Abstract
This study focused on a haloduric BTEX-degrading microbial consortium EC20 enriched from Bohai Sea sediment. EC20 degraded 87% of BTEX at 435 mg L−1 initial concentration (benzene, toluene, ethylbenzene, and xylenes in equal proportions) in the presence of 3.4% NaCl. 16S rRNA gene-based PCR-DGGE profiles revealed that the dominant bacteria in EC20 were Proteobacteria, Actinobacteria, Bacteroidetes, and Firmicutes at the phylum level, and Pseudomonas, Mesorhizobium, Achromobacter, Stenotrophomonas, and Halomonas at the genus level. PCR detection of genes coding the key enzymes which participated in BTEX degradation pathways showed that the enriched consortium EC20 contained TOL pathway and TOD pathway to initiate biodegradation of BTEX.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
BTEX is a commonly used abbreviation for the compounds benzene, toluene, ethylbenzene, and xylenes; all are volatile aromatic hydrocarbons containing a benzene ring. They are found in a variety of petroleum-related products such as tar, crude petroleum, diesel, and petrol fuels, and they account for about 18% (w/w) in standard gasoline and nearly 90% of its water-soluble fraction [1, 2]. BTEX are widespread in groundwater, soil, and sediments as contaminants caused by oil spills or leaking of underground storage tanks [2], and they naturally occur in seawater around areas with deposits of natural gas, petroleum, and coal deposits, as well as in gas emissions from volcanoes. BTEX compounds are known to possess toxic, mutagenic, and carcinogenic properties even in very low concentrations, which can cause adverse effects on human health, such as drowsiness, dizziness, rapid heart rate, headaches, and so on. For this reason, they are classified as priority pollutants by the U.S. Environmental Protection Agency [3–5].
Bioremediation is a cost-effective and environmental strategy to remove the BTEX from contaminated environment [6]. Many bacteria can degrade BTEX under aerobic conditions, such as Pseudomonas, Acinetobacter, Serratia, Comamonas, Coccobacillus, Bacillus, Burkholderia, Terrimonas, Sphingomonas, Chryseobacterium, and Fulvimonas [5, 7–10]. Pseudomonas-related microorganisms are the most widespread BTEX degraders in the environment [11]. Previous studies on BTEX biodegradation tended to focus on single isolates; however, pure cultures are usually powerless when all BTEX compounds exist simultaneously [12, 13]. It has been reported that microbial consortium is more efficient on BTEX-degrading than the single strains [14]. Bioaugmentation is one of the bioremediation strategies by adding targeted microorganisms to the contaminated environment for enhancing the bioremediation efficiency [15, 16]. However, the selected exogenous BTEX degrader(s) might not be so competitive compared with the intrinsic bacteria, which may weaken the efficiency of biodegradation [17]. Therefore, screening intrinsic degraders from contaminated sites is a better way to improve bioremediation efficiency and it will not destroy the original bacterial diversity [13, 17, 18].
The key steps of the BTEX metabolic pathways are the ring activation and cleavage [19, 20], including that the aromatic ring is directly attacked by dioxygenase and monooxygenase respectively in the TOD and TOM pathways, and the alkyl side chain is oxidized by monooxygenase in the TOL pathway [11, 20]. Then the intermediate products such as catechol and catechol-like intermediates will undergo ring cleavage catalyzed by catechol 1,2-dioxygenase (1,2-CTD) or catechol 2,3-dioxygenase (2,3-CTD), and the products will participate in the Kreb’s cycle via central reactions [20, 21].
The Bohai Sea was an important fishing area for Chinese fishermen, but the fish resource has been almost exhausted by heavy contamination and now this area is also an important petroleum field. Previously, BTEX at about 7 mg L−1 in sediments from unpolluted areas and 200–300 mg L−1 in sediments at the oil fields have been detected in Bohai Sea, and BTEX concentration in sediments at a seriously polluted area even reached 450 mg L−1 [22, 23]. The BTEX in high concentration might threaten the survival of the microorganisms and weaken the degradation ability because of their toxicity [24]. Therefore, removing BTEX in the Bohai Sea is of both ecological and economical importance.
Considering the effects of environmental conditions and the contaminant compositions on the microbial communities, the BTEX-degrading microbes varied according to the geological location. Therefore, the effective degraders of the BTEX should be also adapted to the local conditions. In the present study, to obtain a microbial consortium capable of degrading BTEX efficiently in Bohai Sea, a series of enrichment experiments were carried out from the sediments of Bohai Sea in mineral saline medium (MSM) supplied with BTEX as the sole energy source. Then, the bacterial consortium was characterized using 16S ribosomal RNA (rRNA) gene-based PCR-DGGE methods, and the BTEX-degrading pathways were determined by detecting key catabolic genes in the bacterial consortium.
