Abstract
The aim of this study was to evaluate the frequency (as qualitative analysis) and level (as quantitative analysis) of promoter hypermethylation of four genes, P16, TSHR, RASSF1A and RARβ2, and to assess their diagnostic or prognostic values in papillary thyroid tumors. Fifty formalin-fixed paraffin-embedded (FFPE) samples consisting of 25 malignant tumors and 25 non-malignant thyroid tumors were analyzed using COBRA method. Promoter hypermethylation of P16, TESH, RASSF1A and RARB2 genes was noted not only in 10, 7, 19 and 13 cases of malignant tumors, but also it was detected in 7, 11, 23 and 8 cases of benign tumors, respectively, limiting its diagnostic usefulness. The quantitative hypermethylation level was significantly higher in malignant tumors compared to benign tumors for P16 (P < 0.004), TSHR (P < 0.006) and RASSF1A (P < 0.001), but the methylation level of RARβ2 (P < 0.31) showed considerable overlap between the two groups. The mean levels of hypermethylation of P16, TSHR and RASSF1A genes were significantly higher in malignant papillary thyroid tumors compared to benign tumors and by choosing the appropriate cutoff for each gene, we could distinguish 9, 9 and 8 PTCs from 25 cases by P16, RASSF1A and TSHR methylation analysis, respectively. According to our results, these three genes, in combination, may be useful as molecular markers. The findings of present study imply that the P16, TSHR and RASSF1A gene promoter hypermethylation may play important roles in the pathogenesis of PTC and can be a potential biomarker for selecting patients with PTC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Thyroid cancers are the most common malignancies of the endocrine system. PTCs (papillary thyroid cancer) are the most frequent histological types of thyroid malignancies. It is estimated that 5–10% of the general population will develop a clinically significant thyroid nodule during their lifetime [1]. Fine needle aspiration (FNA) biopsy is a widely used diagnostic procedure for thyroid nodules. It is the best and cost-effective method for primary evaluation of the thyroid nodules, but unfortunately, about 20% of the cytological findings are reported as indeterminate, especially for follicular subtypes, and cannot differentiate malignant from benign tumors [2, 3]. At present, most of these patients must undergo surgery, but only minorities of them are found to harbor malignant tumor. To reduce unnecessary thyroid surgery, molecular biomarkers could help to distinguish malignancies from benign lesions. Some types of molecular biomarkers that could help to better assess thyroid nodules include BRAF mutations [4], promoter hypermethylation [5], mRNA expression [6] and IHC [7] in FNA or tissue biopsy of thyroid nodules.
Abnormal methylation of the CpG islands located in the promoter region of genes has been described as an alternative mechanism to gene inactivation in many cancer types. In this regard, the number of hypermethylated genes is estimated to be high in some tumors [8].
The P16 (CDKN2a) gene encodes a tumor suppressor protein that controls two critical cell-cycle control pathways, P16-Rb and P14-P53 [9]. The P16 gene is one of the most commonly studied candidates in the pathogenesis of human neoplasm, and its defects have been demonstrated in different human malignancies, including thyroid cancers [10]. A mechanism of the P16 gene inactivation is promoter hypermethylation [11]. The results of previous studies have shown that methylation of P16 gene is observed in malignant and benign thyroid tumors [11, 12], although some researchers have reported it only in malignant thyroid tumors [10, 13].
TSHR plays a fundamental role in the regulation of thyrocyte function and growth. It stimulates several key steps in thyrocyte concentration of iodine, including its uptake by NIS and oxidation before incorporation into thyroglobulin by thyroid peroxidase. The normal iodine intake is stimulated by TSHR [14]. Previous reports have indicated that TSHR promoter hypermethylation occurs frequently in thyroid tumors [12, 15]. Smith et al. have reported that TSHR promoter methylated more frequently in well-differentiated thyroid cancer than in benign controls [5].
The RASSF1A gene product functions as a negative regulator of cell proliferation through inhibition of G1/S-phase progression [16]. The growth arrest correlates with inhibition of cyclin-D1 protein accumulation, which likely prevents RASSF1A-expressing cells from passing through the RB family cell-cycle restriction point and entering S-phase. RASSF1A knockout mice are susceptible to develop different type of cancers [17]. The RASSF1A locus is epigenetically inactivated at high frequency in a variety of tumors, including malignant and benign thyroid tumors [13, 18, 19].
