ, Post JG, Mensink RG, Verlind E, Sluis T, Kempinga C, Sijmons RH, Zee AG, Hollema H, et al. Germline mutations of EXO1 gene in patients with hereditary nonpolyposis colorectal cancer (HNPCC) and atypical HNPCC forms. Gastroenterology. 2001;120(7):1580–7.
Valle L, Hernández-Illán E, Bellido F, Aiza G, Castillejo A, Castillejo MI, Navarro M, Seguí N, Vargas G, Guarinos C, et al. New insights into POLE and POLD1 germline mutations in familial colorectal cancer and polyposis. Hum Mol Genet. 2014;23(13):3506–12. https://doi.org/10.1093/hmg/ddu058.
Li D, Li R, Zhang J, Li K, Wu Y. Association between the LIG1 polymorphisms and lung cancer risk: a meta-analysis of case-control studies. Cell Biochem Biophys. 2015;73(2):381–7. https://doi.org/10.1007/s12013-015-0619-3.
Takahashi K, Satoh F, Sone M, Totsune K, Arihara Z, Noshiro T, Mouri T, Murakami O. Expression of adrenomedullin mRNA in adrenocortical tumors and secretion of adrenomedullin by cultured adrenocortical carcinoma cells. Peptides. 1998;19(10):1719–24.
Holwell TA, Schweitzer SC, Reyland ME, Evans RM. Vimentin-dependent utilization of LDL-cholesterol in human adrenal tumor cells is not associated with the level of expression of apoE, sterol carrier protein-2, or caveolin. J Lipid Res. 1999;40(8):1440–52.
Kiiveri S, Siltanen S, Rahman N, Bielinska M, Lehto VP, Huhtaniemi IT, Muglia LJ, Wilson DB, Heikinheimo M. Reciprocal changes in the expression of transcription factors GATA-4 and GATA-6 accompany adrenocortical tumorigenesis in mice and humans. Mol Med. 1999;5(7):490–501.
Liu-Chittenden Y, Jain M, Gaskins K, Wang S, Merino MJ, Kotian S, Kumar Gara S, Davis S, Zhang L, Kebebew E. RARRES2 functions as a tumor suppressor by promoting β-catenin phosphorylation/degradation and inhibiting p38 phosphorylation in adrenocortical carcinoma. Oncogene. 2017;36(25):3541–52. https://doi.org/10.1038/onc.2016.497.
Ma T, Zhao Y, Wei K, Yao G, Pan C, Liu B, Xia Y, He Z, Qi X, Li Z, et al. MicroRNA-124 functions as a tumor suppressor by regulating CDH2 and epithelial-mesenchymal transition in non-small cell lung cancer. Cell Physiol Biochem. 2016;38(4):1563–74. https://doi.org/10.1159/000443097.
Nasu T, Oku Y, Takifuji K, Hotta T, Yokoyama S, Matsuda K, Tamura K, Ieda J, Yamamoto N, Takemura S, et al. Predicting lymph node metastasis in early colorectal cancer using the CITED1 expression. J Surg Res. 2013;185(1):136–42. https://doi.org/10.1016/j.jss.2013.05.041.
Gupta S, Iljin K, Sara H, Mpindi JP, Mirtti T, Vainio P, Rantala J, Alanen K, Nees M, Kallioniemi O. FZD4 as a mediator of ERG oncogene-induced WNT signaling and epithelial-to-mesenchymal transition in human prostate cancer cells. Cancer Res. 2010;70(17):6735–45. https://doi.org/10.1158/0008-5472.CAN-10-0244.
Tang F, Zhang R, He Y, Zou M, Guo L, Xi T. MicroRNA-125b induces metastasis by targeting STARD13 in MCF-7 and MDA-MB-231 breast cancer cells. PLoS ONE. 2012;7(5):e35435. https://doi.org/10.1371/journal.pone.0035435.
Li H, Chen X, Gao Y, Wu J, Zeng F, Song F. XBP1 induces snail expression to promote epithelial- to-mesenchymal transition and invasion of breast cancer cells. Cell Signal. 2015;27(1):82–9. https://doi.org/10.1016/j.cellsig.2014.09.018.
Abd Elmageed ZY, Moroz K, Kandil E. Clinical significance of CD146 and latexin during different stages of thyroid cancer. Mol Cell Biochem. 2013;381(1–2):95–103. https://doi.org/10.1007/s11010-013-1691-x.
