Abstract
Many of the 25 members of the selenoprotein family function as enzymes that utilize their selenocysteine (Sec) residues to catalyze redox-based reactions. However, some selenoproteins likely do not exert enzymatic activity by themselves and selenoprotein K (SELENOK) is one such selenoprotein family member that uses its Sec residue in an alternative manner. SELENOK is an endoplasmic reticulum (ER) transmembrane protein that has been shown to be important for ER stress and for calcium-dependent signaling. Molecular mechanisms for the latter have recently been elucidated using knockout mice and genetically manipulated cell lines. These studies have shown that SELENOK interacts with an enzyme in the ER membrane, DHHC6 (letters represent the amino acids aspartic acid, histidine, histidine, and cysteine in the catalytic domain), and the SELENOK/DHHC6 complex catalyzes the transfer of acyl groups such as palmitate to cysteine residues in target proteins, i.e., palmitoylation. One protein palmitoylated by SELENOK/DHHC6 is the calcium channel protein, the inositol 1,4,5-trisphosphate receptor (IP3R), which is acylated as a means for stabilizing the tetrameric calcium channel in the ER membrane. Factors that lower SELENOK levels or function impair IP3R-driven calcium flux. This role for SELENOK is important for the activation and proliferation of immune cells, and recently, a critical role for SELENOK in promoting calcium flux for the progression of melanoma has been demonstrated. This review provides a summary of these findings and their implications in terms of designing new therapeutic interventions that target SELENOK for treating cancers like melanoma.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Selenium (Se) is an essential dietary trace mineral that is important for various aspects of human health [1]. The biological effects of Se are mainly exerted through its incorporation into selenoproteins as the 21st amino acid, selenocysteine (Sec) [2]. The 25 members of the selenoprotein family exhibit a wide variety of functions including the control of reactive oxygen species and cellular redox tone, regulating thyroid hormone metabolism, facilitating sperm maturation/protection, and promoting optimal immunity [3]. Many selenoproteins exhibit a common thioredoxin-like fold within their catalytic domains comprised of a conserved cysteine (Cys)-X-X-Sec (CXXU) or Cys-X-X-Ser [4, 5]. Many of these selenoenzymes carry out well-defined catalytic activities (e.g., glutathione peroxidases, thioredoxin reductases, iodothyronine deiodinase, methionine sulfoxide reductases), while other CXXU-containing selenoproteins have been proposed to serve as oxidoreductase enzymes [2]. Selenoprotein K (SELENOK) does not contain a CXXU motif nor does it appear to catalyze redox reactions like other established selenoenzymes. In fact, the intrinsically disordered nature of SELENOK [6] suggests its function is dependent on the partner proteins with which it complexes. The role of intrinsically disordered domains in proteins serving as docking platforms for signaling molecules or binding proteins has been established for other nonenzymatic proteins [7]. Indeed, much of what has been learned about SELENOK function has been gained by examining its binding partners [8,9,10]. This review will summarize what has been discovered regarding SELENOK function and possible therapeutic approaches that may center on this selenoprotein.
Tissue Distribution, Localization, Se Dependence of SELENOK
Dietary [Se] regulates the expression of selenoproteins at the levels of mRNA and protein. Under conditions of low Se status, the translation of selenoproteins stalls at the Sec-encoding UGA codon and both the mRNA and truncated protein get degraded through nonsense-mediated decay and destruction via C-end degrons, respectively [11, 12]. Moreover, dietary [Se] impacts expression levels of different selenoproteins to different degrees [13]. This concept is part of what is referred to as “the hierarchy of selenoproteins,” and the ranking of least to most sensitive Se-sensitive selenoproteins in this hierarchy depends on the tissue involved and varies between cell lines [13, 14]. Differences between selenoproteins in sensitivity to limiting [Se] were first demonstrated by experiments comparing the activities of glutathione peroxidases 1 and 4 (GPX1 and GPX4) in rat liver, kidney, and heart [15]. Under conditions of Se deficiency, GPX1 activity fell sharply while GPX4 activity was better maintained, and tissues showed varied differences between these two GPXs. It must be noted that, in addition to the tissue involved, sex also influences the selenoprotein hierarchy [16]. Our data in mice suggests that SELENOK levels are similar between male and females [17]. Also, there is high sequence homology between mouse and human SELENOK at the protein level. SELENOK amino acid sequences from human and mouse share 91% identity [18], which supports functional investigations using mouse models in combination with human cell lines.
While many selenoproteins exhibit some degree of tissue specificity, SELENOK mRNA and protein are widely expressed throughout mouse tissues, although particularly high levels have been found in lymphoid tissues [17, 19]. These results are consistent with findings from the Human Protein Atlas, where SELENOK was shown to be widely distributed but highest in the gastrointestinal tract that contains many lymph nodes, the immune system, and lung that is rich in mucosal lymphoid tissues [20]. Northern blot data from one report suggested high levels of expression in the heart [21], but this has not been supported by subsequent studies. SELENOK has been shown to exhibit altered levels of expression in response to changes in Se status in humans and mice [17, 22], but it is not known exactly where SELENOK lies within the selenoprotein hierarchy. In some tissues like mouse colon and spleen, SELENOK was not found to be Se sensitive [23, 24]. One study using human lymphocytes isolated from human donors before and after Se supplementation showed that SELENOK mRNA was among three selenoprotein transcripts upregulated after supplementation [25]. SELENOK has not been found to be upregulated in cancers like melanoma, despite an apparent dependence on SELENOK function for growth and metastasis [26]. The sensitivity of SELENOK levels to bioavailable Se very likely depends on tissue and cell type, with tissues such as the brain, muscle, and testes receiving “priority” for bioavailable Se to express SELENOK under Se deficiency at a cost to other tissues such as those comprising the immune system [13]. The “bioactivity” of SELENOK has not yet been compared between tissues or under different [Se] conditions. Overexpression of GFP-tagged SELENOK in HeLa cells showed localization in the endoplasmic reticulum (ER) [21]. Subsequently, immunofluorescent staining of primary human monocytes confirmed colocalization of SELENOK with the ER marker KDEL and cell fractionation data also showed ER localization [17].
