Abstract
In this study, we cloned a full-length cDNA and the genomic DNA sequence of SmCCoAOMT (GenBank ID JQ007585) from Salvia miltiorrhiza. The 744-bp open-reading frame encodes a protein of 247 amino acids that shares 95 % similarity with one in Vitis vinifera. Real-time quantitative PCR analysis revealed that SmCCoAOMT is most highly expressed in the stems and can be induced by methyl jasmonate (MeJA) and XC-1 treatment. To evaluate its function in vivo, we generated RNA interference transgenic plants through Agrobacterium tumefaciens-mediated gene transfer. Compared with untransformed control plants, the transgenics had significantly less lignin and the expression of lignin-biosynthetic genes SmCCR and SmCOMT was depressed. In 90-day-old roots from plants of transgenic line M5, accumulations of rosmarinic acid and salvianolic acid B (Sal B) were greatly reduced by 0.89- and 0.69-fold, respectively. This low-Sal B phenotype was stable in the roots, with the level of accumulation being approximately 43.58 mg g−1 dry weight, which was 52 % of the amount measured in the untransformed control. Our results suggest that SmCCoAOMT is involved in lignin biosynthesis and affects the accumulation of phenolic acids. This study also provides potential guidance for using lignin-related genes to genetically engineer Salvia miltiorrhiza.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
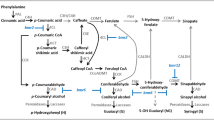
Lignin is a major component of secondary cell walls, providing mechanical support and also defending the plant against pathogen attack through increased lignification [1, 25]. Three types of lignin occur: guaiacyl (G), syringyl (S), and hydroxyphenyl (H) monolignol subunits, which are derived from p-coumaryl, coniferyl, and sinapyl alcohols [9]. The enzymes that catalyze lignin biosynthesis have been analyzed, and the genes involved in this biosynthetic pathway have been cloned and characterized in Arabidopsis thaliana, Nicotiana tabacum (tobacco), and Salvia miltiorrhiza [2, 6, 27].
Salvia miltiorrhiza is one of the most famous traditional Chinese medicines because its constituent phenolic acids (especially salvianolic acid B, or Sal B) are effective in the treatment of cardiovascular diseases [13, 40]. The biosynthesis of lignin and phenolic acids follows the same phenylpropanoid pathway, converting phenylalanine to 4-coumaric acid. In Salvia miltiorrhiza, suppression of cinnamoyl CoA reductase (CCR) and caffeic acid/5-hydroxyferulic acid O-methyltransferase (COMT) results in a reduction in lignin production while the concentrations of Sal B and another phenolic acid, rosmarinic acid (RA), are enhanced [34]. This process is consistent with that reported for other species [23, 31]. However, research with another lignin-related gene for caffeoyl-CoA O-methyltransferase (CCoAOMT) in Arabidopsis thaliana has shown that isorhamnetin is less abundant in the ccomt 1 mutant, suggesting that the gene is involved in the methylation of flavonoids and phenolic acids [4]. Thus, all of those study results strongly imply that the role of CCoAOMT in the production of lignin and phenolic acids differs from that of other lignin biosynthesis-related genes.
CCoAOMT is specific for the methylation of caffeoyl CoA, a key metabolite in the biosynthesis of guaiacyl lignin. Various CCoAOMTs have been cloned from Arabidopsis thaliana, tobacco, Zea mays, and Corchorus capsularis [3, 16, 37]. However, little is known about such genes in Salvia miltiorrhiza.
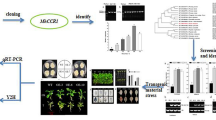
In this study, we isolated and characterized the expression of SmCCoAOMT from Salvia miltiorrhiza. To gain insight into lignin and phenolic biosynthesis in this species, we applied RNA interference (RNAi) technology to suppress expression of that gene. Our goal was to provide a new perspective that could enable the engineering of improved phenolic acid production in this medicinal plant.
