Abstract
Numerous reports suggest the involvement of oxidative stress in cadmium toxicity, but the nature of the reactive species and the mechanism of Cd-induced oxidative damage are not clear. In this study, E. coli mutants were used to investigate mechanisms of Cd toxicity. Effects of Cd on metabolic activity, production of superoxide radical by the respiratory chain, and induction of enzymes controlled by the soxRS regulon were investigated. In E. coli, the soxRS regulon controls defense against O2·−and univalent oxidants. Suppression of metabolic activity, inability of E. coli to adapt to new environment, and slow cell division were among the manifestations of Cd toxicity. Cd increased production of O2·− by the electron transport chain and prevented the induction of soxRS-controlled protective enzymes, even when the regulon was induced by the redox-cycling agent, paraquat. The effect was not limited to soxRS-dependent proteins and can be attributed to previously reported suppression of protein synthesis by Cd. Increased production of superoxide, combined with inability to express protective enzymes and to replace damaged proteins by de novo protein synthesis, seems to be the main reason for growth stasis and cell death in Cd poisoning.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Increased production of reactive oxygen species (ROS) by cadmium has been reported for various organisms and cellular model systems, but very few studies applied direct, reliable methods for identification of the generated species [1, 2]. Cadmium is not a redox-active metal and cannot directly generate reactive species by reducing oxygen or by catalyzing decomposition of peroxides [3, 4]. The availability of superoxide dismutase (SOD)-deficient E. coli mutants makes this microorganism a suitable cellular model for studying the contribution of superoxide radical in Cd toxic action. It has been previously reported that sodA sodB E. coli mutants are more sensitive to Cd than their isogenic, SOD-proficient parents [5, 6]. Whether this indicates increased superoxide production by Cd or simply reflects an additive effect of two toxic agents, Cd and superoxide (O2·−), each acting by its own mechanism, is not clear. Furthermore, it has been speculated that SOD may protect against Cd toxicity not by scavenging O2·−, but by metal sequestration [5], as shown for Saccharomyces cerevisiae [7, 8]. Therefore, higher sensitivity of SOD-deficient cells to Cd might not be related to superoxide radical contribution.
The aim of this study was to clarify the involvement of superoxide radical and to shed more light on the mechanism of cadmium-induced oxidative stress. Using E. coli as a model system, we obtained direct evidence that Cd increases the production of superoxide radical by the electron transport chain. At the same time, Cd prevented the expression of protective enzymes controlled by the soxRS regulon and suppressed protein synthesis. Together with the increased production of superoxide radical, failure to induce adequate protection leads to irreparable oxidative cell damage. Since translation was suppressed, cells were unable to replace inactivated enzymes and damaged proteins by de novo protein synthesis.
Materials and Methods
Strains and Media
Luria–Bertani (LB) medium contained 10 g of bactotryptone, 5 g of yeast extract, and 10 g of NaCl per liter. M9CA medium was made up of M9 salts, 0.2% casamino acids, 0.2% glucose, 3 mg pantothenate, and 5 mg of thiamine per liter [9]. Low-phosphate 3-(N-morpholino)propanesulfonic acid (MOPS) medium was prepared as described by LaRossa and coworkers [6], except that casamino acids were added as in the M9CA medium.
The following E. coli strains were used in this study: GC4468 (F−Δlac U169 rpsL), QC1799 (same as GC4468 plus Δ sodA3, Δ sodB-kan) [10]; DJ 901 = GC4468 Δ (soxR-Zjc2204) Zjc2205::Tn10 K m; QC1817 = GC4468 ΔsodA 3, ΔsodB-kan, Δsox8::cat [11]; AB1157 [F− thr-1 leuB6 proA2 his-4 thi-1 argE2 lacY1 galK2 rpsL surE44 ara-14 xyl-15 mtl-1 tsx-33]; JI 132 [Same as AB1157 plus (sodA::Mu d PR13)25 (sodB-kan)1-Δ2] [12]; and KK196 = AB1157 (sodA::Mu d PR13)25, sodB-kan)1-Δ2 + pFeSOD, KK198 = (sodA::Mu d PR13)25, sodB-kan)1-Δ2 + pMnSOD. Strain XA90 containing the plasmid pKOXR (kindly provided by J. Imlay) was used for induction of β-galactosidase and soxR protein [13]. The strain was grown aerobically in LB medium and was induced with isopropyl-β-d-thiogalactopyranoside (IPTG) as described by Demple et al. [13].