The RARβ2 gene encodes retinoic acid receptor beta, a member of the thyroid–steroid hormone receptor family of nuclear transcriptional regulators. It binds retinoic acid, which mediates cellular signaling during embryonic morphogenesis, cell growth and differentiation [20]. Previous studies had shown that RARβ2 promoter hypermethylation takes place frequently in malignant and benign thyroid neoplasm [12, 18], although the results of a study have been shown that promoter hypermethylation of RARβ2 gene take place just in malignant thyroid tumors [13].
In the present study, we aimed to investigate the utility of a set of four independent genes (P16, TSHR, RASSF1A and RARβ2) promoter hypermethylation analysis as a diagnostic or prognostic tool in papillary thyroid tumors. These genes were chosen because of their role in thyroid function and tumorigenesis.
Materials and method
Thyroid samples
This work was conducted from 2007 through 2008. A total of 50 formalin-fixed paraffin-embedded (FFPE) samples were obtained from patients who underwent thyroidectomy surgery from 2006 to 2007 in referral Shariati Hospital, Tehran University of Medical Sciences, Tehran, Iran. The tissue samples of total of 50 patients, consisting of 25 malignant tumors and 25 non-malignant samples, were reviewed by a pathologist.
This study was approved by the Ethics and Research Committee of Tehran University of Medical Sciences and was conducted in accordance with the Declaration of Helsinki principles.
Laser capture microdissection (LCM) and DNA extraction
Six-μM FFPE tissue sections were deparaffinized and stained with methyl green (Sigma–Aldrich, St. Louis, MO) and were then dissected manually or by using laser capture microdissection system (Leica AS LMD, Wetzler, Germany). DNA from dissected FFPE samples was extracted by using QIAamp DNA Micro Kit (QIAGEN, Hilden, Germany) according to the manufacturer’s instructions. Quality and quantity of the extracted DNAs were determined by spectrophotometry (NanoDrop ND-1000, Wilmington, Delaware USA).
Bisulfite treatment
Sodium Bisulfite modification was performed using the EZ DNA methylation kit (Zymogen, CA, USA) according to the manufacturer’s protocol. The CpGenome™ universal methylated DNA (Millipore) and peripheral blood lymphocyte (PBL) DNA were used as positive and negative controls, respectively.
COBRA
Combined bisulfite restriction analysis (COBRA) method was used for qualitative and quantitative promoter hypermethylation detection. We amplified P16, TSHR, RASSF1A and RARβ2 CpG islands by using primers listed in Table 1. We have used the Methprimer (http://www.urogene.org/methprimer/) and MethBLAST (http://medgen.ugent.be/methBLAST/) Web sites for designing and checking of the primers, respectively.
PCR was carried out with the following conditions: initial denaturation at 95°C for 5 min, followed by 50 cycles of denaturation at 95°C for 30 s, annealing temperature at different degrees for each gene for 30 s, and elongation at 72°C for 30 s, and a final extension at 72°C for 5 min. The PCR products were visualized on 8% polyacrylamide gels with ethidium bromide staining.
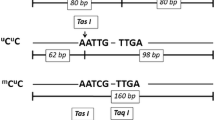
Then, the amplification products were digested by TaqI, RsaI or BstUI (New England Biolabs, Ipswich, MA) restriction endonuclease as noted in Table 1 and Fig. 1. These enzymes cut only methylated alleles. After that, digested products were runs on PAGE (8% 19:1 polyacrylamide, TAE 1×) and stained by CyberGold (invitrogen). Presence of any digested bands was supposed as positive for qualitative promoter hypermethylation, and the quantitative level of hypermethylation was measured by density analysis of bands and by using the following formula, percent of methylation = 100% * (band density of digested band/band density of undigested band) [21].
Statistical analysis
All the statistical analyses were performed by the SPSS software (SPSS version 11.0, Chicago, IL, USA). T-test and chi-square test were used for association of qualitative and quantitative promoter methylation status and clinicopathologic data. P-value of less than 0.05 was considered to be significant.