Materials and Methods
Plant Material and Culture Conditions
The mature seeds of Salvia miltiorrhiza Bunge were obtained from Shangluo (Shaanxi Province) and cultured on an MS basal medium. Plants of Salvia miltiorrhiza were grown in the greenhouse at 25 ± 2 °C with a 16-h light/8-h dark photoperiod provided by cool-white fluorescent lamps (25 μmol m−2 s−1).
DNA and RNA Isolations
Genomic DNA and total RNA were extracted by the CTAB method [5] and Trizol reagent (BioFlux, China), respectively. cDNA was synthesized by reverse-transcription with oligo-dT primers (Takara, Japan).
Cloning of SmCCoAOMT and Sequence Analysis
The full-length sequences of DNA and RNA for SmCCoAOMT were obtained from RNA-seq through PCR [12]. The 5′ sequence end was cloned by DNA walking technology using a kit (no. 636405, Clontech, USA) (Table 1). PCR was performed as follows: 95 °C for 5 min; 34 cycles at 95 °C for 30 s, 52 °C for 30 s, and 72 °C for 2 min with a final extension step of 72 °C for 10 min. Amplified fragments were then purified and sequenced (Takara, Japan). Similarity search of SmCCoAOMT were performed by BLASTP (http://www.ncbi.nlm.nih.gov/BLAST/). Multiple-alignment analyses were conducted using ClustalW (http://www.ebi.ac.uk/Tools/clustalw/index.html), and a neighbor-joining tree was built using PhyML [10]. Promoter was analyzed according to the PlantCARE database (http://bioinformatics.psb.ugent.be/webtools/plantcare.html).
Expression Analysis and Quantitative Real-Time PCR
Two-month-old plants were treated with either methyl jasmonate (MeJA, 5 μM) or the bacterium Xanthomonas campestris pv. campestris (XC-1). The growing conditions for XC-1 were previously described [18]. The expression of SmCCoAOMT was analyzed by real-time quantitative PCR, with glyceraldehyde-3-phosphate dehydrogenase (GAPDH) being amplified as a control (Table 1). PCR condition in the iQ5 thermocycler (Bio-Rad, USA) was as follows: 94 °C for 1 min, followed by 35 cycles at of 94 °C for 10 s, 60 °C for 20 s, and 82 °C for 15 s of collection fluorescence. Expression was quantified by the comparative CT method [24].
RNAi Construct and Plant Transformation
A 275-bp fragment corresponding to the 3′ region of SmCCoAOMT cDNA was amplified by PCR, using upstream primer 5′-CCATCGATGGTACCCCTTGCAGATGGGTTCTCATACACCAG-3′ (sites underlined for Cla I and Kpn I, respectively) plus downstream primer 5′-CGGGATCCTCGAGGCTGGGCTGATGAGGAATACTGCAAG-3′ (sites underlined for BamH I and Xho I, respectively). The SmCCoAOMT fragment was inserted upstream and downstream of the intron in the pKANNIBAL vector. The recombinant vector and pART27 [8] were ligated to generate the pART27-CCoAOMTi vector, and then introduced into Agrobacterium tumefaciens strain EHA105 [11].
Gene transfer was performed as described by Wang and Wang [33, 34]. To test for the putative transgenic plants, we amplified a 427-bp fragment from the intron, using primer PDK F (5′-GTGATGTGTAAGACGAAGAAG-3′) and PDK R (5′-GATAGATCTTGCGCTTTG-3′). The plasmid pART27-CCoAOMTi was used as positive control. The PCR-positive lines were used for further analysis.
