Abstract
Mesenchymal chondrosarcoma is a rare but deadly form of chondrosarcoma that typically affects adolescents and young adults. While curative intent is possible for patients with localized disease, few options exist for patients in the unresectable/metastatic setting. Thus, it is imperative to understand the fusion-driven biology of this rare malignant neoplasm so as to lead to the future development of better therapeutics for this disease. This manuscript will briefly review the clinical and pathologic features of mesenchymal chondrosarcoma followed by an appraisal of existing data linked to the fusions, HEY1-NCOA2 and IRF2BP2-CDX1, and the associated downstream pathways.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mesenchymal chondrosarcoma is a rare malignant neoplasm known for exhibiting features of primitive appearing mesenchymal cells mixed with islands of cartilage differentiation. This entity was first described by Lichtenstein and Bernstein in 1959 and represents less than 5% of all chondrosarcoma cases diagnosed each year in the USA [1]. The peak incidence occurs in the third decade of life, though the range involves children as young as 7 years and elderly individuals as old as 80 years [2, 3•]. This disease generally arises from bone, but extra-osseous variants have been known to involve areas such as visceral organs and meninges (Fig. 1). Frequent sites of involvement include the vertebrae, ribs, pelvic bones, and craniofacial bones (mandible and maxilla), with a predilection toward late local and distant recurrences [4, 5]. For many centers, the treatment of this tumor is still a subject of debate [6]. Localized disease should be managed surgically with wide margin resection with or without radiation, and consideration should be given to the use of systemic chemotherapy [2, 4, 7]. While multimodal strategies reported in the past did not appear to substantially improve prognosis, a recent European Musculoskeletal Oncology Society study reported a reduced risk of recurrence and superior survival metrics (progression-free and overall survivals) with the use of cytotoxic chemotherapy in the neoadjuvant and/or adjuvant setting for localized disease [7, 8]. For patients with metastatic disease, systemic chemotherapy is the primary form of therapy with surgical resection considered for cases in which surgery may result in remission. The approach of our center to localized mesenchymal chondrosarcoma involves neoadjuvant, anthracycline-based chemotherapy, radiation therapy (unless contraindicated), and surgical resection with wide margins. Unfortunately, there is scarcity of available data on mesenchymal chondrosarcoma, which is limited to small-sample retrospective studies without prospective mesenchymal chondrosarcoma-specific trial data [2, 4, 9,10,11]. A recent evaluation of 205 mesenchymal chondrosarcoma patients through the Surveillance, Epidemiology, and End Results (SEER) database demonstrated 5- and 10-year overall survival rates of 51 and 43%, respectively [3•]. Clearly, better therapeutic options are needed for the treatment of patients with mesenchymal chondrosarcoma.
Spectrum of imaging appearance of mesenchymal chondrosarcoma. a Fourteen-year-old boy with a lesion arising from the left temporal bone. b Nine-year-old girl with a soft tissue mass centered in the left middle cranial fossa compressing the temporal lobe. c Twenty-one-year-old woman with a mass arising from a left anterior rib. Note internal calcifications. d Thirty-five-year-old man with a soft tissue mass in the left hemithorax. e Eighteen-year-old woman with a mass arising from the right ilium. f Thirty-six-year-old man with a soft tissue mass in the left gluteal musculature. Note internal calcifications. g Forty-six-year-old woman with a lytic lesion of the proximal tibial metaphysis. h Twenty-four-year-old man with a soft tissue nodule in the anterior compartment of the thigh
A recurrent translocation, HEY1-NCOA2, has been recently identified in the great majority (at least 80%) of mesenchymal chondrosarcomas [12, 13]. This finding is of interest since molecular characterization of this disease may shed light on its pathogenesis and the potential therapeutic avenues that could be explored. The following review aims to (1) summarize the different structural components of the HEY1-NCOA2 fusion, (2) identify potential pathways that might be modulated by it, and (3) suggest future therapeutic strategies for the management of mesenchymal chondrosarcoma.
Pathology of Mesenchymal Chondrosarcoma
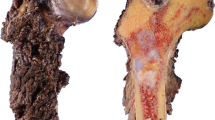
Mesenchymal chondrosarcoma is characterized by having a prominent primitive component comprised of round to spindled cells punctated with islands of mature cartilage (Fig. 2a). The combination of these two components is virtually diagnostic, but a small core needle biopsy sampling only one component or the other can be diagnostically challenging. The primitive cell component has a non-specific immunohistochemical profile but shows SOX9 nuclear reactivity indicative of a chondroid lineage (Fig. 2b). Demonstration of the characteristic HEY1-NCOA2 fusion by a variety of methodologies can be useful in diagnostically challenging cases.
Structure and Function of HEY1
HEY1 (hairy/enhancer-of-split related with YRPW motif 1) is a gene located on the long arm of human chromosome 8 (Fig. 3). It encodes an evolutionary conserved protein of the hairy and enhancer of split-related family of basic helix-loop-helix (bHLH) transcriptional repressors [14]. Three different regions were isolated in HEY1. They include the bHLH domain which mediates the DNA-binding properties of the protein, the Orange domain, which interacts with the bHLH domain to drive and extend the protein interaction and HEY dimerization, and the C-terminus YRPW domain (Fig. 4) [14, 15].
Through its C-terminus bHLH and N-terminus YRPW motifs, HEY1 is thought to act mainly as a transcriptional repressor (Fig. 5). Inhibition of gene promoters is driven by direct DNA binding and dimerization of HEY1 to gene sequences rich in CACGTG/CACGCG and/or CGCGCG, two E-box-like sequences bound with high affinity to the HEY1 DNA-binding domain [14]. This association initiates the recruitment of corepressors to the DNA area of interest, which inhibits target gene expression. Epigenetic modulation is another mechanism through which HEY1 exerts its inhibitory effects. The amino- and the carboxy-terminal-repressive HEY1 domains depend on histone deacetylase-dependent and independent mechanisms, respectively, to suppress target genes [16]. HEY1 transcriptional activity does not seem to be limited to repression only, since many gene targets are overexpressed upon its stimulation [14]. Of note, E-box motifs have not been found in the activated gene promoters, which raises the possibility of a protein-protein interaction mechanism to mediate gene activation [14].
