Abstract
DNA methylation and demethylation play an important role in neuropathic pain. In general, DNA methylation of CpG sites in the promoter region impedes gene expression, whereas DNA demethylation contributes to gene expression. Here, we evaluated the methylation status of CpG sites in genomic DNA promoter regions in dorsal root ganglions (DRGs) of diabetic neuropathic pain (DNP) mice. In our research, streptozotocin (STZ) was intraperitoneally injected into mice to construct DNP models. The DNP mice showed higher fasting blood glucose (above 11.1 mmol/L), lower body weight, and mechanical allodynia than control mice. Whole-genome bisulfite sequencing (WGBS) revealed an altered methylation pattern in CpG sites in the DNA promoter regions in DRGs of DNP mice. The results showed 376 promoter regions with hypermethylated CpG sites and 336 promoter regions with hypomethylated CpG sites. In addition, our data indicated that altered DNA methylation occurs primarily on CpG sites in DNA promoter regions. The Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis revealed that differentially methylated CpG sites annotated genes were involved in activities of the nervous and sensory systems. Enrichment analysis indicated that genes in these pathways contributed to diabetes or pain. In conclusion, our study enriched the role of DNA methylation in DNP.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
According to the International Diabetes Federation data, 9.3% of adults aged 20 to 79 years—a staggering 463 million people—are living with diabetes, and in 2045, the number will increase to 700 million. About 25% of patients with diabetes mellitus suffer from diabetic neuropathic pain (DNP) (Jolivalt et al. 2016; Shillo et al. 2019; Sloan et al. 2018). As an intractable problem accompanied by diabetes mellitus, DNP causes a huge economic and psychological burden on patients. Although DNP has drawn increasing attention, the underlying mechanisms remain unclear. Available data suggest that the risk factors for the generation of DNP include glycemic burden, female gender, obesity, and genotype (variants of voltage-gated sodium channels [VGSC]) (Feldman et al. 2017; Rosenberger et al. 2020; Shillo et al. 2019). In addition, epigenetic regulation could play a vital role in DNP (Cheng et al. 2015; Ciccacci et al. 2020; Guo et al. 2019; Liu et al. 2018b).
Epigenetic regulation includes DNA modification, histone modification, chromatin remodeling, and non-coding RNAs. DNA modification primarily includes DNA methylation and demethylation. In the mammalian genome, DNA methylation refers to the transfer of a methyl group to the C5 position of cytosine to form 5-methylcytosine (5mC) by DNA methyltransferases (Li and Zhang 2014; Moore et al. 2013). DNA methylation can occur at several genetic locations, primarily including promoters, exons, and introns (Jones 2012; Rauscher et al. 2015). DNA methylation of promoters occurs especially on CpG sites, which impairs transcription factor binding and silences gene expression (Moore et al. 2013). Methylated DNA could be demethylated by ten-eleven translocation proteins (He et al. 2011; Kohli and Zhang 2013; Morgan et al. 2018), which can oxidize 5mC to 5-carboxylcytosine via intermediates 5-hydroxymethylcytosine and 5-formylcytosine and initiate the expression of specific genes. Hence, DNA methylation in the right place at the right time plays an important role in normal physiological functions. Altered DNA methylation may contribute to several disease processes (Ehrlich 2019; Liu et al. 2018a ; Luo et al. 2018).
Altered DNA methylation was also implicated in DNP (Zhang et al. 2015). In this research, the authors found demethylation of CpG sites in the p2×3r gene promoter region in the dorsal root ganglion (DRG) of DNP rats, along with an increased expression of purinergic P2X ligand-gated ion channel 3 receptor (P2X3R), annotated by the p2×3r gene. In type 2 diabetes (T2D), DNA methylation was associated with the progression of diabetic peripheral neuropathy (Guo et al. 2019). In their study, Kai Guo et al. found differentially methylated CpG sites between significant sural nerve regeneration and degeneration. These results indicated that DNA methylation and demethylation occurred during the progression of diabetes. Although there are reports about DNA methylation in DNP, the whole-genome DNA methylation profiling in DNP DRGs has not been reported.