Growth Conditions
Overnight cultures were grown in LB medium. For monitoring the effect of Cd on cell proliferation, overnight cultures were diluted to OD600 nm = 0.05 in M9CA, LB, or MOPS medium and 100 μl aliquots were transferred to triplicate wells in a 96-well plate. CdCl2 was added and growth was monitored turbidimetrically at 600 nm using a plate reader [9].
Assessment of Metabolic Activity
Reduction of 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl-tetrazolium bromide (MTT) to formazan was used as a measure of cell metabolic activity [14,15,16]. One hundred-microliter aliquots of E. coli cultures grown to OD600 nm = 0.3 in either M9CA, LB, or MOPS medium were transferred to triplicate wells in a 96-well plate and CdCl2 was added. After 2 h of incubation on a shaker at 37 °C, 10 μl of 0.5% MTT solution was added to each well. The plates were incubated at 37 °C on a shaker for additional 30 min, and 100 μl of SDS reagent (10% SDS in 10 mM HCl) was added to each well, followed by incubation for 1 h at room temperature [17]. Absorption at 570 nm (formazan) and 700 nm (background) was measured with a plate reader.
Superoxide Production
The effect of Cd on production of superoxide by the electron transport chain was studied using inverted membrane vesicles where substrates are oxidized and O2 is reduced on their outer surface [18]. For preparation of inverted membrane vesicles, overnight cultures of parental and SOD-deficient strains were grown in LB medium with vigorous shaking and were harvested at OD600 nm 0.5–0.6. Cultures were then centrifuged, washed in cold 50 mM phosphate buffer, pH 7.8, and resuspended to ∼3% of the original volume in the same buffer. The cells were disrupted with a French press and were centrifuged for 10 min at 10,000×g to remove cell debris. The supernatant, which contained both cytosolic material and membrane vesicles, was fractionated by a 3-h centrifugation at 85,000×g. The pelleted membrane vesicles were resuspended and recentrifuged for 3 h [18].
For assessing the effect of Cd on production of superoxide, the assay mixture contained 50 μl membrane vesicles, 10 μM acetylated cytochrome c, and 0.3 mM NADH, with or without 50 units bovine CuZnSOD, in 1.0 ml of 50 mM Tris buffer, pH 7.4. Cytochrome c reduction was recorded at 25 °C for 1 min. The difference between the rates of cytochrome c (cyt c) reduction without and with SOD was used to calculate superoxide production. Where CdCl2 was added, samples were preincubated for 5 min at 25 °C before adding NADH.
Induction of the soxRS Regulon
The effect of Cd on the induction of the soxRS regulon was performed by measuring the activities of enzymes coded by genes under the control of the regulon [11, 19, 20]. E. coli cultures were grown to mid-log phase with vigorous shaking, and cell suspensions were divided in 10 ml aliquots. Incubation of aliquots with CdCl2 was performed at 37 °C on a shaking water bath (200 rpm) for 60 and 120 min. When the effect of Cd on induction of enzymes by paraquat was assessed, cultures were preincubated for 15 min with CdCl2 and were then incubated with paraquat for additional 60 or 120 min. At the end of the incubation time, all cultures were washed, resuspended in the respective assay buffers, and disrupted with a French press [21]. The extracts were cleared from debris and unbroken cells by centrifugation at 10,000×g for 5 min and used for determination of protein and activities of enzymes. Protein content of the cell-free extract was estimated by the Lowry assay [22]. Fumarase activity was assayed by measuring the production of fumatrate at 250 nm [19]. One milliliter of assay mixture contained 50 mM l-malate in 50 mM sodium phosphate buffer, pH 7.3 [23]. One unit of fumarase converted 1 μmol/min of l-malate to fumarate, using ε 250 nm = 1.62 mM−1 cm−1 [11]. SOD activity was assayed as described by McCord and Fridovich [24], with slight modification [25]. One milliliter of reaction mixture contained in 50 mM potassium phosphate buffer (pH 7.8), 0.1 mM EDTA, 1 mM xanthine, 10 μM cyt c, and 5–10 μM xanthine oxidase. The initial rate of cyt c reduction (ΔA 550 nm) was recorded for 30 s; then, the sample was added to the cuvette, and the change of the absorbance was monitored for 30 additional seconds. One unit of the enzyme activity is defined as the amount of enzyme that causes 50% inhibition of the O2·−-induced cytochrome c reduction. Nitroreductase A activity [20] was measured using nitrofurantoin as a substrate. Reaction mixture contained in Tris buffer, pH 7.5: nitrofurantoin, 25 μΜ; NADPH, 100 μM; and NADH, 50 μM. Reduction of nitrofurantoin was measured at 378 nm using an extinction coefficient of 2.63 × 104 M−1 cm−1 at 373 nm [20]. Glucose-6-phosphate dehydrogenase activity was assayed as previously described [25]. NADPH production was determined at 340 nm in 1.0 ml of a reaction mixture containing 100 mM Tris–HCl (pH 8), 1 mM glucose-6-phosphate, and 1 mM NADP+. One unit of activity generates 1 μmol of NADPH per minute [25].