Blunt arrows (┴) indicate inhibition while sharp arrows (→) indicate stimulation. a HEY1 bHLH domain binds E-box-like sequences, which leads to recruitment of corepressor factors to mediate HEY1-related gene expression. b HEY1 bHLH inhibits myogenesis through a promoter competition mechanism with MyoD. c This domain inhibits Runx2 to induce and inhibits chondrogenesis and osteogenesis, respectively. d It interacts with GATA to promote the expression of the genes modulated by it. e HEY1 bHLH might interact with other proteins to modify genes expression. f It competes with FBXO45 and promotes a ubiquitination mechanism disruption, which is responsible of some proteins destruction. The g AD1 and h AD2 domains of NCOA2 recruit coactivator elements and histone methyltransferases that lead to epigenetic modifications via chromatin remodeling
HEY1 is a downstream mediator and effector of the activated Notch developmental and stemness pathway. Gene expressions that are modulated by HEY1 up- or downregulation involve multiple embryological processes, including musculoskeletal, neurological, and cardiovascular development [14, 17,18,19,20,21]. Through its interaction with GATA transcription factors, HEY1 inhibits erythropoiesis- and cardiogenesis-related genes [22,23,24]. FBXO45 (F-box only protein 45) is required for proper neural development [25]. It associates with SKP1 (S-phase kinase-associated protein 1) to form an atypical protein-ubiquitin ligase complex [25, 26]. Through its bHLH and Orange sequences, HEY1 indirectly inhibits FBXO45 activity through redirection of the ubiquitination complex toward other proteins (Fig. 5) [27]. In the embryonic murine inner ear, it also maintains the hairy mechano-sensory cell population and stemness [21].
HEY1 dysregulation alters the expression of genes that intervene in musculoskeletal development [14, 16]. The Notch network is important for the prevention of the premature differentiation of myogenic precursors by sustaining stem/progenitor cell self-renewal, whereas myogenin and MyoD are crucial muscle regulatory factors for muscle differentiation [28, 29]. HEY1 exerts its inhibitory effects on myogenesis at least in part through targeting of myogenin promoters by increasing their methylation and compromising MyoD recruitment to its target promoters (Fig. 5) [16]. HEY1 interaction with GATA does not seem to be relevant in myogenesis as compared to erythropoiesis and cardiogenesis, despite the fact that many myogenic gene promoters contain GATA binding sites in their sequences [16].
There are conflicting data in the literature regarding HEY1 involvement in bone and cartilage formation. In C2C12 mesenchymal stem cells, microarray analysis showed HEY1 to be among the most overexpressed gene upon BMP (bone morphogenetic protein)-induced osteogenic differentiation [30]. In pre-osteoblasts MC3T3 cells, HEY1 abrogated Runx2 transcriptional activity, which is an essential transcription factor for bone development [31]. In mice, HEY1 repression did not affect osteogenic maturation and mineralization, while its induction inhibited mesenchymal stem cell osteogenic differentiation and mineralization properties [32]. Mice overexpressing HEY1 exhibited also a 76% increase in the number of hypertrophic chondrocytes [32]. In response to BMP9, which induces mesenchymal stem cells differentiation into osteoblasts precursors, HEY1 was among the most stimulated genes [33]. However, in this study, HEY1 knockdown reduced BMP9-mediated osteogenic differentiation both in vivo and in vitro, while increasing chondrogenic cell numbers and chondroid matrix formation. High HEY1 levels were also linked to high invasive and metastatic potential in mice injected with osteosarcoma cell lines [34••].
Structure and Function of NCOA2
NCOA2 (nuclear receptor coactivator 2) belongs to the p160 nuclear receptor coactivator family and is located on the long arm of human chromosome 8 (Fig. 3). It encodes a transcriptional coactivator protein for nuclear hormone receptors. Three distinct regions are identified in this gene. They encompass an N-terminus bHLH domain that mediates DNA binding and protein dimerization, a central area with three LXLL motifs that drive NCOA2 interaction with nuclear hormone receptors, and a C-terminus region that contains two transcriptional activation sequences with a relevant role in chromatin remodeling (Figs. 4 and 5) [35]. Without directly binding DNA, NCOA2 associates with nuclear receptors, recruits histone methyltransferases to specific sequences, and mediates chromatin remodeling and transcription of specific target genes. AD1 (activation domain 1) interacts with transcriptional coactivators (CBP and P300), whereas AD2 (activation domain 2) interferes with histone methyltransferases, coactivator-associated arginine methyltransferase 1 (CARM1), and protein arginine methyltransferase 1 (PRMT1) [35,36,37].
NCOA2 rearrangements have been identified in several hematologic (Table 1) and solid tumors (Table 2). This gene’s fusion with SRF, TEAD1, and VGLL2 has been found in infantile spindle cell rhabdomyosarcomas [43, 44]. In a rare variant of alveolar rhabdomyosarcoma, a t(2;8) translocation generated a PAX3-NCOA2 chimeric oncogene [45]. Fusions with AHRR and GTF2I have also been found in soft-tissue angiofibromas [48,49,50]. NCOA2 is also involved in MYST3-NCOA2 and ETV6-NCOA2 translocations in leukemias expressing both T-lymphoid and myeloid markers [38, 41]. Carapeti et al. detected and confirmed the presence of the KAT6A-NCOA2 fusion in acute myeloid leukemias [39, 40]. The transforming properties of KAT6A-NCOA2 and PAX3-NCOA2 were biologically elucidated [45, 55]. The former relies on its interaction with CBP through AD1 for acute myeloid leukemia transformation, whereas the latter requires the presence of both activation domains for alveolar rhabdomyosarcoma genesis.