Materials and Methods
Reagents
Analytical reagent grade benzene, toluene, ethylbenzene, and xylenes were purchased from Sinopharm Chemical Reagent Co., Ltd. (Beijing, China). All the other chemicals used in the present study, unless otherwise noted, were analytical grade.
Samples and Enrichment of BTEX-Degrading Microbes
Sediment samples were collected from a near shore site away from the oilfield in Liaodong Bay of Bohai Sea in October 2011 at depths of 10–30 m with the method described by Yang et al. [25]. Samples were transported to the laboratory in sterile bags on ice, and then aliquots of samples were stored at −20 °C for microbial analysis.
The enrichment cultures were set up in 500 mL serum bottle sealed with butyl rubber stoppers and containing 300 mL of liquid MSM as described by Rasheed et al. [26] and supplemented with 3.4% NaCl (equal to the NaCl concentration of the Bohai Sea). Then 10 g of homogenized sediment and 870 mg L−1 BTEX components (benzene, toluene, ethylbenzene, and xylenes in equal proportions) as the sole carbon source were added to each bottle. The bottles were incubated in dark at 30 °C with horizontal shaking at 150 rpm. When significant growth of microorganisms was observed, 3 mL (1%, v/v) of the slurry in the bottles was transferred to another bottle with fresh MSM and BTEX components as mentioned above. The subcultivation was repeatedly executed for 20 times, and at the last three times, all the slurry was centrifuged at 12,000 rpm for 5 min and the precipitation was collected and regarded as bacterial consortium. The bacterial consortia from these three subcultures were named as EC18 (18 times subcultivation), EC19 (19 times subcultivation), and EC20 (20 times subcultivation), respectively. To further investigate the variation of bacterial community structure after incubated with single components of BTEX, EC20 was inoculated to MSM with 50 mg L−1 of each individual substrate (benzene, toluene, ethylbenzene, or xylenes) and subcultured twice with the method mentioned above. Then bacterial consortia named as ECB, ECT, ECE, and ECX (B—benzene, T—toluene, E—ethylbenzene, X—xylenes) were collected, respectively.
Growth Characteristics and BTEX Degradation Assay
The bacterial growth of EC20 was studied on MSM supplemented with 200 and 435 mg L−1 of BTEX mixture (benzene, toluene, ethylbenzene, and xylenes in equal proportion). Growth of the consortium was determined by measuring the optical density at 600 nm (OD600). Abiotic controls with no microbial inoculums were conducted simultaneously to monitor natural losses of BTEX. All of the trials and controls were run in triplicate.
The degrading kinetics of EC20 was researched on MSM supplemented with 435 mg L−1 BTEX mixture (benzene, toluene, ethylbenzene, and xylenes in equal proportion). During 72 h of incubation, the amount of residual BTEX compounds in the culture medium at various time intervals was quantified by gas chromatography (GC) using a SHIMADZU 2010 gas chromatograph equipped with a flame ionization detector and a DM-Wax (DIKMA, CA, USA) 60 m × 0.32 mm × 0.5 μm silica GC capillary column. Column, injector, and detector criteria were operated as reported in a prior study [27]. Uninoculated media (controls) were carried out to eliminate the natural losses of BTEX in the incubation, and two replicates and one control were run in parallel.
PCR-DGGE Analyses
The total genomic DNAs of the consortia were extracted using the TIANamp Bacteria DNA Kit (TIANGEN Biotech, Beijing, China) according to the manufacturer’s protocol. V3 region of 16S rRNA genes were amplified by touchdown PCR using the primers with GC-clamp 357F (5′-CGC CCG CCG CGC GCG GCG GGC GGG GCG GGG GCA CGG GGG GCC TAC GGG AGG CAG CAG-3′; the GC-clamp is italicized) and 518R (5′-ATT ACC GCG GCT GCT GG-3′). The PCR amplification was carried out by a Veriti 96-well Thermal Cycler (Thermo Fisher Scientific, MA, USA) in a 25-μL reaction mix containing 50 ng of genomic DNA, 1.5 mM MgCl2, 0.2 mM dNTPs, 0.2 μM (each) primer, and 1.5 U of Taq polymerase in 1× PCR buffer (Thermo Fisher Scientific, MA, USA). The PCR was performed under the following cycling conditions: initial denaturation at 95 °C for 5 min, 35 amplification cycles at 95 °C for 2 min, 65 °C (annealing temperature decreased by 0.5 °C per cycle for 20 cycles and held at 55 °C for 15 cycles) for 1 min, and 72 °C for 45 s, followed by extension step at 72 °C for 10 min. After confirmed by 1% agarose gel electrophoresis, the PCR products were purified using a Universal DNA Purification kit (TIANGEN Biotech, Beijing, China) following the manufacturer’s instructions.