The Role of SELENOK in Regulating ER Stress and Calcium Flux
The ER is a crucial organelle in which protein folding takes place as well as maturation of secreted and membrane proteins. In many cases, the correct folding and processing of proteins in the ER may not effectively occur and the errant proteins must undergo ER-associated degradation (ERAD) so that ER stress does not build up [27]. ER homeostasis is tightly regulated and may involve one or more of the ER-resident selenoproteins: type 2 iodothyronine deiodinase and selenoproteins F, K, M, N, S, and T [28]. SELENOK has been shown to participate in ERAD by binding in an ERAD complex with other proteins such as Derlin-1 [10]. Importantly, the gene encoding SELENOK contains a functional ER stress response element and SELENOK was found to be upregulated by treatment of cells that cause misfolded protein accumulation in the ER [5]. Along with another selenoprotein, SELENOS, SELENOK participates in ERAD by recruiting cytosolic valosin-containing protein (VCP/p97) to promote translocation of misfolded proteins from the ER lumen [8]. It remains unclear how the Sec residue may or may not participate in retrograde translocation of misfolded proteins, and in SELENOK-knockout mice, there are no apparent signs of ER stress [17, 29]. The latter finding suggests some redundancy or compensation, perhaps involving SELENOS, may occur in the absence of SELENOK expression to mitigate ER stress.
A major finding with the SELENOK-knockout mice was that a deficiency in this selenoprotein led to an impaired flux of calcium from the ER in activated immune cells. In particular, ex vivo experiments in our laboratory suggested that SELENOK was crucial for promoting store-operated calcium entry (SOCE) in T cells, neutrophils, and macrophages stimulated with chemokines and other reagents that trigger G-coupled protein receptor signaling in these immune cells [17]. The impaired SOCE in SELENOK-knockout immune cells resulted in 50% decreased levels of activation as determined by downstream activation markers, and lowered immunity in knockout mice led to higher levels of death during West Nile virus infection compared with controls [17, 29]. The molecular mechanism by which SELENOK regulates SOCE was not clear in these initial studies, but subsequent investigations revealed a specific role for SELENOK in the post-translational modification (i.e., palmitoylation) of the calcium channel protein, the inositol 1,4,5-trisphosphate receptor (IP3R) [30]. SELENOK was found to be crucial for stable expression of IP3R in the ER membrane and thereby a critical regulator of SOCE through this calcium channel protein. The details of these findings are described in more detail below.
The Role of SELENOK in Promoting Protein Palmitoylation
Our investigation into the role that SELENOK plays in the immune system during atherosclerosis revealed that SELENOK was required for lipid raft localization of the oxidized low-density lipoprotein receptor, CD36, on macrophages [31]. Using mouse knockout models and bone-marrow transplants, SELENOK-deficient macrophages were found to have decreased CD36 palmitoylation, which occurs at the ER membrane and is crucial for stabilizing CD36 expression and its clustered expression in the plasma membrane [32]. S-palmitoylation of cysteine residues is a post-translational modification occurring on integral and peripheral membrane proteins that contribute to membrane association [33]. Our laboratory subsequently found that palmitoylation of other proteins required SELENOK including Arf-GAP with Src homology 3 (SH3) domain, ankyrin repeat, and plekstrin homology domain-containing protein 2 (ASAP2) and the IP3R [9, 34]. There are very likely other proteins that required SELENOK for this post-translational modification. Protein palmitoylation involves the addition of the 16-carbon fatty acid, palmitate, to cysteine residues on target substrate proteins through a thioester bond [35]. Since SELENOK itself is not an enzyme, it was apparent to us that it must bind to a partner enzyme to promote palmitoylation. A major clue to this puzzle was the fact that SELENOK is the only selenoprotein with an SH3 binding domain and that one member of the enzyme family involved in palmitoylation reactions contained an SH3 domain. Indeed, SELENOK interacts with the acyltransferase, DHHC6 (letters represent the amino acids aspartic acid, histidine, histidine, and cysteine in the catalytic domain), in the ER membrane through SH3/SH3 binding domain interactions and this is how SELENOK participates in palmitoylation reactions [9].
The specific mechanism by which SELENOK exerts its effects on DHHC6 to increase its catalytic efficiency required new tools to be built so that the enzymatic reaction of acyl transfer could be examined in more detail. The palmitoylation reactions catalyzed by DHHC enzymes have been shown to proceed via a two-step ping-pong mechanism [36]. In the first autopalmitoylation step, a Cys residue within the DHHC catalytic domain reacts with palmitoyl-CoA to form a palmitoyl-PAT intermediate. In the second step, the palmitoyl group is transferred from the PAT to a Cys residue on the target protein. The first step is rapid, whereas the second is much slower. Thus, after the first step, the unstable thioester bond between Cys on the PAT and the palmitic acid may hydrolyze before the transfer of the palmitic acid to a target protein, termed the “futile cycle” [37]. Our laboratory was able to synthesize soluble versions of DHHC6 and has shown that they bind recombinant SELENOK in a complex that exhibited acyltransferase activity. The data from these experiments showed that SELENOK increases the catalytic efficiency of palmitoylation by stabilizing the acyl-DHHC6 intermediate and protecting it from hydrolysis, thereby preventing the futile cycle [38]. This binding of a PAT enzyme with a cofactor to promote intermediate stabilization was previously found in another protein pair in yeast. In particular, the yeast ethylene-responsive factor-2 (ERF2) is an ortholog for mammalian DHHC9 and binds to yeast ERF4 in a manner that stabilizes the acyl-ERF2 intermediate and increases its catalytic efficiency [39]. ERF4 is not a selenoprotein (yeasts do not express selenoproteins), so the manner in which the two cofactors increase catalytic efficiency must be different. As discussed below in more detail, the Sec residue in SELENOK plays a critical role in promoting palmitoylation.