LACTB2-NCOA2 was identified as a rare but recurrent fusion by whole-genome and transcriptome sequencing in colorectal cancer [56••]. In this study, high NCOA2 levels profoundly abolished colorectal cancer cell line proliferation, colony formation, migration, and invasiveness while increasing their apoptotic rate. In nude mice, NCOA2 overexpression repressed colorectal xenograft growth.
In prostate cancer, NCOA2 was respectively upregulated in 8 and 37% of primary and metastatic tumors [57]. In mice, high NCOA2 levels were sufficient to induce early stages of human prostatic cancer [58]. The same study showed that tumor cells with high NCOA2 expression were more invasive compared to their control counterparts. In prostate cancer cells, NCOA2 overexpression was transcriptionally inhibited by the androgen receptor. NCOA2 inhibition in androgen-dependent cells made them more sensitive to androgen deprivation therapy and stopped tumor growth at low grade stages [58].
Structure of HEY1-NCOA2 Fusion
HEY1 and NCOA2 were fused in at least 80% of mesenchymal chondrosarcomas as detected by RT-PCR, FISH, and genome-wide screen of exon-level expression data and appears to be disease defining [13, 51, 59]. The chimeric fusion is generated from an intra-chromosomal deletion between exon 4 of HEY1 and exon 13 of NCOA2 (Figs. 3 and 4) [59]. Similarly to all fusions involving NCOA2, HEY1-NCOA2 bears the DNA-binding domain of HEY1 and the two C-terminal activation domains of NCOA2 (Fig. 4) [12, 51]. The histone methyltransferase-interacting domain of NCOA2 is also conserved.
Genomics of Mesenchymal Chondrosarcoma
Despite being specific to mesenchymal chondrosarcoma, HEY1-NCOA2 fusion does not seem to be the only recurrent translocation identified in this disease. Therefore, fusion heterogeneity should not be excluded even though one translocation might be more frequent compared to others. Nyquist et al. reported a case of mesenchymal chondrosarcoma with an in-frame t(1;5)(q42;q32) fusion resulting in an IRF2BP2-CDX1 translocation between exon 1 of the IRF2BP2 gene on chromosome 1 and intron 1 of the CDX1 gene on chromosome 5 [60]. CDX1 encodes a transcription factor, with an irregular expression in intestinal cancer [61,62,63]. IRF2BP2 contains a zinc finger motif that can bind DNA. It has the ability to interact with TP53 tumor suppressor and IRF2 oncogene to affect tumorigenesis [64, 65]. Nevertheless, TP53 anomalies have been reported in previous studies including one report that showed 39% of these alterations (loss of protein expression) involving the small cell area while only 7% of the cartilaginous portions of mesenchymal chondrosarcoma samples harbored similar aberrations [66]. Within this same report, the retinoblastoma pathway was the most altered network with 3 of 11 (70%) samples having a homozygous loss of the CDKN2A/p16 locus [66]. Contrary to conventional chondrosarcomas and dedifferentiated chondrosarcomas, no mutations in the IDH1 or IDH2 genes were noted in this same study. Aside from the presence of fusions, no consistent additional genetic abnormalities have been discovered for this disease.
Potentially Activated Pathways in Mesenchymal Chondrosarcoma
Notch Signaling Pathway
As a member of the Notch signaling network, HEY1 is involved in multiple processes. The effects of the Notch pathway are cell dependent but often promote oncogenesis through apoptosis repression, proliferation, and epithelial-to-mesenchymal stimulation [67,68,69]. Nuclear HEY1 expression, in particular, has been associated with local lymph node and neurovascular bundle invasion, as well as adverse prognosis in pancreatic adenocarcinoma [70]. HEY1 has been shown to dramatically reduce gene expression levels of COL2A1, which is responsible for encoding the alpha 1 chain of type II collagen, an essential component of the cartilaginous extracellular matrix [71]. This occurs by binding to the N-box domains in intron 1 of COL2A1, thus modulating the interaction between SOX9 and COL2A1. In line with these findings, Notch pathway inhibition in general, HEY1 more specifically, might be of interest in mesenchymal chondrosarcoma.
Chromatin Remodeling
HEY1-NCOA2 exerts, at least to some extent, its pathogenetic impact by affecting chromatin configuration. Pathogenesis may involve recruitment of coactivators or corepressors through NCOA2 or HEY1 domains to HEY1 target genes, respectively (Fig. 5). Therefore, in mesenchymal chondrosarcoma, chromatin modulation through DNA methylation and HDAC inhibitors might also be useful as epigenetic modifier treatments.
Apoptosis
A recent report showed that malignant mesenchymal chondroblasts have higher expression of CD99, PKC-α, PDGFR-α, and Bcl-2 antigens, with proliferation pathways centering around PKC-α and PDGFR-α networks [72]. A separate study confirmed the high protein expression of Bcl-2 and Bcl-xL in mesenchymal chondrosarcoma [73]. Activated PKC-α phosphorylates Bcl-2 and subsequently slows apoptosis. CD99 is a mediator of MAPK pathway activation via the PKC pathway and has been shown to be positive in mesenchymal chondrosarcoma [74,75,76]. Compared to malignant chondrocytes, malignant mesenchymal chondroblasts exhibit higher expression of Akt and mTOR [77]. PDGFR-α is an upstream inducer of Akt signaling in mesenchymal chondrosarcoma. Therefore, it is worth mentioning the potential role of mTOR and/or PDGFR inhibitors in the management of mesenchymal chondrosarcoma.