Lignin Analysis
Air-dried stems of transgenic and wild-type (WT) control plants were used for lignin analysis after being grown under the same conditions for 180 days. The stems were exhaustively extracted for 24 h with methanol and acetone separately in a Soxhlet apparatus. The main G and S lignin monomers (269 m/z for G and 299 m/z for S) were analyzed via gas chromatography–mass spectrometry (GC–MS; QP2010; Shimadzu, Japan) [38, 39]. Separation was achieved on an RTX-5MS column (30 m × 250 μm, 0.25 μm, Agilent J&W Scientific), with helium as the carrier gas at a constant flow rate of 1.19 mL/min. The temperature of injection, transfer interface, and ion source was set to 250, 250, and 200 °C, respectively. The GC temperature programming was set at 50 °C, followed by 35 °C/min oven temperature ramps to 220 °C, 0.5 °C/min to 230 °C, and 50 °C/min to 280 °C and a final 7-min maintenance at 280 °C. For relative quantification, peak areas from selected unique fragment ions for every identified compound were used. Levels of Klason lignin (acid-insoluble) and acid-soluble lignin were measured based on previous methods [38, 39].
Extraction of Phenolic Acids and HPLC Analysis
Roots were collected from plants grown in the greenhouse for 90 or 180 days. Briefly, 20 mg of air-dried sample tissue was extracted with 75 % methanol (1 mL). Extractions of the phenolic compounds and analysis via high-performance liquid chromatography (HPLC; LC-2010A; Shimadzu) were performed as described by Zhang and Wang [34, 39].
Statistical Analysis
Significant differences among results were evaluated by Tukey’s pairwise comparison tests with SPSS 19.0 software. Results were considered to be statistically significant when p < 0.05.
Results
Isolation and Characterization of CCoAOMT
Our PCR-based EST library and transcriptome analysis produced a 1037-bp sequence that we designated Salvia miltiorrhiza CCoAOMT. The full-length SmCCoAOMT was assigned with GenBank accession number JQ007585 (2783 bp). The open-reading frame was 744 bp long and the gene contained five exons and four introns in sizes of 549 bp (Fig. 1a).
a Structure of SmCCoAOMT. Exons are denoted by black boxes; introns, by lines. Lengths of exons are shown in base pairs. b Protein structure diagram: DEFGH conserved domains for Mg2+ and substrate binding; ABC conserved domains for methyl donor. c Phylogenetic tree of SmCCoAOMT and nine OMTs characterized from dicots and monocots. The tree was built using PhyML and neighbor-joining tree, with MEGA4. Accession numbers of sequences used for constructing tree are as follows: Arabidopsis thaliana, AtCCoAOMT (NP_974104.1); Arabidopsis thaliana, AtCOMT (AT5G54160); Carthamus tinctorius, CtCCoAOMT (BAG71891.1); Glycine max, GmCCoAOMT (XP_003518701); Gossypium hirsutum, GhCCoAOMT (ACQ59095); Nicotiana tabacum, NtCCoAOMT (Q42945.1); Salvia miltiorrhiza, SmCOMT (JF693491); Vitis vinifera, VvCCoAOMT (XP_002282867); and Zea mays, ZmCCoAOMT ( AAP37884.1)
The BLASTP analysis showed that the predicted protein shared high sequence identity with other CCoAOMTs, such as 95 % similarity with the protein from Vitis vinifera. The motifs of homologous CCoAOMT protein sequences were highly conserved and a comparison of active site residues presented five conserved domains and three catalytic domains (Fig. 1b). Molecular phylogenetic analysis demonstrated that SmCCoAOMT is very closely related to CCoAOMTs, including Vitis vinifera and Carthamus tinctorius. However, SmCCoAOMT and SmCOMT occupied separate categories, implying that they were involved in different methylation reactions (Fig. 1c).
Using genome walking, we obtained a 1320-bp 5′ flanking sequence for SmCCoAOMT. Promoter analysis revealed several cis-acting regulator elements involved in light-responsiveness, including 4cl-CMA2b, an AE-box, ATCT-motif, G-box, L-box, and Sp1. The 5′ flanking regions also contained an AuxRR-box for auxin-responsiveness, plus TC-rich repeats that participate in defense and stress responses.