Results
Characteristics of patients
Fifty FFPE samples including 25 cases of malignant papillary tumors (20 CV-PCT, 2 TC-PTC, 1 FV-PTC and 2 UTC) and 25 benign cases (11 GT, 9 HT, 6 FA and 3 normal) were included in this study. The mean age in malignant and benign group was 56 and 45 years, respectively.
Distribution of qualitative promoter hypermethylation
The frequencies of aberrant promoter methylation of the four genes in benign and malignant groups are shown in Table 2. Promoter hypermethylation of P16, TESH, RASSF1A and RARB2 genes was noted not only in 10, 7, 19 and 13 cases of malignant tumors, but also it was detected in 7, 11, 23 and 8 cases of benign tumors, respectively, limiting its diagnostic usefulness (Table 2).
The detailed stratified analysis was done to determine the distributions of methylation status according to the selected demographic and clinical characteristics, including age, gender, tumor invasion and lymph node metastasis, and the results showed that there is not any statistically difference in promoter hypermethylation between benign and malignant groups.
Distribution of quantitative promoter hypermethylation
The quantitative hypermethylation level was significantly higher in malignant tumors, compared to benign tumors for P16 (P < 0.004), TSHR (P < 0.006) and RASSF1A (0.001), but the methylation level of RARβ2 (P < 0.31) showed considerable overlap between the two groups (Fig. 2).
Table 2 shows the analysis results of the level of quantitative methylation of four genes according to age, gender, invasion and metastasis status at the time of diagnosis.
Mean level of promoter hypermethylation of P16, TSHR, RASSF1A and RARβ2 was 27.2, 5.4, 28.3 and 4.2% for malignant tumors and 3.6, 1.0, 11.4 and 5.7% for benign tumors, respectively (Table 3).
The promoter hypermethylation levels of the four genes in malignant group were analyzed according to gender, age, recurrent of malignancy and lymph node metastasis status, and generally, the results showed that there was no prominent significant difference between methylation level and the clinicopathologic and demographic status (Table 3). However, promoter hypermethylation in older patients showed a higher level for P16 (P = 0.02), but there was no association between P16 promoter hypermethylation level and sex of patients (Table 3).
In the three malignant cases that two of them were UTCs, the three genes (P16, RASSF1A and TSHR) were highly methylated. This suggests a possible role for hypermethylation of these three genes in the more aggressiveness of tumors such as UTCs.
Discussion
Methylation of DNA is important in the genetic regulation of mammalian cells’ activity. The methylation status of CpG islands has been shown to be involved in inactivation of tumor suppressor genes [22].
The promoter CpG islands of genes in normal cells are generally protected from hypermethylation, but this protection may be lost early in tumorigenesis. This has led to the use of hypermethylation of tumor suppressor genes, in particular, as biomarkers for the early diagnosis of cancers [23].
The aim of this study was to evaluate the frequency (as qualitative analysis) and level (as quantitative analysis) of promoter hypermethylation of a set of four independent genes, P16, TSHR, RASSF1A and RARβ2, and to assess their diagnostic or prognostic values in papillary thyroid tumors. The RASSF1A gene was found to be the most frequently methylated gene in the thyroid neoplasms.
The qualitative analysis of promoter hypermethylation showed that epigenetic alteration of genomic DNA is a common finding in thyroid tissue, regardless of histological tumor subtype.
We showed that there were prominent overlaps in hypermethylation in all the four genes analyzed in this study between malignant and benign tumors, and unfortunately, many benign tumors showed promoter hypermethylation too, limiting their diagnostic or prognostic utility in papillary thyroid tumors. The qualitative results of the present study are in agreement with previous studies that detected promoter hypermethylation in P16 [11, 24], TSHR [5], RASSF1A [18] and RARβ2 genes in both benign and malignant papillary thyroid tumors. The presence of TSHR methylation, however, in benign tumors is in contrast to the finding by Xing et al. who have reported that TSHR promoter hypermethylation to occur only in malignant thyroid tumors [15].