Statistics
Each experiment was repeated at least three times. Except where otherwise indicated, mean ± standard error is presented. One-way analysis of variance (ANOVA) was performed using SigmaPlot version 11.0, and p value ≤0.05 was accepted as statistically significant.
Results
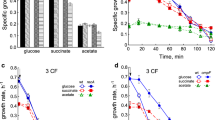
It had been previously reported that SODs protect E. coli against heavy metal toxicity [5]. In media normally used for growth of E. coli, Cd binding to phosphates and chelation by organic material decreases its active concentration. To avoid that, low-phosphate MOPS medium [6, 26] was used. When stationary phase cultures were diluted in MOPS medium, 2 μM CdCl2 substantially slowed the growth of the sodA sodB strain, and 5 μM prevented cell division for at least 6 h (Fig. 1a). Restoring either FeSOD or MnSOD abolished the hypersensitivity of the SOD-deficient mutant to Cd (Supplementary Fig. S1). Overexpression of SODs, however, did not make the overproducers more resistant to Cd than their parent. This result indicates that SODs do not protect against Cd by metal sequestration, as was previously proposed [5].
Stationary phase cultures were diluted to OD600 nm 0.05 in low-phosphate MOPS medium (a, b) or M9CA medium (c). CdCl2 was added to the indicated final concentrations, and cultures were grown on a shaking water bath at 37 °C and 200 rpm. Growth was monitored turbidimetrically by measuring the OD at 600 nm. c Optical density at 600 nm after 6 h of growth is presented as mean ± SE of three separate experiments
The parental, SOD-proficient strain grew better in MOPS medium and was much more resistant to Cd. At 5 μM CdCl2 did not retard growth and at 10 μM only extended the lag phase to 4 h, but did not decrease the growth rate. Complete suppression of growth of the parental GC4468 strain was observed at 50 μM CdCl2 (Fig. 1b).
As seen in Fig. 1a, the SOD-deficient strain did not grow well in MOPS medium, and for 8 h, OD600 nm reached only slightly above 0.25. Growth of the mutant almost completely stopped after 8 h even in the absence of Cd (not shown). To be sure that the effect was not due to inability of the sodA sodB mutant to grow in a low-phosphate medium [27], the experiment was repeated in M9CA medium. When stationary phase LB cultures were transferred to M9CA medium containing Cd, the sodA sodB mutant was about tenfold more sensitive to Cd than the SOD-proficient parent (Fig. 1c). Similar results were obtained when strains with different genetic background were tested (Supplementary Fig. S2) and when the experiment was performed in phosphate-free LB medium (not shown).
It has been suggested that the effect of Cd depends on cell density and growth phase [4, 28]. To test if Cd affects only the ability of stationary phase SOD-deficient cells to adapt and resume growth in a fresh medium, parental and SOD-deficient cultures were grown to mid-log phase in MOPS medium and were then diluted in fresh MOPS medium containing CdCl2. When both parental and sodA sodB mid-log cultures were diluted to OD600 = 0.05, the sodA sodB strain did not resume growth for up to 8 h in a medium containing 10 μM CdCl2, while 50 μM CdCl2 was needed to achieve the same effect for the parental strain (Supplementary Fig. S3). These results indicate that Cd suppresses both the ability of E. coli to adapt to new environment and division of cells that were already in mid-log phase. In both situations, the sodA sodB strain was more sensitive to Cd than its isogenic parent.
Since Cd has high affinity for sulfur, it can be expected that effects of Cd will be associated with inhibition of functions depending on sulfur-containing proteins [4, 29]. Among such functions is catalysis in metabolic pathways. Therefore, inhibition of metabolic enzymes and consequently suppression of metabolism is expected to be among the first manifestations of Cd toxicity. Superoxide radical in turn interferes with sulfur utilization by E. coli [30] and suppresses aerobic metabolism. It seems reasonable to expect that high steady-state O2·− concentrations in sodA sodB cells would add to the suppression of metabolism by Cd, which could explain the higher sensitivity to Cd of the SOD-deficient cultures. We used the MTT assay, based on enzymatic reduction of MTT to colored formazan, as an integrative measure of the effect of Cd on cell metabolism. Figure 2 shows that Cd inhibits MTT reduction by both parental and SOD-deficient cells, but no statistically significant differences in sensitivity to Cd between the two strains were observed at any of the tested concentrations. Similar results were obtained when strains with different genetic backgrounds were tested (not shown). Based on these data, it can be concluded that higher sensitivity of SOD− strains to Cd is not a consequence of stronger inhibition of metabolism.