Analysis of CCoAOMT Expression
To gain further information about the spatiotemporal transcript abundance of CCoAOMT in Salvia miltiorrhiza, we performed tissue- and age-specific profiling via real-time quantitative PCR. Overall, SmCCoAOMT was expressed predominantly in the stems and at different growth stages. Expression was also high in the flowers (Fig. 2a). When compared with the untreated control, transcript levels were 11-fold higher in plants exposed to MeJA (Fig. 2b). Upon treatment with the XC-1 bacterium, transcription decreased in the first 6 h before rising at 24 and 48 h (Fig. 2c). This suggested that SmCCoAOMT participates in the plant defense mechanism.
Expression patterns of SmCCoAOMT in different organs (a) and under treatment with MeJA (b) or XC-1 (c). (1) Overwintering Salvia miltiorrhiza plant and (2) 2-month-old plants. Data represent averages of three experiments and error bars represent standard deviations. Asterisk represents values significantly different from untreated control at p < 0.05
Development of RNAi Lines in Salvia miltiorrhiza
To understand the function of SmCCoAOMT in lignin biosynthesis, we constructed an RNAi vector that produced dsRNA under the control of the cauliflower mosaic virus (CaMV) 35S promoter (Fig. 3a). The Agrobacterium-transformed plants were cultured on a kanamycin medium, and 11 independent transgenic lines were selected. Leaves from 1-month-old plants were used for PCR analysis. An amplified 427-bp fragment of the intron confirmed that the transgenics from eight of those lines were positive (Fig. 3b, c). Expression of SmCCoAOMT was significantly decreased in all transgenic lines, especially lines M4 and M5, when compared with the control. However, no differences in phenotype were found among any of those lines (data not shown).
Vector construct and molecular analysis of transgenic plants. a PCR screening of SmCCOAoMT-suppressed Salvia miltiorrhiza. PDK intron was amplified to screen positive transgenic lines. Lane M, DL2000 DNA marker. Band sizes from top: 250, 500, 750, 1000, and 2000 bp; − control lines, B blank vector (without fragment of SmCCoAOMT), + positive control; lanes 1–11, different transgenic lines; C no-template control. b Sketch map of interfering box, including restriction sites and target gene. c Real-time quantitative PCR analysis of SmCCOAoMT in RNAi Salvia miltiorrhiza. All data are means of three replicates, with error bars showing standard deviations. Asterisk represents values significantly different from untransformed control at p < 0.05
Impact on Lignin Contents and Compositions
Our examination of the RNAi-transgenic lines and the control revealed changes in lignin levels and compositions (i.e., the relative proportions of S- and G-type monomers) (Table 2). Both the yield and relative frequency of the G monomers were reduced in the RNAi-transgenic stems while the S-lignin units were significantly higher. Consequently, the S/G thioacidolysis ratio was substantially higher in the transgenic tissues (0.89 for line M5 versus 0.69 for control). Finally, the contents of total lignin were decreased by 18.9 and 24.8 % in lines M4 and M5, respectively, when compared with the control plants (Table 2).
Effect of CCoAOMT Inhibition on the Expression of COMT and CCR
The functional consequences of altering the expression of CCoAOMT was tested by measuring transcript levels of two genes related to the lignin-biosynthetic pathway in transgenic Salvia miltiorrhiza. Compared with activity in control plants, expression of SmCOMT and SmCCR was markedly reduced in the transgenics (Fig. 4). That pattern was consistent with the trend found for lignin contents.