In the present study, we used COBRA method that could detect methylation status in two kinds, qualitative and quantitative. The quantitative analysis showed better results for the discrimination of malignant tumors in three genes including P16, RASSF1A and TSHR genes.
The mean levels of hypermethylation of P16, TSHR and RASSF1A genes were significantly higher in malignant papillary thyroid tumors compared to benign tumors and by choosing the appropriate cutoff for each gene, we could distinguish 9, 9 and 8 PTCs from 25 cases by P16, RASSF1A and TSHR methylation analysis, respectively. Interestingly, by the combination of their hypermethylation profile, we could clearly distinguish 17 (66%) of 25 papillary malignant tumors.
We found a considerable overlap in quantitative methylation level of RARβ2 gene between benign and malignant tumors, limiting its diagnosis utility in papillary thyroid tumors.
Interestingly, in three malignant cases, all the three genes were hypermethylated and 2 (66%) of them were UTCs, suggesting that the hypermethylation of these three genes might account for the severity of this carcinoma, which is among the most aggressive types of human cancer.
The quantitative analysis of promoter hypermethylation of the RARβ2 gene showed that there is prominent overlap between benign and malignant thyroid tumor and the results are in consistent with previous reports [12].
The results of this study also showed that quantitative analysis of promoter hypermethylation was better than qualitative analysis and could differentiate between thyroid malignant and benign tumors.
As shown by previous studies, the methylation analysis can be performed on serum DNA to detect malignancies [23]. According to our results, these three genes, in combination, may be useful as molecular markers. We plan to continue the investigation by performing promoter hypermethylation assay on extracted DNA from serum of patients to assess their values for early detection or monitoring of metastasis in patients after total thyroidectomy.
We could detect promoter hypermethylation in both benign and malignant tumors, while in a higher level in malignant tumors. This supports the theory of adenoma to carcinoma sequence in thyroid tumorigenesis. The results of this study suggest that promoter methylation of P16, RASSF1A and TSHR genes are early events in the multistep tumorigenesis in papillary thyroid cancers. These results are consistent with reports that have shown graduate increment of promoter hypermethylation during adenoma to carcinoma progress in thyroid tumors [11]. We were unable to find any relationship between promoter hypermethylation of P16, TSHR, RASSF1A and RARβ2 genes and prognostic factors, possibly due to our small sample size. Future studies are required to clarify the role of aberrant hypermethylation patterns in the genesis and progression of thyroid cancer.
The findings of present study imply that the P16, TSHR and RASSF1A gene promoter hypermethylation may play important roles in the pathogenesis of PTC and can be a potential biomarker for selecting patients with PTC.
References
Mazzaferri EL. Management of a solitary thyroid nodule. N Engl J Med. 1993;328(8):553–9.
Belfiore A, La Rosa GL. Fine-needle aspiration biopsy of the thyroid. Endocrinol Metab Clin North Am. 2001;30(2):361–400.
McCaffrey TV. Evaluation of the thyroid nodule. Cancer Control. 2000;7(3):223–8.
Mohammadi-asl J, Larijani B, Khorgami Z, Tavangar SM, Haghpanah V, Mehdipour P. Prevalence of BRAFV600E mutation in iranian patients with papillary thyroid carcinoma: a single-center study. J Appl Sci. 2009;9(19):3593–7.
Smith JA, Fan CY, Zou C, Bodenner D, Kokoska MS. Methylation status of genes in papillary thyroid carcinoma. Arch Otolaryngol Head Neck Surg. 2007;133(10):1006–11.
Fryknas M, Wickenberg-Bolin U, Goransson H, Gustafsson MG, Foukakis T, Lee JJ, et al. Molecular markers for discrimination of benign and malignant follicular thyroid tumors. Tumor Biol. 2006;27(4):211–20.
Haghpanah V, Shooshtarizadeh P, Heshmat R, Larijani B, Tavangar SM. Immunohistochemical analysis of survivin expression in thyroid follicular adenoma and carcinoma. Appl Immunohistochem Mol Morphol. 2006;14(4):422–5.
Herman JG. Epigenetic changes in cancer and preneoplasia. Cold Spring Harb Symp Quant Biol. 2005;70:329–33.