TGF-β1 Signaling
Transforming growth factor beta (TGF-β) superfamily represents a large group of conserved genes that encode ligands and receptors which interact with Smad transcription factors to regulate gene transcription [78]. Along with BMP, TGF-β signaling assists in the regulation and maintenance of SOX9, which is considered the master regulator of cartilage development. Van Oosterwijk et al. have demonstrated that mesenchymal chondrosarcoma expresses high levels of p-SMAD2 by immunohistochemistry, particularly in the small cell components of the tumor [73]. This protein directly interacts with TGF-β. Furthermore, p-SMAD1 and PAI-1 were highly expressed in approximately half of the small cell component and in the third of the cartilaginous components. These findings suggest a possible therapeutic role for small molecule inhibitors of the TGF-β signaling pathway.
Limitations
Efforts have been made to generate mesenchymal chondrosarcoma cell lines that will provide researchers tools to study this disease. It was not until recently that a novel mesenchymal chondrosarcoma cell line (MCS170) was reported by the Leiden group in the Netherlands, in which the HEY1-NCOA2 translocation was identified by FISH, RT-PCR, and sequencing analyses [79••]. A crucial driver role of a genetic translocation is implied by its recurrence, its association with few or no other genetic abnormalities, and its current restriction to one tumor phenotype [80]. Despite the fact that the HEY1 DNA-binding and the NCOA2 transcriptional activation domains are always preserved in the generated HEY1-NCOA2 translocation, the contribution of each region to the effects of the fusion may be context and cell dependent (Fig. 5). The area that is responsible for the normal function of the wild-type protein might also be absent in the genetic fusion [80]. This is the case of NCOA2 that is characterized by its DNA-binding domain, which is lacking in the HEY1-NCOA2 translocation. There also might be a difference in the genes regulated by the wild-type transcription factor compared to its respective chimeric counterpart. Moreover, the activity of the amino-terminal end seems more complicated than being a simple potentiator of the transcription of the carboxy-terminal partner targets [81].
Conclusions
Discovery of the HEY1-NCOA2 translocation as a specific and recurrent fusion in mesenchymal chondrosarcoma is an important breakthrough for characterizing and understanding the pathogenesis of this disease. The subsequent identification of the pathways modulated by this fusion would help guide and develop drugs to assess their efficacy in treating mesenchymal chondrosarcoma. Based on the available data on individual HEY1 and NCOA2, HEY1-NCOA2 fusion evokes many different mechanisms to promote sarcomagenesis, such as direct DNA binding, protein-protein interaction, and epigenetic modification. It is likely that the combination of these pathway dysregulations is what allows this single translocation and resulting chimeric fusion protein to drive the biology of this rare and aggressive sarcoma.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
Lichtenstein L, Bernstein D. Unusual benign and malignant chondroid tumors of bone. A survey of some mesenchymal cartilage tumors and malignant chondroblastic tumors, including a few multicentric ones, as well as many atypical benign chondroblastomas and chondromyxoid fibromas. Cancer. 1959;12(6):1142–57. https://doi.org/10.1002/1097-0142(195911/12)12:6<1142::AID-CNCR2820120610>3.0.CO;2-D.
Frezza AM, Cesari M, Baumhoer D, Biau D, Bielack S, Campanacci DA, et al. Mesenchymal chondrosarcoma: prognostic factors and outcome in 113 patients. A European musculoskeletal oncology society study. Eur J Cancer. 2015;51(3):374–81. https://doi.org/10.1016/j.ejca.2014.11.007.
• Schneiderman BA, Kliethermes SA, Nystrom LM. Survival in mesenchymal chondrosarcoma varies based on age and tumor location: a survival analysis of the SEER database. Clin Orthop Relat Res. 2017;475(3):799–805. This is a study that evaluates prognosticators of mesenchymal chondrosarcoma in 205 patients identified via the SEER database. It showed that the main poor prognostic factors were tumor size and presence of metastases at diagnosis. Young patients and/or cranial disease had better survival when compared to older patients and/or axial or appendicular tumors
Cesari M, Bertoni F, Bacchini P, Mercuri M, Palmerini E, Ferrari S. Mesenchymal chondrosarcoma. An analysis of patients treated at a single institution. Tumori. 2007;93(5):423–7.
Kim MJ, et al. Chondrosarcoma: with updates on molecular genetics. Sarcoma. 2011;2011:405437.
Riedel RF, Larrier N, Dodd L, Kirsch D, Martinez S, Brigman BE. The clinical management of chondrosarcoma. Curr Treat Options in Oncol. 2009;10(1–2):94–106. https://doi.org/10.1007/s11864-009-0088-2.
Kawaguchi S, Weiss I, Lin PP, Huh WW, Lewis VO. Radiation therapy is associated with fewer recurrences in mesenchymal chondrosarcoma. Clin Orthop Relat Res. 2014;472(3):856–64. https://doi.org/10.1007/s11999-013-3064-x.
Dantonello TM, Int-Veen C, Leuschner I, Schuck A, Furtwaengler R, Claviez A, et al. Mesenchymal chondrosarcoma of soft tissues and bone in children, adolescents, and young adults: experiences of the CWS and COSS study groups. Cancer. 2008;112(11):2424–31. https://doi.org/10.1002/cncr.23457.
Bishop MW, et al. Mesenchymal chondrosarcoma in children and young adults: a single institution retrospective review. Sarcoma. 2015;2015:608279.
Shakked RJ, Geller DS, Gorlick R, Dorfman HD. Mesenchymal chondrosarcoma: clinicopathologic study of 20 cases. Arch Pathol Lab Med. 2012;136(1):61–75. https://doi.org/10.5858/arpa.2010-0362-OA.
• Xu J, et al. Mesenchymal chondrosarcoma of bone and soft tissue: a systematic review of 107 patients in the past 20 years. PLoS One. 2015;10(4):e0122216. The authors of this study searched medical libraries for all mesenchymal chondrosarcoma cohort reports available in the literature. They showed that surgery constitutes the mainstay therapy for localized disease. Survival was not affected with adjuvant radiation or chemotherapy.
