Abstract
The neuron-restrictive silencer factor (NRSF) a transcriptional regulator that function as a hub that coordinately regulates multiple aspects of neurogenesis, orchestrates neural differentiation, and preserves the unique neural phenotype. NRSF also acts as an oncogene in neural tumorigenesis, although its effect differs depending on the cell type and tissues. Intriguingly, far more than above functions, potential roles for NRSF and its target genes have also been implicated in the pathogenesis and therapeutic mechanism of neurodegenerative diseases. NRSF acts as a flexible and complicated regulator in nervous system, from transcriptional repressor to activator or modulator, and plays a part in neuronal survival or neuronal death. Here, we present the mechanisms proposed to account for the multiple roles of NRSF in neurogenesis and neurological diseases and discuss the therapeutic perspective of recent advances. The mechanisms underlying this duality of NRSF are helpful to understanding the physiological and pathological conditions of neurons and provide new therapeutic approaches to neurological disorders and diseases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In 1995, two groups independently identified a gene encoding a zinc finger protein that was suggested to function as a master negative regulator of neurogenesis. The transcription factor REST, an RE1-silencing transcription factor (Chong et al. 1995), also known as neuron-restrictive silencer factor (NRSF) (Schoenherr and Anderson 1995) and X2 Box Repressor (XBR) [3] blocks transcription of its target genes by binding to a specific consensus 21 bp RE1 binding site/neuron-restrictive silencer element (RE1/NRSE) that is present in the target genes’ regulatory regions (Schoenherr et al. 1996; Valouev et al. 2008). Occasionally, a non-canonical bipartite RE1/NRSE sequence has been described, consisting in two half-sites separated by 10–16 base pairs (Johnson et al. 2007). Nowadays, more and more studies have expanded the functions of NRSF much beyond its initial role. At the molecular level, NRSF acts cooperatively with other proteins to execute its multiple and broad regulatory roles in neuronal differentiation and development (Soldati et al. 2012; Covey et al. 2012; Gao et al. 2011; Qureshi et al. 2010), such as fine-tuning neural gene expression (Qureshi et al. 2010), modulating synaptic plasticity (Rodenas-Ruano et al. 2012), and keeping maintenance of self-renewal capacity of neural stem cells (NSCs) (Covey et al. 2012). Moreover, NRSF has also been implicated as a suppressor in non-neuronal tumors and as an oncogene in neuronal tumors, such as neuroblastomas, medulloblastomas, and pheochromocytomas (Negrini et al. 2013). Intriguingly, dysfunction of NRSF and aberrations in the regulation of NRSF target genes are closely related to neurological diseases,especially in neurodegeneration (Table 1). Consist with the gene-environment interactions mechanism (Quinn et al. 2013), these findings show that NRSF plays diverse roles in multiple cellular processes in nervous system.
The Biological Aspects of NRSF
As the master regulator of neural cell differentiation in normal physiological condition, NRSF is highly expressed in embryonic stem cells but reduced rapidly in neural progenitors and maintained at very low levels after differentiation (Ooi and Wood 2007; Ballas and Mandel 2005). The low expression of NRSF in mature neurons allows the transcription of a large panel of NRSF target genes, which are necessary for the acquisition of the unique phenotype of neural cells. PC12 pheochromocytoma cells are almost devoid of NRSF (Bruce et al. 2006; D’Alessandro et al. 2008) in which NRSF as a hub of the neurosecretory function (Zhang et al. 2014). In addition, pancreatic β-cells express almost undetectable levels of NRSF (Ballas and Mandel 2005), allowing transcriptional activators to bind and initiate the chromatin of NRSF target genes. In contrast, in neural cell tumors, high levels of NRSF were expressed in medulloblastomas (Fuller et al. 2005), neuroblastomas (Singh et al. 2011), and multiform glioblastomas (Kamal et al. 2012) and were correlated with the proliferation and severity of these tumors (Kamal et al. 2012; Conti et al. 2012). However, in some epithelial cell types, such as in human mammary carcinoma cells (Lv et al. 2010; Wagoner et al. 2010), various colon carcinoma lines (Hatano et al. 2011) and in small-cell carcinomas (SCLCs) of the lung (Kreisler et al. 2010; Coulson et al. 2000), NRSF was identified as a tumor suppressor and expressed in low levels. Therefore, NRSF acts as an oncogene in neural tumors and as a tumor suppressor of carcinomas in the breast, colon, and lung. Taken together, NRSF plays dual, opposing roles in different conditions, which depend not only on the cell type and tissue specific, but also on the concentration of NRSF, the chromatin architecture of the particular genes (Negrini et al. 2013; Thiel et al. 2014). Thus, it is necessary for us to realize the basic structure and binding partners of NRSF (Ooi and Wood 2007) as well as the dynamics of its expression.
The Dynamics of NRSF Expression
Although NRSF is a giant communication hub for the neurons, precisely how NRSF itself is regulated still remains an open question. Changes in its expression can be due to both transcriptional and posttranscriptional processes. At the same time, the dynamics of NRSF levels in neural and non-neural tumors with respect to their cells of origin could be due to miscellaneous regulatory mechanisms. In human embryonic kidney (HEK) cells and neural progenitors, rapid NRSF turnover is mediated by targeting to a proteasomal pathway (Ballas et al. 2005), which is kept in equilibrium by the enzyme regulating its ubiquitination, the ubiquitin ligase SCFβ-TRPC (beta-transducin repeat containing E3 ubiquitin protein ligase) (Guardavaccaro et al. 2008; Westbrook et al. 2008) involving casein kinase 1 (Kaneko et al. 2014), and the deubiquitinase HAUSP (the herpesvirus-associated ubiquitin-specific protease, also known as USP7) (Huang and Bao 2012; Huang et al. 2011a, b). TRF2 (telomere repeat-binding factor 2) functions as a key component of the so called telomere ‘shelterin’ complex in maintaining telomere integrity at chromosome ends (D’Adda et al. 2003). However, recent findings suggests that TRF2 binds to and stabilizes NRSF thereby facilitating the physiological self-renewal of neural progenitor cells and the pathological uncontrolled proliferation of cancer cells (Ning et al. 2006; Zhang et al. 2008). As a consequence, reduced TRF2 binding to NRSF, and increased SCFβ-TRPC activity, target NRSF for proteasomal degradation and thereby inhibit cancer stem cell proliferation, especially in glioblastoma (Zhang et al. 2009). What’s more, the β-catenin/TCF system is reported to regulate the synthesis of NRSF and play a critical role in cell proliferation (Tomasoni et al. 2011). At the transcription level, a transcription factor Yin Yang (YY1) can activate NRSF transcription in SHSY5Y neuroblastoma cells (Jiang et al. 2008). In addition, in the Wnt pathway, there are several genes influence NRSF transcription via β-catenin (Nishihara et al. 2003). The dysregulation of the hedgehog pathway could induce changes in NRSF levels (Gates et al. 2010). CTDSP1 (the RNA polymerase C-terminal domain small phosphatase 1), a phosphatase, activity stabilizes NRSF in stem cells and that ERK-dependent phosphorylation combined with Pin1 (peptidylprolyl cis/trans isomerase) activity promotes NRSF degradation in neural progenitors (Nesti et al. 2014). Moreover, microRNAs (miRNAs) are excellent candidates for regulating cellular phenotype (He and Hannon 2004; Bartel 2004) (Table 2). They change the expression of NRSF with a regulatory feedback mechanism, in which the reciprocal action of miR-9 (Laneve et al. 2010) or miR-124a (Conaco et al. 2006) with NRSF may be relevant for the maintenance of the neuronal differentiation program. Finally, in addition to the regulation of neuronal NRSF concentrations at the level of protein stability, up-regulation of NRSF expression by nutrient or neuronal activity has been proposed. Transgenic mouse model that allows an inducible expression of NRSF in neurons would be very valuable to clarify the dynamics of NRSF expression and its role in nervous system.