Biktasova AK, Dudimah DF, Uzhachenko RV, Park K, Akhter A, Arasada RR, Evans JV, Novitskiy SV, Tchekneva EE, Carbone DP, et al. Multivalent forms of the Notch ligand DLL-1 enhance antitumor T-cell immunity in lung cancer and improve efficacy of EGFR-targeted therapy. Cancer Res. 2015;75(22):4728–41. https://doi.org/10.1158/0008-5472.CAN-14-1154.
Jones A, Teschendorff AE, Li Q, Hayward JD, Kannan A, Mould T, West J, Zikan M, Cibula D, Fiegl H, et al. Role of DNA methylation and epigenetic silencing of HAND2 in endometrial cancer development. PLoS Med. 2013;10(11):e1001551. https://doi.org/10.1371/journal.pmed.1001551.
Okazaki S, Ishikawa T, Iida S, Ishiguro M, Kobayashi H, Higuchi T, Enomoto M, Mogushi K, Mizushima H, Tanaka H, et al. Clinical significance of UNC5B expression in colorectal cancer. Int J Oncol. 2012;40(1):209–16. https://doi.org/10.3892/ijo.2011.1201.
Cooper AB, Wu J, Lu D, Maluccio MA. Is autotaxin (ENPP2) the link between hepatitis C and hepatocellular cancer? J Gastrointest Surg. 2007;11(12):1628–34. https://doi.org/10.1007/s11605-007-0322-9.
Falck E, Klinga-Levan K. Expression patterns of Phf5a/PHF5A and Gja1/GJA1 in rat and human endometrial cancer. Cancer Cell Int. 2013;13(1):43. https://doi.org/10.1186/1475-2867-13-43.
Gao F, Zhang Y, Wang S, Liu Y, Zheng L, Yang J, Huang W, Ye Y, Luo W, Xiao D. Hes1 is involved in the self-renewal and tumourigenicity of stem-like cancer cells in colon cancer. Sci Rep. 2014;4:3963. https://doi.org/10.1038/srep03963.
Logan M, Anderson PD, Saab ST, Hameed O, Abdulkadir SA. RAMP1 is a direct NKX3.1 target gene up-regulated in prostate cancer that promotes tumorigenesis. Am J Pathol. 2013;183(3):951–63. https://doi.org/10.1016/j.ajpath.2013.05.021.
Huffman DM, Grizzle WE, Bamman MM, Kim JS, Eltoum IA, Elgavish A, Nagy TR. SIRT1 is significantly elevated in mouse and human prostate cancer. Cancer Res. 2007;67(14):6612–8. https://doi.org/10.1158/0008-5472.CAN-07-0085.
Fu L, Shi K, Wang J, Chen W, Shi D, Tian Y, Guo W, Yu W, Xiao X, Kang T, et al. TFAP2B overexpression contributes to tumor growth and a poor prognosis of human lung adenocarcinoma through modulation of ERK and VEGF/PEDF signaling. Mol Cancer. 2014;13:89. https://doi.org/10.1186/1476-4598-13-89.
Wang X, Liu Y, Shao D, Qian Z, Dong Z, Sun Y, Xing X, Cheng X, Du H, Hu Y, et al. Recurrent amplification of MYC and TNFRSF11B in 8q24 is associated with poor survival in patients with gastric cancer. Gastric Cancer. 2016;19(1):116–27. https://doi.org/10.1007/s10120-015-0467-2.
Milan E, Lazzari C, Anand S, Floriani I, Torri V, Sorlini C, Gregorc V, Bachi A. SAA1 is over-expressed in plasma of non small cell lung cancer patients with poor outcome after treatment with epidermal growth factor receptor tyrosine-kinase inhibitors. J Proteomics. 2012;76:91–101. https://doi.org/10.1016/j.jprot.2012.06.022.
Stasinopoulos IA, Mironchik Y, Raman A, Wildes F, Winnard P Jr, Raman V, et al. HOXA5-twist interaction alters p53 homeostasis in breast cancer cells. J Biol Chem. 2005;280(3):2294–9. https://doi.org/10.1074/jbc.M411018200.
Ota T, Gilks CB, Longacre T, Leung PC, Auersperg N. HOXA7 in epithelial ovarian cancer: interrelationships between differentiation and clinical features. Reprod Sci. 2007;14(6):605–14. https://doi.org/10.1177/1933719107307781.