Wang L, Motoi T, Khanin R, Olshen A, Mertens F, Bridge J, et al. Identification of a novel, recurrent HEY1-NCOA2 fusion in mesenchymal chondrosarcoma based on a genome-wide screen of exon-level expression data. Genes Chromosomes Cancer. 2012;51(2):127–39. https://doi.org/10.1002/gcc.20937.
Nakayama R, Miura Y, Ogino J, Susa M, Watanabe I, Horiuchi K, et al. Detection of HEY1-NCOA2 fusion by fluorescence in-situ hybridization in formalin-fixed paraffin-embedded tissues as a possible diagnostic tool for mesenchymal chondrosarcoma. Pathol Int. 2012;62(12):823–6. https://doi.org/10.1111/pin.12022.
Heisig J, Weber D, Englberger E, Winkler A, Kneitz S, Sung WK, et al. Target gene analysis by microarrays and chromatin immunoprecipitation identifies HEY proteins as highly redundant bHLH repressors. PLoS Genet. 2012;8(5):e1002728. https://doi.org/10.1371/journal.pgen.1002728.
Taelman V, van Wayenbergh R, Sölter M, Pichon B, Pieler T, Christophe D, et al. Sequences downstream of the bHLH domain of the Xenopus hairy-related transcription factor-1 act as an extended dimerization domain that contributes to the selection of the partners. Dev Biol. 2004;276(1):47–63. https://doi.org/10.1016/j.ydbio.2004.08.019.
Buas MF, Kabak S, Kadesch T. The Notch effector Hey1 associates with myogenic target genes to repress myogenesis. J Biol Chem. 2010;285(2):1249–58. https://doi.org/10.1074/jbc.M109.046441.
Fischer A, Schumacher N, Maier M, Sendtner M, Gessler M. The Notch target genes Hey1 and Hey2 are required for embryonic vascular development. Genes Dev. 2004;18(8):901–11. https://doi.org/10.1101/gad.291004.
Kokubo H, Miyagawa-Tomita S, Johnson RL. Hesr, a mediator of the Notch signaling, functions in heart and vessel development. Trends Cardiovasc Med. 2005;15(5):190–4. https://doi.org/10.1016/j.tcm.2005.05.005.
Kokubo H, Miyagawa-Tomita S, Nakazawa M, Saga Y, Johnson RL. Mouse hesr1 and hesr2 genes are redundantly required to mediate Notch signaling in the developing cardiovascular system. Dev Biol. 2005;278(2):301–9. https://doi.org/10.1016/j.ydbio.2004.10.025.
Rutenberg JB, Fischer A, Jia H, Gessler M, Zhong TP, Mercola M. Developmental patterning of the cardiac atrioventricular canal by Notch and hairy-related transcription factors. Development. 2006;133(21):4381–90. https://doi.org/10.1242/dev.02607.
Benito-Gonzalez A, Doetzlhofer A. Hey1 and Hey2 control the spatial and temporal pattern of mammalian auditory hair cell differentiation downstream of hedgehog signaling. J Neurosci. 2014;34(38):12865–76. https://doi.org/10.1523/JNEUROSCI.1494-14.2014.
Elagib KE, Xiao M, Hussaini IM, Delehanty LL, Palmer LA, Racke FK, et al. Jun blockade of erythropoiesis: role for repression of GATA-1 by HERP2. Mol Cell Biol. 2004;24(17):7779–94. https://doi.org/10.1128/MCB.24.17.7779-7794.2004.
Kathiriya IS, King IN, Murakami M, Nakagawa M, Astle JM, Gardner KA, et al. Hairy-related transcription factors inhibit GATA-dependent cardiac gene expression through a signal-responsive mechanism. J Biol Chem. 2004;279(52):54937–43. https://doi.org/10.1074/jbc.M409879200.
Fischer A, Klattig J, Kneitz B, Diez H, Maier M, Holtmann B, et al. Hey basic helix-loop-helix transcription factors are repressors of GATA4 and GATA6 and restrict expression of the GATA target gene ANF in fetal hearts. Mol Cell Biol. 2005;25(20):8960–70. https://doi.org/10.1128/MCB.25.20.8960-8970.2005.
Tada H, Okano HJ, Takagi H, Shibata S, Yao I, Matsumoto M, et al. Fbxo45, a novel ubiquitin ligase, regulates synaptic activity. J Biol Chem. 2010;285(6):3840–9. https://doi.org/10.1074/jbc.M109.046284.
Saiga T, Fukuda T, Matsumoto M, Tada H, Okano HJ, Okano H, et al. Fbxo45 forms a novel ubiquitin ligase complex and is required for neuronal development. Mol Cell Biol. 2009;29(13):3529–43. https://doi.org/10.1128/MCB.00364-09.
Salat D, Winkler A, Urlaub H, Gessler M. Hey bHLH proteins interact with a FBXO45 containing SCF ubiquitin ligase complex and induce its translocation into the nucleus. PLoS One. 2015;10(6):e0130288. https://doi.org/10.1371/journal.pone.0130288.
Schuster-Gossler K, Cordes R, Gossler A. Premature myogenic differentiation and depletion of progenitor cells cause severe muscle hypotrophy in Delta1 mutants. Proc Natl Acad Sci U S A. 2007;104(2):537–42. https://doi.org/10.1073/pnas.0608281104.
Vasyutina E, Lenhard DC, Wende H, Erdmann B, Epstein JA, Birchmeier C. RBP-J (Rbpsuh) is essential to maintain muscle progenitor cells and to generate satellite cells. Proc Natl Acad Sci U S A. 2007;104(11):4443–8. https://doi.org/10.1073/pnas.0610647104.
de Jong DS, Steegenga WT, Hendriks JMA, van Zoelen EJJ, Olijve W, Dechering KJ. Regulation of Notch signaling genes during BMP2-induced differentiation of osteoblast precursor cells. Biochem Biophys Res Commun. 2004;320(1):100–7. https://doi.org/10.1016/j.bbrc.2004.05.150.