In this study, we studied the changes in DNA methylation in genomic DNA promoter regions in DNP mice. We believe this study will provide a better understanding of the role of DNA methylation in DNP.
Materials and Methods
Animals. Male C57BL/6 wild-type mice (18–22 g) were purchased from Capital Medical University. Four mice were housed per cage in a temperature- and light-controlled room with a 12-h:12-h light: dark cycle. Mice were provided water and food ad libitum. All animal experimental procedures were approved by the experimental animal ethics committee of Capital Medical University.
Establishment of Type 1 Diabetics in Mice. Mice were fasted overnight and provided free access to water. Next, mice were administered an intraperitoneal injection of freshly prepared streptozotocin (STZ, 50 mg/kg/day; Sigma, S0130) for five consecutive days. Control mice received an intraperitoneal injection of citrate buffer (5 mL/kg/day) (Zheng et al. 2018) for five days. The fasting blood glucose (mmol/L) was detected at the fourth week using the blood sample collected from the tail vein using the ACCU-CHEK® Mobile blood glucose meter (Roche). When the blood glucose meter showed “high,” the data were recorded as 33.3 mmol/L. Mice with fasting blood glucose over 11.1 mmol/L were considered diabetic and used for subsequent studies. In addition, the body weight of mice was recorded.
Paw Withdrawal Threshold. The 50% paw withdrawal threshold of hind paws was measured using calibrated von Frey hairs (Stoelting, Wood Dale, IL, USA). Briefly, mice were placed on a metal mesh floor and covered with transparent plastic cages. Before the test, mice were allowed to habituate for at least 30 min for three consecutive days. Eight von Frey hairs (0.02, 0.04, 0.07, 0.16, 0.40, 0.60, 1.00, 1.40 g) were selected (Zheng et al. 2017). Each test began with 0.16 g von Frey filament that was applied perpendicularly to the plantar surface in the center of either hind paw for 3 to 5 s. Responses were recorded as positive when mice showed paw withdrawal, flinching, or paw licking. In addition, 50% paw withdrawal threshold (PWT) was measured using the “up-and-down” method as previously described (Chaplan et al. 1994; Chen et al. 2018) and calculated using the following formula: 50% PWT (g) = 10X + kd/104, where X is the value of the final von Frey hair used, k is the value measured from the pattern of positive/negative responses, and d is the average increment (in log units) between von Frey hairs used. If a mouse responded to the lowest von Frey hair, 0.02 g was recorded. If a mouse did not respond to the highest von Frey hair, 2.00 g was recorded.
Whole-Genome Bisulfite Sequencing. Harvested L3–L5 DRGs of mice and mixed DRGs from three mice to obtain sufficient tissue and considered as one sample. The control and DNP groups included three samples. Genomic DNA was extracted from DRGs using the Wizard Genomic DNA Purification Kit (Promega) and sequenced in Igenecode Technology (Beijing, China). The quality of extracted DNA was measured and evaluated. Only high-quality genomic DNA was used for the subsequent study. The genomic DNA was fragmented to about 200 bp by sonication. Bisulfite was used to convert the unmethylated cytosine to uracil, whereas methylated cytosine was unaffected. Finally, PCR amplification was performed. The libraries were quantified and sequenced on Illumina HiSeq. To better understand the function of differentially methylated CpG site annotated genes, Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis was performed (Kanehisa and Goto 2000). The KEGG database is updated daily and is freely available (http://www.genome.ad.jp/kegg/). A hypergeometric test was used for enrichment analysis of the KEGG pathway, and p < 0.05 was considered significant.
Data Analysis. Data are expressed as mean ± standard error of the mean (SEM). GraphPad Prism 7 software was used for statistical analysis. Unpaired Student’s t-test was used for comparison between two groups (Fig. 1). A p < 0.05 was considered significant.
