Abstract
Hyperspectral imaging is built with the aggregation of imaging, spectroscopy and radiometric techniques. This technique observes the sample behaviour when it is exposed to light and interprets the properties of the biological samples. As hyperspectral imaging helps in interpreting the sample at the molecular level, it can distinguish very minute changes in the sample composition from its scatter properties. Hyperspectral data collection depends on several parameters such as electromagnetic spectrum wavelength range, imaging mode and imaging system. Spectral data acquired using a hyperspectral imaging system contain variations due to external factors and imaging components. Moreover, food samples are complex matrices with conditions of surface and internal heterogeneities, which may lead to variations in acquired data. Hence, before extracting information, these variations and noises must be reduced from the data using reference-dependent or reference-independent spectral pre-processing techniques. Using of the entire hyperspectral data for information extraction is tedious and time-consuming. In order to overcome this, exploratory data analysis techniques are used to select crucial wavelengths from the excessive hyperspectral data. Using appropriate chemometric techniques (supervised or unsupervised learning techniques) on this pre-processed hyperspectral data, qualitative or quantitative information from sample can be obtained. Qualitative information for analysing of the chemical composition, detecting of the defects and determining the purity of the food product can be extracted using discriminant analysis techniques. Quantitative information including variation in chemical constituents and contamination levels in food and agricultural sample can be extracted using categorical regression techniques. In combination with appropriate spectra pre-processing and chemometric technique, hyperspectral imaging stands out as an advanced quality evaluation system for food and agricultural products.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The revolutions in computer, optical and image sensor technologies and their applications in the field of agriculture have led to sensing techniques like imaging systems which are capable of automated and quick quality analysis on the processing line, with very little human interference (Zayas et al. 1985; Sapirstein et al. 1987; Symonds and Fulcher 1988; Brosnan and Sun 2004; Du and Sun 2004; Gowen et al. 2007). Initial applications of image processing to the food and agricultural products were the use of red-blue-green colour vision system for colour and size grading and later for identifying defects (Chen et al. 2002; Paliwal et al. 2005; Ramalingam et al. 2011; Singh et al. 2012). Although colour imaging can perform the quality analysis, when it comes to the identification of minor constituents or contaminants, it is ineffective. Multispectral imaging systems have been developed using few narrow wavelengths generally less than 20, to detect features of interest (Gowen et al. 2007; Yang 2012). These wavelengths may range from visible to near infrared. The number of wavelengths used to constitute a multispectral image cube may range based on the imaging system, number of filters used and features of interest. For instance, if the feature of interest is already known, the research acquires images in those wavelength ranges that can accurately provide information about that feature. Novales et al. (1999) used multispectral fluorescence imaging for the identification of maize, peas, soybean and wheat. Their multispectral imaging system was built to take 12 images and was able to identify each of these food products with classification accuracy up to 98.8 %. Liu et al. (2014) used multispectral imaging to take images at 19 wavelengths ranging from 405 to 970 nm for determining various quality attributes (firmness, total soluble solids (TSS)) and ripeness of strawberry fruits. Multispectral imaging has proven to be advantageous over conventional chemical analysis routines like gas chromatography (GC) and high-performance liquid chromatography (HPLC) which are destructive in nature. Multispectral imaging finds itself as the basis for the evolution of hyperspectral imaging system, which is capable of acquiring images over a wide range of wavelengths along the electromagnetic spectrum and correlating the spectral variation with the chemical constituent of interest. The multispectral imaging typically involves acquiring images in the three to six spectral bands that range from visible to near-infrared (NIR) range (Jensen 2007). Acquiring images in such a few narrow wavelengths is the primary disadvantage of multispectral imaging. Acquiring images in this small wavelength range provides gaps in the spectral band to exploit the entire signature. Hyperspectral imaging fills this gap by acquiring images in a broader wavelength range of more than 200 spectral bands that range from visible to NIR section. Hyperspectral imaging is an advanced technique for the quality evaluation in the food and agricultural industry, which is built with the combination of regular imaging, radiometry and spectroscopic principles. Radiometry is the measure of the amount of electromagnetic energy (generally expressed in common energy units, W) present within a specific wavelength range. Typical radiometer is designed with only one sensor with a filter installed to just to select the intended wavelength range. Spectrometry is a measure of light’s intensity (generally expressed in W/m2) within specific wavelength range. Unlike radiometer, spectrometers use diffraction grating or prisms or multiple sensors to divide the wavelength range into different wavelength bands. Goetz et al. (1985) used the word ‘hyperspectral imaging’ for the first time when they were discussing their remote sensing of earth using spectroscopic imaging techniques. The term ‘hyper’, meaning too much, has a negative implication in the medical sense, like hypertension, but in image processing, ‘hyperspectral’ imaging means acquiring images at ‘many bands’ along the electromagnetic spectrum. Chemicals when available in pure form and to be identified uniquely do not require hundreds of spectral bands spread over several octaves of the electromagnetic spectrum (Goetz 2009). But biological materials exist as a complex system of chemical compounds with interactions and bonds between them. Application of image processing in combination with spectroscopic technique makes it possible for automatic target detection and measurement of the analytical composition of that material; hence, hyperspectral imaging can be effectively applied for quantitative and qualitative analyses, as it can identify the presence of the material as well as their spatial location.
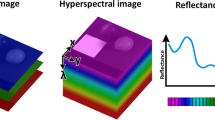
Hyperspectral imaging is an appropriate technique for many operations as it can generate both a spatial map and spectral variation (ElMasry and Sun 2010). Data acquired from hyperspectral imaging system are three-dimensional structures made up of one spectral and two spatial dimensions, commonly known as the ‘hypercubes’ or the ‘datacubes’ (Fig. 1) (Schweizer and Moura 2001; Chen et al. 2002; Kim et al. 2002; Mehl et al. 2004).
Goetz et al. (1985) originally developed the hyperspectral imaging for the remote sensing; later, its applications were expanded to various fields like agriculture (Du et al. 2013; Suzuki et al. 2012; Gracia and León 2011; Pérez-Marín et al. 2011), environment (Wang et al. 2016), food science (Lu 2003; Cheng et al. 2004; Nicolai et al. 2006; Gowen et al. 2007; ElMasry et al. 2007; Qu et al. 2016; Xu et al. 2016a,b; Xie et al. 2016, Wold 2016, Cheng et al. 2016), material science (Neville et al. 2003; Tatzer et al. 2005), biomedicine (Zheng et al. 2004; Kellicut et al. 2004; Liu et al. 2012; Akbari et al. 2012; Kiyotoki et al. 2013), astronomy (Zhang et al. 2008; Soulez et al. 2011; Nguyen et al. 2013) and pharmacy (Amigo and Ravn 2009; Lopes et al. 2010; Amigo 2010; Vajna et al. 2011). In the field of food and agriculture, hyperspectral imaging is used to determine the quality in fruits (Lu 2003; Mehl et al. 2004; Xing et al. 2005; Lee et al. 2005; Nicolai et al. 2006; Lefcourt et al. 2006; Qin et al. 2008; Bulanon et al. 2013; Pu and Sun 2016; Pu et al. 2016), vegetables (Cheng et al. 2004; Ariana et al. 2006; Gowen et al. 2009; Taghizadeh et al. 2011; Wang et al. 2012; Su and Sun 2016), meat and meat products (Park et al. 2002; Park et al. 2006; Sivertsen et al. 2009; Qu et al. 2016; Xu et al. 2016a,b; Xie et al. 2016; Wold 2016; Cheng et al. 2016; Chen et al. 2016; Ma et al. 2016), cereals (Cogdill et al. 2004; Zhang et al. 2007; Mahesh et al. 2008; Choudhary et al. 2009; Singh et al. 2010) and diseases in crops (Smith et al. 2004; Keulemans et al. 2007; Qin et al. 2009; Kumar et al. 2012; Xie et al. 2015; Kuska et al. 2015). Hyperspectral data collection is the most important step to understand the properties of the sample. Significant scientific literature has been published in recent years regarding the data acquisition using hyperspectral imaging system (Varshney and Arora 2004; Borengasser et al. 2007; Park and Lu 2015; Vadivambal and Jayas 2016).
Hyperspectral Data Collection
Understanding the behaviour of food sample at molecular level on its exposure to the light photons is the basis for the hyperspectral imaging (ElMasry and Sun 2010). Hyperspectral imaging deals with the sample at the molecular level; therefore, it can distinguish even slight changes in the sample composition. Biological materials continuously absorb or emit energy (radiation) by lowering or raising its molecules from one energy level to another. The strength and wavelength of this radiation depend on the nature of the material (ElMasry and Sun 2010). Hyperspectral imaging generally measures the intensity of the absorbed or emitted radiation over a range of spectral band along the electromagnetic spectrum. Data collected using a hyperspectral imaging system are the absorbed or emitted radiation values and are stored in the form of a hypercube. The hypercube is a complex data unit, which contains abundant information about the physical and chemical properties of the sample. Data collection using a hyperspectral imaging system depends on various parameters such as electromagnetic spectrum wavelength range used for imaging, type of the imaging system and type of the imaging mode.
Imaging Spectrum Range
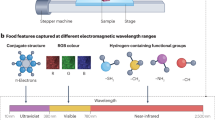
Electromagnetic spectrum (Fig. 2) is the distribution of all types of electromagnetic radiation arranged in one spatial scale ranging from very short wavelength (very high frequency and energy) gamma radiation to the long wavelength (very low frequency and energy) radio waves. Electromagnetic radiation is a wave of energy packets called photons with both magnetic and electrical properties which when interact with matter produce a spectrum. Gamma radiation (wavelength less than 0.01 nm) has very high energy and good penetration potential. In food and agricultural industry, it is used for microbial safety as it can kill the microorganisms by disrupting their DNA bonds. X-rays are classified into two forms, hard X-rays (wavelength between 0.01 and 0.1 nm) and soft X-rays (wavelength between 0.1 and 10 nm), and are used in food applications for detection of hidden infestation in grains (Karunakaran et al. 2003) and defects in fruits and vegetables (Schatzki et al. 1997). There is a limitation in the use of gamma rays and X-rays for biological materials as their high energy damages the structure and components of biological materials. The use of ultraviolet light (wavelength between 10 and 100 nm) is well established in the food industry for water treatment, air disinfection and surface decontamination due to its low penetration power (Koutchma 2008). Microwaves (wavelength range from 106 to 109 nm) and radio waves (wavelength range from 109 to 1017 nm) have been used for grain disinfestation (Vadivambal et al. 2007) and to study the grain moisture distribution (Ghosh et al. 2007) due to their high transmission power. Visible range (400 to 700 nm) imaging is used mainly for colour-based sorting of agricultural and food materials. Visible light is generally used in combination with near-infrared radiations (400–1000 nm range) to achieve a robust hyperspectral range for quality identification in various foods. This range is called the visible-infrared (Vis-IR) region. Kim et al. (2002) effectively detected faecal contamination on apples using hyperspectral imaging in the Vis-IR region (450 to 851 nm). Mehl et al. (2004) applied Vis-IR hyperspectral imaging within the spectrum of 430 to 900 nm for the detection of surface defects such as side rots, bruises, flyspeck, scrubs and moulds, fungal infection and soil contamination on apples. Delwiche et al. (2011) applied Vis-IR hyperspectral imaging (400–1000 nm) to find Fusarium head blight damage in wheat kernels. Infrared radiation with a wavelength range of 900 to 12,000 nm falls within electromagnetic spectrum between the visible and microwave regions and is the most explored region for hyperspectral image analysis. Infrared spectrum that is closest to visible light is called NIR (ranging from 900 to 1700 nm) spectrum. Hyperspectral instrument designed for use in NIR region will have a very high signal-to-noise ratio (10,000:1) (Hans 2003). In recent years, NIR hyperspectral imaging has been extensively applied for understanding the quality characteristics of food and agricultural materials. Vermeulen et al. (2011) applied NIR hyperspectral imaging (900–1700 nm) to identify the presence of ergot bodies in wheat kernels. NIR hyperspectral spectral imaging was used to identify the level of Fusarium infection in maize using 720 to 940 nm range (Firrao et al. 2010) and in wheat using 400 to 1000 nm range (Bauriegel et al. 2011). Dacal-Nieto et al. (2011) applied NIR hyperspectral imaging for the identification of hollow heart in potato using 900 to 1700 nm range. Short-wave infrared (SWIR) (970–3000 nm) hyperspectral imaging technique has been used for identification of undesirable substances in food and feed (Pierna et al. 2012), wheat kernels damaged by insects (Singh et al. 2010), moisture content of cereals like barley, wheat and sorghum grain (McGoverin et al. 2011). Mid-wave infrared (MWIR) (3000 to 5000 nm) and long-wave infrared (LWIR) (8000 to 12,000 nm) hyperspectral imaging has been used for the identification of hot gases and minerals, respectively, from the earth, using remote sensing techniques (Anonymous 2013).
Imaging System
The three-dimensional hypercube (x, y, λ), which is a pile of two-dimensional spatial images acquired at various wavelengths can be obtained in any one of these three approaches:
-
1.
Intensity data collected at all the wavelengths for one pixel at a time.
-
2.
Intensity data collected at all the wavelengths for one row of pixels at a time.
-
3.
Intensity data collected at one wavelength for all the pixels at a time.
In a hypercube, a spatially arranged image at each wavelength contains an equal number of pixels. Each pixel contains the spectrum, which relates to the chemical composition of the sample at that individual pixel. The three-dimensional hyperspectral image datacube can be acquired using any one of these four approaches (Wang and Paliwal 2007):
-
1.
A point-to-point spatial pattern (whiskbroom method).
-
2.
Fourier transform imaging.
-
3.
A line-by-line spatial scan pattern (line scan or pushbroom imaging system).
-
4.
Wavelength tuning with filters (area scan or staring array imaging system).
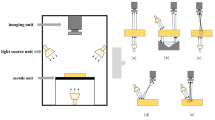
The line scan and area scan imaging system are better suited for quality analysis of food and agricultural materials (Mehl et al. 2004) (Fig. 3).
Line Scan or Pushbroom Imaging System
Line scan or pushbroom imaging system is proficient by collecting spectral data of single spatial line at a time and progressively constructing the hypercube (Wang and Paliwal 2007). The light intensity of the narrow line of the sample along all the wavelength range is imaged onto one row of the hypercube using two-dimensional dispersing element and a two-dimensional detector array. This imaging system generally operates with an imaging lens having a slit-shaped opening aperture. The pushbroom hyperspectral imaging system uses wavelength dispersive system that uses a diffraction grating (transmissive or reflective) or prism techniques. Pushbroom imaging works either by physically moving the sample (Martinsen et al. 1999) or by directing the beam and detector to the region of interest (Nicolai et al. 2007). Unlike area scan imaging system, spectral record of the entire imaging session can be controlled in the pushbroom technique. Pushbroom imaging system has been adopted in food and agricultural industries because of its speed and adaptability (Manley et al. 2009).
Area Scan or Staring Array Imaging System
Area scan or staring array imaging system acquires the entire spatial image of the sample at one wavelength and steps through the next wavelength. This technique is also called wavelength scanning or tunable filter scanning or focal plane scanning. Generally, it uses tunable filters to collect images at one wavelength at one time. Tran (2005) had provided an extensive review of the principles, instrumentation and application of filters in infrared multispectral imaging systems. The most frequently used tuning filters in staring array hyperspectral imaging system are filter wheels, liquid crystal tunable filter (LCTF) and acousto-optic tunable filter (AOTF) (Fig. 4).
Filter wheels are used in the most basic form of multispectral imaging systems, where a mechanical filter wheel is incorporated into the optical path during imaging. The number of imaging wavelengths is determined by the number of windows in the filter wheel. This technique is restricted to a limited number of fixed wavelengths, which is initially determined by the prior knowledge about the sample’s spectral behaviour (Fong and Wachman 2008). This technique is mechanically restricted to change wavelength at a relatively slow pace. Filter wheels are only used in inexpensive imaging systems (Wang and Paliwal 2007). AOTFs are crystals with optical properties, and the wavelengths are controlled by sending acoustic waves through them. These sound waves are created by applying radiofrequency electrical signals, AOTF wavelength and bandwidth that can be regulated electronically. LCTFs, also known as Lyot filters, can select various band passes from the sample (band-sequential scanning system), as they are generated by alternative layers of crystal quartz plates and liquid crystal polarizer. LCTFs generally work in between visible and SWIR of the electromagnetic spectra. Use of these filters with hyperspectral imaging system increased the speed of data acquisition, made auto wavelength change and random wavelength selection possible, but, in turn, increased the overall cost of the system. The staring array imaging system is applicable for many operations where the sample is fixed at one point, for example, these tunable filters are used to change the excitation and emission wavelengths electronically to achieve excitation-emission matrix in florescence imaging (El Masry and Sun 2010). The two main drawbacks of staring array imaging systems when compared to the pushbroom imaging system are
-
1.
Because of lengthy imaging time and continuous exposure to the illumination system, the sample gets heated up and sometimes loses moisture.
-
2.
It is not useful for online sensing or real-time application in industrial scale, as stationary samples are needed.
Imaging Mode
Understanding the theory of interaction of light with the surface of the studied material is important for understanding the concept of hyperspectral imaging. Generally, hyperspectral imaging is carried out in any one of the optical modes (reflectance, absorbance and transmittance) or in florescence mode. According to the optical property of the sample, when the electromagnetic radiation is incident on this sample, the radiation is reflected, absorbed and transmitted. The absorbed light can sometime be re-emitted at lower energy and longer wavelength, and this aspect is known as fluorescence. The fluorescence and optical properties are the unified functions of the wavelength and angle of the incident radiation as well as physical and chemical characteristics of the sample (Chen et al. 2002; ElMasry et al. 2012a). According to Beer-Lambert law, the absorbance of the sample is directly proportional to the concentration of the chemical constituent of that sample. This law had laid the base for the concept of chemical composition analysis using hyperspectral imaging.
