Abstract
The sucrose isomerase (SIase) gene from an efficient strain of Erwinia rhapontici NX-5 for isomaltulose hyperproduction was cloned and overexpressed in Escherichia coli. Protein sequence alignment revealed that SIase was a member of the glycoside hydrolase 13 family. The molecular mass of the purified recombinant protein was estimated at 66 kDa by SDS-PAGE. The SIase had an optimal pH and temperature of 5.0 and 30 °C, respectively, with a K m of 257 mmol/l and V max of 48.09 μmol/l/s for sucrose. To the best of our knowledge, the recombinant SIase has the most acidic optimum pH for isomaltulose synthesis. When the recombinant E. coli (pET22b- palI) cells were used for isomaltulose synthesis, almost complete conversion of sucrose (550 g/l solution) to isomaltulose was achieved in 1.5 h with high isomaltulose yields (87%). The immobilized E. coli cells remained stable for more than 30 days in a “batch”-type enzyme reactor. This indicated that the recombinant SIase could continuously and efficiently produce isomaltulose.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Isomaltulose (commonly referred to as palatinose), a structural isomer of sucrose, has been suggested as a non-cariogenic alternative to sucrose and is widely used in health products and in the food industry [1–3]. It is produced from sucrose by bacteria that contain an enzyme designated sucrose isomerase (SIase) or isomaltulose synthase. SIase activity has been reported in a wide range of bacterial species, and in addition to catalyzing the isomerization of sucrose to isomaltulose, it also produces another sucrose isomer, trehalulose, as well as small amounts of glucose and fructose as byproducts [4–9]. The ratios of these enzymatic products vary from mainly isomaltulose (75–85%) to predominantly trehalulose (−90%), depending on the bacterial strain [10–12] and the reaction conditions. A small quantity of other products, such as isomaltose and isomelezitose, may be formed under certain conditions [13].
SIase has been purified and characterized from microorganisms, including Erwinia rhapontici NCPPB 1578, Serratia plymuthica ATCC 15928, Pantoea dispersa UQ68J, Klebsiella sp., Enterobacter sp. FMB-1, Pseudomonas mesoacidophila MX-45 and also from a species of whitefly [4, 6–10, 12, 13]. Although a number of the corresponding genes derived from different microorganisms have been cloned and expressed, the exception has been the SIase gene from E. rhapontici WAC2928, which was cloned in Escherichia coli but showed unstable expression [12]. The SIases from Klebsiella sp. LX3, P. mesoacidophila MX-45 and Protaminobacter rubrum are currently the only three with available crystal structural information and these show a (β/α)8 catalytic domain at N-terminal with highly conserved catalytic residues. The catalytic mechanism of SIase from P. mesoacidophila MX-45 and P. rubrum have also been studied in recent years [8, 14–16].
The conversion of sucrose into isomaltulose has been accomplished with both free and immobilized cells. However, for the industrial production of isomaltulose, major factors to consider are the ratio of final products and the stability of the immobilized cells. In addition, there are still issues with the use of immobilized cells, such as slow conversion, low productivity, which limits the industrial efficiency of isomaltulose production [17–20]. We have recently isolated an isomaltulose-producing strain E. rhapontici NX-5 that shows a high initial SIase activity. In the present paper, the gene encoding the SIase of E. rhapontici NX-5 was cloned and expressed in E. coli. The recombinant enzyme was purified and characterized and demonstrated a continuous and efficient production of isomaltulose.
Materials and Methods
Materials
Reagents for the polymerase chain reaction (PCR), Ex-TaqDNA polymerase, T4 DNA ligase and restriction enzymes were purchased from TAKARA. Plasmids of pUC18 and pET22b were obtained from Novagen Co., USA. Genomic DNA extraction kit, plasmid isolation kits, and nickel–nitrilo triacetic acid (Ni–NTA) superflow columns for purification were from Qiagen (Hilden, Germany). Oligonucleotide primers were obtained from Jinsiter Co. (China). Electrophoresis reagents were from Bio-Rad.