Intraperitoneal injection of STZ to induce DNP in mice. a The level of fasting blood glucose in control (ctrl) and DNP mice. b The body weight of ctrl and DNP mice. c and d Mechanical allodynia in the left and right paws of DNP mice. All data are expressed as mean ± SEM. ***p < 0.001 when compared with ctrl, unpaired t-test
Results
Induction of DNP in Mice
To establish DNP in mice, diabetes was induced in male C57BL/6 wild-type mice by intraperitoneal injection of STZ for five days. After four weeks, mice with STZ treatment showed higher fasting blood glucose (mmol/L) when compared with control mice (Fig. 1a), and the values were 29.30 ± 1.64 mmol/L in STZ-treated mice and 8.58 ± 0.31 mmol/L in control mice (n = 9). STZ-treated mice were significantly lighter in weight (g) than control mice (Fig. 1b), with values 22.71 ± 0.65 g in STZ-treated mice and 27.99 ± 0.56 g in control mice. Frey hairs method used to test the sensitivity to mechanical stimuli revealed that STZ-treated mice showed reduced threshold (g) to mechanical stimulation in both left and right paws (Fig. 1c, d). The values were 0.19 ± 0.04 g in STZ-treated mice and 1.57 ± 0.17 g in control mice for left paws and 0.18 ± 0.05 g in STZ-treated mice versus 1.66 ± 0.15 g in control mice for right paws (n = 9). For all the aforementioned cases, the data are expressed as mean ± SEM, *** p < 0.001, and unpaired t-test. Therefore, STZ-treated mice showed DNP behavior.
Distribution of Differentially Methylated Regions
The methylation of cytosine in different regions had varying effects on gene expression. The analysis of our data indicated that 386 promoter regions, 473 exon regions, and 8896 intron regions were hypermethylated in cytosines, whereas 347 promoter regions, 421 exon regions, and 7858 intron regions were hypomethylated in cytosines in STZ-induced DNP mice when compared with control mice (Fig. 2a). Further analysis of our data revealed that among 386 hypermethylated promoter regions, 376 promoter regions were CpG hypermethylated, and among 347 hypomethylated promoter regions, 336 promoter regions were CpG hypomethylated (Fig. 2b). These results indicate that altered DNA methylation in promoter regions primarily occurred in CpG sites. Based on these findings, we conclude that differentially methylated CpG sites play an important role in the regulation of gene expression.
Kyoto Encyclopedia of Genes and Genomes Enrichment Analysis of Differentially Methylated CpG Sites in Promoter Regions
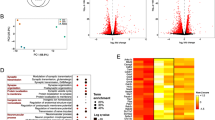
To better understand the function of differentially methylated CpG sites in promoter regions, their annotated genes were analyzed by the KEGG pathway. We found annotated genes in the KEGG pathway that were involved in the sensory system, nervous system, neurodegenerative diseases, and energy metabolism (Fig. 3). Enrichment analysis showed that 13 pathways were significantly enriched (p < 0.05), including “starch and sugar metabolism” and “glycerophospholipid metabolism,” which are involved in diabetes and diabetic peripheral neuropathy (Fig. 4). In addition, we noticed that the “asthma” pathway includes a gene (IL-10) related to pain (Fig. 4; Table 1) (Iwasa et al. 2019; Khan et al. 2015). The CpG sites in the IL-10 gene promoter regions were hypermethylated in DNP mice, which may inhibit IL-10 expression and contribute to DNP. This finding was consistent with that reported in certain previous studies (Khan et al. 2015). The results indicated that altered DNA methylation occurs in genes associated with diabetes or pain in DRG of DNP mice.