Reflectance
Reflection is a process where the fraction of the electromagnetic radiation (radiant flux) incident on the surface of the sample returns into the same hemisphere whose base is the surface of the sample and which contains the incident radiation. The reflectance of light occurs in three main types: specular (reflects back from the surface at the same angle as that of incoming light), diffuse (scattered or reflected at many different angles) or a combination of both. Most of the biological materials reflect light in the diffusion mode. The general mathematical definition for reflectance ρ is given by the radiant flux reflected ϕ r divided by the radiant flux incident ϕ i :
Reflectance is the most common hyperspectral imaging mode used for the analyses of food quality and safety. Reflectance mode measurements are easy to conduct without any contact with the substance, and light level is reasonably high in relation to the sample (El Masry et al. 2012a). Hyperspectral reflectance imaging is commonly applied in Vis-NIR (400–1000 nm) or NIR (1000–1700 nm) spectrum and has been applied to identify defects, quantify contaminants and analyse quality attributes of fruits, vegetables and meat products (Gowen et al. 2007; Wu and Sun 2013b). In particular, it has been used to classify cereals (Mohan et al. 2005; Gorretta et al. 2006; Mahesh et al. 2008), identify fungal infection in agricultural products (Wang et al. 2003; Pearson et al. 2001) and determine quality of fruits and vegetables (Kim et al. 2002).
Transmittance
Transmittance is a process by which the fraction of the electromagnetic radiation (radiant flux) leaves the surface of the sample from the side (usually opposite) other than that of incident radiation. Within the electromagnetic spectral range, transmittance is more commonly measured in the 700 to 1100-nm NIR region (Givens et al. 1997; Ariana and Lu 2010). Although in transmittance, the extent of light passing through the sample is relatively small; it contains significant valuable information to detect internal defects and estimate the internal ingredient concentration of the food more accurately (Schaare and Fraser 2000; Shenderey et al. 2010). As it requires a strong light source and a sensitive detector, transmission mode is not always preferred when compared to the reflectance mode (ElMasry et al. 2012a). In agricultural and food applications, transmittance hyperspectral system was used to extract the data from cucumbers for detection of insects, using a moving sample (Lu and Ariana 2013) and other vegetables and soybeans (Huang et al. 2012b), from cod fish sample for detection of parasite load (Heia et al. 2007; Sivertsen et al. 2011), from tart cherries for detection of pits (Qin and Lu 2005) and from corn for estimation of moisture content and oil of kernels (Cogdill et al. 2004).
Interactance
Interactance is an image sensing mode where the light is allowed to fall on the sample and detected using a detector lying on the same side of the sample (Wu and Sun 2013a). The part of light that interacts with the sample and re-emitted is analysed in interactance mode. Due to exploration of this property of light in interactance hyperspectral imaging, it can deliver more valuable information about the deeper properties of the sample compared to reflectance. It is also advantageous than transmittance due to reduced influence of thickness of the sample. Unlike transmittance, where a special setup is required for sealing the light to prevent it from entering directly into the detector, interactance is also advantageous in its simple setup (Nicolai et al. 2007). Interactance imaging is widely used for the determination of fat content of humans and animals. Interactance hyperspectral imaging system application for food and agricultural products has been limited, but it was used for detection of fat content of beef (Wold et al. 2011), ham (Gou et al. 2013) and pork (O’Farrell et al. 2010) and nematodes in cod fish (Sivertsen et al. 2012). Wang et al. (2013) and Schaare and Fraser (2000) compared the reflectance, interactance and transmittance hyperspectral imaging for the quality assessment of the fruits and vegetables.
Absorbance
Absorbance is the process by which a portion of incident electromagnetic radiation (radiant flux) is retained in the sample without complete reflectance or transmittance. In this case, the incident energy is converted into different form and preserved within the sample. Amount of energy converted into different form is dependent on chemical constituents of the sample. The general mathematical definition of absorbance α is the ratio of the radiant flux absorbed ϕ a to the radiant flux incident ϕ i :
Extrinsic characteristics like size, geometry, appearance and colour and chemical compositions like water content, fat, protein and carbohydrate of the sample can be identified using hyperspectral absorbance imaging (Lu and Chen 1999). Most biological materials have stronger absorption in the spectral range of 600–1000 nm (Kavdir et al. 2009). Although the use of hyperspectral absorbance imaging mode is very limited when compared to the reflectance mode, effective literature in the field of food and agriculture was identified. For cereal grain research, spectroscopic absorbance imaging was used for categorization of vitreous and non-vitreous kernels (Dowell 2000), detecting insect infestation (Baker et al. 1999) and quantifying essential nutrients and chemical constituents (Wang et al. 2004) of various cereals. In general, hyperspectral data acquired using any of the radiometric properties (reflectance, absorbance or transmittance) can be exchanged between each other.
Fluorescence
Fluorescence is a technique in which light absorbed at a given wavelength by a chromophore of the sample is retained and later emitted at higher wavelength (Kim et al. 2001). This difference in absorption and emission wavelengths is called Stokes shift. The phenomenon of fluorescence is observed in various samples in the visible region (400–700 nm), when excited with ultraviolet (UV) region radiation (Chappelle et al. 1991; Lang et al. 1992). In the areas of cell biology, medicine, forensic and environmental sciences, steady-state fluorescence imaging is regarded as a sensitive optical technique and used for scientific research (Harris and Hartly 1976; Chappelle et al. 1984; Albers et al. 1995). Changes in fluorescence emission of food due to contaminations with faecal material and infection with pathogens serve as a basis for application of fluorescence hyperspectral imaging for foods. Despite these advantages, the use of fluorescence imaging for the online quality and safety assessment in food industry is not fully explored due to the complexity in this phenomenon (Kim et al. 2001). Fluorescence hyperspectral imaging system was used for assessing various quality parameters like skin and flesh colour, firmness, soluble solids and titrateable acidity (TA) in apples (Noh and Lu 2007), for detecting skin tumours in chicken carcasses (Kim et al. 2004). This technique was also used for detection of the faecal contamination on apples (Kim et al. 2002; Lefcourt et al. 2006) and cantaloupes (Vargas et al. 2005).
Hyperspectral Imaging Sample Complexities
In comparison with the traditional imaging, hyperspectral imaging is complex, as it involves working with more number of images at the same time in both spatial and spectral dimensions. In order to say that hyperspectral imaging results are the perfect representation of the objective results, there is a need to consider the drawbacks and issues that arise due to sample complexity. In this section, the most common sample complexities that are to be considered while working with hyperspectral images are as follows:
Variations in Sample Surface
Food materials are complex and show very high variations in the sample surface. In reflectance mode, incident radiation in the form of photons falls on the sample surface. Based on the sample surface condition, this radiation may enter the sample or get reflected from the surface. The amount of the photons that reflected back to the sensor is based on the incident radiation wavelength and the nature of the sample surface (Clark and Sasic 2006). The reflected radiation that reached the detector contains both the sample composition at different depths and location within the sample. This depth of penetration depends on the condition of the sample surface. Hence, while conducting hyperspectral imaging, it is important to consider the sample surface variations to arrive at an accurate analysis in the later stages.
Asymmetry in Sample Surface
Considering the focus nature of the hyperspectral imaging systems, the use of flat-surfaced materials for imaging is preferable. The use of flat-surfaced sample makes the sample surface parallel to the imaging plane (Amigo 2010). In contrast to these food materials, sample surfaces are irregular with variations in colour and surface conditions. Most of the dry food and agricultural materials are having large variations in the surface roughness, which creates incident light to scatter. Hence, the collected information may not be the exact representative of the sample spectra. While acquiring hyperspectral images, it is important to keep in mind the asymmetry and roughness of sample surface.
Sample Representativeness
Hyperspectral imaging involves extraction of information due to the interaction of incident light mostly with the surface and very little with the penetration of the food sample. In most cases, penetration depth is negligible as the food samples are complex and opaque. As the food sample matrix is very complex, it can be imagined that if the sample is sliced into infinite number of thin slices, the composition of each sliced surface will show that each sliced surface itself will be very different due to variation in composition and particle sizes in each layer. Keeping this sample complexity in mind, it can be concluded that the one single sample spectra cannot be considered as the signature of the sample at that particular condition (Amigo 2010). Spectral signature can only be constructed by acquiring hyperspectral images of large number of sample (replicates) under that particular condition, to increase the sample representativeness.
Hyperspectral Data Pre-Processing Before Extraction of the Information
Unlike conventional imaging, hyperspectral imaging is more subjected to variations due to external factors and imaging components and complexities in food. Food samples being complex matrices with conditions like surface inhomogeneities bring variations in acquired data. Incident light on the food material experiences both scattering and absorption effect due to the complexity in its interaction with various components in food material. This scatter effect is due to physical properties of food material like cellular structure, particle size, density, etc., and absorption effect is due to chemical composition like carbohydrates, protein, fat, etc. Hence, before applying any modelling method, the homogeneity among the input data should be ascertained, as data are affected by outliers, subgroups or clusters (Centner et al. 1996). To reduce these variations and to extract the useful information from hyperspectral images, spectral pre-processing tools are generally used. In spectral data analysis, the most important function of the pre-processing techniques is to reduce the undesirable variation and noise during hyperspectral data acquisition and make the data analysis more meaningful. If fair results are expected from the hyperspectral data analysis, the pre-processing step is often of vital importance (ElMasry and Sun 2010).
Hyperspectral imaging involves the application of Beer-Lambert law which shows the interaction between the incident light and chemical properties of the sample through which this light passes. Beer-Lambert law can be mathematically represented as
where ‘a’ is the absorbance of the sample, ‘ε’ is the molar absorptivity of the sample, ‘b’ is the effective path length and ‘c’ is the constituent concentration in the sample. As the product of ‘ε × b’ remains constant, spectral pre-processing techniques aim to maintain the relationship between absorbance a and constituent concentration c to be linear and making the measured absorbance a perfect representation of constituent concentration. This linear relationship is affected by undesirable phenomena such as particle size effects, scattering of light, morphological differences like surface roughness and detector artefacts. For instance, if the spectrum of a sample is affected by light scattering, Beer-Lambert law is not true and this kind of spectrum need to be pre-processed in order to eliminate this effect before model development (Stordrange et al. 2002). Pre-processing techniques are applied on spectral data to promote the linear relationship between hyperspectral data and concentration of sample constituent and to compensate for these deviations. Pre-processing techniques that are established on feature selection or extraction always intend to reduce or transform the primary feature space into another space of lower dimensionality (Melgani and Bruzzone 2004). Pre-processing and collection of accurate auxiliary data are the most important requirements to extract qualitative information from the hyperspectral data.
Pre-processing techniques are used on the hyperspectral data for the following reasons:
-
1.
Identification and removal of trends, outliers and noise in data.
-
2.
Improvement of the performance of the subsequent qualitative and quantitative analyses.
-
3.
Enhancement of data interpretation.
-
4.
Simplified machine learning using pre-processed data.
-
5.
Removal of inappropriate and unnecessary information from the data and reduction of the scale of data mining step.
For more systematic understanding, different spectral pre-processing techniques can be categorized into reference-dependent and reference-independent groups.
Reference-Dependent Pre-Processing Techniques
Reference-dependent techniques use pre-determined reference values for spectral pre-processing. The main categories of reference-dependent techniques are orthogonalization, optimized scaling and spatial interface subtraction.
Orthogonal Signal Correction
Hyperspectral data are generally analysed as multivariate data. Application of partial least squares (PLS) to obtain projections for the latent structures is one of the main methods to analyse these multivariate data, where measurable relationship between a raw data matrix X and the response matrix Y is needed. PLS is a useful tool in various areas like multivariate calibration, classification, discriminate analysis and pattern recognition (Martens and Naes 1992). PLS is very successful in the multivariate data analysis because it can handle the data with collinearity among the variables, missing data and noisy data. Wold et al. (1998) had proposed the orthogonal signal correction (OSC) as a reference-dependent pre-processing technique. OSC technique removes all the variations in X that are not corresponding to Y, in order to attain exceptional models in multivariate data analysis (Trygg and Wold 2002). From the general understanding of hyperspectral dataset, most of the spectral variance is of very little or even nil analytical importance. Therefore, for better analysis, it is important to remove the variance that is orthogonal to the component of interest from the dataset. The number of orthogonal factors to be eliminated from the dataset must be decisive, and the more the number of factors eliminated, the greater is the reduction of orthogonal variance (Boysworth and Booksh 2007). In order to achieve orthogonal correction models, three important criteria must be met:
-
1.
Component of interest should involve the large systematic variation in X.
-
2.
Component of interest must be predictive by X.
-
3.
Component of interest must be orthogonal to Y (Trygg and Wold 2002).
These three criteria are met by performing an OSC on X, which will remove most of the components of X that are unrelated to the model developed from Y. The main disadvantages of OSC pre-processing technique are
-
1.
Time-consuming, as OSC methods follow a number of internal iterations to identify the orthogonal components of X.
-
2.
Additional time for cross-validation.
-
3.
Difficult to estimate the correct number of components that are needed to be eliminated from X, those are not correlated to Y.
Raw hyperspectral data before PLS modelling are generally pre-processed using OSC method to eliminate the variance from X that is not related to Y. OSC is also used for instrument standardization by removing the variability which is inappropriate to the predicted variables.
Orthogonal Projections to Latent Structures
Orthogonalization is not a single pre-processing technique; rather, it is a group of algorithms and programs, with a goal to separate the variation in the sample spectrum, which will not correlate with the reference value (Rinnan et al. 2009). When the factors are too many and well collinear, PLS is used to develop predictive models. PLS was first developed by Herman Wold in 1960, as a statistical technique for economics. Since then, it was subjected to continuous improvement and has been used in many different fields for multivariate calibration, regression, classification, discriminate analysis and pattern recognition. Projections to latent structures by means of PLS is one of the important methods to analyse the multivariate data, where measurable relationship between a raw data matrix X and the response matrix Y is needed (Trygg and Wold 2002). PLS is a very robust technique in handling collinearity, noises and missing data in both descriptor and response matrices. PLS was subjected to continuous improvement like development of non-linear iterative partial least square (NIPALS) method, partial least square discriminate analysis (PLS-DA), OSC, etc., since the time it was proposed. Orthogonal projections to latent structures (O-PLS) is a relatively new technique, which is developed as a spectral pre-processing technique to enhance the interpretation of PLS models and relatively reduce the model complexities. Similar to OSC, the objective of O-PLS is to eliminate the precise variations in X that are not correlated to Y. In O-PLS, systematic variability in matrix X is separated into Y predictive components and Y orthogonal components. Components containing the variation that is commonly correlated with X and Y are represented as Y predictive components; these variations are in linear correlation with Y. Components that have specific variation for X that is uncorrelated or orthogonal to Y are represented as Y orthogonal (Cloarec et al. 2005; Trygg et al. 2007; Stenlund et al. 2009).
The main advantages of O-PLS method are
-
1.
O-PLS-treated data are easier to interpret, as they have limited components.
-
2.
Data are more relevant because it not only separates non-correlated variation from the dataset but also provides an opportunity to study and analyse this non-correlated variation.
-
3.
Removal of non-correlated variance from the data before modelling makes the prediction of component of interest simple.
-
4.
Data prediction ability of the model also increases.
Optimized Scaling
Optimized scaling (OS) was first introduced by Karstang and Manne (1992), as a theoretical-based method for the linear calibration of spectral data, when it does not have a fixed intensity range. They proposed two calibration methods for spectral datasets. Initially, this method gained little attention, as it is advantageous only when one constituent calibration data are available. But the second method is more generalized. Multiplicative scatter correction method uses reference spectrum (usually mean spectrum) for the spectral calibration, but these problems are not faced when optimized scaling is used. Karstang and Manne (1992) had shown the successful application of optimized scaling for the data from X-ray diffraction data of mixture of minerals, infrared spectra of organic liquids and NIR spectra of various food products. They also proposed that care must be taken while applying optimized scaling on the data with additive or baseline effects.
Spectral Interference Subtraction
Target chemical constituent identification using hyperspectral imaging is a demanding task. Hyperspectral data obtained for the analysis of chemical components is generally affected by unidentified components, component interaction, temperature, light scattering, etc. Spectral interfaces are generally caused by interference (impurities) that interacts with the analyte or constituent of interest. Spectral interference subtraction (SIS) method involves the removal or elimination of certain additive interferences from the input spectra. Martens and Stark (1991) developed SIS method as a spectral pre-processing technique for near-infrared spectroscopy and then used it for the interference correction in the field of biomedicine for speech and language processing (Hu and Wang 2011), electrocardiography (Levkov et al. 2005) and trace elements using X-rays (Donovan et al. 1993). The three main purposes of this technique are
-
1.
To remove the additive interferences caused by the presence of known constituents from the spectra.
-
2.
To maintain the changes as small as possible to the spectral data.
-
3.
To make the later regression analysis effortless to interpret.
The use of SIS as a pre-processing technique not only makes the data more effective for chemical analysis but also decreases the analysis charges and enhances the accountability of the subsequent multivariate regression analysis technique. The effective use of SIS pre-processing technique can be identified in online process control (industrial scale) where there is a difficulty to generate real calibration samples for spectral correction.
Reference-Independent Pre-Processing Techniques
Reference-independent pre-processing techniques provide more generalized tools suitable for exploratory studies, where there is no reference value available (Rinnan et al. 2009). The most commonly used spectral pre-processing techniques are broadly divided into four categories: scatter correction, normalization, derivatives and baseline correction.
Scatter Correction
Scatter correction is a statistical method to remove the scatter variations in the spectral data. Multiplicative scatter correction, standard normal variate and detrending are the most common scatter correction techniques.