Parental (GC4468) and SOD-deficient (QC1799) cultures were grown to OD600 nm = 0.3, and CdCl2 was added to the specified final concentrations. After 2 h of incubation, 100 μl aliquots were transferred to triplicate wells in a 96-well plate and the MTT assay was performed. Mean ± SEM is presented (n = 4). No statistically significant differences were found when sensitivity of parental and SOD− cells was compared for each CdCl2 concentration (p = 0.245–0.294)
It has been previously reported that Cd inhibits cellular respiration and that a ΔubiE mutant displaying low respiration rate was more resistant to Cd [26]. It is possible that by blocking the electron flow in the electron transport chain, Cd diverts more electrons to one-electron reduction of O2 resulting in increased O2·− production.
In isolated mammalian mitochondria, inhibition of the electron transport chain by Cd was accompanied with increased production of ROS [2]. Judging by the inhibition of the ESR signals by SOD (0.5 mg/ml) [2], superoxide production was increased by Cd. In order to find if Cd increases the release of O2·− by the electron transport chain of E. coli and if more superoxide is produced by the SOD-deficient mutant, inverted membrane vesicles prepared from SOD-deficient and SOD-proficient cells were used [18]. Superoxide production was measured by the SOD-inhibitable reduction of cytochrome c.
Addition of Cd increased the rate of SOD-inhibitable cytochrome c reduction, which reflects the increase of superoxide radical production [18]. The effect reached a maximum at 2.0 μM CdCl2 (Fig. 3). Further increase of Cd concentration up to ∼5.0 μM did not increase O2·− production, and concentrations above 5.0 μM caused suppression of cytochrome c reduction. At all tested concentrations of CdCl2, no statistically significant differences in the rates of O2·− production between the SOD-proficient parent and the SOD-deficient mutant were observed.
Superoxide production by inverted membrane vesicles. The reaction mixture contained in 1.0 ml of 50 mM Tris buffer, pH 7.4: 50 μl membrane vesicles, 10 μM acetylated cytochrome c, and 0.3 mM NADH, with or without 50 units of bovine CuZnSOD. Cytochrome c reduction at 25 °C was recorded for 1 min. Samples were preincubated for 5 min with CdCl2 at 25 °C before adding NADH. The experiment was repeated three times in triplicates with separate vesicle preparations. Mean ± SE is presented
Based on results obtained by reverse transcription polymerase chain reaction (RT-PCR), it has been concluded that Cd can induce the soxRS regulon [3, 31], which in turn would upregulate the translation of superoxide-resistant metabolic enzymes and enzymes involved in protection against oxidative stress. Among them are MnSOD, endonuclease IV, fumarase C, glucose-6-phosphate dehydrogenase (G6PD), nitroreductase A, and aconitase A. It has been previously reported that Cd induces the genes coding for fumarase C and aconitase A, but data were obtained by microarray and activities of the gene products have not been determined [31]. Our results showed that incubation of parental and sodA sodB cells with CdCl2 at a wide concentration range (1.0–100 μM) did not cause induction of any of the members of the soxRS regulon. No increase of the activities of fumarase C, MnSOD (parental), nitroreductase A, or G6PD was observed irrespective of the presence or absence of growth inhibition by Cd (Supplementary Fig. S4).
To test if such lack of response is due to inhibition of the enzymes by Cd, soxRS was induced by paraquat and cell-free extracts were obtained. The cell-free extracts were aliquoted, and CdCl2 was added at concentrations ranging from 20 to 100 μM. After short (5 min) preincubation, the extracts were assayed for activities of fumarase C, G6PD, and SOD. No inhibition of the enzymes by Cd was detected (data not shown).
Another possible explanation for the lack of induction of soxRS-controlled enzymes is prevention by Cd of the activation of the soxRS regulon. To test this possibility, soxRS-dependent enzymes were induced by paraquat. CdCl2 was added at varying concentrations 15 min before the addition of paraquat, and the cells were incubated with paraquat for 1 or 2 h. Results demonstrated that Cd prevented the induction by paraquat of fumarase C (Fig. 4), MnSOD, nitroreductase A, and G6PD (not shown). Varying the concentration of paraquat from 20 to 100 μM did not change the inhibitory effect of Cd (not shown). Similar result was obtained when the sodA sodB strain was tested (Supplementary Fig. S5).