Kusinska R, Górniak P, Pastorczak A, Fendler W, Potemski P, Mlynarski W, Kordek R. Influence of genomic variation in FTO at 16q12.2, MC4R at 18q22 and NRXN3 at 14q31 genes on breast cancer risk. Mol Biol Rep. 2012;39(3):2915–9. https://doi.org/10.1007/s11033-011-1053-2.
Sharifi N, Hurt EM, Kawasaki BT, Farrar WL. TGFBR3 loss and consequences in prostate cancer. Prostate. 2007;67(3):301–11. https://doi.org/10.1002/pros.20526.
Olakowski M, Tyszkiewicz T, Jarzab M, Król R, Oczko-Wojciechowska M, Kowalska M, Kowal M, Gala GM, Kajor M, Lange D, et al. NBL1 and anillin (ANLN) genes over-expression in pancreatic carcinoma. Folia Histochem Cytobiol. 2009;47(2):249–55. https://doi.org/10.2478/v10042-009-0031-1.
Hayward DG, Clarke RB, Faragher AJ, Pillai MR, Hagan IM, Fry AM. The centrosomal kinase Nek2 displays elevated levels of protein expression in human breast cancer. Cancer Res. 2004;64(20):7370–6. https://doi.org/10.1158/0008-5472.CAN-04-0960.
Yoshida K, Sato M, Hase T, Elshazley M, Yamashita R, Usami N, Taniguchi T, Yokoi K, Nakamura S, Kondo M, et al. TIMELESS is overexpressed in lung cancer and its expression correlates with poor patient survival. Cancer Sci. 2013;104(2):171–7. https://doi.org/10.1111/cas.12068.
Warner SL, Stephens BJ, Nwokenkwo S, Hostetter G, Sugeng A, Hidalgo M, Trent JM, Han H, Von Hoff DD. Validation of TPX2 as a potential therapeutic target in pancreatic cancer cells. Clin Cancer Res. 2009;15(21):6519–28. https://doi.org/10.1158/1078-0432.CCR-09-0077.
Maire V, Baldeyron C, Richardson M, Tesson B, Vincent-Salomon A, Gravier E, Marty-Prouvost B, De Koning L, Rigaill G, Dumont A, et al. TTK/hMPS1 is an attractive therapeutic target for triple-negative breast cancer. PLoS ONE. 2013;8(5):e63712. https://doi.org/10.1371/journal.pone.0063712.
Jiao DC, Lu ZD, Qiao JH, Yan M, Cui SD, Liu ZZ. Expression of CDCA8 correlates closely with FOXM1 in breast cancer: public microarray data analysis and immunohistochemical study. Neoplasma. 2015;62(3):464–9. https://doi.org/10.4149/neo_2015_055.
Nagahara M, Nishida N, Iwatsuki M, Ishimaru S, Mimori K, Tanaka F, Nakagawa T, Sato T, Sugihara K, Hoon DS, et al. Kinesin 18A expression: clinical relevance to colorectal cancer progression. Int J Cancer. 2011;129(11):2543–52. https://doi.org/10.1002/ijc.25916.
Nakamura Y, Tanaka F, Nagahara H, Ieta K, Haraguchi N, Mimori K, Sasaki A, Inoue H, Yanaga K, Mori M. Opa interacting protein 5 (OIP5) is a novel cancer-testis specific gene in gastric cancer. Ann Surg Oncol. 2007;14(2):885–92. https://doi.org/10.1245/s10434-006-9121-x.
Chen J, Chen H, Yang H, Dai H. SPC25 upregulation increases cancer stem cell properties in non-small cell lung adenocarcinoma cells and independently predicts poor survival. Biomed Pharmacother. 2018;100:233–9. https://doi.org/10.1016/j.biopha.2018.02.015.
Yuan W, Xie S, Wang M, Pan S, Huang X, Xiong M, Xiao RJ, Xiong J, Zhang QP, Shao L. Bioinformatic analysis of prognostic value of ZW10 interacting protein in lung cancer. Onco Targets Ther. 2018;11:1683–95. https://doi.org/10.2147/ott.s149012.
Lorenzi PL, Llamas J, Gunsior M, Ozbun L, Reinhold WC, Varma S, Ji H, Kim H, Hutchinson AA, Kohn EC, Goldsmith PK, et al. Asparagine synthetase is a predictive biomarker of l-asparaginase activity in ovarian cancer cell lines. Mol Cancer Ther. 2008;7(10):3123–8. https://doi.org/10.1158/1535-7163.MCT-08-0589.