The Relationship between NREF and its Isoforms
Due to alternative splicing, nrsf produces different transcripts. REST4, a neuron-specific truncated form of NRSF in rodent, could partly resist the silencing function of NRSF (Shimojo et al. 1999; Tabuchi et al. 2002), promoting neural gene expression and neurogenesis (Raj et al. 2011; Uchida et al. 2010) as it lacks the C-terminal repression domain of REST and is unable to interact with CoREST (Andres et al. 1999; Ooi and Wood 2007). In human, hREST4 protein, a truncated REST generated by the alternative splicing of exon N62 from the REST gene, was mainly expressed in neural tissues/cells as rodent REST4 (Palm et al. 1999). The functions of REST4 in the brain are much more complex. Recent studies demonstrate that TRF2 interacts with hREST4 to protect hREST4 from ubiquitin-mediated degradation by the proteasome, hence positively regulating neural progenitor formation and maintenance (Ovando-Roche et al. 2014). In addition, the network of REST4-mediated genes in the mPFC during the early postnatal period but not adult mice plays an important role in the development of stress vulnerability (Uchida et al. 2010). The neural-specific Ser/Arg repeat-related protein of 100 kDa (nSR100/SRRM4) directly promotes alternative splicing of REST transcripts to produce a REST isoform (REST4) with greatly reduced repressive activity, thereby activating expression of REST targets in neural cells which required for neurogenesis (Raj et al. 2011). REST4 gives assistance to REST and plays a role in neurological disorder, including epilepsy (Spencer et al. 2006), Parkinson’s disease (Yu et al. 2009), mood disorders (Warburton et al. 2015), and fetal alcohol syndrome (Cai et al. 2011). Splicing of REST mRNA into its REST4 form also occurs in tumor, such as small cell lung cancer (SCLC) (Coulson et al. 2000) and breast (Wagoner et al. 2010). In SCLC, the increasing expression of the NRSF isoform impart a neuroendocrine phenotype on the cells (Coulson et al. 2000). In breast cancer, at least in part via the alternative splicing to REST4, REST function is lost (Wagoner et al. 2010). Taken together, the balance of REST4 and REST is important not only for neural differentiation and NPC maintenance, but also for steady neurological network. What’s more, NRSF and its splice variant may be attractive therapeutic targets and represent specific clinical markers (Coulson et al. 2000; Wagoner et al. 2010) for these neurological diseases.
The Role of NRSF in Neurogenesis
By maintaining neural progenitor cells in a self-renewing state, NRSF plays a crucial role in both embryonic development and adult neurogenesis (Ballas and Mandel 2005). Perturbation of NRSF expression or function results in early embryonic lethality (Chen et al. 1998) or tumors (Negrini et al. 2013). NRSF works as a key regulator of the fates of neural stem cells and cancer cells. However, both neurogenesis and tumorigenesis are complex and multifactorial process, governed by a cascade of genetic and epigenetic events. Fully understanding them requires consideration of the cooperative effects.
Originally, NRSF was proposed to be a master regulator of neurogenesis (Schoenherr and Anderson 1995; Chong et al. 1995). Based on this original hypothesis, many studies focus on performing the role of NRSF during the development of nervous system. On the one hand, when embryonic stem cells (ESC) differentiate in vitro to neural stem cells, NRSF expression is down-regulated (Ballas et al. 2005). miR-124 is highly and specially expressed in neurons (Sempere et al. 2004). Its expression is induced by falling levels of NRSF during differentiation of neural progenitor (Yoo et al. 2009). By this way, the induction of miR-124 remodels the terminal differentiation by the repression of transcriptional repressor cofactors (Visvanathan et al. 2007), initiation of neural-specific splicing (Makeyev et al. 2007), and the repression of the neural progenitor npBAF complex (Yoo et al. 2009). NRSF also controls the differentiation and gene transcription of human and rat neural stem cells along the neuronal lineage (Ekici et al. 2008; Gao et al. 2011). Similarly, although the expression of the pluripotency genes is decreased by the neural induction, in the NRSF-deficient embryonic stem cells, the down-regulation of pluripotency genes expression is delayed (Soldati et al. 2012). What’s more, the reduced self-renewal capacity as well as precocious neuronal differentiation has been tested in NRSF-deficient neural progenitor cells (Covey et al. 2012). Finally, reduced proliferation capacity (Gao et al. 2011) and depression of neuronal genes (Aoki et al. 2012) have been measured in the brains of NRSF-deficient mice.
On the other hand, other researchers demonstrated that depletion of NRSF from embryonic stem cells did not change their differentiation status (Buckley et al. 2009; Jorgensen et al. 2009a, b; Yamada et al. 2010; Soldati et al. 2012). NRSF-deficient embryonic stem cells remained pluripotent and were able to differentiate into cells of the three germ layers mesoderm, endoderm, and ectoderm (Jorgensen et al. 2009b; Covey et al. 2012). NRSF regulates ESC pluripotency in culture condition- and ESC line-dependent fashion, and ESC pluripotency needs to be evaluated in a context dependent manner. Extracellular matrix components, such as feeder cells and laminin, can rescue the role of NRSF in ESC pluripotency (Singh et al. 2012). Taken together, these data demonstrate that NRSF is not required to maintain the pluripotency state of embryonic stem cells while the neuronal gene expression program is suppressed by NRSF in these cells. The establishment and maintenance of neuronal identity require both derepression of NRSF-regulated genes as well as posttranscriptional down-regulation of non-neuronal transcripts by microRNAs (Laneve et al. 2010; Conaco et al. 2006),such as a human miR-9-2 gene, expressed almost exclusively in the brain (Dietrich et al. 2012; Zheng et al. 2009). miR-9 has been implicated in nervous system development, physiology, and pathology in several organisms (Greenway et al. 2007; Yu et al. 2011), that is also under NRSF control. miR-9-2 contribute to neural differentiation, neural fate determination, and cell cycle exit through the repression of a number of neural transcription factors including TLX (an orphan nuclear receptor), HES-1 (a Notch signaling effector), FOXG1 (a forebrain-specific transcription factor) (Rockowitz et al. 2014). TLX is critical for maintaining neural progenitor cells (NPCs) in their undifferentiated state (Shi et al. 2004). HES-1 is required for NSC homeostasis/maintenance (Bonev et al. 2012), as its repression accelerates, while its overexpression inhibits, neurogenesis (Kageyama et al. 2008). FoxG1 maintains NPC self-renewal (Fasano et al. 2009) and suppresses the formation of early-born neurons (Hanashima et al. 2004). In addition, previous research demonstrates the existence of a feedback mechanism in which the reciprocal action of miR-9 and NRSF (Rockowitz et al. 2014) may be responsible for the maintenance of the neuronal differentiation program. These results indicate that NRSF plays an important role in neurogenesis by both directly targeting key neuronal transcription factors and regulating the transcription of neuronal miRNAs to controls neurogenesis synergistically by fine-tuning the expression of individual components to maintain a balance, which is necessary for the proper development of multiple neuronal lineages and for maintaining some level of developmental plasticity. On the other hand, what should be highlighted is that the analysis of neural stem/progenitor cells through either deleting of NRSF or overexpressing NRSF revealed that NRSF is not the master regulator that is solely responsible for the acquisition of the neuronal fate. Rather, NRSF provides a regulatory hub that coordinately regulates multiple tiers of neuronal development (Soldati et al. 2012).
The Role of NRSF in Tumorigenesis
In neural tumors, the role of NRSF is oncogenic. High NRSF, present in relevant percentages of these tumors, such as medulloblastomas (Fuller et al. 2005), neuroblastomas (Singh et al. 2011), multiform glioblastomas (Kamal et al. 2012), and pheochromacytoma (Tomasoni et al. 2011), stimulates their proliferation and worsens prognosis. Several mechanisms have been proposed to explain how high NRSF results in high proliferation. First of all, in medulloblastomas, NRSF-dependent repression of the deubiquitylase USP37 (ubiquitin-specific peptidase 37) decreased the levels of CDKNIB/p27, a cyclin-dependent kinase inhibitor, thus decreasing its cell cycle inhibition, resulting in an increase in cell proliferation (Das et al. 2013). Second, TRF2 binds to and stabilizes NRSF in nuclear foci that colocalize with PML (promyelocytic leukemia) nuclear bodies in human neuroblastoma and glioblastomas cells thereby facilitating the pathological uncontrolled proliferation of cancer cells (Zhang et al. 2009, 2008). Third, in glioblastomas, high NRSF induces a decrease in expression of the miRNA miR-124a (Kamal et al. 2012; Conti et al. 2012; Fowler et al. 2011), thereby increasing the expression level of the NRSF targets transcription factor SNAI-1 (Snail homolog 2) (Xia et al. 2012), a transcription factor that promotes cell invasion and tumor metastases, and two small phosphatases, Scp1 (Small C-terminal domain phosphatase 1) and PTPN12 (Protein-tyrosine phosphatase, non-receptor type 12) (Conti et al. 2012). All three of these miR-124a targets stimulate cell proliferation. What’s more, illustrates part of the signaling loop operative in high-NRSF pheochromocytoma cells. High levels of NRSF induce a decrease of tuberous sclerosis complex 2 (TSC2), a hub that governs various intracellular signaling pathways. Low TSC2 results in decreased turnover and increased transfer to the nucleus of β-catenin (Nishihara et al. 2003), which stimulates the transcription of oncogenes such as cMyc and Cyclin D1 (Tomasoni et al. 2011). Moreover, evidence for a role of TSC2 and β-catenin in proliferation has been reported in medulloblastomas (Baryawno et al. 2010) (Figs. 1 and 2).