Parental, GC4468 culture was grown in MOPS medium to mid-log phase and was divided in 10 ml aliquots. The aliquots were preincubated for 15 min with CdCl2 at 37 °C on a shaking water bath (200 rpm). After that, paraquat was added to a final concentration of 100 μM, and cultures were left on the shaker for additional 60 min. At the end of the incubation time, the cultures were washed, resuspended in phosphate buffer, and disrupted with a French press. Fumarase C and protein were determined in the cell-free extracts. Data are presented as mean ± SE (n = 3)
Among the reasons behind such an effect could be prevention by Cd of the induction of the soxRS regulon. Induction of the soxRS regulon depends on activation of soxS by a transcription factor, soxR. Both, reduced form of soxR and apo-soxR (soxR stripped of Fe-S clusters), bind to DNA, but only the oxidized soxR activates soxS [32, 33]. It has been reported that Cd inactivates [4S-4Fe] cluster-containing proteins by disrupting the clusters and releasing free Fe [4]. By analogy, it may be suspected that Cd is capable of disrupting the fully coordinated [2Fe-2S] clusters of SoxR, thus preventing the activation of the signaling cascade. This means that the effect of Cd would be specific, limited to the induction of soxRS-dependent, but not to other genes products. To test the specificity of Cd interaction, its effect on induction of β-galactosidase by isopropyl-β-d-thiogalactoside (IPTG) has been determined. As seen (Fig. 5a), Cd suppressed the induction of β-galactosidase the same way that it suppressed the induction of fumarase C (Fig. 5b). Among the reasons could be prevention of protein translation by Cd [3].
Effect of Cd on IPTG-induced a β-galactosidase activity and b paraquat-induced fumarase C. Strain XA90/pKOXR was grown to mid-log phase in aerobic LB medium, and CdCl2 was added to the indicated final concentrations. After 15 min, IPTG and paraquat (100 μM) were added and cultures were incubated for 60 min at 37 °C with shaking
Results obtained so far indicate that Cd increases O2·− production, which eventually explains the higher sensitivity of the sodA sodB mutants, and prevents the induction of soxRS-dependent protection. It is reasonable to expect that if SODs are present, proteins would be protected against oxidative damage, and the inability to induce soxRS, and to synthesize proteins in general, would not be so critical. To test this assumption, the effect on Cd on soxRS-deficient mutants with either SOD-positive or SOD-negative background has been investigated. It was found that absence of soxRS did not make a SOD-proficient mutant more sensitive to Cd than the respective parental strain (Fig. 6a). This is not unexpected since soxRS-dependent proteins are not synthesized anyway in the presence of toxic concentrations of Cd. A triple sodA sodB soxRS − mutant, QC1817, was much more sensitive to Cd than the SOD-proficient soxRS − strain DJ901. It grew very poorly in MOPS medium and needed 10 h to reach OD600 of only 0.11 (Fig. 6b). Its growth was completely prevented by CdCl2 at a concentration as low as 1 μM. This result suggests that in the SOD-negative background, members of soxRS, already expressed before the addition of Cd [11], exert protection and make the cells more resistant to Cd-induced damage.
Effect of deletion of soxR on sensitivity to Cd. Two SoxRS-negative mutants, a DJ901 (SOD-proficient) and b QC1817 (SOD-deficient), were grown to stationary phase in LB medium. The cultures were diluted to OD600 nm ∼0.05 in low-phosphate MOPS medium and CdCl2 was added. Growth was followed at 600 nm
Discussion
Oxidative stress has been reported for organisms exposed to cadmium [1, 26, 31], and various indirect mechanisms of Cd prooxidant action have been proposed: depletion of cellular antioxidants and inhibition of antioxidant enzymes [31], release of redox-active metals from Fe-S clusters [4, 31], and stimulation of ROS release by interference with the mitochondrial electron transport chain [2]. The importance of Cd-induced oxidative damage has been supported by the finding that SOD-deficient E. coli mutants are more sensitive to Cd than the SOD-proficient parents [5, 6]. Using MTT reduction as an integrative measure of metabolic activity, we found that the inhibition of cell metabolism by Cd is similar in both the SOD− and the SOD+ strains. At the same time, Cd increased O2·− production by inverted membrane vesicles which contain the components of the electron transport chain, but again, no difference between parental and SOD-deficient preparations was observed. This is not surprising, because SODs are soluble and membrane preparations were free of SOD.