Bacterial Strains, Plasmids, Growth Media, and Culture Condition
E. rhapontici NX-5, an isomaltulose-producing strain, was isolated from soil in China and used for cloning the sucrose isomerase gene (palI). It was deposited in the China General Microbiological Culture Collection Center with accession number CGMCC No.2222 [21]. E. rhapontici NX-5 was inoculated into 50 ml culture medium containing sucrose 50 g/l, yeast extract 10 g/l, and Na2HPO4·12 H2O 5 g/l in a 500 ml flask and aerobically incubated at 30 °C for 12 h with shaking at 200 r/min. E. coli DH5α and E. coli BL21(DE3) were used as the host strains for cloning and expression of palI, respectively. Both strains were routinely grown overnight in Luria–Bertani (LB) medium. Ampicillin was added at 100 μg/ml when required.
DNA Isolation and Cloning of the SIase Gene
The palI gene was amplified by PCR from E. rhapontici NX-5 using two oligonucleotide primers, 5′-TTAAGCTT CCATGGATTCTCAAGGATT -3′ (HindIII and Nco I restriction sites are underlined) and 5′-TTGGATCC CTCGAGCGGATTAAGTTTATAAATG-3′ (BamHI and XhoI restriction sites are underlined). HindIII and BamHI sites were incorporated in the forward and reverse primers for cloning into vector pUC18. The amplified fragment was inserted into pUC18 digested by HindIII and BamHI, generating pUC18-palI for analysis of the nucleotide sequence. The pUC18-palI and pET22b were then digested with restriction enzymes XhoI and NcoI. The palI gene released from pUC18-palI vector was ligated with pET22b to give pET22b-palI, which was under the control of the T7 promoter. SIase was expressed as a protein fused to the N terminus of a 6 His-tag. The recombinant plasmid was then transformed into E. coli BL21 (DE3).
Expression of SIase in E. Coli
E. coli BL21 (DE3) harboring pET22b-palI was grown at 37 °C in LB medium containing ampicillin at 100 μg/ml. The expression of recombinant enzyme was performed using 0.5 mmol/l lactose. The cells were incubated for an additional 11 h at 24 °C and harvested by centrifugation (10,000×g, 15 min, 4 °C). The level of SIase expression was determined by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE).
Purification of the Recombinant SIase
Recombinant E. coli BL21(DE3) cells were collected and washed twice with 50 ml of 0.85% NaCl solution. Harvested cells were suspended in 10 ml sodium phosphate buffer (PBS, 25 mmol/l, pH 6.0), and then disrupted by sonication at 4 °C for 15 min. The cell debris was removed by centrifugation (10,000×g, 15 min, 4 °C). The resulting crude extract was retained for purification. The cell-free extract was applied onto a Ni–NTA superflow column (5 cm, Qiagen) which was previously equilibrated with a binding buffer (50 mmol/l PBS, pH 6.0). Unbound proteins were washed out from the column with a washing buffer (50 mmol/l PBS, 20 mmol/l imidazole, pH 6.0). The SIase proteins were then eluted from the column with a linear gradient from 0 to 500 mmol/l imidazole in an elution buffer which contained 50 mmol/l PBS (pH 6.0). The purified protein was concentrated by ultrafiltration after dialysis against 50 mmol/l PBS (pH 6.0) and analyzed by 12% SDS-PAGE after visualizing by staining with Coomassie blue R250.
Characterization of SIase
Effects of Temperature and pH on SIase Activity
The effect of temperature on the activity of SIase was investigated in the range of 10–60 °C for 60 min. The optimum pH of SIase was determined as 30 °C using the standard assay conditions with three buffer systems (50 mmol/l), which were disodium hydrogen phosphate-citric acid buffer at pH 4.0–6.0, sodium phosphate buffer at pH 6.0–8.0, and glycine–NaOH at pH 8.0–9.0.
