Abstract
Origin of cancer stem cells (CSCs) and mechanisms by which oncogene PTTG1 contributes to tumor progression via CSCs is not known. Ovarian CSCs exhibit characteristics of self-renewal, tumor-initiation, growth, differentiation, drug resistance, and tumor relapse. A common location of putative origin, namely the ovarian surface epithelium, is shared between the normal stem and CSC compartments. Existence of ovarian stem cells and their co-expression with CSC signatures suggests a strong correlation between origin of epithelial cancer and CSCs. We hereby explored a putative oncogene PTTG1 (Securin), reported to be overexpressed in various tumors, including ovarian. We report a previously overlooked role of PTTG1 as a marker of CSCs thereby modulating CSC, germline, and stemness-related genes. We further characterized PTTG1’s ability to regulate (cancer) stem cell-associated self-renewal and epithelial-mesenchymal transition pathways. Collectively, the data sheds light on a potential target expressed during ovarian tumorigenesis and metastatically disseminated ascites CSCs in the peritoneal cavity. Present study highlights this unconventional, under-explored role of PTTG1 in regulation of stem and CSC compartments in ovary, ovarian cancer and ascites and highlights it as a potential candidate for developing CSC specific targeted therapeutics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ovarian cancer is a deadly gynecological disease accounting for approximately 22,240 new cases and 14,070 deaths in the USA in 2018 [1]. Recently, a vast body of research has indicated cancer stem cells (CSCs) as putative entities responsible for cancer initiation and progression. Parallel characteristic properties are shared between stem cells and CSCs, which make them bona fide candidates responsible for the resurgence of tumors. Oncogenes are at the “epicenter” for putative cellular transformation, the formation of precancerous lesions, and their subsequent clinical manifestation into mature cancer [2,3,4]. Several oncogenes are reported to be overexpressed in CSCs. Pituitary tumor transforming gene 1 (PTTG1), also known as securin, is a multi-domain, multi-functional proto-oncogene overexpressed in various tumors, including ovarian cancer [5,6,7,8,9,10].
Cloning of this novel oncogene, namely PTTG1, from the testis [5] and ovarian tumors [8] was previously reported by our group. PTTG1/securin acts as a regulator of sister chromatid separation during cell division under physiological conditions [9]. Its overexpression induces cellular transformation and tumor development in nude mice [5,6,7, 10]. It is linked to genetic instability, aneuploidy, tumor progression, invasion, metastasis, and cancer recurrence [11,12,13,14,15,16,17,18]. PTTG1 also directly regulates gene transcriptional activity and induces mitogenic [19] and angiogenic genes [20,21,22], such as c-Myc [23] and VEGF and bFGF [19, 21, 24] respectively. It also regulates cell division, cell cycle, transactivation of growth factors, acting as an initiator and promoter of tumorigenesis [9, 10, 25]. Of note, the overexpression of PTTG1 stimulates basic fibroblast growth factor (b-FGF) expression and secretion [21, 24], which is known to regulate human embryonic stem cells and may influence the stem cell compartment under normal physiological conditions [26]. Various methods of PTTG1 down-regulation have been shown to inhibit ovarian cell proliferation and suppression of tumor growth in nude mice [27,28,29].
Tumorigenic function of PTTG1 was further demonstrated by the overexpression of PTTG1 in ovarian surface epithelial (OSE) cells, which resulted in enlarged ovaries accompanied by an increase in corpora lutea, abnormal fallopian tubes, and endothelium with early signs of hyperplasia and neoplasia [30]. Similarly, over-expression of PTTG1 in pituitary cells resulted in the development of GH-cell focal hyperplasia and adenoma, associated with increased serum levels of LH, GH, testosterone, and/or IGF-I, and enlargement of the prostate [31]. In contrast, crossbreeding of animals (Rb± ) with PTTG null (PTTG−/−) animals reduced the pituitary tumor development from 86% in Rb± /PTTG+/+ to 30% in Rb± /PTTG−/−, suggesting an important role of PTTG1 in tumorigenesis [32]. Overexpression of PTTG1 has been unanimously reported in several oncological settings [5,6,7,8, 10, 33]. Yoon et al. (2012) (ref. 34) have demonstrated the role of PTTG1 in human breast cancer in comparison with normal tissues and reported that its expression levels correlated with the migratory and invasive potential of breast cancer cells by modulating EMT process. PTTG1 mediated AKT activation which is implicated in stemness and EMT properties of cancer cells was studied, thus recognizing PTTG1 oncogene as a potential therapeutic target. However, the precise expression of PTTG1 in stem cells/CSCs and its role in the regulation of stem cell and cancer stem cell compartments has not been explored. In the present study, we initially investigated co-expression of PTTG1 with stem cell/CSC markers in normal ovarian samples and ovarian tumors at various stages of tumorigenesis as well as in the ascites-derived CSCs collected from patients with recurrent ovarian cancer. We further delineated its role in the regulation of CSC populations by studying molecules specific for self-renewal and epithelial-mesenchymal transition (EMT) signaling pathways. Our study demonstrated co-expression of PTTG1 with several stem cell/CSC markers, pluripotent and germinal lineage markers in normal ovary (NO), benign tumor (BN), borderline tumor (BL) and high grade tumors (HG), as well as ascites-derived CSCs. Knockdown and overexpression of PTTG1 in ovarian cancer cells (A2780) showed a differential expression of key stem/CSC genes and those related to self-renewal and EMT signaling pathways. Collectively, our study results suggest a key and underexplored role of the oncogene PTTG1 in regulating stem cells and CSCs in ovary and ovarian cancer, respectively.
Results
PTTG1 Co-Localizes with Stem/CSCs in Normal Ovarian and Ovarian Tumor Samples
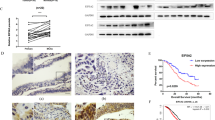
Our previous studies and those from other research groups clearly showed overexpression of PTTG1 in various tumors [5,6,7,8, 33]. However, to the best of our knowledge, there is no report showing the expression of PTTG1 gene in normal stem cells or CSCs in oncological settings, such as ovarian cancer. Therefore, we initially examined the expression of PTTG1 in normal ovarian tissues and ovarian tumors at various stages of tumorigenesis. Using real-time PCR analysis, we detected the expression of PTTG1 and various stem cell/CSC markers (stemness related: NANOG, OCT4A and SOX2; CSC-specific: ALDH1, CD24, CD44, CD117, CD133, and LGR5; and germinal lineage specific: DDX4/VASA and IFITM3/FRAGILIS) transcripts in NO samples and ovarian tumors. RNA from 7 normal individuals and 7 patients at each stage of tumorigenesis (BN, BL and HG) was used for analysis. Our analysis showed a detectable expression of PTTG1 and each marker gene in NO samples and in BN, BL, and HG samples (Table 1). High variation in inter-patient expression levels was observed for PTTG1, as well as stem cell/cancer stem cell markers in NO and across the tumor stages. However, a consistent trend of increased expression levels of PTTG1 and other marker genes such as CD133 and CD24, in all the stages of tumorigenesis was observed compared to NO. ALDH1 reflected a decreased trend of expression in BN and HG, but an increase in BL compared to NO. DDX4 and IFITM3 did not reveal a substantial and consistent increased trend in expression levels in BN, BL, or HG samples compared to NO samples; instead increased expression of DDX4 in BN whereas that of IFITM3 in BL were noted. The pluripotency-related triad of genes OCT4A, NANOG, and SOX2 exhibited a clear increased trend of expression in BN, BL, and HG compared to NO (except for HG NANOG). LGR5 also showed increase in expression in BN, BL, and HG tumors compared to NO. However, a large degree of variation in gene expression fold change values across tumor stages was observed (Table 1). Specific genes revealing a decreased expression in HG tumors might suggest a differentiated state of the tumors.