Based on the concentration of cytoplasmic SODs (∼20 μM) and the rate of O2·− production (∼5 μM/s), it has been calculated that in SOD-proficient E. coli exponentially growing in air-saturated medium, O2·− concentration is ∼0.1 nM [34], which is enough to inactivate half of the [4Fe-4S] enzymes within ∼30 min. Increase in the rate of production of superoxide or decrease of SODs would cause damage of O2·− targets (mainly [4F-4S] cluster containing enzymes), which explains the slow aerobic growth of the SOD-deficient strains even in complete media. In our experiments, Cd increased O2·− production by about 2.5-fold. Based on the rate constant of reaction between [4Fe-4S] clusters and O2·−, 106–107 M−1 s−1 (ignoring all other potential superoxide targets), it has been concluded that considerable enzyme damage and consequently suppression of growth would occur if [O2·−] increases by more than twofold [34]. Such a deleterious effect can be eventually prevented by induction of the soxRS regulon [11, 35].
It has been reported, based on data obtained by microarray and RT-PCR, that soxR is upregulated after up to 25-min incubation of E. coli with low Cd concentration (∼9 μM) [3]. Based on the same techniques, activation of soxR and soxS at higher Cd concentration (100 μM for 10 min) has also been observed [31]. Genes induced by soxRS include sodA (MnSOD), to scavenge O2·−; zwf (G6PD), to generate NADPH needed for reduction of glutathione disulfide; fumC (fumarase C), to replace the O2·−-sensitive fumarases A and B; acnA (aconitase A); nfo (endonuclease IV), to repair oxidative DNA damage; ntrA (nitroreductase A), fur (regulator of Fe metabolism), etc. (reviewed in [36]). Among the members of the soxRS regulon, fur [3], acnA [31], and fumC [31] were reportedly activated by Cd exposure. It has been speculated that exposure to Cd leads to damage and oxidation of the SoxR Fe-S clusters, which activates soxS and consequently upregulates fumC [31]. Such induction, however, might be following different mechanisms, as found for genes under Fe deprivation [37]. Furthermore, the previously listed studies did not measure the activities of the protein products coded by the genes controlled by the soxRS regulon. Our data show that none of the tested enzymes was induced by Cd when applied in a wide concentration range (1–100 μM). What is more, Cd prevented the induction of the soxRS-controlled enzymes by the redox-cycling agent paraquat. This appeared not to be due to specific inactivation of soxRS and most probably resulted from generalized suppression of protein synthesis [3].
Based on the knowledge obtained so far, the following scenario for the toxic action of Cd could be envisaged. Cadmium-induced distortion of protein complexes [31] seems to be the most probable cause of alterations of complexes of the electron transport chain and has two important consequences: (a) switch to inefficient anaerobic metabolism and (b) production of superoxide radical. Superoxide in turn reacts with [4Fe-4S] cluster-containing proteins, liberating free Fe [38,39,40]. This adds to the Fe released form [4Fe-4S] as a result of direct Cd-protein interaction [4]. Hydrogen peroxide generated as a result of increased O2·− production and Fe released from [4Fe-4S] clusters interact in generating hydroxyl radical by a site-specific Fenton reaction [41]. Hydroxyl radical not only is capable of modifying practically all biomolecules, but also initiates free radical chain reactions that augment the damage by generating reactive toxic products. Oxidative inactivation of enzymes, together with direct Cd-protein interactions, and suppression of protein translation [3] will impair metabolism and will lead to growth stasis; oxidatively damaged DNA which is not repaired will cause loss of viability, and these are well-recognized consequences of Cd intoxication.
Proteins that are oxidatively modified are subjected to proteolytic degradation [42, 43]. As a result, the content of some proteins in E. coli exposed to superoxide or hydrogen peroxide drops below half of their normal level [42]. In order to keep metabolic activity and survive, cells replace damaged proteins by de novo protein synthesis. Cadmium, however, blocks translation [3] and prevents protein synthesis. This affects much stronger the SOD-deficient cells, which sustain enhanced oxidative damage due to higher steady-state concentration of O2·−.