Impact on Phenolic Acids
In addition to its established role in lignin formation, suppression of CCoAOMT can affect the metabolism of other participants in the phenylpropanoid pathway. To investigate whether such metabolism was modified in the RNAi-transgenic plants, we extracted their phenolic acids and separated them via HPLC. Levels of RA and Sal B, the two major hydrophilic and active pharmaceutical ingredients in Salvia miltiorrhiza, were decreased to varying degrees in 90-day-old plants from lines M4 and M5. Concentrations were lowest in the latter, i.e., 8.73 ± 1.23 mg g−1 dry weight (DW) for RA and 38.90 ± 0.72 mg g−1 DW for Sal B. By comparison, those levels in control plants were 9.88 ± 0.78 mg g−1 DW for RA and 55.60 ± 2.31 mg g−1 DW for Sal B (Fig. 5a). To determine whether this low-RA/low-Sal B phenotype was stable during the stages of vegetative development, we examined the roots from genotypes after 180 days of growth and found that levels of RA and Sal B were further decreased by 34 and 47 %, respectively, in line M5 when compared with the control (Fig. 5b).
Discussion
We have cloned a lignin-biosynthetic gene, SmCCoAOMT, from Salvia miltiorrhiza and evaluated its functioning in vivo. Cluster and conserved domain analyses indicated that roles vary between CCoAOMT and COMT genes, confirming that their roles differ for catalytic methylation during lignin biosynthesis in this species [14]. The expression pattern of SmCCoAOMT demonstrated that lignification was more active during that stage of development, with changes in transcripts following a pattern similar to that described for other CCoAOMTs [7, 16, 19]. We found it noteworthy that exposure to MeJA or XC-1 treatments enhanced the expression of CCoAOMT, thereby suggesting that lignin production is affected by signal transduction and plant biodefenses, consistent with results from promoter analysis. Similar findings have been reported for tobacco [16].
As observed previously with cinnamyl alcohol dehydrogenase (CAD), CCR, COMT, and phenylalanine ammonia lyase (PAL) [22, 30], lignin concentrations were reduced and compositions were altered in transgenic lines [29, 34]. However, a comparison of RNAi-SmCCoAOMT lines and control plants revealed no differences in phenotype. This is in contrast to the dwarfing phenotypes manifested by transgenic lines of SmCCR [34], indicating that SmCCoAOMT is not associated with lodging resistance [37]. In parallel, our investigation showed that downregulation of SmCCoAOMT led to decreased amounts of total lignin but an increase in the S/G ratio, which is consistent with previous study results [21, 32].
In contrast to other lignin-related genes in Salvia miltiorrhiza [4, 15], downregulation of CCoAOMT significantly decreased the accumulation of phenolic acids (RA and Sal B) in the transgenic plants, evidence that precursors not used in the lignin pathway were not reallocated. Those results from our experiments with RNAi-SmCCoAOMT are of interest because similar effects have been reported for other species [17]. One possible explanation is that SmCCoAOMT participates in the biosynthesis of phenolic acids by influencing the process of methylation. However, different genes in the lignin-biosynthetic pathway are not necessarily suppressed, implying that another potential metabolic branch may occur in the phenylpropanoid pathway [26, 28]. That aspect was not focused on in this current examination of Salvia miltiorrhiza, but the mechanism by which this phenomenon of lignin and phenolic acid deposition occurs will require further investigation.
Using genetic engineering to improve the production of effective components is becoming a new research goal with traditional Chinese medicines [35, 36, 39]. For example, some researchers have demonstrated that decreasing the flux through competitive pathways is an efficient strategy for increasing the levels of plant secondary metabolites [20]. Likewise, the synthesis of phenolic acids has been promoted by downregulating the expression of lignin synthesis-related genes in Salvia miltiorrhiza [34, 39]. Based on our studies, we propose that the functions of those genes should be comprehensively evaluated before taking this approach because blind application of similar strategies may lead to unexpected results. Nevertheless, our findings provide new insight into how SmCCoAOMT participates in the formation of health-promoting phenolic acids.
In summary, we cloned and characterized SmCCoAOMT from Salvia miltiorrhiza and analyzed its role in lignin biosynthesis. Our data showed that this gene influences the accumulation of phenolic acids, especially Sal B. Therefore, we believe that CCoAOMT is a candidate gene for genetically engineering plants of this species to enhance their production of valuable phenolic acids.