Multiplicative scatter correction (MSC) also called multiplicative signal correction is the most common pre-processing technique used for scatter correction of NIR and IR spectra (Fig. 5a). Martens et al. (1983) developed MSC as a multivariate linearity transformation technique; later, it was elaborated and applied to NIR reflectance of meat by Geladi et al. (1985). The main purpose of MSC is to reduce the scatter in the spectra that is caused by various particles in the sample. Initially, it was developed as a multiplicative scatter correction technique to handle the multiplicative problems that arise from scatter alone, but later, it was used to handle diverse problems, and the abbreviation was changed to multiplicative signal correction (Rinnan et al. 2009). The MSC technique initially calculates the correction factor for the original spectra (Fig. 5b) using reference spectra (generally mean spectrum) and then corrects the original spectra using this correction factor by back transformation. It is mathematically represented as
Near-infrared reflectance intensity spectra of 10 and 18 % moisture content of Canada Western Red Spring (CWRS) wheat: a raw reflectance intensity, b multiplicative scatter corrected reflectance intensity, c standard normal variate reflectance intensity, d normalized reflectance intensity and e Savitzky-Golay smoothing and derivative corrected reflectance intensity
where x i ORG, x REF and x i CORR are the original, reference and corrected spectrum, respectively.
a i and b i are the correction coefficients of the ith sample, that are estimated by least square regression of the sample’s original and reference spectrum. In other words, a i and b i are the intercept and slope of the correction coefficient curve drawn between reference spectrum on x-axis and original spectrum on y-axis. From the mathematical derivation, it is clear that MSC does not eliminate the scatter but decreases the inter-sample variance of the scatter by implementing an additive and multiplicative transformation of the individual spectrum into the simple idealized reference or mean spectrum (Andersson 1999). Initial use of MSC was to choose the range from the spectra with no chemical information, but in NIR spectra, it is difficult to identify such a region without chemical information at all (Stordrange et al. 2002), so Martens et al. (1983) proposed that whole spectral range should be used to determine the correction coefficients. MSC is restricted by assuming that there is no chemical variation between original and reference spectrum (Wang et al. 2011). If this assumption is not made, there might be error in the estimation of correction coefficient (Chen et al. 2006). This problem was addressed by Martens and Stark (1991) and Martens et al. (2003) in their extended MSC (EMSC) technique by including the wavelength correction to the MSC. Isaksson and Kowalski (1993) had proposed piecewise multiplicative scatter correction (PMSC) by introducing spectral value correction at each wavelength and independent intercept and slope correction terms to the traditional MSC technique.
Standard normal variate (SNV) was first proposed by Barnes et al. (1989) as a spectral pre-processing technique, with an aim to remove the multiplicative interferences due to particle size of the sample and scatter and to account for the differences in the curvilinearity and baseline shift in the reflectance spectra (Buddenbaum and Steffens 2012) (Fig. 5c). The mathematical model for SNV is given as
where x i CORR and x i ORG are the SNV corrected and original spectrum, respectively.
a i and b i are the mean and standard deviation of the ith sample, respectively.
As the correction factors used in this technique are the mean and standard deviation, SNV is also called a z-transformation, centering or scaling (Otto 1998). Although SNV-corrected spectrum is technically similar to MSC (Dhanoa et al. 1994), SNV does not need any reference spectrum for the calculation of correction factors; hence, the user loses the grip on computation. As there is no least square step, SNV technique is more affected by noise than the traditional MSC (Rinnan et al. 2009). This technique makes the data more understandable as corrected spectrum has a greater linear relationship between spectrum and constituent concentration.
Trend is a statistical technique that helps to interpret the spectral data. If the data follows a particular trend, this may strongly confuse or superimpose the changes of interest. Detrending is the most popular mathematical or statistical method of removing the trends from the data. Barnes et al. (1989) introduced the detrending along with SNV technique for the NIR reflectance spectra pre-processing and which was further applied by Buddenbaum and Steffens (2012). For instance, detrending is applied for removing the long-term spectral changes in spectral time series. One of the simple detrending methods is by subtracting mean values of spectrum from each column of that spectrum. More complicated detrending methods aim at disintegrating the spectral changes into the trend (low-frequency components) and the spectra of interest (which are generally characterized by the higher frequency component) (Lasch 2012).
Normalization
Normalization is a scatter correction technique applied to scale up the spectra within its similar range (Fig. 5d). Normalization was used in vibrational spectroscopy for correcting the spectra that was affected with differences in optical path length (light path) or difference in sample quality (Lasch 2012). In normalization to equal length, spectra are generally divided by their own norms, and eventually, the length of the spectra becomes 1 (Stordrange et al. 2002):
Another method of normalization is to sum the spectral data to 1. This technique is not commonly used as the data can be misinterpreted. For example, when summing up the positive and negative regions, improper zeros result in the corrected data. Normalization had shown an increase in accuracy and efficiency of the spectral distance measurement modelling (Heraud et al. 2006).
Derivative
Resolution of IR spectra is well enhanced by spectral derivatives. Savitzky-Golay smoothing and derivative and Norris-Williams derivation are the most common spectral derivative techniques used with an aim to resolve and remove the overlapping bands in complex IR spectral signals.
Savitzky-Golay (SG) smoothing and derivative technique was first proposed by Savitzky and Golay (1964) and further elaborated by Steinier et al. (1972). Initially, this technique was used for continuous and equally spaced data. Evaluation of derivative is achieved by executing the data through a window-wise symmetric filter (Brown et al. 2000) (Fig. 5e). In this technique, the spectrum is elaborated with a window having 2d + 1 points, where each window is used for the estimation of the centre point and with d points on each side. These 2d + 1 points are fitted by a polynomial of a given order, and the coefficients found by this fit are used for the estimation of the new value at this wavelength (either just by smoothing or by both smoothing and differentiation) (Rinnan et al. 2009). The main benefit of SG technique is that it can carry out smoothing, noise reduction and computing derivatives in one single step. The method proposed by Savitzky and Golay (1964) contains some numerical errors which were addressed and corrected by Steinier et al. (1972).
Norris (1983) introduced the derivative technique, which was later modified by Norris and Williams (1984). This technique was named the Norris-Williams (NW) spectral derivative technique (Norris 1983; Norris and Williams 1984). The principle behind NW technique is to smoothen the spectral data based on a moving average (Massart et al. 1997) over data points, and the gap between these data points is used to estimate the derivatives, and then, finite difference is calculated based on this smoothing spectrum. Unlike the SG method, the NW method is less prone to high-frequency noise, as it uses both smoothing by moving average and gap size for derivative. Application of NW pre-processing technique (gap size for derivatives part) for spectral data is not properly defended as the data are not presented in a time domain.
Baseline Correction
Reflectance or transmittance IR spectra often contain unwanted background features or noise additionally to the desired signal (Schulze et al. 2005). Spectral noise is caused due to scattering, external factors causing variations in data acquisition (illumination, temperature, etc.) and instrumental noises. To acquire proper information from the spectral data, it is important to remove this noise or background features from the signal. Baseline correction is a pre-processing technique used to eliminate these background noises from the spectral data, thus making a more easy illustratable signal. It also generates more accurately predictable spectral parameters like band positions and intensity values (Mazet et al. 2005). Most common baseline correction techniques are offset correction, detrending (Barnes et al. 1989), SG technique (Savitzky and Golay 1964; Steinier et al. 1972) and iterative polynomial baseline correction (IPBC) (Lieber and Mahadevan-Jansen 2003). As SG technique and detrending are discussed in previous sessions, the focus is on remaining techniques in this section. Offset correction is the easiest baseline correction technique. Offset correction is conducted by subtracting a linear horizontal baseline from the original spectrum. This horizontal baseline value is selected in such a way that no less than one point of the corrected spectrum equals to 1 and spectra remain unscaled (Lasch 2012). Instead of using simple straight horizontal baseline as offset correction, an nth-order iterative polynomial function is used by IPBC in order to fit the spectral data points. To prevent baseline correction artefacts, a lower order polynomial is recommended for IPBC (Lasch 2012).
Types of Information Extracted from Hyperspectral Data
The hypercube or datacube collected using high-performance hyperspectral imaging can be visualized in two different ways: one along all wavelengths of the imaging spectrum to obtain spectral data and another at each wavelength to obtain spatial data. Using these spectral and spatial data, either sample qualitative (e.g., detection of defects) or quantitative information (e.g., quantification of chemical composition and contamination) can be gathered.
Qualitative Analysis
Qualitative analysis of food and agricultural products includes analysis of chemical composition of the materials, detection of defects and purity of materials present. Hyperspectral imaging technique has proved to be a non-destructive analytical technique for the non-destructive analysis of food quality and safety (Wu and Sun 2013b). Qualitative information about the sample can be obtained when spectral data of a hypercube are analysed using proper chemometric techniques. Using the entire spectra for the qualitative analysis is time-consuming and burdensome on the statistical software. In order to overcome this, exploratory data analysis techniques are used to select indispensable wavelengths from the excessive data for multispectral analysis. When selecting the indispensable wavelengths, care should be taken to avoid loss of crucial data. Qualitative parameters are generally analysed using data exploratory techniques or supervised or unsupervised classification techniques. Out of all chemometric techniques, classification methods are the basis for the qualitative analysis of chemical composition and purity of the component in the food and agricultural sample. Classification methods are machine learning techniques that develop mathematical models that can recognize each member of the sample to its appropriate class on the grounds of some set model parameters. Once the model is developed, it has the capability to read the unknown sample using its model parameters and later assign it to one of the classes. Classification techniques are extensively used in the fields of chemistry, process monitoring, medicine, pharmacy, social sciences, economics and food science (Ballabio and Todeschini 2009). Rapid quality inspection is of major concern in automated food and agricultural industries. Classification modelling is the most appropriate technique to handle these concerns.
Quantitative Analysis
Initially, the use of hyperspectral imaging in the field of food and agriculture started to assess the quality of the product. Quantification of various chemical constituents in food and agricultural samples can be analysed using categorical regression techniques. Hyperspectral regression models are developed using known concentrations of chemical constituents, and once the model is developed and validated, analysis of an unknown sample is simple using this model. Detection of various materials using hyperspectral imaging involves identification and quantification of those in a spatial image using their respective spectral features. Every pixel of the identical sample in an image has identical spectral signature. Identification and quantification of various materials using hyperspectral imaging is a challenging task. Hyperspectral imaging applied along with image processing and chemometrics makes the detection and quantification of materials possible. Chemometric techniques like principal component regression (PCR), multilinear regression (MLR) and partial least square regression (PLSR) are generally used to measure various food components quantitatively. Regression models are quantitative feedbacks on the basis of a set of descriptive variables (Ballabio and Todeschini 2009). Regression analysis helps to develop an association between the hyperspectral data and the measured property of the material. This mathematical expression relating the systematic responses or signals to the constituent concentration of the sample is also known as the calibration equation (Mark and Workman 2003).
Chemometric Tools for the Extraction of Information from Hyperspectral Data
Exploratory Data Analysis
Exploratory data analysis technique when used as a statistical method can recognize the main characteristics of the data and see what the data tells us beyond classification modelling and regression analysis. Exploratory data analysis technique was promoted by John Tukey as a method to see what the data can offer for the researcher other than the general modelling for performing qualitative analysis (Tukey 1980). In a broad sense, these methods are used for detecting the data errors, cross validating the assumptions and selecting a suitable model. In hyperspectral imaging, these are also used to reduce the amount of information in the data. Most commonly used exploratory data analysis methods are principal component analysis (PCA) and independent component analysis (ICA).
Principal component analysis is a method to transform the dataset linearly into smaller dimension dataset with variables that are uncorrelated with each other. It was proposed for the first time by Pearson (1901) and Hotelling (1933) to explore the invisible structures in the data that assist in more accurate classification. In order to achieve this, the data are rotated in the sample space of observation. PCA is an important step for the identification of key wavelengths and reduces the massive hyperspectral imaging dataset. This reduction in data makes it possible to use in online sensing. PCA is used for feature reduction by changing data to a new set of axes and creating subsets, and these subsets show higher differentiability (when compared to the original data subsets) (Jiang et al. 2010). Projection-based methods like PCA are generally applied for data dimension reduction and feature selection problems.
Principal components are the new components that are generated by orthogonal linear transformation of the original dataset. The new components are formed by rotating the data so that the first principal component has the highest variance by any projections of the dataset; the second principal component has the second highest variance by any projection of the dataset and so on. Hyperspectral dataset (hypercube) H with X × Y × λ dimensions is decomposed using PCA into a set of scores and loadings (Amigo 2010). Transformed dataset can be represented in matrix form as
where H I the transformed dataset with dimension XY × λ, S is the score surface with dimension XY × F, L S is the loading profiles with dimension F × λ and E is the residual matrix with dimension XY × λ. The main advantage of PCA is that the number of principal components can be ascertained from the principal component score, variance and factor loading results. The loading profile and the score surface of each principal component can be an association of pure principal component, a physical effect or even a combination of both (Amigo et al. 2008). These properties of PCA make it a very robust technique to handle the massive hyperspectral data. PCA is the most useful exploratory data analysis technique for food and agricultural materials (Xing et al. 2007; Vargas et al. 2005; Barbin et al. 2012; Cho et al. 2013). Van Der Maaten et al. (2009) provided a comparative review between linear (PCA) and non-linear (multidimensional scaling, maximum variance unfolding, isomap, diffusion maps, multilayer autoencoders, kernel PCA, manifold charting and locally linear coordination) dimensionality reduction techniques and concluded that non-linear techniques, although they have large variance, are not capable of outperforming the PCA technique.
Independent component analysis (ICA) is another exploratory data analysis and hyperspectral image feature selection method. It is also used for feature extraction, pattern recognition and unsupervised classification. It was proposed for the first time by Herault and Jutten (1986) and further developed by Comon (1994). It is a method to separate spectral signal into a number of additive subcomponents. These subcomponents are believed to be statistically independent (occurrence on a subcomponent does not affect the other) of one another and are not following Gaussian function. It obtains the independent source signals by searching for a linear or non-linear transformation that maximizes statistical independence between components (Jiang et al. 2010). Before application of the ICA algorithm on the hyperspectral data, it is recommended to centre, whiten and reduce the dimension of the data (Polder et al. 2003). Data centring is done to simplify the ICA algorithm and is attained by subtracting the mean of the components of the vector from every component. Whitening transformation technique transforms the data so that they have the identity covariance which suggests that all dimensions of the matrix are statistically independent and the variance along each of the dimension is equal to 1. Whitening and data reduction can be done using PCA; hence, data are processed using PCA before applying ICA. Du et al. (2003) and Botchko et al. (2003) had used ICA for hyperspectral image processing primarily to reduce the dimensionality and select the specific bands for feature extraction. Polder et al. (2003) had applied ICA for tomato sorting using visible spectral images in the spectral range of 400 to 710 nm. Application of ICA for hyperspectral images was not as successful as PCA (Table 1), as it is necessary for defining the number of independent components prior to the computational analysis for ICA, which is not a usual practice with PCA.
Unsupervised Learning
Unsupervised machine learning techniques are used to explore the hidden structures in a hyperspectral dataset. The main difference that separates unsupervised from supervised machine learning techniques is that unsupervised techniques when dealing with the unlabelled data deliver an underlying structure of that data. As there are no generated response data for comparison of results, this technique cannot generate the error terms. Unsupervised learning will only have the original input data to work on. Two most common unsupervised machine learning methods are dimensionality reduction and clustering. PCA and ICA are the classic examples of dimensionality reduction techniques. PCA and ICA disintegrate the spectral data into a number of components (principal or independent) in order to distinguish the key directions of variability in the high-dimensional data space (Wu and Sun 2013b). The first few components of PCA and ICA carry most of the information that can accurately distinguish between the samples.
The K-means, fuzzy and hierarchical clustering are the classic examples of clustering techniques. The K-means clustering technique clusters input data into k cluster groups of equal variance. Samples belong to their respective cluster group by minimizing their distance to the cluster centroid. K-means uses the centroid of the cluster as the criterion to assign the cluster for each sample and is achieved by minimizing the sum of squared errors (Ding and He 2004). K-means is also called hard clustering as it verifies whether the object belongs to the cluster or not and the number of clusters is initially specified. In contrast, fuzzy clustering method assigns the samples to different clusters simultaneously with varying degrees of membership (Amigo et al. 2008). Hierarchical clustering is a method to build a hierarchy of clusters that is normally presented in the form of a binary tree diagram, commonly known as dendrogram. The main principle of hierarchical clustering is to group the sample to a cluster by measuring the distance between the two consecutive samples. The hierarchical clustering is not suitable for large datasets (Wu and Sun 2013b). Among the clustering techniques, fuzzy clustering is more prominently used for hyperspectral data analysis. Although clustering techniques are very robust, their use for hyperspectral data analysis is very limited as they lack the ability to guess the correct number of clusters to be developed (Amigo et al. 2008; Lopes and Wolff 2009). Table 2 shows the limited application of unsupervised learning techniques for food and agricultural products using hyperspectral spectral imaging technique.
Supervised Learning
Supervised learning is a machine leaning technique which groups the unknown samples into the known predefined groups as per their measured features (Wu and Sun 2013b). Supervised learning is a process of understanding a set of rules from the instance (training set) with an aim to create a classifier that can be used upon new instance (Kotsiantis 2007). A training set consists of pairs of input vectors and desired output. The output of the classifier can be a model to predict unknown samples (for regression) or can anticipate a class label of the input vectors (for classification). In other words, supervised learning evaluates input testing set to develop function that can be used for mapping the new set (testing set). Supervised learning is the most successful statistical technique used along with hyperspectral imaging in food and agricultural applications. Kotsiantis (2007) provided an extensive review on different supervised machine learning techniques and their use in classification of various real-world problems. Table 3 shows the extensive application of supervised learning techniques for food products using hyperspectral spectral imaging technique.
Discriminant Analysis
Discriminant analysis is by far the well-known and extensively used classification method (McLachlan 1992). It is a supervised learning technique which uses discriminant analysis function to assign a dataset to one of the previously established groups. Discriminant analyses are very robust, but they overfit the multicollinear data (correlated among themselves) (Hand 1997). Data reduction using stepwise discriminant analysis (SWDA) (Jennrich 1977) or PCA (Pearson 1901; Hotelling 1933) can resolve this problem. Quadratic, linear and partial least square discriminant analyses are a few important discriminant analysis techniques.
Like PCA, discriminant analysis technique separates samples into different classes by maximizing the variance between classes and minimizing the variance within a class. Mathematical relationship of this statement can be framed from considering a classic classification problem, where a test sample x i is designated to one of the prior defined class C based on j measurements [x i = (x i1 , x i2 .....x ij )T]. The discriminant function (d f ) (classification score) is given below, and the test sample is designated to the class which has the minimum classification score (Wu et al. 1996):
The first two terms of this equation express the between the test sample x i and centroid x̄ c (Mahalanobis 1936) where Σ c is the class C’s covariance matrix and calculated by
x̄ c is the centroid of class C and calculated by
πc is the class probability of class C and calculated by
where n c and n are the total number of samples in class C and training set.
Quadratic discriminant analysis (QDA) classifies the samples into the classes with quadratic-shaped boundaries and assuming that multivariate normal distribution is common in each class (Ballabio and Todeschini 2009). Mahesh et al. (2008) and Choudhary et al. (2009) applied QDA for the classification of wheat classes using hyperspectral imaging, and Singh et al. (2009, 2010) applied QDA for the classification of midge-damaged and insect-damaged wheat using hyperspectral imaging.