Enrichment analysis of KEGG pathways and all 13 significantly enriched pathways are shown. The size of the dot means the number of genes in each pathway. The color means the p-value. Rich factor means the percentage of annotated genes to all genes belonging to the KEGG pathway. A hypergeometric test was used for enrichment analysis of the KEGG pathway. A p < 0.05 was considered significant
Discussion
DNA methylation and demethylation are important regulators of gene expression and play a major role in pain (Chidambaran et al. 2017; Garriga et al. 2018; Gombert et al. 2020; Sun et al. 2015). Although certain reports indicate that under pain conditions, altered DNA methylation occurs in DRG and contributes to pain sensitivity (Garriga et al. 2018; Zhang et al. 2015), genomic DNA methylation status in DRG with DNP has not been reported. We examined the DNA methylation status in the DRG of DNP mice at four weeks using whole-genome bisulfite sequencing (WGBS). The results indicate that altered DNA methylation occurs primarily in intron regions; however, in the upstream regions, namely, promoter regions, altered DNA methylation preferentially occurs in CpG sites (Fig. 2). These findings are consistent with those reported earlier (Guo et al. 2019). We identified 712 promoter regions with differentially methylated CpG sites; among these, 376 were hypermethylated, and 336 were hypomethylated. DNA methylation at different sites had a varying effect on gene expression; for example, methylation in the promoter regions usually impeded gene transcription, whereas in intragenic regions increased the activity of the gene (Guo et al. 2019; Rideout III et al. 1990).
To better understand the function of annotated genes with differentially methylated CpG sites, KEGG analysis was performed. As expected, the annotated genes were involved in the functions of nervous and sensory systems. Enrichment analysis of KEGG pathways revealed that “starch and sugar metabolism” and “glycerophospholipid metabolism” were significantly enriched. Similar to our results, Kai Guo et al., who performed KEGG pathway enrichment analysis, found that “glycerophospholipid metabolism” pathway was significantly enriched. However, they had compared significant sural nerve regeneration and degeneration in type 2 diabetes, whereas we compared healthy and DNP mice in DRG. Reduced sugar and starchy carbohydrate intake can reverse diabetic disorder (Feinmann 2016). In the “glycerophospholipid metabolism” pathway, glycerophospholipids are components of cell membranes and are involved in several neurological diseases, such as Alzheimer’s disease, ischemia, and spinal cord trauma (Farooqui and Horrocks 2001; Frisardi et al. 2011). Another significantly enriched pathway that attracted our attention was the “asthma” pathway, including an annotated gene IL-10. Previous studies reported the involvement of IL-10 in pain. Changes in the IL-10 expression in different tissues under different pain conditions varied. For example, the expression of IL-10 in the paw (skin, muscle, and fascia) increased significantly in Complete Freund’s Adjuvant-induced inflammatory pain (Martins et al. 2016). However, the levels of IL-10 decreased significantly in the affected side of DRGs in chronic constriction injury- and partial sciatic ligation-induced neuropathic pain (Khan et al. 2015). Further analysis of our data revealed that promoter regions of IL-10 were hypermethylated, which could inhibit gene expression. These results suggest the involvement of decreased IL-10 expression in DRG in DNP mice, similar to that observed in chronic constriction injury and partial sciatic ligation neuropathic pain models. Although we found that only the IL-10 gene was directly involved in pain, a report indicated that curcumin alleviated chronic pain induced-depression by modulating glycerophospholipid metabolism and ether lipid metabolism (Zhang et al. 2020). These findings suggested that genes involved in “glycerophospholipid metabolism” and “ether lipid metabolism,” significantly enriched pathways in our research, might also be involved in pain.
Conclusions
We observed altered genomic DNA methylation status in DRGs of DNP mice. Certain genes were found to be related to diabetes, and others were involved in pain conditions. We believe that our work will further enrich the knowledge of epigenetic regulation in DNP and provide potential targets for DNP treatment.
Data Availability
All data and materials described in the study are available from the corresponding author on reasonable request.