The Sec Residue in SELENOK Is Responsible for Its Bioactivity
SELENOK is a relatively small protein (94 amino acids) that is localized to the ER membrane (Fig. 1). This single spanning transmembrane has a short ER luminal domain and a longer cytosolic domain containing the Sec residue. The topology of this transmembrane protein was first described for Drosophila SELENOK (G-rich protein) showing the N-terminus in the lumen with no N-terminal signal peptide [40]. The Sec residue is located near the C-terminus of SELENOK (amino acid 92), which is a feature found in approximately one-third of the selenoproteins [41]. While the Sec residue has been shown to exhibit some antioxidant activity in vitro [21, 42], a role for regulating redox reactions in vivo has not been established. As described above, we generated SELENOK-knockout mice and found impaired SOCE in T cells, neutrophils, and macrophages stimulated with chemokines and other reagents that trigger G-coupled protein receptor signaling in these immune cells [17]. Importantly, when these immune cells were stimulated with thapsigargin, a drug that induces SOCE in a manner that bypasses the IP3R, the wild-type and knockout cells did not differ. This implicated the IP3R as a molecule somehow involved in SELENOK-dependent SOCE.
The exact mechanism by which SELENOK regulated SOCE was not initially clear, although the requirement of the Sec residue in this process became evident when we subsequently found that a calpain protease (calpain-2) that cleaved the C-terminal end of SELENOK (residues 81–94, including Sec92) away from the remaining protein abolished the SOCE-promoting activity of SELENOK [43]. Interestingly, resting macrophages actively cleave SELENOK via calpain-2 to maintain low levels of calcium flux and upon activation, the endogenous inhibitor of calpains, calpastatin, is upregulated to increase levels of full-length SELENOK and endow the cells with full potential of calcium flux via IP3R. We subsequently used CRISPR/Cas9 to generate cell lines lacking the 81–94 amino acid portion to confirm that this region is the functional domain of SELENOK [26]. Cells lacking the functional domain of SELENOK were defective in generating calcium flux through the IP3R in a manner similar to those with active calpain-2 cleavage of SELENOK. In this sense, dietary Se and the calpain-2/calpastatin system are both regulators of SELENOK function. In fact, the endogenous inhibitor of calpain-2, calpastatin, protects SELENOK from cleavage and thereby serves as a positive regulator of SELENOK function [43, 44]. Dietary Se levels regulate the levels of Sec incorporation into SELENOK while calpain-2 regulates the cleavage of the functional domain. To date, no other selenoproteins have been described as being proteolytically modulated by calpain enzymes, although investigations into this issue have not been extensive.
The definitive experiments showing the requirement of the Sec residue at position 92 of SELENOK for its function were recently published as described above. In particular, recombinant versions of SELENOK containing either Sec92, Cys92, or Ala92 complexed to a soluble version of DHHC6 were used in PAT reactions to test the bioactivity of each. Our results showed that only Sec92 improved the catalytic efficiency of DHHC6, while Cys92 or Ala92 were equally insufficient to allow SELENOK to increase the catalytic efficiency of DHHC6. Selenoester bonds have been shown to have high reactivity in acyl transfer reactions [45]. This is consistent with the Sec residue in SELENOK being particularly suited to combine with DHHC6 for catalyzing protein palmitoylation as an important post-translational modification, and the fact that our studies showed that Sec cannot be replaced with Cys for protecting the thioester bond in the acyl-DHHC6 intermediate from hydrolysis. The role SELENOK plays in palmitoylation reactions and how it is regulated are illustrated in Fig. 2.
SELENOK in Cancer
The exact role of SELENOK in carcinogenesis or tumor progression has not been well studied. The effects of SELENOK loss-of-function or gain-of-function in different tumor cell lines have been studied to a limited extent. For example, reducing SELENOK in human choriocarcinoma cells increased the proliferative, migratory, and invasive capacity of these cells while over-expressing SELENOK had the opposing effect [46]. Similarly, overexpression of SELENOK in human gastric cancer BGC-823 cells inhibited cell adhesion and migration [47]. Studies in other cancer cells have shown that reduced SELENOK levels reduced proliferation and migration [26]. Micro RNAs may also play a role in regulating SELENOK levels in certain cancers [48], and this may depend on the type of cancer or may even vary between individual cancer cells within tumors. Given that SELENOK complexes with other protein partners to exert its functions, manipulating the levels of SELENOK may perturb the stoichiometry of the members within complexes. Thus, it may be that the increases or decreases in SELENOK that alter the optimum level within complexes can disrupt functions required by different cancer cells at different stages, or different cell lines immortalized at different stages. Investigation into the mutational status of SELENOK in various cancers using The Cancer Genome Atlas Program (TCGA) revealed 37 somatic mutations in SELENOK from 39 cancer-associated cases [49]. Most mutations occurred within either the 5′ or 3′ UTR. However, several mutations were observed in the protein-coding region of SELENOK, resulting in 1 frameshift mutation, 9 missense mutations, and 1 mutation that occurred within a splice donor site of SELENOK. Additionally, 853 individual cases were affected by 819 copy number variation events in SELENOK whereby most resulted in loss of function. On the other hand, patients involved in a study of germ cell neoplasms demonstrated mostly an increase in SELENOK copy number.