Linear discriminant analysis (LDA) is considered as the special case of discriminant analysis, first proposed by Fisher (1936). LDA follows all the principles of discriminant analysis, but it uses pooled covariance Σ p as it assumes that the covariance matrices of the class Σ c are equal.
Pooled covariance matrix of the class is given as
For classification of samples into different classes, LDA develops linear projections of independent variables (Wu and Sun 2013b). Although it is similar to PCA in considering the independent variables, LDA also contains class information of samples. Like QDA, LDA also assumes that multivariate normal distribution is common in each class, but the covariance matrices of the classes are equal; hence, both LDA and QDA are assumed to work well if the sample classes are normal. The main disadvantage of LDA is that it does not hold well with the condition where the number of samples is less than that of number of variables, as in this condition, the inversion of covariance matrices becomes difficult. Hence with LDA, it is always preferred to have more samples than the number of variables. As QDA generates covariance for each class, it can handle or require more number of samples than LDA. QDA and LDA are the most commonly used classification techniques of hyperspectral imaging data from food and agricultural products. Wang and Paliwal (2006) used LDA, QDA, k-nearest neighbour (KNN), probabilistic neural network (PNN) and least squares support vector machines (LS-SVM) for the classification of hyperspectral images of six Canadian wheat classes and confirmed that LDA gave the highest classification accuracy. Gowen et al. (2009) developed a technique with the combination of LDA and hyperspectral imaging method for accurately identifying freeze-damaged mushrooms.
PLS-DA is a supervised learning technique that combines the regression power of PLS and classification power of discriminant analysis. PLS-DA based on their spectral fingerprint assigns the unknown samples to one (and only one) of the available categories. This technique is derived from PLS regression (Wold et al. 1983) and is referred as a secondary data analytical step involving the construction of statistical classification models (Baker and Rayens 2003). PLS was designed for regression analysis (Wold 1975), but recently, it was combined with LDA and used as a discriminant technique. PLS-DA coincides with the inverse least square approach to LDA and gives the same result as that of LDA but with less noise and the advantage of variable selection (Ballabio and Todeschini 2009). There are two kinds of PLS algorithms: PLS1 algorithm deals with one dependent Y variable, and PLS2 deals with more than one dependent Y variable. PLS-DA is a PLS2 algorithm (many Y variables) that hunts for abstract (or latent) variables that have maximum covariance with Y variables (qualitative in nature). The Y block represents whether the sample is classified correctly or not. In PLS-DA classification with two classes, Y variable will be given 1 if the sample is classified correctly or else given 0 if it is wrongly classified. When dealing with multiclass classification, for each sample, a PLS2 algorithm for multivariate qualitative response is applied, and it will return prediction value between 0 and 1 for each class. If the value is closer to 1, then it indicates that the sample belongs to that particular class or else not. A threshold between 0 and 1 is used to assign a class for the sample. The use of PLS-DA as a classification tool for hyperspectral imaging of food and agricultural products is a recent development. Pearson et al. (2001) used NIR spectroscopy and PLS-DA for accurately (95 %) detecting aflatoxin-contaminated corn kernels. Serranti et al. (2013) used PLS-DA for accurate (100 %) classification of groat and oat kernels using hyperspectral imaging in the near-infrared range from 1006 to 1650 nm. Vermeulen et al. (2011) used PLS-DA for the online identification of contaminants in cereals by near-infrared hyperspectral imaging.
Categorical Regression
Regression is a statistical procedure which determines the relation between dependent variable and independent variable. Categorical regression calibrates the data by accrediting numerical values to classes, which results in optimal linear regression equation for the transformed variables. PCR and PLSR are the best-known categorical regression techniques used for extracting information from hyperspectral imaging data.
Multicollinearity is a statistical procedure where several independent variables participating in multiple regression modelling are highly correlated to one another. Due to multicollinearity, least square of the variables is unbiased, but they have more variance between them. PCR is a multivariate regression analysis technique for analysing data with high multicollinearity between their variables. PCR reduces the standard error by adding bias to the regression estimates and is hoped that more reliable estimates will be achieved due to this overall effect. The simple matrix notation of PCR is given as follows:
where Y is the dependent variable matrix (concentration matrix), X is the independent variable matrix (hyperspectral imaging data), Ḇ is the coefficient of regression and E is the error or the residual matrix. As PCR considers the variances between variables, more freedom is provided in variable selection, and hence, it avoids the problem during further mathematical calculations. PCR model developed using the independent and dependent variables sometimes gives a random error or noise rather than giving the anticipated relationship (model outfitting). This can be avoided by choosing optimal number of PCs. The main problem with PCR is that the PCs that describe the independent variable matrix (hyperspectral imaging data) may not be the perfect PCs for predicting the concentration of the sample. This problem can be resolved by applying an intermediate correlation coefficient calculation step between sample concentrations and the PC scores and selecting those components that are significantly correlated for subsequent regression (Romia and Bernardez 2009). Cogdill et al. (2004) developed PCR models to accurately predict the moisture content and oil content of single corn kernels using hyperspectral absorbance spectra in the wavelength region of 750–1090 nm.
PLSR is a well-known chemometric tool that is used to estimate the biological and chemical properties of the sample from their hyperspectral spectrum. PLSR is extensively used for the study of massive hyperspectral data and was introduced for the first time by Wold (1975) for the field of econometric as an alternative to the general least square regression to handle the data with variables showing high collinearity. Similar to PCs of PCR, when applied to create a relationship between concentration matrix Y and hyperspectral matrix X, PLSR develops latent variables known as PLSR components (PLSRCs) which are in linear combinations with variables in matrix X. Unlike PCR, PLSR ensures that the first few of its PLSRCs contain as much information as possible to classify the matrices X and Y. They also manage to explain most of the X and Y variances during calibration and compression. As PLSRCs are corrected for maximizing the prediction capability of matrix Y, they will not match fully with the direction of maximum variation (Romia and Bernardez 2009). The simple matrix notation of PLSR is given as follows:
where Y is the dependent variable matrix (concentration matrix), X is the independent variable matrix (hyperspectral imaging data), X p and Y p are the matrices of the X scores and Y scores, respectively, P and Q are the orthogonal loading matrices and E 1 and E 2 are the error or residual matrices. Similar to PLS-DA, PLSR also has two kinds of algorithms: PLS1 to handle only one response variable and PLS2 to handle more than two (multiple) response variables. PLS1 is generally used to determine concentration of only one component of Y, and PLS2 is used to determine the concentration of multiple components of Y. In the recent past, extensive literature on the application of PLSR for extraction of useful information from hyperspectral images of food and agricultural materials is available. Wu and Sun (2013b) reviewed the use of PLSR with hyperspectral imaging system for meat and meat products, fish (Xu et al. 2016a,b), fruits and vegetables’ quality and safety analysis.
Naïve Bayes Classifier
Naïve Bayes (NB) classifier is one of the most popular probabilistic statistical classification methods. NB classifier is based on application of Bayes theorem (Bayesian statistics) with few firm assumptions. The main assumption made for this classifier is that all the features are independent of other features specifying the state of class variables (Langley et al. 1992; Friedman et al. 1997). Decision in NB classifier is made by assuming that the availability or non-availability of a particular feature is always unrelated to the availability or non-availability of any other feature of a class. NB considers that these individual features contribute to the probabilistic theory in order to judge whether the sample should be categorized to a particular class or not. Under supervised learning conditions, NB classifier can be trained very precisely as it depends on the effective nature of probability model. Maximum likelihood method is used for the estimation of parameters for NB models. Due to this special character of NB, one can use it for classification without believing in Bayesian probability or using any Bayesian methods. NB classifier had performed very well in solving various real-world problems using hyperspectral imaging despite of its speculation that all the features are independent. Main advantages of NB classifier is that very small training set is enough to determine the parameters (e.g., mean and variance) needed for classification. As the main assumption is to have independent variables, variance of the variables for each class is required but not the covariance of the entire matrix.
Support Vector Machine
Support vector machine (SVM) is a linear supervised machine learning technique which relies on the concept of hyperplanes that define decision boundaries. Hyperplane is a decision plane that separates set of samples to different classes. The concept of SVM was introduced by Cortes and Vapnik (1995) and later used for data mining by Burges (1998) and Duda et al. (2001). In SVM, the hyperplane separating two classes is supported by subsets called support vectors. Classification rule is drawn using these support vectors (subsets of training set that are lying in the nearness of the boundary between two classes). Due to this, the result of the SVM classification is contingent only on the support vectors, and this result does not vary when some of the samples other than these support vectors are removed. SVM algorithm during optimization searches for the hyperplane with maximum margin (distance between hyperplane and the support vectors) to give the ideal separating plane for classification. Generally, SVMs are the linear classifiers, if non-linear separable classes are available; it is difficult to use the linear classifier for accurate classification (Ballabio and Todeschini 2009). In order to handle this problem, SVMs are integrated with one of the non-linear kernel functions (polynomial, Gaussian and sigmoid kernels). SVMs are generally applied for binary classification problems; multiclass classification is possible by taking into account one class at a time and developing classifier that separates this class from all the other classes. Then during testing, a sample is assigned to the nearest class and decision function is used to formulate the distance from each class.
The SVMs are very successful as supervised classifiers for object recognition (Guo et al. 2000) and face detection (Osuna et al. 1997). Growth of interest in SVMs is revealed by their successful application in food and agricultural analysis for classification of starch (Pierna et al. 2005), compound feeds (Pierna et al. 2006), wheat (Wang and Paliwal 2006) and corn (Zhang et al. 2012).
Artificial Neural Networks
An artificial neural network (ANN) is the most popular chemometric technique that has gained importance in the chemical analysis application. ANNs are robust and can handle unsupervised (clustering) and supervised (modelling) problems and can work with both qualitative and quantitative analysis. It is important to review the nature of a problem and then consider the best neural network (NN) strategy to solve it, since different NN constructions and different NN learning programs have been suggested in the literature (Zupan 1994). ANN works like the human nervous system, where each neuron receives a signal from neighbouring neurons, later executes them and finally gives out the output signal (Eluyode and Akomolafe 2013). The function which computes the output vector from the input signal consists of two parts:
-
1.
Evaluation of net input, and it is the linear product of input variables and their weights.
-
2.
Transferring the net input to output vector in a non-linear way (Ballabio and Todeschini 2009).
The number of neurons used in ANN may vary from ten to several thousands and are based on the analysed data (training set) (Zupan and Gasteiger 1993). ANN has been successfully applied for hyperspectral imaging in the field of food and agriculture for classification of wheat classes (Mahesh et al. 2008; Choudary et al. 2009) and detection of fungal-damaged wheat (Wang et al. 2003).
Advantages and Disadvantages of Hyperspectral Imaging
Hyperspectral imaging has revolutionized the quality assessment for food and agricultural materials by providing both the quantity (spatial) and quality (spectral) of the product at the same time. The advantages of using hyperspectral imaging setup for quality assessment are as follows:
-
1.
Hyperspectral imaging does not require any separate sample preparation.
-
2.
It is a non-damaging technique; hence, sample can be saved for further analysis.
-
3.
Once the model is developed, calibrated and validated, analysis of unknown is simple.
-
4.
Relatively safe to the environment, as no chemicals are required for analysis.
-
5.
Economical when compared to conventional chemical analysis methods, as it reduces additional costs for reagents, labour and waste treatment.
-
6.
Collection of massive spectral information for every pixel provides more accurate information about the sample chemical constituents and provides a chance to refine the data and perform critical analysis.
-
7.
As every pixel in the hypercube has similar number of spectral bands, selection of region of interest is flexible.
-
8.
Both subjective and measurable study can be done using the same images.
-
9.
Multiple constituents can be analysed from the same images.
-
10.
Further processing allows us to understand the chemical constituents of the material and generally called chemical imaging (ElMasry and Sun 2010).
-
11.
Although samples are similar in morphology, colour, etc. (e.g., Canadian wheat classes), very small chemical variations are sufficient enough to classify them accurately (Mahesh et al. 2010).
-
12.
If the most prominent wavelengths for classification are identified, data processing time can be reduced drastically.
Despite its advantages, hyperspectral imaging also has some constraints or disadvantages:
-
1.
Hyperspectral imaging system is expensive when compared to other image processing techniques.
-
2.
Because the hyperspectral data are massive, the process requires large capacity drives for data storage and high-speed computers for data processing.
-
3.
As many images are collected all along the spectral range, this technique requires more time than traditional digital imaging system.
-
4.
Requires efficient and powerful correlation models to understand the minor chemical constituents.
-
5.
Development of online monitoring system is difficult as data acquisition and analysis is time-consuming.
-
6.
Imaging is affected by external environment like illumination, scattering, etc., hence produces poor signal-to-noise ratio.
-
7.
Spectral pre-processing techniques are required in order to curtail the consequences of external factors like light scattering, noises, etc.
-
8.
Liquid samples are difficult to handle as the imaging is affected by heterogeneity within the sample.
-
9.
From analyst point of view, regulated calibration and precise model generation are needed as hyperspectral imaging is an indirect analysis technique (ElMasry and Sun 2010).
-
10.
Identification and detection of different objects within the same image using spectral data is challenging unless different objects have different absorption features.
-
11.
Cannot detect microcompounds as accurately as GC and HPLC.
Hyperspectral imaging is an effective technique, but this system is highly influenced by various external factors like variations in illumination system, noises in instrumentation, etc.
Applications of Hyperspectral Imaging for Food and Agro Products
Hyperspectral imaging technique can be used to obtain spectrum of sample over a certain wavelength range. This spectrum delivers the complete chemical signature of the sample at the pixel level. The meaningful information from the spectrum can be obtained using different chemometric techniques. In this section, hyperspectral imaging technique applications for food or agro products and infield agricultural operations were summarized.
Food and Agro Products
Hyperspectral imaging technique was successfully applied to solve various qualitative and quantitative problems for food and agro products.
Xing et al. (2010) compared the SWIR hyperspectral imaging with Fourier-transformed near-infrared (FTIR) spectrophotometer in the 1235–2450 nm wavelength range for predicting the alpha-amylase activity in the individual wheat kernels. The SWIR imaging system was used to acquire the reflectance spectra, and FTIR was used to acquire absorbance spectra of the same wheat kernels. Both SWIR and FTIR spectra were pre-processed using area normalization, SG second derivative and EMSC pre-processing techniques. The unprocessed SWIR hyperspectral imaging system gave similar r 2 and root mean square error (RMSE) values as that of the FTIR imaging system. The SWIR hyperspectral imaging when used along with SG second derivative pre-processing technique gave the highest r 2 value of 0.88 when compared with any other combination of imaging system and pre-processing technique.
McGoverin et al. (2011) investigated the viability of barley, wheat and sorghum grains using NIR hyperspectral imaging. Hyperspectral images of cereal grains were obtained in the 1000–2498 nm wavelength range. The acquired reflectance spectra were transformed to pseudo-absorbance before applying the SNV data pre-processing technique. The PCA was applied to this pre-processed data to identify and remove unwanted data points, and partial least square discriminant analysis (PLSDA) was conducted to discriminate the non-viable from viable cereal grain samples. It was identified that the use of PLSDA eliminated subjective assessment of the patterns in the PCA score plots. To overcome this, PLSR was applied to the dataset. It was identified that the PLSR model required larger datasets to improve the viability prediction.
Bauriegel et al. (2011) applied hyperspectral imaging technique for the early identification of the Fusarium infection in wheat. Hyperspectral images of wheat ear infected with Fusarium were acquired when the wheat ears were still on the plant in 400–1000 nm range. PCA was applied to differentiate the infected and healthy wheat ear tissue spectra. The healthy and diseased samples were successfully classified with 87 % accuracy when the Spectral Angle Mapper image analysis technique was applied to the data.
Ariana and Lu (2010) used hyperspectral imaging technique for identifying the surface defects of whole pickles. Hyperspectral images of the pickles were acquired in the reflectance and transmittance mode in the 400–675 and 675–1000 nm, respectively. The first three principal component (PC) scores obtained by applying PCA were used to select the most contributing factors for the discrimination of the good and defective pickles. These PC scores contributed for 95 % of variation between the good and defective pickles.
Kamruzzaman et al. (2013) used NIR hyperspectral imaging technique in the 900–1700 nm range for the discriminating three types of lamb muscles (semitendinosus, longissimus dorsi and psoas major). PCA technique was applied on the hyperspectral data, and the six most prominent wavelengths (934, 974, 1074, 1141, 1211 and 1308 nm) that show variation between lamb muscles were selected. Using these wavelengths, the NIR hyperspectral imaging accurately (100 %) discriminated the lamb muscles.
Cho et al. (2013) investigated the feasibility of hyperspectral fluorescence imaging applied in 400–700 nm wavelength range to identify the cuticle crack defects on cherry tomatoes. Defective cherry tomatoes with existing cracks were harvested from the field and used for acquiring fluorescence emission spectra using pushbroom imaging system. PCA was applied on the spectra, and it was identified that PC1 through PC4 loadings had contributed to 99.21 % of variation between good and defective areas of tomatoes. The wavelengths 503, 670 and 689 nm were determined to be the best wavelengths for detection of defective areas on tomato surface.
Kong et al. (2004) used hyperspectral fluorescence imaging for the detection of skin tumours on poultry carcasses. Sixty-five spectral bands were acquired in the visible region (425–711 nm) to construct the hypercube. PCA was applied for reducing the dimensionality of the data, and later, fuzzy hierarchy technique was applied for identifying the skin tumours on the poultry skin. Fuzzy hierarchy technique in combination with hyperspectral imaging system was 82 % accurate in the detection of the skin tumour regions on the carcass skin.
Cogdill et al. (2004) evaluated the quality of single corn kernels using NIR hyperspectral imaging in the Vis-NIR region. Vis-NIR absorbance spectra were obtained in the wavelength range of 750–1090 nm and pre-processed using SNV, detrending (DET), MSC and wavelength selection by generic algorithm techniques. PLSR and PCR models were developed using Vis-NIR absorbance spectra to analyse the moisture and oil content. Results showed the cross-validation standard error of 1.20 % for moisture and 1.38 % for oil content.