Park SA, Platt J, Lee JW, López-Giráldez F, Herbst RS, Koo JS. E2F8 as a novel therapeutic target for lung cancer. J Natl Cancer Inst. 2015. https://doi.org/10.1093/jnci/djv151.
Yang J, Bielenberg DR, Rodig SJ, Doiron R, Clifton MC, Kung AL, Strong RK, Zurakowski D, Moses MA. Lipocalin 2 promotes breast cancer progression. Proc Natl Acad Sci USA. 2009;106(10):3913–8. https://doi.org/10.1073/pnas.0810617106.
Hu F, Deng X, Yang X, Jin H, Gu D, Lv X, Wang C, Zhang Y, Huo X, Shen Q, et al. Hypoxia upregulates Rab11-family interacting protein 4 through HIF-1α to promote the metastasis of hepatocellular carcinoma. Oncogene. 2015;34(49):6007–17. https://doi.org/10.1038/onc.2015.49.
Wang C, Zheng X, Shen C, Shi Y. MicroRNA-203 suppresses cell proliferation and migration by targeting BIRC5 and LASP1 in human triple-negative breast cancer cells. J Exp Clin Cancer Res. 2012;31:58. https://doi.org/10.1186/1756-9966-31-58.
Grabsch H, Takeno S, Parsons WJ, Pomjanski N, Boecking A, Gabbert HE, Mueller W. Overexpression of the mitotic checkpoint genes BUB1, BUBR1, and BUB3 in gastric cancer–association with tumour cell proliferation. J Pathol. 2003;200(1):16–22. https://doi.org/10.1002/path.1324.
Ding K, Li W, Zou Z, Zou X, Wang C. CCNB1 is a prognostic biomarker for ER + breast cancer. Med Hypotheses. 2014;83(3):359–64. https://doi.org/10.1016/j.mehy.2014.06.013.
Shi Q, Wang W, Jia Z, Chen P, Ma K, Zhou C. ISL1, a novel regulator of CCNB1, CCNB2 and c-MYC genes, promotes gastric cancer cell proliferation and tumor growth. Oncotarget. 2016;7(24):36489–500. https://doi.org/10.18632/oncotarget.9269.
Shen Z, Yu X, Zheng Y, Lai X, Li J, Hong Y, Zhang H, Chen C, Su Z, Guo R. CDCA5 regulates proliferation in hepatocellular carcinoma and has potential as a negative prognostic marker. Onco Targets Ther. 2018;11:891–901. https://doi.org/10.2147/OTT.S154754.
Li T, Xue H, Guo Y, Guo K. CDKN3 is an independent prognostic factor and promotes ovarian carcinoma cell proliferation in ovarian cancer. Oncol Rep. 2014;31(4):1825–31. https://doi.org/10.3892/or.2014.3045.
Ueda S, Kondoh N, Tsuda H, Yamamoto S, Asakawa H, Fukatsu K, Kobayashi T, Yamamoto J, Tamura K, Ishida J, et al. Expression of centromere protein F (CENP-F) associated with higher FDG uptake on PET/CT, detected by cDNA microarray, predicts high-risk patients with primary breast cancer. BMC Cancer. 2008;8:384. https://doi.org/10.1186/1471-2407-8-384.
Abdel-Fatah TMA, Agarwal D, Liu DX, Russell R, Rueda OM, Liu K, Xu B, Moseley PM, Green AR, Pockley AG, et al. SPAG5 as a prognostic biomarker and chemotherapy sensitivity predictor in breast cancer: a retrospective, integrated genomic, transcriptomic, and protein analysis. Lancet Oncol. 2016;17(7):1004–18. https://doi.org/10.1016/S1470-2045(16)00174-1.
Sakamoto H, Friel AM, Wood AW, Guo L, Ilic A, Seiden MV, Chung DC, Lynch MP, Serikawa T, Munro E, et al. Mechanisms of Cables 1 gene inactivation in human ovarian cancer development. Cancer Biol Ther. 2008;7(2):180–8.
Wang W, Huper G, Guo Y, Murphy SK, Olson JA Jr, Marks JR. Analysis of methylation-sensitive transcriptome identifies GADD45a as a frequently methylated gene in breast cancer. Oncogene. 2005;24(16):2705–14. https://doi.org/10.1038/sj.onc.1208464.