The schematic of the relationships among REST, binding partners, and its isoforms. Two repressor domains are located on the N- and C-termini of REST (Ooi and Wood 2007). The repressor element 1 (RE1) sites are recognized by the zinc-finger domain of REST, and the interaction with DNA is stabilized by the ATP-dependent chromatin-remodeling enzyme, BRG1 (Ooi et al. 2006). REST-mediated gene repression is achieved by the recruitment of two separate corepressor complexes, mSin3 and CoREST (Naruse et al. 1999). The N-terminus of REST interacts with the mSin3 complex, which contains two class I histone deacetylases (HDACs), HDAC1, and HDAC2. In myocytes, REST’s N-terminus also recruits the class II HDACs ,HDAC4, and HDAC5 (Nakagawa et al. 2006). The C-terminus of REST interacts with the CoREST complex, which contains HDAC1, HDAC2, BRG1, the H3K4 demethylase LSD1, and the H3K9 methylase G9a (Roopra et al. 2004a). The methyl-CpG2 binding protein MeCP2 has also been found in the REST corepressor complex. CoREST binding might be stabilized by its ability to bind DNA79 and/or by its interaction with MeCP2 (Lunyak et al. 2002). The NADH-sensitive corepressor C-terminal binding protein CtBP is recruited in the presence of low levels of NADH, but dissociates from the REST complex when NADH levels are high (Roopra et al. 2004b). In vitro, the N- and C-termini of REST seem to form distinct repression domains that interact with different corepressor complexes (Grimes et al. 2000). The diagram is not meant to necessarily imply direct interactions. nSR100 mediates alternative splicing switch from REST to REST4 splice isoforms in neurons thereby promoting neural gene expression. TRF2 also interacts with REST4 in human neural progenitors to protect hREST4 from ubiquitin-mediated degradation by the proteasome for positively regulating neural progenitor formation and maintenance
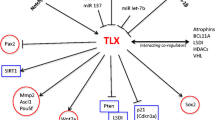
Mechanisms of high expression of REST induced proliferation in neural tumors. REST-dependent repression of USP37 (ubiquitin-specific peptidase 37) accelerates the turnover of p27 (a cyclin-dependent kinase inhibitor), thus decreasing its cell cycle inhibition, resulting in an increase in cell proliferation. TRF2 binds to and stabilizes REST, thereby preventing its degradation and facilitating the pathological uncontrolled proliferation of cancer cells, whereas activity of the ubiquitin E3 ligase SCFβ-TrCP accelerates proteasomal degradation of REST. High REST induces a decrease in expression of the miRNA miR-124a, thereby increasing the expression level of the REST targets transcription factor SNAI-1 (Snail homolog 2), Scp1(Small C-terminal domain phosphatase 1), localized in the nucleus, and PTPN12(Protein-tyrosine phosphatase, non-receptor type 12), distributed primarily in the cytoplasm. In addition, high levels of REST induce a decrease of TSC2 (tuberous sclerosis complex 2). Low TSC2 results in decreased turnover and increased transfer to the nucleus of β-catenin, which stimulates the transcription of oncogenes such as cMyc and Cyclin D1
In summary, various mechanisms have been identified to support the oncogenic role of NRSF in neural tumors. Interestingly, these mechanisms are not specific to the tumor type in which they were first identified. Initial evidence suggests that they may also operate in other types of neural tumors (Huang et al. 2011a, b; Negrini et al. 2013). In contrast, transfection with neither NRSF nor the oncogene cMyc is sufficient to induce neural stem cells to form a medulloblastoma; when NRSF and cMyc are cotransfected, however, they do induce tumors, although only if injected in the cerebellum, the site of human medulloblastoma formation, as opposed to the cerebral cortex. Alternatively, NRSF and the other factors could operate in parallel, via distinct but synergistic pathways. The complexity of tumorigenesis requires the cooperation of different mechanisms that may be governed by a variety of additional factors. Future studies should be aimed at revealing the comprehensive cell biological processes by which NRSF cooperates with other factors to govern cell proliferation, cell transformation, and tumor growth.
The Role of NRSF in Neurological Disorder Diseases
NRSF has been implicated in diverse neurological disorder diseases, highlighting the importance of NRSF-mediated regulation to the integrity of the cell, especially in neurodegeneration.
Ischemic insults promote NRSF specially binding and epigenetic remodeling at the miR-132 promoter and silencing of miR-132 expression in selectively vulnerable hippocampal CA1 neurons. A substantial decrease in two marks of active gene transcription, dimethylation of lysine 4 on core histone 3 (H3K4me2) and acetylation of lysine 9 on H3 (H3K9ac) at the miR-132 promoter are induced by ischemia, documenting a role for NRSF-dependent repression of miR-132 in the neuronal death associated with global ischemia (Hwang et al. 2014). Moreover, increased levels of NRSF are associated with a down-regulation of the transcriptionally responsive genes Gria2, which leads to increased calcium entry through GluR2-lacking AMPA receptors and subsequent neuronal cell death (Calderone et al. 2003). Polycomb group proteins serve as global enforcers of epigenetically repressed states in an array of cell types, including neurons (Zukin 2010). Recent studies indicate that polycomb repressive complex 2 (PRC2) is recruited to RE1-containing genes by NRSF via the non-coding RNA HOTAIR (Tsai et al. 2010) and that PRC1 interacts with NRSF at RE1 sites (Ren and Kerppola 2011). NRSF had opposite effects on PRC1 occupancy as well as on transcription at genes that contained distal versus proximal RE1 elements in differentiating neurons (Ren and Kerppola 2011). Moreover, polycomb proteins are activated and afford neuroprotection in the setting of ischemic preconditioning (Stapels et al. 2010). Casein Kinase 1 (CK1), an upstream effector that bidirectionally regulates NRSF cellular abundance, associates with and phosphorylates NRSF at two neighboring, but distinct motifs within the C terminus of NRSF critical for binding of β-rTCP and targeting of NRSF for proteasomal degradation. Global ischemia in rats in vivo triggers a decrease in CK1 and an increase in NRSF in selectively vulnerable hippocampal CA1 neurons. CK1 activation protects against ischemia-induced neuronal death via promoting β-rTCP stability by targeting of NRSF for proteasomal degradation and unsilencing of GluA2. However, NRSF also regulates an additional gene target, OPRM1 (opioid receptor 1 or MOR-1), by directly inhibition via histone deacetylation and methylation in the CA1 region of the hippocampus. Repression of Opmr1 seems to be neuroprotective, possibly because of an increased GABA (γ-aminobutyric acid) release that reduces the level of neuronal activity (Fig. 3a).