Pierna et al. (2006) used hyperspectral imaging in NIR range to develop a procedure to screen compound animal feeds. The SVM model was used to generate discriminant equations using hyperspectral images. Classification tree method was used to identify the composition of the sample. Higher classification accuracy (99–100 %) was obtained in classifying the composition of the sample.
Zhang et al. (2007) used NIR hyperspectral imaging to identify the wheat kernels infected with three storage fungi (Aspergillus glaucus, Aspergillus niger van Tieghem and Penicillium spp.). Hyperspectral images were acquired in the NIR range of 1000–1600 nm; PCA was used to reduce data and extract the most useful wavelengths. Classification model was developed using SVM, and classification accuracies of 100, 87.2, 92.9 and 99.3 % were obtained for identification of healthy, A. glaucus, A. niger van Tieghem and Penicillium spp.-infected samples, respectively.
Mahesh et al. (2008) used NIR hyperspectral imaging to classify Western Canadian wheat classes. NIR reflectance spectra were obtained in 960–1700 nm wavelength range at 10-nm intervals. Statistical and ANN models were developed using NIR reflectance spectra, and classification accuracies greater than 94 and 90 % were obtained for statistical and ANN classifier, respectively.
Choudhary et al. (2009) identified wheat classes using NIR hyperspectral imaging. NIR reflectance spectra were obtained in the wavelength range of 960–1700 nm. Wavelet features of the bulk sample images were extracted and used to develop LDA, QDA and BPNN models. Wheat classes were classified with accuracy of 99.1 and 92.1 % using LDA and BPNN models, respectively.
Singh et al. (2009) used NIR hyperspectral imaging for detection of insect infestation in wheat. Hyperspectral images of single wheat kernels healthy and infested with Sitophilus oryzae (L.), Rhyzopertha dominica (F.) and Cryptolestes ferrugineus (Stephens) were acquired in the wavelength range of 1000–1600 nm. Multivariate image analysis (MVI) was used to reduce the dimensions of hyperspectral data. Spectral features (statistical and histogram) were used to develop LDA and QDA models. The infested and uninfested wheat kernels were identified with the classification accuracies in the range of 85–100 %.
Williams et al. (2009) classified the maize kernels based on the hardness using two IR hyperspectral imaging systems. NIR hyperspectral images were acquired in the wavelength range of 960–1662 nm, and SWIR hyperspectral images were acquired in the wavelength range of 1000–2498 nm. PCA was used to remove the background, bad pixels and shading effects from absorbance images. PLSDA models were developed using PC2 of SWIR hyperspectral images and PC3 of NIR hyperspectral images. The developed models gave better differences between glassy and floury endosperms of maize kernels. PLSDA model gave a root mean square error of prediction of 0.18, 0.18 and 0.29 for 12-kernel, 24-kernel NIR hyperspectral image and SWIR image, respectively.
Singh et al. (2010) used NIR hyperspectral imaging and conventional digital imaging to identify the midge-damaged wheat kernels. LDA, QDA and Mahalanobis models were developed using the statistical and histogram features of hyperspectral images in the wavelength range of 700–1100 nm. The highest classification accuracies of 95.3–99.3 % were obtained to classify healthy and midge-damaged wheat kernels when histogram features were combined with ten colour image features.
Pierna et al. (2012) conducted two separate case studies using NIR hyperspectral imaging system and chemometric techniques for the detection of impurities in cereals and contamination of plants by pathogens. For the identification of the impurities in cereals, hyperspectral images of wheat, spelt, barley and rapeseed in combination with various impurities like the cellulose waste (wood and straw), animal contaminants (insects) and other contaminants (paintings, plastics and stones) using pushbroom NIR hyperspectral camera in the 1100–2400 nm spectral range. Quadratic SVM algorithm successfully differentiated the contaminants from the cereal grains with greater than 92 % accuracy. For discriminating the plants into tolerant and susceptible base in the presence of nematode cysts on the sugar beet plants, NIR hyperspectral images of the plant roots with varying numbers of cysts were acquired using whiskbroom NIR hyperspectral camera in the 900–1700 nm range. The spectra were pre-processed using SNV and second derivative SG techniques. On an average of 40 tolerant and 69 susceptible plants whose cysts were identified under microscope, the NIR hyperspectral imaging system with SVM discriminant technique correctly identified 21 tolerant and susceptible plants.
Hyperspectral imaging system was successful in qualitative and quantitative analysis of various food and agricultural products. Each study used different combination of imaging mode, imaging system, imaging wavelength range and chemometric technique. Various applications of hyperspectral imaging for food and agro products were summarized in Tables 1, 2 and 3. A separate section (Tables 4 and 5) was given to summarize the applications of hyperspectral imaging system for real situations where the hyperspectral imaging system and its models can be applied to the real situations directly or with minor modifications.
Agriculture and Agricultural Operations
Hyperspectral imaging was very successful for solving various problems in the field of agriculture. This technique was applied for both infield and laboratory studies. The fact that banana skin colour changes from green to yellow based on its maturity was used to develop a mobile phone-based two-dimensional spectral image analysis.
Du et al. (2013) investigated the carbonate content in soil samples using FTIR photoacoustic spectroscopy (FTIR-PAS) in the mid-infrared range (2500–20,000 nm). PCA, PLSR and generalized regression NN models were developed and tested to predict the soil carbonate content. Out of the three chemometric techniques, the generalized regression NN performed gave better prediction of the carbonate content results with validation error percentage or the root mean square error percentage of 1.21 %, determination coefficient of 0.899 and ratio of standard deviation to prediction error of 3.83.
Kemps et al. (2010) investigated the application of visible near-infrared (Vis-NIR) reflectance spectroscopy for the assessment of various chemical parameters of grapes during the growing season. The Vis-NIR reflectance spectra of the intact grapes of four varieties were acquired using spectrophotometer in the visible and near-infrared range of 320–1660 nm during the growing season. Predictive models were developed using PLSR for the prediction of anthocyanin, polyphenol, sugar and density of the intact grapes. The anthocyanin content remained fairly unchanged, and hence, it was not possible to estimate the anthocyanin content. The polyphenols and density were accurately predicted using this technique, but the prediction of sugar content was possible as the grapes also shown variation in the colour as they mature.
Suzuki et al. (2008) presented a study to determine various chemical properties like protein, fibre, calcium, phosphorus, magnesium and potassium using hyperspectral imaging system in the 360–1010 nm wavelength range. A staring array imaging system was used to acquire the images of the grass field in Aomori, Japan. Various samples of grass were collected from the same area of imaging for the chemical analysis. The MLR, multilayer NN and PLSR techniques were used to develop the prediction models to predict the chemical components of the grass. Cross-validation results had suggested that the PLSR model gave the higher accuracy when compared to the MLR and multilayer NN.
Suzuki et al. (2012) used hyperspectral imaging system for mapping the spatial distribution of plant species or species groups and herbage mass of the grass pastures. A field-scale and small-scale area scan imaging systems was used to conduct this study. Images were acquired in the reflectance mode in the Vis-NIR range of 360–1010 nm. LDA was used to classify in to three plant species or species groups, and PLSR was used to estimate the herbage mass of the grass pasture. Using LDA, the overall success rate for the discrimination of plant species was 91.6 %, and the herbage mass was accurately estimated with a R 2 of 0.60.
Pérez-Marín et al. (2009) investigated the ability of spectroscopy for the determination of major quality parameters like soluble solid content, flesh firmness, fruit weight and fruit diameter of nectarines using one instrument infield during ripening of fruits on tree and other instrument during post-harvest storage. The microelectromechanical system (MEMS) spectrometer was used in 1600–2400 nm wavelength range for the measurement of quality parameters in the field study, and a diode array Vis-NIR spectrophotometer was used in 400–1700 nm range to determine these qualities during post-harvest storage. Both the instruments gave good precision for the determination of soluble solid content (r 2 of 0.89) and firmness (r 2 of 0.84–0.86). The diode array Vis-NIR spectrophotometer gave good prediction results of fruit weight (r 2 of 0.98) and diameter (r 2 of 0.75).
Gracia and León (2011) investigated the ability of NIR technology for the assessment of olive oil and moisture content of olive fruits both on tree infield and in laboratory conditions. Prediction models were developed using PLSR separately for each trial. The laboratory test gave better results when compared to the infield test. The prediction model accurately predicted the oil content with r = 0.89 and moisture content with r = 0.88.
Pérez-Marín et al. (2011) evaluated the ability of NIR system for the classification of nectarines by their internal quality during storage as a function of the irrigation methods during the growing stage and post-harvest storage period. The study was conducted using one instrument infield during ripening of fruits on tree and another instrument during post-harvest storage. Classification models were developed using PLSDA technique to classify the nectarines based on their internal quality. The PLSDA classification model accurately classified the samples with 57–84 % accuracy based on the irrigation method during the growing period.
Vigneau et al. (2011) investigate the use of hyperspectral imaging system for the prediction of leaf nitrogen content in wheat both infield and greenhouse. The entire study was conducted by acquiring the hyperspectral images of wheat leaves in the 400–1000 nm range. In the first study, the PLSR model was calibrated to estimate the nitrogen content of the flat leaves using the reflectance spectra (R 2 0.903). Later, this model was used to predict the nitrogen content of the leaves both infield (R 2 = 0.881) and greenhouse (0.889).
Conclusions
Hyperspectral imaging has proven to be a highly accurate technique for quality assessment of food and agricultural products as the data acquired using this technique contain adequate information about the properties of the sample. Depending on the equipment availability and the type of study it is intended for, hyperspectral data can be acquired using different imaging modes, systems and wavelength ranges. The hyperspectral data can be affected by various environment and equipment factors. Spectral pre-processing techniques are generally used alone or in combinations to reduce these effects. As the hyperspectral imaging provides massive amount of data, exploratory data analysis techniques like PCA can be used to extract the valuable data subsets that contain most of the information. Hyperspectral data can be explored for qualitative properties of the sample using discriminant analysis techniques (QDA, LDA and PLS-DA) or for quantitative properties using regression analysis techniques (PCR and PLSR) or for both using advance chemometric techniques (SVM, NB and ANN).
References
Akbari H., Halig L.V., Zhang H., Wang D., Chen Z.G. & Fei B. (2012) Detection of cancer metastasis using a novel macroscopic hyperspectral method. In: Proceedings of the SPIE Volume 8317 Medical Imaging 2012: Biomedical Applications in Molecular, Structural, and Functional Imaging, 23 March 2012, Paper No. 831711, San Diego, California, USA.
Albers, B., DiBenedetto, J., Lutz, S., & Purdy, C. (1995). More efficient environmental monitoring with laser-induced fluorescence imaging. Biophotonics International Magazine, 2(6), 42–54.
Amigo, J. M., & Ravn, C. (2009). Direct quantification and distribution assessment of major and minor components in pharmaceutical tablets by NIR-chemical imaging. European Journal of Pharmaceutical Sciences, 37(2), 76–82.
Amigo, J. M. (2010). Practical issues of hyperspectral imaging analysis of solid dosage forms. Analytical Bioanalytical Chemistry, 398(1), 93–109.
Amigo, J. M., Cruz, J., Bautista, M., Maspoch, S., Coello, J., & Blanco, M. (2008). Study of pharmaceutical samples by NIR chemical-image and multivariate analysis. Trends in Analytical Chemistry, 27(8), 696–713.
Anderson, N. M., & Walker, P. N. (2003). Measuring fat content of ground beef stream using on-line visible/NIR spectroscopy. Transactions of the ASAE, 46(1), 117–124.
Andersson, C. A. (1999). Direct orthogonalization. Chemometrics and Intelligent Laboratory Systems, 47(1), 51–63.
Anonymous (2013) Hyperspectral imaging. Hyperspectral imaging: components and systems catalog, Middleton Research, Middleton, WI, USA, Available at: www.middletonresearch.com/pdfs/3-HSI.pdf. Accessed 21 November 2013.
Ariana, D. P., & Lu, R. (2010). Evaluation of internal defect and surface color of whole pickles using hyperspectral imaging. Journal of Food Engineering, 96(4), 583–590.
Ariana, D. P., Lu, R., & Guyer, D. E. (2006). Near-infrared hyperspectral reflectance imaging for detection of bruises on pickling cucumbers. Computers and Electronics in Agriculture, 53(1), 60–70.
Baker, J. E., Dowell, F. E., & Throne, J. E. (1999). Detection of parasitized rice weevils in wheat kernels with near-infrared spectroscopy. Biological Control, 16, 88–90.
Baker, M., & Rayens, W. (2003). Partial least squares for discrimination. Journal of Chemometrics, 17(3), 166–173.
Ballabio D & Todeschini R (2009) Multivariate classification for qualitative analysis. Sun Infrared Spectroscopy for Food Quality Analysis and Control, pp 83–104. Academic press, Burlington, USA.
Baranowski, P., Mazurek, W., Wozniak, J., & Majewska, U. (2012). Detection of early bruises in apples using hyperspectral data and thermal imaging. Journal of Food Engineering, 110(3), 345–355.
Barbin, D., ElMasry, G., Sun, D.-W., & Allen, P. (2012). Near-infrared hyperspectral imaging for grading and classification of pork. Meat Science, 90(1), 259–268.
Barbin, D. F., ElMasry, G., Sun, D.-W., & Allen, P. (2013). Non-destructive determination of chemical composition in intact and minced pork using near-infrared hyperspectral imaging. Food Chemistry, 138(2–3), 1162–1171.
Barnes, R. J., Dhanoa, M. S., & Lister, S. J. (1989). Standard normal variate transformation and de-trending of near-infrared diffuse reflectance spectra. Applied Spectroscopy, 43(5), 772–777.
Bauriegel, E., Giebel, A., Geyer, M., Schmidt, U., & Herppich, W. B. (2011). Early detection of Fusarium infection in wheat using hyperspectral imaging. Computers & Electronics in Agriculture, 75(2), 304–312.
Beghi, R., Spinardi, A., Bodria, L., Mignani, I., & Guidetti, R. (2013). Apples nutraceutic properties evaluation through a visible and near-infrared portable system. Food and Bioprocess Technology, 6(9), 2547–2554.
Borengasser, M., Hungate, W. S., & Watkins, R. (2007). Hyperspectral remote sensing: principles and applications. Boca Raton, USA: CRC Press.
Botchko V, Berina E, Korotkaya Z, Parkkinen J & Jaaskelainen, T (2003) Independent component analysis in spectral images. 4th International Symposium on Independent Component Analysis and Blind Signal Separation (ICA 2003), 1–4 April 2003, Nara, Japan.
Boysworth, M. K., & Booksh, K. S. (2007). Aspects of multivariate calibration applied to near-infrared spectroscopy. In Ciurczak & Burns (Eds.), Handbook of near-infrared analysis (3rd ed., pp. 207–229). Boca Raton, USA: CRC Press.
Brosnan, T., & Sun, D.-W. (2004). Improving quality inspection of food products by computer vision-a review. Journal of Food Engineering, 61(1), 3–16.
Brown, C. D., Vega-Montoto, L., & Wentzell, P. D. (2000). Derivatives preprocessing and optimal corrections for baseline drift in multivariate calibration. Applied Spectroscopy, 54(7), 1055–1068.
Buddenbaum, H., & Steffens, M. (2012). The effect of spectral pretreatments on chemometrics analyses of soil profile using laboratory imaging spectroscopy. Applied and Environmental Soil Science, Article ID, 274903, 1–12.
Bulanon, D. M., Burks, T. F., Kim, D. G., & Ritenour, M. A. (2013). Citrus black spot detection using hyperspectral image analysis. Agricultural Engineering International: CIGR Journal, 15(3), 171–180.
Burges, C. J. C. (1998). A tutorial on support vector machines for pattern recognition. Data Mining and Knowledge Discovery, 2(2), 121–167.
Centner, V., Massart, D. L., & de Noord, O. E. (1996). Detection of inhomogeneities in sets of NIR spectra. Analytica Chimica Acta, 330(1), 1–17.
Du, C., Ma Z., Zhou J., Goyne K.W. (2013). Application of mid-infrared photoacoustic spectroscopy inmonitoring carbonate content in soils. Sensors and Actuators B, 188, 1167–1175.
Chappelle, E. W., McMurtrey III, J. E., & Kim, M. S. (1991). Identification of the pigment responsible for the blue fluorescence band in laser induced fluorescence (LIF) spectra of green plants, and the potential use of this band in remotely estimating rates of photosynthesis. Remote Sensing of Environment, 36(3), 213–218.
Chappelle, E. W., Wood, F. M., McMurtrey III, J. E., & Newcomb, W. W. (1984). Laser induced fluorescence of green plants. 1: a technique for the remote detection of plant stress and species differentiation. Applied Optics, 23(1), 134–138.
Chen, Y.-N., Sun, D.-W., Cheng, J.-H., & Gao, W.-H. (2016). Recent advances for rapid identification of chemical information of muscle foods by hyperspectral imaging analysis. Food Engineering Reviews, 8(3), 336–350.
Chen, Y.-R., Chao, K., & Kim, M. S. (2002). Machine vision technology for agricultural applications. Computers and Electronics in Agriculture, 36(2–3), 173–191.
Chen, Z. P., Morris, J., & Martin, E. (2006). Extracting chemical information from spectral data with multiplicative light scattering effects by optical path-length estimation and correction. Analytical Chemistry, 78(22), 7674–7681.
Cheng, W., Sun, D.-W., & Cheng, J.-H. (2016). Pork biogenic amine index (BAI) determination based on chemometric analysis of hyperspectral imaging data. LWT-Food Science and Technology, 73, 13–19.
Cheng, X., Chen, Y. R., Tao, Y., Wang, C. Y., Kim, M. S., & Lefcourt, A. M. (2004). A novel integrated PCA and FLD method on hyperspectral image feature extraction for cucumber chilling damage inspection. Transactions of the ASAE, 47(4), 1313–1320.
Cho, B.-K., Kim, M. S., Baek, I.-S., Kim, D.-Y., Lee, W.-H., Kim, J., Bae, H., & Kim, Y.-S. (2013). Detection of cuticle defects on cherry tomatoes using hyperspectral fluorescence imagery. Postharvest Biology and Technology, 76, 40–49.
Choudhary, R., Mahesh, S., Paliwal, J., & Jayas, D. S. (2009). Identification of wheat classes using wavelet features from near infrared hyperspectral images of bulk samples. Biosystems Engineering, 102(2), 115–127.