Syed N, Coley HM, Sehouli J, Koensgen D, Mustea A, Szlosarek P, McNeish I, Blagden SP, Schmid P, Lovell DP, et al. Polo-like kinase Plk2 is an epigenetic determinant of chemosensitivity and clinical outcomes in ovarian cancer. Cancer Res. 2011;71(9):3317–27. https://doi.org/10.1158/0008-5472.CAN-10-2048.
Lind GE, Kleivi K, Meling GI, Teixeira MR, Thiis-Evensen E, Rognum TO, Lothe RA. ADAMTS1, CRABP1, and NR3C1 identified as epigenetically deregulated genes in colorectal tumorigenesis. Cell Oncol. 2006;28(5–6):259–72.
Kalmyrzaev B, Pharoah PD, Easton DF, Ponder BA, Dunning AM. Hyaluronan-mediated motility receptor gene single nucleotide polymorphisms and risk of breast cancer. Cancer Epidemiol Biomark Prev. 2008;17(12):3618–20. https://doi.org/10.1158/1055-9965.EPI-08-0216.
Lee SJ, Jeon HS, Jang JS, Park SH, Lee GY, Lee BH, Kim CH, Kang YM, Lee WK, Kam S, et al. DNMT3B polymorphisms and risk of primary lung cancer. Carcinogenesis. 2005;26(2):403–9. https://doi.org/10.1093/carcin/bgh307.
Cauchi S, Stücker I, Solas C, Laurent-Puig P, Cénée S, Hémon D, Jacquet M, Kremers P, Beaune P, Massaad-Massade L. Polymorphisms of human aryl hydrocarbon receptor (AhR) gene in a French population: relationship with CYP1A1 inducibility and lung cancer. Carcinogenesis. 2001;22(11):1819–24.
Frank B, Hoeft B, Hoffmeister M, Linseisen J, Breitling LP, Chang-Claude J, Brenner H, Nieters A. Association of hydroxyprostaglandin dehydrogenase 15-(NAD) (HPGD) variants and colorectal cancer risk. Carcinogenesis. 2011;32(2):190–6. https://doi.org/10.1093/carcin/bgq231.
Samowitz WS, Wolff RK, Curtin K, Sweeney C, Ma KN, Andersen K, Levin TR, Slattery ML. Interactions between CYP2C9 and UGT1A6 polymorphisms and nonsteroidal anti-inflammatory drugs in colorectal cancer prevention. Clin Gastroenterol Hepatol. 2006;4(7):894–901. https://doi.org/10.1016/j.cgh.2006.04.021.
Percy MJ, Myrie KA, Neeley CK, Azim JN, Ethier SP, Petty EM. Expression and mutational analyses of the human MAD2L1 gene in breast cancer cells. Genes Chromosomes Cancer. 2000;29(4):356–62.
Hashizume R, Fukuda M, Maeda I, Nishikawa H, Oyake D, Yabuki Y, Ogata H, Ohta T. The RING heterodimer BRCA1-BARD1 is a ubiquitin ligase inactivated by a breast cancer-derived mutation. J Biol Chem. 2001;276(18):14537–40. https://doi.org/10.1074/jbc.C000881200.
Kiseljak-Vassiliades K, Zhang Y, Kar A, Razzaghi R, Xu M, Gowan K, Raeburn CD, Albuja-Cruz M, Jones KL, Somerset H, et al. Elucidating the role of the maternal embryonic leucine zipper kinase in adrenocortical carcinoma. Endocrinology. 2018;159(7):2532–44. https://doi.org/10.1210/en.2018-00310.
Grolmusz VK, Karászi K, Micsik T, Tóth EA, Mészáros K, Karvaly G, Barna G, Szabó PM, Baghy K, Matkó J, et al. Cell cycle dependent RRM2 may serve as proliferation marker and pharmaceutical target in adrenocortical cancer. Am J Cancer Res. 2016;6(9):2041–53.
El-Khairi R, Martinez-Aguayo A, Ferraz-de-Souza B, Lin L, Achermann JC. Role of DAX-1 (NR0B1) and steroidogenic factor-1 (NR5A1) in human adrenal function. Endocr Dev. 2011;20:38–46. https://doi.org/10.1159/000321213.
Jarmalaite S, Laurinaviciene A, Tverkuviene J, Kalinauskaite N, Petroska D, Böhling T, Husgafvel-Pursiainen K. Tumor suppressor gene ZAC/PLAGL1: altered expression and loss of the nonimprinted allele in pheochromocytomas. Cancer Genet. 2011;204(7):398–404. https://doi.org/10.1016/j.cancergen.2011.07.002.