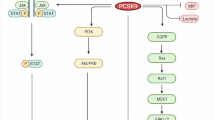
When different cell types transform into malignant tumor cells, calcium homeostasis and calcium-dependent signaling may be altered in a process referred to as calcium remodeling [50]. SOCE and the downstream calcium-dependent signaling cascades are exploited by cancer cells to increase their capacity to proliferate, invade, and migrate. Because SOCE is important for both primary tumor progression and metastasis to other tissues, molecules regulating SOCE may serve as important targets for developing anti-cancer therapeutics. In fact, cancer cells may develop a stringent dependence on highly efficient SOCE along with the signaling pathways and gene expression programs shaped by calcium remodeling [51, 52]. The importance of calcium flux in the regulation of melanoma proliferation and cell migration was previously suggested by the knockdown of stromal-interacting molecule 1 (STIM1) or Orai1 in human melanoma cell lines as well the use of a pharmacological inhibitor of SOCE [53]. Similar effects were shown for breast tumor cells and glioblastoma cells [54, 55]. Our findings using cell-based assays and animal models of melanoma indicated that SELENOK was required for progression of melanoma [26]. In particular, our in vitro and in vivo findings suggest that melanoma requires SELENOK expression for the IP3R-dependent maintenance of stemness, tumor growth, and metastasic potential. This suggests that SELENOK may represent a new potential therapeutic target for treating melanoma and possibly other cancers.
Designing and Testing Drugs That Inhibit SELENOK Function
Although published data suggest that targeting SELENOK with inhibitors may be an effective approach to treating cancers that rely on SOCE for disease progression, this approach presents several challenges. Our unpublished work has established that anti-SELENOK antibodies targeting the C-terminal portion of SELENOK can interfere with the palmitoylation reaction catalyzed by the SELENOK/DHHC6 complex. Thus, antibody-based inhibitors (i.e., immunotherapeutics) may represent an effective form of therapy to treat melanoma or other cancers. However, SELENOK is an intracellular protein and this renders standard immunotherapeutic approaches that rely on antibody-based treatment unrealistic since targets for these drugs are conventionally extra-cellular proteins. Reassessment of intracellular proteins as immunotherapeutic targets like SELENOK may need to be prioritized since intracellular molecules constitute nearly half of the human proteome and represent an untapped reservoir of potential therapeutic targets [56]. Even though antibodies may exist that recognize an intracellular protein of interest discovered through mechanistic studies, the promising new field of immunotherapy is not a viable option due to the hurdles of delivering large antibodies (Ab ~ 150 kDa) or other antibody fragments (Fab ~ 50 kDa, scFv ~ 25 kDa) into cells to inhibit these proteins. In addition, the multiple chains comprising Ab reagents also make recombinant production of biologics very challenging.
One possible way to overcome these barriers involves a novel strategy that combines two emerging technologies: single domain antibodies (sdAbs, a.k.a. nanobodies) from camelids [57, 58] and cell penetrating peptides (CPPs) [59,60,61]. In theory, these small versions of Abs exhibit the high specificity of conventional Abs required for effective therapeutics. In fact, studies have demonstrated the use of humanized sdAb fused to CPP to effectively inhibit their intracellular targets. For example, CPP-sdAb directed against different hepatitis C virus proteins inhibited viral replication within cells [62,63,64]. Our laboratory is currently exploring this approach for the development of intracellular immunotherapeutics to inhibit SELENOK, and this approach may yield a novel set of inhibitory reagents.
Synthesis of small molecules that bind to vital regions of target proteins is a more conventional approach for developing inhibitors of intracellular proteins like SELENOK. However, this approach also presents challenges, such as a need for insight on the structure of the target protein. Gaining structural information on SELENOK for designing small molecules to inhibit its function is limited by the fact that the exposed, cytosolic portion of SELENOK is intrinsically disordered [42]. Intrinsically disordered proteins have long been challenging for the field of inhibitor development, but some new technologies are emerging to overcome the lack of structural detail. Small molecules that bind to intrinsically disordered proteins have been successfully identified through screens utilizing surface plasmon resonance, nuclear magnetic resonance (NMR) spectroscopy, and fluorescence methods [65]. Identifying small molecule inhibitors specific for blocking SELENOK-dependent palmitoylation would provide a very useful research tool and, perhaps, a lead compound for therapy development in the context of cancers such as melanoma. In addition, techniques have been developed for the synthesis of recombinant SELENOK [66], and our laboratory has shown that this recombinant SELENOK bound to soluble DHHC6 is functional in cell-free palmitoylation enzyme assays [38]. Thus, screens that successfully identify small molecules with high-affinity binding to SELENOK would be able to use downstream in vitro assays to confirm inhibitory activity.
A third possible approach for targeting SELENOK for functional inhibition could take advantage of the SH3 binding domain in its cytosolic region. Our previous work demonstrated that the SH3 binding domain in SELENOK binds to the SH3 domain of DHHC6 [9, 38], and this interaction is required for the function of the SELENOK/DHHC6 complex. SH3/SH3 binding domains are an abundant mechanism used within cells for promoting protein-protein interactions [67]. Although a multitude of SH3/SH3 binding domain interactions have been identified, there may be ways to include “context” or to modify residues adjacent to the motifs to provide specificity to inhibitor design [68, 69]. The most successful efforts to increase SH3 ligand specificity have focused primarily on replacing sequences flanking the PxxP motif with natural or non-natural moieties [70, 71]. Since the SH3 binding domain is well defined in SELENOK, this approach may prove successful for peptide inhibitors designed to disrupt the interaction between two binding domains of SELENOK and DHHC6. This is another area of research that we have focused our efforts, with the goal of developing the smallest peptide or peptide mimetic that can specifically and effectively interrupt interaction between SELENOK and DHHC6.