Abbreviations
- DNP:
-
Diabetic neuropathic pain
- DRG:
-
Dorsal root ganglion
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- P2X3R:
-
Purinergic P2X ligand-gated ion channel 3 receptor
- PWT:
-
Paw withdrawal threshold
- STZ:
-
Streptozotocin
- WGBS:
-
Whole-genome bisulfite sequencing
References
Chaplan SR, Bach FW, Pogrel JW, Chung JW, Yaksh TL (1994) Quantitative assessment of tactile allodynia in the rat paw. J Neurosci Methods 53(1):55–63. https://doi.org/10.1016/0165-0270(94)90144-9
Chen W, Chi YN, Kang XJ, Liu QY, Zhang HL, Li ZH, Zhao ZF, Yang Y, Su L, Cai J, Liao FF, Yi M, Wan Y, Liu FY (2018) Accumulation of CaV3.2 T-type calcium channels in the uninjured sural nerve contributes to neuropathic pain in rats with spared nerve injury. Front Mol Neurosci 11: 24. https://doi.org/10.3389/fnmol.2018.00024
Cheng C, Kobayashi M, Martinez JA, Ng H, Moser JJ, Wang XL, Singh V, Pharm M, Fritzler MJ, Zochodne DW (2015) Evidence for epigenetic regulation of gene expression and function in chronic experimental diabetic neuropathy. J Neuropathol Exp Neurol 74(8):804–817. https://doi.org/10.1097/NEN.0000000000000219
Chidambaran V, Zhang X, Martin LJ, Ding L, Weirauch MT, Geisler K, Stubbeman BL, Sadhasivam S, Ji H (2017) DNA methylation at the mu-1 opioid receptor gene (OPRM1) promoter predicts preoperative, acute, and chronic postsurgical pain after spine fusion. Pharmgenomics Pers Med 10:157–168. https://doi.org/10.2147/PGPM.S132691
Ciccacci C, Latini A, Colantuono A, Politi C, D’Amato C, Greco C, Rinaldi ME, Lauro D, Novelli G, Spallone V, Borgiani P (2020) Expression study of candidate miRNAs and evaluation of their potential use as biomarkers of diabetic neuropathy. Epigenomics 12(7):575–585. https://doi.org/10.2217/epi-2019-0242
Ehrlich M (2019) DNA hypermethylation in disease: mechanisms and clinical relevance. Epigenetics 14(12):1141–1163. https://doi.org/10.1080/15592294.2019.1638701
Farooqui AA, Horrocks LA (2001) Plasmalogens: workhorse lipids of membranes in normal and injured neurons and glia. Neuroscientist 7(3):232–245. https://doi.org/10.1177/107385840100700308
Feinmann J (2016) Advice on sugar and starch is urged in type 2 diabetes counselling. BMJ 355:i6543. https://doi.org/10.1136/bmj.i6543
Feldman EL, Nave KA, Jensen TS, Bennett DLH (2017) New horizons in diabetic neuropathy: mechanisms, bioenergetics, and pain. Neuron 93(6):1296–1313. https://doi.org/10.1016/j.neuron.2017.02.005
Frisardi V, Panza F, Seripa D, Farooqui T, Farooqui AA (2011) Glycerophospholipids and glycerophospholipid-derived lipid mediators: a complex meshwork in Alzheimer’s disease pathology. Prog Lipid Res 50(4):313–330. https://doi.org/10.1016/j.plipres.2011.06.001
Garriga J, Laumet G, Chen SR, Zhang Y, Madzo J, Issa JPJ, Pan HL, Jelinek J (2018) Nerve injury-induced chronic pain is associated with persistent DNA methylation reprogramming in dorsal root ganglion. J Neurosci 38(27):6090–6101. https://doi.org/10.1523/JNEUROSCI.2616-17.2018
Gombert S, Rhein M, Winterpacht A, Münster T, Hillemacher T, Leffler A, Frieling H (2020) Transient receptor potential ankyrin 1 promoter methylation and peripheral pain sensitivity in Crohn’s disease. Clin Epigenetics 12(1):1. https://doi.org/10.1186/s13148-019-0796-9
Guo K, Elzinga S, Eid S, Figueroa-Romero C, Hinder LM, Pacut C, Feldman EL, Hur J (2019) Genome-wide DNA methylation profiling of human diabetic peripheral neuropathy in subjects with type 2 diabetes mellitus. Epigenetics 14(8):766–779. https://doi.org/10.1080/15592294.2019.1615352