Robertson KD, Jones PA. Tissue-specific alternative splicing in the human INK4a/ARF cell cycle regulatory locus. Oncogene. 1999;18(26):3810–20.
Lam AK, Lo CY, Leung P, Lang BH, Chan WF, Luk JM. Clinicopathological roles of alterations of tumor suppressor gene p16 in papillary thyroid carcinoma. Ann Surg Oncol. 2007;14(5):1772–9.
Boltze C, Zack S, Quednow C, Bettge S, Roessner A, Schneider-Stock R. Hypermethylation of the CDKN2/p16INK4A promotor in thyroid carcinogenesis. Pathol Res Pract. 2003;199(6):399–404.
Hoque MO, Rosenbaum E, Westra WH, Xing M, Ladenson P, Zeiger MA, et al. Quantitative assessment of promoter methylation profiles in thyroid neoplasms. J Clin Endocrinol Metab. 2005;90(7):4011–8.
Schagdarsurengin U, Gimm O, Dralle H, Hoang-Vu C, Dammann R. CpG island methylation of tumor-related promoters occurs preferentially in undifferentiated carcinoma. Thyroid. 2006;16(7):633–42.
Kopp P. The TSH receptor and its role in thyroid disease. Cell Mol Life Sci. 2001;58(9):1301–22.
Xing M, Usadel H, Cohen Y, Tokumaru Y, Guo Z, Westra WB, et al. Methylation of the thyroid-stimulating hormone receptor gene in epithelial thyroid tumors: a marker of malignancy and a cause of gene silencing. Cancer Res. 2003;63(9):2316–21.
Shivakumar L, Minna J, Sakamaki T, Pestell R, White MA. The RASSF1A tumor suppressor blocks cell cycle progression and inhibits cyclin D1 accumulation. Mol Cell Biol. 2002;22(12):4309–18.
Tommasi S, Dammann R, Zhang Z, Wang Y, Liu L, Tsark WM, et al. Tumor susceptibility of Rassf1a knockout mice. Cancer Res. 2005;65(1):92–8.
Schagdarsurengin U, Gimm O, Hoang-Vu C, Dralle H, Pfeifer GP, Dammann R. Frequent epigenetic silencing of the CpG island promoter of RASSF1A in thyroid carcinoma. Cancer Res. 2002;62(13):3698–701.
Nakamura N, Carney JA, Jin L, Kajita S, Pallares J, Zhang H, et al. RASSF1A and NORE1A methylation and BRAFV600E mutations in thyroid tumors. Lab Invest. 2005;85(9):1065–75.
Soprano DR, Qin P, Soprano KJ. Retinoic acid receptors and cancers. Annu Rev Nutr. 2004;24:201–21.
Xiong Z, Laird PW. COBRA: a sensitive and quantitative DNA methylation assay. Nucleic Acids Res. 1997;25(12):2532–4.
Konishi N, Nakamura M, Kishi M, Nishimine M, Ishida E, Shimada K. Heterogeneous methylation and deletion patterns of the INK4a/ARF locus within prostate carcinomas. Am J Pathol. 2002;160(4):1207–14.
Hu S, Ewertz M, Tufano RP, Brait M, Carvalho AL, Liu D, et al. Detection of serum deoxyribonucleic acid methylation markers: a novel diagnostic tool for thyroid cancer. J Clin Endocrinol Metab. 2006;91(1):98–104.
Ishida E, Nakamura M, Shimada K, Higuchi T, Takatsu K, Yane K, et al. DNA hypermethylation status of multiple genes in papillary thyroid carcinomas. Pathobiology. 2007;74(6):344–52.
Acknowledgments
Authors would like to thank School of Medicine, Tehran University of Medical Sciences for the financial supports.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mohammadi-asl, J., Larijani, B., Khorgami, Z. et al. Qualitative and quantitative promoter hypermethylation patterns of the P16, TSHR, RASSF1A and RARβ2 genes in papillary thyroid carcinoma. Med Oncol 28, 1123–1128 (2011). https://doi.org/10.1007/s12032-010-9587-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12032-010-9587-z