References
Bi, C., Chen, F., Jackson, L., Gill, B. S., & Li, W. (2010). Expression of lignin biosynthetic genes in wheat during development and upon infection by fungal pathogens. Plant Molecular Biology Reporter, 29, 149–161.
Bo, W., Wei, S., Li, Q., Ying, L., Luo, H., Song, J., Chao, S., Qian, J., Zhu, Y., & Hayward, A. (2014). Genome-wide identification of phenolic acid biosynthetic genes in Salvia miltiorrhiza. Planta, 241, 711–725.
Chen, Y., Zein, I., Brenner, E. A., Andersen, J. R., Landbeck, M., Ouzunova, M., & Lübberstedt, T. (2010). Polymorphisms in monolignol biosynthetic genes are associated with biomass yield and agronomic traits in European maize (Zea mays L. BMC Plant Biology, 10, 12.
Christin, F., Maike, V. O., Vinzenz, H., & Thomas, V. (2012). The role of CCoAOMT1 and COMT1 in Arabidopsis anthers. Planta, 236, 51–61.
Doyle, J., & Doyle, J. (1986). A rapid DNA isolation procedure from small quantities of fresh leaf tissues. Phytochemistry, 19, 11–15.
Ehlting, J., Mattheus, N., Aeschliman, D. S., Li, E., Hamberger, B., Cullis, I. F., Zhuang, J., Kaneda, M., Mansfield, S. D., & Samuels, L. (2005). Global transcript profiling of primary stems from Arabidopsis thaliana identifies candidate genes for missing links in lignin biosynthesis and transcriptional regulators of fiber differentiation. Plant Journal, 42, 618–640.
Fossdal, C. G., Nagy, N. E., Hietala, A. M., Kvaalen, H., Slimestad, R., Woodward, S., & Solheim, H. (2012). Indications of heightened constitutive or primed host response affecting the lignin pathway transcripts and phenolics in mature Norway spruce clones. Tree Physiology, 32, 1137–1147.
Gleave, A. P. (1993). A versatile binary vector system with a T-DNA organizational-structure conducive to efficient integration of cloned DNA into the plant genome. Plant Molecular Biology, 20, 1203–1207.
Grabber, J. H., Ralph, J., Hatfield, R. D., & Quideau, S. (1997). P-hydroxyphenyl, guaiacyl, and syringyl lignins have similar inhibitory effects on wall degradability. Journal of Agricultural & Food Chemistry, 45, 2530–2532.
Guindon, S., & Gascuel, O. (2003). PhyML—a simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Systematic Biology, 52, 696–704.
Holsters, M., Waele, D. D., Depicker, A., Messens, E., Montagu, M. V., & Schell, J. (1978). Transfection and transformation of Agrobacterium tumefaciens. Molecular & General Genetics, 163, 181–187.
Hua, W., Zhang, Y., Song, J., Zhao, L., & Wang, Z. (2011). De novo transcriptome sequencing in Salvia miltiorrhiza to identify genes involved in the biosynthesis of active ingredients. Genomics, 98, 272–279.
Lei, X., & Chiou, G. C. Y. (2012). Studies on cardiovascular actions of Salvia miltiorrhiza. American Journal of Chinese Medicine, 14, 26–32.
Li, L., Zhou, Y., Cheng, X., Sun, J., Marita, J. M., Ralph, J., & Chiang, V. L. (2003). Combinatorial modification of multiple lignin traits in trees through multigene cotransformation. Proceedings of the National Academy of Sciences of the United States of America, 100, 4939–4944.
Li, X., Chen, W., Zhao, Y., Xiang, Y., Jiang, H., Zhu, S., & Cheng, B. (2013). Downregulation of caffeoyl-CoA O-methyltransferase (CCoAOMT) by RNA interference leads to reduced lignin production in maize straw. Genetics & Molecular Biology, 36, 540–546.
Martz, F., Maury, S., Pinçon, G., & Legrand, M. (1998). cDNA cloning, substrate specificity and expression study of tobacco caffeoyl-CoA 3-O-methyltransferase, a lignin biosynthetic enzyme. Plant Molecular Biology, 36, 427–437.