He YF, Li BZ, Li Z, Liu P, Wang Y, Tang Q, Ding J, Jia Y, Chen Z, Li L, Sun Y, Li X, Dai Q, Song CX, Zhang K, He C, Xu GL (2011) Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science 333(6047):1303–1307. https://doi.org/10.1126/science.1210944
Iwasa T, Afroz S, Inoue M, Arakaki R, Oshima M, Raju R, Waskitho A, Inoue M, Baba O, Matsuka Y (2019) IL-10 and CXCL2 in trigeminal ganglia in neuropathic pain. Neurosci Lett 703:132–138. https://doi.org/10.1016/j.neulet.2019.03.031
Jolivalt CG, Frizzi KE, Guernsey L, Marquez A, Ochoa J, Rodriguez M, Calcutt NA (2016) Peripheral neuropathy in mouse models of diabetes. Curr Protoc Mouse Biol 6(3):223–255. https://doi.org/10.1002/cpmo.11
Jones PA (2012) Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat Rev Genet 13(7):484–492. https://doi.org/10.1038/nrg3230
Kanehisa M, Goto S (2000) KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28(1):27–30. https://doi.org/10.1093/nar/28.1.27
Khan J, Ramadan K, Korczeniewska O, Anwer MM, Benoliel R, Eliav E (2015) Interleukin-10 levels in rat models of nerve damage and neuropathic pain. Neurosci Lett 592:99–106. https://doi.org/10.1016/j.neulet.2015.03.001
Kohli RM, Zhang Y (2013) TET enzymes, TDG and the dynamics of DNA demethylation. Nature 502(7472):472–479. https://doi.org/10.1038/nature12750
Li E, Zhang Y (2014) DNA methylation in mammals. Cold Spring Harb Perspect Biol 6(5):a019133. https://doi.org/10.1101/cshperspect.a019133
Liu C, Jiao C, Wang K, Yuan N (2018a) DNA methylation and psychiatric disorders. Prog Mol Biol Transl Sci 157:175–232. https://doi.org/10.1016/bs.pmbts.2018.01.006
Liu CL, Li CC, Deng ZY, Du E, Xu CS (2018b) Long non-coding RNA BC168687 is involved in TRPV1-mediated diabetic neuropathic pain in rats. Neuroscience 374:214–222. https://doi.org/10.1016/j.neuroscience.2018.01.049
Luo C, Hajkova P, Ecker JR (2018) Dynamic DNA methylation: in the right place at the right time. Science 361(6409):1336–1340. https://doi.org/10.1126/science.aat6806
Martins DF, Turnes BL, Cidral-Filho FJ, Bobinski F, Rosas RF, Danielski LG, Petronilho F, Santos ARS (2016) Light-emitting diode therapy reduces persistent inflammatory pain: Role of interleukin 10 and antioxidant enzymes. Neuroscience 324:485–495. https://doi.org/10.1016/j.neuroscience.2016.03.035
Moore LD, Le T, Fan G (2013) DNA methylation and its basic function. Neuropsychopharmacology 38(1):23–38. https://doi.org/10.1038/npp.2012.112
Morgan AE, Davies TJ, Mc Auley MT (2018) The role of DNA methylation in ageing and cancer. Proc Nutr Soc 77(4):412–422. https://doi.org/10.1017/S0029665118000150
Rauscher GH, Kresovich JK, Poulin M, Yan L, Macias V, Mahmoud AM, Al-Alem U, Kajdacsy-Balla A, Wiley EL, Tonetti D, Ehrlich M (2015) Exploring DNA methylation changes in promoter, intragenic, and intergenic regions as early and late events in breast cancer formation. BMC Cancer 15:816. https://doi.org/10.1186/s12885-015-1777-9
Rideout WM III, Coetzee GA, Olumi AF, Jones PA (1990) 5-methylcytosine as an endogenous mutagen in the human LDL receptor and p53 genes. Science 249(4974):1288–1290. https://doi.org/10.1126/science.1697983
Rosenberger DC, Blechschmidt V, Timmerman H, Wolff A, Treede RD (2020) Challenges of neuropathic pain: focus on diabetic neuropathy. J Neural Transm (Vienna) 127(4):589–624. https://doi.org/10.1007/s00702-020-02145-7