References
Liu J, Qu W, Kadiiska MB (2009) Role of oxidative stress in cadmium toxicity and carcinogenesis. Toxicol Appl Pharmacol 238(3):209–214. doi:10.1016/j.taap.2009.01.029
Wang Y, Fang J, Leonard SS, Rao KMK (2004) Cadmium inhibits the electron transfer chain and induces reactive oxygen species. Free Radic Biol Med 36(11):1434–1443
Wang A, Crowley DE (2005) Global gene expression responses to cadmium toxicity in Escherichia coli. J Bacteriol 187(9):3259–3266
Xu FF, Imlay JA (2012) Silver(I), mercury(II), cadmium(II), and zinc(II) target exposed enzymic iron-sulfur clusters when they toxify Escherichia coli. Appl Environ Microbiol 78(10):3614–3621
Geslin C, Llanos J, Prieur D, Jeanthon C (2001) The manganese and iron superoxide dismutases protect Escherichia coli from heavy metal toxicity. Res Microbiol 152(10):901–905
LaRossa RA, Smulski DR, Van Dyk TK (1995) Interaction of lead nitrate and cadmium chloride with Escherichia coli K- 12 and Salmonella typhimurium global regulatory mutants. J Ind Microbiol 14(3–4):252–258 Publication year 1995
Ciriolo MR, Civitareale P, Carri MT, Demartino A, Galiazzo F, Rotilio G (1994) Purification and characterization of Ag,Zn-superoxide dismutase from Saccharomyces cerevisiae exposed to silver. J Biol Chem 269(41):25783–25787
Culotta VC, Joh HD, Lin SJ, Slekar KH, Strain J (1995) A physiological role for Saccharomyces cerevisiae copper/zinc superoxide dismutase in copper buffering. J Biol Chem 270(50):29991–29997
Tovmasyan A, Reboucas JS, Benov L (2014) Simple biological systems for assessing the activity of superoxide dismutase mimics. Antioxid Redox Signal 20(15):2416–2436. doi:10.1089/ars.2013.5576
Touati D, Jacques M, Tardat B, Bouchard L, Despied S (1995) Lethal oxidative damage and mutagenesis are generated by iron in delta fur mutants of Escherichia coli—protective role of superoxide dismutase. J Bacteriol 177(9):2305–2314
Liochev SI, Benov L, Touati D, Fridovich I (1999) Induction of the soxRS regulon of Escherichia coli by superoxide. J Biol Chem 274(14):9479–9481. doi:10.1074/jbc.274.14.9479
Imlay JA, Linn S (1987) Mutagenesis and stress responses induced in Escherichia coli by hydrogen peroxide. J Bacteriol 169(7):2967–2976
Demple B, Ding H, Jorgensen M (2002) Escherichia coli SoxR protein: sensor/transducer of oxidative stress and nitric oxide. Methods Enzymol 348. doi:10.1016/S0076-6879(02)48623-5
Tsukatani T, Higuchi T, Suenaga H, Akao T, Ishiyama M, Ezoe T, Matsumoto K (2009) Colorimetric microbial viability assay based on reduction of water-soluble tetrazolium salts for antimicrobial susceptibility testing and screening of antimicrobial substances. Anal Biochem 393(1):117–125. doi:10.1016/j.ab.2009.06.026
Berridge MV, Herst PM, Tan AS (2005) Tetrazolium dyes as tools in cell biology: new insights into their cellular reduction. Biotechnol Annu Rev 11:127–152
Berridge MV, Tan AS (1993) Characterization of the cellular reduction of 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT): subcellular localization, substrate dependence, and involvement of mitochondrial electron transport in MTT reduction. Arch Biochem Biophys 303(2):474–482
Thomas M, Benov L (2011) Cell-based bioassay for compounds with prooxidant activity. In: Brebbia CA, Eglite M, Knets I, Miftahof R, Popov V (eds) WIT transactions on biomedicine and health, vol 15. WIT Press, Southampton, pp 171–182. doi:10.2495/EHR110161
Imlay JA, Fridovich I (1991) Assay of metabolic superoxide production in Escherichia coli. J Biol Chem 266(11):6957–6965
Liochev SI, Fridovich I (1992) Fumarase C, the stable fumarase of Escherichia coli, is controlled by the Soxrs regulon. P Natl Acad Sci USA 89(13):5892–5896. doi:10.1073/pnas.89.13.5892
Liochev SI, Hausladen A, Fridovich I (1999) Nitroreductase A is regulated as a member of the soxRS regulon of Escherichia coli. P Natl Acad Sci USA 96(7):3537–3539. doi:10.1073/pnas.96.7.3537
Benov L, Al-Ibraheem J (2002) Disrupting Escherichia coli: a comparison of methods. J Biochem Mol Biol 35(4):428–431