Concluding Remarks
Much progress has been made in understanding the role that SELENOK plays in the immune system and in cancer cells. Insight into the molecular mechanisms by which this selenoprotein regulates calcium-dependent signaling was provided with the discovery that SELENOK interacts with the DHHC6 enzyme to promote protein palmitoylation. A major target of SELENOK/DHHC6 palmitoylation is the calcium channel protein, the IP3R. SELENOK deficiency results in impaired calcium flux in immune cells that leads to ~ 50% maximal activation of immune cells including T cells, macrophages, and neutrophils. However, SELENOK deficiency appears to have a more dramatic effect on cancer cells that rely on calcium-dependent signaling to grow and metastasize. Thus, therapeutics that inhibit SELENOK will likely impair immunity but this may not be as dramatic as the static effect they may exert on tumors. Furthermore, combining cancer-static SELENOK inhibitors with cancer-toxic chemotherapeutics may prove to be an effective strategy for treating cancers that are particularly dependent on calcium signaling. It is important to note that the anti-cancer effects of SELENOK have thus far only been investigated in a mouse melanoma model and a human melanoma cell line. Much more needs to be determined in terms of the broader range of cancer types for which SELENOK inhibitors may be effective. Also, effects that inhibiting SELENOK have on ER stress or other physiological processes need to be considered. The potential of SELENOK as a therapeutic target is only beginning to be studied and challenges remain regarding its “drugability,” but basic science investigations have been revealing and SELENOK is now much less of an enigmatic member of the selenoprotein family.
References
Rayman MP (2012) Selenium and human health. Lancet 379(9822):1256–1268. https://doi.org/10.1016/S0140-6736(11)61452-9
Reeves MA, Hoffmann PR (2009) The human selenoproteome: recent insights into functions and regulation. Cell Mol Life Sci 66(15):2457–2478. https://doi.org/10.1007/s00018-009-0032-4
Schweizer U, Fradejas-Villar N (2016) Why 21? The significance of selenoproteins for human health revealed by inborn errors of metabolism. FASEB J 30(11):3669–3681. https://doi.org/10.1096/fj.201600424
Qi Y, Grishin NV (2005) Structural classification of thioredoxin-like fold proteins. Proteins 58(2):376–388. https://doi.org/10.1002/prot.20329
Du S, Zhou J, Jia Y, Huang K (2010) SelK is a novel ER stress-regulated protein and protects HepG2 cells from ER stress agent-induced apoptosis. Arch Biochem Biophys 502(2):137–143. https://doi.org/10.1016/j.abb.2010.08.001
Polo A, Guariniello S, Colonna G, Ciliberto G, Costantini S (2016) A study on the structural features of SELK, an over-expressed protein in hepatocellular carcinoma, by molecular dynamics simulations in a lipid-water system. Mol BioSyst 12(10):3209–3222. https://doi.org/10.1039/c6mb00469e
Simister PC, Feller SM (2012) Order and disorder in large multi-site docking proteins of the Gab family--implications for signalling complex formation and inhibitor design strategies. Mol BioSyst 8(1):33–46. https://doi.org/10.1039/c1mb05272a
Lee JH, Park KJ, Jang JK, Jeon YH, Ko KY, Kwon JH, Lee SR, Kim IY (2015) Selenoprotein S-dependent Selenoprotein K binding to p97(VCP) protein is essential for endoplasmic reticulum-associated degradation. J Biol Chem 290(50):29941–29952. https://doi.org/10.1074/jbc.M115.680215
Fredericks GJ, Hoffmann PR (2015) Selenoprotein K and protein palmitoylation. Antioxid Redox Signal 23(10):854–862. https://doi.org/10.1089/ars.2015.6375
Shchedrina VA, Everley RA, Zhang Y, Gygi SP, Hatfield DL, Gladyshev VN (2011) Selenoprotein K binds multiprotein complexes and is involved in the regulation of endoplasmic reticulum homeostasis. J Biol Chem 286(50):42937–42948. https://doi.org/10.1074/jbc.M111.310920
Seyedali A, Berry MJ (2014) Nonsense-mediated decay factors are involved in the regulation of selenoprotein mRNA levels during selenium deficiency. RNA 20(8):1248–1256. https://doi.org/10.1261/rna.043463.113
Lin HC, Yeh CW, Chen YF, Lee TT, Hsieh PY, Rusnac DV, Lin SY, Elledge SJ, Zheng N, Yen HS (2018) C-terminal end-directed protein elimination by CRL2 ubiquitin ligases. Mol Cell 70(4):602–613 e603. https://doi.org/10.1016/j.molcel.2018.04.006
Burk RF, Hill KE (2015) Regulation of selenium metabolism and transport. Annu Rev Nutr 35:109–134. https://doi.org/10.1146/annurev-nutr-071714-034250
Touat-Hamici Z, Bulteau AL, Bianga J, Jean-Jacques H, Szpunar J, Lobinski R, Chavatte L (2018) Selenium-regulated hierarchy of human selenoproteome in cancerous and immortalized cells lines. Biochim Biophys Acta Gen Subj 1862:2493–2505. https://doi.org/10.1016/j.bbagen.2018.04.012