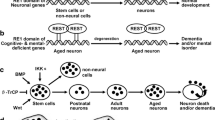
The multiple role of REST in neurological disorder diseases. a In ischemia, ischemic insults promote up-regulation of REST, thereby accelerating REST specially binding and silencing of miR-132 expression in selectively vulnerable hippocampal CA1 neurons. A substantial decrease in dimethylation of lysine 4 on core histone 3 (H3K4me2) and acetylation of lysine 9 on H3 (H3K9ac) also increase the repression of miR-132 expression. Increased levels of REST are associated with a down-regulation of the transcriptionally responsive genes Gria2, which leads to increased calcium entry through GluA2-lacking AMPA receptors and subsequent neuronal cell death. The increase in H3K9me2 and in binding of MeCP2 to methylated DNA corresponds to an increase in repressive GluA2 and a reciprocal potential decrease in transcription. Moreover, global ischemia decreases CK1 and β-TrCP. CK1 associates with and phosphorylates REST, thereby promoting β-TrCP-mediated ubiquitination. CK1 activation protects against ischemia-induced neuronal death. b In huntingtin disease (HD), wild-type huntingtin (wtHtt) sequesters REST in the cytoplasm denying access of REST to its regulatory elements on its target genes and permitting activated gene expression. wtHtt interacts with REST indirectly: a complex that contains p150Glued that binds HAP1 that, in turn, interacts with RILP. In HD, mutant huntingtin (muHtt)/HAP1/p150Glued complex is disrupted leading to the release of REST, which is free to migrate to the nucleus and consequently repress transcription of neuronal genes, including BDNF. In addition, muHtt triggers a pathogenic cascade involving Sp1 activation, which leads to REST up-regulation and repression of neuronal gene. c In epilepsies, under normal circumstances, NADH generated by glycolysis destabilizes the interaction of CtBP and REST, allowing transcription of REST target genes such as brain-derived neurotrophic factor (BDNF) and its receptor, TrkB, and maintaining normal neuronal excitability. The glycolytic inhibitor 2-deoxy-d-glucose (2DG) inhibits glycolysis, reducing NADH concentrations. The corepressor CtBP is recruited to form the REST-CtBP complex on REST target genes, reducing their transcription. Lower expression of BDNF and TrkB leads to reduced neuronal excitability, increasing seizure threshold, and inhibiting progression of kindling. Furthermore, the derepression of BDNF is associated with the activation of PLCγ and PI(3)K signaling pathways. d In Alzheimer’s disease(AD) and aging, the loss of neuroprotective REST functions contributes to neuronal vulnerability in the brains of those with Alzheimer’s. Both the Wnt signaling and the REST induction of Patients with AD are suppressed in, leading to neurodegeneration. REST is lost from the nucleus and appears in autophagosomes together with pathological misfolded proteins. During normal ageing, REST is induced in part by cell non-autonomous Wnt signaling. In the brains of healthy aged individuals, increased expression of nuclear REST both targets and suppresses several pro-apoptotic genes, as well as certain genes that encode enzymes involved in the pathology of Alzheimer’s
Potential roles for NRSF and its target genes have also been implicated in the pathogenesis of Huntington disease. One of the factors that contribute to the disease phenotype is the inability of mutant huntingtin (Htt) protein to interact with NRSF (Zuccato et al. 2003). Wild-type Htt sequesters NRSF in the cytoplasm of mouse striatal neurons, thereby inhibiting its function. NRSF is therefore prevented from binding to its cognate cis RE1 regulatory elements. Htt does not seem to interact with NRSF directly, but rather it is part of a complex that contains HAP1 and NRSF-interacting LIM domain protein (RILP), a protein that directly binds REST/NRSF and promotes its nuclear translocation. REST/NRSF, dynactin p150Glued, huntingtin, HAP1, and RILP form a complex involved in the translocation of REST/NRSF into the nucleus and that HAP1 controls REST/NRSF cellular localization in neurons (Shimojo 2008; Shimojo and Hersh 2006). However, the mutant HD protein cannot interact with NRSF, resulting in higher levels of NRSF in the nucleus and repression of its target genes. One such target is the neuronal survival factor, BDNF, low levels of which are thought to contribute to neuronal degeneration in Huntington disease (Zuccato et al. 2003; Shimojo and Hersh 2006). In addition, mHtt triggers a pathogenic cascade involving Sp1 activation, which leads to NRSF up-regulation and repression of neuronal genes (Ravache et al. 2010). A dominant-negative form of NRSF restored the BDNF level in HD cells (Zuccato et al. 2007). Other studies suggest that NRSF regulation is altered by polyglutamine (polyQ) toxicity, which expansion at the N-terminus of huntingtin (Htt). mHtt has an amino-terminal fragment (Nter) corresponding to the first 171 amino acids of human Htt with 142Q (thereafter called Nter-142Q) or with 15Q as control (Nter-15Q). mHtt fragment of this size has been shown to cause a neurological phenotype in mice (Schilling et al. 1999), the Nter of mHtt induces aberrant expression of NRSF (Ravache et al. 2010). The global level of NRSF proteins is increased in the brain of the R6/1 mouse model of HD (Smith et al. 2006). Also, NRSF can regulate its own expression level through a double negative feedback loop involving NRSF-dependent expression of a specific microRNA, MiR-9, a microRNA that regulates NRSF expression level, is down-regulated in HD and may account for the observed increase of NRSF expression (Packer et al. 2008). Thus, mHtt and its Nter (an amino-terminal fragment) fragments could trigger the activation of NRSF through three different mechanisms: by increasing NRSF transcription, decreasing microRNA regulation, and increasing NRSF nuclear translocation. Previous researches have shown that the expression of a dominant negative cDNA construct comprising the eight zinc fingers that represent the DNA binding domain of NRSF is able to reduce the binding of NRSF to its cognate genomic binding sites, leading to a consequent increase in transcription of BDNF and other NRSF-regulated genes (Belyaev et al. 2004; Bruce et al. 2004; Greenway et al. 2007; Zuccato and Cattaneo 2007). Decoys are double-stranded oligodeoxynucleotides corresponding to the DNA-binding element of a transcription factor and act to sequester it, thereby abrogating its transcriptional activity. Delivery of the decoy in cells expressing mutant Huntingtin leads to its specific interaction with NRSF, a reduction in NRSF occupancy of RE1s and rescue of target gene expression, including Bdnf (Soldati et al. 2011). A combination of virtual screening and biological approaches can lead to compounds reducing NRSF complex formation, which may be useful in HD and in other pathological conditions (Conforti et al. 2013) (Fig. 3b).
Increased levels of NRSF are also important in rat hippocampal and cortical neurons in response to epileptic seizures (Palm et al. 1998). To explain how the ‘ketogenic diet’ treatment works for drug-resistant epilepsy, researchers demonstrate that the glycolytic inhibitor 2-deoxy-d-glucose (2DG) potently reduces the progression of kindling and blocks seizure-induced increases in the expression of brain-derived neurotrophic factor (BDNF) and its receptor, TrkB. This reduced expression is mediated by the transcription factor NRSF, which recruits the NADH-binding co-repressor C-terminal binding protein (CtBP) to generate a repressive chromatin environment around the BDNF promoter (Garriga-Canut et al. 2006; Huang and McNamara 2006). Moreover, the conditional NRSF knockout mice with a Cre-loxp system to specifically delete NRSF in excitatory neurons of the postnatal mouse forebrain exhibited a dramatically accelerated seizure progression in an animal model of epilepsy, indicating that NRSF functions as a repressor of epileptogenesis (Hu et al. 2011). Taken together, above suggest that NRSF functions as an intrinsic repressor of epileptogenesis and a low expression level of NRSF in neurons is essential for maintaining neuronal functions (Fig. 3c).
It has also been shown that NRSF expression protects mature hippocampal neurons against hyperexcitability (Pozzi et al. 2013). NRSF is almost absent from the nuclei of cortical and hippocampal neurons of individuals with Alzheimer’s disease and is found in autophagosomes together with misfolded proteins. In the brains of healthy aged individuals, nuclear NRSF both targets and suppresses several pro-apoptotic genes, as well as certain genes that encode enzymes involved in the pathology of Alzheimer’s. During normal aging, NRSF is induced in part by cell non-autonomous Wnt signaling. In contrast, in the mouse brain, the conditional deletion of NRSF expression increased degeneration and cell death of neurons (Lu et al. 2014) (Fig. 3d). Moreover, neuron-specific conditional NRSF knockout mice were shown to be more vulnerable to the neurotoxin 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine (MPTP), which is frequently used to induce a Parkinson’s disease model in mice. Disturbance of the homeostasis of NRSF and its target genes, gliogenesis, and inflammation may contribute to the higher MPTP sensitivity in NRSF/REST neuronal cKO mice (Yu et al. 2013). What’s more, the apoptotic effect through cleaved-Caspase 3 induced by ethanol is increased in the brains of neuron-specific conditional NRSF knockout mice on neurons, providing new evidence that NRSF can be a therapeutic target in fetal alcohol syndrome (FAS) (Cai et al. 2011). Together, these data show that NRSF is neuroprotective and essential for maintaining neuronal viability. The neuroprotective activity is based on the repression of genes that encode pro-apoptotic genes or genes involved in the pathology of Alzheimer’s disease. Moreover, NRSF increases the expression of FOXO transcription factors that mediate oxidative stress resistance (Lu et al. 2014). Interestingly, NRSF levels are increased in cortical and hippocampal neurons of the aging healthy brain (Lu et al. 2014), supporting the preservation of cognitive function during aging. NRSF was also found to repress the u-opioid receptor in neuronal cells, and thus may have a role in opium addiction (Kim et al. 2004). Similarly, NRSF was found to repress the serotonin 1A receptor, which is implicated in depression and anxiety (Lemonde et al. 2004). NRSF dysfunction may contribute to the pathogenesis of a number of different neurodegenerative disorders. In addition to AD, NRSF was also significantly depleted in frontotemporal dementia and dementia with Lewy bodies. In each of these disorders, NRSF was lost from the nucleus and appeared in autophagosomes together with pathological misfolded proteins, including A β, phosphorylated tau, TDP-43, and a synuclein (Lu et al. 2014). This may represent a common pathogenic mechanism that links altered proteostasis to aberrant gene expression.