Lowry OH, Rosebrough NJ, Farr AL, Randall RJ (1951) Protein measurement with the Folin phenol reagent. J Biol Chem 193(1):265–275
Benov L, Fridovich I (2002) Induction of the soxRS regulon of Escherichia coli by glycolaldehyde. Arch Biochem Biophys 407(1):45–48
McCord JM, Fridovich I (1969) Superoxide dismutase. An enzymic function for erythrocuprein (hemocuprein). J Biol Chem 244(22):6049–6055
Al-Mutairi DA, Craik JD, Batinic-Haberle I, Benov LT (2007) Inactivation of metabolic enzymes by photo-treatment with zinc meta N-methylpyridylporphyrin. Biochim Biophys Acta Gen Subj 1770(11):1520–1527
Pacheco CC, Passos JF, Castro AR, Moradas-Ferreira P, De Marco P (2008) Role of respiration and glutathione in cadmium-induced oxidative stress in Escherichia coli K-12. Arch Microbiol 189(3):271–278
Al-Maghrebi MA, Benov LT (2001) Polyphosphate accumulation and oxidative DNA damage in superoxide dismutase-deficient Escherichia coli. Free Radic Biol Med 31(11):1352–1359
Ferianc P, Farewell A, Nyström T (1998) The cadmium-stress stimulon of Escherichia coli K-12. Microbiology 144(4):1045–1050
Nies DH (1999) Microbial heavy-metal resistance. Appl Microbiol Biotechnol 51(6):730–750
Benov L, Kredich NM, Fridovich I (1996) The mechanism of the auxotrophy for sulfur-containing amino acids imposed upon Escherichia coli by superoxide. J Biol Chem 271(35):21037–21040
Helbig K, Grosse C, Nies DH (2008) Cadmium toxicity in glutathione mutants of Escherichia coli. J Bacteriol 190(15):5439–5454. doi:10.1128/Jb.00272-08
Hidalgo E, Demple B (1994) An iron-sulfur center essential for transcriptional activation by the redox-sensing Soxr protein. EMBO J 13(1):138–146
Gaudu P, Weiss B (1996) SoxR, a [2Fe-2S] transcription factor, is active only in its oxidized form. P Natl Acad Sci USA 93(19):10094–10098. doi:10.1073/pnas.93.19.10094
Gort AS, Imlay JA (1998) Balance between endogenous superoxide stress and antioxidant defenses. J Bacteriol 180(6):1402–1410
Baez A, Shiloach J (2013) Escherichia coli avoids high dissolved oxygen stress by activation of SoxRS and manganese-superoxide dismutase. Microb Cell Factories 12:23. doi:10.1186/1475-2859-12-23
Touati D (2000) Sensing and protecting against superoxide stress in Escherichia coli—how many ways are there to trigger soxRS response? Redox Rep 5(5):287–293
Fuentes AM, Díaz-Mejía JJ, Maldonado-Rodríguez R, Amábile-Cuevas CF (2001) Differential activities of the SoxR protein of Escherichia coli: SoxS is not required for gene activation under iron deprivation. FEMS Microbiol Lett 201(2):271–275. doi:10.1016/S0378-1097(01)00283-X
Benov L (2001) How superoxide radical damages the cell. Protoplasma 217(1–3):33–36. doi:10.1007/Bf01289410
Liochev SI (1996) The role of iron-sulfur clusters in in vivo hydroxyl radical production. Free Radic Res 25(5):369–384. doi:10.3109/10715769609149059
Imlay JA (2003) Pathways of oxidative damage. Annu Rev Microbiol 57:395–418. doi:10.1146/annurev.micro.57.030502.090938
Fridovich I (2013) Oxygen: how do we stand it? Med Princ Pract 22(2):131–137. doi:10.1159/000339212
Davies KJ, Lin SW (1988) Oxidatively denatured proteins are degraded by an ATP-independent proteolytic pathway in Escherichia coli. Free Radic Biol Med 5(4):225–236
Davies KJ, Lin SW (1988) Degradation of oxidatively denatured proteins in Escherichia coli. Free Radic Biol Med 5(4):215–223
Acknowledgements
This work was supported by Kuwait University grant MB 03/07. We thank Dr. J. Imlay (University of Illinois at Urbana-Champaign, Urbana, IL) for the helpful discussions. We are also grateful to Dr. J. Imlay and Dr. D. Touati (Institute Jacques Monod, CNRS, Paris, France) for providing the E. coli strains used in this study.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
ESM 1
(DOCX 125 kb).
Rights and permissions
About this article
Cite this article
Thomas, M., Benov, L. The Contribution of Superoxide Radical to Cadmium Toxicity in E. coli . Biol Trace Elem Res 181, 361–368 (2018). https://doi.org/10.1007/s12011-017-1048-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12011-017-1048-5