DGGE of the purified PCR products and recovery of the DGGE bands were performed with the reagents and methods described by Wang et al. [28]. The relative intensity of each DGGE band to the overall signal in the identical lane were analyzed with the Quantity One software (Bio-Rad Laboratories, CA, USA). The recovered DGGE bands served as DNA template separately for re-amplification, and the same primers (without GC-clamp), reaction systems, and thermal cycling program were used as mentioned above. Then, the purified PCR products were ligated into pGEM®-T vector (Promega, WI, USA) and transformed into Escherichia coli DH5α (Biomed, Beijing, China) using the manufacturer’s guidelines. The transformed E. coli clones were spread on Luria-Bertani (LB) medium containing 50 μg mL−1 ampicillin and grew for 15 h at 37 °C. About ten colonies of each clone library were randomly picked using sterilized toothpicks and transferred to new ampicillin-contained LB plates. Clones were verified for DNA inserts by colony PCR with the M13 primers; a total of 10 μL PCR mixture contained 1× Taq PCR StarMix (GenStar, Beijing, China), 0.2 μM M13 (each) primer, and clones picked by sterilized toothpicks, and the PCR program included initial denaturation at 95 °C for 5 min, 25 amplification cycles at 95 °C for 30 s, 55 °C for 30 s, and 72 °C for 1 min, and extension step at 72 °C for 10 min. Then about five positive colonies of each clone library were selected randomly and sequenced. The nucleotide sequences (vector sequences removed) were compared with extant sequences in GenBank via BLAST searches. Molecular evolutionary genetics analysis was performed by MEGA 6.0.
Catabolic Gene Detection and Identification
The total genomic DNA was extracted from EC20 as mentioned above and genes coding for the key catabolic enzymes involved in the initial attacking and ring cleavage were amplified, including TMOA, TBMD, and TODC1 (monooxygenase and dioxygenase involved in the activation of benzene ring); TOL and XYLA (side chain monooxygenase involved in initial oxidizing); and XYLE1, XYLE2, CDO, TBUE, and TODE (subfamilies of catechol extradiol dioxygenases catalyzing ring cleavage). The PCR primers described by Hendrickx et al. [21] were listed in the supplementary Table S1. A 25-μL PCR mixture contained 1× Taq PCR StarMix (GenStar, Beijing, China), 0.2 μM forward primer, 0.2 μM reverse primer, and 100 ng of genomic DNA. The PCR program used for all primers was initial denaturation at 95 °C for 10 min, 35 amplification cycles at 95 °C for 1 min, 62 °C (annealing temperature decreased by 0.5 °C per cycle for 20 cycles and held at 52 °C for 15 cycles) for 1 min, and 72 °C for 1 min, with a final extension at 72 °C for 10 min. The purification and clone library construction of PCR products were carried out as described above. About 30 positive colonies of each clone library were selected randomly and sequenced. The nucleotide sequences (vector sequences removed) were compared with extant sequences in GenBank via BLAST searches. Molecular evolutionary genetics analysis was performed by MEGA 6.0.
Nucleotide Sequence Accession Numbers
The obtained nucleotide sequences of V3 region of 16S rRNA genes (accession numbers KP747006–KP747133) and partial BTEX catabolic genes (accession numbers KP746941–KP746971) are accessible in GenBank.
Results and Discussion
Enrichment and Determination of Growth
By the enrichment and 20 times of subculture in liquid MSM supplemented with 3.4% NaCl and 870 mg L−1 BTEX, a consortium named as EC20 was obtained. The growth curves of EC20 in MSM with different concentrations of BTEX showed no lag phase at low initial substrate concentrations (<50 mg L−1) (data not shown), while obvious lag time was observed for the growth of EC20 at higher substrate concentrations (200 and 435 mg L−1) of BTEX (Fig. 1). The results also showed that the higher the substrate concentration of BTEX, the longer the lag time and the greater the biomass (reflected by OD600). Since numerous enzymes were needed for bacteria to initiate oxidizing BTEX [29], the lag phase at high concentration of BTEX might be related to additional time required for induction of related enzymes.