Co-expression of PTTG1 using dual fluoro-immunological analysis was performed using specific-antibodies for PTTG1 and each of the stem cell/CSC markers [35, 36]. We observed co-expression of PTTG1 with: ALDH1, CD34, CD44, LGR5, NANOG, OCT4, and SSEA4 in NO samples, as well as in BN, BL, and HG samples. Cells co-expressing PTTG1 and stem cell/CSC-specific markers were consistently distributed in both the OSE and cortex regions in NO and ovarian tumors at all stages of tumorigenesis (BN, BL, and HG) (Fig. 1a-h). We observed a graded increase in the expression of PTTG1 and CSC markers from NO samples throughout the tumorigenic stages. In addition, we observed the existence of tumor-like cell clusters positive for PTTG1 and each marker in both OSE and cortex regions in the tumor samples, suggesting the importance of PTTG1 and CSC genes in tumor progression. These results are consistent with recent findings from our group investigating the existence of various stem/CSC populations across normal ovary (NO) and ovarian tumors (BN, BL and HG) [35, 36].
Co-localization of PTTG1 with stem cell/cancer stem cell markers in normal ovary (NO), benign tumor (BN), borderline tumor (BL) and high grade tumor HG). Anti-ALDH1, CD34, CD44, LGR5, NANOG, OCT4 and SSEA4 specific antibodies (green color) were co-localized with PTTG1 (red color) in OSE (ovarian surface epithelium) and cortex in normal ovarian and tumor tissues sections. Sections were counter-stained with nuclear specific dye DAPI (blue). Overlap of green, red and blue represents (overlap). Alexafluor-labelled secondary antibody (anti-rabbit or anti-mouse) were employed for detection. Scale bar = 50 μm. a and b = normal ovary (NO), c and d = benign tumor (BN), e and f = borderline tumor (BL), and g and h = high grade tumor (HG). a, c, e and g = Ovarian surface epithelium (OSE), and b, d, f and h = Cortex. Results shown are representative of two samples from independent patients
Further, to confirm co-expression of PTTG1 with CSC markers, we isolated the ALDH1+ population from the ovarian cancer cell line A2780 and analysed the expression of both ALDH1 and PTTG1 using real-time PCR. As shown in Fig. 2, PTTG1 was found to be highly expressed in ALDH1+ cells compared to unsorted A2780 cells and ALDH1− cells. ALDH1 expression was relatively lower in A2780 cells compared to ALDH1+ cells, and undetectable in ALDH1− cells (Fig. 2), corroborating the co-expression of PTTG with CSC markers. Taken together, our results demonstrate co-expression of PTTG1 with stem cell/CSC markers in NO samples and ovarian cancer at various stages of tumorigenesis (BN, BL and HG), as well as in an ovarian cancer cell line (A2780), suggesting that PTTG1 may serve as a previously overlooked marker for stem cells and CSCs.
Expression ofPTTG1and ALDH1 in A2780 cells, and isolated ALDH1+and ALDH1−populations. ALDH1+ and ALDH1− cells from A2780 cells were isolated as described in materials and methods. Primers specific for ALDH1 and PTTG1 genes were employed in real-time PCR analysis. GAPDH primers were used as an internal control. Values shown are average of two independent experiments. A = ALDH1 and B = PTTG1
Ascites-Derived CSCs Reveal Co-Expression of PTTG1 and CSC Genes
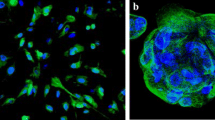
Cancer cells and CSCs are known to disseminate into the peritoneal cavity at certain stage of tumorigenesis, metastasize to the omentum, and grow as a “liquid cancer” (ascites fluid), thereby imposing a clinical problem noted as a major cause of death amongst affected patients [37,38,39]. To determine if CSCs present in ascites cells are indeed similar to those observed in ovarian tumors and could serve as future therapeutic targets, we analysed various CSC populations in ascites cells. Enriched CSCs collected from ascites of recurrent ovarian cancer patients were subjected to immuno-fluorescence analysis for PTTG1 and CSC-specific markers, as described previously [35, 36]. As evident in Fig. 3, PTTG1 was found to be highly co-expressed with the markers: ALDH1, CD34, CD44, EpCAM, LGR5, NANOG, OCT4, and SSEA4 in ascites-derived CSCs (Fig. 3a, b). Expression of PTTG1 and each of the CSC makers was found in single and clustered cells (normally found in ascites fluid) [37]. In addition, we observed differential expression levels of PTTG1 and each of the aforementioned markers in single cells and clustered cells found in the ascites fluid. Expression of PTTG1 and each of the CSC marker genes was confirmed by RT-PCR (Fig. 3c) using gene specific primers (Supplement Table S1). A high inter-patient variation was observed, indicating heterogeneity of CSC populations in ascites samples (Fig. 3c). Our results suggest the presence of similar populations of CSCs in ovarian tumors and ascites, indicating a clinically relevant role of PTTG1 and its proposed role as a unique CSC marker that could explain the association of ascites fluid-derived CSCs with poor prognosis and patient survival, and thus ultimately the mortality due to tumor relapse.
Co-localization of stem cell/cancer stem cell markers with PTTG1 and their detection in ascites derived CSCs from patients with recurrent ovarian cancer. Co-expression of PTTG1 with each of the CSC markers such as ALDH1, CD34, CD44, EpCAM, LGR5, NANOG, OCT4 and SSEA4 was detected as detailed in Fig. 1a-h using a specific antibody for each marker. Alexafluor-labelled secondary antibody (anti-rabbit or anti-mouse) were employed for detection purpose. Specific expression of markers was detected in ascites derived tumor CSCs either as single cell (a) or clusters of cells (b). CSC markers (green), PTTG1 (red), DAPI (blue) and overlap/merged image of green, red and blue (Overlap). Results represent cells from two patients with recurrent ovarian cancer. Scale bar = 50 μm. (c) Expression of PTTG1 and various CSCs genes in ascites derived CSCs using RT-PCR. Total RNA was purified from ascites CSCs from four patients and was used to determine the expression of PTTG1 and various CSC genes using specific primers (Supplement Table S1) for each gene. PCR amplicons of expected size were detected. High expression levels for PTTG1 and variable expression levels for CSCs genes among four patients were observed. GAPDH primers were used as control primers. Experiments were repeated at least two times
Existence of Self-Renewal and EMT Signaling in Ascites-Derived CSCs
Active self-renewal and EMT signaling mechanisms are frequently reported in studies assessing CSCs [34, 40,41,42,43,44,45,46,47,48,49,50]. In our previous study, we showed that the overexpression of PTTG1 in ovarian epithelial tumor cell line (A2780) induced the EMT process through the regulation of: TGF-β, Twist, Snail, Slug, E-cadherin and Vimentin expression, whereas a knockdown of PTTG1/Securin reversed the expression of genes responsible for EMT [50]. These pathways are linked to the regulation of CSC populations in various cancers, including ovarian cancer [42,43,44,45]. In the present study, we investigated the expression of various genes belonging to stem cell-specific self-renewal pathways: WNT1/β-catenin, NOTCH1, and SHH in ascites-derived CSCs (Fig. 4). Our studies showed high levels of expression of self-renewal related genes (Fig. 4a-c). In addition, our studies also revealed low expression levels of E-cadherin, and high levels of N-cadherin, Vimentin, TGF-β, Snail, Slug and Zeb1 (Fig. 4d), suggesting the persistence of highly active self-renewal mechanisms and EMT related genes in ascites-derived CSCs. High expression of PTTG1 in ascites-derived CSCs with a concomitant increase in self-renewal mechanisms and EMT related genes suggest the capability of PTTG1 to regulate self-renewal and EMT mechanisms. This further suggests a role of PTTG1 in maintaining stem cell/CSC populations in NO, ovarian cancer, and ascites-derived CSCs, respectively.