Shillo P, Sloan G, Greig M, Hunt L, Selvarajah D, Elliott J, Gandhi R, Wilkinson ID, Tesfaye S (2019) Painful and painless diabetic neuropathies: what is the difference? Curr Diab Rep 19(6):32. https://doi.org/10.1007/s11892-019-1150-5
Sloan G, Shillo P, Selvarajah D, Wu J, Wilkinson ID, Tracey I, Anand P, Tesfaye S (2018) A new look at painful diabetic neuropathy. Diabetes Res Clin Pract 144:177–191. https://doi.org/10.1016/j.diabres.2018.08.020
Sun Y, Sahbaie P, Liang DY, Li W, Shi X, Kingery P, Clark JD (2015) DNA methylation modulates nociceptive sensitization after incision. PLoS ONE 10(11):e0142046. https://doi.org/10.1371/journal.pone.0142046
Zhang L, Ma Z, Wu Z, Jin M, An L, Xue F (2020) Curcumin improves chronic pain induced depression through regulating serum metabolomics in a rat model of trigeminal neuralgia. J Pain Res 13:3479–3492. https://doi.org/10.2147/JPR.S283782
Zhang HH, Hu J, Zhou YL, Qin X, Song ZY, Yang PP, Hu S, Jiang X, Xu GY (2015) Promoted interaction of nuclear factor-kB with demethylated purinergic P2X3 receptor gene contributes to neuropathic pain in rats with diabetes. Diabetes 64(12):4272–4784. https://doi.org/10.2337/db15-0138
Zheng J, Jiang YY, Xu LC, Ma LY, Liu FY, Cui S, Cai J, Liao FF, Wan Y, Yi M (2017) Adult hippocampal neurogenesis along the dorsoventral axis contributes differentially to environmental enrichment combined with voluntary exercise in alleviating chronic inflammatory pain in mice. J Neurosci 37(15):4145–4157. https://doi.org/10.1523/JNEUROSCI.3333-16.2017
Zheng J, Wang Y, Han S, Luo Y, Sun X, Zhu N, Zhao L, Li J (2018) Identification of protein kinase C isoforms involved in type 1 diabetic encephalopathy in mice. J Diabetes Res 2018:8431249. https://doi.org/10.1155/2018/8431249
Funding
This work was supported by the National Natural Science Foundation of China (grant no. 81971037 to F. Yang, 81870865 to W. Cui), the Beijing Natural Science Foundation Program and Scientific Research Key Program of Beijing Municipal Commission of Education (KZ201910025026 to F. Yang), Support Project of High-level Teachers in Beijing Municipal Universities in the Period of 13th Five–year Plan (CIT&TCD201904092 to Q. Li), and Beijing Postdoctoral Research Foundation (grant no. ZZ 2019–01 to W. Chen).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Wen Chen, Fei Yang, and Weihua Cui prepared the manuscript. Wen Chen, Ting Lan, and Qingyu Sun conducted most of the experiments. Yurui Zhang, Danmin Shen, Tingting Hu, Jing Liu, Yingzi Chong, Peipei Wang, and Qian Li analyzed the data and commented on previous publications. All authors approved the final manuscript.
Corresponding author
Ethics declarations
Ethics Approval and Consent to Participate
All animal procedures were approved by the experimental animal ethics committee of Capital Medical University and were performed in accordance with the National Institute of Health Guidelines for the Treatment Animals. The manuscript does not contain any study on human participants.
Consent for Publication
All co-authors have agreed to the submission of the final manuscript.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Chen, W., Lan, T., Sun, Q. et al. Whole Genomic DNA Methylation Profiling of CpG Sites in Promoter Regions of Dorsal Root Ganglion in Diabetic Neuropathic Pain Mice. J Mol Neurosci 71, 2558–2565 (2021). https://doi.org/10.1007/s12031-021-01847-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12031-021-01847-1