Meyermans, H., Morreel, K., Lapierre, C., Pollet, B., Bruyn, A., De Busson, R., Herdewijn, P., Devreese, B., Beeumen, J., Van Marita, J. M., John, R., Cuiying, C., Bart, B., Marc, V. M., Eric, M., & Wout, B. (2000). Modifications in lignin and accumulation of phenolic glucosides in poplar xylem upon down-regulation of caffeoyl-coenzyme A O-methyltransferase, an enzyme involved in lignin biosynthesis. Journal of Biological Chemistry, 275, 36899–36909.
Miguens, F. C. (1993). Identification of compatible and incompatible interactions between Arabidopsis thaliana and Xanthomonas campestris pv. campestris and characterization of the hypersensitive response. Journal of Crystal Growth, 310, 4451–4455.
Negi, S., Tak, H., & Ganapathi, T. R. (2015). In vitro xylem vessel elements formation from banana embryogenic cells and expression analysis of vessel development-related genes. Plant Biotechnology Reports, 9, 47–54.
Oksman-Caldentey, K. M., & Inze, D. (2004). Plant cell factories in the post-genomic era: new ways to produce designer secondary metabolites. Trends in Plant Science, 9, 433–440.
Pang, S. L., Ong, S. S., Lee, H. H., Zamri, Z., Kandasamy, K. I., Choong, C. Y., & Wickneswari, R. (2014). Isolation and characterization of CCoAOMT in interspecific hybrid of Acacia auriculiformis x Acacia mangium—a key gene in lignin biosynthesis. Genetics and Molecular Research, 13, 7217–7238.
Rahantamalala, A., Rech, P., Martinez, Y., Chaubet-Gigot, N., Grima-Pettenati, J., & Pacquit, V. (2010). Coordinated transcriptional regulation of two key genes in the lignin branch pathway—CAD and CCR—is mediated through MYB-binding sites. BMC Plant Biology, 10, 107–113.
Rest, B. V. D., Danoun, S., Boudet, A. M., & Rochange, S. F. (2006). Down-regulation of cinnamoyl-CoA reductase in tomato (Solanum lycopersicum L.) induces dramatic changes in soluble phenolic pools. Journal of Experimental Botany, 57, 1399–1411.
Schmittgen, T. D., & Livak, K. J. (2008). Analyzing real-time PCR data by the comparative CT method. Nature Protocols, 3, 1101–1108.
Senthil-Kumar, M., Hema, R., Suryachandra, T. R., Ramegowda, H. V., Gopalakrishna, R., Rama, N., Udayakumar, M., & Mysore, K. S. (2010). Functional characterization of three water deficit stress-induced genes in tobacco and Arabidopsis: an approach based on gene down regulation. Plant Physiology and Biochemistry, 48, 35–44.
Seok, B., Kim, B. G., Na, R. J., Lee, Y., Lim, Y., Chong, Y., & Ahn, J. H. (2009). Structural modeling and biochemical characterization of flavonoid O-methyltransferase from rice. Bulletin of the Korean Chemical Society, 30, 2803–2805.
Sewalt, V., Ni, W., Blount, J. W., Jung, H. G., Masoud, S. A., Howles, P. A., Lamb, C., & Dixon, R. A. (1997). Reduced lignin content and altered lignin composition in transgenic tobacco down-regulated in expression of L-phenylalanine ammonia-lyase or cinnamate 4-hydroxylase. Plant Physiology, 115, 41–50.
Shaipulah, N. F. M., Muhlemann, J. K., Woodworth, B. D., Moerkercke, A. V., Verdonk, J. C., Ramirez, A. M., Haring, M. A., Dudareva, N., & Schuurink, R. (2015). CCoAOMT downregulation activates anthocyanin biosynthesis in petunia. Plant Physiology, 170.