Detection of stem cell/ cancer stem cell specific-signaling pathways for self-renewal and EMT in ascites derived CSCs. Expression of WNT1 and downstream signaling genes (B-catenin, TCF-4, LEF) (a), NOTCH1 and its downstream signaling genes (Hes1 and Hey1) (b), and SHH and its downstream signaling gene (GLI) (c) were amplified from ascites derived CSCs collected from four different patients with recurrent ovarian cancer by RT/PCR using the specific primers for each gene. GAPDH primers were used as control. Variable levels of gene expression among patients was observed for each gene. Experiments were repeated at least twice. EMT signaling pathway specific genes E-Cadherin, N-Cadherin, Vimentin, TGF-β, Snail, Slug and Zeb1 (D) were amplified from ascites derived CSCs collected from four patients with recurrent ovarian cancer using RT-PCR. GAPDH primers were used as internal control. Experiments were repeated at least twice
PTTG1 Regulates Expression of Stem Cell/Cancer Stem Cell Genes
To assess if PTTG1 regulates the expression of CSC marker genes, in addition to being concomitantly expressed as shown in prior figures, we employed a gene-specific siRNA to knockdown PTTG1 mRNA in the ovarian cancer cell line A2780 and a scramble (negative) siRNA as a negative control. After 48 h of transfection, the transfected cells were harvested and analyzed for the expression of CSC genes by real-time PCR. As shown in Fig. 5, A2780 cells transfected for 48 h with PTTG1 siRNA (at a concentration of 25 nM or 50 nM), showed a significant (90 to 95%) reduction in PTTG1 mRNA expression levels compared to un-transfected cells or cells transfected with scramble siRNA. Western blotting for PTTG1 was performed and showed a minimally present immuno-reactive band in the control and control siRNA groups, with no detectable bands present for the PTTG1 siRNA groups (Data not shown). Analysis of various CSC markers upon treatment with PTTG1 siRNA exhibited a differential regulation in transcript expression. We observed a significant decreased expression of ALDH1, CD44, CD133, LGR5, IFITM3, and SOX2, and a significant increased expression of CD117, DDX4/VASA, NANOG and OCT4A (Table 2A) compared to control (un-transfected) or scramble transfected cells.
Down-regulation and up-regulation ofPTTG1in A2780 cells. a = A2780 cells in growing phase were transfected with scramble (control) siRNA or PTTG1-specific siRNA at a final concentration of 25 nM or 50 nM. After 48 h of transfection RNA was purified and subjected to PTTG1 gene amplification using real-time PCR. GAPDH primers were used as control. The results shown are fold change values compared to control un-transfected cells and are representative of at least 4 independent experiments (n = 4–6 in each group). b and c = A2780 cells were infected with adenovirus expressing PTTG1 or control adenovirus at MOI of 1:4.5 or 1:9.0. After 48 h of infection, cells were harvested to purify RNA or protein. RNA was used for amplification of PTTG1 using real-time PCR (b) and protein was used for western blot analysis (c). Significantly high levels of expression of PTTG1 was observed in cells infected with adenovirus expressing both MOI of 1:4.5 or 1:9 compared to cells infected with control adenovirus. $p ≤ 0.05; $$p ≤ 0.01; $$$p ≤ 0.001 compared to control siRNA (25 nM) as determined by one-way ANOVA and Tukey’s HSDT post hoc analysis. *p ≤ 0.05 compared to control siRNA (50 nM). @p ≤ 0.05 compared to control virus (MOI 1:4.5). #p ≤ 0.05 compared to control virus (MOI 1:9.0)
In contrast, overexpression of PTTG1 using an adenovirus expression system resulted in a significant increase in PTTG1 expression at both the mRNA and protein levels (Fig. 5b and c). Overexpression of PTTG1 resulted in a significant increase in expression of ALDH1, CD133, LGR5, IFITM3, and SOX2, a significant decrease in the expression of CD117, CD44, DDX4 and NANOG and no significant change in the expression of OCT4A compared to control virus transfected cells (Table 2B). Some of the genes that showed down-regulation upon knockdown of PTTG1 did not reveal a transcript expression reversal upon up-regulation of PTTG1, which could be due to the saturation of PTTG1 levels in CSCs or the recruitment of different factors and mechanisms used by PTTG1 upon its up-regulation, as opposed to its down-regulation. At present, it remains unknown if the regulatory effects of PTTG1 on CSC-specific gene expression are direct or indirect as well as the mechanisms involved in their differential regulation.
Effect of PTTG1 on Regulation of Self-Renewal Mechanisms
Stem cells and CSCs undergo self-renewal and share similar pathways modulating this facet, such as WNT/β-catenin, NOTCH1, Sonic hedgehog (SHH), STAT3, and NF-κB [44,45,46,47]. Crosstalk between various cell signaling pathway components forms a complex biological network that is functionally dys-regulated in cancer and CSCs. This dys-regulation leads to tumorigenic self-renewal and a subsequent heterogeneous cellular differentiation producing different tumor cell types within a tumor. This complex scenario offers a diverse range of cell surface receptors, ligands, and thus potential targets worth exploration for therapeutic purposes [48]. With this in mind, we focused on studying the regulation of self-renewal mechanisms with respect to the oncogene PTTG1. To determine the effect of PTTG1 on the regulation of self-renewal pathways, we performed knockdown of PTTG1 using gene-specific siRNA as described above. Knockdown of PTTG1 in A2780 cells resulted in a down-regulation of TGF-β, Snail, Slug and Zeb1 (Fig. 6a), suggesting a contribution of PTTG1 in maintaining CSC populations and a CSC-phenotype. In addition, knockdown of PTTG1 in A2780 cells resulted in a significant down-regulation of β-catenin and its downstream effector genes (TCF4, c-Myc, and cyclin D1) (Fig. 6b). Similarly, we observed a significant down-regulation of NOTCH1 and its downstream effector genes (Hes1 and Hey1) (Fig. 6c). Collectively our results suggest that the regulation of CSC marker genes by PTTG1 could possibly be achieved through the regulation of key CSC self-renewal mechanisms and EMT specific pathways. Direct, indirect regulation and other compensatory pathways operating to execute stemness and uninterrupted cellular functioning seems to be a more complex phenomenon and a Systems biology based approach to decipher various questions regarding the functioning and regulation of PTTG1 oncogene is anticipated.
Effect of knockdown ofPTTG1on EMT and self-renewal (WNT1/β-catenin and NOTCH1) genes. A2780 cells were transfected with control siRNA or PTTG1-specific RNA. After 48 h of transfection, RNA was purified and subjected to amplification for EMT genes (TGF-β, Snail, Slug and Zeb 1) (a), Self-renewal specific (WNT1/β-catenin signaling genes (β-catenin, TCF-4, cMYC and Cyclin D1) (b), NOTCH1 signaling genes (NOTCH1, Hes1 and Hey 1) (c) using specific primers in real-time PCR. GAPDH primers were used as control. Results shown are fold change in expression levels for each gene. PTTG1-specific siRNA greatly reduced the expression levels of each gene compared to control un-transfected cells or cells transfected with control siRNA. Results shown are representative of at least five independent experiments (n = 5–7 in each group). *p ≤ 0.05; **p ≤ 0.01; ***p ≤ 0.001; ****p ≤ 0.0001 compared to control siRNA (50 nM) as determined by one-way ANOVA and Tukey’s HSDT post hoc analysis
Discussion
The results of this study suggest PTTG1/securin as a novel marker for stem cells/CSCs that regulates the expression of several stem cell and CSC-related genes through the regulation of self-renewal and EMT specific pathways, underlying a previously overlooked functionality of PTTG1. To the best of our knowledge, this is the first study proposing this unique role of oncogene PTTG1/securin and its potential influence on both the stem cell/CSC compartments in the ovary, ovarian tumors and ascites.
In the current study, we showed co-expression of PTTG1 with several pluripotent stem cell, CSC, and germ stem cell markers in normal ovary and at various stages of ovarian tumorigenesis (BN, BL, and HG), and in ascites-derived CSCs collected from patients with recurrent ovarian cancer, underscoring a conserved functionality shared between normal and pathological states. Since stem cells and CSCs share similar markers, our results suggest a compelling possibility that stem cells present within the OSE regions in normal ovary may be undergoing malignant transformation and spread across the cortex explaining the origin of epithelial ovarian tumors [51, 52]. In addition, our study revealed the regulation of self-renewal mechanisms by PTTG1, suggesting that PTTG1 may serve as i) a novel marker for stem cells and CSCs and ii) a possible switch to maintain stem/CSC populations. Therefore, it could be responsible for the transformation of normal stem cells into CSCs related to change in expression levels of PTTG1 and possible alterations in the microenvironment [53, 54]. Another study has reported the co-expression of stem cell and CSC-specific genes (OCT4, NANOG, SOX2, BMI-1, NESTIN, CD24, CD44, CD117, CD133, ALDH1, and ABCG2) in high grade serous ovarian carcinoma and tumor derived spheres/CSCs [55], thus pointing towards a (cancer) stem cell hierarchy responsible for metastasis, malignancy and chemo resistance, highlighting the possibility of malignant cell migration from HG tumors to secondary targets in metastasis. Similarly, expression of common markers in ovarian tumor stages (BN, BL and HG) and ascites-derived CSCs suggests a possible origin of metastatic CSCs present in ascites fluid from the disseminated HG ovaries reported in present study.