The growth of EC20 at different concentrations of BTEX including 200 mg L−1 (▲) and 435 mg L−1 (◆), hollow symbols represent the unincubated controls on 200 mg L−1 (△) and 435 mg L−1 (◇) BTEX. Data are average of three replicates. Error bars represent standard deviation, some error bars are smaller than the symbols
Biodegradation of BTEX Compounds
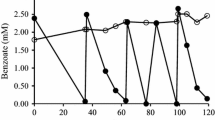
Figure 2 shows that the BTEX level in the culture was reduced to 50.4 mg L−1 from 435 mg L−1 (87.1% degradation) within 72 h. As to single components, degradation rates of benzene, toluene, ethylbenzene, and xylenes were 91.5, 72.9, 67, and 66.8% within 38 h, and 92.8, 85.7, 80.2, and 88.2% within 72 h, respectively. These results suggested significant biodegradation ability of EC20. A pure culture of Janibacter sp. SB2 isolated from a sea-tidal flat could degrade 45.5% BTEX after 60 h when the initial concentration was 240 mg L−1 [12]. Jesús et al. reported that BTEX at 200 mg L−1 was degraded by 50% within 36 h and 85% within 60 h by Pseudomonas FMB08 isolated from a microbial consortium, which could degrade 60% benzene, 78% toluene, 81.5% ethylbenzene, and 50% xylenes after 36 h when the initial concentration for each component was 50 mg L−1 [2]. The enrichment culture EC20 could degrade 87.1% of BTEX, although the initial concentration equaled to the BTEX level of the seriously contaminated area of Bohai Sea. The excellent BTEX-degrading capability and salinity adaptability of EC20 implied its great potential of bioremediation for the sea environment.
Biodegradation (×) of BTEX mixture and the removal rate of single BTEXs by EC20 at 435 mg L−1 BTEX containing equal proportions of each components: benzene, ■; toluene, ♦; ethylbenzene, ▲; xylenes, ●. Data are average of two replicates. Error bars represent standard deviation, some error bars are smaller than the symbols
Microbial Community Composition and Dynamics
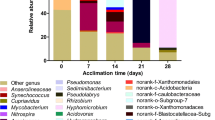
Previous studies suggested that BTEX could alter microbial community structure in the contaminated site [30, 31]. To investigate the stability of microbial community in the process of enrichment, EC18 and EC19 (subcultured for 18 and 19 generations, respectively) were also collected and compared with EC20 using DGGE based on 16S rRNA gene of bacteria. DGGE bands of EC18, EC19, and EC20 had similarities and differences both in quantities and location distribution (Fig. 3). The amounts of bands decreased with the increased times of subculturing, but the dominant bands were unchanged in all the enrichment cultures. Since microbial diversity was decreased and the growth of bacteria was inhibited on the condition of high concentration of petroleum hydrocarbons [32], it was reasonable that some bands existing in EC18 and EC19 disappeared in EC20. Ten bands of DGGE profiles were shared by EC18, EC19, and EC20, such as B2, B4, B6, B7, B12, B13, B16, B18, B20, and B25. Among those bands, six (B2, B4, B12, B13, B18, and B20) were always prominent in the process of enrichment with high average relative abundance (over 5%, Table S2), suggesting that the microorganisms represented by those bands might be responsible for the bioremediation of BTEX.
EC20 subcultured in liquid MSM supplied separately with only 50 mg L−1 of benzene, toluene, ethylbenzene, or xylenes (ECB, ECT, ECE, and ECX) showed slight differences from each other in DGGE patterns, but they still shared ten bands (B2, B4, B8, B9, B10, B12, B13, B16, B18, and B20) (Fig. 3). Interestingly, seven bands among the ten shared bands in EC18, EC19, and EC20 were also detected in ECB, ECT, ECE, and ECX, indicating that the bacterial community structure of EC20 was stabilized basically, and the predominant bacteria might play an important role in the biodegradation of all the BTEX compounds.
Phylogenetic analyses of the clone libraries of DGGE bands showed that the bacteria in enrichment cultures are mainly composed of four phyla, Proteobacteria, Actinobacteria, Bacteroidetes, and Firmicutes, representing 85.95, 10.15, 3.13, and 0.78%, respectively (Fig. 4 and Table S3), which contained many members capable of degrading oil hydrocarbons in a variety of environments [33–35]. At the genus level, the dominant microorganisms in enriched cultures were Pseudomonas, Mesorhizobium, Achromobacter, Stenotrophomonas, and Halomonas (Table S3). Members of the Pseudomonas assumed absolute superiority of the total clones, implying their vital role in degrading BTEX, since Pseudomonas strains possessed the ability to degrade a wide range of organic compounds such as alkane and polycyclic aromatic hydrocarbons (PAHs) [36–38]. Firouz et al. [39] reported that Pseudomonas putida owned metabolic versatility to different aromatic hydrocarbons, which was benefited from the plasmids containing arene degradation genes. A strain of Pseudomonas aeruginosa isolated from several areas of Hokkaido could remove 90–95% of total diesel oil [17]. Li et al. also showed that BTEX was degraded effectively by Pseudomonas plecoglossicida [40]. The genus Mesorhizobium also existed in abundance second only to Pseudomonas. Mesorhizobium were known as symbiotic nitrogen-fixing bacteria [41], and only a few researches proved their abilities for degradation of PAHs [42, 43]; however, the BTEX degradation capacity has not been reported previously. Achromobacter were widespread in freshwater, marine environment, and soil. Bacosa et al. [44] found that Achromobacter sp. strain was inhibited by p-xylene, but it could grow together with a p-xylene-degrading Alcaligenes sp. strain, suggesting that their isolates of Achromobacter may only utilize the intermediate products of aromatic hydrocarbons degraded by other microorganism. Stenotrophomonas were tolerant of heavy metals and widely distributed in water and soil [45]. Perhaps members of Stenotrophomonas in EC20 were guarantees of its survival when there was heavy metal pollution in the Bohai Sea. Halomonas were able to degrade PAHs, phenols, and salicylate in high salinity environment [46–49]; their presence in EC20 indicated that Halomonas might have the potential for BTEX degradation.