Lei XG, Evenson JK, Thompson KM, Sunde RA (1995) Glutathione peroxidase and phospholipid hydroperoxide glutathione peroxidase are differentially regulated in rats by dietary selenium. J Nutr 125(6):1438–1446. https://doi.org/10.1093/jn/125.6.1438
Cao L, Zhang L, Zeng H, Wu RT, Wu TL, Cheng WH (2017) Analyses of Selenotranscriptomes and selenium concentrations in response to dietary selenium deficiency and age reveal common and distinct patterns by tissue and sex in telomere-dysfunctional mice. J Nutr 147(10):1858–1866. https://doi.org/10.3945/jn.117.247775
Verma S, Hoffmann FW, Kumar M, Huang Z, Roe K, Nguyen-Wu E, Hashimoto AS, Hoffmann PR (2011) Selenoprotein K knockout mice exhibit deficient calcium flux in immune cells and impaired immune responses. J Immunol 186(4):2127–2137. https://doi.org/10.4049/jimmunol.1002878
Hoffmann PR (2012) An emerging picture of the biological roles of selenoprotein K. In: Hatfield DL, Berry MJ, Gladyshev VN (eds) Selenium: its molecular biology and role in human health, 3rd edn. Springer, New York, pp 335–344
Hoffmann PR, Hoge SC, Li PA, Hoffmann FW, Hashimoto AC, Berry MJ (2007) The selenoproteome exhibits widely varying, tissue-specific dependence on selenoprotein P for selenium supply. Nucleic Acids Res 35(12):3963–3973. https://doi.org/10.1093/nar/gkm355
Uhlen M, Fagerberg L, Hallstrom BM, Lindskog C, Oksvold P, Mardinoglu A, Sivertsson A, Kampf C, Sjostedt E, Asplund A, Olsson I, Edlund K, Lundberg E, Navani S, Szigyarto CA, Odeberg J, Djureinovic D, Takanen JO, Hober S, Alm T, Edqvist PH, Berling H, Tegel H, Mulder J, Rockberg J, Nilsson P, Schwenk JM, Hamsten M, von Feilitzen K, Forsberg M, Persson L, Johansson F, Zwahlen M, von Heijne G, Nielsen J, Ponten F (2015) Proteomics. Tissue-based map of the human proteome. Science 347(6220):1260419. https://doi.org/10.1126/science.1260419
Lu C, Qiu F, Zhou H, Peng Y, Hao W, Xu J, Yuan J, Wang S, Qiang B, Xu C, Peng X (2006) Identification and characterization of selenoprotein K: an antioxidant in cardiomyocytes. FEBS Lett 580(22):5189–5197. https://doi.org/10.1016/j.febslet.2006.08.065
Meplan C, Johnson IT, Polley AC, Cockell S, Bradburn DM, Commane DM, Arasaradnam RP, Mulholland F, Zupanic A, Mathers JC, Hesketh J (2016) Transcriptomics and proteomics show that selenium affects inflammation, cytoskeleton, and cancer pathways in human rectal biopsies. FASEB J 30(8):2812–2825. https://doi.org/10.1096/fj.201600251R
Kipp A, Banning A, van Schothorst EM, Meplan C, Schomburg L, Evelo C, Coort S, Gaj S, Keijer J, Hesketh J, Brigelius-Flohe R (2009) Four selenoproteins, protein biosynthesis, and Wnt signalling are particularly sensitive to limited selenium intake in mouse colon. Mol Nutr Food Res 53(12):1561–1572. https://doi.org/10.1002/mnfr.200900105
Kipp AP, Banning A, van Schothorst EM, Meplan C, Coort SL, Evelo CT, Keijer J, Hesketh J, Brigelius-Flohe R (2012) Marginal selenium deficiency down-regulates inflammation-related genes in splenic leukocytes of the mouse. J Nutr Biochem 23(9):1170–1177. https://doi.org/10.1016/j.jnutbio.2011.06.011
Pagmantidis V, Meplan C, van Schothorst EM, Keijer J, Hesketh JE (2008) Supplementation of healthy volunteers with nutritionally relevant amounts of selenium increases the expression of lymphocyte protein biosynthesis genes. Am J Clin Nutr 87(1):181–189. https://doi.org/10.1093/ajcn/87.1.181
Marciel MP, Khadka VS, Deng Y, Kilicaslan P, Pham A, Bertino P, Lee K, Chen S, Glibetic N, Hoffmann FW, Matter ML, Hoffmann PR (2018) Selenoprotein K deficiency inhibits melanoma by reducing calcium flux required for tumor growth and metastasis. Oncotarget 9(17):13407–13422. https://doi.org/10.18632/oncotarget.24388
Bagola K, Mehnert M, Jarosch E, Sommer T (2011) Protein dislocation from the ER. Biochim Biophys Acta 1808(3):925–936. https://doi.org/10.1016/j.bbamem.2010.06.025
Addinsall AB, Wright CR, Andrikopoulos S, van der Poel C, Stupka N (2018) Emerging roles of endoplasmic reticulum-resident selenoproteins in the regulation of cellular stress responses and the implications for metabolic disease. Biochem J 475(6):1037–1057. https://doi.org/10.1042/BCJ20170920
Huang Z, Hoffmann FW, Fay JD, Hashimoto AC, Chapagain ML, Kaufusi PH, Hoffmann PR (2012) Stimulation of unprimed macrophages with immune complexes triggers a low output of nitric oxide by calcium-dependent neuronal nitric-oxide synthase. J Biol Chem 287(7):4492–4502. https://doi.org/10.1074/jbc.M111.315598
Fredericks GJ, Hoffmann FW, Rose AH, Osterheld HJ, Hess FM, Mercier F, Hoffmann PR (2014) Stable expression and function of the inositol 1,4,5-triphosphate receptor requires palmitoylation by a DHHC6/selenoprotein K complex. Proc Natl Acad Sci U S A 111(46):16478–16483. https://doi.org/10.1073/pnas.1417176111