PTTG1 has transforming activity in vitro and tumorigenic activity in vivo. This gene is highly expressed in various tumors, including ovarian [5,6,7,8, 33], but has not been related to CSCs. Its role in tumor initiation, growth, angiogenesis, and progression is well documented in several tumor types but molecular mechanisms remain unknown. Some studies delineated PTTG1 role in induction of cell proliferation by down-regulation of oncogenes v-Jun and v-maf and up-regulation of the histone family of genes [56].The encoded protein of PTTG1 is a homolog of yeast securin proteins, which prevent separins from promoting sister chromatid separation [9] and plays a central role in chromosome stability through the regulation of p53/TP53 pathway, and DNA repair known as key processes in the manifestation of cancer. Therefore, CSCs expressing the biomarker PTTG1 along with other bio markers could be important for the isolation and targeting of ovarian CSCs with respect to treatment, drug resistance, and tumor relapse in ovarian cancer patients.
Activation of oncogenes/oncogenic pathways coupled with a self-renewal and multi-lineage differentiation properties of CSCs and further endowment with metastatic nature and resistance to chemo and radio therapy are reported so far [57]. It is also hypothesized that putative perturbations (genetic mutations and epigenetic regulations) in the stem cell compartment and its immediate microenvironment could possibly lead to ovarian tumor development [50] which requires further investigation. Since CSC populations exhibit stemness properties linked with the key embryonic signaling pathways, such as: STAT3, NANOG, NOTCH, WNT, and SHH [58], understanding their dys-regulation may help to elucidate the distinction between normal stem cells and CSCs and thus help to improve the patient/clinical outcome [59].
In this study, gene specific siRNA based knockdown of PTTG1 in the ovarian cancer cell line A2780 revealed an overall 90% down-regulation, accompanied by down-regulation of several stem cell and CSC genes, self-renewal and EMT signaling pathways along with all other complex activities, as proposed by other studies [34, 60]. Results also implicate hampered metastatic potential of CSCs due to down-regulation of PTTG1, which in turn underlines the putative role of PTTG1 in regulating EMT and metastasis. An earlier study from our group involving adenoviral-vector based overexpression of PTTG1 resulted in a significant increase in expression of integrins αV and β3 of αVβ3-FAK signaling pathway and downstream signaling genes RAC1, RHOA, CDC42, and DOCK180 specific for EMT signaling. On the contrary adenovirus expressing PTTG1-specific siRNA reversed the process thus implicating PTTG1/securin oncogene in the regulation of integrins αV and β3 and adhesion complex proteins leading to induction of EMT [61] In the present study, ectopic expression of PTTG1 by adeno-viral expression system led to the over-expression of most of the CSC specific genes, thus implicating the role of PTTG1 gene in regulating CSC populations in ovarian cancer cells.
While overlapping transcriptomic signatures comprising of TGFβ, WNT1/β-catenin, HEDGEHOG (HH), NOTCH between EMT and CSC phenotypes are reported [62], molecular pathways implicated in metastatic spread and malignancy are also known to regulate both EMT and CSCs [63]. NOTCH1, WNT1 and HH pathways known to regulate embryonic and adult stem cells, also regulate stem cells expressing self-renewal property and OCT4, SOX2 and even CD44 genes. Overexpression of pluripotency related transcription factors OCT4, NANOG, and SOX2 are associated with tumor transformation, inhibition of apoptosis, tumorigenicity and tumor metastasis [64], while stemness state of tumor resonated with elevated risk of worse disease-free survival and poor outcome [65]. Other recent study in breast cancer cells has exemplified the role of PTTG1 in modulating EMT and CSC population thus highlighting the therapeutic significance of this onco protein [34].
The expression of CSC markers, including PTTG1, in our study reflects the presence of CSCs in ovarian tissues, which were present both in the OSE and cortex regions. Stem-like cells have already been reported in the OSE lining, oviductal epithelium, fallopian tube, fimbrial cells and hilum regions of the ovary [52, 53, 66]. More than a decade back, mitotically active germ stem cells in adult rodent [67, 68], human [69,70,71,72] and ovine [69] ovaries were also demonstrated with the potential to be involved in the cancer manifestation [35, 36, 59, 61, 73, 74]. In our study, the expression of germ lineage-related markers in CSCs were found in all samples tested, thus indicating the presence of stem cell related to the germinal lineage in ovarian tissues which may be involved in the manifestation of ovarian cancer. Isolation of PTTG1-positive cells in future from ovarian tumors may better explain the phenotype and characteristics of ovarian CSCs in humans.
Ovarian cancer is the seventh most common cancer among women worldwide and is potentially the most lethal form of all gynaecological cancers affecting females in almost all ages, especially menopause [75]. A number of therapies are under clinical trials [76, 77]. Although, to date, a limited numbers of FDA approved drugs for ovarian cancer are available that neither result in complete remission, nor an appreciable improvement in overall survival [78]. Ovarian CSC-specific therapies appear to be of great interest due to the possibility of reducing drug resistance and tumor relapse, with fewer negative side effects than conventional therapies. Due to the lack of fundamental knowledge regarding the CSC population downstream targets, and drivers of signaling mechanisms regulating their self-renewal, further concerted efforts are warranted. Therefore, the insights gleaned in this study indicating PTTG1 as a unique marker for stem cells and/or CSCs is of significant advance and may lay the foundations for and present excellent leads towards developing new therapies against ovarian cancer (stem cells) in the near future.
Materials and Methods
Ethical Approval for Use of Human Tissues
Tissue and ascites samples were collected from ovarian cancer patients (≥ 18 years of age) admitted to the James Graham Brown Cancer Center, University of Louisville (under biorepository’s IRB number 08.0388 and IRB exempt protocol number 16.0490 for specimen used) by the University of Louisville. Informed patient consent was obtained prior to surgery. All protocols and associated ethics were reviewed and approved by the University of Louisville’s Institutional Review Board (IRB) before the start of the study.
Cell Line and Cell Culture
The A2780 ovarian epithelial cancer cell line was maintained in Roswell Park Memorial Institute (RPMI) Medium-1640 supplemented with: 10% Fetal Bovine Serum (FBS, Hyclone Laboratories Inc., GE Healthcare), 100 U/ml Penicillin, and 10 μg/ml Streptomycin (MilliporeSigma, Burlington, Massachusetts, USA). Cells were cultured in a humidified atmosphere of 5% CO2 at 37 °C, and the medium was changed every 48 h as described previously [79]. Cell line was a generous gift from Denise Connolly (Fox Chase Cancer Center).
Collection of Ascites from Patients with Ovarian Cancer and Isolation of CSCs
Ascites fluid samples were collected from patients (≥ 18 years of age) diagnosed with recurrent ovarian cancer that had completed their first round of chemotherapy (usually cisplatin or a combination of carboplatin and paclitaxel) and were scheduled for surgery or a second round of chemotherapy. An appropriately signed informed consent was obtained from each patient prior to surgery. Subjects with HIV and those treated with antineoplastic drugs before being hospitalized or with pregnancy were excluded from the study. Approval of the protocol and ethical oversights were obtained from the IRB of the University of Louisville. Independent staff pathologists determined histological diagnosis as part of the patient’s clinical diagnosis. Approximately one liter of ascites fluid was processed to collect cells following the procedures described by Latifi et al., [37]. Briefly, ascites fluid was centrifuged to collect all the nucleated cells. Contaminating red blood cells were removed by treating with red blood cell lysis buffer [ammonium-chloride-potassium (ACK)]. Ascites cells were seeded on ultra-low attachment plates (Corning, NY, USA) in RPMI 1640 growth medium supplemented with fetal bovine serum (10%), glutamine (2 mM) and penicillin/streptomycin. Cells were maintained at 37 °C in the presence of 5% CO2. Under these conditions, non-adhering (NAD) cells float as spheroids (characteristic of ascites derived CSCs), while adhering (AD) non-CSCs attach to the plate [37,38,39]. After 2 to 3 days of incubation, floating NAD spheroids were collected and fixed in 10% buffered formalin for use in subsequent experimentation.