Taken together, these data support the view that the concentration of NRSF plays an important role in neurodegeneration and suggest that the NRSF concentration in adult neurons has to be tightly regulated. These findings indicate that under different conditions, in different cell types, and during different stages of development, NRSF regulates different networks of target genes. Understanding the mechanisms that underlie the involvement of NRSF and its copartners in diseases should identify putative therapeutic targets.
Clinical Perspectives
Progress of research on NRSF-dependent neurological disorder diseases, with identification of mechanisms that trigger and sustain their growth, has already offered new perspectives in terms of diagnostics, prognosis, and therapy.
To date, there is one laboratory test proposed for the presurgical identification of low-NRSF carcinomas, which uses the appearance in peripheral blood of NRSF-regulated transcripts as biomarkers (Moss et al. 2009). Other, similar tests could be developed in the near future. In terms of prognosis, both neural and non-neural NRSF-dependent tumors appear more aggressive than NRSF independent tumors of the same organ. This conclusion is based on gene profiling studies of surgically removed tumors, which identified specific molecular signatures and thus potentially useful prognostic markers (Wagoner and Roopra 2012; Sanson et al. 2006), combined with detailed analysis of pathology archives (Lv et al. 2010; Taylor et al. 2012).
In glioblastomas, low TRF2 and high SCFβ-TRCP levels accelerate NRSF turnover and thus play critical roles in cell proliferation (Zhang et al. 2009). This finding stimulated the search for agents that can specifically increase TRF2 or decrease SCFβ-TRCP. These agents are expected to induce fewer side effects than conventional chemotherapeutic drugs (Zhang et al. 2009). Combining a NRSF/TRF2-based treatment with low doses of existing chemotherapeutic agents might further improve the outcome in patients with glioblastoma. The identification of SCFβ-TRCP as a regulator of NRSF stability also provides a new perspective on the molecular mechanisms that regulate the fate of neural progenitor cells and cancer cells. However, when considering SCFβ-TRCP as a potential therapeutic target, it is important to recognize that NRSF is not the only target of the SCFβ-TRCP pathway. In a few cases, glioblastoma remission has been reported following treatment with valproic acid (VPA), an old drug used for decades in the treatment of epilepsy that has recently been recognized as an inhibitor of class I histone deacetylases, key NRSF effectors (Warburton et al. 2015). The drug induces differentiation of tumor cells can prevent their invasion into surrounding tissues and may inhibit tumor angiogenesis. Despite the broad substrate specificity of VPA, which hyperacetylates numerous proteins including some not associated with epigenetic regulation, the toxicity of VPA can be kept to a minimum (Taylor et al. 2012; Berendsen et al. 2012). VPA, as well as a few analogs and other molecules that block the action of NRSF, has been tested in medulloblastoma and glioblastoma cell lines with encouraging results (Taylor et al. 2012; Berendsen et al. 2012). At this stage, the combination of VPA or analogs with low doses of chemotherapeutic drugs and/or radiation appears a rational option that deserves investigation by well-designed prospective clinical trials. Two other approaches based on recent developments of NRSF studies also appear promising: first, the therapeutic potential of miRNAs that so far has been tested mostly in glioblastomas (Hummel et al. 2011), and second, the use of small peptides competing with full-length NRSF for RE-1 binding (Donev et al. 2008). In the future, it is possible that the combinations of histone deacetylase inhibitors with specific miRNAs may be introduced in clinical settings (Hummel et al. 2011), and the use of small peptides could strengthen immunotherapy in neuroblastomas by repressing the expression of membrane complement regulators such as CD59 (Donev et al. 2008). Moreover, the research of the factors that control the regulators of NRSF is another method to mediate the expression of NRSF. For example, recent findings identify DEAD-box RNA helicase (DDX3) as an essential upstream regulator mediate CK1 and Wnt–β-catenin signaling (Cruciat et al. 2013), raising the possibility that DDX3 may serve to regulate other CK1 targets such as NRSF. In addition to the regulation of neuronal NRSF concentrations at the level of protein stability, up-regulation of NRSF expression by nutrient or neuronal activity has been proposed (Pozzi et al. 2013; Garriga-Canut et al. 2006). What’s more, Yalda Sedaghat et al. employ second-generation antisense oligonucleotides (ASOs) to study the impact of NRSF-mediated suppression on gene expression. They suggested that the antisense approach may by a viable strategy for selectively modulating NRSF activity in vivo (Sedaghat et al. 2013).
There are hundreds of targets of NRSF, both direct and indirect, and several could be involved in the stimulation or repression of cell proliferation in tumors. The development of a new NRSF-based therapy for tumor may be worthwhile, if its use would allow decreases in chemotherapy doses and thus attenuation of the present serious problems of drug resistance and toxicity. What should be highlighted is that NRSF may trigger distinct cellar pathways in different neurological diseases to act as a stimulator or suppressor and to play a part in neuronal survival or neuronal death. It has been reported that NRSF-dependent silencing of miR-132 is causally related to ischemia-induced neuronal death and that overexpression of miR-132 in the CA1 of living rats affords robust protection against ischemia-induced neuronal death in a clinically relevant model of ischemic stroke (Hwang et al. 2014). In contrast, in Alzheimer’s disease, NRSF (Lu et al. 2014) and miR-132 (Wong et al. 2013) both appear to have prosurvival function. When point to a role for NRSF as a novel therapeutic target to tumor and neurodegeneration, we should have a comprehensive understanding depend on diversity molecular regulative pathways. On the basis of precious research, one strategy would be to activate Wnt signaling in aged individuals in Alzheimer (Lu et al. 2014; Tsai and Madabhushi 2014). However, such activation is also implicated in the development of various cancers, and so this approach would probably require careful targeting of Wnt activation in the brain (Anastas and Moon 2013). Alternative strategies include finding either Wnt-independent NRSF activators or small molecules that prevent the export of NRSF from the nucleus. A deeper understanding of the molecular mechanisms that govern NRSF activation in the aging brain will be crucial for such efforts to be successful.
Concluding Remarks
NRSF provides a regulatory hub that coordinately regulates multiple physiology and pathology of neuronal development and neurological diseases in vitro and vivo. As one of the parameters, long post-mortem delay that may have caused ischaemic damaged and increased NRSF level in neurons. More detailed description or universally accepted standard of the experiments may help researchers to have a comprehensive understanding and comparison of NRSF in different neurodegeneration and distinct mechanisms.
It is currently unclear how NRSF selectively represses distinct target genes in different cellular contexts, and why changes in NRSF expression or activity in many of these diseases result in changes in expression of only a subset of target genes. Even though the answers are still not known, it is becoming clear that the NRSF-mediated regulation of its target genes is not an all-or-none function and will depend on the cellular context, on the amount of NRSF protein present in the cell, and on the affinity of the NRSF protein complex toward its specific target gene in the given cellular environment, including the cell’s chromatin architecture. As the concentration of NRSF during development and in adult neuronal and endocrine cells is important for the biological function of NRSF, studies addressing the regulation of NRSF gene transcription and NRSF protein stability are essential to understand the regulation of NRSF expression in neurons during development. Gain of function experiments, i.e., should give us information about which concentration of NRSF is tolerated by neurons without loss of their cell type-specific phenotype and function.