Phylogenetic tree of V3 region of 16S rRNA gene retrieved from DGGE bands. Sequences with 97% similarity are classified into an OTU (operational taxonomic unit). The tree was constructed by the neighbor-joining method with 1000 bootstrap replications using MEGA 6.0 software, bootstrap values over 50% are shown at the branch nodes. The scale bar represents 0.05 base changes per nucleotide position
Detection and Identification of Key Genes in BTEX-Degrading Pathway
In this analysis, five of the ten selected key catabolic genes, including tol, xylA, todC1, xylE1, and todE, were detected in the total genomic DNA of EC20 (Fig. 5). All of them shared high similarities with those reported in the genus Pseudomonas, consistent with the dominance of Pseudomonas (60.94%) revealed by the 16S rRNA gene analysis (Table S3). Tol and xylA coded the enzymes involved in the initial oxidation of side chain by the monooxygenases, and their existence suggested that TOL pathway was used to degrade BTEX in EC20. TodC1 encodes dioxygenase TODC1, which activates the benzene ring in the initial degradation; therefore, the EC20 consortium also contained TOD pathway. Also, EC20 carried xylE1 and todE for catechol 2,3-dioxygenases (C23O) to catalyze ring cleavage. Only Pseudomonas putida CM23 and Pseudomonas sp. KA were reported possessing two BTEX degradation pathways concurrently [50, 51], but it is common that different pathways presented simultaneously in bacterial consortia. Fathepure et al. [29] detected two pathways in an enriched culture from sediments of Great Salt Lake, and multiple pathways were discovered in a microbial enrichment from the East China Sea [11]. None of the benzene monooxygenases (TMOA and TBMD) was detected in the EC20 consortium, which might suggest the absence of TOM degradation pathway in EC20, but also might be caused by the low concentrations of their encoding genes in samples.
Phylogenetic trees of key catabolic genes detected in EC20 and reference sequences from GenBank. a tol. b xylA. c todC1. d xylE1. e todE. The clones are shown in bold, and accession numbers of all sequences are given in parentheses. The trees were constructed by neighbor-joining method with 1000 bootstrap replications using MEGA 6.0, bootstrap values over 50% are shown at the branch nodes. The scale bars indicating corresponding sequence divergences are shown in the lower-left corner of every tree
Conclusions
A microbial consortium EC20 was obtained from Bohai Sea sediment, which possessed excellent ability to degrade BTEX mixture at equivalent salinity to Bohai Sea, suggesting its bioaugmentation potential of BTEX at the contaminant sites. Pseudomonas, Mesorhizobium, Achromobacter, Stenotrophomonas, and Halomonas were identified as the dominant bacteria in EC20. Detection of the key catabolic enzyme-coding genes revealed that EC20 had TOL pathway and TOD pathway for the initial attack of BTEX. These results are significant for bioremediation of petroleum hydrocarbon pollution especially in the marine environment.
References
Doherty, V. F., & Otitoloju, A. A. (2016). Occurrence and distribution of monocyclic aromatic hydrocarbons (BTEX) and the impact on macrobenthic community structure in Lagos lagoon, Nigeria. Environmental Monitoring and Assessment, 188, 571.
Morlett-Chávez, J. A., Ascacio-Martínez, J. Á., Rivas-Estilla, A. M., Velázquez-Vadillo, J. F., Haskins, W. E., Barrera-Saldaña, H. A., & Acuña-Askar, K. (2010). Kinetics of BTEX biodegradation by a microbial consortium acclimatized to unleaded gasoline and bacterial strains isolated from it. Int. Biodeterior. Biodegradation, 64, 581–587.