Preparation of Tissues Blocks, Immunofluorescence Staining and Confocal Microscopy
Freshly collected normal ovarian and ovarian tumor tissues (BN, BL, and HG) were fixed in 10% buffered formalin, then processed and embedded in paraffin using protocols as described previously [35, 36]. NAD ascites cell pellet was centrifuged, fixed in 10% buffered formalin, and then transferred to molten 2% agarose gel. A cellular “button” was obtained upon solidifying of the Agarose gel, which was subsequently embedded in paraffin wax and processed as described previously [35, 36]. Five μm thick sections of the embedded normal ovarian tissues, ovarian cancer tumors, and ascites cells were de-paraffinized in xylene and rehydrated in decreasing graded series of ethanol as described previously [80]. Sections were rinsed three times with PBS followed by antigen retrieval as optimized previously. Next, the sections were blocked with 5% normal goat serum for 60 min at room temperature. Following, sections were incubated with PTTG1 polyclonal antibody (1:1500) and CSC marker-specific monoclonal antibodies for ALDH1, CD44, LGR5, OCT4, SSEA4, CD34 and NANOG at appropriate dilution as described previously (35, 36) and incubated overnight at 4 °C. Sections were rinsed thrice with PBS (5 min each) and incubated with labeled secondary antibody [Alexa Fluor 488/568 labeled goat anti-mouse IgG and/or goat anti rabbit-IgG (1:1000) (Molecular Probes, Thermo Fisher Scientific, Life Technologies Corp., NY, USA)] for 45–60 min and counterstained with nuclear dye 4′, 6-diamidino-2-phenylindole (DAPI; MilliporeSigma, USA). The sections were rinsed with PBS, dehydrated and mounted in mounting medium (Eukitt Quick-hardening; MilliporeSigma, USA). Approximately 10 representative images covering both the OSE and cortex regions of the ovarian tissues, ascites cell clusters and single cells were captured (at 40x magnification) using Nikon (Eclipse TI) laser scanning confocal microscope and NIS Elements AR software (version 4.5.1). Background noise interference during confocal microscopy was maintained at minimum threshold limits by setting imaging parameters precisely and maintaining uniform parameters throughout imaging. Experiments were repeated at least three times to attain reproducible results. Images were processed uniformly on Adobe Photoshop CS3 (version 10) to prepare final image panels with a resolution of at least 300 dpi.
RNA Purification and Real-Time PCR
RNA was purified from various tissues and cells using an RNeasy Mini Kit (Qiagen Inc., Maryland, USA; Catalog # 74104) and the accompanying manufacturer’s protocol, as previously detailed [80]. First strand cDNA was synthesized using 1 μg of purified RNA and a commercially available kit (iScript™ cDNA synthesis, Bio-Rad Catalog # 170–8891). Quantification of mRNA expression was performed similar to previously described protocol using the SYBR Green dye method on a StepOnePlus™ system (Applied Biosystems, California, USA) using gene-specific primers [80]. Specific primers used for each gene are listed in Supplement Table S1.
Isolation of ALDH1 Positive CSCs
We isolated ALDH1-positive CSCs from A2780 cells as described previously [80]. Briefly, A2780 cells growing in log phase were rinsed with PBS and harvested by using non-enzymatic cell dissociation solution (MilliporeSigma, USA) followed by incubation at 37 °C for 45 min. After centrifugation at 1500 rpm for 3 min, cells were resuspended in binding buffer from Aldefluor kit (Stem Cell Technologies, Vancouver, BC, Canada) at 2 × 106 cells/ml and incubated with Aldefluor substrate (1 μM) at 37 °C for 45 min followed by centrifugation at 1500 rpm for 3 min and finally resuspended in binding buffer at a concentration of 10 × 106/ml. Negative control samples were treated with 50 mmol/L of diethylaminobenzaldehyde (DEAB, an ALDH1 inhibitor) before adding Adelfluor substrate as described by Ginestier et al. (2007) (ref. 81). Highly bright (ALDH1+/hi, considered to be ALDH1+) and those with very low brightness (ALDH1+/low, considered to be ALDH1−) were detected in the green fluorescence channel (520–540 nm) using Beckman Coulter MoFlo XDP and collected in RPMI-1640 medium containing 10% FBS. The ALDH1+ cells were plated on ultra-low attachment plates in a RPMI medium containing 1% fetal bovine serum, 0.5% BSA, 5 μg/ml insulin, 20 ng/ml epidermal growth factor and 20 ng/ml basic fibroblast growth factor. The ALDH1− cells were plated on regular tissue culture plates in RPMI-1640 medium containing insulin and 10% FBS. Cells were incubated at 37 °C and 5% CO2 for 3 to 5 days. Spheroids formed by ALDH1+ cells were collected after 5 days of plating, whereas ALDH1− cells were harvested after two days of plating. Total RNA from ALDH1+ and ALDH1− cells were used for RNA purification as described above.
Knockdown of PTTG1
siRNA was employed to knockdown PTTG1 mRNA in the ovarian cancer cell line A2780 to assess if PTTG1 regulates expression of all or some of the CSC marker genes. Scramble (negative) siRNA (Ambion, Thermo Fisher Scientific, Waltham, Massachusetts, United States cat # 4390843) and PTTG1-specific siRNA (cat # 4390824) (sense 5’-CCCGUGUGGUUGCUAAGGATT-3′) and (antisense 5’-UCCUUAGCAACCACACGGGTG-3′) were used. A2780 cells were plated at a concentration of ~2 × 105 cells/well of 6 well plates for 24 h and were transfected with siRNA to a final concentration of 25 nM or 50 nM using the transfectin reagent (BioRad Laboratories, California, USA) in a growth medium containing 5% fetal bovine serum as described previously [27]. After 48 h of transfection, cells were harvested and total RNA was purified.
Overexpression of PTTG1
Ovarian cancer cells (A2780) were seeded in six well plates (2 × 105 cells/well of 6 well plates). After 24 h of plating, cells were infected with adenovirus expressing PTTG1 or control adenovirus at MOI 1:4.5 or 9.0, similar to protocol described previously [51]. After 48 h of infection, cells were harvested to examine the overexpression of PTTG1 mRNA using real-time PCR or protein analysis.
Western Blot Analysis
A2780 cells were harvested in PBS and lysed in a chilled RIPA lysis buffer (MilliporeSigma) supplemented with a protease inhibitor cocktail (MilliporeSigma). Protein in each sample was quantitated using Bradford Reagent (BioRad Laboratories, California, USA), and bovine serum albumin as a standard. An equal amount of protein (40 μg) from each sample was denatured with 1× SDS sample buffer at 95 °C for 5 min and resolved on a 10% SDS-PAGE gel and subsequently transferred onto a nitrocellulose membrane. Blots were probed with anti-PTTG1 antibody [82] at a dilution of 1:1500 in tris-buffered saline containing 0.1% Tween-20 (TBST). Immuno-reactive proteins were visualized using the Enhanced Chemiluminescent Detection system kit from MilliporeSigma, according to manufacturer’s instructions. The membrane was stripped off and re-probed with 1% horseradish peroxidase-labeled β-actin monoclonal antibody (MilliporeSigma, USA) to normalize the variation in loading of samples.
Statistical Analysis
Student’s t test was performed to calculate the statistical differences between the control and experimental groups. For comparisons between three groups, a one-way analysis of variance was performed, followed by Tukey’s Honestly Significant Difference Test post hoc analysis where indicated. p ≤ 0.05 was considered statistically significant. The error bars represent the standard deviation of independent experiments.
Data Availability
All relevant data related to this manuscript are available from the authors.
References
Torre, L. A., Trabert, B., DeSantis, C. E., Miller, K. D., Samimi, G., Runowicz, C. D., Gaudet, M. M., Jemal, A., & Siegel, R. L. (2018). Ovarian cancer statistics, 2018. CA: A Cancer J Clin, 68(4), 284–296.