Lin M, Morrison CD, Jones S, Mohamed N, Bacher J, Plass C. Copy number gain and oncogenic activity of YWHAZ/14-3-3zeta in head and neck squamous cell carcinoma. Int J Cancer. 2009;125(3):603–11. https://doi.org/10.1002/ijc.24346.
Luo X, Yao J, Nie P, Yang Z, Feng H, Chen P, Shi X, Zou Z. FOXM1 promotes invasion and migration of colorectal cancer cells partially dependent on HSPA5 transactivation. Oncotarget. 2016;7(18):26480–95. https://doi.org/10.18632/oncotarget.8419.
Chakraborty PK, Zhang Y, Coomes AS, Kim WJ, Stupay R, Lynch LD, Atkinson T, Kim JI, Nie Z, Daaka Y. G protein-coupled receptor kinase GRK5 phosphorylates moesin and regulates metastasis in prostate cancer. Cancer Res. 2014;74(13):3489–500. https://doi.org/10.1158/0008-5472.CAN-13-2708.
Zhang Z, Borecki I, Nguyen L, Ma D, Smith K, Huettner PC, Mutch DG, Herzog TJ, Gibb RK, Powell MA, et al. CD83 gene polymorphisms increase susceptibility to human invasive cervical cancer. Cancer Res. 2007;67(23):11202–8. https://doi.org/10.1158/0008-5472.CAN-07-2677.
Rosette C, Roth RB, Oeth P, Braun A, Kammerer S, Ekblom J, Denissenko MF. Role of ICAM1 in invasion of human breast cancer cells. Carcinogenesis. 2005;26(5):943–50. https://doi.org/10.1093/carcin/bgi070.
Gonzalez-Herrera A, Salgado-Bernabe M, Velazquez-Velazquez C, Salcedo-Vargas M, Andrade-Manzano A, Avila-Moreno F, Pina-Sanchez P. Increased expression of HOXB2 and HOXB13 proteins is associated with HPV infection and cervical cancer progression. Asian Pac J Cancer Prev. 2015;16(4):1349–53.
Pressinotti NC, Klocker H, Schäfer G, Luu VD, Ruschhaupt M, Kuner R, Steiner E, Poustka A, Bartsch G, Sültmann H. Differential expression of apoptotic genes PDIA3 and MAP3K5 distinguishes between low- and high-risk prostate cancer. Mol Cancer. 2009;8:130. https://doi.org/10.1186/1476-4598-8-130.
Sasaki A, Yumita S, Kimura S, Miura Y, Yoshinaga K. Immunoreactive corticotropin-releasing hormone, growth hormone-releasing hormone, somatostatin, and peptide histidine methionine are present in adrenal pheochromocytomas, but not in extra-adrenal pheochromocytoma. J Clin Endocrinol Metab. 1990;70(4):996–9. https://doi.org/10.1210/jcem-70-4-996.
Humphries B, Wang Z, Li Y, Jhan JR, Jiang Y, Yang C. ARHGAP18 downregulation by miR-200b suppresses metastasis of triple-negative breast cancer by enhancing activation of RhoA. Cancer Res. 2017;77(15):4051–64. https://doi.org/10.1158/0008-5472.CAN-16-3141.
Chai SJ, Yap YY, Foo YC, Yap LF, Ponniah S, Teo SH, Cheong SC, Patel V, Lim KP. Identification of four-jointed box 1 (FJX1)-specific peptides for immunotherapy of nasopharyngeal carcinoma. PLoS ONE. 2015;10(11):e0130464. https://doi.org/10.1371/journal.pone.0130464.
Bae SY, Choi SK, Kim KR, Park CS, Lee SK, Roh HK, Shin DW, Pie JE, Woo ZH, Kang JH. Effects of genetic polymorphisms of MDR1, FMO3 and CYP1A2 on susceptibility to colorectal cancer in Koreans. Cancer Sci. 2006;97(8):774–9. https://doi.org/10.1111/j.1349-7006.2006.00241.x.
Ding YB, Chen GY, Xia JG, Zang XW, Yang HY, Yang L. Association of VCAM-1 overexpression with oncogenesis, tumor angiogenesis and metastasis of gastric carcinoma. World J Gastroenterol. 2003;9(7):1409–14. https://doi.org/10.3748/wjg.v9.i7.1409.