Meiler S, Baumer Y, Huang Z, Hoffmann FW, Fredericks GJ, Rose AH, Norton RL, Hoffmann PR, Boisvert WA (2013) Selenoprotein K is required for palmitoylation of CD36 in macrophages: implications in foam cell formation and atherogenesis. J Leukoc Biol 93(5):771–780. https://doi.org/10.1189/jlb.1212647
Thorne RF, Ralston KJ, de Bock CE, Mhaidat NM, Zhang XD, Boyd AW, Burns GF (2010) Palmitoylation of CD36/FAT regulates the rate of its post-transcriptional processing in the endoplasmic reticulum. Biochim Biophys Acta 1803(11):1298–1307. https://doi.org/10.1016/j.bbamcr.2010.07.002
Rocks O, Gerauer M, Vartak N, Koch S, Huang ZP, Pechlivanis M, Kuhlmann J, Brunsveld L, Chandra A, Ellinger B, Waldmann H, Bastiaens PI (2010) The palmitoylation machinery is a spatially organizing system for peripheral membrane proteins. Cell 141(3):458–471. https://doi.org/10.1016/j.cell.2010.04.007
Norton RL, Fredericks GJ, Huang Z, Fay JD, Hoffmann FW, Hoffmann PR (2017) Selenoprotein K regulation of palmitoylation and calpain cleavage of ASAP2 is required for efficient FcgammaR-mediated phagocytosis. J Leukoc Biol 101(2):439–448. https://doi.org/10.1189/jlb.2A0316-156RR
Bijlmakers MJ, Marsh M (2003) The on-off story of protein palmitoylation. Trends Cell Biol 13(1):32–42
Jennings BC, Linder ME (2012) DHHC protein S-acyltransferases use similar ping-pong kinetic mechanisms but display different acyl-CoA specificities. J Biol Chem 287(10):7236–7245. https://doi.org/10.1074/jbc.M111.337246
Mitchell DA, Mitchell G, Ling Y, Budde C, Deschenes RJ (2010) Mutational analysis of Saccharomyces cerevisiae Erf2 reveals a two-step reaction mechanism for protein palmitoylation by DHHC enzymes. J Biol Chem 285(49):38104–38114. https://doi.org/10.1074/jbc.M110.169102
Fredericks GJ, Hoffmann FW, Hondal RJ, Rozovsky S, Urschitz J, Hoffmann PR (2017) Selenoprotein K increases efficiency of DHHC6 catalyzed protein palmitoylation by stabilizing the acyl-DHHC6 intermediate. Antioxidants (Basel) 7(1). https://doi.org/10.3390/antiox7010004
Mitchell DA, Hamel LD, Ishizuka K, Mitchell G, Schaefer LM, Deschenes RJ (2012) The Erf4 subunit of the yeast Ras palmitoyl acyltransferase is required for stability of the Acyl-Erf2 intermediate and palmitoyl transfer to a Ras2 substrate. J Biol Chem 287(41):34337–34348. https://doi.org/10.1074/jbc.M112.379297
Chen CL, Shim MS, Chung J, Yoo HS, Ha JM, Kim JY, Choi J, Zang SL, Hou X, Carlson BA, Hatfield DL, Lee BJ (2006) G-rich, a Drosophila selenoprotein, is a Golgi-resident type III membrane protein. Biochem Biophys Res Commun 348(4):1296–1301. https://doi.org/10.1016/j.bbrc.2006.07.203
Kryukov GV, Castellano S, Novoselov SV, Lobanov AV, Zehtab O, Guigo R, Gladyshev VN (2003) Characterization of mammalian selenoproteomes. Science 300(5624):1439–1443. https://doi.org/10.1126/science.1083516
Liu J, Zhang Z, Rozovsky S (2014) Selenoprotein K form an intermolecular diselenide bond with unusually high redox potential. FEBS Lett 588(18):3311–3321. https://doi.org/10.1016/j.febslet.2014.07.037
Huang Z, Hoffmann FW, Norton RL, Hashimoto AC, Hoffmann PR (2011) Selenoprotein K is a novel target of m-calpain, and cleavage is regulated by Toll-like receptor-induced calpastatin in macrophages. J Biol Chem 286(40):34830–34838. https://doi.org/10.1074/jbc.M111.265520
Huang Z, Rose AH, Hoffmann FW, Hashimoto AS, Bertino P, Denk T, Takano J, Iwata N, Saido TC, Hoffmann PR (2013) Calpastatin prevents NF-kappaB-mediated hyperactivation of macrophages and attenuates colitis. J Immunol 191(7):3778–3788. https://doi.org/10.4049/jimmunol.1300972
Reich HJ, Hondal RJ (2016) Why nature chose selenium. ACS Chem Biol 11(4):821–841. https://doi.org/10.1021/acschembio.6b00031
Li M, Cheng W, Nie T, Lai H, Hu X, Luo J, Li F, Li H (2018) Selenoprotein K mediates the proliferation, migration, and invasion of human choriocarcinoma cells by negatively regulating human chorionic gonadotropin expression via ERK, p38 MAPK, and Akt signaling pathway. Biol Trace Elem Res 184(1):47–59. https://doi.org/10.1007/s12011-017-1155-3
Ben SB, Peng B, Wang GC, Li C, Gu HF, Jiang H, Meng XL, Lee BJ, Chen CL (2015) Overexpression of selenoprotein SelK in BGC-823 cells inhibits cell adhesion and migration. Biochemistry (Mosc) 80(10):1344–1353. https://doi.org/10.1134/S0006297915100168
Potenza N, Castiello F, Panella M, Colonna G, Ciliberto G, Russo A, Costantini S (2016) Human MiR-544a modulates SELK expression in hepatocarcinoma cell lines. PLoS One 11(6):e0156908. https://doi.org/10.1371/journal.pone.0156908
The Cancer Genome Atlas Program, National Cancer Institute, National Institutes of Health. https://www.cancer.gov/about-nci/organization/ccg/research/structural-genomics/tcga. Accessed 31 May 2019