Feramisco, J. R., Gross, M., Kamata, T., Rosenberg, M., & Sweet, R. W. (1984). Microinjection of the oncogene form of the human H-ras (t-24) protein results in rapid proliferation of quiescent cells. Cell, 38(1), 109–117.
Bollig-Fischer, A., Dewey, T. G., & Ethier, S. P. (2011). Oncogene activation induces metabolic transformation resulting in insulin-independence in human breast cancer cells. PLoS One, 6(3), e17959. https://doi.org/10.1371/journal.pone.0017959.
Shortt, J., & Johnstone, R. W. (2012). Oncogenes in cell survival and cell death. Cold Spring Harbor Perspectives in Biology, 4(12), a009829. https://doi.org/10.1101/cshperspect.a009829.
Kakar, S. S., & Jennes, L. (1999). Molecular cloning and characterization of the tumor transforming gene (TUTR1): A novel gene in human tumorigenesis. Cytogenetics and Cell Genetics, 84(3–4), 211–216.
Pei, L., & Melmed, S. (1997). Isolation and characterization of a pituitary tumor-transforming gene (PTTG). Molecular Endocrinology, 11(4), 433–441.
Domínguez, A., Ramos-Morales, F., Romero, F., Rios, R. M., Dreyfus, F., Tortolero, M., & Pintor-Toro, J. A. (1998). Hpttg, a human homologue of rat pttg, is overexpressed in hematopoietic neoplasms. Evidence for a transcriptional activation function of hPTTG. Oncogene, 7(17), 2187–2193.
Puri, R., Tousson, A., Chen, L., & Kakar, S. S. (2001). Molecular cloning of pituitary tumor transforming gene 1 from ovarian tumors and its expression in tumors. Cancer Letters, 163(1), 131–139.
Zou, H. (1999). Identification of a vertebrate sister-chromatid separation inhibitor involved in transformation and tumorigenesis. Science, 285(5426), 418–422. https://doi.org/10.1126/science.285.5426.418.
Hamid, T., Malik, M. T., & Kakar, S. S. (2005). Ectopic expression of PTTG1/securin promotes tumorigenesis in human embryonic kidney cells. Molecular Cancer, 4(1), 3. https://doi.org/10.1186/1476-4598-4-3.
Kim, D., Pemberton, H., Stratford, A. L., Buelaert, K., Watkinson, J. C., Lopes, V., et al. (2005). Pituitary tumour transforming gene (PTTG) induces genetic instability in thyroid cells. Oncogene, 24(30), 4861–4866. https://doi.org/10.1038/sj.onc.1208659.
Yu, R., Heaney, A. P., Lu, W., Chen, J., & Melmed, S. (2000). Pituitary tumor transforming gene causes aneuploidy and p53-dependent and p53-independent apoptosis. Journal of Biological Chemistry, 275(47), 36502–36505. https://doi.org/10.1074/jbc.c000546200.
Wierinckx, A., Auger, C., Devauchelle, P., Reynaud, A., Chevallier, P., Jan, M., et al. (2007). A diagnostic marker set for invasion, proliferation, and aggressiveness of prolactin pituitary tumors. Endocrine-Related Cancer, 14(3), 887–900. https://doi.org/10.1677/erc-07-0062.
Zhang, J., Yang, Y., Chen, L., Zheng, D., & Ma, J. (2014). Overexpression of pituitary tumor transforming gene (PTTG) is associated with tumor progression and poor prognosis in patients with esophageal squamous cell carcinoma. Acta Histochemica, 116(3), 435–439. https://doi.org/10.1016/j.acthis.2013.09.011.
Yan, S., Zhou, C., Lou, X., Xiao, Z., Zhu, H., Wang, Q., & Xu, N. (2009). PTTG overexpression promotes lymph node metastasis in human esophageal squamous cell carcinoma. Cancer Research, 69(8), 3283–3290. https://doi.org/10.1158/0008-5472.can-08-0367.
Zhou, C., Tong, Y., Wawrowsky, K., & Melmed, S. (2014). PTTG acts as a STAT3 target gene for colorectal cancer cell growth and motility. Oncogene, 33(7), 851–861. https://doi.org/10.1038/onc.2013.16.
Liu, J., Wang, Y., He, H., Jin, W., & Zheng, R. (2015). Overexpression of the pituitary tumor transforming gene upregulates metastasis in malignant neoplasms of the human salivary glands. Experimental and Therapeutic Medicine, 10(2), 763–768. https://doi.org/10.3892/etm.2015.2566.
Solbach, C., Roller, M., Eckerdt, F., Peters, S., & Knecht, R. (2006). Pituitary tumor-transforming gene expression is a prognostic marker for tumor recurrence in squamous cell carcinoma of the head and neck. BMC Cancer, 6, 242. https://doi.org/10.1186/1471-2407-6-242.
Hunter, J., Skelly, R., Aylwin, S., Geddes, J., Evanson, J., Besser, G., & Burrin, J. (2003). The relationship between pituitary tumour transforming gene (PTTG) expression and in vitro hormone and vascular endothelial growth factor (VEGF) secretion from human pituitary adenomas. European Journal of Endocrinology, 203–211. https://doi.org/10.1530/eje.0.1480203.
Kim, D., Buchanan, M., Stratford, A., Watkinson, J., Eggo, M., Franklyn, J., & Mccabe, C. (2006). PTTG promotes a novel VEGF-KDR-ID3 autocrine mitogenic pathway in thyroid cancer. Clinical Otolaryngology, 31(3), 246–246. https://doi.org/10.1111/j.1749-4486.2006.01236_6.x.
Ishikawa, H. (2001). Human pituitary tumor-transforming gene induces angiogenesis. Journal of Clinical Endocrinology & Metabolism, 86(2), 867–874. https://doi.org/10.1210/jc.86.2.867.
Malik, M. T., & Kakar, S. S. (2006). Regulation of angiogenesis and invasion by human pituitary tumor transforming gene (PTTG) through increased expression and secretion of matrix metalloproteinase-2 (MMP-2). Molecular Cancer, 5, 61. https://doi.org/10.1186/1476-4598-5-61.
Pei, L. (2000). Identification of c-mycas a Down-stream target for pituitary tumor-transforming gene. Journal of Biological Chemistry, 276(11), 8484–8491. https://doi.org/10.1074/jbc.m009654200.
Heaney, A. P., Horwitz, G. A., Wang, Z., Singson, R., & Melmed, S. (1999). Early involvement of estrogen-induced pituitary tumor transforming gene and fibroblast growth factor expression in prolactinoma pathogenesis. Nature Medicine, 5(11), 1317–1321. https://doi.org/10.1038/15275.
Yu, R. (2000). Pituitary tumor transforming gene (PTTG) regulates placental JEG-3 cell division and survival: Evidence from live cell imaging. Molecular Endocrinology, 14(8), 1137–1146. https://doi.org/10.1210/me.14.8.1137.
Levenstein, M. E., Ludwig, T. E., Xu, R., Llanas, R. A., Vandenheuvel-Kramer, K., Manning, D., & Thomson, J. A. (2006). Basic fibroblast growth factor support of human embryonic stem cell self-renewal. Stem Cells, 24(3), 568–574. https://doi.org/10.1634/stemcells.2005-0247.
Kakar, S., & Malik, M. (2006). Suppression of lung cancer with siRNA targeting PTTG. International Journal of Oncology. https://doi.org/10.3892/ijo.29.2.387.
El-Naggar, S., Malik, M., & Kakar, S. (2007). Small interfering RNA against PTTG: A novel therapy for ovarian cancer. International Journal of Oncology. https://doi.org/10.3892/ijo.31.1.137.
Jung, C., Yoo, J., Jang, Y. J., Kim, S., Chu, I., Yeom, Y. I., & Im, D. (2006). Adenovirus-mediated transfer of siRNA against PTTG1 inhibits liver cancer cell growthin vitroandin vivo. Hepatology, 43(5), 1042–1052. https://doi.org/10.1002/hep.21137.
El-Naggar, S. M., Malik, M. T., Martin, A., Moore, J. P., Proctor, M., Hamid, T., & Kakar, S. S. (2007). Development of cystic glandular hyperplasia of the endometrium in Mullerian inhibitory substance type II receptor-pituitary tumor transforming gene transgenic mice. Journal of Endocrinology, 194(1), 179–191. https://doi.org/10.1677/joe-06-0036.