Alfreider, A., & Vogt, C. (2007). Bacterial diversity and aerobic biodegradation potential in a BTEX-contaminated aquifer. Water, Air, and Soil Pollution, 183, 415–426.
Keith, L. H., & Telliard, W. A. (1979). Priority pollutants: I. A perspective view. Environmental Science & Technology, 13, 416–423.
Jiang, B., Zhou, Z., Dong, Y., Tao, W., Wang, B., Jiang, J., & Guan, X. (2015). Biodegradation of benzene, toluene, ethylbenzene, and o-, m-, and p-xylenes by the newly isolated bacterium Comamonas sp. JB. Appl. Biochem. Biotech., 176, 1700–1708.
Shim, H., Shin, E., & Yang, S. (2002). A continuous fibrous-bed bioreactor for BTEX biodegradation by a co-culture of Pseudomonas putida and Pseudomonas fluorescens. Advances in Environmental Research, 7, 203–216.
Zhou, Y., Huang, H., & Shen, D. (2016). Multi-substrate biodegradation interaction of 1, 4-dioxane and BTEX mixtures by Acinetobacter baumannii DD1. Biodegradation, 27, 37–46.
Avanzi, I. R., Gracioso, L. H., Baltazar, M. D. P. G., Karolski, B., Perpetuo, E. A., & Nascimento, C. A. O. (2015). Aerobic biodegradation of gasoline compounds by bacteria isolated from a hydrocarbon-contaminated soil. Environmental Engineering Science, 32, 990–997.
Mesgari Shadi, A., Yaghmaei, S., Vafaei, F., Khataee, A. R., & Hejazi, M. S. (2015). Degradation of benzene, toluene, and xylene (BTX) from aqueous solution by isolated bacteria from contaminated sites. Research on Chemical Intermediates, 41, 265–275.
Huang, Y., & Li, L. (2014). Biodegradation characteristics of naphthalene and benzene, toluene, ethyl benzene, and xylene (BTEX) by bacteria enriched from activated sludge. Water Environment Research, 86, 277–284.
Li, H., Zhang, Q., Wang, X., Ma, X., Lin, K., Liu, Y., Gu, J., Lu, S., Shi, L., Lu, Q., & Shen, T. (2012). Biodegradation of benzene homologues in contaminated sediment of the East China Sea. Bioresource Technology, 124, 129–136.
Jin, H. M., Choi, E. J., & Jeon, C. O. (2013). Isolation of a BTEX-degrading bacterium, Janibacter sp. SB2, from a sea-tidal flat and optimization of biodegradation conditions. Bioresource Technology, 145, 57–64.
Prenafeta-Boldu, F. X., Ballerstedt, H., Gerritse, J., & Grotenhuis, J. (2004). Bioremediation of BTEX hydrocarbons: effect of soil inoculation with the toluene-growing fungus Cladophialophora sp strain T1. Biodegradation, 15, 59–65.
Mukherjee, A. K., & Bordoloi, N. K. (2012). Biodegradation of benzene, toluene, and xylene (BTX) in liquid culture and in soil by Bacillus subtilis and Pseudomonas aeruginosa strains and a formulated bacterial consortium. Environmental Science and Pollution Research, 19, 3380–3388.
Vogel, T. M. (1996). Bioaugmentation as a soil bioremediation approach. Current Opinion in Biotechnology, 7, 311–316.
Daghio, M., Tatangelo, V., Franzetti, A., Gandolfi, I., Papacchini, M., Careghini, A., Sezenna, E., Saponaro, S., & Bestetti, G. (2015). Hydrocarbon degrading microbial communities in bench scale aerobic biobarriers for gasoline contaminated groundwater treatment. Chemosphere, 130, 34–39.
Wongsa, P., Tanaka, M., Ueno, A., Hasanuzzaman, M., Yumoto, I., & Okuyama, H. (2004). Isolation and characterization of novel strains of Pseudomonas aeruginosa and Serratia marcescens possessing high efficiency to degrade gasoline, kerosene, diesel oil, and lubricating oil. Current Microbiology, 49, 415–422.
Hendrickx, B., Dejonghe, W., Boenne, W., Brennerova, M., Cernik, M., Lederer, T., Bucheli-Witschel, M., Bastiaens, L., Verstraete, W., Top, E. M., Diels, L., & Springael, D. (2005). Dynamics of an oligotrophic bacterial aquifer community during contact with a groundwater plume contaminated with benzene toluene, ethylbenzene, and xylenes: an in situ mesocosm study. Appl. Environ. Microb., 71, 3815–3825.