Effects of Metal Ions and EDTA on SIase Activity
Before studying the effects of metal ions on SIase activity, the purified enzyme was dialyzed against 50 mmol/l PBS (pH 6.0) containing 10 mmol/l ethylenediaminetetraacetic acid (EDTA) for 24 h at 4 °C. Subsequently, the enzyme was dialyzed against 50 mmol/l EDTA-free PBS (pH 6.0). The enzyme activity was then assessed under standard conditions in the presence of several metal ions (MgCl2, MnCl2, CoCl2, ZnCl2, CaCl2, CuCl2) and EDTA with a final concentration of 1 and 5 mmol/l. The measured activities of the enzyme were compared with those obtained in the absence of added ions under the same conditions.
Determination of Kinetic Parameters
Samples with different concentrations of sucrose (10–60 g/l) were prepared in 50 mmol/l PBS (pH 6.0), and incubated with the purified SIase at 30 °C for 10 min. The K m and V max values were determined by Michaelis–Menten plots.
Conversion of Sucrose into Isomaltulose by E. Coli (pET22b-palI) Cell Suspension
The E. coli cells (1 g wet wt.) expressing the SIase gene were washed two times with 50 mmol/l PBS (pH 6.0). The cell pellets were resuspended in the same solution at the desired concentrations. Reactions were conducted in 50 ml flasks containing 10 ml 550 g/l sucrose solution at 30 °C and shaken at 150 r/min for about 5 h. Aliquots of the reaction mixture were sampled and analyzed for the amounts of isomaltulose formed. The reactions were terminated by heating the flasks for 10 min in a boiling water bath.
Cell Immobilization and Production of Isomaltulose Using a Repeated-Batch Process
To immobilize the E. coli cells expressing SIase gene (or E. rhapontici NX-5 cells), cell pellets were washed twice with 50 mmol/l PBS (pH 6.0), treated with 0.05% (v/v) glutaraldehyde solution for 20 min and finally washed with sterile distilled water [22]. Then Five grams of treated cells were suspended in 10 ml PBS (50 mmol/l, pH 6.0) and mixed with 15 ml of 2% (w/v) sodium alginate solution. Immobilized beads were prepared by dropwise extrusion of the cell-alginate suspension into 3% (w/v) calcium chloride solution using silicone tubing.
Repeated-batch processes were carried out for isomaltulose production. The “batch”-type enzyme reactor containing 30 ml 550 g/l sucrose solution and 4 g immobilized cells of E. coli were kept at 30 °C for the desired time interval. At the end of the incubation of each batch, samples were collected and analyzed. The immobilized cells were then used for the next batch with addition of fresh substrate. The batch conversion processes were conducted for 30 days.
Assay of Enzyme Activity and Analysis of the Isomerized Product
The reaction mixture (1 ml) consisted of 0.5 ml appropriately diluted enzyme, and 0.5 ml 500 g/l (w/v) sucrose in 50 mmol/l PBS (pH 6.0) to obtain a final concentration of 250 g/l sucrose. The reactions were carried out at 30 °C for 60 min, and stopped by boiling for 10 min. One unit (U) of enzyme activity was defined as the amount of enzyme that formed 1 μmol of isomaltulose per min under the specified assay conditions. Enzyme activity of the immobilized cells was measured by incubating 1 g of immobilized cells with 10 ml of 500 g/l sucrose solution in 50 mmol/l PBS (pH 6.0) at 37 °C with gentle agitation. Thus, One unit of immobilized cells activity was determined as the amount of enzyme that can release 1 μmol isomaltulose per min at the initial stage under the standard assay conditions. All enzyme assays were performed in triplicate.
Reaction products were filtered through 0.22 μm membrane filters before HPLC analysis (Agilent 1200, USA system equipped with a refractive index detector). The samples were diluted 10-fold and 20 μl of diluted sample was injected onto a Rezex RCM-Monosaccharide Ca2+ column (Phenomenex, USA) for measurement of the sugar composition. The mobile phase was water with a flow rate of 0.5 ml/min at 80 °C. Glucose, fructose, sucrose, isomaltulose and trehalulose were used as standard sugars.