References
Anastas JN, Moon RT (2013) WNT signalling pathways as therapeutic targets in cancer. Nat Rev Cancer 13:11–26
Andres ME, Burger C, Peral-Rubio MJ et al (1999) CoREST: a functional corepressor required for regulation of neural-specific gene expression. Proc Natl Acad Sci U S A 96:9873–9878
Aoki H, Hara A, Era T et al (2012) Genetic ablation of Rest leads to in vitro-specific derepression of neuronal genes during neurogenesis. Development 139:667–677
Bahn S, Mimmack M, Ryan M et al (2002) Neuronal target genes of the neuron-restrictive silencer factor in neurospheres derived from fetuses with Down’s syndrome: a gene expression study. Lancet 359:310–315
Ballas N, Mandel G (2005) The many faces of REST oversee epigenetic programming of neuronal genes. Curr Opin Neurobiol 15:500–506
Ballas N, Grunseich C, Lu DD et al (2005) REST and its corepressors mediate plasticity of neuronal gene chromatin throughout neurogenesis. Cell 121:645–657
Bartel DP (2004) MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 116:281–297
Baryawno N, Sveinbjornsson B, Eksborg S et al (2010) Small-molecule inhibitors of phosphatidylinositol 3-kinase/Akt signaling inhibit Wnt/beta-catenin pathway cross-talk and suppress medulloblastoma growth. Cancer Res 70:266–276
Belyaev ND, Wood IC, Bruce AW et al (2004) Distinct RE-1 silencing transcription factor-containing complexes interact with different target genes. J Biol Chem 279:556–561
Berendsen S, Broekman M, Seute T et al (2012) Valproic acid for the treatment of malignant gliomas: Review of the preclinical rationale and published clinical results. Expert Opin Investig Drugs 21:1391–1415
Bonev B, Stanley P, Papalopulu N (2012) MicroRNA-9 Modulates Hes1 ultradian oscillations by forming a double-negative feedback loop. Cell Rep 2:10–18
Bruce AW, Donaldson IJ, Wood IC et al (2004) Genome-wide analysis of repressor element 1 silencing transcription factor/neuron-restrictive silencing factor (REST/NRSF) target genes. Proc Natl Acad Sci U S A 101:10458–10463
Bruce AW, Krejci A, Ooi L et al (2006) The transcriptional repressor REST is a critical regulator of the neurosecretory phenotype. J Neurochem 98:1828–1840
Buckley NJ, Johnson R, Sun YM et al (2009) Is REST a regulator of pluripotency? Nature 457:E5–E6, E7
Cai L, Bian M, Liu M et al (2011) Ethanol-induced neurodegeneration in NRSF/REST neuronal conditional knockout mice. Neuroscience 181:196–205
Calderone A, Jover T, Noh KM et al (2003) Ischemic insults derepress the gene silencer REST in neurons destined to die. J Neurosci 23:2112–2121
Chen ZF, Paquette AJ, Anderson DJ (1998) NRSF/REST is required in vivo for repression of multiple neuronal target genes during embryogenesis. Nat Genet 20:136–142
Chong JA, Tapia-Ramirez J, Kim S et al (1995) REST: a mammalian silencer protein that restricts sodium channel gene expression to neurons. Cell 80:949–957
Conaco C, Otto S, Han JJ et al (2006) Reciprocal actions of REST and a microRNA promote neuronal identity. Proc Natl Acad Sci U S A 103:2422–2427
Conforti P, Zuccato C, Gaudenzi G et al (2013) Binding of the repressor complex REST-mSIN3b by small molecules restores neuronal gene transcription in Huntington’s disease models. J Neurochem 127:22–35
Conti L, Crisafulli L, Caldera V et al (2012) REST controls self-renewal and tumorigenic competence of human glioblastoma cells. PLoS One 7:e38486
Coulson JM, Edgson JL, Woll PJ et al (2000) A splice variant of the neuron-restrictive silencer factor repressor is expressed in small cell lung cancer: a potential role in derepression of neuroendocrine genes and a useful clinical marker. Cancer Res 60:1840–1844
Covey MV, Streb JW, Spektor R et al (2012) REST regulates the pool size of the different neural lineages by restricting the generation of neurons and oligodendrocytes from neural stem/progenitor cells. Development 139:2878–2890
Cruciat CM, Dolde C, de Groot RE et al (2013) RNA helicase DDX3 is a regulatory subunit of casein kinase 1 in Wnt-beta-catenin signaling. Science 339:1436–1441
D’Adda DFF, Reaper PM, Clay-Farrace L et al (2003) A DNA damage checkpoint response in telomere-initiated senescence. Nature 426:194–198
D’Alessandro R, Klajn A, Stucchi L et al (2008) Expression of the neurosecretory process in PC12 cells is governed by REST. J Neurochem 105:1369–1383
Das CM, Taylor P, Gireud M et al (2013) The deubiquitylase USP37 links REST to the control of p27 stability and cell proliferation. Oncogene 32:1691–1701
Dietrich N, Lerdrup M, Landt E et al (2012) REST-mediated recruitment of polycomb repressor complexes in mammalian cells. PLoS Genet 8:e1002494
Donev RM, Gray LC, Sivasankar B et al (2008) Modulation of CD59 expression by restrictive silencer factor-derived peptides in cancer immunotherapy for neuroblastoma. Cancer Res 68:5979–5987
Ekici M, Hohl M, Schuit F et al (2008) Transcription of genes encoding synaptic vesicle proteins in human neural stem cells: chromatin accessibility, histone methylation pattern, and the essential role of rest. J Biol Chem 283:9257–9268
Fasano CA, Phoenix TN, Kokovay E et al (2009) Bmi-1 cooperates with Foxg1 to maintain neural stem cell self-renewal in the forebrain. Genes Dev 23:561–574
Fowler A, Thomson D, Giles K et al (2011) miR-124a is frequently down-regulated in glioblastoma and is involved in migration and invasion. Eur J Cancer 47:953–963
Fuller GN, Su X, Price RE et al (2005) Many human medulloblastoma tumors overexpress repressor element-1 silencing transcription (REST)/neuron-restrictive silencer factor, which can be functionally countered by REST-VP16. Mol Cancer Ther 4:343–349
Gao Z, Ure K, Ding P et al (2011) The master negative regulator REST/NRSF controls adult neurogenesis by restraining the neurogenic program in quiescent stem cells. J Neurosci 31:9772–9786
Garriga-Canut M, Schoenike B, Qazi R et al (2006) 2-Deoxy-D-glucose reduces epilepsy progression by NRSF-CtBP-dependent metabolic regulation of chromatin structure. Nat Neurosci 9:1382–1387
Gates KP, Mentzer L, Karlstrom RO et al (2010) The transcriptional repressor REST/NRSF modulates hedgehog signaling. Dev Biol 340:293–305
Greenway DJ, Street M, Jeffries A et al (2007) RE1 Silencing transcription factor maintains a repressive chromatin environment in embryonic hippocampal neural stem cells. Stem Cells 25:354–363
Grimes JA, Nielsen SJ and Battaglioli E, et al. (2000) The co-repressor mSin3A is a functional component of the REST-CoREST repressor complex. Journal Of Biological Chemistry 275: 9461-9467
Guardavaccaro D, Frescas D, Dorrello NV et al (2008) Control of chromosome stability by the beta-TrCP-REST-Mad2 axis. Nature 452:365–369
Hanashima C, Li SC, Shen L et al (2004) Foxg1 suppresses early cortical cell fate. Science 303:56–59
Hatano Y, Yamada Y, Hata K et al (2011) Genetic ablation of a candidate tumor suppressor gene, rest, does not promote mouse colon carcinogenesis. Cancer Sci 102:1659–1664
He L, Hannon GJ (2004) MicroRNAs: Small RNAs with a big role in gene regulation. Nat Rev Genet 5:522–531
Hu XL, Cheng X, Cai L et al (2011) Conditional deletion of NRSF in forebrain neurons accelerates epileptogenesis in the kindling model. Cereb Cortex 21:2158–2165
Huang Z, Bao S (2012) Ubiquitination and deubiquitination of REST and its roles in cancers. Febs Lett 586:1602–1605
Huang YZ, McNamara JO (2006) Inhibiting glycolysis to reduce seizures: how it might work. Nat Neurosci 9:1351–1352
Huang X, Summers MK, Pham V et al (2011a) Deubiquitinase USP37 is activated by CDK2 to antagonize APC(CDH1) and promote S phase entry. Mol Cell 42:511–523
Huang Z, Wu Q, Guryanova OA et al (2011b) Deubiquitylase HAUSP stabilizes REST and promotes maintenance of neural progenitor cells. Nat Cell Biol 13:142–152
Hummel R, Maurer J, Haier J (2011) MicroRNAs in brain tumors: a new diagnostic and therapeutic perspective? Mol Neurobiol 44:223–234