Clark, D., & Sasic, S. (2006). Chemical images: technical approaches and issues. Cytometry Part A, 69A(8), 815–824.
Cloarec, O., Dumas, M. E., Craig, A., Barton, R. H., Trygg, J., Hudson, J., Blancher, C., Gauguier, D., Lindon, J. C., Holmes, E., & Nicholson, J. (2005). Statistical total correlation spectroscopy: an exploratory approach for latent biomarker identification from metabolic 1H NMR data sets. Analytical Chemistry, 77(5), 1282–1289.
Cogdill, R. P., Hurburgh Jr., C. R., Rippke, G. R., Bajic, S. J., Jones, R. W., McClelland, J. F., Jensen, T. C., & Liu, J. (2004). Single-kernel maize analysis by near-infrared hyperspectral imaging. Transactions of the ASAE, 47(1), 311–320.
Comon, P. (1994). Independent component analysis, a new concept? Signal Processing, 36(3), 287–314.
Cortes, C., & Vapnik, V. (1995). Support vector networks. Machine Learning, 20(3), 273–297.
Dacal-Nieto A, Formella A, Carrión P, Vazquez-Fernandez E & Fernández-Delgado M (2011) Non-destructive detection of hollow heart in potatoes using hyperspectral Imaging. In: Berciano et al. (eds) Proceedings of the 14th International Conference on Computer Analysis of Images and Pattern (CAIP 2011), Part II, 29–31 August 2011, pp 180–187, Seville, Spain.
De Temmerman, J., Saeys, W., Nicolai, B., & Ramon, H. (2007). Near infrared reflectance spectroscopy as a tool for the in-line determination of the moisture concentration in extruded semolina pasta. Biosystems Engineering, 97(3), 313–321.
Delwiche, S. R., Kim, M. S., & Dong, Y. (2011). Fusarium damage assessment in wheat kernels by Vis/NIR hyperspectral imaging. Sensing and Instrumentation for Food Quality and Safety, 5(2), 63–71.
Dhanoa, M. S., Lister, S. J., Sanderson, R., & Barnes, R. J. (1994). The link between multiplicative scatter correction (MSC) and standard normal variate (SNV) transformations of NIR spectra. Journal of Near Infrared Spectroscopy, 2(1), 43–47.
Ding C & He X (2004) K-means clustering via principal component analysis. In: Brodley (ed) Proceedings of the 21st International Conference on Machine Learning, 4–8 July 2004, Banff, Canada.
Donovan, J. J., Snyder, D. A., & Rivers, M. L. (1993). An improved interference correction for trace element analysis. Microbeam Analysis, 2, 23–28.
Dowell, F. E. (2000). Differentiating vitreous and nonvitreous of durum wheat kernels by using near-infrared spectroscopy. Cereal Chemistry, 77(2), 155–158.
Du, C.-J., & Sun, D.-W. (2004). Recent developments in the applications of image processing techniques for food quality evaluations. Trends in Food Science & Technology, 15(5), 230–249.
Du H, Qi H, Wang X, Ramanath R & Snyder, WE (2003) Band selection using independent component analysis for hyperspectral image processing. In: Proceedings of the 32nd Applied Imagery Pattern Recognition Workshop, 15–17 October 2003, Washington, USA.
Duda, R. O., Hart, P. E., & Stork, D. G. (2001). Pattern classification (2nd edition). New York, USA: Wiley–Interscience.
ElMasry, G. & Sun D.-W. (2010) Principles of hyperspectral imaging technology. In: Sun Hyperspectral Imaging for Food Quality Analysis and Control (1st edition), pp 3–43. Academic Press, London, UK.
ElMasry, G., Iqbal, A., Sun, D.-W., Allen, P., & Ward, P. (2011). Quality classification of cooked, sliced Turkey hams using NIR hyperspectral imaging system. Journal of Food Engineering, 103(3), 333–344.
ElMasry, G., Kamruzzaman, M., Sun, D.-W., & Allen, P. (2012a). Principles and applications of hyperspectral imaging in quality evaluation of agro-food products: a review. Critical Reviews in Food Science and Nutrition, 52(11), 999–1023.
ElMasry, G., Sun, D.-W., & Allen, P. (2012b). Near-infrared hyperspectral imaging for predicting colour, pH and tenderness of fresh beef. Journal of Food Engineering, 110(1), 127–140.
ElMasry, G., Wang, N., ElSayed, A., & Ngadi, M. (2007). Hyperspectral imaging for nondestructive determination of some quality attributes for strawberry. Journal of Food Engineering, 81(1), 98–107.
Eluyode, O. S., & Akomolafe, D. T. (2013). Comparative study of biological and artificial neural networks. European Journal of Applied Engineering and Scientific Research, 2(1), 36–46.
Feng, Y.-Z., & Sun, D.-W. (2012). Determination of total viable count (TVC) in chicken breast fillets by near-infrared hyperspectral imaging and spectroscopic transforms. Talanta, 105, 244–249.
Fernández-Ahumada, E., Garrido-Varo, A., & Guerrero-Ginel, J. E. (2008). Feasibility of diode-array instruments to carry near-infrared spectroscopy from laboratory to feed process control. Journal of Agricultural and Food Chemistry, 56(9), 3185–3192.
Firrao, G., Torelli, E., Gobbi, E., Raranciuc, S., Bianchi, G., & Locci, R. (2010). Prediction of milled maize fumonisin contamination by multispectral image analysis. Journal of Cereal Science, 52(2), 327–330.
Fisher, R. A. (1936). The use of multiple measurements in taxonomic problems. Annals of Eugenics, 7(2), 179–188.
Fletcher JT & Kong SG (2003) Principal component analysis for poultry tumor inspection using hyperspectral fluorescence imaging. In Proceedings of the IEEE International Joint Conference on Neural Networks, Volume 1, 20–24 July 2003, pp 149–153, Portland, USA.
Foca, G., Salvo, D., Cino, A., Ferrari, C., Lo Fiego, D. P., Minelli, G., & Ulrici, A. (2013). Classification of pig fat samples from different subcutaneous layers by means of fast and non-destructive analytical techniques. Food Research International, 52(1), 185–197.
Fong AY & Wachman E (2008) Advanced photonic tools for hyperspectral imaging for life sciences. SPIE Newsroom: Electronic Imaging & Signal Processing, Available at: https://spie.org/documents/Newsroom/Imported/1051/1051-2008-03-20.pdf. Accessed 29 November 2013.
Friedman, N., Geiger, D., & Goldszmidt, M. (1997). Bayesian network classifiers. Machine Learning, 29, 131–163.
Geladi, P., MacDougall, D., & Martens, H. (1985). Linearization and scatter-corrections for near-infrared reflectance spectra of meat. Applied Spectroscopy, 39(3), 491–500.
Ghosh, P. K., Jayas, D. S., Gruwel, M. L. H., & White, N. D. G. (2007). A magnetic resonance imaging study of wheat drying kinetics. Biosystems Engineering, 97(2), 189–199.
Givens, D. I., De Boever, J. L., & Deaville, E. R. (1997). The principles, practices and some future applications of near infrared spectroscopy for predicting the nutritive value of foods for animals and humans. Nutrition Research Reviews, 10(1), 83–114.
Goetz, A. F. H. (2009). Three decades of hyperspectral remote sensing of the earth: a personal review. Remote sensing of environment, 113. Supplement, 1, S5–S16.
Goetz, A. F. H., Vane, G., Solomon, J. E., & Rock, B. N. (1985). Imaging spectrometry for earth remote sensing. Science, 228(4704), 1147–1153.
Golic, M., & Walsh, K. B. (2006). Robustness of calibration models based on near infrared spectroscopy for the in-line grading of stonefruit for total soluble solids content. Analytica Chimica Acta, 555(2), 286–291.
Gorretta, N., Roger, J., Aubert, M., Bellon-Maurel, V., Campan, F., & Roumet, P. (2006). Determining vitreousness of durum wheat kernels using near infrared hyperspectral imaging. Journal of Near Infrared Spectroscopy, 14(1), 231–239.
Gou, P., Santos-Garcés, E., Høy, M., Wold, J. P., Liland, K. H., & Fulladosa, E. (2013). Feasibility of NIR interactance hyperspectral imaging for on-line measurement of crude composition in vacuum packed dry-cured ham slices. Meat Science, 95(2), 250–255.
Gowen, A. A., O’Donnell, C. P., Cullen, P. J., Downey, G., & Frias, J. M. (2007). Hyperspectral imaging—an emerging process analytical tool for food quality and safety control. Trends in Food Science and Technology, 18(12), 590–598.
Gowen, A. A., Taghizadeh, M., & O’Donnell, C. P. (2009). Identification of mushrooms subjected to freeze damage using hyperspectral imaging. Journal of Food Engineering, 93(1), 7–12.
Gracia, A., & León, L. (2011). Non-destructive assessment of olive fruit ripening by portable near infrared spectroscopy. Grasas y Aceites, 62(3), 268–274.
Guo, G., Li, S. Z., & Chan, K. L. (2000). Support vector machines for face recognition. Image and Vision Computing, 19(9–10), 631–638.
Hand, D. J. (1997). Construction and assessment of classification rules. New York, USA: John Wiley and Sons.
Hans, B.-P. (2003). Analysis of water in food by near infrared spectroscopy. Food Chemistry, 82(1), 107–115.
Harris, P. J., & Hartley, R. D. (1976). Detection of bound ferulic acid in cell walls of the Gramineae by ultraviolet fluorescence microscopy. Nature, 259, 508–510.
Heia, K., Sivertsen, A. H., Stormo, S. K., Elvevoll, E., Wold, J. P., & Nilsen, H. (2007). Detection of nematodes in cod (Gadus morhua) fillets by imaging spectroscopy. Journal of Food Science, 72(1), E011–E015.
Heraud, P., Wood, B. R., Beardall, J., & McNaughton, D. (2006). Effects of pre-processing of Raman spectra on in vivo classification of nutrient status of microalgal cells. Journal of Chemometrics, 20(5), 193–197.
Herault J & Jutten C (1986) Space or time adaptive signal processing by neural network models. In: Proceedings of the American Institute of Physics Conference, Neural Networks for Computing, 13–16 April 1986, Snowbird, USA.
Hottelling, H. (1933). Analysis of a complex of statistical variables into principal components. Journal of Education Psychology, 24(6), 417–441.
Hu, K., & Wang, D. (2011). Unvoiced speech segregation from nonspeech interference via CASA and spectral subtraction. IEEE Transactions on Audio, Speech, and Language Processing, 19(6), 1600–1609.
Huang, M., & Lu, R. (2010). Optimal wavelength selection for hyperspectral scattering prediction of apple firmness and soluble solids content. Transactions of the ASABE, 53(4), 1175–1182.
Huang, M., Zhu, Q., Wang, B., & Lu, R. (2012a). Analysis of hyperspectral scattering images using locally linear embedding algorithm for apple mealiness classification. Computers and Electronics in Agriculture, 89, 175–181.
Huang, Y. B., Thomson, S. J., Molin, W. T., Reddy, K. N., & Yao, H. B. (2012b). Early detection of soybean plant injury from glyphosate by measuring chlorophyll reflectance and fluorescence. Journal of Agricultural Science, 4(5), 117–124.
Iqbal, A., Sun, D.-W., & Allen, P. (2013). Prediction of moisture, color and pH in cooked, pre-sliced Turkey hams by NIR hyperspectral imaging system. Journal of Food Engineering, 117(1), 42–51.
Isaksson, T., & Kowalski, B. (1993). Piece-wise multiplicative scatter correction applied to near-infrared diffuse transmittance data from meat products. Applied Spectroscopy, 47(6), 702–709.
Jennrich RJ (1977) Stepwise discriminant analysis. In: Enslein et al (eds) Statistical Methods for Digital Computers, Wiley and Sons, New York, USA.
Jensen JR (2007) Introductory to digital image processing: a remote sensing perspective. Prentice Hall Series in Geographic Information Science.
Jiang, L., Zhu, B. & Tao, Y. (2010) Hyperspectral image classification methods. In: Sun (ed) Hyperspectral Imaging for Food Quality Analysis and Control (1st edition), pp 79–98, Academic Press is an imprint of Elsevier, London, UK.
Jun, W., Kim, M. S., Cho, B.-K., Millner, P. D., Chao, K., & Chan, D. E. (2010). Microbial biofilm detection on food contact surfaces by macro-scale fluorescence imaging. Journal of Food Engineering, 99(3), 314–332.
Kaliramesh, S., Chelladurai, V., Jayas, D. S., Alagusundaram, K., White, N., & Fields, P. (2013). Detection of infestation by Callosobruchus maculatus in mung bean using near-infrared hyperspectral imaging. Journal of Stored Products Research, 52, 107–111.
Kamruzzaman, M., ElMasry, G., Sun, D.-W., & Allen, P. (2011). Application of NIR hyperspectral imaging for discrimination of lamb muscles. Journal of Food Engineering, 104(3), 332–340.
Kamruzzaman, M., ElMasry, G., Sun, D.-W., & Allen, P. (2012). Non-destructive prediction and visualization of chemical composition in lamb meat using NIR hyperspectral imaging and multivariate regression. Innovative Food Science and Emerging Technologies, 16, 218–226.
Kamruzzaman, M., Sun, D.-W., ElMasry, G., & Allen, P. (2013). Fast detection and visualization of minced lamb meat adulteration using NIR hyperspectral imaging and multivariate image analysis. Talanta, 103, 130–136.
Karstang, T. V., & Manne, R. (1992). Optimized scaling: a novel approach to linear calibration with closed data sets. Chemometrics and Intelligent Laboratory Systems, 14(1–3), 165–173.
Karunakaran, C., Jayas, D. S., & White, N. D. G. (2003). Soft X-ray inspection of wheat kernels infested by Sitophilus oryzae. Transactions of the ASAE, 46(3), 739–745.
Kavdir, I., Buyukcan, M. B., Lu, R., Kocabiyik, H., & Seker, M. (2009). Prediction of olive quality using FT-NIR spectroscopy in reflectance and transmittance modes. Biosystems Engineering, 103(3), 304–312.
Kellicut, D., Weiswasser, J., Arora, S., Freeman, J., Lew, R., Shuman, C., Mansfield, J. R., & Sidewy, A. N. (2004). Emerging technology: hyperspectral imaging. Perspectives in Vascular Surgery and Endovascular Therapy, 16(1), 53–57.
Kemps, B., Leon, L., Best, S., De Baerdemaeker, J., & Ketelaere, D. (2010). Assessment of the quality parameters in grapes using VIS/NIR spectroscopy. Biosystems Engineering, 105(4), 507–513.
Keulemans, W., Delalieux, S., Aardt, J., & Coppin, P. (2007). Detection of biotic stress (Venturia inaequalis) in apple trees using hyperspectral analysis: nonparametric statistical approaches and physiological implications. European Journal of Agronomy, 27(1), 130–143.
Kim, I., Kim, M. S., Chen, Y. R., & Kong, S. G. (2004). Detection of skin tumors on chicken carcasses using hyperspectral fluorescence imaging. Transactions of the ASAE, 47(5), 1785–1792.
Kim, M. S., Chen, Y. R., & Mehl, P. M. (2001). Hyperspectral reflectance and fluorescence imaging system for food quality and safety. Transactions of the ASAE, 44(3), 721–729.
Kim, M. S., Lefcourt, A. M., Chao, K., Chen, Y. R., Kim, I., & Chan, D. E. (2002). Multispectral detection of fecal contamination on apples based on hyperspectral imagery: part I. Application of visible and near-infrared reflectance imaging. Transactions of the ASAE, 45(6), 2027–2037.
Kim, T., Cho, B.-K., & Kim, M. S. (2010). Emission filter design to detect poultry skin tumors using fluorescence hyperspectral imaging. Revista Colombiana de Ciencias Pecuarias, 23(1), 9–16.
Kiyotoki, S., Nishikawa, J., Okamoto, T., Hamabe, K., Saito, M., Goto, A., Fujita, Y., Hamamoto, Y., Takeuchi, Y., Satori, S., & Sakaida, I. (2013). New method for detection of gastric cancer by hyperspectral imaging: a pilot study. Journal of Biomedical Optics, 18(2), 26010.
Kong SG (2003) Inspection of poultry skin tumor using hyperspectral fluorescence imaging. In Proceedings of the SPIE 6th International Conference on Quality Control by Artificial Vision, Paper No 5132, pp 455–463, 30 April 2003, Gatlinburg, USA.
Kong, S. G., Chen, Y.-R., Kim, I., & Kim, M. S. (2004). Analysis of hyperspectral fluorescence images for poultry skin tumor inspection. Applied Optics, 43(4), 824–833.
Kotsiantis, S. B. (2007). Supervised machine learning: a review of classification techniques. Informatica, 31, 249–268.
Koutchma, T. (2008). UV light for processing foods. IUVA news, 10(4), 24–29.
Kumar, A., Lee, W. S., Ehsani, R., Albrigo, L. G., Yang, C., & Mangan, R. L. (2012). Citrus greening disease detection using aerial hyperspectral and multispectral imaging techniques. Journal of Applied Remote Sensing, 6(1), 063542.
Kuska, M., Wahabzada, M., Leucker, M., Dehne, H.-W., Kersting, K., Oerke, K.-C., Steiner, U., & Mahlein, A.-K. (2015). Hyperspectral phenotyping on the microscopic scale: towards automated characterization of plant-pathogen interactions. Plant Methods, 11, 28.
Lang, M., Stiffel, P., Braunova, Z., & Lichtenthaler, H. K. (1992). Investigation of the blue–green fluorescence emission of plant leaves. Botanica Acta, 105, 395–468.
Langley, P., Iba, W., & Thompson, K. (1992). An analysis of Bayesian classifiers. In Proceedings of the 10th National Conference on artificial intelligence (pp. 223–228). San Jose, USA: AAAI Press.
Lara, M. A., Lleó, L., Diezma-Iglesias, B., Roger, J. M., & Ruiz-Altisent, M. (2013). Monitoring spinach shelf-life with hyperspectral image through packaging films. Journal of Food Engineering, 119(2), 353–361.
Lasch, P. (2012). Spectral pre-processing for biomedical vibrational spectroscopy and microspectroscopic imaging. Chemometrics and Intelligent Laboratory Systems, 117, 100–114.