Zamurovic N, Cappellen D, Rohner D, Susa M. Coordinated activation of notch, Wnt, and transforming growth factor-beta signaling pathways in bone morphogenic protein 2-induced osteogenesis. Notch target gene Hey1 inhibits mineralization and Runx2 transcriptional activity. J Biol Chem. 2004;279(36):37704–15. https://doi.org/10.1074/jbc.M403813200.
Salie R, Kneissel M, Vukevic M, Zamurovic N, Kramer I, Evans G, et al. Ubiquitous overexpression of Hey1 transcription factor leads to osteopenia and chondrocyte hypertrophy in bone. Bone. 2010;46(3):680–94. https://doi.org/10.1016/j.bone.2009.10.022.
Sharff KA, Song WX, Luo X, Tang N, Luo J, Chen J, et al. Hey1 basic helix-loop-helix protein plays an important role in mediating BMP9-induced osteogenic differentiation of mesenchymal progenitor cells. J Biol Chem. 2009;284(1):649–59. https://doi.org/10.1074/jbc.M806389200.
•• Tsuru A, et al. Hairy/enhancer-of-split related with YRPW motif protein 1 promotes osteosarcoma metastasis via matrix metallopeptidase 9 expression. Br J Cancer. 2015;112(7):1232–40. This basic science report aimed to detect the relationship between HEY1 and osteosarcoma. The authors used human cell lines and a murine xenograft model for this purpose. They found that HEY1 promotes osteosarcoma’s metastatic potential by upregulating matrix metalloproteinase 9 expression.
Xu J, Wu RC, O'Malley BW. Normal and cancer-related functions of the p160 steroid receptor co-activator (SRC) family. Nat Rev Cancer. 2009;9(9):615–30. https://doi.org/10.1038/nrc2695.
Chen D, Ma H, Hong H, Koh SS, Huang SM, Schurter BT, et al. Regulation of transcription by a protein methyltransferase. Science. 1999;284(5423):2174–7. https://doi.org/10.1126/science.284.5423.2174.
Koh SS, Chen D, Lee YH, Stallcup MR. Synergistic enhancement of nuclear receptor function by p160 coactivators and two coactivators with protein methyltransferase activities. J Biol Chem. 2001;276(2):1089–98. https://doi.org/10.1074/jbc.M004228200.
Troke PJ, et al. MOZ fusion proteins in acute myeloid leukaemia. Biochem Soc Symp. 2006;73:23–39. https://doi.org/10.1042/bss0730023.
Carapeti M, Aguiar RC, Goldman JM, Cross NC. A novel fusion between MOZ and the nuclear receptor coactivator TIF2 in acute myeloid leukemia. Blood. 1998;91(9):3127–33.
Carapeti M, Aguiar RCT, Watmore AE, Goldman JM, Cross NCP. Consistent fusion of MOZ and TIF2 in AML with inv(8)(p11q13). Cancer Genet Cytogenet. 1999;113(1):70–2. https://doi.org/10.1016/S0165-4608(99)00007-2.
Strehl S, Nebral K, Konig M, Harbott J, Strobl H, Ratei R, et al. ETV6-NCOA2: a novel fusion gene in acute leukemia associated with coexpression of T-lymphoid and myeloid markers and frequent NOTCH1 mutations. Clin Cancer Res. 2008;14(4):977–83. https://doi.org/10.1158/1078-0432.CCR-07-4022.
Brown RA, Kwong BY, McCalmont TH, Ragsdale B, Ma L, Cheung C, et al. ETV3-NCOA2 in indeterminate cell histiocytosis: clonal translocation supports sui generis. Blood. 2015;126(20):2344–5. https://doi.org/10.1182/blood-2015-07-655530.
Mosquera JM, Sboner A, Zhang L, Kitabayashi N, Chen CL, Sung YS, et al. Recurrent NCOA2 gene rearrangements in congenital/infantile spindle cell rhabdomyosarcoma. Genes Chromosomes Cancer. 2013;52(6):538–50. https://doi.org/10.1002/gcc.22050.
Alaggio R, Zhang L, Sung YS, Huang SC, Chen CL, Bisogno G, et al. A molecular study of pediatric spindle and Sclerosing rhabdomyosarcoma: identification of novel and recurrent VGLL2-related fusions in infantile cases. Am J Surg Pathol. 2016;40(2):224–35. https://doi.org/10.1097/PAS.0000000000000538.
Sumegi J, Streblow R, Frayer RW, Dal Cin P, Rosenberg A, Meloni-Ehrig A, et al. Recurrent t(2;2) and t(2;8) translocations in rhabdomyosarcoma without the canonical PAX-FOXO1 fuse PAX3 to members of the nuclear receptor transcriptional coactivator family. Genes Chromosomes Cancer. 2010;49(3):224–36. https://doi.org/10.1002/gcc.20731.
Meloni-Ehrig A, Smith B, Zgoda JA, Greenberg J, Perdahl-Wallace E, Zaman S, et al. Translocation (2;8)(q35;q13): a recurrent abnormality in congenital embryonal rhabdomyosarcoma. Cancer Genet Cytogenet. 2009;191(1):43–5. https://doi.org/10.1016/j.cancergencyto.2009.01.010.
Hosoi H, Kakazu N, Konishi E, Tsuchihashi Y, Hada S, Amaya E, et al. A novel PAX3 rearrangement in embryonal rhabdomyosarcoma. Cancer Genet Cytogenet. 2009;189(2):98–104. https://doi.org/10.1016/j.cancergencyto.2008.10.016.