Results and Discussion
Cloning and Sequence Analysis of the E. Rhapontici NX-5 SIase Gene
The SIase gene was amplified from E. rhapontici NX-5 and five colonies containing about 2 kb foreign DNA fragments were screened out from approximately 100 colonies. After proceeding restriction enzyme treatment and DNA sequence analysis, an open reading frame (ORF) composed of 1803 nucleotides was determined to harbor a complete SIase gene fragment (Fig. 1). The ORF encoded a protein of 601 amino acid residues with a calculated isoelectric point of pH 6.0 and a molecular mass of 70 kDa.
Cloning of the SIase gene from Erwinia rhapontici NX-5 by PCR. Lane 1, DL2000 marker. Lane 2, SIase gene. Lane 3, pUC18 digested with HindIII (2,686 bp). Lane 4, pUC18-palIdigested with HindIII and BamH I to release the vector DNA (upper band) along with the insert DNA (1,803 bp, lower band). Lane 5, pET22b digested with Xho I (5,493 bp). Lane 6, pET22b- palIdigested with Xho I and Nco I to release the vector DNA (upper band) along with the insert DNA (1803 bp, lower band). Lane 7, DL15000 marker
The E. rhapontici NX-5 enzyme shared 92% homology at the nucleotide sequence level and 92.8% homology at the peptide sequence level with the SIase encoded by the SIase gene from E. rhapontici WAC2928 [12]. The SIase of E. rhapontici NX-5 differed in seven residues (Asp2, Gly5, Thr8, Ser236, Gln449, Asp527, Leu540) from that of strain WAC2928 (Ser2, Glu5, Ala8, X236, X449, Ala527, Phe540). In addition, there were substantial differences in the C-terminal regions of the two enzymes. The SIase of E. rhapontici WAC2928 contained an additional 32 amino acids that were absent from the SIase of E. rhapontici NX-5. Recombinant SIase from E. rhapontici WAC2928 has been reported to be unstable [12]. Further research should examine whether the C-terminal is concerned with the stability of the enzyme.
PalI of E. rhapontici NX-5 exhibited 82% identity to the SIase from P. rubum and S. plymuthica, and 71% identity with the SIase from Klebsiella sp. LX3 and Klebsiella pneumoniae NK33-98-8. A further homology search revealed that the deduced SIase gene product SIase from E. rhapontici NX-5 showed 75%, 72%, and 66% amino acid identity with SIase from Erwinia tasmaniensis, P. dispersa UQ68J, P. mesoacidophila MX-45, respectively (Fig. 2).
Amino acid sequence alignments of GH13 enzymes using ESPript tool [29]. Sequences are Erwinia rhapontici NX-5 (PalI, GenBank Accession No.HM461324) in this study, E. rhapontici WAC2298 (PalI, GenBank Accession No.AY223550), Serratia plymuthica (SIase, GenBank Accession No.CQ765973), Protaminobacter rubrum (SmuA, GenBank Accession No.CQ765969), Klebsiella sp. (PalI, GenBank Accession No.CQ765979), Pseudomonas mesoacidophila (MutB, GenBank Accession No.DQ304536), Bacillus cereus (OGL, GenBank Accession No.NZ_ACLU01000088). Secondary structure elements of SmuA (PDB Accession No. 3GBD) are indicated on top of the sequence with arrows for β-stands and cylinders for α-helices. The six conserved active sites are denoted by a green box
Domain architecture analysis using the SMART program (http://smart.embl-heidelberg.de) revealed that SIase contained an α-amylase domain spanning residues 55–460, with an E value of 3.32E-146. The main structural feature of the α-amylase domain was the central catalytic (β/α)8-barrel. Alignment with the amino acid sequences of other α-amylase family enzymes (glycosyl hydrolase family, GH 13), amylosucrase from Neisseria polysacchareais, and oligo-1,6-glucosidase from Bacillus cereus [23, 24], revealed six highly conserved residues Asp241, Glu295, Asp369, Tyr57, His145, His368 in the catalytic pocket that bound the substrate and hydrolyzed the glycosidic bond. The similarity of the active site architecture suggested that SIase of E. rhapontici NX-5 was a member of GH13 (Fig. 2).