Lee, K.-J., Kang, S., Kim, M.S., Noh S.H. (2005) Hyperspectal imaging for detecting defect on apples. ASABE conference paper, Paper No. 053075. St. Joseph, MI, USA.
Lefcourt, A. M., Kim, M. S., Chen, Y. R., & Kang, S. (2006). Systematic approach for using hyperspectral imaging data to develop multispectral imaging systems: detection of feces on apples. Computers and Electronics in Agriculture, 54(1), 22–35.
Leiva-Valenzuela, G. A., Lu, R., & Aguilera, J. M. (2012). Prediction of firmness and soluble solids content of blueberries using hyperspectral reflectance imaging. Journal of Food Engineering, 115(1), 91–98.
Lelong, C. C. D., Pinet, P. C., & Poilve, H. (1998). Hyperspectral imaging and stress mapping in agriculture: a case study on wheat in Beauce (France). Remote Sensing of Environment, 66(2), 179–191.
Levkov, C., Mihov, G., Ivanov, R., Daskalov, I., Christov, I., & Dotsinsky, I. (2005). Removal of power-line interference from the ECG: a review of the subtraction procedure. Biomedical Engineering Online, 4(50), 1–18.
Lieber, C. A., & Mahadevan-Jansen, A. (2003). Automated method for subtraction of fluorescence from biological Raman spectra. Applied Spectroscopy, 57(11), 1363–1367.
Liu, C., Liu, W., Lu, X., Ma, F., Chen, W., Yang, J., & Zheng, L. (2014). Application of multispectral imaging to determine quality attributes and ripeness stage in strawberry fruit. PloS One, 9(2), e87818.
Liu, L., Ngadi, M. O., Prasher, S. O., & Gariépy, C. (2010). Categorization of pork quality using Gabor filter-based hyperspectral imaging technology. Journal of Food Engineering, 99(3), 284–293.
Liu, Z., Wang, H., & Li, Q. (2012). Tongue tumor detection in medical hyperspectral images. Sensors, 12(1), 162–174.
Lopes, M. B., & Wolff, J. C. (2009). Investigation into classification/sourcing of suspect counterfeit Heptodintrade mark tablets by near infrared chemical imaging. Analytica Chimica Acta, 633(1), 149–155.
Lopes, M. B., Wolff, J. C., Bioucas-Dias, J. M., & Figueiredo, M. A. (2010). Near-infrared hyperspectral unmixing based on a minimum volume criterion for fast and accurate chemometric characterization of counterfeit tablets. Analytical Chemistry, 82(4), 1462–1469.
Lü, Q., & Tang, M. (2012). Detection of hidden bruise on kiwi fruit using hyperspectral imaging and parallelepiped classification. Procedia Environmental Sciences, 12(B), 1172–1179.
Lu, R., & Ariana, D. P. (2013). Detection of fruit fly infestation in pickling cucumbers using a hyperspectral reflectance/transmittance imaging system. Postharvest Biology and Technology, 81, 44–50.
Lu R & Chen YR (1999). Hyperspectral imaging for safety inspection of food and agricultural products. In: Proceedings of the SPIE Conference on Pathogen Detection and Remediation for Safe Eating, Volume 3544, 12 January 1999, pp 121–133, Boston, USA.
Lu, R. (2003). Detection of bruises on apples using near-infrared hyperspectral imaging. Transactions of the ASAE, 46(2), 523–530.
Ma, J., Sun, D.-W., & Pu, H. (2016). Spectral absorption index in hyperspectral image analysis for predicting moisture contents in pork longissimus dorsi muscles. Food Chemistry, 197, 848–854.
Mahalanobis, P. C. (1936). On the generalised distance in statistics. Proceedings of the National Institute of Sciences of India, 2(1), 49–55.
Mahesh S, Jayas DS, Paliwal J & White NDG (2010) Near-infrared (NIR) hyperspectral imaging—an emerging analytical tool for classification of western Canadian wheat classes from different locations and crop years. ASABE conference paper, Paper No. MB-SK 10–302. St. Joseph, MI, USA.
Mahesh, S., Jayas, D. S., Paliwal, J., & White, N. D. G. (2011). Identification of wheat classes at different moisture levels using near-infrared hyperspectral images of bulk samples. Sensing and Instrumentation for Food Quality and Safety, 5(1), 1–9.
Mahesh, S., Manickavasagan, A., Jayas, D. S., Paliwal, J., & White, N. D. G. (2008). Feasibility of near-infrared hyperspectral imaging to differentiate Canadian wheat classes. Biosystems Engineering, 101(1), 50–57.
Manley, M., Williams, P., Nilsson, D., & Geladi, P. (2009). Near infrared hyperspectral imaging for the evaluation of endosperm texture in whole yellow maize (Zea maize L.) kernels. Journal of Agricultural and Food Chemistry, 57(19), 8761–8769.
Mark, H., & Workman, J. (2003). Statistics in spectroscopy (2nd edition). San Diego: Academic Press.
Marquez, A. J., AM, D., & MIP, R. (2005). Using optical NIR sensor for on-line virgin olive oils characterization. Sensors and Actuators B, 107(1), 64–68.
Martens, H., & Naes, T. (1992). Multivariate calibration. New York: John Wiley and Sons.
Martens, H., & Stark, E. (1991). Extended multiplicative signal correction and spectral interference subtraction: new preprocessing methods for near infrared spectroscopy. Journal of Pharmaceutical and Biomedical Analysis, 9(8), 625–635.
Martens H, Jensen SA & Geladi P (1983) Multivariate linearity transformations for near infrared reflectance spectroscopy. In: Christie (ed) Proceedings of the Nordic Symposium on Applied Statistics, pp 205–234, 12–14 June 1983, Stokkand Forlag Publishers, Stavanger, Norway.
Martens, H., Nielsen, J. P., & Engelsen, S. B. (2003). Light scattering and light absorbance separated by extended multiplicative signal correction. Application to near-infrared transmission analysis of powder mixtures. Analytical Chemistry, 75(3), 394–404.
Martinsen, P., Schaare, P., & Andrews, M. (1999). A versatile near infrared imaging spectrometer. Journal of Near Infrared Spectroscopy, 7(1), 17–25.
Massart DL, Vandeginste BGM, Buydens LMC, de Jong S, Lewi PJ & Smeyers-Verbeke J (1997) Handbook of chemometrics and qualimetrics: Part A. Elsevier Science B. V., Amsterdam, The Netherlands.
Mazet, V., Carteret, C., Brie, D., Idier, J., & Humbert, B. (2005). Background removal from spectra by designing and minimising a non-quadratic cost function. Chemometrics and Intelligent Laboratory Systems, 76(2), 121–133.
McGoverin, C. M., Engelbrecht, P., Geladi, P., & Manley, M. (2011). Characterisation of non-viable whole barley, wheat and sorghum grains using near-infrared hyperspectral data and chemometrics. Analytical and Bioanalytical Chemistry, 401(7), 2283–2289.
McLachlan, G. (1992). Discriminant analysis and statistical pattern recognition. Hoboken: John Wiley & Sons, Inc..
Mehl, P. M., Chen, Y.-R., Kim, M. S., & Chan, D. E. (2004). Development of hyperspectral imaging technique for the detection of apple surface defects and contaminations. Journal of Food Engineering, 61(1), 67–81.
Melgani, F., & Bruzzone, L. (2004). Classification of hyperspectral remote sensing images with support vector machines. IEEE Transactions on Geoscience and Remote Sensing, 42(8), 1778–1790.
Mendoza, F., Lu, R., Ariana, D., Cen, H., & Bailey, B. (2011). Integrated spectral and image analysis of hyperspectral scattering data for prediction of apple fruit firmness and soluble solids content. Postharvest Biology and Technology, 62(2), 149–160.
Mohan, L. A., Karunakaran, C., Jayas, D. S., & White, N. D. G. (2005). Classification of bulk cereals using visible and NIR reflectance characteristics. Canadian Biosystems Engineering, 47, 7.7–7.14.
Nagata M, Tallada JG, Kobayashi T & Toyada H (2005) NIR hyperspectral imaging for measurement of internal quality in strawberries. ASABE conference paper, Paper No. 053131, St. Joseph, MI, USA.
Nagata M, Tallada JG, Kobayashi T, Cui Y & Gejima Y (2004) Predicting maturity quality parameters of strawberries using hyperspectral imaging. ASABE conference paper, Paper No. 043033, St. Joseph, MI, USA.
Neville, R. A., Levesque, J., Staenz, K., Nadeau, C., Hauff, P., & Borstad, G. A. (2003). Spectral unmixing of hyperspectral imagery for mineral exploration: comparison of results from SFSI and AVIRIS. Canadian Journal of Remote Sensing, 29(1), 99–110.
Nguyen NH, Chen J, Richard C, Honeine P & Theys C (2013) Supervised nonlinear unmixing of hyperspectral images using a pre-image methods. In: Mary et al (eds) New Concepts in Imaging: Optical and Statistical Models, EAS Publications Series, 59, 417–437.
Nicolai, B. M., Beullens, K., Bobelyn, E., Peirs, A., Saeys, W., Theron, K. I., & Lammertyn, J. (2007). Nondestructive measurement of fruit and vegetable quality by means of NIR spectroscopy: a review. Postharvest Biology and Technology, 46(2), 99–118.
Nicolai, B. M., Lotze, E., Peirs, A., Scheerlinck, N., & Theron, K. I. (2006). Non-destructive measurement of bitter pit in apple fruit using NIR hyperspectral imaging. Postharvest Biology and Technology, 40(1), 1–6.
Noh HK & Lu R (2005) Hyperspectral reflectance and fluorescence for assessing apple quality. ASABE conference paper, Paper No 053069, St. Joseph, MI, USA.
Noh, H. K., & Lu, R. (2007). Hyperspectral laser-induced fluorescence imaging for assessing apple fruit quality. Postharvest Biology and Technology, 43(2), 193–201.
Noh, H. K., Peng, Y., & Lu, R. (2007). Integration of hyperspectral reflectance and fluorescence imaging for assessing apple maturity. Transactions of the ASABE, 50(3), 963–971.
Norris, K. H., & Williams, P. C. (1984). Optimization of mathematical treatments of raw near-infrared signals in the measurement of protein in hard red spring wheat: 1. Influence of particle size. Cereal Chemistry, 61(2), 158–165.
Norris KH (1983) Extraction information from spectrophotometric curves. Predicting chemical composition from visible and near-infrared spectra. In: Martens & Russwurm Jr (eds) Food Research and Data Analysis, pp 95–113, Applied Science, London, UK.
Novales, B., Bertrand, D., Devaux, M. F., Robert, P., & Sire, A. (1999). Multispectral fluorescence imaging for the identification of food products. Journal of the Science of Food and Agriculture, 71(3), 376–382.
O’Farrell, M., Wold, J. P., Hoy, M., Tschudi, J., & Schulerud, H. (2010). On-line fat content classification of in homogeneous pork trimmings using multispectral near infrared interactance imaging. Journal of Near Infrared Spectroscopy, 18(2), 135–146.
Okamoto, H., & Lee, W. S. (2009). Green citrus detection using hyperspectral imaging. Computers and Electronics in Agriculture, 66(2), 201–208.
Osuna E, Freund R & Girosi F (1997) Training support vector machines: an application to face detection. In: Proceedings of the IEEE Computer Society Conference on Computer Vision and Pattern Recognition, 17–19 June 1997, pp 130–136, Puerto Rico, USA.
Ottavian, M., Fasolato, L., Serva, L., Facco, P., & Barolo, M. (2014). Data fusion for food authentication: fresh/frozen-thawed discrimination in west African goatfish (Pseudupeneus prayensis) fillets. Food and Bioprocess Technology, 7(4), 1025–1036.
Otto, O. (1998). Chemometrics: statistics and computer application in analytical chemistry. Wiley-VCH Verlag GmbH and Co. Weinheim: KGaA.
Paliwal, J., Jayas, D. S., Visan, N. S., & White, N. D. G. (2005). Quantification of variations in machine-vision-computed features of cereal grains. Canadian Biosystems Engineering, 47, 7.1–7.6.
Park B & Lu R (2015) Hyperspectral Imaging Technology in Food and Agriculture. Gustavo VB (eds.). Springer, New York, USA.
Park, B., Lawrence, K. C., Windham, W. R., & Buhr, R. J. (2002). Hyperspectral imaging for detecting fecal and ingesta contamination on poultry carcasses. Transactions of the ASAE, 45(6), 2017–2026.
Park, B., Lawrence, K. C., Windham, W. R., & Smith, D. P. (2006). Performance of hyperspectral imaging system for poultry surface fecal contaminant detection. Journal of Food Engineering, 75(3), 340–348.
Pearson, K. (1901). On lines and planes of closest fit to systems of points in space. Philosophical Magazine, 2, 559–572.
Pearson, T. C., Wicklow, D. T., Maghirang, E. B., Xie, F., & Dowell, F. E. (2001). Detecting aflatoxin in single corn kernels by using transmittance and reflectance spectroscopy. Transactions of the ASAE, 44(5), 1247–1254.
Pérez-Marín D, Sánchez M-T, Paz P & González-Dugo V (2011) Postharvest shelf-life discrimination of nectarines produced under different irrigation strategies using NIR-spectroscopy. Postharvest shelf-life discrimination of nectarines produced under different irrigation strategies using NIR-spectroscopy, 44(6), 1405–1414.
Pérez-Marín, D., Sánchez, M.-T., Paz, P., Soriano, M.-A., Guerrero, J.-E., & Garrido-Varo, A. (2009). Non-destructive determination of quality parameters in nectarines during on-tree ripening and postharvest storage. Postharvest Biology and Technology, 52(2), 180–188.
Pierna, J. A. F., Baeten, V., & Dardenne, P. (2006). Screening of compound feeds using NIR hyperspectral data. Chemometrics and Intelligent Laboratory Systems, 84(1–2), 114–118.
Pierna, J. A. F., Vermeulen, P., Amand, O., Tossens, A., Dardenne, P., & Baeten, V. (2012). NIR hyperspectral imaging spectroscopy and chemometrics for the detection of undesirable substances in food and feed. Chemometrics and Intelligent Laboratory Systems, 117, 233–239.
Pierna, J. A. F., Volery, P., Besson, R., Baeten, V., & Dardenne, P. (2005). Classification of modified starches by Fourier transform infrared spectroscopy using support vector machines. Journal of Agricultural and Food Chemistry, 53(17), 6581–6585.
Polder, G., Van Der Heijden, G. W. A. M., & Young, I. T. (2003). Tomato sorting using independent component analysis on spectral images. Real-Time Imaging, 9, 253–259.
Polder, G., Van Der Heijden, G. W. A. M., Waalwijk, C., & Young, I. T. (2005). Detection of Fusarium in single wheat kernels using spectral imaging. Seed Science and Technology, 33, 655–668.
Pu, H., Liu, D., Wang, L., & Sun, D.-W. (2016). Soluble solids content and pH prediction and maturity discrimination of lychee fruits using visible and near infrared hyperspectral imaging. Food Analytical Methods, 9(1), 235–244.
Pu, Y.-Y., & Sun, D.-W. (2016). Prediction of moisture content uniformity of microwave-vacuum dried mangoes as affected by different shapes using NIR hyperspectral imaging. Innovative Food Science and Emerging Technologies, 33, 348–356.
Qiao, J., Ngadi, M. O., Wang, N., Gariépy, C., & Prasher, S. O. (2007). Pork quality and marbling level assessment using a hyperspectral imaging system. Journal of Food Engineering, 83(1), 10–16.
Qin, J., & Lu, R. (2005). Detection of pits in tart cherries by hyperspectral transmission imaging. Transactions of the ASABE, 48(5), 1963–1970.
Qin, J., Burks, T. F., Kim, M. S., Chao, K., & Ritenour, M. A. (2008). Citrus canker detection using hyperspectral reflectance imaging and PCA-based image classification method. Sensing and Instrumentation for Food Quality and Safety, 2(3), 168–177.
Qin, J., Burks, T. F., Ritenour, M. A., & Bonn, W. G. (2009). Detection of citrus canker using hyperspectral reflectance imaging with spectral information divergence. Journal of Food Engineering, 93(2), 183–191.
Qu J-H, Sun D-W, Cheng J-H & Pu H (2016) Mapping moisture contents in grass carp (Ctenopharyngodon idella) slices under different freeze drying periods by Vis-NIR hyperspectral imaging. LWT-Food Science and Technology, In Press, Accepted Manuscript. Available online 19 September 2016.
Rajkumar, P., Wang, N., Eimasry, G., Raghavan, G. S. V., & Gariepy, Y. (2012). Studies on banana fruit quality and maturity stages using hyperspectral imaging. Journal of Food Engineering, 108(1), 194–200.
Ramalingam, G., Neethirajan, S., Jayas, D. S., & White, N. D. G. (2011). Characterization of the influence of moisture content on single wheat kernels using machine vision. Applied Engineering in Agriculture, 27(3), 403–409.
Rinnan A, Norgaard L, Van Der Berg FWJ, Thygesen J, Bro R & Engelsen SB (2009) Data pre-processing. In: Sun (ed) Infrared Spectroscopy for Food Quality Analysis and Control, pp 29–50, Academic press, Burlington, USA.
Rodríguez-Pulido, F. J., Barbin, D. F., Sun, D.-W., Gordillo, B., González-Miret, M. L., & Heredia, F. J. (2013). Grape seed characterization by NIR hyperspectral imaging. Postharvest Biology and Technology, 76, 74–82.
Romia MB & MA Bernardez (2009) Multivariate calibration for quantitative analysis. In: Sun (ed) Infrared spectroscopy for food quality analysis and control, pp 51–82, Academic Press, Burlington, USA.
Sapirstein, H. D., Neuman, M., Wright, E. H., Shwedyk, E., & Bushuk, W. (1987). An instrumental system for cereal grain classification using digital image analysis. Journal of Cereal Science, 6(1), 3–14.
Savitzky, A., & Golay, M. J. E. (1964). Smoothing and differentiation of data by simplified least squares procedures. Analytical Chemistry, 36(8), 1627–1639.
Schaare, P. N., & Fraser, D. G. (2000). Comparison of reflectance, interactance and transmission modes of visible-near infrared spectroscopy for measuring internal properties of kiwifruit (Actinidia chinensis). Postharvest Biology and Technology, 20(2), 175–184.