Song, J., & Wang, Z. (2011). RNAi-mediated suppression of the phenylalanine ammonia-lyase gene in Salvia miltiorrhiza causes abnormal phenotypes and a reduction in rosmarinic acid biosynthesis. Journal of Plant Research, 124, 183–192.
Thévenin, J., Pollet, B., Letarnec, B., Saulnier, L., Gissot, L., Maia-Grondard, A., Lapierre, C., & Jouanina, L. (2011). The simultaneous repression of CCR and CAD, two enzymes of the lignin biosynthetic pathway, results in sterility and dwarfism in Arabidopsis thaliana. Molecular Plant, 4, 70–82.
Tschaplinski, T. J., Standaert, R. F., Engle, N. L., Martin, M. Z., Sangha, A. K., Parks, J. M., Smith, J. C., Samuel, R., Nan, J., & Pu, Y. (2012). Down-regulation of the caffeic acid O-methyltransferase gene in switchgrass reveals a novel monolignol analog. Biotechnology for Biofuels, 5, 1–15.
Wagner, A., Tobimatsu, Y., Phillips, L., Flint, H., Torr, K., Donaldson, L., Pears, L., & Ralph, J. (2011). CCoAOMT suppression modifies lignin composition in Pinus radiata. The Plant Journal, 67, 119–129.
Wang, D., Yao, W., Song, Y., Liu, W., & Wang, Z. (2012). Molecular characterization and expression of three galactinol synthase genes that confer stress tolerance in Salvia miltiorrhiza. Journal of Plant Physiology, 169, 1838–1848.
Wang, Z., Cui, L., Chen, C., Liu, X., Yan, Y., & Wang, Z. (2012). Downregulation of cinnamoyl CoA reductase affects lignin and phenolic acids biosynthesis in Salvia miltiorrhiza Bunge. Plant Molecular Biology Reporter, 30, 1229–1236.
Wilson, S. A., & Roberts, S. C. (2014). Metabolic engineering approaches for production of biochemicals in food and medicinal plants. Current Opinion in Biotechnology, 26, 174–182.
Yan, Q., Shi, M., Ng, J., & Wu, J. Y. (2006). Elicitor-induced rosmarinic acid accumulation and secondary metabolism enzyme activities in Salvia miltiorrhiza hairy roots. Plant Science, 170, 853–858.
Zhang, G., Zhang, Y., Xu, J., Niu, X., Qi, J., Tao, A., Zhang, L., Fang, P., Lin, L. H., & Su, J. (2014). The CCoAOMT1 gene from jute (Corchorus capsularis L.) is involved in lignin biosynthesis in Arabidopsis thaliana. Gene, 546, 398–402.
Zhang, Y., Yan, Y., & Wang, Z. (2010). The Arabidopsis PAP1 transcription factor plays an important role in the enrichment of phenolic acids in Salvia miltiorrhiza. Journal of Agricultural & Food Chemistry, 58, 565–574.
Zhang, Y., Yan, Y., Wu, Y., Hua, W., Chen, C., Ge, Q., & Wang, Z. (2013). Pathway engineering for phenolic acid accumulations in Salvia miltiorrhiza by combinational genetic manipulation. Metabolic Engineering, 21, 71–80.
Zhou, Y., & Fu, R. (2015). Continuing treatment with Salvia miltiorrhiza injection attenuates myocardial fibrosis in chronic iron-overloaded mice. PloS One, 10, 659–663.
Acknowledgments
This work was supported by the Natural Science Foundation of Shaanxi Province, China (2012JQ4013) and by the National Natural Science Foundation of China (Grant No. 31300256).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, Z., Ge, Q., Chen, C. et al. Function Analysis of Caffeoyl-CoA O-Methyltransferase for Biosynthesis of Lignin and Phenolic Acid in Salvia miltiorrhiza . Appl Biochem Biotechnol 181, 562–572 (2017). https://doi.org/10.1007/s12010-016-2231-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-016-2231-4