Vandera, E., Samiotaki, M., Parapouli, M., Panayotou, G., & Koukkou, A. I. (2015). Comparative proteomic analysis of Arthrobacter phenanthrenivorans Sphe3 on phenanthrene, phthalate and glucose. Journal of Proteomics, 113, 73–89.
Lima-Morales, D., Jauregui, R., Camarinha-Silva, A., Geffers, R., Pieper, D. H., & Vilchez-Vargas, R. (2016). Linking microbial community and catabolic gene structures during the adaptation of three contaminated soils under continuous long-term pollutant stress. Applied and Environmental Microbiology, 82, 2227–2237.
Hendrickx, B., Junca, H., Vosahlova, J., Lindner, A., Ruegg, I., Bucheli-Witschel, M., Faber, F., Egli, T., Mau, M., Schlomann, M., Brennerova, M., Brenner, V., Pieper, D. H., Top, E. M., Dejonghe, W., Bastiaens, L., & Springael, D. (2006). Alternative primer sets for PCR detection of genotypes involved in bacterial aerobic BTEX degradation: distribution of the genes in BTEX degrading isolates and in subsurface soils of a BTEX contaminated industrial site. Journal of Microbiological Methods, 64, 250–265.
Fu, J., Ai, X., Liu, H., Han, D., Chen, D., & Ma, W. (2005). Determination of trace benzene, toluene, ethylbenzene and xylenes in seabed sediments and seawater by purge and trap gas chromatography. Chinese Journal of Analytical Chemistry, 33, 1753–1756.
Qin, J., Wang, S. Q., Sun, W. L., Yang, J. J., & Shen, B. (2013). BTEX content of marine surface sediments used as indicators of marine oil and gas. Rock & Mineral Analysis, 32, 785–790.
Chiu, H. Y., Hong, A., Lin, S. L., Surampalli, R. Y., & Kao, C. M. (2013). Application of natural attenuation for the control of petroleum hydrocarbon plume: mechanisms and effectiveness evaluation. Journal of Hydrology, 505, 126–137.
Yang, F. L., Yang, J. S., Deng, C. P., Chen, N., Wang, S. Q., Wang, E., & Yuan, H. L. (2015). Bacterial communities and their hydrocarbon bioremediation potential in the Bohai Sea. China. Mar. Ecol. Prog. Ser., 538, 117–130.
Rasheed, M. A., Patil, D. J., and Dayal, A. M. (2013) In Hydrocarbon in microbial techniques for hydrocarbon exploration 195–210.
Wang, S., Qin, J., Sun, W., Shen, B., Yang, J., & Yan, K. (2012). Determination of benzene series in soil and sediment with combined thermal desorption gas chromatography. Petroleum Geology & Experiment, 34, 94–98.
Wang, X., Wang, W., Gao, L., & Cui, Z. (2006). Protocols of application of denaturing gradient gel electrophoresis (DGGE) in studies of environmental microorganism. Journal of China Agricultural University, 11, 1–7.
Sei, A., & Fathepure, B. Z. (2009). Biodegradation of BTEX at high salinity by an enrichment culture from hypersaline sediments of Rozel Point at Great Salt Lake. Journal of Applied Microbiology, 107, 2001–2008.
El-Naas, M. H., Acio, J. A., & El Telib, A. E. (2014). Aerobic biodegradation of BTEX: progresses and prospects. Journal of Environmental Chemical Engineering, 2, 1104–1122.
Lin, C., Wu, C., Tang, C., & Chang, S. (2012). Novel oxygen-releasing immobilized cell beads for bioremediation of BTEX-contaminated water. Bioresource Technology, 124, 45–51.
Chiu, H. Y., Hong, A., Lin, S. L., Surampalli, R. Y., & Kao, C. M. (2013). Application of natural attenuation for the control of petroleum hydrocarbon plume: mechanisms and effectiveness evaluation. Journal of Hydrology, 505, 126–137.
Jiao, S., Liu, Z., Lin, Y., Yang, J., Chen, W., & Wei, G. (2016). Bacterial communities in oil contaminated soils: biogeography and co-occurrence patterns. Soil Biology and Biochemistry, 98, 64–73.
Yuan, J., Lai, Q., Sun, F., Zheng, T., & Shao, Z. (2015). The diversity of PAH-degrading bacteria in a deep-sea water column above the Southwest Indian Ridge. Frontiers in Microbiology, 6, 853.
Deng, C., Yu, X., Yang, J., Li, B., Sun, W., & Yuan, H. (2016). Universal indicators for oil and gas prospecting based on bacterial communities shaped by light-hydrocarbon microseepage in China. Journal of Microbiology and Biotechnology, 26, 1320–1332.