Schatzki, T. F., Haff, R. P., Young, R., Can, I., Le, L.-C., & Toyofuku, N. (1997). Defect detection in apples by means of X-ray imaging. Transactions of the ASAE, 40(5), 1407–1415.
Schulze, G., Jirasek, A., Yu, M. M., Lim, A., Turner, R. F., & Blades, M. W. (2005). Investigation of selected baseline removal techniques as candidates for automated implementation. Applied Spectroscopy, 59(5), 545–574.
Schweizer, S. M., & Moura, J. M. F. (2001). Efficient detection in hyperspectral imagery. IEEE Transactions on Image Processing, 10(4), 584–597.
Serranti, S., Cesare, D., Marini, F., & Bonifazi, G. (2013). Classification of oat and groat kernels using NIR hyperspectral imaging. Talanta, 103, 276–284.
Shenderey, C., Shmulevich, I., Alchanatis, V., Egozi, H., Hoffman, A., Ostrovsky, V., Lurie, S., Arie, B., & Schmilovitch, Z. (2010). NIRS detection of moldy core in apples. Food and Bioprocess Technology, 3(1), 79–86.
Singh, C. B., Jayas, D. S., Paliwal, J., & White, N. D. G. (2007). Fungal detection in wheat using near-infrared hyperspectral imaging. Transactions of the ASABE, 50(6), 2171–2176.
Singh, C. B., Jayas, D. S., Paliwal, J., & White, N. D. G. (2009). Detection of insect-damaged wheat kernels using near-infrared hyperspectral imaging. Journal of Stored Products Research, 45(3), 151–158.
Singh, C. B., Jayas, D. S., Paliwal, J., & White, N. D. G. (2010). Detection of midge-damaged wheat kernels using short-wave near-infrared hyperspectral and digital colour imaging. Biosystems Engineering, 105(3), 380–387.
Singh, C. B., Jayas, D. S., Paliwal, J., & White, N. D. G. (2012). Fungal damage detection in wheat using short-wave near-infrared hyperspectral and digital colour imaging. International Journal of Food Properties, 15(1), 11–24.
Siripatrawan, U., Makino, Y., Kawagoe, Y., & Oshita, S. (2011). Rapid detection of Escherichia coli contamination in packaged fresh spinach using hyperspectral imaging. Talanta, 85(1), 276–281.
Sivertsen, A. H., Chu, C.-K., Wang, L.-C., Godtliebsen, F., Heia, K., & Nilsen, H. (2009). Ridge detection with application to automatic fish fillet inspection. Journal of Food Engineering, 90(3), 317–324.
Sivertsen, A. H., Heia, K., Hindberg, K., & Godtliebsen, F. (2012). Automatic nematode detection in cod fillets (Gadus morhua L.) by hyperspectral imaging. Journal of Food Engineering, 111(4), 675–681.
Sivertsen, A. H., Heia, K., Stormo, S. K., Elvevoll, E., & Nilsen, H. (2011). Automatic nematode detection in cod fillets (Gadus morhua) by transillumination hyperspectral imaging. Journal of Food Science, 76(1), s77–s83.
Smith, K. L., Steven, M. D., & Colls, J. J. (2004). Use of hyperspectral derivative ratios in the red-edge region to identify plant stress responses to gas leaks. Remote Sensing of Environment, 92(2), 207–217.
Soulez F, Bongard S, Thiebaut E & Bacon R (2011) Restoration of hyperspectral astronomical data from integral field spectrograph. In: Proceedings of the 3rd IEEE-GRSS Workshop on Hyperspectral Image and Signal Processing: Evolution in Remote Sensing WHISPERS, 6–9 June 2011, pp 1–4, Lisbonne, Portugal.
Steinier, J., Termonia, Y., & Deltour, J. (1972). Comments on smoothing and differentiation of data by simplified least squares procedures. Analytical Chemistry, 44(11), 1906–1909.
Steinmetz, V., Roger, J. M., Molto, E., & Blasco, J. (1999). On-line fusion of colour camera and spectrophotometer for sugar content prediction of apples. Journal of Agricultural Research, 73(2), 207–216.
Stenlund, H., Johansson, E., Gottfries, J., & Trygg, J. (2009). Unlocking interpretation in near infrared multivariate calibrations by orthogonal partial least squares. Analytical Chemistry, 81(1), 203–209.
Stordrange, L., Libnau, F. O., Malthe-Sørenssen, D., & Kvalheim, O. M. (2002). Feasibility study of NIR for surveillance of a pharmaceutical process, including a study of different preprocessing techniques. Journal of Chemometrics, 16(8–10), 529–541.
Su, W.-H., & Sun, D.-W. (2016). Potential of hyperspectral imaging for visual authentication of sliced organic potatoes from potato and sweet potato tubers and rapid grading of the tubers according to moisture proportion. Computers and Electronics in Agriculture, 125, 113–124.
Suzuki, Y., Okamoto, H., & Kataoka, T. (2008c). Image segmentation between crop and weed using hyperspectral imaging for weed detection in soybean field. Environment Control in Biology, 46(3), 163–173.
Suzuki, Y., Okamoto, H., Takahashi, M., Kataoka, T., & Shibata, Y. (2012). Mapping the spatial distribution of botanical composition and herbage mass in pastures using hyperspectral imaging. Japanese Society of Grassland Science, 58(1), 1–7.
Suzuki, Y., Okamoto, H., Tanaka, K., Kato, W., & Kataoka, T. (2008a). Estimation of chemical composition of grass in meadows using hyperspectral imaging. Environment Control in Biology, 46(2), 129–137.
Suzuki, Y., Tanaka, K., Kato, W., Okamoto, H., Kataoka, T., Shimada, H., Sugiura, T., & Shima, E. (2008b). Field mapping of chemical composition of forage using hyperspectral imaging in a grass meadow. Japanese Society of Grassland Science, 54(4), 179–188.
Symons, S. J., & Fulcher, R. G. (1988). Determination of wheat kernel morphological variation by digital image analysis: I. Variation in eastern Canadian milling quality wheats. Journal of Cereal Science, 8(3), 211–218.
Taghizadeh, M., Gowen, A. A., & O’Donnell, C. P. (2011). The potential of visible-near infrared hyperspectral imaging to discriminate between casing soil, enzymatic browning and undamaged tissue on mushroom (Agaricus bisporus) surfaces. Computers and Electronics in Agriculture, 77(1), 74–80.
Tatzer, P., Wolf, M., & Panner, T. (2005). Industrial application for inline material sorting using hyperspectral imaging in the NIR range. Real-Time Imaging, 11(2), 99–107.
Tøgersen, G., Arnesen, J. F., Nilsena, B. N., & Hildruma, K. I. (2003). On-line prediction of chemical composition of semi-frozen ground beef by non-invasive NIR spectroscopy. Meat Science, 63(4), 515–523.
Tøgersen, G., Isaksson, T., Nilsen, B. N., Bakkerb, E. A., & Hildruma, K. I. (1999). On-line NIR analysis of fat, water and protein in industrial scale ground meat batches. Meat Science, 51(1), 97–102.
Tran, C. D. (2005). Principles, instrumentation and applications of infrared multispectral imaging, an overview. Analytical Letters, 38(5), 735–752.
Trygg, J., & Wold, S. (2002). Orthogonal projections to latent structures (O-PLS). Journal of Chemometrics, 16(3), 119–128.
Trygg, J., Holmes, E., & Lundstedt, T. (2007). Chemometrics in metabonomics. Journal of Proteome Research, 6(2), 469–479.
Tukey, J. W. (1980). We need both exploratory and confirmatory. The American Statistician, 34(1), 23–25.
Vadivambal, R., & Jayas, D. S. (2016). Bio-imaging: principles, techniques and applications. Oxford: CRC Press, Taylor and Francis Group Ltd.
Vadivambal, R., Jayas, D. S., & White, N. D. G. (2007). Wheat disinfestation using microwave energy. Journal of Stored Products Research, 43(4), 508–514.
Vajna, B., Patyi, G., Nagy, Z., Farkas, A., & Marosi, G. (2011). Comparison of chemometric methods in the analysis of pharmaceuticals with hyperspectral Raman imaging. Journal of Raman Spectroscopy, 42(11), 1977–1986.
Van Der Maaten L, Postma E & Van Der Herik J (2009) Dimensionality reduction: a comparative review. Tilburg University Technical Report, TiCC-TR 2009–005.
Vargas AM, Kim MS, Tao Y, Lefcourt A & Chen Y-R (2004) Safety inspection of cantaloupes and strawberries using multispectral fluorescence imaging techniques. ASABE conference paper, Paper No. 043056. St. Joseph, MI, USA.
Vargas, A. M., Kim, M. S., Tao, Y., Lefcourt, A. M., Chen, Y.-R., Luo, Y., Song, Y., & Buchanan, R. (2005). Detection of fecal contamination on cantaloupes using hyperspectral fluorescence imagery. Journal of Food Science, 70(8), e471–e476.
Varshney, P. K., & Arora, M. K. (2004). Advanced image processing techniques for remotely sensed hyperspectral data. Berlin Heidelberg: Springer-Verlag.
Vermeulen, P., Pierna, J. A., Van Egmond, H. P., Dardenne, P., & Baeten, V. (2011). Online detection and quantification of ergot bodies in cereals using infrared hyperspectral imaging. Food additives and contaminants. Part A, Chemistry, Analysis, Control, Exposure & Risk Assessment, 29(2), 232–240.
Vigneau, N., Ecarnot, M., Rabatel, G., & Roumet, P. (2011). Potential of field hyperspectral imaging as a non destructive method to assess leaf nitrogen content in wheat. Field Crops Research, 122(1), 25–31.
Wang, D., Dowell, F. E., Ram, M. S., & Schapaugh, W. T. (2003). Classification of fungal-damaged soybean seeds using near-infrared spectroscopy. International Journal of Food Properties, 7(1), 75–82.
Wang, H., Li, C., & Wang, M. (2013). Qualitative determination of onion internal quality using hyperspectral imaging with reflectance, interactance, and transmittance modes. Transactions of ASABE, 56(4), 1–14.
Wang, K., Chi, G., Lau, R., & Chen, T. (2011). Multivariate calibration of near infrared spectroscopy in the presence of light scattering effect: a comparative study. Analytical Letters, 44(5), 824–836.
Wang, L., Pu, H., & Sun, D.-W. (2016). Estimation of chlorophyll-a concentration of different seasons in outdoor ponds using hyperspectral imaging. Talanta, 147, 422–429.
Wang, W., & Paliwal, J. (2006). Spectral data compression and analyses techniques to discriminate wheat classes. Transactions of the ASABE, 49(5), 1607–1612.
Wang, W., & Paliwal, J. (2007). Near-infrared spectroscopy and imaging in food quality and safety. Sensing and Instrumentation for Food Quality and Safety, 1(4), 193–207.
Wang, W., Li, C., Tollner, E. W., Gitaitis, R. D., & Rains, G. C. (2012). Shortwave infrared hyperspectral imaging for detecting sour skin (Burkholderiacepacia)-infected onions. Journal of Food Engineering, 109(1), 38–48.
Wang W, Paliwal J & Jayas DS (2004). Determination of moisture content of ground wheat using near-infrared spectroscopy. ASABE conference paper, Paper No. MB04–200, St. Joseph, MI, USA.
Williams, P., Geladi, P., Fox, G., & Manley, M. (2009). Maize kernel hardness classification by near infrared (NIR) hyperspectral imaging and multivariate data analysis. Analytica Chimica Acta, 653(2), 121–130.
Wold, H. (1975). Soft modeling by latent variables. In Gani (Ed.), Nonlinear iterative partial least squares approach. Perspectives in probability and statistics (pp. 520–540). London: Academic Press.
Wold, J. P. (2016). On-line and non-destructive measurement of core temperature in heat treated fish cakes by NIR hyperspectral imaging. Innovative Food Science and Emerging Technologies, 33, 431–437.
Wold, J. P., O’Farrell, M., Hoy, M., & Tschudi, J. (2011). On-line determination and control of fat content in batches of beef trimmings by NIR imaging spectroscopy. Meat Science, 89(3), 317–324.
Wold, S., Antti, H., Lindgren, F., & Ohman, J. (1998). Orthogonal signal correction of near-infrared spectra. Chemometrics and Intelligent Laboratory Systems, 44(1–2), 175–185.
Wold, S., Martens, H., & Wold, H. (1983). The multivariate calibration problem in chemistry solved by the PLS method. In Ruhe & Kagstrom (eds) proceedings of the conference on matrix pencils. Lecture notes in mathematics, 973 (pp. 286–293). Heidelberg: Springer.
Wu, D., & Sun, D.-W. (2013a). Advanced applications of hyperspectral imaging technology for food quality and safety analysis and assessment: a review-part I: fundamentals. Innovative Food Science and Emerging Technologies, 19, 1–14.
Wu, D., & Sun, D.-W. (2013b). Advanced applications of hyperspectral imaging technology for food quality and safety analysis and assessment: a review-part II: applications. Innovative Food Science and Emerging Technologies, 19, 15–28.
Wu, D., Shi, H., Wang, S., He, Y., Bao, Y., & Liu, K. (2012). Rapid prediction of moisture content of dehydrated prawns using online hyperspectral imaging system. Analytica Chimica Acta, 726, 57–66.
Wu, W., Mallet, Y., Walczak, B., Penninckx, W., Massart, D. L., Heuerding, S., & Erni, F. (1996). Comparison of regularized discriminant analysis, linear discriminant analysis and quadratic discriminant analysis, applied to NIR data. Analytica Chimica Acta, 329(3), 257–265.
Xie, A., Sun, D.-W., Zhu, Z., & Pu, H. (2016). Nondestructive measurements of freezing parameters of frozen porcine meat by NIR hyperspectral imaging. Food and Bioprocess Technology, 9(9), 1444–1454.
Xie, C., Shao, Y., Li, X., & Hea, Y. (2015). Detection of early blight and late blight diseases on tomato leaves using hyperspectral imaging. Scientific Reports, 5, 16564.
Xing, J., Bravo, C., Jancsok, P. T., Ramon, H., & De Baerdemaeker, J. (2005). Detecting bruises on ‘golden delicious’ apples using hyperspectral imaging with multiple wavebands. Biosystems Engineering, 90(1), 27–36.
Xing, J., Hung, P. V., Symons, S., Shahin, M., & Hatcher, D. (2009). Using a short wavelength infrared (SWIR) hyperspectral imaging system to predict alpha amylase activity in individual Canadian western wheat kernels. Sensing and Instrumentation for Food Quality and Safety, 3(4), 211–218.
Xing, J., Saeys, W., & De Baerdemaeker, J. (2007). Combination of chemometric tools and image processing for bruise detection on apples. Computers and Electronics in Agriculture, 56(1), 1–13.
Xing, J., Symons, S., Hatcher, D., & Shahin, M. (2011). Comparison of short-wavelength infrared (SWIR) hyperspectral imaging system with an FT-NIR spectrophotometer for predicting alpha-amylase activities in individual Canadian western red spring (CWRS) wheat kernels. Biosystems Engineering, 108(4), 303–310.
Xu, J.-L., Riccioli, C., & Sun, D.-W. (2016a). Development of an alternative technique for rapid and accurate determination of fish caloric density based on hyperspectral imaging. Journal of Food Engineering, 190, 185–194.
Xu, J.-L., Riccioli, C., & Sun, D.-W. (2016b). Efficient integration of particle analysis in hyperspectral imaging for rapid assessment of oxidative degradation in salmon fillet. Journal of Food Engineering, 169, 259–271.
Yang, C. (2012). A high-resolution airborne four-camera imaging system for agricultural remote sensing. Computers and Electronics in Agriculture, 88, 13–24.
Zayas, I., Pomeranz, Y., & Lai, F. S. (1985). Discrimination between Arthur and Arkan wheats by image analysis. Cereal Chemistry, 62(6), 478–480.
Zhang, H., Paliwal, J., Jayas, D. S., & White, N. D. G. (2007). Classification of fungal infected wheat kernels using near-infrared reflectance hyperspectral imaging and support vector machine. Transactions of the ASABE, 50(5), 1779–1785.
Zhang, Q., Wang, H., Plemmons, R. J., & Pauca, V. P. (2008). Tensor methods for hyperspectral data analysis: a space object material identification study. Journal of the Optical Society of America. A, Optics, Image Science, and Vision, 25(12), 3001–3012.
Zhang, X., Liu, F., He, Y., & Li, X. (2012). Application of hyperspectral imaging and chemometric calibrations for variety discrimination of maize seeds. Sensors, 12(12), 17234–17246.
Zheng, G., Chen, Y., Intes, X., Chance, B., & Glickson, J. D. (2004). Contrast-enhanced near-infrared (NIR) optical imaging for subsurface cancer detection. Journal of Porphyrins and Phthalocyanines, 8(9), 1106–1117.
Zhu, Q. B., Huang, M., Zhao, X., & Wang, S. (2013). Wavelength selection of hyperspectral scattering image using new semi-supervised affinity propagation for prediction of firmness and soluble solid content in apples. Food Analytical Methods, 6(1), 334–342.
Zude, M., Herold, B., Roger, J.-M., Bellon-Maurel, V., & Landahl, S. (2006). Non-destructive tests on the prediction of apple fruit flesh firmness and soluble solids content on tree and in shelf life. Journal of Food Engineering, 77(2), 254–260.
Zupan J & Gesteinger J (1993) Neural networks for chemists: an introduction. Wiley, VCH, Weinheim, Germany.
Zupan, J. (1994). Introduction of artificial neural network (ANN) methods: what they are and how to use them. Acta Chimica Slovenica, 41(3), 327–352.
Acknowledgments
The authors thank the University of Manitoba Graduate Fellowship, the Graduate Enhancement of Tri-Council Stipends (GETS) program and the Natural Sciences and Engineering Research Council of Canada for funding this study. This research was also supported by the International S&T Cooperation Program of China (2015DFA71150) and the International S&T Cooperation Program of Guangdong Province, China (2013B051000010).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ravikanth, L., Jayas, D.S., White, N.D.G. et al. Extraction of Spectral Information from Hyperspectral Data and Application of Hyperspectral Imaging for Food and Agricultural Products. Food Bioprocess Technol 10, 1–33 (2017). https://doi.org/10.1007/s11947-016-1817-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11947-016-1817-8