Expression and Purification of the Recombinant SIase
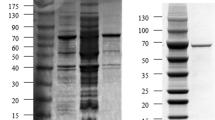
The SIase gene of E. rhapontici NX-5 was subcloned into the expression vector pET22b to generate the pET22b-palI fusion gene under the control of a lactose-inducible promoter. To maximize the yield of the fusion gene, different induction conditions were tested. When the cells were induced with lactose at a lower temperature, rather than at 37 °C to produce SIase, palI NX-5 activity was detected. Higher activity (15 U/ml) was achieved after induction with 0.5 mmol/l lactose at 24 °C for 11 h. The activity of recombinant SIase was over ten times higher than that in the wild strain. (1.3 U/ml). The enzyme was expressed with a fused 6 His-tag and purified by Ni–NTA chromatography. The protein was eluted at about 250 mmol/l imidazole and was purified up to 20-fold by Ni–NTA affinity chromatography in a single step. SDS-PAGE analysis of the purified recombinant SIase revealed a single band with a molecular mass of about 66 kDa (Fig. 3). Purified His-tagged SIase was collected for further enzymatic characterization.
Enzymatic Characterization of the Recombinant Sucrose Isomerase
The optimum pH for purified recombinant E. rhapontici NX-5 SIase was 5.0, with 89%, 86%, 80%, 34%, and 30% of maximum activity at pH 4.0, 6.0, 7.0, 8.0, and 9.0, respectively (Fig. 4). For SIase activity, this pH range was broad and more than 80% activity was obtained when pH varied from 4.0 to 7.0. The optimum temperature for SIase activity was 30 °C, SIase was stable for longer than 48 h at this temperature (Fig. 5). However, the stability of the enzyme decreased drastically at 50 °C and 60 °C with a half-life duration of 90 min and 5 min, respectively (data not shown).
This recombinant SIase has the most acidic optimum pH for isomaltulose synthesis compared with other SIases; moreover, it retained 85.2% of its maximum activity at pH 4.0. In contrast, three other well-characterized SIases from Klebsiella sp. LX3, P. Mesoacidophila MX-45, and Enterobacter sp. FMB-1, showed significant decreases in enzyme activity at pH below 5.0 [4, 6, 9]. Thus, the stability of the present SIase would be less likely to be affected when cells experienced lower pH and pO2 conditions [10], and enzymes that could tolerate acidic environment would help to reduce the chemical reaction between sugars and proteins [25]. Furthermore, the optimal pH of recombinant SIase was much closer to the pH value of the substrate solution, which might prove beneficial for biocatalysis.
Effects of Metal Ions on the Recombinant Sucrose Isomerase Activity and Kinetic Analysis
SIase activity was measured in the presence of metal ions (1 and 5 mmol/l). The SIase activities were inhibited by the metal ions to various degrees. Most of the tested metal ions inhibited enzyme activity to a level of 20–30%, except for Ca2+. SIase activity was greatly affected by the Ca2+, Cu2+, Zn2+, and Co2+, while the concentration of Mg2+ and Zn2+ showed no significant influence on SIase activity. The chelating agent EDTA inhibited 18% of the SIase activity (data not shown). Consequently, metal ions might not be required for the activity of SIase NX-5, which was consistent with previous studies [10]. The non-linear regression fitting of the Michaelis–Menten equation for the conversion of sucrose under standard assay conditions yielded K m and V max values for sucrose of 257 mmol/l and 48.09 μmol/l/s, respectively.