Abbud, R. A., Takumi, I., Barker, E. M., Ren, S., Chen, D., Wawrowsky, K., & Melmed, S. (2005). Early multipotential pituitary focal hyperplasia in the α-subunit of glycoprotein hormone-driven pituitary tumor-transforming gene transgenic mice. Molecular Endocrinology, 19(5), 1383–1391. https://doi.org/10.1210/me.2004-0403.
Chesnokova, V., Kovacs, K., Castro, A. V., Zonis, S., & Melmed, S. (2005). Pituitary hypoplasia in Pttg−/− mice is protective for Rb+/− pituitary tumorigenesis. Molecular endocrinology (Baltimore, Md.), 19(9), 2371–2379. https://doi.org/10.1210/me.2005-0137.
Lewy, G. D., Sharma, N., Seed, R. I., Smith, V. E., Boelaert, K., & McCabe, C. J. (2012). The pituitary tumor transforming gene in thyroid cancer. Journal of Endocrinological Investigation, 35(4), 425–433 Review.
Yoon, C. H., Kim, M. J., Lee, H., Kim, R. K., Lim, E. J., Yoo, K. C., et al. (2012). PTTG1 oncogene promotes tumor malignancy via epithelial to mesenchymal transition and expansion of cancer stem cell population. The Journal of Biological Chemistry, 287(23), 19516–19527. https://doi.org/10.1074/jbc.M111.337428.
Parte, S. C., Smolenkov, A., Batra, S. K., Ratajczak, M. Z., & Kakar, S. S. (2017). Ovarian Cancer stem cells: Unraveling a germline connection. Stem Cells and Development, 26(24), 1781–1803. https://doi.org/10.1089/scd.2017.0153.
Parte, S. C., Batra, S. K., & Kakar, S. S. (2018). Characterization of stem cell and cancer stem cell populations in ovary and ovarian tumors. Journal of Ovarian Research, 11(1), 69. https://doi.org/10.1186/s13048-018-0439-3.
Latifi, A., Luwor, R. B., Bilandzic, M., Nazaretian, S., Stenvers, K., Pyman, J., & Ahmed, N. (2012). Isolation and characterization of tumor cells from the ascites of ovarian cancer patients: Molecular phenotype of chemoresistant ovarian tumors. PLoS One, 7(10), e46858. https://doi.org/10.1371/journal.pone.0046858.
Bapat, S. A., Mali, A. M., Koppikar, C. B., & Kurrey, N. K. (2005). Stem and progenitor-like cells contribute to the aggressive behavior of human epithelial ovarian cancer. Cancer Research., 65(8), 3025–3029.
Ahmed, N., & Stenvers, K. L. (2013). Getting to know ovarian cancer ascites: Opportunities for targeted therapy-based translational research. Frontiers in Oncology, 3, 256. https://doi.org/10.3389/fonc.2013.00256.
Yadav, A. K., & Desai, N. S. (2019). Cancer stem cells: Acquisition, characteristics, therapeutic implications. Targeting Strategies and Future Prospects. Stem Cell Rev., 15(3), 331–355. https://doi.org/10.1007/s12015-019-09887-2.
Moustakas, A., & Heldin, C. H. (2007). Signaling networks guiding epithelial–mesenchymal transitions during embryogenesis and cancer progression. Cancer Science, 98(10), 1512–1520.
Borah, A., Raveendran, S., Rochani, A., Maekawa, T., & Kumar, D. S. (2015). Targeting self-renewal pathways in cancer stem cells: Clinical implications for cancer therapy. Oncogenesis, 4(11), e177. https://doi.org/10.1038/oncsis.2015.35.
Marquardt, S., Solanki, M., Spitschak, A., Vera, J., & Pützer, B. M. (2018). Emerging functional markers for cancer stem cell-based therapies: Understanding signaling networks for targeting metastasis. Seminars in Cancer Biology, 53, 90–109. https://doi.org/10.1016/j.semcancer.2018.06.006.
Reguart, N., He, B., Taron, M., You, L., Jablons, D. M., & Rosell, R. (2005). The role of Wnt signaling in cancer and stem cells. Future Oncology, 1(6), 787–797. https://doi.org/10.2217/14796694.1.6.787.
Ingham, P. W., & Placzek, M. (2006). Orchestrating ontogenesis: Variations on a theme by sonic hedgehog. Nature Reviews Genetics, 7(11), 841–850. https://doi.org/10.1038/nrg1969.
Abel, E. V., Kim, E. J., Wu, J., Hynes, M., Bednar, F., Proctor, E., & Simeone, D. M. (2014). The Notch pathway is important in maintaining the cancer stem cell population in pancreatic cancer. PLoS One, 9(3), e91983. https://doi.org/10.1371/journal.pone.0091983.
Lin, J. C., Tsai, J. T., Chao, T. Y., Ma, H. I., & Liu, W. H. (2018). The STAT3/slug Axis enhances radiation-induced tumor invasion and Cancer stem-like properties in Radioresistant glioblastoma. Cancers, 10(12), 512. https://doi.org/10.3390/cancers10120512.
Koury, J., Zhong, L., & Hao, J. (2017). Targeting signaling pathways in Cancer stem cells for Cancer treatment. Stem Cells International, 2017, 2925869–2925810. https://doi.org/10.1155/2017/2925869.
Udoh, K., Parte, S., Carter, K., Mack, A., & Kakar, S. S. (2019). Targeting of lung Cancer stem cell self-renewal pathway by a small molecule Verrucarin J. Stem Cell Reviews, 15(Mar 5), 601–611.
Bao, B., Ahmad, A., Azmi, A. S., Ali, S., & Sarkar, F. H. (2013). Overview of cancer stem cells (CSCs) and mechanisms of their regulation: Implications for cancer therapy. Current Protocols in Pharmacology, chapter 14, unit–14.25. https://doi.org/10.1002/0471141755.ph1425s61.
Shah, P. P., & Kakar, S. S. (2011). Pituitary tumor transforming gene induces epithelial to mesenchymal transition by regulation of twist, snail, slug, and E-cadherin. Cancer Letters, 311(1), 66–76. https://doi.org/10.1016/j.canlet.2011.06.033.
Ng, A., & Barker, N. (2015). Ovary and fimbrial stem cells: Biology, niche and cancer origins. Nature Reviews Molecular Cell Biology, 16(10), 625–638. https://doi.org/10.1038/nrm4056.
Flesken-Nikitin, A., Hwang, C. I., Cheng, C. Y., Michurina, T. V., Enikolopov, G., & Nikitin, A. Y. (2013). Ovarian surface epithelium at the junction area contains a cancer-prone stem cell niche. Nature, 495(7440), 241–245. https://doi.org/10.1038/nature11979.
Polyak, K., Haviv, I., & Campbell, I. G. (2009). Co-evolution of tumor cells and their microenvironment. Trends in Genetics, 25, 30–38.
He, Q. Z., Luo, X. Z., Wang, K., Zhou, Q., Ao, H., Yang, Y., Li, S. X., Li, Y., Zhu, H. T., & Duan, T. (2014). Isolation and characterization of cancer stem cells from high-grade serous ovarian carcinomas. Cellular Physiology and Biochemistry, 33, 173–184.
Panguluri, S. K., & Kakar, S. S. (2009). Effect of PTTG on endogenous gene expression in HEK 293 cells. BMC Genomics, 10, 577. https://doi.org/10.1186/1471-2164-10-577.
Willis, R. E. (2016). Targeted Cancer therapy: Vital oncogenes and a new molecular genetic paradigm for Cancer initiation progression and treatment. International Journal of Molecular Sciences, 17(9), 1552. https://doi.org/10.3390/ijms17091552.
Oren, O., & Smith, B. D. (2017). Eliminating Cancer stem cells by targeting embryonic signaling pathways. Stem Cell Rev. and Rep., 3(1), 17–23. https://doi.org/10.1007/s12015-016-9691-3.
Lathia, J. D., & Liu, H. (2017). Overview of Cancer stem cells and Stemness for community oncologists. Targeted Oncology, 12(4), 387–399. https://doi.org/10.1007/s11523-017-0508-3.