Hwang JY, Kaneko N, Noh KM et al (2014) The gene silencing transcription factor REST represses miR-132 expression in hippocampal neurons destined to die. J Mol Biol 426:3454–3466
Jiang L, Yao M, Shi J et al (2008) Yin yang 1 directly regulates the transcription of RE-1 silencing transcription factor. J Neurosci Res 86:1209–1216
Johnson DS, Mortazavi A, Myers RM et al (2007) Genome-wide mapping of in vivo protein-DNA interactions. Science 316:1497–1502
Jorgensen HF, Chen ZF et al (2009a) Is REST required for ESC pluripotency? Nature 457:E4–E5, E7
Jorgensen HF, Terry A et al (2009b) REST selectively represses a subset of RE1-containing neuronal genes in mouse embryonic stem cells. Development 136:715–721
Kageyama R, Ohtsuka T, Kobayashi T (2008) Roles of Hes genes in neural development. Develop Growth Differ 50(Suppl 1):S97–S103
Kamal MM, Sathyan P, Singh SK et al (2012) REST regulates oncogenic properties of glioblastoma stem cells. Stem Cells 30:405–414
Kaneko N, Hwang JY, Gertner M et al (2014) Casein kinase 1 suppresses activation of REST in insulted hippocampal neurons and halts ischemia-induced neuronal death. J Neurosci 34:6030–6039
Kim CS, Hwang CK, Choi HS et al (2004) Neuron-restrictive silencer factor (NRSF) functions as a repressor in neuronal cells to regulate the mu opioid receptor gene. J Biol Chem 279:46464–46473
Kreisler A, Strissel PL, Strick R et al (2010) Regulation of the NRSF/REST gene by methylation and CREB affects the cellular phenotype of small-cell lung cancer. Oncogene 29:5828–5838
Laneve P, Gioia U, Andriotto A et al (2010) A minicircuitry involving REST and CREB controls miR-9-2 expression during human neuronal differentiation. Nucleic Acids Res 38:6895–6905
Lemonde S, Rogaeva A, Albert PR (2004) Cell type-dependent recruitment of trichostatin A-sensitive repression of the human 5-HT1A receptor gene. J Neurochem 88:857–868
Loe-Mie Y, Lepagnol-Bestel AM, Maussion G et al (2010) SMARCA2 and other genome-wide supported schizophrenia-associated genes: Regulation by REST/NRSF, network organization and primate-specific evolution. Hum Mol Genet 19:2841–2857
Lu T, Aron L, Zullo J et al (2014) REST and stress resistance in ageing and Alzheimer’s disease. Nature 507:448–454
Lunyak VV, Burgess R and Prefontaine GG, et al. (2002) Corepressordependent silencing of chromosomal regions encoding neuronal genes. Science 298: 1747–1752
Lv H, Pan G, Zheng G et al (2010) Expression and functions of the repressor element 1 (RE-1)-silencing transcription factor (REST) in breast cancer. J Cell Biochem 110:968–974
Makeyev EV, Zhang J, Carrasco MA et al (2007) The MicroRNA miR-124 promotes neuronal differentiation by triggering brain-specific alternative pre-mRNA splicing. Mol Cell 27:435–448
Moss AC, Jacobson GM, Walker LE et al (2009) SCG3 transcript in peripheral blood is a prognostic biomarker for REST-deficient small cell lung cancer. Clin Cancer Res 15:274–283
Nakagawa Y, Kuwahara K and Harada M, et al. (2006) Class II HDACs mediate CaMK-dependent signaling to NRSF in ventricular myocytes. Journal Of Molecular And Cellular Cardiology 41: 1010–1022
Naruse Y, Aoki T and Kojima T, et al. (1999) Neural restrictive silencer factor recruits mSin3 and histone deacetylase complex to repress neuron-specific target genes. Proc Natl Acad Sci U S A 96: 13691–13696
Negrini S, Prada I, D’Alessandro R et al (2013) REST: an oncogene or a tumor suppressor? Trends Cell Biol 23:289–295
Nesti E, Corson GM, McCleskey M et al (2014) C-terminal domain small phosphatase 1 and MAP kinase reciprocally control REST stability and neuronal differentiation. Proc Natl Acad Sci U S A 111:E3929–E3936
Ning H, Li T, Zhao L et al (2006) TRF2 promotes multidrug resistance in gastric cancer cells. Cancer Biol Ther 5:950–956
Nishihara S, Tsuda L, Ogura T (2003) The canonical Wnt pathway directly regulates NRSF/REST expression in chick spinal cord. Biochem Biophys Res Commun 311:55–63
Noh KM, Hwang JY, Follenzi A et al (2012) Repressor element-1 silencing transcription factor (REST)-dependent epigenetic remodeling is critical to ischemia-induced neuronal death. Proc Natl Acad Sci U S A 109:E962–E971
Ohnuki T, Nakamura A, Okuyama S et al (2010) Gene expression profiling in progressively MPTP-lesioned macaques reveals molecular pathways associated with sporadic Parkinson’s disease. Brain Res 1346:26–42
Ooi L, Belyaev ND and Miyake K, et al. (2006) BRG1 chromatin remodeling activity is required for efficient chromatin binding by repressor element 1-silencing transcription factor (REST) and facilitates REST-mediated repression. Journal Of Biological Chemistry 281: 38974–38980
Ooi L, Wood IC (2007) Chromatin crosstalk in development and disease: Lessons from REST. Nat Rev Genet 8:544–554
Ovando-Roche P, Yu JS, Testori S et al (2014) TRF2-mediated stabilization of hREST4 is critical for the differentiation and maintenance of neural progenitors. Stem Cells 32:2111–2122
Packer AN, Xing Y, Harper SQ et al (2008) The bifunctional microRNA miR-9/miR-9* regulates REST and CoREST and is downregulated in Huntington’s disease. Journal Of Neuroscience 28:14341–14346
Palm K, Belluardo N, Metsis M et al (1998) Neuronal expression of zinc finger transcription factor REST/NRSF/XBR gene. Journal Of Neuroscience 18:1280–1296
Palm K, Metsis M, Timmusk T (1999) Neuron-specific splicing of zinc finger transcription factor REST/NRSF/XBR is frequent in neuroblastomas and conserved in human, mouse and rat. Brain Res Mol Brain Res 72:30–39
Pozzi D, Lignani G, Ferrea E et al (2013) REST/NRSF-mediated intrinsic homeostasis protects neuronal networks from hyperexcitability. Embo J 32:2994–3007
Quinn JP, Warburton A, Myers P et al (2013) Polymorphic variation as a driver of differential neuropeptide gene expression. Neuropeptides 47:395–400
Qureshi IA, Gokhan S, Mehler MF (2010) REST and CoREST are transcriptional and epigenetic regulators of seminal neural fate decisions. Cell Cycle 9:4477–4486
Raj B, O’Hanlon D, Vessey JP et al (2011) Cross-regulation between an alternative splicing activator and a transcription repressor controls neurogenesis. Mol Cell 43:843–850
Ravache M, Weber C, Merienne K et al (2010) Transcriptional activation of REST by Sp1 in Huntington’s disease models. PLoS One 5:e14311
Ren X, Kerppola TK (2011) REST interacts with Cbx proteins and regulates polycomb repressive complex 1 occupancy at RE1 elements. Mol Cellu Biol 31:2100–2110
Rockowitz S, Lien WH, Pedrosa E et al (2014) Comparison of REST cistromes across human cell types reveals common and context-specific functions. PLoS Comput Biol 10:e1003671
Rodenas-Ruano A, Chavez AE, Cossio MJ et al (2012) REST-dependent epigenetic remodeling promotes the developmental switch in synaptic NMDA receptors. Nat Neurosci 15:1382–1390
Roopra A, Qazi R and Schoenike B, et al. (2004a) Localized domains of G9a-mediated histone methylation are required for silencing of neuronal genes. Molecular Cell 14: 727–738
Roopra A, Qazi R and Schoenike B, et al. (2004b) Localized domains of G9a-mediated histone methylation are required for silencing of neuronal genes. Molecular Cell 14: 727–738
Sanson M, Laigle-Donadey F, Benouaich-Amiel A (2006) Molecular changes in brain tumors: Prognostic and therapeutic impact. Curr Opin Oncol 18:623–630
Schilling G, Becher MW, Sharp AH et al (1999) Intranuclear inclusions and neuritic aggregates in transgenic mice expressing a mutant N-terminal fragment of huntingtin. Hum Mol Genet 8:397–407
Schoenherr CJ, Anderson DJ (1995) The neuron-restrictive silencer factor (NRSF): a coordinate repressor of multiple neuron-specific genes. Science 267:1360–1363
Schoenherr CJ, Paquette AJ, Anderson DJ (1996) Identification of potential target genes for the neuron-restrictive silencer factor. Proc Natl Acad Sci U S A 93:9881–9886
Sedaghat Y, Bui HH, Mazur C et al (2013) Identification of REST-regulated genes and pathways using a REST-targeted antisense approach. Nucleic Acid Ther 23:389–400