Conversion of Sucrose into Isomaltulose by Recombinant E. Coli (pET22b-palI)
The typical profile for the conversion of sucrose into isomaltulose is shown in Fig. 6. Sucrose conversion by E. rhapontici NX-5 was very rapid during the first 3 h and had gone nearly to completion in 7 h. In contrast, the recombinant E. coli (pET22b-palI) cells took only 1.5 h to convert the same amount of sucrose. The enzymatic reactants contained 87% isomaltulose and 10.5% trehalulose as well as slight amounts of glucose and fructose (Fig. 7). The productivity of E. coli (pET22b-palI) cells was 315.8 g/l/h, while that of E. rhapontici NX-5 was 62.23 g/l/h. Based on these results, recombinant E. coli (pET22b-palI) significantly shortened the conversion time and improved the isomaltulose productivity compared to the original E. rhapontici NX-5 cells, which would be beneficial for isomaltulose production.
Obtaining isomaltulose with immobilized cells containing recombinant SIase has not been previously reported, but the method looks promising. This is not only because of the high SIase activity that was retained by the immobilized cells, but also because of the simple medium and the short adaptation time needed for the immobilized E. coli. This method could increase access to the source of the enzyme. The recombinant SIase was located in the periplasmic space, unlike SIases of Enterobacter sp. FMB-1, P. rubrum, and S. plymuthica [9, 13, 26]. The facility of substrate transfer and product export could result in less product inhibition.
As shown in Table 1, Tsuyuki et al. obtained isomaltulose (63.9%) from a 200 g/l sucrose solution using immobilized K. planticola MX10 cells [27]. Krastanov and Yoshida also used immobilized cells of S. plymuthica 15928 for isomaltulose production and obtained a complete conversion of sucrose (400 g/l solution) into 80% isomaltulose and 7% trehalulose with small amount of glucose (2.89%), fructose (5.67%), isomaltose (1.20%) as byproducts [17]. Kawaguti et al. studied a continuous process using immobilized E. sp. D12 cells in a packed-bed reactor, and produced 55–66% of isomaltulose from 200 to 300 g/l sucrose solutions over 17 days, however, the rate of isomaltulose production could not be maintained for prolonged period [22]. Li observed that immobilized Klebsiella sp. LX-3 cells converted 99.7% of supplied sucrose (100 g/l solution) into 87% of isomaltulose, but increasing the sucrose concentration up to 300 g/l resulted in a decrease in the sucrose conversion rate to 71.8% [28]. In the present study, the immobilized E. coli (pET22b-palI) cells remained viable after 30 days conversions of concentrated sucrose solution (up to 550 g/l), with high isomaltulose yields (86 ± 1%) and sucrose conversion rates (>99%). In addition, this medium with concentrated sucrose might prevent the growth of other contaminating microorganisms, and achieve a high degree of conversion of sucrose into product. This would allow a great reduction in the size of equipment and the volume of sucrose required for isomaltulose production.
Conclusions
In this work, the SIase from E. rhapontici NX-5 was cloned and expressed in E. coli. The SIase NX-5 is distinguished from others by its most acidic optimum pH for isomaltulose synthesis, high catalytic efficiency for sucrose, and better product specificity. The immobilized E. coli (pET22b-palI) cells could continuously and efficiently produce isomaltulose with relative high isomaltulose yields (87%) and sucrose conversion rates (>99%). Furthermore, the successful cloning and overexpression of the SIase gene from E. rhapontici NX-5 has now set the stage for more detailed investigations of this enzyme, such as X-ray crystallography and protein engineering studies, which will permit further optimization of the catalytic properties of the SIase to improve its industrial use.
References
Oshima, T., Izumitani, A., Sobue, S., et al. (1983). Infection and Immunity, 39, 43–49.
Schiweck, H., Munir, M., Rapp, K. M., et al. (1991). Schneider. In Carbohydrates as Organic Raw Materials (Ed.), Lichtenthaler, F.W (pp. 57–94). Weinheim: Wiley-VCH.
Lina, B. A., Jonker, D., Kozianowski, G., et al. (2002). Food and Chemical Toxicology, 40, 1375–1381.
Nagai, Y., Sugitani, T., & Tsuyuki, K. (1994). Bioscience, Biotechnology, and Biochemistry, 58, 1789–1793.
Bornke, F., Hammad, M., & Sonnewald, U. (2001). Journal of Bacteriology, 183, 2425–2430.