Kenda Suster, N., Smrkolj, S., & Virant-Klun, I. (2017). Putative stem cells and epithelial-mesenchymal transition revealed in sections of ovarian tumor in patients with serous ovarian carcinoma using immunohistochemistry for vimentin and pluripotency-related markers. Journal of Ovarian Research, 10(1), 11. https://doi.org/10.1186/s13048-017-0306-7.
Shah, P. P., Fong, M. Y., & Kakar, S. S. (2012). PTTG induces EMT through integrin αVβ3-focal adhesion kinase signaling in lung cancer cells. Oncogene, 31(26), 3124–3135. https://doi.org/10.1038/onc.2011.488.
Nguyen, L. V., Vanner, R., Dirks, P., & Eaves, C. J. (2012). Cancer stem cells: An evolving concept. Nature Reviews. Cancer, 12, 133–143.
Wang, S. S., Jiang, J., Liang, X. H., & Tang, Y. L. (2015). Links between cancer stem cells and epithelial-mesenchymal transition. OncoTargets and Therapy, 8, 2973–2980. https://doi.org/10.2147/OTT.S91863.
Ben-Porath, I., Thomson, M. W., Carey, V. J., Ge, R., Bell, G. W., Regev, A., & Weinberg, R. A. (2008). An embryonic stem cell-like gene expression signature in poorly differentiated aggressive human tumors. Nature Genetics, 40(5), 499–507. https://doi.org/10.1038/ng.127.
Schwede, M., Spentzos, D., Bentink, S., Hofmann, O., Haibe-Kains, B., Harrington, D., et al. (2013). Stem cell-like gene expression in ovarian cancer predicts type II subtype and prognosis. PLoS One, 8(3), e57799. https://doi.org/10.1371/journal.pone.0057799.
Auersperg, N. (2013). The stem-cell profile of ovarian surface epithelium is reproduced in the oviductal fimbriae, with increased stem-cell marker density in distal parts of the fimbriae. International Journal of Gynecological Pathology, 32, 444–453.
Zou, K., Yuan, Z., Yang, Z., Luo, H., Sun, K., Zhou, L., Xiang, J., Shi, L., Yu, Q., Zhang, Y., Hou, R., & Wu, J. (2009). Production of offspring from a germline stem cell line derived from neonatal ovaries. Nature Cell Biology, 11(5), 631–636.
Johnson, J., Canning, J., Kaneko, T., Pru, J. K., & Tilly, J. L. (2004). Germline stem cells and follicular renewal in the postnatal mammalian ovary. Nature, 428(6979), 145–150.
Parte, S., Bhartiya, D., Patel, H., Daithankar, V., Chauhan, A., Zaveri, K., & Hinduja, I. (2014). Dynamics associated with spontaneous differentiation of ovarian stem cells in vitro. Journal of Ovarian Research, 7, 25. https://doi.org/10.1186/1757-2215-7-25.
White, Y. A., Woods, D. C., Takai, Y., Ishihara, O., Seki, H., & Tilly, J. L. (2012). Oocyte formation by mitotically active germ cells purified from ovaries of reproductive-age women. Nature Medicine, 18(3), 413–421. https://doi.org/10.1038/nm.2669.
Parte, S., Bhartiya, D., Telang, J., Daithankar, V., Salvi, V., Zaveri, K., & Hinduja, I. (2011). Detection, characterization, and spontaneous differentiation in vitro of very small embryonic-like putative stem cells in adult mammalian ovary. Stem Cells and Development, 20(8), 1451–1464. https://doi.org/10.1089/scd.2010.0461.
Virant-Klun, I., Rozman, P., Cvjeticanin, B., Vrtacnik-Bokal, E., Novakovic, S., Rülicke, T., Dovc, P., & Meden-Vrtovec, H. (2009). Parthenogenetic embryo-like structures in the human ovarian surface epithelium cell culture in postmenopausal women with no naturally present follicles and oocytes. Stem Cells and Development, 18, 137–149.
Virant-Klun, I., & Stimpfel, M. (2016). Novel population of small tumour-initiating stem cells in the ovaries of women with borderline ovarian cancer. Scientific Reports, 6, 34730. https://doi.org/10.1038/srep34730.
Virant-Klun, I., Kenda-Suster, N., & Smrkolj, S. (2016). Small putative NANOG, SOX2, and SSEA-4-positive stem cells resembling very small embryonic-like stem cells in sections of ovarian tissue in patients with ovarian cancer. Journal of ovarian research, 9, 12. https://doi.org/10.1186/s13048-016-0221-3.
Webb, P. M., & Jordan, S. J. (2017). Epidemiology of epithelial ovarian cancer. Best Practice & Research. Clinical Obstetrics & Gynaecology, 41, 3–14.
Kaur, T., Slavcev, R. A., & Wettig, S. D. (2009). Addressing the challenge: Current and future directions in ovarian cancer therapy. Current Gene Therapy, 9(6), 434–458.
Palmirotta, R., Silvestris, E., D'Oronzo, S., Cardascia, A., & Silvestris, F. (2017). Ovarian cancer: Novel molecular aspects for clinical assessment. Critical Reviews in Oncology/Hematology, 117, 12–29.
Liu, J. F., Konstantinopoulos, P. A., & Matulonis, U. A. (2014). PARP inhibitors in ovarian cancer: Current status and future promise. Gynecologic Oncology, 133(2), 362–369.
Kakar, S. S., Ratajczak, M. Z., Powell, K. S., Moghadamfalahi, M., Miller, D. M., Batra, S. K., & Singh, S. K. (2014). Withaferin a alone and in combination with cisplatin suppresses growth and metastasis of ovarian cancer by targeting putative cancer stem cells. PLoS One, 9(9), e107596. https://doi.org/10.1371/journal.pone.0107596.
Kakar, S. S., Parte, S., Carter, K., Joshua, I. G., Worth, C., Rameshwar, P., & Ratajczak, M. Z. (2017). Withaferin A (WFA) inhibits tumor growth and metastasis by targeting ovarian cancer stem cells. Oncotarget, 8(43), 74494–74505. https://doi.org/10.18632/oncotarget.20170.
Ginestier, C., Hur, M. H., Charafe-Jauffret, E., Monville, F., Dutcher, J., Brown, M., et al. (2007). ALDH1 is a marker of normal and malignant human mammary stem cells and a predictor of poor clinical outcome. Cell Stem Cell, 1(5), 555–567. https://doi.org/10.1016/j.stem.2007.08.014.
Kakar, S. S., Chen, L., Puri, R., Flynn, S. E., & Jennes, L. (2001). Characterization of a polyclonal antibody to human pituitary tumor transforming gene 1 (PTTG1) protein. The Journal of Histochemistry and Cytochemistry, 49(12), 1537–1546.
Acknowledgements
The authors would like to thank the Confocal Microscopy facility (University of Louisville) for allowing extensive use of the confocal microscope and Dr. V. Jala (Department of Microbiology & Immunology, University of Louisville) for his technical help during confocal microscopy.
Funding
This study was funded by a grant from NIH/NCI UO1CA2177798 (SKB, SSK) and T32HL134644 (SSK).
Author information
Authors and Affiliations
Contributions
SSK conceived and designed the work. SSK, IVK, MP, and ARS performed the experiments and data acquisition. SSK and ARS quantified data and performed statistical analyses. SP provided intellectual inputs for the study, data analysis and interpretation and prepared the first to final drafts of the manuscript with SSK. SP, IVK, and SSK wrote the manuscript and all authors (SP, ARS, IVK, MP, SKB, SSK) edited the manuscript and approved the final version.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interest.
Disclaimer Note
The authors regret if they have missed quoting any work wherever they should have, due to space constraints or oversight. The overwhelming information available in the knowledge base is truly appreciated by us as we humbly join the war against cancer with this publication.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 14 kb)
Rights and permissions
About this article
Cite this article
Parte, S., Virant-Klun, I., Patankar, M. et al. PTTG1: a Unique Regulator of Stem/Cancer Stem Cells in the Ovary and Ovarian Cancer. Stem Cell Rev and Rep 15, 866–879 (2019). https://doi.org/10.1007/s12015-019-09911-5
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12015-019-09911-5