Villalobos C, Sobradillo D, Hernandez-Morales M, Nunez L (2016) Remodeling of calcium entry pathways in cancer. Adv Exp Med Biol 898:449–466. https://doi.org/10.1007/978-3-319-26974-0_19
Cui C, Merritt R, Fu L, Pan Z (2017) Targeting calcium signaling in cancer therapy. Acta Pharm Sin B 7(1):3–17. https://doi.org/10.1016/j.apsb.2016.11.001
Stanisz H, Vultur A, Herlyn M, Roesch A, Bogeski I (2016) The role of Orai-STIM calcium channels in melanocytes and melanoma. J Physiol 594(11):2825–2835. https://doi.org/10.1113/JP271141
Umemura M, Baljinnyam E, Feske S, De Lorenzo MS, Xie LH, Feng X, Oda K, Makino A, Fujita T, Yokoyama U, Iwatsubo M, Chen S, Goydos JS, Ishikawa Y, Iwatsubo K (2014) Store-operated Ca2+ entry (SOCE) regulates melanoma proliferation and cell migration. PLoS One 9(2):e89292. https://doi.org/10.1371/journal.pone.0089292
Yang S, Zhang JJ, Huang XY (2009) Orai1 and STIM1 are critical for breast tumor cell migration and metastasis. Cancer Cell 15(2):124–134. https://doi.org/10.1016/j.ccr.2008.12.019
Motiani RK, Hyzinski-Garcia MC, Zhang X, Henkel MM, Abdullaev IF, Kuo YH, Matrougui K, Mongin AA, Trebak M (2013) STIM1 and Orai1 mediate CRAC channel activity and are essential for human glioblastoma invasion. Pflugers Arch 465(9):1249–1260. https://doi.org/10.1007/s00424-013-1254-8
Trenevska I, Li D, Banham AH (2017) Therapeutic antibodies against intracellular tumor antigens. Front Immunol 8:1001. https://doi.org/10.3389/fimmu.2017.01001
Gonzalez-Sapienza G, Rossotti MA, Tabares-da Rosa S (2017) Single-domain antibodies as versatile affinity reagents for analytical and diagnostic applications. Front Immunol 8:977. https://doi.org/10.3389/fimmu.2017.00977
Lauwereys M, Arbabi Ghahroudi M, Desmyter A, Kinne J, Holzer W, De Genst E, Wyns L, Muyldermans S (1998) Potent enzyme inhibitors derived from dromedary heavy-chain antibodies. EMBO J 17(13):3512–3520. https://doi.org/10.1093/emboj/17.13.3512
Joliot A, Pernelle C, Deagostini-Bazin H, Prochiantz A (1991) Antennapedia homeobox peptide regulates neural morphogenesis. Proc Natl Acad Sci U S A 88(5):1864–1868
Frankel AD, Pabo CO (1988) Cellular uptake of the tat protein from human immunodeficiency virus. Cell 55(6):1189–1193
Green M, Loewenstein PM (1988) Autonomous functional domains of chemically synthesized human immunodeficiency virus tat trans-activator protein. Cell 55(6):1179–1188
Glab-Ampai K, Malik AA, Chulanetra M, Thanongsaksrikul J, Thueng-In K, Srimanote P, Tongtawe P, Chaicumpa W (2016) Inhibition of HCV replication by humanized-single domain transbodies to NS4B. Biochem Biophys Res Commun 476(4):654–664. https://doi.org/10.1016/j.bbrc.2016.05.109
Phalaphol A, Thueng-In K, Thanongsaksrikul J, Poungpair O, Bangphoomi K, Sookrung N, Srimanote P, Chaicumpa W (2013) Humanized-VH/VHH that inhibit HCV replication by interfering with the virus helicase activity. J Virol Methods 194(1–2):289–299. https://doi.org/10.1016/j.jviromet.2013.08.032
Thueng-in K, Thanongsaksrikul J, Srimanote P, Bangphoomi K, Poungpair O, Maneewatch S, Choowongkomon K, Chaicumpa W (2012) Cell penetrable humanized-VH/V(H)H that inhibit RNA dependent RNA polymerase (NS5B) of HCV. PLoS One 7(11):e49254. https://doi.org/10.1371/journal.pone.0049254
Dobrev VS, Fred LM, Gerhart KP, Metallo SJ (2018) Characterization of the binding of small molecules to intrinsically disordered proteins. Methods Enzymol 611:677–702. https://doi.org/10.1016/bs.mie.2018.09.033
Zhang Z, Liu J, Rozovsky S (2018) Preparation of selenocysteine-containing forms of human SELENOK and SELENOS. Methods Mol Biol 1661:241–263. https://doi.org/10.1007/978-1-4939-7258-6_18
Pawson T (1995) Protein modules and signalling networks. Nature 373(6515):573–580. https://doi.org/10.1038/373573a0
Cohen GB, Ren R, Baltimore D (1995) Modular binding domains in signal transduction proteins. Cell 80(2):237–248
Nguyen JT, Porter M, Amoui M, Miller WT, Zuckermann RN, Lim WA (2000) Improving SH3 domain ligand selectivity using a non-natural scaffold. Chem Biol 7(7):463–473
Pisabarro MT, Serrano L (1996) Rational design of specific high-affinity peptide ligands for the Abl-SH3 domain. Biochemistry 35(33):10634–10640. https://doi.org/10.1021/bi960203t
Feng S, Kapoor TM, Shirai F, Combs AP, Schreiber SL (1996) Molecular basis for the binding of SH3 ligands with non-peptide elements identified by combinatorial synthesis. Chem Biol 3(8):661–670
Funding
This research was supported by NIAID/NIH grant R01AI089999.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Marciel, M.P., Hoffmann, P.R. Molecular Mechanisms by Which Selenoprotein K Regulates Immunity and Cancer. Biol Trace Elem Res 192, 60–68 (2019). https://doi.org/10.1007/s12011-019-01774-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12011-019-01774-8