Zhang, D., Li, X., & Zhang, L. H. (2002). Applied and Environmental Microbiology, 68, 2676–2682.
Wu, L., & Birch, R. G. (2005). Journal of Applied Microbiology, 97, 93–103.
Ravaud, S., Watzlawick, H., Haser, R., et al. (2006). Acta crystallographica section F structural biology and crystallization communications, 62, 74–76.
Cha, J., Jung, J. H., Park, S. E. et al. (2009). Journal of Applied Microbiology. dio:10.1111/j.1365-2672.2009.04295.x
Cheetham, P. S. J. (1984). The Biochemical Journal, 220, 213–220.
Salvucci, M. E. (2003). Comparative Biochem Physiol B, 135, 385–395.
Wu, L., & Birch, R. G. (2005). Applied and Environmental Microbiology, 71, 1581–1590.
Veronese, T., & Perlot, P. (1999). Enzyme and Microbial Technology, 24, 263–269.
Zhang, D., Li, N., Lok, S. M., et al. (2003). The Journal of Biological Chemistry, 278, 35428–35434.
Ravaud, S., Robert, X., Watzlawick, H., et al. (2007). The Journal of Biological Chemistry, 282, 28126–28136.
Ravaud, S., Robert, X., Watzlawick, H., et al. (2009). FEBS Letters, 583, 1964–1968.
Krastanov, A., & Yoshida, T. (2003). Journal of Industrial Microbiology and Biotechnology, 30, 593–598.
Krastanov, A., Blazheva, D., & Stanchev, V. (2007). Process Biochemistry, 42, 1655–1659.
Kawaguti, H. Y., & Sato, H. H. (2007). Biochemical Engineering Journal, 36, 202–208.
Kawaguti, H. Y., Buzzato, M. F., & Sato, H. H. (2007). Journal of Industrial Microbiology and Biotechnology, 34, 261–269.
Xu, H., Li, S., Huang, M. X. et al. (2007). CN patent 200710190755.2.
Kawaguti, H. Y., Buzzato, M. F., Orsi, D. C., et al. (2006). Process Biochemistry, 41, 2035–2040.
Montalk, G. P., Magali, R. S., Willemot, R. M., et al. (2000). FEBS Letters, 471, 219–223.
Watanabe, K., Miyake, K., & Suzuki, Y. (2001). Bioscience, Biotechnology, and Biochemistry, 65, 2058–2064.
Dworschak, E. (1980). Critical Reviews in Food Science and Nutrition, 13, 1–40.
Mcallister, M., Kelly, C. T., Doyle, E., et al. (1990). Biotechnological Letters, 12, 667–672.
Tsuyuki, K., Sugitani, Y., Miyata, Y., et al. (1992). The Journal of General and Applied Microbiology, 38, 483–490.
Li, X., Zhao, C., An, Q., et al. (2003). Journal of Applied Microbiology, 95, 521–527.
Gouet, P., Robert, X., & Courcelle, E. (2003). Nucleic Acids Research, 31, 3320–3323.
Cho, M. H., Park, S. E., Lim, J. K., et al. (2007). Biotechnological Letters, 29, 453–458.
Nagai, Y., Tsuyuki, K., Sugitani, T., et al. (1993). Bioscience, Biotechnology, and Biochemistry, 57, 2049–2053.
Acknowledgements
This work was supported by Key Projects in the National Science and Technology Pillar Program during the Eleventh 5-Year Plan Period (2008BAI63B07), National Natural Science Foundation of China (20906050), Natural Science Foundation of Jiangsu Province (BK2009357), The Natural Science Foundation of the Jiangsu Higher Education Institutions of China (08KJA180001), The University Nature Science Research of Jiangsu Province (09KJB530007).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Li, S., Cai, H., Qing, Y. et al. Cloning and Characterization of a Sucrose Isomerase from Erwinia rhapontici NX-5 for Isomaltulose Hyperproduction. Appl Biochem Biotechnol 163, 52–63 (2011). https://doi.org/10.1007/s12010-010-9015-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-010-9015-z