Yan DW, Li DW, Yang YX, Xia J, Wang XL, Zhou CZ, Fan JW, Wen YG, Sun HC, Wang Q, et al. Ubiquitin D is correlated with colon cancer progression and predicts recurrence for stage II-III disease after curative surgery. Br J Cancer. 2010;103(7):961–9. https://doi.org/10.1038/sj.bjc.6605870.
Tsukishiro S, Suzumori N, Nishikawa H, Arakawa A, Suzumori K. Use of serum secretory leukocyte protease inhibitor levels in patients to improve specificity of ovarian cancer diagnosis. Gynecol Oncol. 2005;96(2):516–9. https://doi.org/10.1016/j.ygyno.2004.10.036.
Mullarky E, Mattaini KR, Vander Heiden MG, Cantley LC, Locasale JW. PHGDH amplification and altered glucose metabolism in human melanoma. Pigment Cell Melanoma Res. 2011;24(6):1112–5. https://doi.org/10.1111/j.1755-148X.2011.00919.x.
Wegman P, Elingarami S, Carstensen J, Stål O, Nordenskjöld B, Wingren S. Genetic variants of CYP3A5, CYP2D6, SULT1A1, UGT2B15 and tamoxifen response in postmenopausal patients with breast cancer. Breast Cancer Res. 2007;9(1):R7. https://doi.org/10.1186/bcr1640.
Shulby SA, Dolloff NG, Stearns ME, Meucci O, Fatatis A. CX3CR223-fractalkine expression regulates cellular mechanisms involved in adhesion, migration, and survival of human prostate cancer cells. Cancer Res. 2004;64(14):4693–8. https://doi.org/10.1158/0008-5472.CAN-03-3437.
Yuan F, Zhang Y, Ma L, Cheng Q, Li G, Tong T. Enhanced NOLC1 promotes cell senescence and represses hepatocellular carcinoma cell proliferation by disturbing the organization of nucleolus. Aging Cell. 2017;16(4):726–37. https://doi.org/10.1111/acel.12602.
Kleeff J, Ishiwata T, Friess H, Büchler MW, Israel MA, Korc M. The helix-loop-helix protein Id2 is overexpressed in human pancreatic cancer. Cancer Res. 1998;58(17):3769–72.
Ko HL, Wang YS, Fong WL, Chi MS, Chi KH, Kao SJ. Apolipoprotein C1 (APOC1) as a novel diagnostic and prognostic biomarker for lung cancer: a marker phase I trial. Thorac Cancer. 2014;5(6):500–8. https://doi.org/10.1111/1759-7714.12117.
Yang Z, Zhuang L, Szatmary P, Wen L, Sun H, Lu Y, Xu Q, Chen X. Upregulation of heat shock proteins (HSPA12A, HSP90B1, HSPA4, HSPA5 and HSPA6) in tumour tissues is associated with poor outcomes from HBV-related early-stage hepatocellular carcinoma. Int J Med Sci. 2015;12(3):256–63. https://doi.org/10.7150/ijms.10735.
Vedeld HM, Andresen K, Eilertsen IA, Nesbakken A, Seruca R, Gladhaug IP, Thiis-Evensen E, Rognum TO, Boberg KM, Lind GE. The novel colorectal cancer biomarkers CDO1, ZSCAN18 and ZNF331 are frequently methylated across gastrointestinal cancers. Int J Cancer. 2015;136(4):844–53. https://doi.org/10.1002/ijc.29039.
Awadallah SM, Atoum MF. Haptoglobin polymorphism in breast cancer patients form Jordan. Clin Chim Acta. 2004;341(1–2):17–21. https://doi.org/10.1016/j.cccn.2003.10.032.
Szczyrba J, Nolte E, Hart M, Döll C, Wach S, Taubert H, Keck B, Kremmer E, Stöhr R, Hartmann A, et al. Identification of ZNF217, hnRNP-K, VEGF-A and IPO7 as targets for microRNAs that are downregulated in prostate carcinoma. Int J Cancer. 2013;132(4):775–84. https://doi.org/10.1002/ijc.27731.
Vasiljević N, Ahmad AS, Beesley C, Thorat MA, Fisher G, Berney DM, Møller H, Yu Y, Lu YJ, Cuzick J, et al. Association between DNA methylation of HSPB1 and death in low Gleason score prostate cancer. Prostate Cancer Prostatic Dis. 2013;16(1):35–40. https://doi.org/10.1038/pcan.2012.47.
Unoki M, Daigo Y, Koinuma J, Tsuchiya E, Hamamoto R, Nakamura Y. UHRF1 is a novel diagnostic marker of lung cancer. Br J Cancer. 2010;103(2):217–22. https://doi.org/10.1038/sj.bjc.6605717.