Sempere LF, Freemantle S, Pitha-Rowe I et al (2004) Expression profiling of mammalian microRNAs uncovers a subset of brain-expressed microRNAs with possible roles in murine and human neuronal differentiation. Genome Biol 5:R13
Shi Y, Chichung LD, Taupin P et al (2004) Expression and function of orphan nuclear receptor TLX in adult neural stem cells. Nature 427:78–83
Shimojo M (2008) Huntingtin regulates RE1-silencing transcription factor/neuron-restrictive silencer factor (REST/NRSF) nuclear trafficking indirectly through a complex with REST/NRSF-interacting LIM domain protein (RILP) and dynactin p150 Glued. J Biol Chem 283:34880–34886
Shimojo M, Hersh LB (2006) Characterization of the REST/NRSF-interacting LIM domain protein (RILP): Localization and interaction with REST/NRSF. J Neurochem 96:1130–1138
Shimojo M, Paquette AJ, Anderson DJ et al (1999) Protein kinase A regulates cholinergic gene expression in PC12 cells: REST4 silences the silencing activity of neuron-restrictive silencer factor/REST. Mol Cell Biol 19:6788–6795
Singh SK, Kagalwala MN, Parker-Thornburg J et al (2008) REST maintains self-renewal and pluripotency of embryonic stem cells. Nature 453:223–227
Singh A, Rokes C, Gireud M et al (2011) Retinoic acid induces REST degradation and neuronal differentiation by modulating the expression of SCF(beta-TRCP) in neuroblastoma cells. Cancer 117:5189–5202
Singh SK, Veo BL, Kagalwala MN et al (2012) Dynamic status of REST in the mouse ESC pluripotency network. PLoS One 7:e43659
Smith R, Chung H, Rundquist S et al (2006) Cholinergic neuronal defect without cell loss in Huntington’s disease. Hum Mol Genet 15:3119–3131
Soldati C, Bithell A, Conforti P et al (2011) Rescue of gene expression by modified REST decoy oligonucleotides in a cellular model of Huntington’s disease. J Neurochem 116:415–425
Soldati C, Bithell A, Johnston C et al (2012) Repressor element 1 silencing transcription factor couples loss of pluripotency with neural induction and neural differentiation. Stem Cells 30:425–434
Spencer EM, Chandler KE, Haddley K et al (2006) Regulation and role of REST and REST4 variants in modulation of gene expression in in vivo and in vitro in epilepsy models. Neurobiol Dis 24:41–52
Stapels M, Piper C, Yang T et al (2010) Polycomb group proteins as epigenetic mediators of neuroprotection in ischemic tolerance. Sci Signal 3:a15
Tabuchi A, Yamada T, Sasagawa S et al (2002) REST4-mediated modulation of REST/NRSF-silencing function during BDNF gene promoter activation. Biochem Biophys Res Commun 290:415–420
Tahiliani M, Mei P, Fang R et al (2007) The histone H3K4 demethylase SMCX links REST target genes to X-linked mental retardation. Nature 447:601–605
Taylor P, Fangusaro J, Rajaram V et al (2012) REST is a novel prognostic factor and therapeutic target for medulloblastoma. Mol Cancer Ther 11:1713–1723
Thiel G, Ekici M, Rossler OG (2014) RE-1 silencing transcription factor (REST): a regulator of neuronal development and neuronal/endocrine function. Cell Tissue Res. doi:10.1007/s00441-014-1963-0
Tomasoni R, Negrini S, Fiordaliso S et al (2011) A signaling loop of REST, TSC2 and beta-catenin governs proliferation and function of PC12 neural cells. J Cell Sci 124:3174–3186
Tsai LH, Madabhushi R (2014) Alzheimer’s disease: a protective factor for the ageing brain. Nature 507:439–440
Tsai MC, Manor O, Wan Y et al (2010) Long noncoding RNA as modular scaffold of histone modification complexes. Science 329:689–693
Uchida S, Hara K, Kobayashi A et al (2010) Early life stress enhances behavioral vulnerability to stress through the activation of REST4-mediated gene transcription in the medial prefrontal cortex of rodents. J Neurosci 30:15007–15018
Valouev A, Johnson DS, Sundquist A et al (2008) Genome-wide analysis of transcription factor binding sites based on ChIP-Seq data. Nat Methods 5:829–834
Visvanathan J, Lee S, Lee B et al (2007) The microRNA miR-124 antagonizes the anti-neural REST/SCP1 pathway during embryonic CNS development. Genes Dev 21:744–749
Wagoner MP, Roopra A (2012) A REST derived gene signature stratifies glioblastomas into chemotherapy resistant and responsive disease. BMC Genomics 13:686
Wagoner MP, Gunsalus KT, Schoenike B et al (2010) The transcription factor REST is lost in aggressive breast cancer. PLoS Genet 6:e1000979
Warburton A, Breen G, Rujescu D et al (2014) Characterization of a REST-regulated internal promoter in the schizophrenia genome-wide associated gene MIR137. Schizophr Bull. doi:10.1093/schbul/sbu117
Warburton A, Savage AL, Myers P et al (2015) Molecular signatures of mood stabilisers highlight the role of the transcription factor REST/NRSF. J Affect Disord 172:63–73
Westbrook TF, Hu G, Ang XL et al (2008) SCFbeta-TRCP controls oncogenic transformation and neural differentiation through REST degradation. Nature 452:370–374
Wong HK, Veremeyko T, Patel N et al (2013) De-repression of FOXO3a death axis by microRNA-132 and -212 causes neuronal apoptosis in Alzheimer’s disease. Hum Mol Genet 22:3077–3092
Xia H, Cheung WK, Ng SS et al (2012) Loss of brain-enriched miR-124 microRNA enhances stem-like traits and invasiveness of glioma cells. J Biol Chem 287:9962–9971
Yamada Y, Aoki H, Kunisada T et al (2010) Rest promotes the early differentiation of mouse ESCs but is not required for their maintenance. Cell Stem Cell 6:10–15
Yoo AS, Staahl BT, Chen L et al (2009) MicroRNA-mediated switching of chromatin-remodelling complexes in neural development. Nature 460:642–646
Yu M, Cai L, Liang M et al (2009) Alteration of NRSF expression exacerbating 1-methyl-4-phenyl-pyridinium ion-induced cell death of SH-SY5Y cells. Neurosci Res 65:236–244
Yu HB, Johnson R, Kunarso G et al (2011) Coassembly of REST and its cofactors at sites of gene repression in embryonic stem cells. Genome Res 21:1284–1293
Yu M, Suo H, Liu M et al (2013) NRSF/REST neuronal deficient mice are more vulnerable to the neurotoxin MPTP. Neurobiol Aging 34:916–927
Zhang P, Pazin MJ, Schwartz CM et al (2008) Nontelomeric TRF2-REST interaction modulates neuronal gene silencing and fate of tumor and stem cells. Curr Biol 18:1489–1494
Zhang P, Lathia JD, Flavahan WA et al (2009) Squelching glioblastoma stem cells by targeting REST for proteasomal degradation. Trends Neurosci 32:559–565
Zhang K, Biswas N, Gayen JR et al (2014) Chromogranin B: Intra- and extra-cellular mechanisms to regulate catecholamine storage and release, in catecholaminergic cells and organisms. J Neurochem 129:48–59
Zheng D, Zhao K, Mehler MF (2009) Profiling RE1/REST-mediated histone modifications in the human genome. Genome Biol 10:R9
Zuccato C, Cattaneo E (2007) Role of brain-derived neurotrophic factor in Huntington’s disease. Prog Neurobiol 81:294–330
Zuccato C, Tartari M, Crotti A et al (2003) Huntingtin interacts with REST/NRSF to modulate the transcription of NRSE-controlled neuronal genes. Nat Genet 35:76–83
Zuccato C, Belyaev N, Conforti P et al (2007) Widespread disruption of repressor element-1 silencing transcription factor/neuron-restrictive silencer factor occupancy at its target genes in Huntington’s disease. J Neurosci 27:6972–6983
Zukin RS (2010) Eradicating the mediators of neuronal death with a fine-tooth comb. Sci Signal 3:e20
Acknowledgments
This work was supported by the Natural Science Foundation of China (Project No. 31172293, No. 31272532, and No.31472166), 948 projects (2013-S11; 2014-S9), and the Program for Cheung Kong Scholars and Innovative Research Team in University of China (No. IRT0866).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Song, Z., Zhao, D., Zhao, H. et al. NRSF: an Angel or a Devil in Neurogenesis and Neurological Diseases. J Mol Neurosci 56, 131–144 (2015). https://doi.org/10.1007/s12031-014-0474-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12031-014-0474-5