Jin Y, Möller E, Nord KH, Mandahl N, von Steyern FV, Domanski HA, et al. Fusion of the AHRR and NCOA2 genes through a recurrent translocation t(5;8)(p15;q13) in soft tissue angiofibroma results in upregulation of aryl hydrocarbon receptor target genes. Genes Chromosomes Cancer. 2012;51(5):510–20. https://doi.org/10.1002/gcc.21939.
Panagopoulos I, Gorunova L, Viset T, Heim S. Gene fusions AHRR-NCOA2, NCOA2-ETV4, ETV4-AHRR, P4HA2-TBCK, and TBCK-P4HA2 resulting from the translocations t(5;8;17)(p15;q13;q21) and t(4;5)(q24;q31) in a soft tissue angiofibroma. Oncol Rep. 2016;36(5):2455–62. https://doi.org/10.3892/or.2016.5096.
Arbajian E, Magnusson L, Mertens F, Domanski HA, Vult von Steyern F, Nord KH. A novel GTF2I/NCOA2 fusion gene emphasizes the role of NCOA2 in soft tissue angiofibroma development. Genes Chromosomes Cancer. 2013;52(3):330–1. https://doi.org/10.1002/gcc.22033.
Panagopoulos I, et al. Chromosome aberrations and HEY1-NCOA2 fusion gene in a mesenchymal chondrosarcoma. Oncol Rep. 2014;32(1):40–4. https://doi.org/10.3892/or.2014.3180.
Yoshihara K, Wang Q, Torres-Garcia W, Zheng S, Vegesna R, Kim H, et al. The landscape and therapeutic relevance of cancer-associated transcript fusions. Oncogene. 2015;34(37):4845–54. https://doi.org/10.1038/onc.2014.406.
Kalyana-Sundaram S, Shankar S, DeRoo S, Iyer MK, Palanisamy N, Chinnaiyan AM, et al. Gene fusions associated with recurrent amplicons represent a class of passenger aberrations in breast cancer. Neoplasia. 2012;14(8):702–8. https://doi.org/10.1593/neo.12914.
Robinson DR, Kalyana-Sundaram S, Wu YM, Shankar S, Cao X, Ateeq B, et al. Functionally recurrent rearrangements of the MAST kinase and Notch gene families in breast cancer. Nat Med. 2011;17(12):1646–51. https://doi.org/10.1038/nm.2580.
Deguchi K, Ayton PM, Carapeti M, Kutok JL, Snyder CS, Williams IR, et al. MOZ-TIF2-induced acute myeloid leukemia requires the MOZ nucleosome binding motif and TIF2-mediated recruitment of CBP. Cancer Cell. 2003;3(3):259–71. https://doi.org/10.1016/S1535-6108(03)00051-5.
•• Yu J, et al. Disruption of NCOA2 by recurrent fusion with LACTB2 in colorectal cancer. Oncogene. 2016;35(2):187–95. The authors identified a LACTB2-NCOA2 fusion in colorectal cancer cases with low NCOA2 expression. Inhibiting and expressing NCOA2 in normal colonocytes and colorectal cancer cells induced and repressed their tumorigenic properties, respectively. LACTB2-NCOA2 is a recurrent fusion responsible of disrupting wild-type NCOA2 in colorectal cancer.
Taylor BS, Schultz N, Hieronymus H, Gopalan A, Xiao Y, Carver BS, et al. Integrative genomic profiling of human prostate cancer. Cancer Cell. 2010;18(1):11–22. https://doi.org/10.1016/j.ccr.2010.05.026.
Qin J, Lee HJ, Wu SP, Lin SC, Lanz RB, Creighton CJ, et al. Androgen deprivation-induced NCoA2 promotes metastatic and castration-resistant prostate cancer. J Clin Invest. 2014;124(11):5013–26. https://doi.org/10.1172/JCI76412.
Moriya K, Katayama S, Onuma M, Rikiishi T, Hosaka M, Watanabe M, et al. Mesenchymal chondrosarcoma diagnosed on FISH for HEY1-NCOA2 fusion gene. Pediatr Int. 2014;56(5):e55–7. https://doi.org/10.1111/ped.12407.
Nyquist KB, Panagopoulos I, Thorsen J, Haugom L, Gorunova L, Bjerkehagen B, et al. Whole-transcriptome sequencing identifies novel IRF2BP2-CDX1 fusion gene brought about by translocation t(1;5)(q42;q32) in mesenchymal chondrosarcoma. PLoS One. 2012;7(11):e49705. https://doi.org/10.1371/journal.pone.0049705.
Kang JM, Lee BH, Kim N, Lee HS, Lee HE, Park JH, et al. CDX1 and CDX2 expression in intestinal metaplasia, dysplasia and gastric cancer. J Korean Med Sci. 2011;26(5):647–53. https://doi.org/10.3346/jkms.2011.26.5.647.
Chan CW, et al. Gastrointestinal differentiation marker cytokeratin 20 is regulated by homeobox gene CDX1. Proc Natl Acad Sci U S A. 2009;106(6):1936–41. https://doi.org/10.1073/pnas.0812904106.
Wong NA, et al. Loss of CDX1 expression in colorectal carcinoma: promoter methylation, mutation, and loss of heterozygosity analyses of 37 cell lines. Proc Natl Acad Sci U S A. 2004;101(2):574–9. https://doi.org/10.1073/pnas.0307190101.
Childs KS, Goodbourn S. Identification of novel co-repressor molecules for interferon regulatory factor-2. Nucleic Acids Res. 2003;31(12):3016–26. https://doi.org/10.1093/nar/gkg431.
Koeppel M, van Heeringen SJ, Smeenk L, Navis AC, Janssen-Megens EM, Lohrum M. The novel p53 target gene IRF2BP2 participates in cell survival during the p53 stress response. Nucleic Acids Res. 2009;37(2):322–35. https://doi.org/10.1093/nar/gkn940.