Chen JH, Liang YX, He HC, Chen JY, Lu JM, Chen G, Lin ZY, Fu X, Ling XH, Han ZD, et al. Overexpression of PDZ-binding kinase confers malignant phenotype in prostate cancer via the regulation of E2F1. Int J Biol Macromol. 2015;81:615–23. https://doi.org/10.1016/j.ijbiomac.2015.08.048.
Alrfaei BM, Vemuganti R, Kuo JS. microRNA-100 targets SMRT/NCOR2, reduces proliferation, and improves survival in glioblastoma animal models. PLoS ONE. 2013;8(11):e80865. https://doi.org/10.1371/journal.pone.0080865.
Morrison JA, Pike LA, Sams SB, Sharma V, Zhou Q, Severson JJ, Tan AC, Wood WM, Haugen BR. Thioredoxin interacting protein (TXNIP) is a novel tumor suppressor in thyroid cancer. Mol Cancer. 2014;13:62. https://doi.org/10.1186/1476-4598-13-62.
Ishida M, Sunamura M, Furukawa T, Lefter LP, Morita R, Akada M, Egawa S, Unno M, Horii A. The PMAIP1 gene on chromosome 18 is a candidate tumor suppressor gene in human pancreatic cancer. Dig Dis Sci. 2008;53(9):2576–82. https://doi.org/10.1007/s10620-007-0154-1.
Maacke H, Opitz S, Jost K, Hamdorf W, Henning W, Krüger S, Feller AC, Lopens A, Diedrich K, Schwinger E, et al. Over-expression of wild-type Rad51 correlates with histological grading of invasive ductal breast cancer. Int J Cancer. 2000;88(6):907–13.
Nadin SB, Cuello-Carrión FD, Sottile ML, Ciocca DR, Vargas-Roig LM. Effects of hyperthermia on Hsp27 (HSPB1), Hsp72 (HSPA1A) and DNA repair proteins hMLH1 and hMSH2 in human colorectal cancer hMLH1-deficient and hMLH1-proficient cell lines. Int J Hyperthermia. 2012;28(3):191–201. https://doi.org/10.3109/02656736.2011.638962.
Gao ZH, Suppola S, Liu J, Heikkilä P, Jänne J, Voutilainen R. Association of H19 promoter methylation with the expression of H19 and IGF-II genes in adrenocortical tumors. J Clin Endocrinol Metab. 2002;87(3):1170–6. https://doi.org/10.1210/jcem.87.3.8331.
Pong WW, Walker J, Wylie T, Magrini V, Luo J, Emnett RJ, Choi J, Cooper ML, Griffith M, Griffith OL, et al. F11R is a novel monocyte prognostic biomarker for malignant glioma. PLoS ONE. 2013;8(10):e77571. https://doi.org/10.1371/journal.pone.0077571.
Rogers CD, Heijden MS, Brune K, Yeo CJ, Hruban RH, Kern SE, Goggins M. The genetics of FANCC and FANCG in familial pancreatic cancer. Cancer Biol Ther. 2004;3(2):167–9.
Moullan N, Cox DG, Angèle S, Romestaing P, Gérard JP, Hall J. Polymorphisms in the DNA repair gene XRCC1, breast cancer risk, and response to radiotherapy. Cancer Epidemiol Biomark Prev. 2003;12(11 Pt 1):1168–74.
Acknowledgements
I thank Demeure, Arizona State University, 1001 S. McAllister Ave, Tempe, AZ, USA very much, the author who deposited their microarray dataset, GSE19775, into the public GEO database.
Author information
Authors and Affiliations
Contributions
Ali Mohamed Alshabi associated with project administration, methodology, and review and editing; Ali Mohamed Alshabi, Ibrahim Ahmed Shaikh, and Basavaraj Vastrad participated in writing original draft and investigation; and Chanabasayya Vastrad performed software, supervision, formal analysis, and validation. Basavaraj Vastrad and Ali Mohamed Alshabi made substantial contributions to acquisition of resources, formal analysis, and validation.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed consent
No informed consent because this study does not contain any human or animal participants.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Alshabi, A.M., Vastrad, B., Shaikh, I.A. et al. Identification of important invasion and proliferation related genes in adrenocortical carcinoma. Med Oncol 36, 73 (2019). https://doi.org/10.1007/s12032-019-1296-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12032-019-1296-7