Wallisch, S., Gril, T., Dong, X., Welzl, G., Bruns, C., Heath, E., Engel, M., Suhadolc, M., & Schloter, M. (2014). Effects of different compost amendments on the abundance and composition of alkB harboring bacterial communities in a soil under industrial use contaminated with hydrocarbons. Frontiers in Microbiology, 5, 96.
Dong, C., Bai, X., Sheng, H., Jiao, L., Zhou, H., & Shao, Z. (2015). Distribution of PAHs and the PAH-degrading bacteria in the deep-sea sediments of the high-latitude Arctic Ocean. Biogeosciences, 12, 2163–2177.
Hemidouche, S., Favier, L., Sadaoui, Z., & Amrane, A. (2016). Degradation of clofibric acid by a phenol resistant Pseudomonas aeruginosa strain. Journal of Biotechnology, 231S, S71.
Abbasian, F., Lockington, R., Megharaj, M., & Naidu, R. (2016). A review on the genetics of aliphatic and aromatic hydrocarbon degradation. Applied Biochemistry and Biotechnology, 178, 224–250.
Li, J., de Toledo, R. A., Chung, J., & Shim, H. (2014). Removal of mixture of cis-1,2-dichloroethylene/trichloroe-thylene/benzene, toluene, ethylbenzene, and xylenes from contaminated soil by Pseudomonas plecoglossicida. Journal of Chemical Technology & Biotechnology, 89, 1934–1940.
Teixeira, H., & Rodríguez-Echeverría, S. (2016). Identification of symbiotic nitrogen-fixing bacteria from three African leguminous trees in Gorongosa National Park. Systematic and Applied Microbiology, 39, 350–358.
Jiménez, N., Viñas, M., Guiu-Aragonés, C., Bayona, J. M., Albaigés, J., & Solanas, A. M. (2011). Polyphasic approach for assessing changes in an autochthonous marine bacterial community in the presence of Prestige fuel oil and its biodegradation potential. Appl. Microbiol. Biot., 91, 823–834.
Zhang, S. Y., Wang, Q. F., Wan, R., & Xie, S. G. (2011). Changes in bacterial community of anthracene bioremediation in municipal solid waste composting soil. Journal of Zhejiang University-SCIENCE B, 12, 760–768.
Bacosa, H. P., Suto, K., & Inoue, C. (2012). Bacterial community dynamics during the preferential degradation of aromatic hydrocarbons by a microbial consortium. Int. Biodeterior. Biodegradation, 74, 109–115.
Berg, G., Roskot, N., & Smalla, K. (1999). Genotypic and phenotypic relationships between clinical and environmental isolates of Stenotrophomas maltophilia. Journal of Clinical Microbiology, 37, 3594–3600.
Castillo-Carvajal, L. C., Sanz-Martín, J. L., & Barragán-Huerta, B. E. (2014). Biodegradation of organic pollutants in saline wastewater by halophilic microorganisms: a review. Environmental Science and Pollution Research, 21, 9578–9588.
Dong, C., Bai, X., Sheng, H., Jiao, L., Zhou, H., & Shao, Z. (2014). Distribution of PAHs and the PAH-degrading bacteria in the deep-sea sediments of the high-latitude Arctic Ocean. Biogeosciences Discussions, 11, 13985–14021.
Gutierrez, T., Biddle, J. F., Teske, A., & Aitken, M. D. (2015). Cultivation-dependent and cultivation-independent characterization of hydrocarbon-degrading bacteria in Guaymas Basin sediments. Frontiers in Microbiology, 6, 695.
Oie, C. S. I., Albaugh, C. E., & Peyton, B. M. (2007). Benzoate and salicylate degradation by Halomonas campisalis, an alkaliphilic and moderately halophilic microorganism. Water Research, 41, 1235–1242.
Colombo, M., Dell'Amico, E., Cavalca, L., & Andreoni, V. (2004). Biodegradation of a BTX mixture by Pseudomonas strains: monitoring of two co-cultured strains by polymerase chain reaction of catabolic genes. Annals of Microbiology, 54, 381–392.
Di Martino, C., López, N. I., & Raiger Iustman, L. J. (2012). Isolation and characterization of benzene, toluene and xylene degrading Pseudomonas sp. selected as candidates for bioremediation. Int. Biodeterior. Biodegradation, 67, 15–20.
Acknowledgment
This work was funded by the National Natural Science Foundation of China (no. 31270533).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interests
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Deng, Y., Yang, F., Deng, C. et al. Biodegradation of BTEX Aromatics by a Haloduric Microbial Consortium Enriched from a Sediment of Bohai Sea, China. Appl Biochem Biotechnol 183, 893–905 (2017). https://doi.org/10.1007/s12010-017-2471-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-017-2471-y