Meijer D, de Jong D, Pansuriya TC, van den Akker BE, Picci P, Szuhai K, et al. Genetic characterization of mesenchymal, clear cell, and dedifferentiated chondrosarcoma. Genes Chromosomes Cancer. 2012;51(10):899–909. https://doi.org/10.1002/gcc.21974.
Zavadil J, Cermak L, Soto-Nieves N, Böttinger EP. Integration of TGF-beta/Smad and Jagged1/Notch signalling in epithelial-to-mesenchymal transition. EMBO J. 2004;23(5):1155–65. https://doi.org/10.1038/sj.emboj.7600069.
Miele L, Osborne B. Arbiter of differentiation and death: Notch signaling meets apoptosis. J Cell Physiol. 1999;181(3):393–409. https://doi.org/10.1002/(SICI)1097-4652(199912)181:3<393::AID-JCP3>3.0.CO;2-6.
Artavanis-Tsakonas S, Rand MD, Lake RJ. Notch signaling: cell fate control and signal integration in development. Science. 1999;284(5415):770–6. https://doi.org/10.1126/science.284.5415.770.
Mann CD, Bastianpillai C, Neal CP, Masood MM, Jones DJL, Teichert F, et al. Notch3 and HEY-1 as prognostic biomarkers in pancreatic adenocarcinoma. PLoS One. 2012;7(12):e51119. https://doi.org/10.1371/journal.pone.0051119.
Grogan SP, Olee T, Hiraoka K, Lotz MK. Repression of chondrogenesis through binding of notch signaling proteins HES-1 and HEY-1 to N-box domains in the COL2A1 enhancer site. Arthritis Rheum. 2008;58(9):2754–63. https://doi.org/10.1002/art.23730.
Brown RE, Boyle JL. Mesenchymal chondrosarcoma: molecular characterization by a proteomic approach, with morphogenic and therapeutic implications. Ann Clin Lab Sci. 2003;33(2):131–41.
van Oosterwijk JG, Meijer D, van Ruler MAJH, van den Akker BEWM, Oosting J, Krenács T, et al. Screening for potential targets for therapy in mesenchymal, clear cell, and dedifferentiated chondrosarcoma reveals Bcl-2 family members and TGFbeta as potential targets. Am J Pathol. 2013;182(4):1347–56. https://doi.org/10.1016/j.ajpath.2012.12.036.
Hahn MJ, Yoon SS, Sohn HW, Song HG, Park SH, Kim TJ. Differential activation of MAP kinase family members triggered by CD99 engagement. FEBS Lett. 2000;470(3):350–4. https://doi.org/10.1016/S0014-5793(00)01330-2.
Kasinrerk W, Tokrasinwit N, Moonsom S, Stockinger H. CD99 monoclonal antibody induce homotypic adhesion of Jurkat cells through protein tyrosine kinase and protein kinase C-dependent pathway. Immunol Lett. 2000;71(1):33–41. https://doi.org/10.1016/S0165-2478(99)00165-0.
Granter SR, Renshaw AA, Fletcher CDM, Bhan AK, Rosenberg AE. CD99 reactivity in mesenchymal chondrosarcoma. Hum Pathol. 1996;27(12):1273–6. https://doi.org/10.1016/S0046-8177(96)90336-6.
Brown RE. Morphoproteomic portrait of the mTOR pathway in mesenchymal chondrosarcoma. Ann Clin Lab Sci. 2004;34(4):397–9.
Samsa WE, Zhou X, Zhou G. Signaling pathways regulating cartilage growth plate formation and activity. Semin Cell Dev Biol. 2017;62:3–15. https://doi.org/10.1016/j.semcdb.2016.07.008.
•• de Jong Y, et al. Inhibition of Bcl-2 family members sensitizes mesenchymal chondrosarcoma to conventional chemotherapy: report on a novel mesenchymal chondrosarcoma cell line. Lab Invest. 2016;96(10):1128–37. This study reports on a derived mesenchymal chondrosarcoma cell line, MCS170, in which the HEY1-NCOA2 translocation was identified. The authors investigated its response to doxorubicin, cisplatin and/or Bcl-2 family members inhibitors. Apoptosis induction and MCS170 sensitization to chemotherapy were achieved when combining Bcl-2 family members inhibitors with conventional chemotherapy.
Mertens F, Antonescu CR, Mitelman F. Gene fusions in soft tissue tumors: recurrent and overlapping pathogenetic themes. Genes Chromosomes Cancer. 2016;55(4):291–310. https://doi.org/10.1002/gcc.22335.
Riggi N, Knoechel B, Gillespie SM, Rheinbay E, Boulay G, Suvà ML, et al. EWS-FLI1 utilizes divergent chromatin remodeling mechanisms to directly activate or repress enhancer elements in Ewing sarcoma. Cancer Cell. 2014;26(5):668–81. https://doi.org/10.1016/j.ccell.2014.10.004.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
Marc El Beaino declares that he has no conflict of interest.
Jason Roszik declares that he has no conflict of interest.
John A. Livingston declares that he has no conflict of interest.
Wei-Lien Wang declares that he has no conflict of interest.
Alexander J. Lazar declares that he has no conflict of interest.
Behrang Amini declares that he has no conflict of interest.
Vivek Subbiah has received research support through grants from Novartis, Bayer, AbbVie, and Roche/Genentech.
Valerae Lewis declares that she has no conflict of interest.
Anthony P. Conley declares that he has no conflict of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
This article is part of the Topical Collection on Sarcomas
Rights and permissions
About this article
Cite this article
El Beaino, M., Roszik, J., Livingston, J.A. et al. Mesenchymal Chondrosarcoma: a Review with Emphasis on its Fusion-Driven Biology. Curr Oncol Rep 20, 37 (2018). https://doi.org/10.1007/s11912-018-0668-z
Published:
DOI: https://doi.org/10.1007/s11912-018-0668-z